Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Protein-encapsulated fluorogenic probes for the selective detection of endogenous O-GlcNAcase (OGA)
Yuan-Hao
Wu
a,
Chen
Guo
ac,
Zi-Ru
Ye
ac,
Xi-Le
Hu
 a,
Tony D.
James
a,
Tony D.
James
 *de,
Jia
Li
*de,
Jia
Li
 *c and
Xiao-Peng
He
*c and
Xiao-Peng
He
 *ab
*ab
aKey Laboratory for Advanced Materials and Joint International Research Laboratory of Precision Chemistry and Molecular Engineering, Feringa Nobel Prize Scientist Joint Research Center, School of Chemistry and Molecular Engineering, East China University of Science and Technology, 130 Meilong Rd, Shanghai 200237, China. E-mail: xphe@ecust.edu.cn
bThe International Cooperation Laboratory on Signal Transduction, Eastern Hepatobiliary Surgery Hospital, National Center for Liver Cancer, Shanghai 200438, China
cNational Center for Drug Screening, State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai 201203, China. E-mail: jli@simm.ac.cn
dDepartment of Chemistry, University of Bath, Bath, BA2 7AY, UK. E-mail: t.d.james@bath.ac.uk
eSchool of Chemistry and Chemical Engineering, Henan Normal University, Xinxiang 453007, China
First published on 17th February 2026
Abstract
Ever-increasing evidence confirms the role of O-GlcNAcase (OGA) in mediating cell growth and development as well as pathology and underscores the importance of developing sensitive and selective chemical tools for the study of OGA biology. Here, based on our previously developed protein-encapsulation strategy, we designed and synthesized a series of fluorogenic probes based on resorufin, and assembled their composites with human serum albumin (HSA) for the selective detection of endogenous OGA activity in live cells. We show that host–guest self-assembly with HSA significantly enhances the OGA sensitivity of the probes in aqueous solution and cells. The structure of the complex of a glycoprobe and HSA was resolved by small-angle X-ray scattering. We demonstrate that the replacement of the acetyl group in GlcNAc with a propionyl group results in selectivity for OGA over hexosaminidases (HEX) that unselectively hydrolyze hexosamines. This allows us to differentiate between two cell lines with different endogenous OGA and HEX expression levels, and selectively detect OGA activity in live cells.
Introduction
Mounting evidence has uncovered the important role of O-linked N-acetylglucosaminylation (O-GlcNAcylation) in mediating cellular actions including cell growth, division, differentiation and development.1 The malfunctioning of this biochemical process is also correlated with a number of pathological processes that can ultimately lead to human diseases such as cancer and neurodegenerative disorders.2,3 Protein O-GlcNAcylation is dynamically tailored by the reverse action of O-GlcNAc transferase (OGT) and O-GlcNAcase (OGA).4 Chemical probes developed for OGT mainly rely on a metabolic labelling strategy using non-natural GlcNAc derivatives (e.g., azide or alkyne-modified GlcNAc probes) in combination with a biorthogonal tag.5,6 This strategy has led to innovative observations indicating that OGT activity is involved in modulating a number of signalling pathways relevant to cellular proteostasis, cardiac function and cancer.7,8On the other hand, OGA removes GlcNAc from the serine/threonine residues of proteins. Previous investigations have shown that OGA is implicated in a series of physiological and disease–relevant processes.9–11 In addition, a recent study suggests that one role of OGA is to promote cancer cell malignancy by modulating the O-GlcNAcylation level on PDZ and LIM domain protein 7 (PDLI7).12 Conventional approaches to detect OGA activity have mainly relied on indirect analysis of the overall O-GlcNAcylation level using immunoblotting. This protocol is easily interfered with by fluctuations in OGT activity as well as the non-specific binding of anti-O-GlcNAc antibodies. As a consequence, colorimetric and fluorogenic substrates for OGA have been developed. For example, bis-acetal-based substrates have been developed as tunable fluorescence-quenched probes for OGA.13 By tuning the intramolecular charge transfer of donor–acceptor-type dye conjugates through GlcNAc modification, fluorogenic probes for OGA have also been developed.14–17 However, molecular probes incorporating GlcNAc as the natural substrate generally lack selectivity for OGA over hexosaminidases (HEX), which unselectively hydrolyze hexosamines.18,19 This limits their use for accurately measuring OGA activity in complicated biological systems.
