Open Access Article
Open Access ArticleNovel chemistry and structural perspectives in vitamin B12-dependent radical SAM enzymes
Olivier
Berteau
 *a,
Magda
Teixeira Nunes
*a,
Magda
Teixeira Nunes
 b,
Nicole
Taraglio
b,
Nicole
Taraglio
 a,
Pierre
Legrand
a,
Pierre
Legrand
 b and
Alhosna
Benjdia
b and
Alhosna
Benjdia
 *a
*a
aUniversité Paris-Saclay, INRAE, AgroParisTech, Micalis Institute, ChemSyBio, 78350, Jouy-en-Josas, France. E-mail: Olivier.berteau@inrae.fr
bSynchrotron SOLEIL, L'Orme des Merisiers, Saint Aubin, France
First published on 24th February 2026
Abstract
Vitamin B12 is one of the most complex vitamins and an essential cofactor for humans, but also for bacteria, which have evolved different strategies for its biosynthesis and acquisition. For decades, vitamin B12 has been mainly assumed to be a source of nucleophilic methyl groups and radical initiators, in the forms of methyl-cobalamin and adenosyl-cobalamin. Recently, the importance and the chemical versatility of vitamin B12 have been reconsidered in light of the emergence of the superfamily of radical SAM enzymes. In this review, we provide an overview of vitamin B12, its role, and its diversity within the human microbiota, and in the emerging family of B12-dependent radical SAM enzymes. We focus on the diversity of B12-dependent radical SAM enzymes from a mechanistic and structural perspective, along with their potential evolutionary links to other B12-dependent enzymes.
1. Introduction
Vitamin B12 (also called cobalamin) is an essential vitamin for humans (Fig. 1). It is also a central metabolite in bacteria and its importance has been recently highlighted in the so-called superfamily of radical SAM enzymes.1–4 This superfamily of enzymes catalyzes unrelated and often chemically challenging transformations, including post-translational modifications,5–10 complex rearrangements,11 and DNA modification,12–14 along with cofactor15,16 and metallic-center17,18 biosynthesis. In the last decade, the importance of these enzymes has been demonstrated in bacterial and human metabolisms,19,20 where they support key functions spanning from the biosynthesis of secondary metabolites to antiviral defense. Originally, the importance of radical SAM enzymes was recognized in a pioneering bioinformatics study, defining them as metalloenzymes containing one unusual [Fe4S4] cluster and using S-adenosyl-L-methionine (SAM) as an essential cofactor to initiate radical catalysis.21 This study also predicted that several of these enzymes would utilize vitamin B12 as a cofactor. However, the in vitro demonstration that some radical SAM enzymes require cobalamin has been reported only relatively recently,22,23 leading to the emergence of a novel enzyme family called the B12-dependent radical SAM enzymes.Being one of the largest classes of radical SAM enzymes,23–33 these biocatalysts delineate a superfamily on their own, comprising more than 220![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 members. Early studies have shown that these enzymes are involved in the alkylation of unreactive carbon atoms,23,25,30,31P-methylation22 and ring contraction.29 More recently, these enzymes have been proposed to catalyze unrelated transformations, such as ring expansion34 and thioether bond formation.35 In all of these reactions, vitamin B12 plays a central but sometimes ill-understood role, underscoring the importance of this vitamin in bacterial metabolism and within the human microbiome.1,36,37 Indeed, it is estimated that the vast majority of bacterial species that constitute this complex ecosystem38,39 require vitamin B12 for their metabolism.36,37 However, only a fraction of them have the biosynthetic machinery required to perform its de novo synthesis, and fierce competition exists to acquire this metabolite.40
000 members. Early studies have shown that these enzymes are involved in the alkylation of unreactive carbon atoms,23,25,30,31P-methylation22 and ring contraction.29 More recently, these enzymes have been proposed to catalyze unrelated transformations, such as ring expansion34 and thioether bond formation.35 In all of these reactions, vitamin B12 plays a central but sometimes ill-understood role, underscoring the importance of this vitamin in bacterial metabolism and within the human microbiome.1,36,37 Indeed, it is estimated that the vast majority of bacterial species that constitute this complex ecosystem38,39 require vitamin B12 for their metabolism.36,37 However, only a fraction of them have the biosynthetic machinery required to perform its de novo synthesis, and fierce competition exists to acquire this metabolite.40
This review focuses on the latest developments regarding vitamin B12 biosynthesis, and explores the structural and functional diversity of B12-dependent radical SAM enzymes, and their catalytic and structural links with other B12-dependent enzymes.
2. Vitamin B12
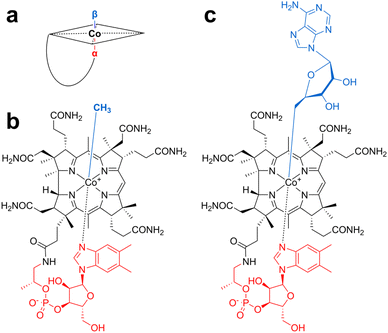
Vitamin B12 (cobalamin) is one of the eight known B vitamins and was identified in 1948 as a red pigment containing cobalt.41,42 Its structure, determined a few years later,43 revealed that it is built based on a substituted tetrapyrrole (corrin) ring containing a central cobalt atom. This ring is generally connected by a three-carbon moiety and a 3′-phosphoribose linked to an unusual nucleotide, 5,6-dimethylbenzimidazole (DMB), which coordinates the cobalt atom in the lower axial position (α face) (Fig. 1). All of these structural features give vitamin B12 unique reactivity and functions. To add to this complexity, vitamin B12 usually coordinates at its upper axial position (β face) a methyl or an adenosyl group in methyl- and 5′-deoxyadenosylcobalamin (MeCbl and AdoCbl), respectively. In humans, these two forms are used by two distinct enzymes: methionine synthase, which catalyzes the conversion of homocysteine to methionine, and the mitochondrial enzyme methylmalonyl CoA mutase, which converts methylmalonyl CoA to succinyl CoA. A deficiency in vitamin B12 leads to severe pathologies, which explains why a sophisticated trafficking pathway has been evolved in humans.44 Interestingly, not only do these two enzymes use two distinct forms of vitamin B12, but they also exploit its two reactivities.45 Indeed, MeCbl is used in SN2-type reactions to transfer a methyl group as a carbocation to a nucleophilic acceptor, while AdoCbl is used exclusively in radical reactions by generating the 5′-deoxyadenosyl radical, following Co–carbon bond homolysis. In bacteria, recent studies exploring the human microbiota have drawn a more complex picture of the role and requirement of vitamin B12.2.1. Vitamin B12 and cobamides in the microbiome
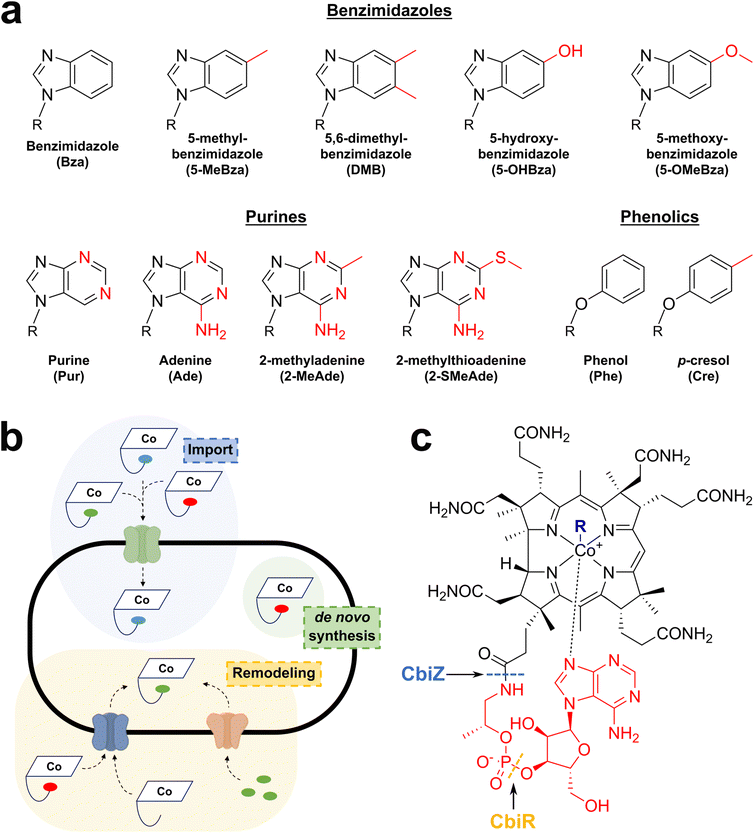
Although vitamin B12 is the most studied, it is part of a larger family of cofactors called cobamides. Indeed, the investigation of several bacterial systems has revealed that, besides the upper axial ligand, the nature of the lower axial ligand is also variable, leading to a wide diversity of cobamide cofactors (Fig. 2a). Surprisingly, in several environments, including the human gut, vitamin B12 has been reported to be a minority among the cobamides produced,36,37,46 in spite of its critical requirement for the host and the microbiome. Indeed, it has been recently recognized that vitamin B12 profoundly impacts the gut microbiome by promoting a more diverse ecosystem with potential health benefits to the host.47,48 A recent systematic review of the literature supports that vitamin B12 is associated with changes in gut microbiota by increasing alpha-diversity and shifting the microbiome composition.48 However, depending on the cobalamin form and uptake, the effects of vitamin B12 appear to differ and to be indirectly mediated by propionate production, which has a positive role in gut integrity and host health.Why do bacteria produce such a diversity of cobamides? One hypothesis is that this structural diversity is connected with vitamin B12 acquisition. Indeed, most bacterial species cannot de novo synthesize cobamides, but instead possess importing systems (Fig. 2b). Altering the cobamide structure might thus provide a fitness advantage by preventing their import by competing species. In support of this evolutionary hypothesis, it has been shown that while Bacteroides thetaiotaomicron can use only benzimidazolyl and purinyl-cobamides,36 other bacteria, such as Sporomusa ovata, favor phenolyl-cobamides.49 Remarkably, to circumvent this issue, many bacteria, including Dehalococcoides mccartyi,50Rhodobacter sphaeroides,51Vibrio cholerae,52 and the gut bacteria Akkermansia muciniphila53 are capable of “cobamide remodeling”. This remodeling is performed by unrelated biosynthetic pathways such as CobS in V. cholerae, the amidohydrolase CbiZ in R. sphaeroides, and the phosphodiesterase CbiR in A. muciniphila53 (Fig. 2c). After the removal of the lower ligand, the imported cobamides can be further processed by the salvaging pathways in order to produce relevant cobamides that support bacterial growth. Although these pathways are likely to provide fitness advantages by repurposing cobamides, the structural and functional diversity of these remodeling enzymes appears to be the result of convergent evolution, underscoring the physiological relevance and importance of vitamin B12.
Another hypothesis regarding the diversity of cobamides is connected to the multiplicity of B12-dependent enzymes and the reactions they catalyze. It is plausible that some cobamides are specifically tailored to fulfill dedicated catalytic functions. To date, this has not been experimentally validated, and further research will be necessary to gain a comprehensive understanding of the structural and functional diversity of cobamides.37,54
2.2. Novel insights into vitamin B12 biosynthesis
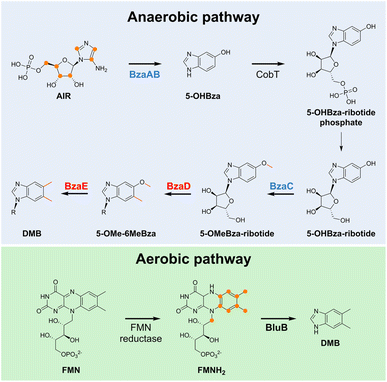
Among its unique features, cobalamin is the only vitamin synthesized exclusively by bacteria.55,56 It is produced via two related though genetically distinct routes called the aerobic and anaerobic pathways.57 The distribution of these pathways is uneven, with anaerobic pathways being the most widespread. Although both pathways require the bismethylation of uroporphyrinogen III to precorrin-2 as the first step, they rapidly diverge. The biosynthesis of vitamin B12 has been described in excellent reviews,44,56,57 and is almost fully elucidated. However, the final steps of its biosynthesis, which involve the synthesis and attachment of the lower nucleotide loop, have not been entirely clarified.58,59In the aerobic pathway, the key role of the oxygen-dependent enzyme BluB, which converts FMN to DMB, has been demonstrated60–62 (Fig. 3). For the anaerobic biosynthesis of DMB, five genes (BzaABCDE) are required, and 5-aminoimidazole ribotide (AIR), an intermediate in the purine biosynthesis pathway, is the central precursor.59 Another difference is that DMB is not synthesized as a free group but likely as a ribotide derivative in the anaerobic pathway. Indeed, it was recently shown that BzaC, the first methyl transferase involved in the conversion of AIR to DMB (Fig. 3), prefers 5-OHBza-ribotide over 5-OHBza as a substrate. Interestingly, the activity of BzaC on this asymmetric precursor provides a structural rationale for the observed regioselectivity of this conversion.59
The conversion of 5-OMeBza into DMB remains an open question. The final steps are predicted to involve two B12-dependent radical SAM enzymes: BzaD and BzaE. BzaD likely catalyzes the methylation of a Csp2-atom to form 5-OMe-6-MeBza, reminiscent of the reaction catalyzed by TsrM23,25 (see below). The BzaE-catalyzed conversion of 5-OMe-6-MeBza to DMB is enigmatic and will require further studies to be clarified. The anaerobic biosynthesis of DMB involves not only unique chemistry but also cobalamin-dependent enzymes, which is a fascinating conundrum.
3. Catalytic functions of vitamin B12
Until recently, vitamin B12-dependent enzymes were classified into three broad classes: the isomerases/mutases, the methyltransferases, and the more recently characterized reductive dehalogenases,63,64 as described below.3.1. Radical B12-dependent enzymes: isomerases, mutases, and reductases
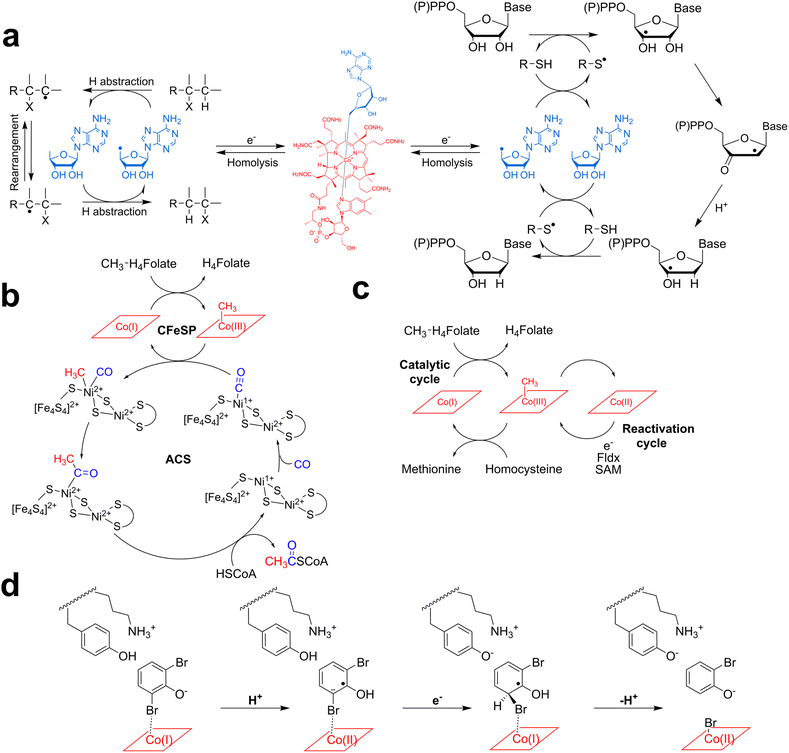
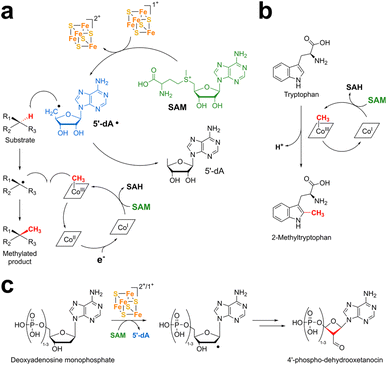
Isomerases/mutases were considered until recently as the largest class of vitamin B12-dependent enzymes.63 They include diol dehydratases,65 ethanolamine ammonia-lyase, lysine 5,6-aminomutase,66 methylmalonyl-CoA mutase, methyleneglutarate mutases,67 glutamate mutase68–71 and ribonucleoside-triphosphate reductase.72 These enzymes use AdoCbl to catalyze radical reactions, and their mechanism can be summarized as follows: upon substrate binding, AdoCbl is homolytically cleaved, generating the 5′-deoxyadenosyl (5′-dA˙) radical and cob(II)alamin. The highly reactive 5′-dA radical abstracts a substrate H-atom and generates a substrate radical that undergoes either a 1,2 rearrangement (isomerases) or carbon skeleton moiety intramolecular migration (mutases). Re-abstraction of the product radical H-atom leads to reaction termination and AdoCbl recombination (Fig. 4 and 5a). Among these enzymes, class II ribonucleotide reductases are outliers, as the 5′-dA radical abstracts a hydrogen atom from a cysteine residue, generating a thiyl radical within the enzyme's active site. This radical species, in turn, abstracts a substrate H-atom and serves as a radical reservoir.12,73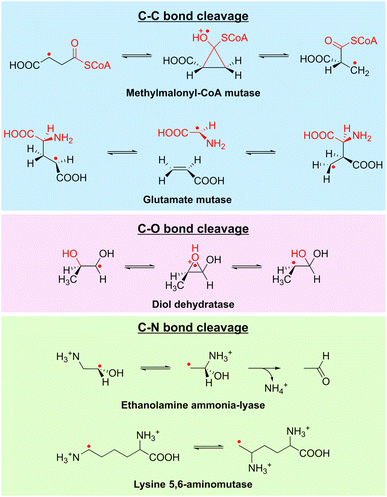 | ||
| Fig. 4 Representative reactions of the isomerase/mutase family. Isomerases and mutases have been shown to be involved in C–C, C–O and C–N bond cleavage and formation. | ||
3.2. Methyltransferases
B12-dependent methyltransferases were identified at the same period as isomerases, more than six decades ago.74 The most widely investigated systems are certainly methionine synthase, which catalyzes the conversion of homocysteine into methionine63,64,67,75–80 and the corrinoid iron-sulfur protein (CFeSP), involved in methyl transfer in the Wood-Ljungdahl carbon fixation pathway.63,81–84 In this pathway, the methyl group is transferred to different metallic sites, with one of two Ni ions85 present in the acetyl-CoA synthase active site serving as its penultimate acceptor, before formation of the final product (Fig. 5b).Both enzymes use the same general mechanism, with methyl being transferred from MeCbl(III) to an acceptor and the formation of the super-nucleophile Cob(I), which is methylated back by methylenetetrahydrofolate. Interestingly, extensive characterization by the Banerjee and Ragsdale laboratories has demonstrated that both enzymes can be inactivated in a Cob(II) state, which needs further reduction and a SAM molecule, as a more efficient methyl donor (Fig. 5c)63,64,76,77,81,86 for enzyme reactivation and subsequent turnovers. This reactivation cycle appears as an impressive trick to prevent the irreversible inactivation of the enzyme and the loss not only of the protein but also of vitamin B12, which is a metabolically costly cofactor. Indeed, it was estimated that oxidative inactivation occurs once every 2000 turnovers, which is significant.
3.3 Reductive dehalogenases
The third group of vitamin B12-dependent enzymes is the reductive dehalogenases.87,88 This group has been identified more recently and proved to usually contain, in addition to a variety of corrinoid cofactors (e.g., norpseudo-B12), iron-sulfur clusters like in the CFeSP protein.89–91 These enzymes play a crucial role in organohalide respiration and have the ability to reductively cleave carbon-halogen bonds found in a variety of environmentally hazardous chemicals. They are, therefore, fascinating biocatalysts for bioremediation. The role of the corrinoid cofactor has been elucidated only recently, thanks to advanced EPR and structural analyses.87,88,92 As a consensus, in these enzymes, the active form of the corrinoid cofactor is Cob(I).For the reductive dehalogenase from Nitratireductor pacificus (NpRdhA), a direct interaction was observed between the cobalt atom of cobalamin and the substrate (Fig. 5d). A mechanism has been proposed with cobalamin directly catalyzing halogen abstraction through the formation of a halogen–cobalt bond, which could be followed either by heterolytic or homolytic bond cleavage, with the transient formation of a halogen-Cob(III)alamin or an aryl radical intermediate. In contrast, a direct interaction with the substrate was not evidenced for the tetrachloroethene reductive dehalogenase from Sulfurospirillum multivorans (PceA).88 Thus, long-range electron transfer in the enzyme's active site may also play a crucial role in catalysis. For this enzyme, a homolytic cleavage of the C–Cl bond was favored because of the strong stability of the C–Cl bond (∼80–100 kcal mol−1) and the detection of Co(II) and radical intermediates in related systems.93,94 Further studies will be required to determine whether there is a common mechanism for dehalogenases or if, as is commonly observed for B12-dependent enzymes, they have evolved distinct reactivity. Of note, the function of the iron-sulfur clusters and the process leading to the reduction of Cob(II) into the catalytically competent Cob(I) remain to be fully clarified.
As illustrated by these three classes of enzymes, vitamin B12 is a powerful cofactor that can support different types of chemistry (i.e., nucleophilic vs. radical) and distinct reactions, from alkylation reactions and isomerization to the cleavage of halogen–carbon bonds. It is, therefore, not surprising that novel roles have emerged for this unique cofactor in the largest and most versatile superfamily of enzymes, the radical SAM enzymes.22,23
4. B12-dependent radical SAM enzymes
The current release of the radical SAM enzyme superfamily includes more than 720![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 proteins.1–3,95–97 These enzymes, recognized 20 years ago as an emerging superfamily,21 form the most diverse and largest superfamily of enzymes. For instance, they catalyze an impressive diversity of radical-based reactions in a myriad of metabolic pathways,3,98,99 including DNA synthesis73 and repair,13,14,100,101 and protein and nucleic acid modifications7,8,102–109 as well as cofactor and vitamin biosynthesis.15,16,110–115 These enzymes also play a central role in the growing family of ribosomally-synthesized and post-translationally modified peptides (RiPPs), which are major metabolites from the human microbiota and represent attractive alternatives to traditional antibiotics.23–25,30,95,116–120 However, as more than half of the different classes have no experimentally validated functions, radical SAM enzymes are still a frontier in biochemistry.
000 proteins.1–3,95–97 These enzymes, recognized 20 years ago as an emerging superfamily,21 form the most diverse and largest superfamily of enzymes. For instance, they catalyze an impressive diversity of radical-based reactions in a myriad of metabolic pathways,3,98,99 including DNA synthesis73 and repair,13,14,100,101 and protein and nucleic acid modifications7,8,102–109 as well as cofactor and vitamin biosynthesis.15,16,110–115 These enzymes also play a central role in the growing family of ribosomally-synthesized and post-translationally modified peptides (RiPPs), which are major metabolites from the human microbiota and represent attractive alternatives to traditional antibiotics.23–25,30,95,116–120 However, as more than half of the different classes have no experimentally validated functions, radical SAM enzymes are still a frontier in biochemistry.
As a general principle, these metalloenzymes are characterized by a [Fe4S4]2+/1+ cluster coordinating the SAM cofactor and employ a radical-based mechanism, with some notable exceptions such as TsrM.23,25,27,121,122 To initiate their reaction, one electron is transferred to the catalytic [Fe4S4]2+/1+ cluster, inducing, like in B12-dependent isomerases and mutases, the homolytic cleavage of a SAM molecule and the generation of the transient 5′-dA radical.123–125 This highly reactive species usually activates a C–H bond by abstracting an H-atom from the substrate, resulting in the formation of a carbon-centered radical intermediate.3,126,127 However, depending on the nature of the substrate, some variations have been evidenced, including the radical addition of 5′-dA to the substrate,128,129 and recently, the generation of alternative radical species.130 How these reactions are controlled by the protein matrix and how they proceed are still matters of debate; however, substrate reactivity likely plays a major role.1,127
Despite more than 30 years of investigation and the early prediction that vitamin B12 could be an essential cofactor for several radical SAM enzymes, this hypothesis has only recently been experimentally validated. The difficulties in expressing and investigating B12-dependent radical SAM enzymes that utilize an [Fe4S4] cluster and cobalamin, which are oxygen- and light-sensitive cofactors, combined with the fact that some of these enzymes are only able to accommodate the B12 cofactor during their translation,25,26,32,33,131 likely explain why our understanding of this superfamily of enzymes remains limited.
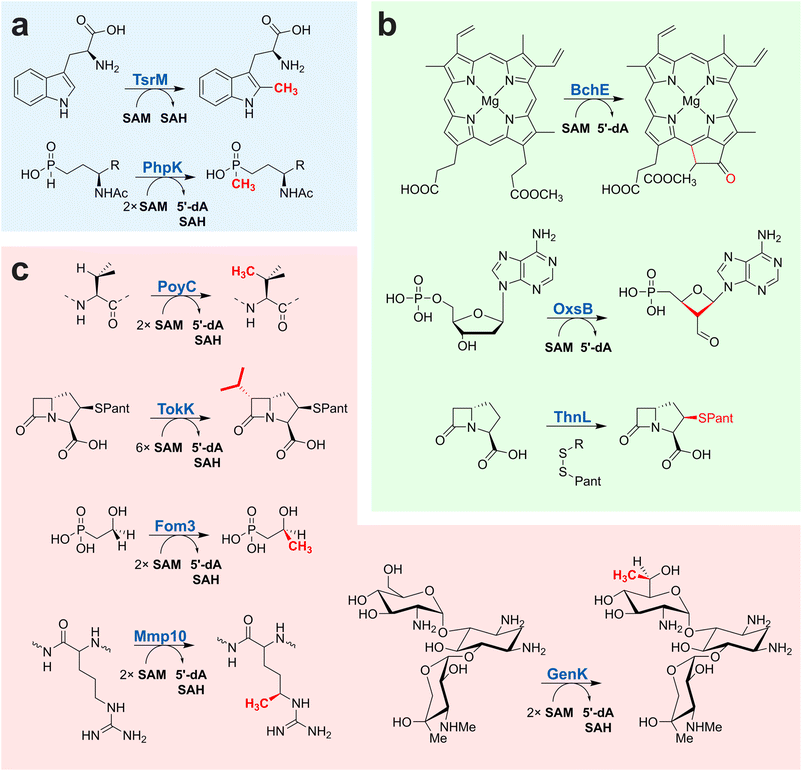
Over the last decade, B12-dependent radical SAM enzymes have been shown to catalyze a diverse range of reactions. Notably, this family includes a large group of methyltransferases that are unique biocatalysts capable of forming a carbon–carbon bond between a methyl group and an unactivated carbon atom23,25,28,31,33,131–136 (Fig. 6). By far, these C-methyltransferases are the most studied and represented group within this superfamily, alkylating usually Csp3-atoms on a broad diversity of molecules from carbohydrates and antibiotics (Fosfomycin,134,137 Gentamicin133) to various metabolites and proteins (Fig. 6). Intriguingly, the first two enzymes investigated in this group were PhpK22 and TsrM,23,25,32 which are a P- and a Csp2-methyltransferase, respectively. In sharp contrast to the methylation of inert Csp3-atoms, the reaction catalyzed by these two enzymes can proceed either by radical or nucleophilic chemistry (Fig. 6 and 7).
Alongside these methyltransferases, B12-dependent radical SAM enzymes that catalyze unrelated transformations, such as ring formation, ring contraction, and thioether bond formation, have been sporadically identified in the bacteriochlorophyll,138 oxetanocin A29 and carbapenem35 biosynthetic pathways, respectively.
In these distinct reactions, the role and nature of the cobalamin cofactor are unclear. For instance, in OxsB, it has been proposed that cobalamin could serve as an electron sink, while in BchE, since two H-atom abstractions are required, AdoCbl might be involved in catalysis.
4.1. Mechanisms of B12-dependent radical SAM enzymes
Known mechanisms of the B12-dependent radical SAM enzymes include both radical and nucleophilic chemistry (Fig. 6). For B12-dependent radical SAM enzymes catalyzing the methylation of a Csp3-atom,23,25,28,31,33,131–136 the consensus mechanism involves a dual use of the SAM cofactor as both a methyl donor and a radical initiator, with the concomitant formation of SAH (S-adenosyl-homocysteine) and 5′-dA, respectively (Fig. 6a). In most cases, methylation occurs with inversion of the stereo-chemistry of the methylated carbon atom (i.e., a double double-inversion process);134,137,139–144 however, some exceptions where the stereochemistry of the alkylated carbon atom is retained are known.139Intriguingly, for the Csp2-methyltransferase TsrM, the currently accepted mechanism implies a nucleophilic displacement without a clear role for the conserved [Fe4S4] cluster, although the latter is functionally critical23,25,121,122 (Fig. 6b). Finally, the other enzyme whose mechanism has been investigated in detail is OxsB, catalyzing oxidative ring contraction. While it has been demonstrated that OxsB radically cleaves SAM and abstracts a substrate H-atom, the function of its cobalamin cofactor is unclear; however, it has been exploited to alter the fate of the reaction, leading to the formation of methylated products.29,145
4.2. A base-off vitamin B12 in the active site of radical SAM enzymes
Thanks to the spectroscopic and structural investigations of several unrelated enzymatic systems in the last decade, significant advances have been made in our understanding of the role of cobalamin in B12-dependent radical SAM enzymes. The first spectroscopic study was published in 2016 and focused on the characterization of TsrM.27 This study not only showed that the [Fe4S4] cluster does not directly interact with SAM, but also that the cobalamin cofactor was likely in a base-off configuration, two conclusions supported by a subsequent structural analysis.121 Thus, while in solution vitamin B12 essentially exists in a hexacoordinated base-on form with the DMB moiety binding the α-face of the corrin-bound cobalt atom (Fig. 2), it is decoordinated in TsrM. Intriguingly, the structural analysis of OxsB146 and the recent structures of three methyltransferases, TokK,147 Mmp10,33 and QCMT,148 revealed a similar base-off configuration of the cobalamin cofactor supporting a general trend within this superfamily (see Section 4.3).The displacement of the DMB moiety has a profound impact on modulating not only the strength, but also the reactivity of the Co–carbon bond localized in trans. It also increases the molecular surface by about 300 Å−2. This binding mode is far from being exclusive to B12-dependent radical SAM enzymes. Indeed, from methionine synthase80 to human and bacterial mutases,149–151 many B12-dependent enzymes have been shown to have a base-off cofactor.152 In these enzymes, the cobalt-atom is coordinated by the protein itself, generally through a histidine residue in the consensus sequence DxHxxG, resulting in a “base-off/His-on” cobalamin. With this conformation, the hexacoordination of the cobalt atom is preserved. Other enzymes, such as the pyridoxal-phosphate (PLP)-dependent enzymes lysine-5,6-amino-mutase153 and ornithine-4,5-aminomutase,154 also share this binding mode. The CFeSP82,83 and the reductive dehalogenases,87,88,92 despite having a base-off cobalamin, do not have a protein residue in direct interaction with the α-face of the corrin-bound Co-atom. Thus, the Co-atom exhibits a penta-coordination with a methyl group or the substrate present on the β-face, a configuration similar to the one encountered in B12-dependent radical SAM enzymes. In contrast, many B12-dependent enzymes, such as diol dehydratases,155,156 ethanolamine ammonia lyase,157 and B12-dependent ribonucleotide reductase,158 possess a base-on cobalamin in their active site.
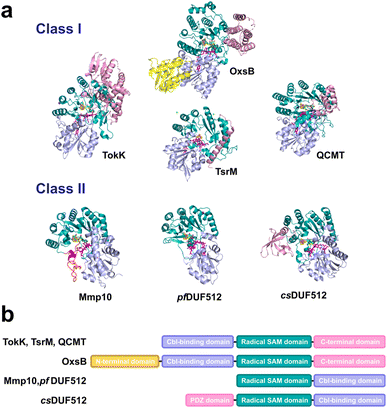
4.3. Structural diversity of B12-dependent radical SAM enzymes
Unraveling the structure of B12-dependent enzymes has proven challenging, with the first structural information published in 2014,159 and the first holo-structure released in 2017.29,160 Recently, the structural characterization of seven B12-dependent radical SAM enzymes, including TsrM,121 TokK,147 Mmp10,33 QCMT,148 and two proteins of unknown function,161 has laid the foundation for comprehending the structural and mechanistic diversity of these enzymes (Fig. 8). Structurally, these enzymes are characterized by a typical radical SAM domain organized in a fold related to the TIM barrel. This domain houses a catalytic [Fe4S4] cluster coordinated by the canonical radical SAM motif CXXXCXXC. In addition to this TIM barrel domain, these enzymes possess a cobalamin-binding domain, which anchors the cobalamin cofactor. This second domain adopts a general Rossmann fold configuration, with some variations regarding the number of beta (β) strands and alpha (α) helices, and their relative orientation to the radical SAM domain, defining two distinct classes of B12-dependent radical SAM enzymes (Fig. 8).The first class encompasses OxsB, TsrM, TokK, and QCMT. Despite catalyzing three distinct reactions (i.e., ring contraction, nucleophilic and radical methylation, respectively), these enzymes share similar structural features with a Rossmann fold constituted of five β-strands and five connecting α helices. In addition, the respective orientation of the cobalamin cofactor and the [Fe4S4] cluster are similar, with a distance of approximately 11 Å between both metallic centers, ideally suited to sandwich the substrate. The only notable difference in these enzymes' active-sites is the tetra-coordination of the [Fe4S4] cluster in TsrM by a glutamate residue, presumed to preclude radical chemistry.
The second class of B12-dependent radical SAM enzymes was initially revealed by the investigation of Mmp10,33 which catalyzes the methylation of Arg285 in the active site of methyl coenzyme M reductase (MCR), the central enzyme in methanogenesis. For this enzyme, the B12-domain is located in the C-terminal position, and the radical SAM domain is in the N-terminal position, in contrast to the majority of B12-dependent radical SAM enzymes (Fig. 8). Moreover, despite catalyzing a radical reaction, Mmp10 possesses a tetra-coordinated cluster like TsrM. In this enzyme, the Rossmann fold comprises 4 β-strands and 4 α-helices. Interestingly, a recent study has shown that unrelated enzymes possessing a DUF512 domain161 share the same overall structure with a notably similar cobalamin binding domain. In these enzymes, the average relative distance between both metallic centers is 12 Å and, in the absence of the substrate, the radical SAM [Fe4S4] cluster is fully coordinated. In the case of Mmp10, it has been shown that upon substrate binding, a unique active-site conformational change allows SAM to bind to the [Fe4S4] cluster, supporting the initiation of the radical reaction. It remains to be shown if DUF512 proteins exhibit a similar behavior.
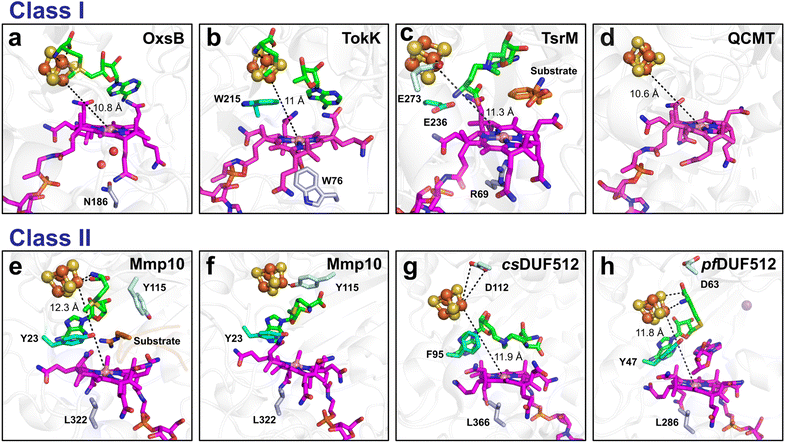
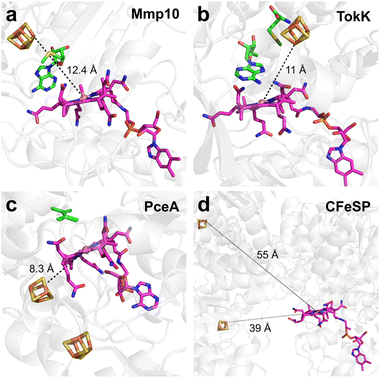
Although there is no direct interaction between the protein and the α-face of the cobalamin cofactor, B12-dependent radical SAM enzymes typically feature a residue in the lower axial position of cobalamin that plays a role in enzyme catalysis. Proteins that catalyze radical-dependent methylations provide a hydrophobic environment to the lower face using distinct residues (i.e., Trp76, Leu322, Leu366, and Leu286 in TokK, Mmp10, csDUF512, and pfDUF512, respectively) (Fig. 9), while enzymes that do not catalyze radical methylation have a more hydrophilic environment (i.e., Arg69 and a water molecule & Asn186 in TsrM and OxsB, respectively). These residues likely prevent the hexacoordination of the cobalamin cofactor, weakening the Co–C bond and increasing its reactivity.
Another intriguing feature is the presence, between the cobalamin cofactor and the [Fe4S4] cluster, of a generally hydrophobic residue (i.e. Trp215, Tyr23, F95 and Y47 in TokK, Mmp10, csDUF512 and pfDUF512 respectively), with the exception of TsrM which possesses a polar residue (Glu236) (Fig. 9). While the function of this residue is unclear, it likely plays an important role in tuning the cobalamin redox potential. Mutation of this or the lower axial residue has been shown to strongly impair enzyme activity,121,147 emphasizing the importance of the cobalamin's surrounding environment. Of note, OxsB, which does not use cobalamin for methyl transfer but most likely as an electron sink, does not possess this stacked residue, and its lower face is solvent accessible. Further studies will be required to ascertain the exact role of these residues, but they likely contribute to finely tuning cobalamin reactivity.
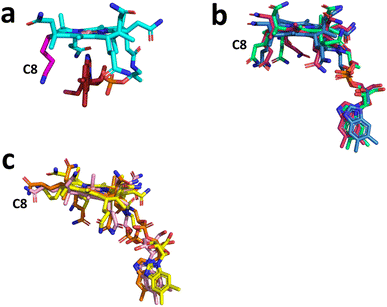
In all the structures solved with a bound cobalamin, the cofactor is in the base-off configuration, with the DMB moiety displaced from the Co-atom (Fig. 10). Intriguingly, among these seven structures, we can define two groups that differ by the orientation of the C8 side chain. While QCMT, TsrM, and OxsB have a configuration similar to base-on cobalamin (Fig. 10a and b), Mmp10, TokK, and DUF512 proteins adopt a more planar configuration of the tetrapyrrole ring (Fig. 10c). Although this conformational constraint is likely to have a substantial effect on cobalt reactivity, it is not generic to enzymes catalyzing radical methyl transfer reactions (e.g. Mmp10 & TokK) since in QCMT, the geometry of the tetrapyrrole ring is similar to the one found in base-on cobalamin.
5. Structural relationships with other iron-sulfur and B12-dependent enzymes
As mentioned above, the association of a [Fe4S4] cluster and a cobalamin cofactor is not unique to B12-dependent radical SAM enzymes and has been previously shown in other enzyme families, including the reductive dehalogenases and the CFeSP. The structures of two distinct types of reductive dehalogenases, NpRdhA87 and PceA,88 have revealed striking structural similarities within this enzyme family, with notably a [Fe4S4] cluster and a cobalamin binding domain located in the C- and N-terminal regions, respectively, along with additional domains. However, in contrast to B12-dependent radical SAM enzymes, the cobalamin cofactor resides in the innermost region of the enzyme's core, whereas two [Fe4S4] clusters are located near the protein surface. Remarkably, one of these clusters is positioned within the van der Waals distance to the tetrapyrrole ring while the other one is located beneath cobalamin, consistent with their function as an electron shuttle (Fig. 11).In CFeSP, the relative organization of the [Fe4S4] cluster and the cobalamin cofactors also differs. Indeed, while both cofactors are located within the same subunit, the distance between the cobalt atom and the nearest iron of the [Fe4S4] cluster is ∼39 Å compared to 8.3 Å in reductive dehalogenases and ∼11–12 Å in B12-dependent radical SAM methyltransferases. Here again, the function of the [Fe4S4] cluster is to shuttle electrons. However, in CFeSP, which catalyzes nucleophilic chemistry, the purpose of the [Fe4S4] cluster is to reactivate the enzyme by reducing the cobalt cofactor from Cob(II) to Cob(I).
Hence, while B12-dependent radical SAM methyltransferases, CFeSP, and reductive dehalogenases use cobalamin and iron-sulfur clusters for catalysis, they use both cofactors for distinct functions and chemistry.
6. Functional perspective on B12-dependent SAM enzymes
B12-dependent radical SAM enzymes have been shown to be involved in diverse transformations with novel functions likely to emerge within this superfamily. Using sequence similarity networks,162 we can have an estimate of their structural and functional diversity (Fig. 12). By analyzing 83![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 589 accession IDs sorted into 37
589 accession IDs sorted into 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 236 “meta-nodes”, it is possible to delineate more than 100 groups. The largest group contains about one-third of the nodes, and the majority of the groups have less than 100 unique protein sequences. To date, structural information has been obtained only for discrete groups (red labels in Fig. 12), with the largest ones still unexplored. All groups, with at least one functionally characterized enzyme, include C-methyltransferases, which appear to be scattered throughout the B12-dependent radical SAM network.
236 “meta-nodes”, it is possible to delineate more than 100 groups. The largest group contains about one-third of the nodes, and the majority of the groups have less than 100 unique protein sequences. To date, structural information has been obtained only for discrete groups (red labels in Fig. 12), with the largest ones still unexplored. All groups, with at least one functionally characterized enzyme, include C-methyltransferases, which appear to be scattered throughout the B12-dependent radical SAM network.
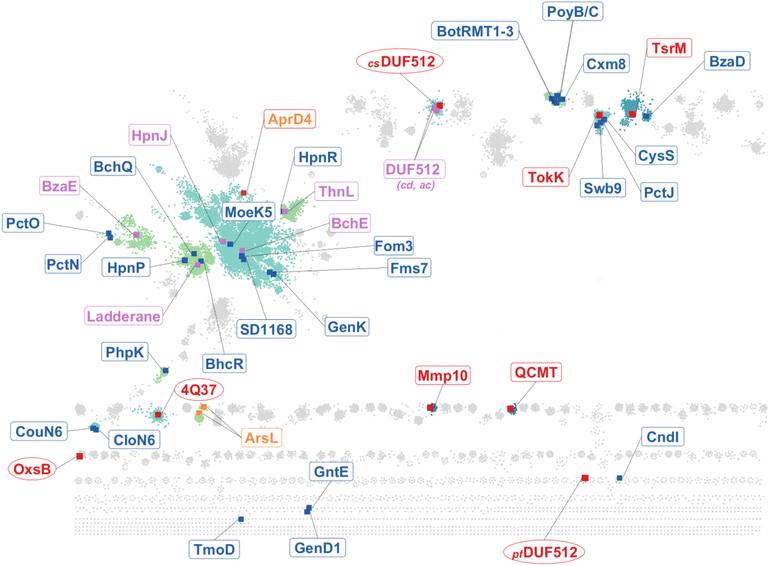 | ||
| Fig. 12 The sequence similarity network of B12-dependent radical SAM enzymes. In blue, non-structurally characterized methyltransferases: Csp2-methyltransferases: CouN6, CloN6,163 BzaD;59 P-methyltransferase: PhpK;22 iterative methyltransferases: PoyB,164 CysS,135 Swb9,165 BchQ,166 TmoD;167 Csp3-methyltransferases: PoyC,131 PctJ,168 BotRMT 1–3,169 HpnR,170 Fms7,171 GenK,133,172 Fom3,31,137 MoeK5,173 SD1168,174 BchR,166 HpnP,170 PctO,175 PctN,175 CndI,176 GntE/GenD1,177 Cmx8. In purple: enzymes catalyzing other or unknown transformations: BchE,178 Ladderane,179 ThnL,35 BzaE,59 HpnJ,180cdDUF512,161acDUF512.161 In red, structurally characterized enzymes with known functions (rounded rectangles): TsrM,23,25,121,181 Mmp10,33,182 QCMT,148,183 TokK,147 and other or unknown transformations (ellipses): OxsB,29 4Q37,159csDUF512,161pfDUF512.161 In orange, unrelated enzymes such as184 AprD4 and ArsL.130 The network was generated from the list of Uniprot IDs of all current B12-dependent radical SAM enzymes (clusters 2.1, 1.4 and 1.8 on radicalsam.org2) using the Enzyme Function Initiative (EFI) enzyme similarity tool (EFI-EST) with an alignment score of 60. To improve the visualization of the network with cytoscape,185 we chose the representative node (“rep node”) representation where sequences sharing > 50% identity are grouped into the same meta-node. | ||
With the exception of OxsB, enzymes involved in non-methylation reactions belong to the largest group of B12-dependent radical SAM enzymes, and show no clear structural features that would allow them to be distinguished from alkylating enzymes. Similarly, some B12-dependent radical SAM enzymes have been shown or predicted to catalyze non-radical methylations, such as TsrM,23,25,121,181 or CloN6.163 However, they belong to distinct groups and appear to share more functional than structural similarities. Further characterization of the structural features and cofactor content of these different groups is necessary before a comprehensive understanding of this emerging enzyme family can be obtained. For instance, Mmp10, with only a radical SAM and a B12-binding domain, might represent one of the simplest scaffolds,33 but its structural analysis revealed unanticipated complexity, with four distinct metallic centers, including the first mononuclear iron identified within this superfamily. Finally, although sequence similarity networks are useful for encapsulating radical SAM enzyme diversity, they may also include unrelated enzyme architectures. For instance, several radical SAM enzymes,184 which do not bind cobalamin (i.e., AprD4 and homologs of the 3-amino-3-carboxypropyl transferase, ArsL130) but have domains remotely related to the Rossmann fold, are present within this SSN (Fig. 12). However, unexpected discoveries are likely to be made along the way while exploring this constellation of biocatalysts.139,186,187
Conclusions and future outlook
B12-dependent radical SAM enzymes, despite their broad distribution and key roles in major biosynthetic pathways from vitamin to antibiotic biosynthesis, have been investigated only recently. While initially mainly regarded as methyltransferases, in the last decade, these enzymes have been shown to catalyze novel reactions that remain to be fully deciphered.139,186 Notably, in non-methyl transfer reactions, the role and exact nature of the cobalamin cofactor remain to be clarified.Structural analysis has provided unexpected insights into the mechanisms of these unique biocatalysts. First, it has recently been discovered that B12-dependent radical SAM enzymes can adopt two distinct architectures, with the B12-binding domain at the N- or C-terminus, delineating two classes of enzymes (Fig. 8).29,33,147,161 More intriguingly, several of these enzymes exhibit tetra-coordination of the radical SAM [Fe4S4] cluster using, in addition to the three canonical cysteine ligands, another protein residue. This conformation, never observed since the inception of this enzyme family 25 years ago,21 is likely important for these enzymes to switch from nucleophilic to radical chemistry. However, due to the limited number of structures solved, notably in the presence of genuine cofactors and substrates, it remains unclear whether these are isolated cases or a general trend within this emerging family of enzymes.
The cobalamin cofactor has been shown to adopt a base-off coordination, as observed in other B12-dependent enzymes. However, in some B12-dependent radical SAM enzymes, the tetrapyrrole ring has a planar conformation with an axial C8 side chain. The purpose of this distinct geometry is unclear, but, together with key residues that interact with both faces of the cobalamin cofactor, it likely tunes the reactivity and redox potential of the Co-center.
Despite the outstanding progress made in the last decade, much remains to be explored within this superfamily. As detailed here, we still lack structurally characterized representatives for the main groups of B12-dependent radical SAM enzymes. Some key aspects of their catalysis, such as the factors that control their stereochemistry or the ability of some of them to catalyze iterative methyl–transfer reactions, have only been marginally investigated. A deeper understanding of these biocatalysts will be necessary to harness their powerful biosynthetic potential, particularly the formation of C–C bonds between unreactive carbon atoms, chemical prowess unique to radical SAM enzymes.95,139 B12-dependent radical SAM enzymes likely hold surprises in store, notably regarding their chemical versatility and their ability to catalyze unprecedented reactions.
Author contributions
O. B., M.T. N & A. B: original draft. All authors contributed to the figures, review, and editing of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data that support some findings of this study, notably the SSN, are available from the corresponding author, O. Berteau, upon reasonable request.Acknowledgements
The authors' work was supported by ANR (ANR-20-CE44-0005, ANR-21-CE11-0030 & ANR-23-CE07-0046).References
- A. Benjdia and O. Berteau, Front. Chem., 2021, 9, 678068 CrossRef CAS PubMed.
- N. Oberg, T. W. Precord, D. A. Mitchell and J. A. Gerlt, ACS Bio Med Chem Au, 2022, 2, 22–35 CrossRef CAS PubMed.
- J. B. Broderick, B. R. Duffus, K. S. Duschene and E. M. Shepard, Chem. Rev., 2014, 114, 4229–4317 CrossRef.
- J. L. Vey and C. L. Drennan, Chem. Rev., 2011, 111, 2487–2506 CrossRef PubMed.
- T. F. Henshaw, J. Cheek and J. B. Broderick, J. Am. Chem. Soc., 2000, 122, 8331–8332 CrossRef.
- J. L. Vey, J. Yang, M. Li, W. E. Broderick, J. B. Broderick and C. L. Drennan, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 16137–16141 Search PubMed.
- A. Benjdia, J. Leprince, A. Guillot, H. Vaudry, S. Rabot and O. Berteau, J. Am. Chem. Soc., 2007, 129, 3462–3463 CrossRef PubMed.
- A. Benjdia, J. Leprince, C. Sandstrom, H. Vaudry and O. Berteau, J. Am. Chem. Soc., 2009, 131, 8348–8349 CrossRef.
- K. Ono, T. Okajima, M. Tani, S. Kuroda, D. Sun, V. L. Davidson and K. Tanizawa, J. Biol. Chem., 2006, 281, 13672–13684 Search PubMed.
- B. P. Anton, L. Saleh, J. S. Benner, E. A. Raleigh, S. Kasif and R. J. Roberts, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 1826–1831 CrossRef PubMed.
- F. Soualmia, M. V. Cherrier, T. Chauvire, M. Mauger, P. Tatham, A. Guillot, X. Guinchard, L. Martin, P. Amara, J. M. Mouesca, M. Daghmoum, A. Benjdia, S. Gambarelli, O. Berteau and Y. Nicolet, J. Am. Chem. Soc., 2024, 146, 6493–6505 CrossRef PubMed.
- B. L. Greene, G. Kang, C. Cui, M. Bennati, D. G. Nocera, C. L. Drennan and J. Stubbe, Annu. Rev. Biochem., 2020, 89, 45–75 CrossRef.
- A. Chandor, O. Berteau, T. Douki, D. Gasparutto, Y. Sanakis, S. Ollagnier-de-Choudens, M. Atta and M. Fontecave, J. Biol. Chem., 2006, 281, 26922–26931 CrossRef PubMed.
- A. Benjdia, K. Heil, T. R. Barends, T. Carell and I. Schlichting, Nucleic Acids Res., 2012, 40, 9308–9318 CrossRef PubMed.
- B. Philmus, L. Decamps, O. Berteau and T. P. Begley, J. Am. Chem. Soc., 2015, 137, 5406–5413 Search PubMed.
- L. Decamps, B. Philmus, A. Benjdia, R. White, T. P. Begley and O. Berteau, J. Am. Chem. Soc., 2012, 134, 18173–18176 Search PubMed.
- J. M. Kuchenreuther, W. K. Myers, T. A. Stich, S. J. George, Y. Nejatyjahromy, J. R. Swartz and R. D. Britt, Science, 2013, 342, 472–475 CrossRef PubMed.
- J. M. Kuchenreuther, W. K. Myers, D. L. Suess, T. A. Stich, V. Pelmenschikov, S. A. Shiigi, S. P. Cramer, J. R. Swartz, R. D. Britt and S. J. George, Science, 2014, 343, 424–427 CrossRef PubMed.
- A. S. Gizzi, T. L. Grove, J. J. Arnold, J. Jose, R. K. Jangra, S. J. Garforth, Q. Du, S. M. Cahill, N. G. Dulyaninova, J. D. Love, K. Chandran, A. R. Bresnick, C. E. Cameron and S. C. Almo, Nature, 2018, 558, 610–614 Search PubMed.
- P. Hanzelmann and H. Schindelin, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 12870–12875 CrossRef PubMed.
- H. J. Sofia, G. Chen, B. G. Hetzler, J. F. Reyes-Spindola and N. E. Miller, Nucleic Acids Res., 2001, 29, 1097–1106 CrossRef PubMed.
- W. J. Werner, K. D. Allen, K. Hu, G. L. Helms, B. S. Chen and S. C. Wang, Biochemistry, 2011, 50, 8986–8988 CrossRef.
- S. Pierre, A. Guillot, A. Benjdia, C. Sandstrom, P. Langella and O. Berteau, Nat. Chem. Biol., 2012, 8, 957–959 CrossRef.
- A. Parent, A. Benjdia, A. Guillot, X. Kubiak, C. Balty, B. Lefranc, J. Leprince and O. Berteau, J. Am. Chem. Soc., 2018, 140, 2469–2477 CrossRef PubMed.
- A. Benjdia, S. Pierre, C. Gherasim, A. Guillot, M. Carmona, P. Amara, R. Banerjee and O. Berteau, Nat. Commun., 2015, 6, 8377 CrossRef PubMed.
- N. D. Lanz, A. J. Blaszczyk, E. L. McCarthy, B. Wang, R. X. Wang, B. S. Jones and S. J. Booker, Biochemistry, 2018, 57, 1475–1490 CrossRef.
- A. J. Blaszczyk, A. Silakov, B. Zhang, S. J. Maiocco, N. D. Lanz, W. L. Kelly, S. J. Elliott, C. Krebs and S. J. Booker, J. Am. Chem. Soc., 2016, 138, 3416–3426 CrossRef.
- D. R. Marous, E. P. Lloyd, A. R. Buller, K. A. Moshos, T. L. Grove, A. J. Blaszczyk, S. J. Booker and C. A. Townsend, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 10354–10358 CrossRef.
- J. Bridwell-Rabb, A. Zhong, H. G. Sun, C. L. Drennan and H. W. Liu, Nature, 2017, 544, 322–326 CrossRef.
- K. D. Allen and S. C. Wang, Arch. Biochem. Biophys., 2014, 543, 67–73 CrossRef.
- B. Wang, A. J. Blaszczyk, H. L. Knox, S. Zhou, E. J. Blaesi, C. Krebs, R. X. Wang and S. J. Booker, Biochemistry, 2018, 57, 4972–4984 CrossRef PubMed.
- F. Soualmia, A. Guillot, N. Sabat, C. Brewee, X. Kubiak, M. Haumann, X. Guinchard, A. Benjdia and O. Berteau, Chem. – Eur. J., 2022, 28, e202200627, DOI:10.1002/chem.202200627.
- C. D. Fyfe, N. Bernardo-Garcia, L. Fradale, S. Grimaldi, A. Guillot, C. Brewee, L. M. G. Chavas, P. Legrand, A. Benjdia and O. Berteau, Nature, 2022, 602, 336–342 CrossRef PubMed.
- M. Wiesselmann, S. Hebecker, J. M. Borrero-de Acuna, M. Nimtz, D. Bollivar, L. Jansch, J. Moser and D. Jahn, Biochem. J., 2020, 477, 4635–4654 CrossRef CAS PubMed.
- E. K. Sinner, R. Li, D. R. Marous and C. A. Townsend, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2206494119 CrossRef PubMed.
- P. H. Degnan, N. A. Barry, K. C. Mok, M. E. Taga and A. L. Goodman, Cell Host Microbe, 2014, 15, 47–57 CrossRef.
- Z. F. Hallberg, E. C. Seth, K. Thevasundaram and M. E. Taga, Biochemistry, 2022, 61, 2791–2796, DOI:10.1021/acs.biochem.2c00367.
- C. Martino, B. P. Kellman, D. R. Sandoval, T. M. Clausen, R. Cooper, A. Benjdia, F. Soualmia, A. E. Clark, A. F. Garretson, C. A. Marotz, S. J. Song, S. Wandro, L. S. Zaramela, R. A. Salido, Q. Zhu, E. Armingol, Y. Vazquez-Baeza, D. McDonald, J. T. Sorrentino, B. Taylor, P. Belda-Ferre, P. Das, F. Ali, C. Liang, Y. Zhang, L. Schifanella, A. Covizzi, A. Lai, A. Riva, C. Basting, C. A. Broedlow, A. S. Havulinna, P. Jousilahti, M. Estaki, T. Kosciolek, R. Kuplicki, T. A. Victor, M. P. Paulus, K. E. Savage, J. L. Benbow, E. S. Spielfogel, C. A. M. Anderson, M. E. Martinez, J. V. Lacey, Jr., S. Huang, N. Haiminen, L. Parida, H. C. Kim, J. A. Gilbert, D. A. Sweeney, S. M. Allard, A. D. Swafford, S. Cheng, M. Inouye, T. Niiranen, M. Jain, V. Salomaa, K. Zengler, N. R. Klatt, J. Hasty, O. Berteau, A. F. Carlin, J. D. Esko, N. E. Lewis and R. Knight, mBio, 2025, 16, e0401524 CrossRef PubMed.
- E. P. Balskus, ACS Infect. Dis., 2018, 4, 1–2 CrossRef PubMed.
- Y. Mathur and A. B. Hazra, Curr. Opin. Struct. Biol., 2022, 77, 102490 CrossRef PubMed.
- E. L. Rickes, N. G. Brink, F. R. Koniuszy, T. R. Wood and K. Folkers, Science, 1948, 108, 134 CrossRef PubMed.
- E. L. Smith, Nature, 1948, 161, 638 CrossRef PubMed.
- D. C. Hodgkin, J. Kamper, M. Mackay, J. Pickworth, K. N. Trueblood and J. G. White, Nature, 1956, 178, 64–66 CrossRef PubMed.
- C. Gherasim, M. Lofgren and R. Banerjee, J. Biol. Chem., 2013, 288, 13186–13193 CrossRef PubMed.
- J. F. Kolhouse and R. H. Allen, Proc. Natl. Acad. Sci. U. S. A., 1977, 74, 921–925 CrossRef PubMed.
- E. E. Putnam and A. L. Goodman, PLoS Pathog., 2020, 16, e1008208 CrossRef PubMed.
- W. Roth and M. Mohamadzadeh, EBioMedicine, 2021, 73, 103676 CrossRef PubMed.
- H. M. Guetterman, S. L. Huey, R. Knight, A. M. Fox, S. Mehta and J. L. Finkelstein, Adv. Nutr., 2021, 13, 530–558 CrossRef PubMed.
- K. C. Mok and M. E. Taga, J. Bacteriol., 2013, 195, 1902–1911 CrossRef PubMed.
- S. Yi, E. C. Seth, Y. J. Men, S. P. Stabler, R. H. Allen, L. Alvarez-Cohen and M. E. Taga, Appl. Environ. Microbiol., 2012, 78, 7745–7752 CrossRef PubMed.
- M. J. Gray and J. C. Escalante-Semerena, Mol. Microbiol., 2009, 74, 1198–1210 Search PubMed.
- A. T. Ma, B. Tyrell and J. Beld, Mol. Microbiol., 2020, 113, 89–102 CrossRef PubMed.
- K. C. Mok, O. M. Sokolovskaya, A. M. Nicolas, Z. F. Hallberg, A. Deutschbauer, H. K. Carlson and M. E. Taga, mBio, 2020, 11 DOI:10.1128/mbio.02507-02520.
- K. C. Mok, Z. F. Hallberg and M. E. Taga, Methods Enzymol., 2022, 668, 61–85 Search PubMed.
- P. H. Degnan, M. E. Taga and A. L. Goodman, Cell Metab., 2014, 20, 769–778 CrossRef PubMed.
- D. A. Bryant, C. N. Hunter and M. J. Warren, J. Biol. Chem., 2020, 295, 6888–6925 CrossRef PubMed.
- A. I. Scott, J. Org. Chem., 2003, 68, 2529–2539 CrossRef PubMed.
- Y. Mathur, S. Sreyas, P. M. Datar, M. B. Sathian and A. B. Hazra, J. Biol. Chem., 2020, 295, 10522–10534 CrossRef PubMed.
- A. B. Hazra, A. W. Han, A. P. Mehta, K. C. Mok, V. Osadchiy, T. P. Begley and M. E. Taga, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 10792–10797 CrossRef PubMed.
- M. E. Taga, N. A. Larsen, A. R. Howard-Jones, C. T. Walsh and G. C. Walker, Nature, 2007, 446, 449–453 CrossRef.
- G. R. Campbell, M. E. Taga, K. Mistry, J. Lloret, P. J. Anderson, J. R. Roth and G. C. Walker, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 4634–4639 CrossRef PubMed.
- M. J. Gray and J. C. Escalante-Semerena, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 2921–2926 Search PubMed.
- R. Banerjee and S. W. Ragsdale, Annu. Rev. Biochem., 2003, 72, 209–247 CrossRef.
- R. Banerjee, Chem. Biol., 1997, 4, 175–186 CrossRef PubMed.
- J. Z. Liu, W. Xu, A. Chistoserdov and R. K. Bajpai, Appl. Biochem. Biotechnol., 2016, 179, 1073–1100 CrossRef.
- F. Parmeggiani, N. J. Weise, S. T. Ahmed and N. J. Turner, Chem. Rev., 2018, 118, 73–118 CrossRef CAS PubMed.
- R. Banerjee, Chem. Rev., 2003, 103, 2083–2094 CrossRef CAS.
- K. Gruber and C. Kratky, Curr. Opin. Chem. Biol., 2002, 6, 598–603 CrossRef CAS PubMed.
- R. J. Sension, A. G. Cole, A. D. Harris, C. C. Fox, N. W. Woodbury, S. Lin and E. N. Marsh, J. Am. Chem. Soc., 2004, 126, 1598–1599 CrossRef CAS PubMed.
- J. B. Rommel and J. Kastner, J. Am. Chem. Soc., 2011, 133, 10195–10203 CrossRef CAS PubMed.
- K. Gruber, V. Csitkovits, A. Lyskowski, C. Kratky and B. Krautler, Angew. Chem., 2022, 134, e202208295 CrossRef PubMed.
- P. Frey, Comprehensive Natural Products II: Chemistry and Biology, 2010, vol. 7, pp. 501–546 Search PubMed.
- J. Stubbe and W. A. van der Donk, Chem. Biol., 1995, 2, 793–801 CrossRef CAS.
- J. R. Guest, S. Friedman, D. D. Woods and E. L. Smith, Nature, 1962, 195, 340–342 CrossRef CAS.
- R. V. Banerjee, N. L. Johnston, J. K. Sobeski, P. Datta and R. G. Matthews, J. Biol. Chem., 1989, 264, 13888–13895 CrossRef CAS.
- R. V. Banerjee, V. Frasca, D. P. Ballou and R. G. Matthews, Biochemistry, 1990, 29, 11101–11109 CrossRef CAS.
- R. V. Banerjee, S. R. Harder, S. W. Ragsdale and R. G. Matthews, Biochemistry, 1990, 29, 1129–1135 CrossRef CAS PubMed.
- R. V. Banerjee and R. G. Matthews, FASEB J., 1990, 4, 1450–1459 CrossRef CAS.
- C. L. Drennan, S. Huang, J. T. Drummond, R. G. Matthews and M. L. Ludwig, Science, 1994, 266, 1669–1674 CrossRef CAS.
- C. L. Drennan, R. G. Matthews and M. L. Ludwig, Curr. Opin. Struct. Biol., 1994, 4, 919–929 CrossRef CAS.
- S. Menon and S. W. Ragsdale, Biochemistry, 1998, 37, 5689–5698 CrossRef CAS.
- Y. Kung, N. Ando, T. I. Doukov, L. C. Blasiak, G. Bender, J. Seravalli, S. W. Ragsdale and C. L. Drennan, Nature, 2012, 484, 265–269 CrossRef CAS.
- T. Svetlitchnaia, V. Svetlitchnyi, O. Meyer and H. Dobbek, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 14331–14336 CrossRef CAS.
- S. W. Ragsdale and M. Kumar, Chem. Rev., 1996, 96, 2515–2540 CrossRef CAS PubMed.
- S. Wiley, C. Griffith, P. Eckert, A. P. Mueller, R. Nogle, S. D. Simpson, M. Kopke, M. Can, R. Sarangi, K. Kubarych and S. W. Ragsdale, J. Biol. Chem., 2024, 300, 107503 CrossRef CAS.
- T. Wongnate, D. Sliwa, B. Ginovska, D. Smith, M. W. Wolf, N. Lehnert, S. Raugei and S. W. Ragsdale, Science, 2016, 352, 953–958 CrossRef CAS.
- K. A. Payne, C. P. Quezada, K. Fisher, M. S. Dunstan, F. A. Collins, H. Sjuts, C. Levy, S. Hay, S. E. Rigby and D. Leys, Nature, 2015, 517, 513–516 CrossRef CAS.
- M. Bommer, C. Kunze, J. Fesseler, T. Schubert, G. Diekert and H. Dobbek, Science, 2014, 346, 455–458 Search PubMed.
- B. A. van de Pas, H. Smidt, W. R. Hagen, J. van der Oost, G. Schraa, A. J. Stams and W. M. de Vos, J. Biol. Chem., 1999, 274, 20287–20292 Search PubMed.
- N. Christiansen, B. K. Ahring, G. Wohlfarth and G. Diekert, FEBS Lett., 1998, 436, 159–162 Search PubMed.
- L. A. Hug, F. Maphosa, D. Leys, F. E. Loffler, H. Smidt, E. A. Edwards and L. Adrian, Philos. Trans. R. Soc. Lond. B Biol. Sci., 2013, 368, 20120322 CrossRef.
- C. Kunze, M. Bommer, W. R. Hagen, M. Uksa, H. Dobbek, T. Schubert and G. Diekert, Nat. Commun., 2017, 8, 15858 CrossRef CAS.
- A. Parthasarathy, T. A. Stich, S. T. Lohner, A. Lesnefsky, R. D. Britt and A. M. Spormann, J. Am. Chem. Soc., 2015, 137, 3525–3532 CrossRef CAS.
- R. P. Schmitz, J. Wolf, A. Habel, A. Neumann, K. Ploss, A. Svatos, W. Boland and G. Diekert, Environ. Sci. Technol., 2007, 41, 7370–7375 CrossRef CAS.
- A. Benjdia, C. Balty and O. Berteau, Front. Chem., 2017, 5, 87 CrossRef.
- N. Mahanta, G. A. Hudson and D. A. Mitchell, Biochemistry, 2017, 56, 5229–5524 CrossRef CAS.
- https://radicalsam.org/ .
- A. Benjdia and O. Berteau, Biochem. Soc. Trans., 2016, 44, 109–115 CrossRef CAS PubMed.
- O. Berteau, J. Biol. Chem., 2018, 293, 8312–8313 CrossRef CAS.
- R. Rebeil, Y. Sun, L. Chooback, M. Pedraza-Reyes, C. Kinsland, T. P. Begley and W. L. Nicholson, J. Bacteriol., 1998, 180, 4879–4885 Search PubMed.
- M. G. Friedel, O. Berteau, J. C. Pieck, M. Atta, S. Ollagnier-de-Choudens, M. Fontecave and T. Carell, Chem. Commun., 2006, 445–447, 10.1039/b514103f.
- S. Arragain, R. Garcia-Serres, G. Blondin, T. Douki, M. Clemancey, J. M. Latour, F. Forouhar, H. Neely, G. T. Montelione, J. F. Hunt, E. Mulliez, M. Fontecave and M. Atta, J. Biol. Chem., 2009, 285, 5792–5801, DOI:10.1074/jbc.M109.065516.
- S. Ollagnier, E. Mulliez, J. Gaillard, R. Eliasson, M. Fontecave and P. Reichard, J. Biol. Chem., 1996, 271, 9410–9416 CrossRef CAS.
- O. Berteau, A. Guillot, A. Benjdia and S. Rabot, J. Biol. Chem., 2006, 281, 22464–22470 Search PubMed.
- A. Benjdia, G. Deho, S. Rabot and O. Berteau, FEBS Lett., 2007, 581, 1009–1014 Search PubMed.
- A. Benjdia, S. Subramanian, J. Leprince, H. Vaudry, M. K. Johnson and O. Berteau, J. Biol. Chem., 2008, 283, 17815–17826 CrossRef CAS.
- A. Benjdia, S. Subramanian, J. Leprince, H. Vaudry, M. K. Johnson and O. Berteau, FEBS J., 2010, 277, 1906–1920 CrossRef CAS.
- F. Pierrel, H. L. Hernandez, M. K. Johnson, M. Fontecave and M. Atta, J. Biol. Chem., 2003, 278, 29515–29524 CrossRef CAS.
- T. L. Grove, J. S. Benner, M. I. Radle, J. H. Ahlum, B. J. Landgraf, C. Krebs and S. J. Booker, Science, 2011, 332, 604–607 CrossRef CAS.
- G. Layer, A. J. Pierik, M. Trost, S. E. Rigby, H. K. Leech, K. Grage, D. Breckau, I. Astner, L. Jansch, P. Heathcote, M. J. Warren, D. W. Heinz and D. Jahn, J. Biol. Chem., 2006, 281, 15727–15734 CrossRef CAS.
- J. E. Harmer, M. J. Hiscox, P. C. Dinis, S. J. Fox, A. Iliopoulos, J. E. Hussey, J. Sandy, F. T. Van Beek, J. W. Essex and P. L. Roach, Biochem. J., 2014, 464, 123–133 CrossRef CAS.
- I. Sanyal, G. Cohen and D. H. Flint, Biochemistry, 1994, 33, 3625–3631 CrossRef CAS.
- P. B. Vander Horn, A. D. Backstrom, V. Stewart and T. P. Begley, J. Bacteriol., 1993, 175, 982–992 CrossRef CAS.
- M. Kriek, F. Martins, R. Leonardi, S. A. Fairhurst, D. J. Lowe and P. L. Roach, J. Biol. Chem., 2007, 282, 17413–17423 CrossRef CAS.
- A. Chatterjee, Y. Li, Y. Zhang, T. L. Grove, M. Lee, C. Krebs, S. J. Booker, T. P. Begley and S. E. Ealick, Nat. Chem. Biol., 2008, 4, 758–765 CrossRef CAS.
- S. C. Wang and P. A. Frey, Biochemistry, 2007, 46, 12889–12895 CrossRef CAS PubMed.
- L. Fluhe, T. A. Knappe, M. J. Gattner, A. Schafer, O. Burghaus, U. Linne and M. A. Marahiel, Nat. Chem. Biol., 2012, 8, 350–357 Search PubMed.
- C. Balty, A. Guillot, L. Fradale, C. Brewee, M. Boulay, X. Kubiak, A. Benjdia and O. Berteau, J. Biol. Chem., 2019, 294, 14512–14525 CrossRef CAS.
- C. Balty, A. Guillot, L. Fradale, C. Brewee, B. Lefranc, C. Herrero, C. Sandström, J. Leprince, O. Berteau and A. Benjdia, J. Biol. Chem., 2020, 295, 16665–16677 CrossRef CAS.
- Y. Imai, K. J. Meyer, A. Iinishi, Q. Favre-Godal, R. Green, S. Manuse, M. Caboni, M. Mori, S. Niles, M. Ghiglieri, C. Honrao, X. Ma, J. J. Guo, A. Makriyannis, L. Linares-Otoya, N. Bohringer, Z. G. Wuisan, H. Kaur, R. Wu, A. Mateus, A. Typas, M. M. Savitski, J. L. Espinoza, A. O'Rourke, K. E. Nelson, S. Hiller, N. Noinaj, T. F. Schaberle, A. D'Onofrio and K. Lewis, Nature, 2019, 576, 459–464 CrossRef CAS.
- H. L. Knox, P. Y. Chen, A. J. Blaszczyk, A. Mukherjee, T. L. Grove, E. L. Schwalm, B. Wang, C. L. Drennan and S. J. Booker, Nat. Chem. Biol., 2021, 17, 485–491 CrossRef CAS PubMed.
- F. Soualmia, A. Guillot, N. Sabat, C. Brewee, X. Kubiak, M. Haumann, X. Guinchard, A. Benjdia and O. Berteau, Chemistry, 2022, 28, e202200627 Search PubMed.
- R. I. Sayler, T. A. Stich, S. Joshi, N. Cooper, J. T. Shaw, T. P. Begley, D. J. Tantillo and R. D. Britt, ACS Cent. Sci., 2019, 5, 1777–1785 CrossRef CAS.
- H. Yang, E. C. McDaniel, S. Impano, A. S. Byer, R. J. Jodts, K. Yokoyama, W. E. Broderick, J. B. Broderick and B. M. Hoffman, J. Am. Chem. Soc., 2019, 141, 12139–12146 CrossRef CAS.
- B. M. Hoffman, W. E. Broderick and J. B. Broderick, Annu. Rev. Biochem., 2023, 92, 333–349, DOI:10.1146/annurev-biochem-052621-090638.
- P. A. Frey, A. D. Hegeman and F. J. Ruzicka, Crit. Rev. Biochem. Mol. Biol., 2008, 43, 63–88 CrossRef CAS PubMed.
- M. Horitani, K. Shisler, W. E. Broderick, R. U. Hutcheson, K. S. Duschene, A. R. Marts, B. M. Hoffman and J. B. Broderick, Science, 2016, 352, 822–825 CrossRef CAS PubMed.
- S. Joshi, N. Mahanta, D. Fedoseyenko, H. Williams and T. P. Begley, J. Am. Chem. Soc., 2017, 139, 10952–10955 CrossRef CAS.
- S. Joshi, D. Fedoseyenko, V. Sharma, M. A. Nesbit, R. D. Britt and T. P. Begley, Biochemistry, 2021, 60, 1642–1646, DOI:10.1021/acs.biochem.1c00181.
- M. Mauger, A. Guillot, M. Seif-Eddine, S. Grimaldi, A. Benjdia and O. Berteau, J. Am. Chem. Soc., 2025, 147, 20381–20393 Search PubMed.
- A. Parent, A. Guillot, A. Benjdia, G. Chartier, J. Leprince and O. Berteau, J. Am. Chem. Soc., 2016, 138, 15515–15518 Search PubMed.
- M. I. Radle, D. V. Miller, T. N. Laremore and S. J. Booker, J. Biol. Chem., 2019, 294, 11712–11725 Search PubMed.
- H. J. Kim, Y. N. Liu, R. M. McCarty and H. W. Liu, J. Am. Chem. Soc., 2017, 139, 16084–16087 Search PubMed.
- M. I. McLaughlin, K. Pallitsch, G. Wallner, W. A. van der Donk and F. Hammerschmidt, Biochemistry, 2021, 60, 1587–1596 CrossRef CAS.
- Y. Wang and T. P. Begley, J. Am. Chem. Soc., 2020, 142, 9944–9954 CrossRef CAS PubMed.
- Y. Wang, B. Schnell, S. Baumann, R. Muller and T. P. Begley, J. Am. Chem. Soc., 2017, 139, 1742–1745 Search PubMed.
- S. Sato, F. Kudo, T. Kuzuyama, F. Hammerschmidt and T. Eguchi, Biochemistry, 2018, 57, 4963–4966 CrossRef CAS PubMed.
- J. Y. Suzuki, D. W. Bollivar and C. E. Bauer, Annu. Rev. Genet., 1997, 31, 61–89 Search PubMed.
- A. Benjdia and O. Berteau, Curr. Opin. Struct. Biol., 2023, 83, 102725 CrossRef CAS PubMed.
- T. Selmer, J. Kahnt, M. Goubeaud, S. Shima, W. Grabarse, U. Ermler and R. K. Thauer, J. Biol. Chem., 2000, 275, 3755–3760 Search PubMed.
- T. Frenzel, P. Zhou and H. G. Floss, Arch. Biochem. Biophys., 1990, 278, 35–40 CrossRef CAS PubMed.
- M. S. Lichstrahl, C. A. Townsend and E. K. Sinner, RSC Chem. Biol., 2022, 3, 1028–1034 RSC.
- F. Kudo, A. Minato, S. Sato, N. Nagano, C. Maruyama, Y. Hamano, J. Hashimoto, I. Kozone, K. Shin-Ya and T. Eguchi, Org. Lett., 2022, 24, 8975–8979 CrossRef CAS.
- P. Zhou, D. O'Hagan, U. Mocek, Z. Zeng, L. D. Yuen, T. Frenzel, C. J. Unkefer, J. M. Beale and H. G. Floss, J. Am. Chem. Soc., 1989, 111, 7274–7276 CrossRef CAS.
- Y. H. Lee, Y. C. Yeh, P. H. Fan, A. Zhong, M. W. Ruszczycky and H. W. Liu, J. Am. Chem. Soc., 2023, 145, 3656–3664 CrossRef CAS.
- M. I. McLaughlin, N. D. Lanz, P. J. Goldman, K. H. Lee, S. J. Booker and C. L. Drennan, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 9446–9450 CrossRef CAS.
- H. L. Knox, E. K. Sinner, C. A. Townsend, A. K. Boal and S. J. Booker, Nature, 2022, 602, 343–348 CrossRef CAS PubMed.
- R. J. Rodriguez Carrero, C. T. Lloyd, J. Borkar, S. Nath, L. M. Mirica, S. Nair, S. J. Booker and W. Metcalf, mBio, 2025, 16, e0354624 CrossRef PubMed.
- D. S. Froese, G. Kochan, J. R. Muniz, X. Wu, C. Gileadi, E. Ugochukwu, E. Krysztofinska, R. A. Gravel, U. Oppermann and W. W. Yue, J. Biol. Chem., 2010, 285, 38204–38213 CrossRef CAS.
- F. Mancia, N. H. Keep, A. Nakagawa, P. F. Leadlay, S. McSweeney, B. Rasmussen, P. Bosecke, O. Diat and P. R. Evans, Structure, 1996, 4, 339–350 CrossRef CAS PubMed.
- M. Tollinger, C. Eichmuller, R. Konrat, M. S. Huhta, E. N. Marsh and B. Krautler, J. Mol. Biol., 2001, 309, 777–791 CrossRef CAS PubMed.
- K. Gruber, B. Puffer and B. Krautler, Chem. Soc. Rev., 2011, 40, 4346–4363 RSC.
- F. Berkovitch, E. Behshad, K. H. Tang, E. A. Enns, P. A. Frey and C. L. Drennan, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 15870–15875 CrossRef CAS PubMed.
- K. R. Wolthers, C. Levy, N. S. Scrutton and D. Leys, J. Biol. Chem., 2010, 285, 13942–13950 CrossRef CAS.
- N. Shibata, J. Masuda, T. Tobimatsu, T. Toraya, K. Suto, Y. Morimoto and N. Yasuoka, Structure, 1999, 7, 997–1008 CrossRef CAS.
- J. Masuda, N. Shibata, Y. Morimoto, T. Toraya and N. Yasuoka, Structure, 2000, 8, 775–788 Search PubMed.
- N. Shibata, H. Tamagaki, N. Hieda, K. Akita, H. Komori, Y. Shomura, S. Terawaki, K. Mori, N. Yasuoka, Y. Higuchi and T. Toraya, J. Biol. Chem., 2010, 285, 26484–26493 CrossRef CAS.
- M. D. Sintchak, G. Arjara, B. A. Kellogg, J. Stubbe and C. L. Drennan, Nat. Struct. Biol., 2002, 9, 293–300 CrossRef CAS PubMed.
- J. A. Farias-Rico, S. Schmidt and B. Hocker, Nat. Chem. Biol., 2014, 10, 710–715 CrossRef CAS PubMed.
- J. Bridwell-Rabb and C. L. Drennan, Curr. Opin. Chem. Biol., 2017, 37, 63–70 CrossRef CAS PubMed.
- B. Wang, A. E. Solinski, M. I. Radle, O. M. Peduzzi, H. L. Knox, J. Cui, R. K. Maurya, N. H. Yennawar and S. J. Booker, ACS Bio Med Chem Au, 2024, 4, 319–330 CrossRef CAS PubMed.
- J. A. Gerlt, J. T. Bouvier, D. B. Davidson, H. J. Imker, B. Sadkhin, D. R. Slater and K. L. Whalen, Biochim. Biophys. Acta, 2015, 1854, 1019–1037 Search PubMed.
- L. Westrich, L. Heide and S. M. Li, Chembiochem, 2003, 4, 768–773 Search PubMed.
- M. F. Freeman, C. Gurgui, M. J. Helf, B. I. Morinaka, A. R. Uria, N. J. Oldham, H. G. Sahl, S. Matsunaga and J. Piel, Science, 2012, 338, 387–390 CrossRef CAS.
- K. Watanabe, K. Hotta, M. Nakaya, A. P. Praseuth, C. C. Wang, D. Inada, K. Takahashi, E. Fukushi, H. Oguri and H. Oikawa, J. Am. Chem. Soc., 2009, 131, 9347–9353 Search PubMed.
- A. Gomez Maqueo Chew, N. U. Frigaard and D. A. Bryant, J. Bacteriol., 2007, 189, 6176–6184 CrossRef PubMed.
- Y. Yu and W. A. van der Donk, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2205285119 CrossRef CAS.
- F. Kudo, Y. Kasama, T. Hirayama and T. Eguchi, J. Antibiot., 2007, 60, 492–503 CrossRef CAS PubMed.
- L. Huo, S. Rachid, M. Stadler, S. C. Wenzel and R. Muller, Chem. Biol., 2012, 19, 1278–1287 Search PubMed.
- P. V. Welander and R. E. Summons, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 12905–12910 Search PubMed.
- T. Dairi, T. Ohta, E. Hashimoto and M. Hasegawa, Mol. Gen. Genet., 1992, 236, 39–48 Search PubMed.
- H. J. Kim, R. M. McCarty, Y. Ogasawara, Y. N. Liu, S. O. Mansoorabadi, J. Levieux and H. W. Liu, J. Am. Chem. Soc., 2013, 135, 8093–8096 Search PubMed.
- B. Ostash, E. H. Doud, C. Lin, I. Ostash, D. L. Perlstein, S. Fuse, M. Wolpert, D. Kahne and S. Walker, Biochemistry, 2009, 48, 8830–8841 Search PubMed.
- K. D. Allen and S. C. Wang, Biochim. Biophys. Acta, 2014, 1844, 2135–2144 CrossRef CAS PubMed.
- F. Kudo, J. Zhang, S. Sato, A. Hirayama and T. Eguchi, Chembiochem, 2019, 20, 2458–2462 CrossRef CAS PubMed.
- S. Rachid, M. Scharfe, H. Blocker, K. J. Weissman and R. Muller, Chem. Biol., 2009, 16, 70–81 Search PubMed.
- C. Huang, F. Huang, E. Moison, J. Guo, X. Jian, X. Duan, Z. Deng, P. F. Leadlay and Y. Sun, Chem. Biol., 2015, 22, 251–261 CrossRef CAS PubMed.
- Z. M. Yang and C. E. Bauer, J. Bacteriol., 1990, 172, 5001–5010 CrossRef CAS PubMed.
- J. E. Rattray, M. Strous, H. J. Op den Camp, S. Schouten, M. S. Jetten and J. S. Damste, Biol. Direct, 2009, 4, 8 Search PubMed.
- C. L. Schmerk, P. V. Welander, M. A. Hamad, K. L. Bain, M. A. Bernards, R. E. Summons and M. A. Valvano, Environ. Microbiol., 2015, 17, 735–750 CrossRef CAS PubMed.
- W. L. Kelly, L. Pan and C. Li, J. Am. Chem. Soc., 2009, 131, 4327–4334 Search PubMed.
- D. Deobald, L. Adrian, C. Schone, M. Rother and G. Layer, Sci. Rep., 2018, 8, 7404 Search PubMed.
- J. Gagsteiger, S. Jahn, L. Heidinger, L. Gericke, J. N. Andexer, T. Friedrich, C. Loenarz and G. Layer, Angew Chem. Int. Ed. Engl., 2022, 61, e202204198 Search PubMed.
- W. Q. Liu, P. Amara, J. M. Mouesca, X. Ji, O. Renoux, L. Martin, C. Zhang, Q. Zhang and Y. Nicolet, J. Am. Chem. Soc., 2018, 140, 1365–1371 Search PubMed.
- P. Shannon, A. Markiel, O. Ozier, N. S. Baliga, J. T. Wang, D. Ramage, N. Amin, B. Schwikowski and T. Ideker, Genome Res., 2003, 13, 2498–2504 CrossRef CAS.
- J. Bridwell-Rabb, B. Li and C. L. Drennan, ACS Bio Med Chem Au, 2022, 2, 173–186 Search PubMed.
- X. Kubiak, I. Polsinelli, L. M. G. Chavas, C. D. Fyfe, A. Guillot, L. Fradale, C. Brewee, S. Grimaldi, G. Gerbaud, A. Thureau, P. Legrand, O. Berteau and A. Benjdia, Nat. Chem. Biol., 2024, 20, 382–391 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |