Open Access Article
Open Access ArticleImplementing the design cues of dissociation dynamics and transmetalation in gallium(III) complexes to promote the anti-proliferative activity of ligands targeting intracellular iron(II) trafficking
Mahendiran
Dharmasivam†
*ab,
Sadia
Faiz†
ad,
Busra
Kaya†
b,
Tharushi P.
Wijesinghe
b,
Mediha
Suleymanoglu
bc,
Mahan Gholam
Azad
ab,
Vera
Richardson
ab,
Ameer Fawad
Zahoor
 d,
Danuta
Kalinowski
a,
William
Lewis
d,
Danuta
Kalinowski
a,
William
Lewis
 e,
Paul V.
Bernhardt
e,
Paul V.
Bernhardt
 f,
Rukhsana
Anjum
a and
Des R.
Richardson
f,
Rukhsana
Anjum
a and
Des R.
Richardson
 *ab
*ab
aMolecular Pharmacology and Pathology Program, Department of Pathology and Bosch Institute, The University of Sydney, Sydney, New South Wales 2006, Australia
bCentre for Cancer Cell Biology and Drug Discovery, Institute for Biomedicine and Glycomics, Griffith University, Southport, Queensland 4215, Australia. E-mail: d.richardson@griffith.edu.au; m.dharmasivam@griffith.edu.au
cDepartment of Medical Biology, Istanbul Faculty of Medicine, Istanbul University, 34093 Fatih-Istanbul, Turkiye
dDepartment of Chemistry, Government College University Faisalabad, Faisalabad 38000, Pakistan
eSchool of Chemistry, The University of Sydney, Sydney, New South Wales 2006, Australia
fSchool of Chemistry and Molecular Biosciences, University of Queensland, Brisbane 4072, Australia
First published on 9th February 2026
Abstract
Designing ligands for cancer has traditionally overlooked complex dissociation and transmetalation in enhancing efficacy. Another neglected criterion is the ligands' ability to intercept the labile Fe(II) pool released after transferrin endocytosis and reduction of transferrin-bound Fe(III). Given iron's essential role in cancer proliferation, disrupting metal homeostasis offers a promising therapeutic strategy. Herein, we introduce a new class of Fe(II)-selective ligands and their Ga(III) complexes for cancer therapy, guided by insights into their dissociative dynamics and transmetalation behavior. Unlike prior approaches focused on static metal coordination, this work integrates dissociation and transmetalation as design features, enabling selective interception of intracellular Fe(II) trafficking. Relative to the ligand, Ga(III) complexation led to a pronounced (p < 0.001–0.0001) enhancement in anti-proliferative activity, with up to a 70-fold increase in potency. This result was in contrast to the modest increase in potency (up to 2.4-fold) observed for the Cu(II) or Zn(II) complexes. Mechanistic dissection demonstrated that, unlike the complete dissociation of the Ga(III) complexes, the relative Zn(II) and Cu(II) complexes underwent only partial dissociation. This difference facilitates complete ligand and Ga(III) release from the complex and may account for the superior cytotoxicity of the Ga(III) complexes versus their Zn(II) and Cu(II) counterparts. Furthermore, their potency was linked to Fe(II) ligation rather than Fe(III), despite electronic similarity to Ga(III). This study introduces three underexplored design principles for anti-cancer ligand engineering: (i) dynamic complex dissociation; (ii) selective intracellular transmetalation using NNO-containing ligands; and (iii) interception of labile Fe(II) generated after endosomal Fe(III) reduction.
1 Introduction
Iron (Fe) is an essential micronutrient that drives tumor progression by supporting DNA synthesis, mitochondrial respiration, and cell cycle regulation.1–3 The elevated metabolic demands of cancer cells result in a pronounced dependency on iron, rendering Fe metabolism a tractable target for anti-cancer therapy.1,4,5 Cellular Fe uptake is predominantly mediated by diferric transferrin (Fe2-Tf), which binds transferrin receptor 1 (TfR1) and undergoes receptor-mediated endocytosis.6–8 Within endosomes, Fe(III) is released under acidic conditions, reduced to Fe(II) by the ferrireductase six-transmembrane epithelial antigen of the prostate 3 (STEAP3),9 and transported into the cytosol via divalent metal transporter 1 (DMT1).10 Cytosolic Fe(II) in the intracellular labile Fe pool11 is utilized by Fe-dependent enzymes or sequestered in ferritin.1,2Targeting intracellular Fe homeostasis12,13 using chelators or Fe(III)-mimetic complexes such as Ga(III)14–16 or Ti(IV)17 has shown promise in disrupting Fe-dependent biological processes.1–3 Among effective scaffolds are the aroylhydrazones and thiosemicarbazones,18–21 which act as chelators12,22–26 or lipophilic metal carriers.21 Their strong Fe-binding capacity has also enabled applications in iron-overload disease.27–30 Extensive structure–activity relationship (SAR) studies have facilitated the specific design of cytotoxic ligands with enhanced potency and selectivity.31–38
In contrast to previous aroylhydrazone-based studies,39–41 this investigation introduces a conceptually distinct strategy that integrates three underexplored design principles into anti-cancer ligand engineering: (i) dynamic dissociation of metal complexes; (ii) selective intracellular transmetalation; and (iii) interception of labile Fe(II), a pool of Fe generated after endosomal reduction of Fe(III) in Fe2-Tf.9 To implement this, we designed a novel series of tridentate naphthofuran-based ligands bearing NNO donor sets, constructed from pyridine, pyrazine, benzylpyridine, or dipyridyl substituents (Fig. 1A and B). These scaffolds offer high Fe(II) affinity, tunable electronics, and favorable lipophilicity and membrane permeability.
Our approach builds upon prior SAR-driven development of hybrid thiosemicarbazones and aroylhydrazones,42 as well as the di-2-pyridylketone thiosemicarbazone (DpT; Fig. 1C) class.43–49 These latter agents include di-2-pyridylketone-4,4-dimethyl-3-thiosemicarbazone (Dp44mT; Fig. 1D), which demonstrates potent in vitro and in vivo anti-tumor activity, and its orally bioavailable analogue, di-2-pyridylketone-4-cyclohexyl-4-methyl-3-thiosemicarbazone (DpC; Fig. 1E), which advanced to multi-centre phase I clinical trials.43–47,50–58
An important aspect of the efficacy of these anti-cancer ligands is the “double punch” mechanism.59 This relates to the tridentate, N,N,S coordination site being capable of binding Fe(III) and Cu(II), which are required for proliferation, but also the appropriate redox potential of these complexes, leading to their redox activity. In particular, the redox-active Cu(II)-thiosemicarbazone complexes generate reactive oxygen species (ROS) and induce lysosomal membrane permeabilization.12,18,49,51,60–66 In contrast, the Fe(III) complexes of the DpT ligand class are far less redox active49 and do not possess marked anti-proliferative efficacy.48 On the other hand, Zn(II) complexes may facilitate anti-cancer activity by in situ transmetalation with Cu(II).62,67–70
Triapine® (3-aminopyridine-2-carboxaldehyde thiosemicarbazone; Fig. 1F) is an older α-N-heterocyclic thiosemicarbazone that has demonstrated anti-tumor activity and has been examined in over 20 human clinical trials.71 More recently, dual-function chelators, namely the Triapine®–deferasirox hybrid, have exemplified the therapeutic potential of pharmacophore combination.13 Of interest, naphthofuran-containing molecules, including natural products such as (±)-laevigatin, (+)-heritol, and balsaminone A, exhibit favorable pharmacokinetic and anti-cancer properties.72,73 In aroylhydrazone and thiosemicarbazone tridentate ligands that form stable, planar complexes,40,41,50,69,70,74 incorporating the naphthofuran moiety can be expected to enhance: (1) electron delocalization, and thus, promote ligand rigidity; and (2) increase lipophilicity that will improve cellular permeability and chelation efficacy.
Herein, we implement a dual-action strategy in which Ga(III) complexes of tridentate ligands undergo intracellular dissociation, releasing free ligands that selectively bind Fe(II). This process enables dynamic transmetalation and targeted disruption of intracellular Fe(II) trafficking. Notably, Ga(III) coordination enhanced the anti-proliferative activity of the ligand by up to 70-fold, substantially exceeding the modest 1.3–2.4-fold increases observed for the corresponding Cu(II) or Zn(II) complexes. This pronounced efficacy was unexpected, as Cu(II) and Zn(II) complexation has previously improved the activity of structurally related ligands,40,67–70,75 but never to this extent.
In line with Ga(III) coordination chemistry,76,77 our results were initially assumed to relate to modulation of atomically similar Fe(III) in cellular pools targeted by Ga(III).78–80 However, again in contrast to expectations, mechanistic dissection revealed the critical roles of robust ligand dissociation and transmetalation with Fe(II), but not Fe(III). We propose that [Ga(L12)2]+ dissociates intracellularly, liberating L12 to bind Fe(II) released during endosomal Fe(III) reduction. Collectively, these results establish dissociation-enabled Fe(II) interception and selective transmetalation as powerful yet underutilized design principles for next-generation Ga(III)-based anti-cancer agents.
2 Results and discussion
Nine bidentate aroylhydrazone and thiosemicarbazone ligands based on benzofuran (L1–L4) and naphthofuran (L5–L9) were synthesized (Fig. 1A), along with four tridentate naphthofuran-based aroylhydrazones (L10–L13; Fig. 1B). The tridentate ligands were designed for Cu(II), Zn(II), and Ga(III) coordination, forming stable, planar complexes via imine nitrogen, carbonyl oxygen, and pyridine- or pyrazine–nitrogen. Fe(II) complexes of these ligands are inherently unstable under aerobic or aqueous conditions, as they readily oxidize to Fe(III), readily dissociate, or undergo rapid ligand exchange, preventing their reproducible isolation and characterization.81 Our studies show that Ga(III) complexes adopt a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 Ga
2 Ga![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) L octahedral geometry, supported by MS and spectroscopy, and consistent with known Ga(III) preferences.76 By contrast, Cu(II) and Zn(II) complexes display four- and five-coordinate geometries from spectroscopy and X-ray analyses, respectively, differences that underpin their distinct dissociation and biological profiles (below).
L octahedral geometry, supported by MS and spectroscopy, and consistent with known Ga(III) preferences.76 By contrast, Cu(II) and Zn(II) complexes display four- and five-coordinate geometries from spectroscopy and X-ray analyses, respectively, differences that underpin their distinct dissociation and biological profiles (below).
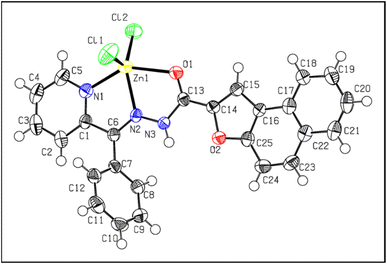
2.1. X-ray crystallography
Understanding the structure of ligands and complexes is crucial for elucidating their medicinal chemistry and biological activity. To this end, X-ray crystallography was employed to deepen our knowledge of the chemistry of L1, L5, L7, and L10–L13, with the ligand structures (Fig. S1A–G) and unit cell packing diagrams shown in Fig. S2A–G. Studies focused on the crystal structures of complexes of the tridentate ligands L10–L13, as these demonstrated the greatest biological activity (described below). Exhaustive attempts to obtain a crystal structure of the Ga(III) complexes of L10–L13 always led to crystals of the ligand alone. This observation reflects critical aspects of the coordination chemistry of these ligands, particularly dissociation, which is key to their biological activity and is described below. However, the crystal structure of [Zn(L11)Cl2]·CH2Cl2 is shown in Fig. 2. The neutral complex is five-coordinate with a distorted square pyramidal geometry (τ5 = 0.18: τ5 = (α − β)/60, where α and β are the two largest coordinate angles). The hydrazone ligand remains protonated at N3.2.2. Inhibition of cellular proliferation
The anti-proliferative activity of the synthesized ligands (L1–L13) and their Cu(II), Zn(II), and Ga(III) complexes was evaluated in SK-N-MC neuroepithelioma cells, MCF-7 (estrogen receptor-positive) and MDA-MB-231 (triple-negative) breast cancer cells following a 72 h/37 °C incubation (Tables 1, 2 and S3). These cell types were selected for their well-characterized responses to metal-binding ligands and their distinct proliferative phenotypes.57,74,82,83 As controls, ligands with well-established clinical or investigational relevance were employed, namely: (1) the clinically used hydrophilic Fe(III) chelator, desferrioxamine (DFO), which shows modest anti-proliferative activity;43,84 (2) the ONO-donor aroylhydrazone, 2-hydroxy-1-naphthylaldehyde-isonicotinoyl-hydrazone (311), which shows moderate anti-proliferative efficacy;30,32,42 and (3) the thiosemicarbazones, DpC and Dp44mT, which demonstrate potent anti-proliferative effects.42,52,67–69,85| Compounds | IC50 (µM) | ||
|---|---|---|---|
| SK-N-MC | MCF-7 | MDA-MB-231 | |
| DFO | 13.15 ± 4.24 | 3.18 ± 0.25 | 27.61 ± 0.14 |
| 311 | 1.06 ± 0.21 | 1.56 ± 0.35 | >12.5 |
| DpC | 0.02 ± 0.01 | 0.03 ± 0.01 | 0.02 ± 0.01 |
| Dp44mT | 0.01 ± 0.01 | 0.02 ± 0.01 | 0.01 ± 0.01 |
| L1 | 8.93 ± 0.78 | 1.26 ± 0.04 | 10.93 ± 0.07 |
| L2 | >12.5 | 6.77 ± 0.84 | >12.5 |
| L3 | >12.5 | 10.13 ± 0.32 | >12.5 |
| L4 | >12.5 | >12.5 | >12.5 |
| L5 | 4.34 ± 0.27 | 11.72 ± 0.46 | 6.40 ± 0.23 |
| L6 | 5.87 ± 0.10 | >12.5 | 5.72 ± 0.12 |
| L7 | >12.5 | 5.89 ± 0.40 | 6.41 ± 0.14 |
| L8 | 6.32 ± 0.14 | >12.5 | 5.78 ± 0.09 |
| L9 | >12.5 | >12.5 | >12.5 |
| L10 | 1.38 ± 0.27 | 1.43 ± 0.03 | 1.62 ± 0.21 |
| L11 | 2.38 ± 0.52 | 3.21 ± 0.49 | 2.52 ± 0.49 |
| L12 | 2.11 ± 0.08 | 1.53 ± 0.35 | 1.79 ± 0.03 |
| L13 | 1.31 ± 0.14 | 1.41 ± 0.11 | 1.17 ± 0.05 |
| Compounds | IC50 (µM) | ||
|---|---|---|---|
| SK-N-MC | MCF-7 | MDA-MB-231 | |
| L10 | 1.38 ± 0.27 | 1.43 ± 0.03 | 1.62 ± 0.21 |
| [Cu(L10)OAc] | 1.17 ± 0.25 | 1.08 ± 0.38 | 1.15 ± 0.02 |
| [Zn(L10)Cl2] | 0.81 ± 0.14 | 1.54 ± 0.12 | 1.28 ± 0.04 |
| [Ga(L10) 2 ] + | 0.04 ± 0.01 | 0.79 ± 0.09 | 0.61 ± 0.21 |
| L11 | 2.38 ± 0.52 | 3.21 ± 0.49 | 2.52 ± 0.49 |
| [Cu(L11)OAc] | 1.60 ± 0.09 | 1.32 ± 0.15 | 1.46 ± 0.03 |
| [Zn(L11)Cl2] | 4.52 ± 0.85 | 1.62 ± 0.04 | 0.81 ± 0.28 |
| [Ga(L11) 2 ] + | 0.13 ± 0.13 | 1.14 ± 0.28 | 0.43 ± 0.01 |
| L12 | 2.11 ± 0.08 | 1.53 ± 0.35 | 1.79 ± 0.03 |
| [Cu(L12)OAc] | 1.61 ± 0.13 | 1.92 ± 0.26 | 1.02 ± 0.29 |
| [Zn(L12)Cl2] | 0.89 ± 0.41 | 1.11 ± 0.24 | 1.08 ± 0.18 |
| [Ga(L12) 2 ] + | 0.03 ± 0.01 | 0.68 ± 0.05 | 0.62 ± 0.14 |
| L13 | 1.31 ± 0.14 | 1.41 ± 0.11 | 1.17 ± 0.05 |
| [Cu(L13)Cl2] | 0.53 ± 0.14 | 0.92 ± 0.45 | 1.00 ± 0.08 |
| [Zn(L13)Cl2] | 2.30 ± 0.42 | 1.08 ± 0.36 | 1.14 ± 0.26 |
| [Ga(L13) 2 ] + | 0.04 ± 0.01 | 0.59 ± 0.14 | 0.83 ± 0.09 |
As expected, DFO exhibited limited activity (IC50: 3.18–27.61 µM),43,84 while 311 displayed moderate potency,30,32,42 particularly against SK-N-MC (IC50: 1.06 µM) and MCF-7 (IC50: 1.56 µM) cells. Relative to these two agents, DpC and Dp44mT exhibited far more potent anti-tumor activity (IC50: 0.01–0.03 µM) across all cell types (Table 1), in agreement with our previous studies.52,67–69,85
Among the new compounds, the bidentate ligands (L1–L9) displayed variable anti-proliferative effects that varied from the moderate (IC50: 1.26 µM) activity of L1 against MCF-7 cells, to the more limited (>4.34–12.5 µM) efficacy observed for all these chelators across the 3 cell-types (Table 1). In contrast, the tridentate naphthofuran-based ligands L10–L13 demonstrated consistent and superior efficacy, with IC50 values of 1.17–3.21 µM across all cell-types. These tridentate ligands were significantly (p < 0.001) more active than DFO in all cell types, except for L11 in MCF-7 cells, where activity was comparable.
Relative to 311, L10–L13 exhibited similar or greater potency, with greater efficacy in the more aggressive MDA-MB-231 triple-negative breast cancer cells (Table 1). This improved activity relative to the bidentate L1–L9 is attributed to their enhanced complex stability via the chelate effect and the NNO donor set, which provides stronger and more geometrically favorable coordination.12
2.3. Potently enhanced anti-proliferative activity of ligands L10–L13 upon Ga(III) complexation
The Cu(II) and Zn(II) complexes of L10–L13 exhibited minimal or inconsistent changes in anti-proliferative activity versus the ligand, with modest enhancements ranging from 1.02- to 3.11-fold and, in some cases, reduced potency (Table 2). In contrast, Ga(III) complexation led to marked and significant (p < 0.001–0.0001) increases in anti-proliferative activity relative to the respective ligands across all cell types. Importantly, [Ga(NO3)3] alone displayed very limited cytotoxicity (IC50 = 88.6 ± 1.2 µM in SK-N-MC and 78.9 ± 2.0 µM in MCF-7 cells), confirming that free Ga(III) is biologically poorly active86 under these conditions (Table S3).31,86 In contrast, [Ga(L12)2]+ markedly and significantly (p < 0.0001) enhances potency in both cell-types (IC50 = 0.099 ± 0.010 µM and 0.292 ± 0.050 µM, respectively), consistent with effective cellular delivery. In fact, the potency of [Ga(L12)2]+ was increased by 270–895-fold relative to [Ga(NO3)3].Most notably, [Ga(L12)2]+ exhibited a 70-fold increase in potency in SK-N-MC cells, far exceeding that of its Cu(II) and Zn(II) analogues (1.3- and 2.4-fold, respectively). Similar enhancements relative to the ligand were also observed for [Ga(L10)2]+, [Ga(L11)2]+, and [Ga(L13)2]+, showing 18–34-fold increases in the same cell-type. In MCF-7 and MDA-MB-231 cells, Ga(III) complexes also exhibited greater activity than the ligands and their Cu(II)/Zn(II) analogues, although the enhancements were more modest (1.4–5.9-fold).
The pronounced increase in anti-proliferative activity observed for the Ga(III) complexes of L10–L13 is likely driven by multiple synergistic factors. These complexes probably act as lipophilic “Trojan horse” shuttles, enabling efficient cellular uptake of Ga(III), followed by intracellular transmetalation.14,68 Complexation significantly increases ligand lipophilicity, thereby promoting membrane diffusion and cellular accumulation.31,87,88 To evaluate this effect, both calculated and experimental n-octanol–water partition coefficients (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) were determined (Table S4).
P) were determined (Table S4).
The calculated log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values of the Ga(III) complexes were consistently and considerably higher than the experimentally determined (n-octanol–water) values (Table S4), because theoretical models treat [Ga(L)2]+ as a fully intact, pseudo-neutral complex in a non-hydrated environment.89 In contrast, under aqueous conditions, the high hydration energy of Ga(III) promotes ligand exchange, dissociation, and extensive solvation, leading to the formation of hydrated and ion-paired species with counter-anions such as NO3−.89,90 These polar, charge-dispersed entities partition preferentially into the aqueous phase, resulting in lower apparent lipophilicity. Consistent with this, UV-Vis analysis of the n-octanol layer showed that the absorption spectra of the Ga(III) complexes closely superimpose with the corresponding free ligands, differing only in intensity, confirming complex dissociation. Consequently, the experimental log
P values of the Ga(III) complexes were consistently and considerably higher than the experimentally determined (n-octanol–water) values (Table S4), because theoretical models treat [Ga(L)2]+ as a fully intact, pseudo-neutral complex in a non-hydrated environment.89 In contrast, under aqueous conditions, the high hydration energy of Ga(III) promotes ligand exchange, dissociation, and extensive solvation, leading to the formation of hydrated and ion-paired species with counter-anions such as NO3−.89,90 These polar, charge-dispersed entities partition preferentially into the aqueous phase, resulting in lower apparent lipophilicity. Consistent with this, UV-Vis analysis of the n-octanol layer showed that the absorption spectra of the Ga(III) complexes closely superimpose with the corresponding free ligands, differing only in intensity, confirming complex dissociation. Consequently, the experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values are much closer to those of the ligands, accurately reflecting the hydrophobicity of the solvated or partially dissociated species. In contrast, the calculated values overestimate lipophilicity by neglecting dissociation, solvent coordination, and hydrolytic equilibria intrinsic to Ga(III) chemistry.
P values are much closer to those of the ligands, accurately reflecting the hydrophobicity of the solvated or partially dissociated species. In contrast, the calculated values overestimate lipophilicity by neglecting dissociation, solvent coordination, and hydrolytic equilibria intrinsic to Ga(III) chemistry.
Notably, similar observations have been reported for related NNO–Ga(III) complexes, where the experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (1.10) differs markedly from the calculated value of 13.09, confirming that hydration and ion-pairing effects dominate the partitioning behaviour of these systems.89,90 Ga(III) closely resembles Fe(III) in ionic radius and charge, but is redox-inactive under physiological conditions, allowing it to interfere with iron-dependent redox processes essential for cancer proliferation.15,91 This chemical similarity facilitates its uptake by Fe-binding proteins such as ribonucleotide reductase, the rate-limiting enzyme in DNA synthesis.92 This ultimately leads to enzyme inhibition and potent anti-proliferative effects.16,78,93
P (1.10) differs markedly from the calculated value of 13.09, confirming that hydration and ion-pairing effects dominate the partitioning behaviour of these systems.89,90 Ga(III) closely resembles Fe(III) in ionic radius and charge, but is redox-inactive under physiological conditions, allowing it to interfere with iron-dependent redox processes essential for cancer proliferation.15,91 This chemical similarity facilitates its uptake by Fe-binding proteins such as ribonucleotide reductase, the rate-limiting enzyme in DNA synthesis.92 This ultimately leads to enzyme inhibition and potent anti-proliferative effects.16,78,93
The Ga(III)-mediated enhancement of the anti-proliferative activity of L10–L13 exceeds that reported for classical ONO-donor ligands such as pyridoxal isonicotinoyl hydrazone (PIH).31,86 For instance, Ga(III)–PIH complexes exhibited only a ∼5-fold increase in anti-proliferative activity in the same SK-N-MC cell type.31 Analogues such as HPKIH and HPKBH, when coordinated to Mn(II), Co(II), Ni(II), Cu(II), or Zn(II), showed no more than a 3- to 7-fold improvement.40 Even Ga(III) complexation with the promising di-2-pyridylketone-4-ethyl-4-methyl-3-thiosemicarbazone (Dp4e4mT) resulted in only a 1.5–1.8-fold increase under identical conditions, comparable to its Cu(II) and Zn(II) complexes.68
These findings establish specially designed Ga(III) complexes, particularly [Ga(L12)2]+, as a highly potent and mechanistically distinct class of anti-cancer agents. Their efficacy is driven not only by targeted metal chelation of critical intracellular pools, but also by a finely tuned set of properties made possible through the strategic incorporation of specific ligand design features. These include, NNO donor identity, lipophilic tuning, and the well-established ability of Ga(III) to mimic Fe(III) and disrupt intracellular iron metabolism.78–80 This work reinforces the therapeutic potential of Ga(III) complexes and highlights the pivotal role of coordination environment and scaffold engineering in driving next-generation metal-based anti-cancer strategies.
2.4. Cellular gallium uptake
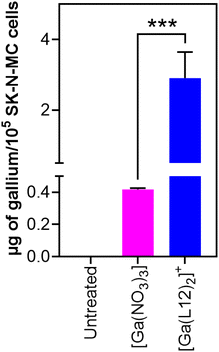
The enhanced cytotoxicity of [Ga(L12)2]+ (Table 2) prompted investigation of whether its activity arises from improved cellular delivery of gallium. Total gallium accumulation in SK-N-MC cells was quantified by inductively coupled plasma–mass spectrometry (ICP-MS; Fig. 3).94 Untreated cells contained negligible gallium,94 whereas treatment with [Ga(NO3)3] led to a modest intracellular level of 0.42 ± 0.01 µg Ga per 105 cells. In contrast, exposure to [Ga(L12)2]+resulted in a significant (p < 0.001) 7-fold higher accumulation (2.91 ± 0.73 µg Ga per 105 cells), confirming its markedly superior ability to deliver gallium into cells (Fig. 3). This enhanced uptake correlates closely with the substantially greater anti-proliferative potency of [Ga(L12)2]+ (IC50 = 0.099 µM versus 88.611 µM (Table S3) for [Ga(NO3)3]). This finding strongly supports the conclusion that Ga(III) coordination to the NNO-donor ligand significantly improves membrane permeability and intracellular transport of Ga(III).2.5. Electrochemistry
As described above in Table 1, among the newly synthesized agents, the tridentate naphthofuran ligands (L10–L13) exhibited the most potent anti-proliferative activity. To investigate the mechanistic basis of this behavior, we first examined the redox properties of their corresponding Cu(II) complexes, given the well-established cytotoxic and redox-active nature of Cu(II)-aroylhydrazone and Cu(II)-thiosemicarbazone complexes.18,40,52,67,68,75,85,95–97 Cyclic voltammetry was performed in MeCN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O (70
H2O (70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 v/v; Fig. 4), a solvent system previously optimized in our laboratory for assessing the electrochemical behavior of hydrazone complexes.40,41
30 v/v; Fig. 4), a solvent system previously optimized in our laboratory for assessing the electrochemical behavior of hydrazone complexes.40,41
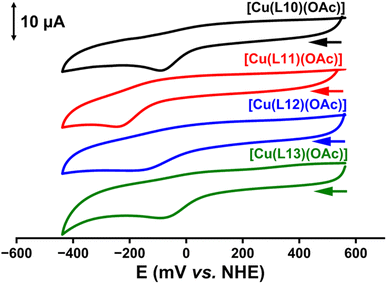
Cyclic voltammograms of [Cu(L10)(OAc)], [Cu(L11)(OAc)], [Cu(L12)(OAc)], and [Cu(L13)(OAc)] (100 µM) exhibited irreversible one-electron reduction waves at −26, −131, −51, and −8 mV versus the normal hydrogen electrode (NHE), respectively (Fig. 4). The observed reduction potential reflects the electronic effects of ligand substituents. For example, replacing the pyridine ring in [Cu(L10)(OAc)] (−26 mV) with a more electron-withdrawing pyrazine in [Cu(L13)(OAc)] (−8 mV) induces a positive shift, consistent with literature reports.74,98 Similarly, substituting the phenyl group in [Cu(L11)(OAc)] (−131 mV) with a pyridyl substituent in [Cu(L12)(OAc)] (−51 mV) results in a more positive potential, in agreement with trends reported for Cu(II) aroylhydrazones.40
To benchmark these findings, we compared the redox behavior of the L10–L13 Cu(II) complexes with the well-characterized, redox-active thiosemicarbazone complex [Cu(DpC)(OAc)], which exhibits a quasi-reversible potential of +15 mV versus NHE in the same solvent system (MeCN![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O, 7
H2O, 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 v/v).67 The more positive potential of [Cu(DpC)(OAc)] correlates with its well-described, marked redox activity and anti-proliferative potency52,67,75 relative to the modest anti-proliferative efficacy of the naphthofuran-based Cu(II) aroylhydrazone complexes (Table 2).
3 v/v).67 The more positive potential of [Cu(DpC)(OAc)] correlates with its well-described, marked redox activity and anti-proliferative potency52,67,75 relative to the modest anti-proliferative efficacy of the naphthofuran-based Cu(II) aroylhydrazone complexes (Table 2).
The irreversible behavior of the aroylhydrazone L10–L13 Cu(II) complexes is likely due to dissociation of the reduced Cu(I)-hydrazone species, in contrast to sulfur-containing ligands such as Dp44mT, which retain S-coordination in the Cu(I) state and thereby facilitate redox cycling.85 This inability to sustain redox cycling may limit ROS generation and contribute to the comparatively lower anti-cancer activity of these naphthofuran Cu(II) complexes (Table 2), relative to the structurally similar and potently cytotoxic Dp44mT Cu(II) complex (IC50 < 0.01 µM) in SK-N-MC cells.85 Nonetheless, the redox potentials of the L10–L13 Cu(II) complexes fall within the biologically relevant range (−0.4 to +0.8 V versus NHE) typically observed for redox-active species in aqueous systems.99
In summary, L10–L13 Cu(II) complexes may exhibit some redox activity, but this is likely to be less pronounced than their thiosemicarbazone counterparts. This is explored in a subsequent mechanistic section, following the examination of potential transmetalation of the Ga(III) complexes with the three most common transition metal ions in cells, namely Cu(II), Fe(III), and Fe(II).
2.6. Transmetalation of [Ga(L12)2]+ with Cu(II), Fe(III), or Fe(II): mechanistic insight into enhanced anti-proliferative activity
The superior anti-cancer activity of Ga(III) complexes of L10–L13 (Table 2) could be driven, in part, by their dissociative behavior and ability to transmetalate with Cu(II), Fe(II), or Fe(III), which are critical for tumor cell growth.1,4,5 We hypothesized that these complexes perturb metal homeostasis and mobilize intracellular Fe(II), contributing to selective cytotoxicity against cancer cells. This proposal was inspired by our previous findings, which showed that structurally related thiosemicarbazone Ga(III) complexes (e.g., [Ga(Dp4e4mT)2]+) readily transmetalate with Cu(II), correlating with their potent anti-proliferative effects.68 Comparable behavior has been observed in thiosemicarbazone and selenosemicarbazone Zn(II) complexes.68,69To explore this, we focused on [Ga(L12)2]+, the most active complex (Table 2), and studied its reactivity with Cu(II), Fe(III), and Fe(II) using UV-Vis spectroscopy, direct mass spectrometry (MS), liquid chromatography-mass spectrometry (LC-MS), and cyclic voltammetry (Fig. 5, 6 and S3–S13). Initial studies employed a 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 DMSO
3 DMSO![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O (v/v) solvent mixture to ensure solubility at relevant concentrations,69 with key experiments also repeated in 100% DMSO. To further mimic physiological conditions, critical experiments were performed in a near-fully aqueous solution (99.75% H2O/0.25% DMSO) in 0.14 M NaCl/20 mM HEPES (pH 7.4), termed “aqueous solution (0.14 M NaCl/pH 7.4)” from hereon.
H2O (v/v) solvent mixture to ensure solubility at relevant concentrations,69 with key experiments also repeated in 100% DMSO. To further mimic physiological conditions, critical experiments were performed in a near-fully aqueous solution (99.75% H2O/0.25% DMSO) in 0.14 M NaCl/20 mM HEPES (pH 7.4), termed “aqueous solution (0.14 M NaCl/pH 7.4)” from hereon.
This transmetalation was further validated by direct MS and LC-MS analyses (Fig. 6A–H). Direct MS of [Ga(L12)2]+ alone confirmed a single intact species (Fig. 6A). Under the labilizing LC-MS conditions,68,69 both [Ga(L12)2]+ and free L12 were detected, consistent with enhanced ligand dissociation (Fig. 6B). Upon addition of equimolar [Cu(OAc)2] (500 µM), direct MS identified [Cu(L12)DMSO]+ as the major product (Fig. 6C), while LC-MS revealed further dissociation, detecting [Cu(L12)DMSO]+ along with [Cu(L12)2] and L12 (Fig. 6D).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
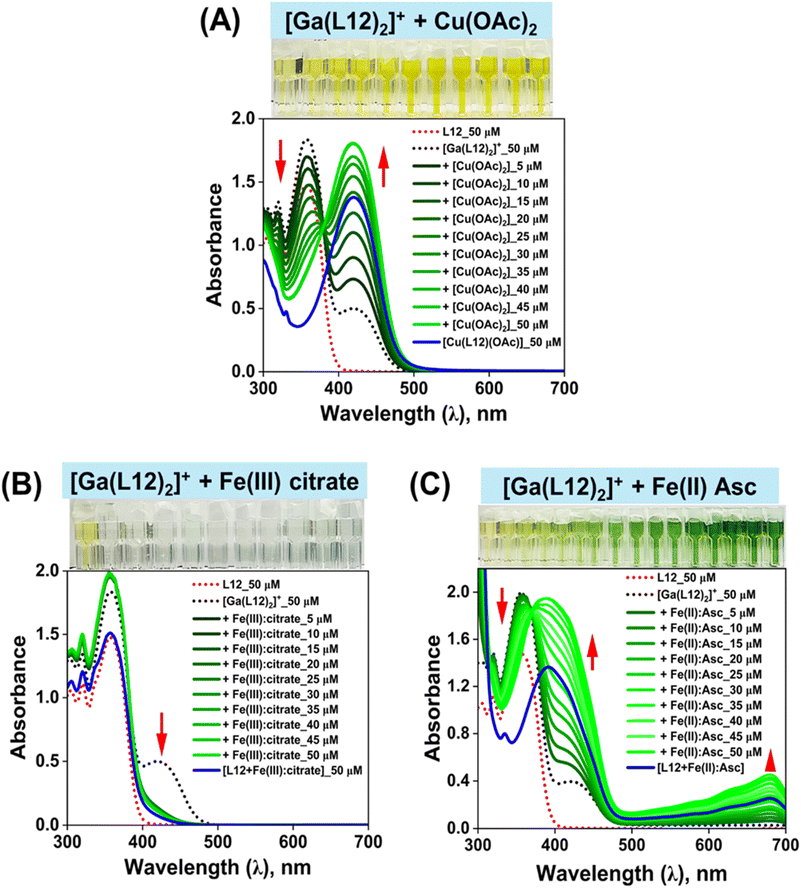
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 metal-to-citrate molar ratio) was performed using a large excess of citrate to prevent Fe(III) hydroxide precipitation under aqueous conditions.101 Citrate was specifically chosen as it is a physiologically relevant Fe(III) ligand implicated in the intracellular labile Fe pool delivered from transferrin,11 and the Fe(III) citrate complex can donate Fe to cells and intracellular proteins.102–105 Hence, Fe(III) citrate is used here to model the intracellular iron pool11 and to investigate its effect on [Ga(L12)2]+ to provide deeper understanding of its potent anti-proliferative activity against cancer cells (Table 2). Upon titration of the pale yellow [Ga(L12)2]+ with Fe(III) citrate, the solution became colorless (Fig. 5B), accompanied by the collapse of the 421 nm band and restoration of the π → π* transition at 356 nm, both indicative of ligand dissociation and release of free L12. This colorimetric and spectral change closely matched those of the free ligand (cf. red dotted line of synthesized and characterized L12 in Fig. 5B), confirming Ga–ligand dissociation without the formation of L12–Fe(III) complexes.
100 metal-to-citrate molar ratio) was performed using a large excess of citrate to prevent Fe(III) hydroxide precipitation under aqueous conditions.101 Citrate was specifically chosen as it is a physiologically relevant Fe(III) ligand implicated in the intracellular labile Fe pool delivered from transferrin,11 and the Fe(III) citrate complex can donate Fe to cells and intracellular proteins.102–105 Hence, Fe(III) citrate is used here to model the intracellular iron pool11 and to investigate its effect on [Ga(L12)2]+ to provide deeper understanding of its potent anti-proliferative activity against cancer cells (Table 2). Upon titration of the pale yellow [Ga(L12)2]+ with Fe(III) citrate, the solution became colorless (Fig. 5B), accompanied by the collapse of the 421 nm band and restoration of the π → π* transition at 356 nm, both indicative of ligand dissociation and release of free L12. This colorimetric and spectral change closely matched those of the free ligand (cf. red dotted line of synthesized and characterized L12 in Fig. 5B), confirming Ga–ligand dissociation without the formation of L12–Fe(III) complexes.
To additionally investigate Fe(III) binding, L12 (50 µM) was titrated with Fe(III) citrate (5–75 µM) in DMSO/H2O (7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 v/v), which resulted in negligible spectral changes (Fig. S3A). Similar outcomes were observed upon adding FeCl3 to L12 in 100% DMSO (Fig. S3B) or physiologically relevant aqueous solution (0.14 M NaCl/pH 7.4; Fig. S3C). This indicated that L12 exhibits minimal ligation of Fe(III) under all tested conditions. To isolate the role of citrate, it was titrated alone into [Ga(L12)2]+, leading to a loss of the 421 nm band, restoration of the 356 nm band (Fig. S4A), and a visible color change from pale yellow to colorless. These changes mirrored those induced by Fe(III) citrate (Fig. 5B), suggesting that both Fe(III) citrate and citrate alone effectively displace L12 from Ga(III), likely forming [Ga(citrate)2] species. This was confirmed by direct MS and LC-MS analyses (Fig. 6E–F and S5C–D), which showed complete ligand dissociation and formation of Ga-citrate species, with no evidence of Fe(III)–L12 complexes.
3 v/v), which resulted in negligible spectral changes (Fig. S3A). Similar outcomes were observed upon adding FeCl3 to L12 in 100% DMSO (Fig. S3B) or physiologically relevant aqueous solution (0.14 M NaCl/pH 7.4; Fig. S3C). This indicated that L12 exhibits minimal ligation of Fe(III) under all tested conditions. To isolate the role of citrate, it was titrated alone into [Ga(L12)2]+, leading to a loss of the 421 nm band, restoration of the 356 nm band (Fig. S4A), and a visible color change from pale yellow to colorless. These changes mirrored those induced by Fe(III) citrate (Fig. 5B), suggesting that both Fe(III) citrate and citrate alone effectively displace L12 from Ga(III), likely forming [Ga(citrate)2] species. This was confirmed by direct MS and LC-MS analyses (Fig. 6E–F and S5C–D), which showed complete ligand dissociation and formation of Ga-citrate species, with no evidence of Fe(III)–L12 complexes.
To further examine the role of citrate, FeCl3 was titrated directly into [Ga(L12)2]+ in 100% DMSO or physiologically relevant aqueous solution (0.14 M NaCl/pH 7.4), with no observable spectral changes or evidence of transmetalation (Fig. S4B–C). These results indicate that Fe(III) alone is unable to displace L12 under either organic or physiological conditions.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 v/v), using a 1
3 v/v), using a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 metal-to-Asc molar ratio to stabilize Fe(II) in aqueous solution (Fig. 5C). An excess of Asc was added to maintain Fe(II) in solution, as it is a well-known physiologically relevant reductant and weak ligand implicated in the intracellular labile Fe pool11 and is involved in cellular Fe metabolism.106,107 Hence, as the labile intracellular iron pool is probably composed of multiple ligands and complexes,11 Fe(II) Asc is used here to examine its effect on [Ga(L12)2]+ to dissect the mechanism of its marked efficacy against cancer cells (Table 2). Upon incremental addition of Fe(II) Asc (5–50 µM) to [Ga(L12)2]+ (50 µM; 70% DMSO/30% H2O), the solution rapidly changed from pale yellow to deep green, with an isosbestic point at 370 nm (Fig. 5C). This was accompanied by a bathochromic shift of the π → π* band (356 nm to 391 nm), loss of the 421 nm Ga–ligand band, and emergence of a 680 nm peak characteristic of Fe(II) → ligand charge transfer (MLCT).108 Similar spectral alterations were observed in a physiologically relevant aqueous solution (0.14 M NaCl/pH 7.4; Fig. S4D), demonstrating the consistency of this reaction across solvent systems.
100 metal-to-Asc molar ratio to stabilize Fe(II) in aqueous solution (Fig. 5C). An excess of Asc was added to maintain Fe(II) in solution, as it is a well-known physiologically relevant reductant and weak ligand implicated in the intracellular labile Fe pool11 and is involved in cellular Fe metabolism.106,107 Hence, as the labile intracellular iron pool is probably composed of multiple ligands and complexes,11 Fe(II) Asc is used here to examine its effect on [Ga(L12)2]+ to dissect the mechanism of its marked efficacy against cancer cells (Table 2). Upon incremental addition of Fe(II) Asc (5–50 µM) to [Ga(L12)2]+ (50 µM; 70% DMSO/30% H2O), the solution rapidly changed from pale yellow to deep green, with an isosbestic point at 370 nm (Fig. 5C). This was accompanied by a bathochromic shift of the π → π* band (356 nm to 391 nm), loss of the 421 nm Ga–ligand band, and emergence of a 680 nm peak characteristic of Fe(II) → ligand charge transfer (MLCT).108 Similar spectral alterations were observed in a physiologically relevant aqueous solution (0.14 M NaCl/pH 7.4; Fig. S4D), demonstrating the consistency of this reaction across solvent systems.
Control experiments confirmed that the latter features were specific to Fe(II) coordination (Fig. S3D–F). Titration of L12 alone with Fe(II) Asc resulted in nearly identical spectral features (391 and 680 nm)108 and a green coloration (Fig. S3D), consistent with Fe(II)–L12 complex formation in 70% DMSO/30%H2O. Similar results were observed upon the addition of Fe(II) (as (NH4)2Fe(SO4)2) to L12 in both the presence (Fig. S3E) and absence (Fig. S3F) of Asc in aqueous solution (0.14 M NaCl/pH 7.4). Direct MS of [Ga(L12)2]+ mixed with equimolar Fe(II) Asc identified [Fe(L12)2], [Ga(Asc)]+, and free L12 (Fig. 6G), while the more dissociative LC-MS68,69 detected [Fe(L12)(Asc)], [Ga(Asc)]+, and Asc (Fig. 6H), confirming complete transmetalation.
To examine the effect of Asc alone, its titration into [Ga(L12)2]+ (50 µM; pale yellow) induced a gradual loss of the 421 nm band and resulted in a colorless solution (Fig. S4E). This result suggests that Asc directly coordinates with Ga(III) and displaces L12. Both direct MS and LC-MS confirmed the formation of [Ga(Asc)2] and free L12 (Fig. S5E–F), paralleling the behavior observed with citrate (Fig. S5C–D). Thus, Asc and citrate, which are both potential ligands of the intracellular iron pool,11 act as oxygen donors capable of sequestering Ga(III) and releasing L12.
Notably, similar Fe(II)-driven transmetalation was observed across the Ga(III) complexes of L10, L11, and L13 (data not shown), suggesting that this behavior is general to this ligand series. Collectively, these results demonstrate a clear preference of L12 for Fe(II) over Fe(III), as [Ga(L12)2]+ undergoes complete ligand exchange with Fe(II) ascorbate, whereas Fe(III) citrate promotes complete ligand dissociation without formation of an Fe(III)–L12 complex. The complete ligand exchange of [Ga(L12)2]+ is likely driven by the high charge density of Ga(III), a hard Lewis acid with a strong thermodynamic preference for oxygen-donor ligands such as citrate.109–111 The lability of Ga(III) complexes to ascorbate and citrate mirrors that of established gallium therapeutics (e.g., gallium nitrate, gallium maltolate), which act as prodrugs, releasing Ga(III) and Fe(II)-intercepting ligands within cells to enhance selectivity and therapeutic potential. These results highlight the role of metal ion identity and redox state in governing ligand exchange and provide mechanistic insight into how [Ga(L12)2]+ perturbs intracellular iron pools to inhibit proliferation (Table 2).
2.7. Comparative transmetalation and ligand liberation studies of Zn(II) and Cu(II) complexes
Unlike [Ga(L12)2]+, [Zn(L12)Cl2] and [Cu(L12)(OAc)] undergo only partial dissociation under the same conditions, forming mixed-ligand species. These differences highlight Ga(III)'s stronger affinity for common oxygen-donor ligands thought to play a role in the intracellular iron pool, e.g., citrate and Asc11 relative to the softer NNO coordination sphere of L12. This difference facilitates complete ligand release and may account for the superior cytotoxicity of [Ga(L12)2]+ relative to its Zn(II) and Cu(II) counterparts (Table 2). Detailed descriptions of these findings are provided in the SI Discussion (Fig. S6–S12).2.8. Electrochemical investigation of [Ga(L12)2]+ transmetalation with Fe and Cu
Building on insights from the UV-Vis and MS studies above, cyclic voltammetry was employed to investigate the transmetalation dynamics and redox behavior of [Ga(L12)2]+ (100 µM) in the presence of 100 µM of either Cu(OAc)2, Fe(II) Asc (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 molar ratio), or Fe(III) citrate (1
100 molar ratio), or Fe(III) citrate (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 molar ratio; Fig. S13A and B). Of note, Zn(II) was not examined as results in a redox inactive complex with a potential of ∼−900 mV versus the NHE, as shown for similar Zn(II) complexes.40,69,70 These electrochemistry studies aimed to assess if Ga(III) is displaced from L12 by Cu(II), Fe(II), or Fe(III). As expected for a redox-inert Ga(III) complex,68 [Ga(L12)2]+ alone showed no significant redox features between −400 and +600 mV (Fig. S13A). Upon addition of Cu(OAc)2, the voltammogram revealed an irreversible one-electron reduction at −51 mV vs. NHE, identical to that of independently synthesised [Cu(L12)OAc] (Fig. 4), confirming Cu(II)-driven transmetalation, as further supported by UV-Vis (Fig. 5A), direct MS (Fig. 6C), and LC-MS (Fig. 6D).
100 molar ratio; Fig. S13A and B). Of note, Zn(II) was not examined as results in a redox inactive complex with a potential of ∼−900 mV versus the NHE, as shown for similar Zn(II) complexes.40,69,70 These electrochemistry studies aimed to assess if Ga(III) is displaced from L12 by Cu(II), Fe(II), or Fe(III). As expected for a redox-inert Ga(III) complex,68 [Ga(L12)2]+ alone showed no significant redox features between −400 and +600 mV (Fig. S13A). Upon addition of Cu(OAc)2, the voltammogram revealed an irreversible one-electron reduction at −51 mV vs. NHE, identical to that of independently synthesised [Cu(L12)OAc] (Fig. 4), confirming Cu(II)-driven transmetalation, as further supported by UV-Vis (Fig. 5A), direct MS (Fig. 6C), and LC-MS (Fig. 6D).
UV-Vis spectrophotometry (Fig. 4B and C) and MS data (Fig. 5E–H) provide compelling evidence that [Ga(L12)2]+ selectively transmetallates with Fe(II) over Fe(III). To probe this electrochemically, the voltammogram scan range for [Ga(L12)2]+ was extended from −1200 to +1200 mV following the addition of either Fe(II) Asc or Fe(III) citrate (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 molar ratio; Fig. S13B). [Ga(L12)2]+ alone exhibited an irreversible reduction at −425 mV vs. NHE, whereas Fe(II) Asc showed a one-electron irreversible reduction at +626 mV. Notably, Fe(II) Asc exhibited two anodic processes, with the lower-potential peak corresponding to Fe(II) oxidation, and the higher-potential peak to direct ascorbate oxidation.112 Mixing L12 with Fe(II) Asc resulted in a green solution, with a shifted potential of +739 mV, representing a +113 mV increase consistent with Fe(II) coordination to L12 (Fig. S13B). This observation was in agreement with UV-Vis data confirming L12 Fe(II) complex formation in the presence of Fe(II) Asc (Fig. S3D).
100 molar ratio; Fig. S13B). [Ga(L12)2]+ alone exhibited an irreversible reduction at −425 mV vs. NHE, whereas Fe(II) Asc showed a one-electron irreversible reduction at +626 mV. Notably, Fe(II) Asc exhibited two anodic processes, with the lower-potential peak corresponding to Fe(II) oxidation, and the higher-potential peak to direct ascorbate oxidation.112 Mixing L12 with Fe(II) Asc resulted in a green solution, with a shifted potential of +739 mV, representing a +113 mV increase consistent with Fe(II) coordination to L12 (Fig. S13B). This observation was in agreement with UV-Vis data confirming L12 Fe(II) complex formation in the presence of Fe(II) Asc (Fig. S3D).
When Fe(II) Asc was added to [Ga(L12)2]+, the pale-yellow solution immediately turned green, and the voltammogram revealed a single irreversible oxidation at +739 mV vs. NHE, corresponding to the Fe(II)/Fe(III) couple (Fig. S13B). This electrochemical signature was identical to that obtained for Fe(II) Asc with free L12 (Fig. S13B), confirming Fe(II) coordination to L12 following transmetalation. The formation of the Fe(II) L12 complex was further substantiated by UV-Vis spectrophotometry (Fig. 5C and S3D) and MS analyses (Fig. 6G and H). Notably, the +739 mV potential of [L12 + Fe(II) Asc] is significantly higher than that of related NNO-type hydrazone Fe(II) complexes, such as [Fe(PCIH)2] (+616 mV vs. NHE in 50% DMF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) H2O), likely reflecting solvent-dependent stabilization effects.113
H2O), likely reflecting solvent-dependent stabilization effects.113
The redox behavior of Fe(III) citrate was next examined alone and in combination with L12 (Fig. S13B). In both cases, the voltammogram profiles were very similar, showing a one-electron irreversible reduction at −374 mV vs. NHE, in agreement with reported values for Fe(III) citrate17 (Fig. S13B). This confirms that L12 does not bind Fe(III) from Fe(III) citrate, consistent with UV-Vis spectrophotometry (Fig. S3A). In contrast, addition of Fe(III) citrate to [Ga(L12)2]+ caused the pale-yellow solution to turn colorless, and the potential to shift markedly to −680 mV vs. NHE (Fig. S13B). Since [Ga(L12)2]+ alone has a potential of −425 mV vs. NHE, this large negative shift indicates a fundamentally different redox process. Given the redox-inert nature of Ga(III),68 the shift is most plausibly attributed to a ligand-centred reduction, consistent with the formation of a Ga(III) citrate complex. This assignment is further supported by direct MS and LC-MS analyses (Fig. 6E and F). In summary, cyclic voltammetry (Fig. S13) provides electrochemical evidence, fully consistent with UV-Vis (Fig. 5A–C) and MS data (Fig. 6A–H), showing that Cu(II) and Fe(II) readily transmetalate with Ga(III) in [Ga(L12)2]+, whereas Fe(III) does not.
2.9. Cu(II) complexes of L10–L13 exhibit lower redox activity than [Cu(DpC)Cl2]
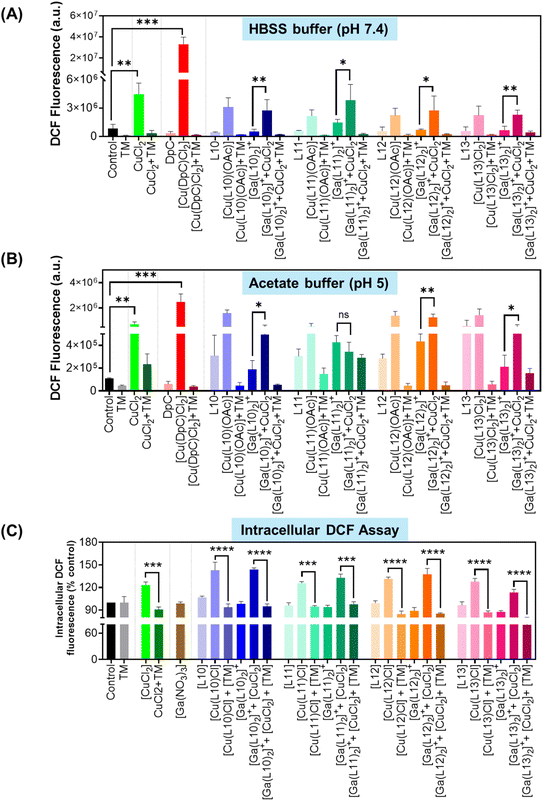
The anti-cancer activity of aroylhydrazones18 and thiosemicarbazones49,67,69 is often attributed to the redox activity of their Cu(II) complexes,49,67,69 which generate reactive oxygen species (ROS) to induce lysosomal membrane permeabilization and cell death.49,67,69 In this context, the Ga(III) complexes of L10–L13 may acquire part of their anti-proliferative activity through intracellular transmetalation with Cu(II), forming redox-active complexes.68 Electrochemical studies showed irreversible redox behavior for these Cu(II) complexes (Fig. 4), suggesting a limited ability to sustain redox cycling.To evaluate this, we assessed 2′,7′-dichlorodihydrofluorescein (H2DCF) oxidation at pH 7.4 (cytosolic) and pH 5.0 (lysosomal) in the presence of Cu(II) and Ga(III) complexes (10 µM), L-cysteine, and H2O2 (100 µM).67–69,75 The generation of ROS was assessed in solution under these controlled conditions to understand intrinsic redox activity in these critical compartments that are known to be important for anti-proliferative efficacy49,67,85 rather than whole cell activity. Moreover, our laboratories have used this assay under the same conditions for many series of similar ligands and their complexes, enabling specific comparison.68–70,75 Fluorescence was recorded and compared to controls, including [Cu(DpC)Cl2] (positive control), CuCl2, DpC (negative control), and the redox-inactive Cu(II) chelator tetrathiomolybdate (TM).67–69,75 TM, DpC, and all free L10–L13 ligands produced no increase in DCF fluorescence under either pH condition, consistent with redox inactivity.67–69,75
In contrast, CuCl2 and [Cu(DpC)Cl2] induced significant fluorescence (p < 0.001–0.01), with greater activity at pH 7.4.67–69 The Cu(II) complexes of L10–L13 showed significantly (p < 0.001–0.05) lower redox activity than [Cu(DpC)Cl2] (Fig. 7A and B), attributable to the NNO donor set, which confers less redox potential than the NNS coordination sphere of DpC.49,67–69,85 These complexes exhibited modest ROS activity, with L10, L12, and L13 showing slight, but significant (p < 0.05) increases versus CuCl2 at pH 5.0, implicating a pH-dependent redox enhancement likely due to ligand protonation and altered Cu(II)/Cu(I) cycling.
Similarly to the ligands alone, the Ga(III) complexes of L10–L13 demonstrated little redox activity across both pH values, as expected from the chemistry of Ga(III).14 However, addition of CuCl2 to the Ga(III) complexes resulted in a significant (p < 0.01–0.05) increase in H2DCF oxidation, indicating transmetalation with Cu(II), although redox activity remained equal to or lower than that of CuCl2 alone (Fig. 7A and B). Notably, [Ga(L12)2]+ at pH 5.0 showed a slight but significant (p < 0.05) increase beyond CuCl2, aligning with transmetalation data from UV-Vis, direct MS, and LC-MS experiments. In contrast, Fe(III) and Fe(II) complexes of L12 resulted in a 1- to 7-fold decrease in ROS generation compared to the Cu(II) complex (data not shown), confirming that iron-mediated redox cycling plays a minimal role in this system.
To complement the extracellular DCF assays (Fig. 7A and B), intracellular ROS production was quantified using DCF-DA (10 µM) upon incubation of SK-N-MC cells for 4 h/37 °C with CuCl2, or the Cu(II) or Ga(III) complexes of L10–L13 (5 µM) in the presence or absence of CuCl2 or TM (5 µM; Fig. 7C). Consistent with their minimal extracellular redox activity, [Ga(NO3)3], the free ligands L10–L13 and their Ga(III) complexes produced no significant increase in fluorescence relative to control (Fig. 7C). In contrast, a marked and significant (p < 0.0001–0.001) increase in intracellular ROS was observed upon incubation with CuCl2 or the Ga(III) complexes with CuCl2. This increase in redox activity observed was similar to that observed for the respective Cu(II) complexes. For all Cu(II) complexes and CuCl2, the addition of TM markedly and significantly (p < 0.0001–0.001) ablated their redox activity (Fig. 7C). Moreover, the activity of the Cu(II) observed was similar to, or only slightly greater than, that of CuCl2 alone, suggesting only minor redox activity. These findings suggest that the oxidative stress originates from Cu-dependent transmetalation of the Ga(III) complex, rather than from Ga(III) itself.
Collectively, these data in Fig. 7A–C demonstrate that the Ga(III) complexes of L10–L13 remain redox-silent themselves, but act as delivery vehicles that may transmetallate with intracellular Cu pools to trigger minor oxidative stress.
2.10. Interception of cellular iron pools: 59Fe mobilization and uptake inhibition by L1–L13
To assess the ability of the new ligands to access intracellular Fe pools, two sensitive and widely validated radionuclide-based assays24,31,32,43,52 were employed: (1) 59Fe mobilization from SK-N-MC cells prelabeled with 59Fe2-transferrin (Tf) (Fig. 8A); and (2) inhibition of 59Fe uptake from 59Fe2-Tf by SK-N-MC cells (Fig. 8B).Given the rapid dissociation of Ga(III) complexes under cellular conditions (as determined in Fig. 5, S4 and S5), iron mobilization and uptake inhibition assays were conducted using the free ligand to directly model the active intracellular species. Comparative analysis included four positive control ligands, namely DFO, Dp44mT, DpC, and 311, previously characterized for their differential Fe-chelation efficiency.31,43,114
Given the short incubation period (3 h), no cytotoxicity was observed using all ligands, with cell viability exceeding 96% (data not shown). Control media alone induced minimal 59Fe mobilization (<5%) from SK-N-MC cells prelabeled with 59Fe2-Tf, consistent with prior studies.32,43,52 Amongst the control ligands,32,43,52 Dp44mT and 311 showed the highest activity, mobilizing 44–47% of total cellular 59Fe, while DpC and DFO were less effective (12–17%; Fig. 8A). Considering the bidentate ligands (L1–L9), L1 displayed the highest 59Fe mobilization activity (31%), significantly (p < 0.01) exceeding that of DFO and DpC. L2 and L6 showed modest efficacy (11–15%), while L3–L5 and L7–L9 were largely inactive, with mobilization comparable to control media (4–6%). In contrast, the tridentate ligands, L10–L13, consistently outperformed most bidentate analogues, mobilizing 20–35% of total cellular 59Fe (Fig. 8A).
The 59Fe uptake assays (Fig. 8B) evaluated the ability of ligands to intercept and prevent internalized 59Fe uptake from 59Fe2-Tf by the cells. Uptake values were normalized to control medium (100%) and showed an inverse correlation with 59Fe mobilization capacity (Fig. 8A). Ligands that promoted strong 59Fe release also suppressed the uptake. Among the positive controls, Dp44mT and 311 were most effective, reducing 59Fe uptake to 8–13%, while DFO and DpC were comparatively weaker (53–66%; Fig. 8B), consistent with prior reports.32,43,52
Of the bidentate ligands tested (L1–L9), L1 was the most effective, reducing 59Fe uptake from 59Fe2-Tf to 28% of control levels. In contrast, the remaining ligands showed minimal or inconsistent effects, with L8 unexpectedly increasing the uptake to 127% (Fig. 8B). The tridentate naphthofuran ligands, L10–L13, consistently outperformed their bidentate counterparts, reducing 59Fe uptake to 44–78% of control. This enhanced efficacy is attributed to their NNO donor motif and rigid naphthofuran scaffold, which promotes selective coordination of Fe(II) observed using UV-Vis and MS (Fig. 5, 6, and S3). As Fe(II) comprises the labile intracellular pool generated after transferrin endocytosis and reduction,9,10 these findings indicate that L10–L13 act by targeting Fe(II) within cells, highlighting a distinct mechanism from that of classical Fe(III)-targeting chelators.
3 Conclusions
Tridentate aroylhydrazones and thiosemicarbazones have long served as versatile scaffolds for the development of therapeutics in both iron overload and oncology contexts.26,31,33,42,84,115,116 Our findings reveal that Ga(III) complexes of novel tridentate NNO-donor naphthofuran ligands (L10–L13) display uniquely potent anti-proliferative activity via a multifaceted mechanism distinct from classical metal chelators. Key to their efficacy is the selective interception of Fe(II), not Fe(III), following endosomal reduction and release from transferrin.12,58 This was demonstrated through highly sensitive 59Fe mobilization and uptake inhibition assays, where L10–L13 robustly disrupted intracellular iron trafficking (Fig. 8). These ligands decreased 59Fe uptake from 59Fe2-Tf to 44–78% of control levels, while promoting intracellular mobilization aligning with their structural design to favor Fe(II) binding (Fig. 9).Mechanistically, the Ga(III) complexes act as lipophilic “Trojan horse” agents, facilitating cellular uptake and rapid intracellular dissociation in the presence of physiologically relevant reducing (e.g., ascorbate) and chelating agents, e.g., citrate and ascorbate (Fig. 9).11,106,107 This releases both cytotoxic Ga(III) and the Fe(II)-targeting ligand (Fig. 5, 6, and 8). As shown in Fig. 3 and Table S3 in SK-N-MC cells, [Ga(L12)2]+ achieved markedly and significantly (p < 0.001) higher gallium accumulation and a significant (p < 0.0001) 895-fold increase in potency (IC50 = 0.099 µM versus 88.611 µM) relative to Ga(NO3)3), indicating that [Ga(L12)2]+ acts as a transport carrier facilitating cell entry. Once inside cells, the Ga(III) complex dissociates, liberating the Fe(II)-binding ligand that results in strong 59Fe mobilization and inhibition of 59Fe uptake from 59Fe-transferrin (Fig. 8).
Multiple lines of spectroscopic evidence confirm Ga(III)–ligand dissociation and subsequent transmetalation with Fe(II) (Fig. 3, 5, 6 and S3–5), underscoring a key and innovative design principle, namely, dissociative lability coupled with selective Fe(II) coordination enables therapeutic amplification. Compared to their Cu(II) and Zn(II) counterparts, which showed only 1.02–3.1-fold improvements, Ga(III) complexes enhanced activity up to 70-fold increase in anti-proliferative efficacy, far exceeding other Ga(III)-based agents, including those tested clinically.14,31,40,68,86,117 Notably, [Ga(L12)2]+ emerged as the most potent agent (Table 2). Unlike redox-active thiosemicarbazone–Cu(II) complexes, the L10–L13 Cu(II) and Ga(III) complexes displayed minimal redox cycling, favoring redox-independent cytotoxicity and reducing off-target oxidative stress.14,78–80,118,119
Our electrochemical and DCF-based ROS assays confirmed that Cu(II) complexes of L10–L13 exhibited irreversible redox behavior and weak ROS generation, while Ga(III) complexes remained inert (Fig. 4 and 7). These observations indicate that cytotoxicity is not markedly driven by redox activity, but by disruption of Fe homeostasis via selective Fe(II) interception and Ga(III)-mediated interference with Fe-dependent enzymes such as ribonucleotide reductase (Fig. 9). Importantly, these properties were preserved across diverse solvent systems, including physiological aqueous solution (0.14 M NaCl/pH 7.4), highlighting their biological relevance. Spectroscopic evidence further confirmed preferential Fe(II) coordination due to favorable electronic, steric, and hard–soft-acid–base factors. The ability of Fe(II) to engage in π-back bonding and adopt low-spin configurations120 contributes to its stabilization with NNO-donor ligands, an interaction not observed with Fe(III).121
Diferric transferrin (Fe2-Tf) delivers Fe(III) into the cell via transferrin receptor 1 (TfR1)-mediated endocytosis. Within the endosome, Fe(III) is reduced to Fe(II) by STEAP3 and transported into the cytoplasm through DMT1, contributing to the labile intracellular Fe pool. The [Ga(L12)2]+ complex enters the cell via a “Trojan horse” mechanism, whereby the lipophilic Ga(III) complex exhibits enhanced permeability. Once inside the cell, it undergoes rapid dissociation triggered by physiological reducing and chelating agents, such as ascorbate and citrate and ascorbate, respectively, disrupting Fe-dependent processes by two mechanisms. This occurs first via the liberated Ga(III) interacting with Fe-containing proteins, including the rate-limiting step of DNA synthesis, ribonucleotide reductase, inhibiting its activity.14,16,78,80 Second, the dissociated tailor designed NNO ligand specifically binds Fe(II) that has entered the cell (as observed in studies demonstrating inhibition of 59Fe uptake from 59Fe2-Tf; Fig. 8B) to form the [Fe(L12)2] complex that leads to cellular Fe release (that is observed as cellular 59Fe mobilization; Fig. 8A).
The differences in potency and mechanism between the classical NNS-donor thiosemicarbazones (DpC and Dp44mT) and the present NNO-donor naphthofuran ligands (L10–L13) arise from their distinct redox properties, donor-atom environments, and intracellular metal-handling behaviour. DpC and Dp44mT possess NNS-donor sets that possess the well-known double-punch mechanism.49,59 That is, after binding the essential cellular nutrients, Fe and Cu, these complexes redox cycle generating high ROS levels that trigger lysosomal membrane permeabilization and oxidative cell death.49,62 In fact, their potency arises primarily from redox-driven toxicity mediated by reactive Cu species, as reflected by the quasi-reversible reduction potential of +15 mV vs. NHE for [Cu(DpC)(OAc)], which supports facile electron transfer and catalytic ROS production.49,52,67,69
In contrast, the NNO-donor framework of L10–L13 was deliberately incorporated to suppress redox activity and operate through a redox-independent, Fe(II)-interception pathway. Cyclic voltammetry (Fig. 4) and solution redox assays (Fig. 7A and B) revealed that their Cu(II) complexes undergo irreversible reductions between −131 mV and −8 mV, indicating far lower redox activity than the Cu(II) complexes of DpC or Dp44mT. Moreover, the Ga(III) complexes are redox-inert, confirming that their biological effects do not arise from redox cycling. This fundamental change in donor atom identity (NNOversus NNS) redirects the mechanism from ROS-driven oxidative stress to metal-ion trafficking disruption.
The coordination of Ga(III) by the NNO ligands, L10–L13, further promotes their anti-proliferative efficacy through a prodrug-like transmetalation process. Upon cellular entry, [Ga(L)2]+ dissociates to release both Ga(III) and the free ligand, which selectively binds Fe(II) generated during endosomal Fe(III) reduction (Fig. 5, 6 and S3–S5). The dissociation of the Ga(III) complex leads to marked cellular accumulation of cytotoxic Ga(III) (Fig. 3), which is well known to inhibit critical Fe-dependent processes, including DNA synthesis.14–16 Furthermore, the interception of the essential nutrient, Fe(II), accounts for the strong 59Fe mobilization and the inhibition of 59Fe uptake from 59Fe2-Tf observed (Fig. 8), demonstrating that the Ga(III)–L10–L13 complexes disrupt cellular iron trafficking without inducing marked oxidative stress.
Therefore, the comparable potency of [Ga(L10)2]+, [Ga(L12)2]+, and [Ga(L13)2]+ to DpC and Dp44mT particularly in SK-N-MC cells (Table 2), reflects a totally different mechanism that reflects the unique chemistries of NNS versus NNO ligands. In fact, while DpC and Dp44mT exploit the double punch mechanism and particularly Cu-mediated redox stress, the L10–L13 Ga(III) complexes achieve similar efficacy through Ga(III) dissociation and accumulation and Fe(II) interception. This latter mechanism minimizes off-target oxidative damage, providing a distinct and possibly safer therapeutic paradigm supported by our electrochemical, spectroscopic, and 59Fe transport data.
Collectively, this investigation establishes that strategic control over ligand donor identity, lipophilicity, metal ion lability, and Fe(II) targeting defines a novel mechanism of action for Ga(III) complexes. This paradigm transcends classical chelation and sets a precedent for the rational development of non-redox active, Fe(II)-intercepting anti-cancer agents that hijack intracellular iron trafficking to exert effective anti-proliferative activity.
Author contributions
D. R. R., S. F., M. D., D. K., and B. K. conceptualized the study, supervised research personnel, analyzed data, secured grant funding, and contributed to manuscript writing and editing. S. F., M. D., D. K., and R. A. were responsible for compound synthesis and characterization, data analysis, cellular and molecular studies, and drafting sections of the manuscript. S. F., M. D., B. K., R. A., D. K., T. P. W., M. S., M. G. A., V. R., and A. F. Z. contributed to manuscript revisions, conducted cellular or molecular experiments, procured materials, and prepared figures. W. L. and P. V. B. performed data analysis, carried out X-ray diffraction studies, and assisted in writing and editing portions of the manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
All compounds were verified to have a purity exceeding 95% through elemental analysis (C, H, N, S) and further confirmed via NMR spectroscopy, MS, and single-crystal X-ray crystallography (Tables S1 and S2). Anti-proliferative activity of [Ga(L12)2]+, L12, [Ga(NO3)3], and DpC in SK-N-MC and MCF-7 cells, and comparison of calculated versus experimental log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values for ligands (L10–L13) and their Ga(III) complexes (Tables S3 and S4).
P values for ligands (L10–L13) and their Ga(III) complexes (Tables S3 and S4).
Supplementary information (SI): details on materials, general methods, crystallographic study methodology, crystallographic refinement data and unit cell packing diagrams, electrochemistry, transmetalation, titration studies via UV-Vis spectrophotometry, synthetic procedures and characterization data for ligands, and their Cu(II), Zn(II), and Ga(III) complexes, experimental protocols for biological assays, the reactivity of Ga(III), Zn(II), and Cu(II) complexes in the presence of citrate or Asc, assessed using UV-Vis spectroscopy, direct MS, and LC-MS. UV-Vis and MS (direct and LC–MS) studies showing spectral and reactivity changes of L12 and [Ga(L12)2]+ upon titration with Fe(III)–citrate or Fe(II)–ascorbate (Fig. S3–S5). Comprehensive transmetalation and titration experiments involving [Zn(L12)Cl2] and [Cu(L12)(OAc)] with Cu(II), Fe(III), and Fe(II) are presented and discussed in Fig. S6–S13. Additionally, it includes the NMR studies as Fig. S14–S55. See DOI: https://doi.org/10.1039/d5sc06084b.
CCDC 1982904–1982909, 1982911 and 2424290 contain the supplementary crystallographic data for this paper.122a–h
Acknowledgements
D. R. R. thanks the National Health and Medical Research Council (NHMRC) of Australia for Senior Principal Research Fellowships (APP1062607 and 1159596), Australian Research Council Discovery Grant (DP200103530), NHMRC Ideas Grant (2010632), and NHMRC Project Grants (APP1144829; APP1128152; and APP1144456). D. R. R., M. D., and P. V. B. thank the NHMRC for an Ideas Grant (2019160) and National Breast Cancer Foundation of Australia (NBCF) for an Investigator Initiated Research Scheme Grant (IIRS-23-004). M. D., B. K. and D. R. R. (mentor) thank the NHMRC MRFF for an Early to Mid-Career Researchers grant (2027365); M. D. and D. R. R. appreciate support from the Thrasher Research Fund USA Early Career Award. M. D. appreciates a Griffith University Postdoctoral Fellowship from Griffith University and an Elaine Henry Postdoctoral Fellowship from the NBCF. B. K. kindly acknowledges support from The Scientific and Technological Research Council of Turkey (TUBITAK) for a 2219-International Postdoctoral Research Fellowship (App. No. 1059B192000031) and Griffith University Postdoctoral Fellowship. M. S. kindly acknowledges support from The Scientific and Technological Research Council of Turkey (TUBITAK) for a 2219-International Postdoctoral Research Fellowship (App. No. 1059B192201197). T. P. W. appreciates a Griffith University Postgraduate Research Scholarship (GUPRS) and Griffith University International Postgraduate Research Scholarship (GUIPRS). M. G. A. and D. R. R. acknowledge the award of a Tour de Cure Postgraduate Research Student PhD Scholarship. M. G. A. appreciates a Griffith University Postgraduate Research Scholarship (GUPRS). ICP-MS measurements were performed with the technical assistance of Mr Ryan Stewart and Dr William Bennett, who provided expert support at the School of Environment and Science, Griffith University, Gold Coast, Australia.References
- N. T. Le and D. R. Richardson, Biochim. Biophys. Acta, 2002, 1603, 31–46 CAS.
- D. R. Richardson, D. J. R. Lane, E. M. Becker, M. L. H. Huang, M. Whitnall, Y. S. Rahmanto, A. D. Sheftel and P. Ponka, Proc. Natl. Acad. Sci. U. S. A, 2010, 107, 10775–10782 CrossRef CAS PubMed.
- Z. Xu, Y.-S. Sung and E. Tomat, J. Am. Chem. Soc., 2023, 145, 15197–15206 CrossRef CAS PubMed.
- D. Richardson, Crit. Rev. Oncol. Hematol., 2002, 42, 267–281 CrossRef CAS PubMed.
- J. C. Kwok and D. R. Richardson, Crit. Rev. Oncol. Hematol., 2002, 42, 65–78 Search PubMed.
- E. Morgan, Mol. Aspect. Med., 1981, 4, 1–123 CrossRef.
- B. J. Iacopetta and E. Morgan, J. Biol. Chem., 1983, 258, 9108–9115 CrossRef CAS PubMed.
- R. D. Klausner, G. Ashwell, J. Van Renswoude, J. B. Harford and K. R. Bridges, Proc. Natl. Acad. Sci. U. S. A, 1983, 80, 2263–2266 CrossRef CAS PubMed.
- R. S. Ohgami, D. R. Campagna, E. L. Greer, B. Antiochos, A. McDonald, J. Chen, J. J. Sharp, Y. Fujiwara, J. E. Barker and M. D. Fleming, Nat. Genet., 2005, 37, 1264–1269 Search PubMed.
- H. Gunshin, B. Mackenzie, U. V. Berger, Y. Gunshin, M. F. Romero, W. F. Boron, S. Nussberger, J. L. Gollan and M. A. Hediger, Nature, 1997, 388, 482–488 CrossRef CAS PubMed.
- A. Jacobs, Blood, 1977, 50, 433–439 CrossRef CAS PubMed.
- D. S. Kalinowski and D. R. Richardson, Pharmacol. Rev., 2005, 57, 547–583 Search PubMed.
- I. Rodríguez, C. Acosta, C. Nieves-Escobar, E. Strangmark, O. Claudio-Ares, A. I. Vargas Figueroa, A. M. Soto-Millán, A. M. Orta-Rivera, A. V. Astashkin and A. D. Tinoco, JACS Au, 2024, 4, 4799–4808 Search PubMed.
- C. R. Chitambar, Biochim. Biophys. Acta, 2016, 1863, 2044–2053 CrossRef CAS PubMed.
- C. R. Chitambar, E. J. Massey and P. A. Seligman, J. Clin. Invest., 1983, 72, 1314–1325 Search PubMed.
- C. R. Chitambar, J. Narasimhan, J. Guy, D. S. Sem and W. J. O'Brien, Cancer Res., 1991, 51, 6199–6201 CAS.
- K. Gaur, S. C. Perez Otero, J. A. Benjamin-Rivera, I. Rodriguez, S. A. Loza-Rosas, A. M. Vazquez Salgado, E. A. Akam, L. Hernandez-Matias, R. K. Sharma, N. Alicea, M. Kowaleff, A. V. Washington, A. V. Astashkin, E. Tomat and A. D. Tinoco, JACS Au, 2021, 1, 865–878 CrossRef CAS PubMed.
- D. K. Johnson, T. B. Murphy, N. J. Rose, W. H. Goodwin and L. Pickart, Inorg. Chim. Acta, 1982, 67, 159–165 CrossRef CAS.
- R. M. Mohareb and F. Al-Omran, Steroids, 2012, 77, 1551–1559 CrossRef CAS PubMed.
- S. Aydın, N. Kaushik-Basu, P. Arora, A. Basu, D. Nichols, T. T. Talele, M. Akkurt, İ. Çelik, O. Büyükgüngör and Ş. G. Küçükgüzel, Marmara Pharm. J., 2013, 17, 26–34 CrossRef.
- T. Nasr, S. Bondock and M. Youns, Eur. J. Med. Chem., 2014, 76, 539–548 CrossRef CAS PubMed.
- J. Dilworth, Coord. Chem. Rev., 1976, 21, 29–62 CrossRef CAS.
- K. Hałdys and R. Latajka, MedChemComm, 2019, 10, 378–389 RSC.
- E. Baker, D. Richardson, S. Gross and P. Ponka, Hepatology, 1992, 15, 492–501 CrossRef CAS PubMed.
- R. R. Mohamed and A. Fekry, Int. J. Electrochem. Sci., 2011, 6, 2488–2508 Search PubMed.
- Y. Yu, D. S. Kalinowski, Z. Kovacevic, A. R. Siafakas, P. J. Jansson, C. Stefani, D. B. Lovejoy, P. C. Sharpe, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2009, 52, 5271–5294 Search PubMed.
- R. J. Bergeron, G. Huang, W. R. Weimar, R. E. Smith, J. Wiegand and J. S. McManis, J. Med. Chem., 2003, 46, 16–24 Search PubMed.
- A. C. Chua, H. A. Ingram, K. N. Raymond and E. Baker, Eur. J. Biochem., 2003, 270, 1689–1698 Search PubMed.
- C. Wong, J. Kwok and D. Richardson, Biochim. Biophys. Acta, 2004, 1739, 70–80 Search PubMed.
- P. Ponka, D. Richardson, E. Baker, H. M. Schulman and J. T. Edward, Biochim. Biophys. Acta, 1988, 967, 122–129 Search PubMed.
- D. Richardson, E. Tran and P. Ponka, Blood, 1995, 86, 4295–4306 CrossRef CAS PubMed.
- D. Richardson and K. Milnes, Blood, 1997, 89, 3025–3038 Search PubMed.
- E. M. Becker, D. B. Lovejoy, J. M. Greer, R. Watts and D. R. Richardson, Br. J. Pharmacol., 2003, 138, 819–830 Search PubMed.
- T. B. Chaston, R. N. Watts, J. Yuan and D. R. Richardson, Clin. Cancer Res., 2004, 10, 7365–7374 CrossRef CAS PubMed.
- J. L. Buss, F. M. Torti and S. V. Torti, Curr. Med. Chem., 2003, 10, 1021–1034 CrossRef CAS PubMed.
- J. Turner, C. Koumenis, T. E. Kute, R. P. Planalp, M. W. Brechbiel, D. Beardsley, B. Cody, K. D. Brown, F. M. Torti and S. V. Torti, Blood, 2005, 106, 3191–3199 Search PubMed.
- S. Torti, F. Torti, S. Whitman, M. Brechbiel, G. Park and R. Planalp, Blood, 1998, 92, 1384–1389 Search PubMed.
- C. A. S. Regino, S. V. Torti, R. Ma, G. P. Yap, K. A. Kreisel, F. M. Torti, R. P. Planalp and M. W. Brechbiel, J. Med. Chem., 2005, 48, 7993–7999 CrossRef CAS PubMed.
- B. Ay, O. Şahin, B. S. Demir, Y. Saygideger, J. M. López-de-Luzuriaga, G. Mahmoudi and D. A. Safin, New J. Chem., 2020, 44, 9064–9072 Search PubMed.
- P. V. Bernhardt, J. Mattsson and D. R. Richardson, Inorg. Chem., 2006, 45, 752–760 Search PubMed.
- P. V. Bernhardt, G. J. Wilson, P. C. Sharpe, D. S. Kalinowski and D. R. Richardson, J. Biol. Inorg. Chem., 2008, 13, 107–119 Search PubMed.
- D. B. Lovejoy and D. R. Richardson, Blood, 2002, 100, 666–676 Search PubMed.
- J. Yuan, D. B. Lovejoy and D. R. Richardson, Blood, 2004, 104, 1450–1458 Search PubMed.
- M. Whitnall, J. Howard, P. Ponka and D. R. Richardson, Proc. Natl. Acad. Sci. U. S. A, 2006, 103, 14901–14906 CrossRef CAS PubMed.
- Y. Yu, Y. Suryo Rahmanto and D. R. Richardson, Br. J. Pharmacol., 2012, 165, 148–166 Search PubMed.
- C. Stefani, G. Punnia-Moorthy, D. B. Lovejoy, P. J. Jansson, D. S. Kalinowski, P. C. Sharpe, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2011, 54, 6936–6948 Search PubMed.
- K. Dixon, G. Lui, Z. Kovacevic, D. Zhang, M. Yao, Z. Chen, Q. Dong, S. J. Assinder and D. R. Richardson, Br. J. Cancer, 2013, 108, 409–419 Search PubMed.
- D. R. Richardson, P. C. Sharpe, D. B. Lovejoy, D. Senaratne, D. S. Kalinowski, M. Islam and P. V. Bernhardt, J. Med. Chem., 2006, 49, 6510–6521 Search PubMed.
- D. B. Lovejoy, P. J. Jansson, U. T. Brunk, J. Wong, P. Ponka and D. R. Richardson, Cancer Res., 2011, 71, 5871–5880 Search PubMed.
- D. S. Kalinowski, P. C. Sharpe, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2007, 50, 6212–6225 Search PubMed.
- Z. Kovacevic, S. Chikhani, D. B. Lovejoy and D. R. Richardson, Mol. Pharmacol., 2011, 80, 598–609 Search PubMed.
- D. B. Lovejoy, D. M. Sharp, N. Seebacher, P. Obeidy, T. Prichard, C. Stefani, M. T. Basha, P. C. Sharpe, P. J. Jansson and D. S. Kalinowski, J. Med. Chem., 2012, 55, 7230–7244 CrossRef CAS PubMed.
- Z. Kovacevic, S. V. Menezes, S. Sahni, D. S. Kalinowski, D.-H. Bae, D. J. Lane and D. R. Richardson, J. Biol. Chem., 2016, 291, 1029–1052 CrossRef CAS PubMed.
- Z.-L. Guo, D. R. Richardson, D. S. Kalinowski, Z. Kovacevic, K. C. Tan-Un and G. C.-F. Chan, J. Hematol. Oncol., 2016, 9, 1–16 Search PubMed.
- B. Geleta, K. C. Park, P. J. Jansson, S. Sahni, S. Maleki, Z. Xu, T. Murakami, M. Pajic, M. V. Apte, D. R. Richardson and Z. Kovacevic, FASEB J., 2021, 35, e21347 Search PubMed.
- S. C. Lim, P. J. Jansson, S. J. Assinder, S. Maleki, D. R. Richardson and Z. Kovacevic, FASEB J., 2020, 34, 11511–11528 CrossRef CAS PubMed.
- F. Shehadeh-Tout, H. H. Milioli, S. Roslan, P. J. Jansson, M. Dharmasivam, D. Graham, R. Anderson, T. Wijesinghe, M. G. Azad and D. R. Richardson, Pharmacol. Res., 2023, 193, 106806 Search PubMed.
- T. P. Wijesinghe, M. Dharmasivam, C. C. Dai and D. R. Richardson, Pharmacol. Res., 2021, 173, 105889 CrossRef CAS PubMed.
- D. S. Kalinowski and D. R. Richardson, Chem. Res. Toxicol., 2007, 20, 715–720 Search PubMed.
- C. Stefani, Z. Al-Eisawi, P. J. Jansson, D. S. Kalinowski and D. R. Richardson, J. Inorg. Biochem., 2015, 152, 20–37 Search PubMed.
- A. M. Merlot, D. S. Kalinowski and D. R. Richardson, Antioxid. Redox Signaling, 2013, 18, 973–1006 CrossRef CAS PubMed.
- A. E. Stacy, D. Palanimuthu, P. V. Bernhardt, D. S. Kalinowski, P. J. Jansson and D. R. Richardson, J. Med. Chem., 2016, 59, 4965–4984 Search PubMed.
- E. Falcone, A. G. Ritacca, S. Hager, H. Schueffl, B. Vileno, Y. El Khoury, P. Hellwig, C. R. Kowol, P. Heffeter and E. Sicilia, J. Am. Chem. Soc., 2022, 144, 14758–14768 CrossRef CAS PubMed.
- J. Lewis, R. Laforest, T. Buettner, S. Song, Y. Fujibayashi, J. Connett and M. Welch, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 1206–1211 CrossRef CAS PubMed.
- A. A. Aruffo, T. B. Murphy, D. K. Johnson, N. J. Rose and V. Schomaker, Inorg. Chim. Acta, 1982, 67, L25–L27 Search PubMed.
- L. Pickart, W. H. Goodwin, W. Burgua, T. B. Murphy and D. K. Johnson, Biochem. Pharmacol., 1983, 32, 3868–3871 CrossRef CAS PubMed.
- M. Dharmasivam, K. Busra, T. Wijesinghe, M. Gholam Azad, M. A. Gonzálvez, M. Hussaini, J. Chekmarev, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2023, 66, 1426–1453 Search PubMed.
- M. Dharmasivam, B. Kaya, T. P. Wijesinghe, V. Richardson, J. R. Harmer, M. A. Gonzalvez, W. Lewis, M. G. Azad, P. V. Bernhardt and D. R. Richardson, Chem. Sci., 2024, 15, 974–990 RSC.
- B. Kaya, M. Gholam Azad, M. Suleymanoglu, J. R. Harmer, T. P. Wijesinghe, V. Richardson, X. Zhao, P. V. Bernhardt, M. Dharmasivam and D. R. Richardson, J. Med. Chem., 2024, 67, 12155–12183 CrossRef CAS PubMed.
- B. Kaya, H. Smith, Y. Chen, M. G. Azad, T. M. Russell, V. Richardson, M. Dharmasivam and D. R. Richardson, Inorg. Chem., 2024, 63, 20840–20858 Search PubMed.
- Y. Yu, J. Wong, D. B. Lovejoy, D. S. Kalinowski and D. R. Richardson, Clin. Cancer Res., 2006, 12, 6876–6883 CrossRef CAS PubMed.
- C. Kirilmis, M. Koca, S. Servi and S. Gür, Turk. J. Chem., 2009, 33, 375–384 Search PubMed.
- M. Halli and R. Sumathi, Arab. J. Chem., 2017, 10, S1748–S1759 CrossRef CAS.
- D. S. Kalinowski, P. C. Sharpe, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2008, 51, 331–344 Search PubMed.
- T. P. Wijesinghe, B. Kaya, M. A. Gonzálvez, J. R. Harmer, M. Gholam Azad, P. V. Bernhardt, M. Dharmasivam and D. R. Richardson, J. Med. Chem., 2023, 66, 15453–15476 CrossRef CAS PubMed.
- G. Bandoli, A. Dolmella, F. Tisato, M. Porchia and F. Refosco, Coord. Chem. Rev., 2009, 253, 56–77 CrossRef CAS.
- B. A. Borgias, S. J. Barclay and K. N. Raymond, J. Coord. Chem., 1986, 15, 109–123 CrossRef CAS.
- N. P. Davies, Y. S. Rahmanto, C. R. Chitambar and D. R. Richardson, J. Pharmacol. Exp. Ther., 2006, 317, 153–162 CrossRef CAS PubMed.
- C. R. Chitambar, J. P. Wereley and S. Matsuyama, Mol. Cancer Ther., 2006, 5, 2834–2843 CrossRef CAS PubMed.
- C. R. Chitambar and W. E. Antholine, Antioxid. Redox Signaling, 2013, 18, 956–972 CrossRef CAS PubMed.
- J. Huang, A. Jones, T. D. Waite, Y. Chen, X. Huang, K. M. Rosso, A. Kappler, M. Mansor, P. G. Tratnyek and H. Zhang, Chem. Rev., 2021, 121, 8161–8233 CrossRef CAS PubMed.
- Z. Wu, D. Palanimuthu, N. Braidy, N. H. Salikin, S. Egan, M. L. Huang and D. R. Richardson, Br. J. Pharmacol., 2020, 177, 1967–1987 Search PubMed.
- S. N. Maqbool, S. C. Lim, K. C. Park, R. Hanif, D. R. Richardson, P. J. Jansson and Z. Kovacevic, Br. J. Pharmacol., 2020, 177, 2365–2380 CrossRef CAS PubMed.
- M. Dharmasivam, S. Zhang, X. Zhao, V. Richardson, T. P. Wijesinghe, M. Suleymanoglu, M. Gholam Azad, P. V. Bernhardt, B. Kaya and D. R. Richardson, J. Med. Chem., 2025, 68, 9594–9622 CrossRef CAS PubMed.
- P. J. Jansson, P. C. Sharpe, P. V. Bernhardt and D. R. Richardson, J. Med. Chem., 2010, 53, 5759–5769 CrossRef CAS PubMed.
- D. R. Richardson, Antimicrob. Agents Chemother., 1997, 41, 2061–2063 CrossRef CAS PubMed.
- K. Staff, M. B. Brown, R. P. Chilcott, R. C. Hider, S. A. Jones and X. L. Kong, Toxicol. Lett., 2011, 202, 155–160 CrossRef CAS PubMed.
- C. Hansch and W. J. Dunn III, J. Pharm. Sci., 1972, 61, 1–19 CrossRef CAS PubMed.
- A. V. Rudnev, L. S. Foteeva, C. Kowol, R. Berger, M. A. Jakupec, V. B. Arion, A. R. Timerbaev and B. K. Keppler, J. Inorg. Biochem., 2006, 100, 1819–1826 CrossRef CAS PubMed.
- A. R. Timerbaev, O. O. Vasylenko, L. S. Foteeva, A. V. Rudnev, O. Semenova and B. K. Keppler, J. Sep. Sci., 2007, 30, 399–406 Search PubMed.
- R. P. Warrell Jr, C. J. Coonley, D. J. Straus and C. W. Young, Cancer, 1983, 51, 1982–1987 CrossRef CAS PubMed.
- L. Thelander, A. Gräslund and M. Thelander, Biochem. Biophys. Res. Commun., 1983, 110, 859–865 Search PubMed.
- P. A. Seligman, P. L. Moran, R. B. Schleicher and E. David Crawford, Am. J. Hematol., 1992, 41, 232–240 Search PubMed.
- S. Rubagotti, S. Croci, E. Ferrari, G. Orteca, M. Iori, P. C. Capponi, A. Versari and M. Asti, J. Inorg. Biochem., 2017, 173, 113–119 CrossRef CAS PubMed.
- V. M. Leovac, M. V. Rodić, L. S. Jovanović, M. D. Joksović, T. Stanojković, M. Vujčić, D. Sladić, V. Marković and L. S. Vojinović-Ješić, Eur. J. Inorg. Chem., 2015, 2015, 882–895 Search PubMed.
- L. Santa Maria de la Parra, A. I. B. Romo, J. Rodriguez-Lopez, O. R. Nascimento, G. A. Echeverria, O. E. Piro and I. E. Leon, Inorg. Chem., 2024, 63, 4925–4938 CrossRef CAS PubMed.
- L. Mazur, B. Modzelewska-Banachiewicz, R. Paprocka, M. Zimecki, U. E. Wawrzyniak, J. Kutkowska and G. Ziółkowska, J. Inorg. Biochem., 2012, 114, 55–64 CrossRef CAS PubMed.
- L. Kohler, A. M. Potocny, J. Niklas, M. Zeller, O. G. Poluektov and K. L. Mulfort, Catalysts, 2021, 11, 75 CrossRef CAS.
- A. Popović-Bijelić, C. R. Kowol, M. E. Lind, J. Luo, F. Himo, É. A. Enyedy, V. B. Arion and A. Gräslund, J. Inorg. Biochem., 2011, 105, 1422–1431 CrossRef PubMed.
- Y.-T. Wang, Y. Fang, M. Zhao, M.-X. Li, Y.-M. Ji and Q.-X. Han, MedChemComm, 2017, 8, 2125–2132 RSC.
- T. G. Spiro, S. E. Allerton, J. Renner, A. Terzis, R. Bils and P. Saltman, J. Am. Chem. Soc., 1966, 88, 2721–2726 Search PubMed.
- D. Richardson and E. Baker, Biochim. Biophys. Acta, 1991, 1093, 20–28 CrossRef CAS PubMed.
- Y. Goto, M. Paterson and I. Listowsky, J. Biol. Chem., 1983, 258, 5248–5255 Search PubMed.
- D. R. Richardson and P. Ponka, Biochim. Biophys. Acta, 1997, 1331, 1–40 Search PubMed.
- M. Grootveld, J. Bell and B. Halliwell, J. Biol. Chem., 1989, 264, 4417–4422 CrossRef CAS PubMed.
- D. J. Lane, S. Chikhani, V. Richardson and D. R. Richardson, Biochim. Biophys. Acta, 2013, 1833, 1527–1541 Search PubMed.
- D.-H. Bae, M. Gholam Azad, D. S. Kalinowski, D. J. Lane, P. J. Jansson and D. R. Richardson, Antioxid. Redox Signaling, 2020, 33, 816–838 CrossRef CAS PubMed.
- S. O. Pehkonen, Y. Erel and M. R. Hoffmann, Environ. Sci. Technol., 1992, 26, 1731–1736 Search PubMed.
- T. W. Price, J. Gallo, V. Kubíček, Z. Böhmová, T. J. Prior, J. Greenman, P. Hermann and G. J. Stasiuk, Dalton Trans., 2017, 46, 16973–16982 Search PubMed.
- A. Schmidtke, T. Läppchen, C. Weinmann, L. Bier-Schorr, M. Keller, Y. Kiefer, J. P. Holland and M. D. Bartholomä, Inorg. Chem., 2017, 56, 9097–9110 CrossRef CAS PubMed.
- I. Velikyan, G. J. Beyer and B. Långström, Bioconjugate Chem., 2004, 15, 554–560 Search PubMed.
- M. Kamyabi, Z. Asgari, H. H. Monfared and A. Morsali, J. Electroanal. Chem., 2009, 632, 170–176 CrossRef CAS.
- P. V. Bernhardt, P. Chin, P. C. Sharpe and D. R. Richardson, Dalton Trans., 2007, 3232–3244 Search PubMed.
- D. Richardson, P. Ponka and E. Baker, Cancer Res., 1994, 54, 685–689 Search PubMed.
- Z. Deng, M. Gholam Azad, D. Richardson and M. G. Azad, Pharmacol. Res., 2025, 19, 107967 Search PubMed.
- B. Kaya, D. Laurencia, M. F. Ayoub, M. G. Azad, M. Dharmasivam and D. R. Richardson, Pharmacol. Rev., 2025, 77, 100087 Search PubMed.
- C. R. Chitambar, D. P. Purpi, J. Woodliff, M. Yang and J. P. Wereley, J. Pharmacol. Exp. Ther., 2007, 322, 1228–1236 Search PubMed.
- S.-r. Choi, M. A. Hassan, B. E. Britigan and P. Narayanasamy, Curr. Issues Mol. Biol., 2024, 46, 9149–9161 CrossRef CAS PubMed.
- Y. Guo, W. Li, H. Li and W. Xia, ACS Infect. Dis., 2019, 5, 1693–1697 Search PubMed.
- P. Verma, Z. Varga, J. E. Klein, C. J. Cramer, L. Que and D. G. Truhlar, Phys. Chem. Chem. Phys., 2017, 19, 13049–13069 Search PubMed.
- C. T. Gutman and T. C. Brunold, Inorg. Chem., 2012, 51, 12729–12737 CrossRef CAS PubMed.
- (a) CCDC 1982904: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcnr; (b) CCDC 1982905: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcps; (c) CCDC 1982906: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcqt; (d) CCDC 1982907: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcrv; (e) CCDC 1982908: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcsw; (f) CCDC 1982909: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kctx; (g) CCDC 1982911: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc24kcwz; (h) CCDC 2424290: Experimental Crystal Structure Determination, 2025, DOI:10.5517/ccdc.csd.cc2mcnxl.
Footnote |
| † Co-first authors contributing equally to this manuscript. |
| This journal is © The Royal Society of Chemistry 2026 |