Previous research has indicated that increasing the steric hindrance of substituents on the 2-amino group of GlcNAc improves substrate affinity for OGA.20 In an earlier study, we have also shown that the replacement of the acetyl (Ac) group in GlcNAc with a propionyl (Pr) group results in a significantly improved selectivity for OGA over HEX in solution-based assays. However, whether this replacement strategy can work for the selective detection of endogenous OGA over HEX in live cells remains unexplored.
Here, we developed a series of fluorogenic probes (GlcNAc-HHPO, GlcNPr-HHPO, GlcNAc-Bn-HHPO and GlcNPr-Bn-HHPO) for the detection of OGA activity in cell lysates and live cells, among which GlcNPr-HHPO with a Pr substitution exhibited outstanding selectivity for OGA over HEX (Fig. 1a). In addition, to improve the sensing performance, our previously developed protein-encapsulation strategy was exploited.21–23 Assemblies of our synthesized fluorogenic probes and human serum albumin (HSA) were prepared through host–guest assembly (Fig. 1b). Small-angle X-ray scattering (SAXS) was used to resolve the complex structure between GlcNPr-HHPO and HSA. Importantly, this ensemble achieved the selective detection of endogenous OGA over HEX activity in two cell lines with the results agreeing with that obtained using quantitative real-time polymerase chain reaction (qPCR) analysis. Furthermore, it could accurately sense OGA activity change in live cells.
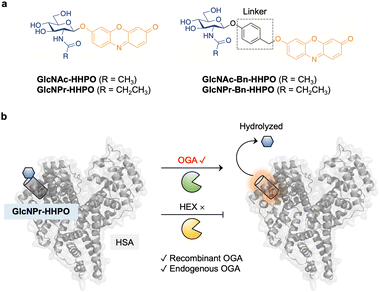 | ||
| Fig. 1 (a) Structure of GlcNAc-HHPO, GlcNPr-HHPO, GlcNAc-Bn-HHPO and GlcNPr-Bn-HHPO. (b) Sensing mechanism of the HSA/GlcNPr-HHPO complex for O-GlcNAcase (OGA) over hexosaminidases (HEX). | ||
Results and discussion
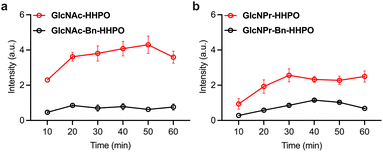
The synthesis of the fluorogenic probes is shown in Scheme S1. Resorufin, which is commonly used for fluorogenic assays, was used as the fluorescent reporter; the introduction of functional groups to its phenolic alcohol quenches the fluorescence due to masking of intramolecular charge transfer.24–26 Two series of fluorogenic probes were constructed; one through the direct glycosylation of resorufin with GlcNAc/GlcNPr (GlcNAc-HHPO/GlcNPr-HHPO), and the other exploiting a benzyl (Bn) linker (GlcNAc-Bn-HHPO/GlcNPr-Bn-HHPO). The introduction of the linker aimed to improve sensitivity,27 while the replacement of Ac with Pr should lead to selectivity for OGA over HEX. To develop the first series of probes, 1-α-Cl-substituted GlcNAc17 and GlcNPr17 were treated with resorufin in the presence of Cs2CO3 and Na2SO4 in MeCN,27 and a subsequent deacetylation using MeONa/MeOH afforded the desired GlcNAc-HHPO and GlcNPr-HHPO, respectively. The second series was similarly constructed by introducing 1-β-O-benzyl-substituted GlcNAc27 and GlcNPr27 to the phenolic alcohol of resorufin, and subsequent deacetylation gave probes GlcNAc-Bn-HHPO and GlcNPr-Bn-HHPO, respectively.With these probes in hand, we established a fluorogenic assay using a purified recombinant human OGA.18,27 The probes (50 µM) were incubated with OGA (1 µg mL−1) in a working buffer (0.1% bovine serum albumin (BSA), 50 mM NaH2PO4, 100 mM NaCl, pH 7.0),18 and the fluorescence of the resulting mixtures was measured using a microplate reader. According to previous studies, the presence of the Bn linker between the glycosyl substrate and the fluorophore could significantly enhance the sensitivity of the resulting probe for the target glycosidase.28,29 However, our analytical results indicated that the fluorescence enhancement of both GlcNAc-HHPO (Fig. 2a) and GlcNPr-HHPO (Fig. 2b) with OGA was much more evident than for GlcNAc-Bn-HHPO and GlcNPr-Bn-HHPO over time, respectively. We attributed this observation to the insufficient solubility of the Bn-containing probes in aqueous solutions. To verify our hypothesis, we obtained the absorption spectra of GlcNPr-HHPO and GlcNPr-Bn-HHPO in phosphate buffered saline (PBS, 0.01 M, pH 7.4) (Fig. S1). Over a concentration range of 10–100 µM, we observed that the absorption of GlcNPr-HHPO enhanced linearly, whereas that of GlcNPr-Bn-HHPO varied slightly. In addition, reddish, insoluble species were generated in a solution of 50 µM of GlcNPr-Bn-HHPO. These results suggest that the suboptimal sensitivity of the probes containing the Bn spacer is probably due to their inferior aqueous solubility, which subsequently compromised enzymatic reactivity.
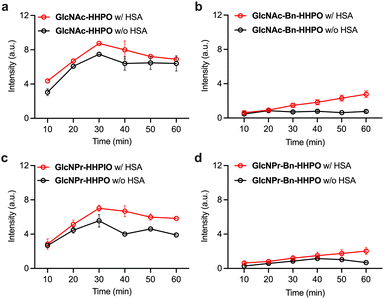
Next, we sought to improve the sensitivity of the probes through our previously developed protein-encapsulation strategy.21–23,30 HSA, which is among the most abundant proteins found in the human body, was used to encapsulate the probes. The probes were incubated with the albumin in the working buffer with mild stirring for 5 min, producing the ensembles HSA/GlcNAc-HHPO, HSA/GlcNAc-Bn-HHPO, HSA/GlcNPr-HHPO and HSA/GlcNPr-Bn-HHPO. Through fluorescence-based measurements, we determined that the presence of HSA consistently improved the sensitivity of all four probes for OGA over time (Fig. 3). The fact that the addition of HSA improved the sensitivity as well as detection linearity of HSA/GlcNAc-Bn-HHPO (Fig. 3c) and HSA/GlcNPr-Bn-HHPO (Fig. 3d) might be a result of an improved dispersibility of these Bn-containing probes in aqueous buffer solutions. However, the probes without the Bn spacer still exhibited a better sensitivity for OGA. In addition, we chose GlcNPr-HHPO to optimize the mixing ratio with HSA. The results indicated that a HSA:probe ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 was optimal for OGA detection (Fig. S2). Subsequently, we used the more sensitive HSA/GlcNAc-HHPO and HSA/GlcNPr-HHPO for selectivity tests. We determined that HSA/GlcNAc-HHPO exhibited a similar level of fluorescence recovery in the presence of OGA and HEX (Fig. S3a). In contrast, while the fluorescence of HSA/GlcNPr-HHPO was observed to significantly enhance when OGA is added, the probe remained quenched with HEX (Fig. S3b). This agrees with our previous study that the replacement of Ac with Pr in GlcNAc substantially improves its selectivity for OGA over HEX.18
1 was optimal for OGA detection (Fig. S2). Subsequently, we used the more sensitive HSA/GlcNAc-HHPO and HSA/GlcNPr-HHPO for selectivity tests. We determined that HSA/GlcNAc-HHPO exhibited a similar level of fluorescence recovery in the presence of OGA and HEX (Fig. S3a). In contrast, while the fluorescence of HSA/GlcNPr-HHPO was observed to significantly enhance when OGA is added, the probe remained quenched with HEX (Fig. S3b). This agrees with our previous study that the replacement of Ac with Pr in GlcNAc substantially improves its selectivity for OGA over HEX.18
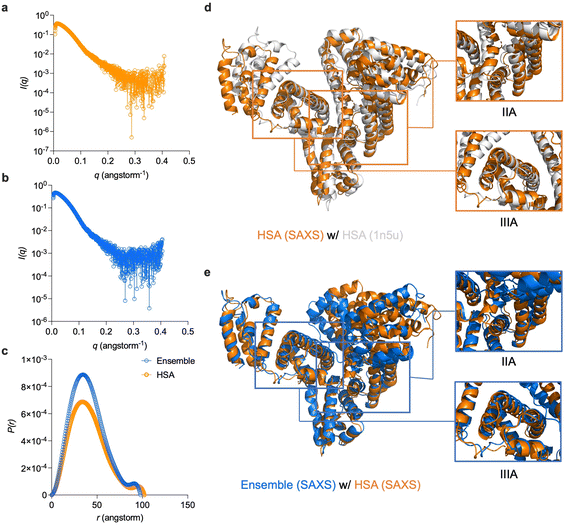
In order to gain deeper understanding of the binding mode between GlcNPr-HHPO and HSA, small-angel X-ray scattering (SAXS) was used.23,28 The SAXS profile for HSA and that for the HSA/GlcNPr-HHPO ensemble is shown in Fig. 4a and b, respectively. SREFLEX,31 a hybrid modelling program that systematically incorporates SAXS data with normal mode analysis was used to simulate the three-dimensional models of HSA and the ensemble. The HSA model was first superimposed onto a reported HSA crystalline structure (PDB entry: 1n5u), and a good overlap between the two models was found (Fig. 4d). Then, the SAXS model of HSA and that of HSA/GlcNPr-HHPO was superimposed to analyze the conformational changes of the latter. As shown in Fig. 4e, while the IIIA region of both models overlapped well, the IIA region of the ensemble was found to position differently from that of HSA. This suggests an inclusion of the probe into the IIA region of HSA, which is known to accommodate hydrophobic molecules.32 We also stacked the distance distribution function (P(r)) curves of both models (Fig. 4c), and found a decreased maximum diameter (Dmax) for HSA/GlcNPr-HHPO (97.9 Å) with respect to HSA (102.1 Å). To corroborate the binding, a competition assay was performed using phenylbutazone (a known IIA-region binder) and ibuprofen (a known IIIA-region binder) as competing agents. The addition of increasing concentrations of phenylbutazone led to the gradual decrease of the tryptophan fluorescence in the IIA site of HSA (Fig. S4),33 which correlated with the probe and was taken as evidence for the inclusion of the probe into the IIA region of HSA. The binding constant (Ka) of probe GlcNPr-HHPO with HSA was determined to be 3.685 × 105 M−1 using the fluorescence double reciprocal method (Fig. S5).
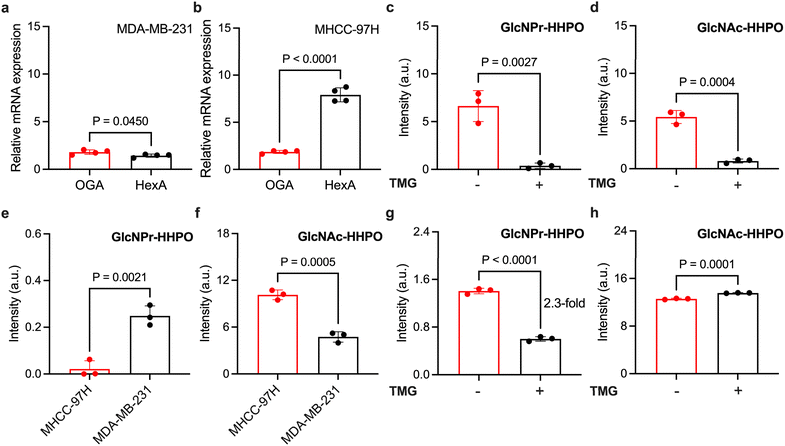
With the promising sensing properties of the ensembles determined, we turned our attention to evaluate their ability to measure endogenous OGA activity in cells. Two cell lines, MDA-MB-231 (human triple-negative breast cancer) and MHCC-97H (human liver cancer), with different OGA and HEX expression levels were used. We first used qPCR to determine the endogenous expression level of OGA and HexA (a predominant isozyme of HEX) in the two cell lines. The results indicated a slightly higher expression level of OGA than that of HexA in MDA-MB-231 (Fig. 5a), and a ∼4-fold higher HexA level than OGA in MHCC-97H (Fig. 5b).
Next, we used both HSA/GlcNAc-HHPO and HSA/GlcNPr-HHPO to treat the cell lysates. Interestingly, we determined a ∼8-fold higher fluorescence for HSA/GlcNPr-HHPO in MDA-MB-231 than MHCC-97H (Fig. 5e). In contrast, a ∼2-fold stronger fluorescence was seen for HSA/GlcNAc-HHPO in MHCC-97H than MDA-MB-231 (Fig. 5f). These results suggest that (1) the fluorescence activation of HSA/GlcNAc-HHPO is largely dependent on the HEX expression of the cells because HexA was determined to be predominantly expressed in MHCC-97H, and (2) HSA/GlcNPr-HHPO can be used to more accurately measure endogenous OGA activity irrespective of HEX expression as evidenced by its minimal fluorescence recovery in the lysate of MHCC-97H cells.
To better corroborate the OGA selectivity of the ensembles, a selective OGA inhibitor, Thiamet-G (TMG) with a nanomolar half-maximal inhibitory concentration, was used.34 We first found that TMG inhibited the fluorescence enhancement of both HSA/GlcNAc-HHPO (Fig. 5d) and HSA/GlcNPr-HHPO (Fig. 5c) when incubated with recombinant OGA. This confirms the effectiveness of the inhibitor. Then, the cell lysate of MDA-MB-231 was collected and pre-treated with TMG, followed by incubation with the ensembles. While treatment of TMG did not interrupt the fluorescence recovery of HSA/GlcNAc-HHPO (Fig. 5h), a ∼2.3-fold fluorescence decrease was determined for HSA/GlcNPr-HHPO in the presence of TMG (Fig. 5g). This demonstrates that HSA/GlcNPr-HHPO but not HSA/GlcNAc-HHPO could selectively detect OGA activity irrespective of the expression of HEX.
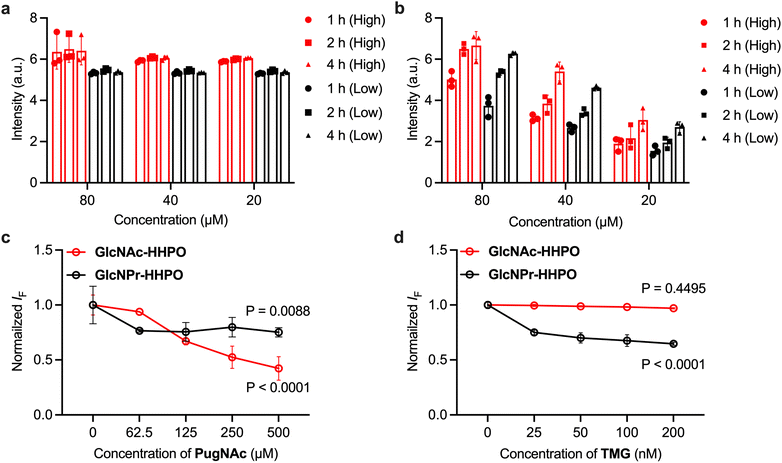
To further evaluate whether our probes could accurately report on OGA activities in live cells, the following experiments were carried out (results are shown in Fig. 6). HeLa (human cervix) cells with a higher endogenous expression level of HexA than that of OGA were used (Fig. S6). The cells were incubated at different densities (40–50% or 80–90% per well) with both HSA/GlcNAc-HHPO (Fig. 6a) and HSA/GlcNPr-HHPO (Fig. 6b). We found that in all test groups the fluorescence of HSA/GlcNAc-HHPO reached equilibrium in 1 h irrespective of cell density and probe concentration. In contrast, the fluorescence enhancement of HSA/GlcNPr-HHPO was seen to be dependent on both the concentration of the probe and cell density. This suggests a quicker intracellular hydrolysis of HSA/GlcNAc-HHPO than that of HSA/GlcNPr-HHPO probably due to the action of both OGA and HEX. A cell viability assay showed that the probes were not toxic to HeLa cells over a concentration range of 0–80 µM (Fig. S7). In addition, a competition assay was carried out by pre-treating HeLa cells with two known glycosidase inhibitors – PugNAc that unselectively inhibits OGA and HEX, and TMG that selectively inhibits OGA.34 Interestingly, the results shown in Fig. S6c (PugNAc) and Fig. S6d (TMG) clearly indicate that the fluorescence of both probes decreased with increasing concentrations of PugNAc; however, in the presence of the OGA selective inhibitor, TMG, only the fluorescence of HSA/GlcNPr-HHPO was seen to decrease in a concentration-dependent manner.
Conclusions
We have developed resorufin-based fluorogenic probes for OGA. We found that the presence of a benzyl spacer compromised the sensitivity of the probes probably due to solubility issues. However, association with HSA was effective in improving the sensitivity of the probes for OGA. Importantly, the substitution of Pr for the Ac group gave rise to selectivity for OGA over HEX. This enables us to differentiate between two cell lines with different endogenous OGA and HexA expression levels. While, comparing with previously established OGA probes (Table S1), the probes also achieved the selective detection of OGA in live cells. As such our research provides valuable chemical tools for the determination of OGA activity in complicated biological systems, and offers insight into the development of molecular glycoprobes for other discrete glycosidases.35,36Author contributions
Yuan-Hao Wu, Guo Chen and Zi-Ru Ye performed experiments, analyzed data and drafted the manuscript. Xi-Le Hu, Tony D. James, Jia Li and Xiao-Peng He supervised research and edited the manuscript. All authors have approved the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting this research are available as part of the supplementary information (SI). Supplementary information: experimental methods, additional figures, and spectra of new compounds. See DOI: https://doi.org/10.1039/d5sc06843f.Acknowledgements
The authors thank the Natural National Science Foundation of China (No. 92253306, 82130099 and 22477030), Science and Technology Commission of Shanghai Municipality (grant No.24DX1400200), the International Cooperation Program of Shanghai Science and Technology (No.23490711600), the Fundamental Research Funds for the Central Universities (222201717003), the Programme of Introducing Talents of Discipline to Universities (B16017), the National Natural Science Foundation of Shanghai Science and Technology (No.24ZR1415400), the Shanghai Oriental Talents youth Program (No. QNKJ2024010),the Shanghai Xuhui District Hospital Local Cooperation Project (23XHYD-20), the Open Funding Project of the State Key Laboratory of Fine Chemicals, Dalian University of Technology (KF 2402), State Key Laboratory of Chemo/Biosensing and Chemometrics, Hunan University, Changsha 410082, P. R. China, Ministry of Education Key Laboratory on Signaling Regulation and Targeting Therapy of Liver Cancer (Naval Medical University) (Grant. 2023-MEKLLC-MS/ZD-00*) and Shandong Laboratory Program (SYS202205) for financial support. T.D.J. wishes to thank the University of Bath and the Open Research Fund of the School of Chemistry and Chemical Engineering, Henan Normal University (2020ZD01) for support. The Research Center of Analysis and Test of East China University of Science and Technology was gratefully acknowledged for assistance in analytical experiments. The authors also thank the staff members of BL19U2 beamline (https://cstr.cn/31129.02.NFPS.BL19U2) at the National Facility for Protein Science in Shanghai (https://cstr.cn/31129.02.NFPS) for providing technical support and assistance in data collection and analysis.Notes and references
- J. C. Zhang and Y. Z. Wang, J. Biol. Chem., 2024, 300, 105677 Search PubMed.
- Y. Zhang, S. Y. Zhou, Y. Kai, Y. Q. Zhang, C. M. Peng, Z. Q. Li, M. J. Mughal, B. Jujlie, X. Y. Zheng, J. F. Ma, C. X. Ma, M. Shen, M. D. Hall, S. Q. Li and W. G. Zhu, Nat. Commun., 2024, 15, 5597 Search PubMed.
- A. T. Balana, A. L. Mahul-Mellier, B. A. Nguyen, M. Horvath, A. Javed, E. R. Hard, Y. Jasiqi, P. Singh, S. Afrin, R. Pedretti, V. Singh, V. M. Y. Lee, K. C. Luk, L. Saelices, H. A. Lashuel and M. R. Pratt, Nat. Chem. Biol., 2024, 20, 646–655 Search PubMed.
- J. F. Ma, C. Y. Hou and C. Wu, Chem. Rev., 2022, 122, 15822–15864 Search PubMed.
- D. J. Vocadlo, H. C. Hang, E. Kim, J. A. Hanover and C. R. Bertozzi, Proc. Natl. Acad. Sci. U. S. A, 2003, 100, 9116–9121 Search PubMed.
- K. Qin, H. Zhang, Z. Q. Zhao and X. Chen, J. Am. Chem. Soc., 2020, 142, 9382–9388 CrossRef CAS PubMed.
- X. Q. Fan, Q. T. Song, D. E. Sun, Y. Hao, J. Y. Wang, C. T. Wang and X. Chen, Nat. Chem. Biol., 2022, 18, 625–633 Search PubMed.
- Y. P. Zhu, L. I. Willems, D. Salas, S. Cecinoi, W. B. Wu, L. J. Foster and D. J. Vocadlo, J. Am. Chem. Soc., 2020, 142, 15729–15739 Search PubMed.
- S. S. Chen, A. C. Mody and C. M. Woo, Chem. Rev., 2024, 124, 12918–13019 Search PubMed.
- A. A. Elbatrawy, E. J. Kim and G. Nam, ChemMedChem, 2020, 15, 1244–1257 Search PubMed.
- A. K. Nagel and L. E. Ball, Amino Acids, 2014, 46, 2305–2316 Search PubMed.
- C. W. Hu, A. Wang, D. C. Fan and J. Y. Jiang, Proc. Natl. Acad. Sci. U. S. A, 2024, 121, e2320867121 Search PubMed.
- S. Cecioni and D. J. Vocadlo, J. Am. Chem. Soc., 2017, 139, 8392–8395 Search PubMed.
- E. J. Kim, D. O. Kang, D. C. Love and J. A. Hanover, Carbohydr. Res., 2006, 341, 971–982 Search PubMed.
- J. Y. Hyun, S. H. Park, C. W. Park, H. B. Kim, J. W. Cho and I. Shin, Org. Lett., 2019, 21, 4439–4442 Search PubMed.
- J. Boo, J. Lee, Y. H. Kim, C. H. Lee, B. Ku and I. Shin, Front. Chem., 2023, 11, 1133018 Search PubMed.
- H. Jung, S. H. Park, W. H. Yang, J. W. Cho and I. Shin, Sens. Actuators, B, 2022, 367, 132093 Search PubMed.
- M. S. Macauley, G. E. Whitworth, A. W. Debowski, D. Chin and D. J. Vocadlo, J. Biol. Chem., 2005, 280, 25313–25322 Search PubMed.
- M. González-Cuesta, P. Sidhu, R. A. Ashmus, A. Males, C. Proceviat, Z. Madden, J. C. Rogalski, J. A. Busmann, L. J. Foster, J. M. G. Fernández, G. J. Davies, C. O. Mellet and D. J. Vocadlo, J. Am. Chem. Soc., 2022, 144, 832–844 Search PubMed.
- E. J. Kim, M. Perreira, C. J. Thomas and J. A. Hanover, J. Am. Chem. Soc., 2006, 128, 4234–4235 Search PubMed.
- Y. X. Fu, H. H. Han, J. J. Zhang, X. P. He, B. L. Feringa and H. Tian, J. Am. Chem. Soc., 2018, 140, 8671–8674 CrossRef CAS PubMed.
- X. Chai, H. H. Han, A. C. Sedgwick, N. Li, Y. Zang, T. D. James, J. Zhang, X. L. Hu, Y. Yu, Y. Li, Y. Wang, J. Li, X. P. He and H. Tian, J. Am. Chem. Soc., 2020, 142, 18005–18013 CrossRef CAS PubMed.
- H. H. Han, A. C. Sedgwick, Y. Shang, N. Li, T. T. Liu, B. H. Li, K. Q. Yu, Y. Zang, J. T. Brewster, II, M. L. Odyniec, M. Weber, S. D. Bull, J. Li, J. L. Sessler, T. D. James, X. P. He and H. Tian, Chem. Sci., 2020, 11, 1107–1113 Search PubMed.
- L. Flamigni, E. Venuti, N. Camainoi and F. Barigelletti, J. Chem. Soc. Faraday. Trans., 1989, 2(85), 1935–1943 Search PubMed.
- C. Bueno, M. L. Villegas, S. G. Bertolotti, C. M. Previtali, M. G. Neumann and M. V. Encinas, Photochem. Photobiol., 2002, 76, 385–390 Search PubMed.
- F. Yan, X. G. Tian, Z. L. Luan, L. Feng, X. C. Ma and T. D. James, Chem. Commun., 2019, 55, 1955–1958 RSC.
- Y. H. Wu, G. J. Wang, C. Guo, P. P. Wang, J. Y. Zhang, X. L. Hu, Y. Zang, T. D. James, J. Li and X. P. He, Chem. Commun., 2024, 60, 8240–8243 Search PubMed.
- J. Yan, S. Lee, A. Zhang and J. Yoon, Chem. Soc. Rev., 2018, 47, 6900–6916 Search PubMed.
- P. L. Carl, P. K. Chakravarty and J. A. Katzenellenbogen, J. Med. Chem., 1981, 24, 479–480 Search PubMed.
- W. T. Dou, P. Qiu, Y. Y. Shi, L. Zhu, C. Guo, N. Li, Y. Zang, T. T. Liu, S. W. Zhao, Y. F. Pan, L. W. Dong, J. L. Sessler, Y. X. Tan, J. Li, H. Y. Wang, H. Tian and X. P. He, J. Am. Chem. Soc., 2023, 145, 17377–17388 Search PubMed.
- A. Panjkovich and D. I. Svergun, Phys. Chem. Chem. Phys., 2016, 18, 5707–5719 Search PubMed.
- Z. B. Liu and X. Y. Chen, Chem. Soc. Rev., 2016, 45, 1432–1456 Search PubMed.
- H. Wataanabe, S. Tanase, K. Nakajou, T. Maruyama, U. Kragh-Hansen and M. Otagiri, Biochem. J., 2000, 349, 813–819 Search PubMed.
- S. A. Yuzwa, M. S. Macauley, J. E. Heinonen, X. Shan, R. J. Dennis, Y. He, G. E. Whitworth, K. A. Stubbs, E. J. McEachern, G. J. Davies and D. J. Vocadlo, Nat. Chem. Biol., 2008, 4, 483–490 Search PubMed.
- H.-H. Han, X.-L. Hu and X.-P. He, Chem. Commun., 2025, 61, 15735–15758 Search PubMed.
- X.-P. He, Chem. Soc. Rev., 2026, 55, 605–618 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |