Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
New benzochromene-based compounds as potential EGFR-TK inhibitors: synthesis, anti-proliferative activity, molecular docking studies, and ADME profiles
Mohamed H. Hekal *a,
Abdullah Yahya Abdullah Alzahranib,
Saad Alrashdi
*a,
Abdullah Yahya Abdullah Alzahranib,
Saad Alrashdi c,
Fatma S. M. Abu El-Azm
c,
Fatma S. M. Abu El-Azm a and
Yasmeen M. Alia
a and
Yasmeen M. Alia
aDepartment of Chemistry, Faculty of Science, Ain Shams University, Abbassia 11566, Cairo, Egypt. E-mail: mohamed.hekal@sci.asu.edu.eg
bDepartment of Chemistry, Faculty of Science, King Khalid University, Abha, Saudi Arabia
cDepartment of Chemistry, College of Science, Jouf University, Sakaka 72341, Aljouf, Saudi Arabia
First published on 30th April 2026
Abstract
In this study, we report the synthesis and biological evaluation of a novel series of benzochromene and benzochromenopyrimidine derivatives employingenaminonitrile compound 1 as a key synthetic precursor. The structures of the synthesized compounds were elucidated using comprehensive analytical and spectroscopic techniques. The antiproliferative activities of the prepared derivatives were evaluated against three cancer cell lines, HCT-116, MCF-7, and HepG2, as well as the normal human lung fibroblast cell line, WI-38, using the MTT assay, with doxorubicin used as the standard reference drug. Among the tested derivatives, compounds 6, 7, and 10 demonstrated notable antiproliferative activity against all examined cancer cell lines. In particular, compound 7 exhibited potent cytotoxic effects, with IC50 values of 5.98, 6.52, and 8.51 µM against HCT-116, MCF-7, and HepG2 cells, respectively, comparable with that of doxorubicin. Importantly, compound 7 displayed low cytotoxicity toward WI-38 cells, resulting in the highest selectivity index (SI = 6.7). Molecular docking analysis further revealed that compound 7 exhibited the promising binding affinity toward EGFR, with a docking score of −9.06 kcal mol−1, comparable with that of gefitinib. Collectively, these findings highlight compound 7 as a potent EGFR-targeting kinase inhibitor with notable anticancer activity.
1. Introduction
Benzochromenes are fused oxygen-containing heterocycles generated by the annulation of a benzene ring. They have attracted considerable interest in medicinal chemistry owing to their structural diversity and wide range of biological activities. Benzochromene derivatives display antifungal and antibacterial activities,1–4 exhibit anti-inflammatory and analgesic effects,5 have antispasmolytic6 and antioxidant properties,7,8 and present anti-HIV activity9 due to the presence of a chromene (benzopyran) moiety. These scaffolds are widely found in natural products and present antianaphylactic,10 antileishmanial,11 estrogenic,12 antihelminthic,13 and CNS effects,14–16 antidepressant and anticonvulsant activities,17 as well as antitubercular properties.18Their potent anticancer properties have established benzochromenes as privileged scaffolds in drug discovery against a wide range of human cancers. Many derivatives induce cell-cycle arrest and apoptosis. They can bind to DNA,19 inhibit key enzymes, such as topoisomerase I and II, and regulate anti-apoptotic proteins such as Bcl-2.20 The identification of HA14-1 marked a significant advancement in the development of 4H-chromene-based anticancer agents. Evidence indicates that it acts synergistically with flavopiridol to downregulate Bcl-2 by disrupting the interaction between Bax and Bcl-2.21–23 Certain structural motifs have gained attention as potential leads in anticancer drug development (Fig. 1). Notably, benzo[f]chromene-2-carbonitrile derivatives (I) induce cell-cycle arrest and apoptosis in human cancer cells through the simultaneous inhibition of tubulin and topoisomerases,24 while 9-hydroxy/methoxy-1H-benzo[f]chromenes (II) show activity against resistant breast cancer cells via P-glycoprotein inhibition and apoptosis.25 Moreover, chromene-based derivatives have gained considerable research attention as attractive and versatile scaffolds for the design of potent antitumour agents.26 Several representatives highlight their therapeutic potential; for instance, 2-amino-4-(3-bromo-4,5-dimethoxyphenyl)-7-(dimethylamino)-4H-chromene-3-carbonitrile (III) has been identified as an effective tubulin polymerization inhibitor.27 In addition, Crolibulin™ (IV) has progressed to phase I/II clinical trials for the treatment of advanced solid tumors,28 while SP-6-27 (V) demonstrates strong antitumour activity against both cisplatin-sensitive and cisplatin-resistant ovarian cancer cell lines.29
Pyrimidine-based heterocycles complement this strategy, being key motifs in clinically used anticancer drugs such as gefitinib, afatinib, idelalisib, ibrutinib, and pralatrexate (Fig. 2).30 These scaffolds modulate diverse molecular targets involved in cancer progression, including kinases and DNA synthesis enzymes, and apoptotic pathways. Pyrimidinones, in particular, have been validated as effective chemotherapeutic agents due to their ability to inhibit tumor cell proliferation, induce apoptosis, and overcome drug resistance.31–33 The incorporation of pyrimidine moieties into chromene-based derivatives thus represents a promising approach for the development of novel multitargeted anticancer agents, providing a clear rationale for the design and synthesis of the compounds investigated in this study. In view of the promising anticancer activities of both benzochromene and pyrimidine scaffolds, this study focuses on the design, synthesis, and biological evaluation of novel chromene–pyrimidine hybrid derivatives. By combining these two pharmacophores, we aim to develop multitargeted agents capable of inhibiting key cancer-related pathways, providing potential leads for further anticancer drug development.
Motivated by the reported anticancer potential of these scaffolds, and as part of our ongoing efforts to synthesize and biologically evaluate a variety of heterocyclic systems,34–62 we herein report the development of a facile synthetic approach to benzochromene-based derivatives, with the aim of assessing their cytotoxic properties and epidermal growth factor receptor (EGFR) kinase inhibitory activity.
1.1. Rational design of the work
Inspired by the established relevance of benzochromeno-pyrimidine scaffolds in kinase inhibition, a series of rationally designed derivatives was developed as prospective EGFR inhibitors. Structure-based design targeted the ATP-binding site, emphasizing hinge-region hydrogen bonding, π–π stacking, and hydrophobic interactions to enhance the affinity and selectivity. The design strategy encompassed the following aspects. (i) A dimethoxyphenyl moiety was incorporated to increase lipophilicity and promote favorable hydrophobic interactions within the non-polar regions of the kinase pocket. (ii) The benzochromeno-pyrimidine core was preserved to maintain molecular planarity, which is essential for effective π–π stacking and hydrogen-bonding interactions. (iii) The cyano (–CN) substituent was strategically incorporated as a strong electron-withdrawing group to increase the electron deficiency of the heterocyclic core, thereby strengthening hydrogen bonding and dipole–dipole interactions with active-site residues. (iv) Integration of the pyrimidine ring was specifically intended to provide multiple hydrogen-bond donor and acceptor sites, thereby strengthening interactions with critical catalytic amino acids and enhancing binding affinity. (v) Systematic modification of the R2 substituent was carried out to fine-tune the electronic and steric environment, allowing modulation of polarity, solubility, and binding geometry (Fig. 3). | ||
| Fig. 3 Design rationale of benzochromene-based scaffolds (A and B), highlighting key substitutions for optimized EGFR binding and selectivity. | ||
2. Results and discussion
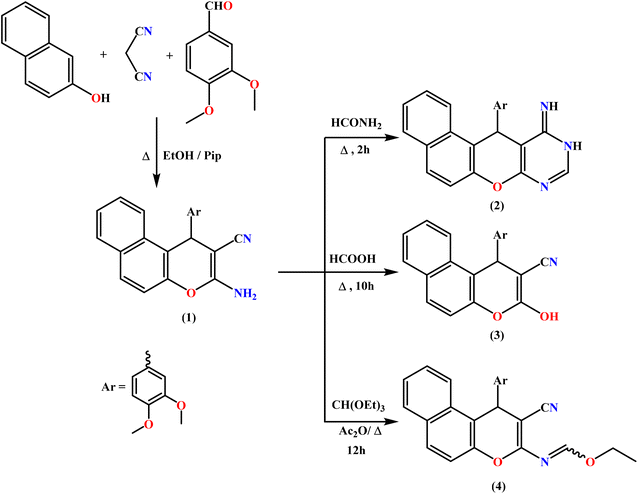
In this context, we focused on the construction of a new class of fused heterocycles based on the benzo[f]chromene framework, namely benzo[f]chromeno[2,3-d]pyrimidines and related derivatives, with the purpose of evaluating their antiproliferative activity. The benzo[f]chromene-2-carbonitrile scaffold 1, which serves as a pivotal synthetic precursor, was prepared via an efficient one-pot multicomponent protocol.63 This reaction involved the condensation of β-naphthol, malononitrile, and 3,4-dimethoxybenzaldehyde in absolute ethanol under basic catalysis using piperidine, affording the desired intermediate 1.Treatment of 1 with formamide under heating promotes cyclocondensation involving the amino and cyano groups, resulting in the formation of a fused pyrimidinone-containing naphthopyran derivative 2 (Scheme 1). The suggested structure was corroborated by the IR spectrum, which showed no absorption band corresponding to the CN group. Furthermore, the 1H NMR and 13C NMR spectra were consistent with the proposed structure of 2.
Heating intermediate 1 in formic acid resulted in deamination of the NH2 group, affording the unexpected compound 1-(3,4-dimethoxyphenyl)-3-hydroxy-1H-benzo[f]chromene-2-carbonitrile (3) instead of the anticipated fused pyrimidinone derivative.
To increase the reactivity of the less active amino group in the enaminonitrile intermediate 1, it was refluxed for 12 hours with triethyl orthoformate in the presence of freshly distilled acetic anhydride. This reaction resulted in the formation of the corresponding ethyl formimidate derivative 4, as illustrated in Scheme 1.
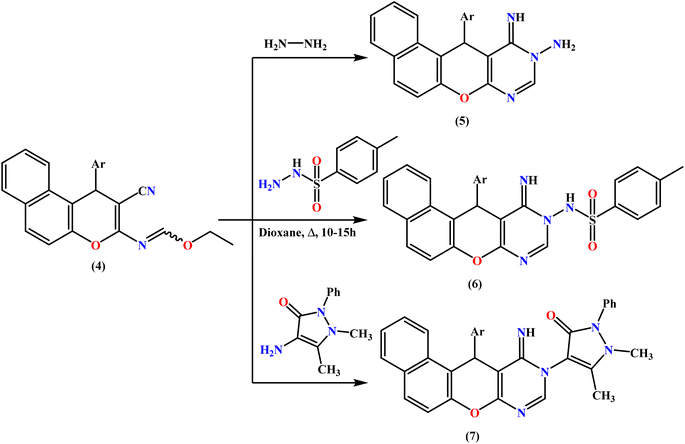
Ethyl formimidate 4 was reacted with different N-nucleophiles in an effort to synthesize novel benzochromenopyrimidines. Thus, treatment of compound 4 with hydrazine hydrate in absolute ethanol at room temperature for 6 hours produced the corresponding amino–imino derivative 5 in 78% yield (Scheme 2), and refluxing compound 4 with p-toluenesulfonohydrazide in 1,4-dioxane for 10 h produced benzochromenopyrimidine 6 as white crystals. The cyclized structure of 6 was confirmed by the expected elemental and spectroscopic analyses. The IR spectrum displayed an absorption band for the NH group at 3328 cm−1, while the absence of the nitrile absorption band was in accordance with the assigned structure 6.
Similarly, heating the tri-heterocyclic compound 4 with heterocyclic amines, such as 4-aminoantipyrine, in boiling 1,4-dioxane afforded the corresponding benzochromenopyrimidine derivative 7 in 68% yield, as illustrated in Scheme 2.
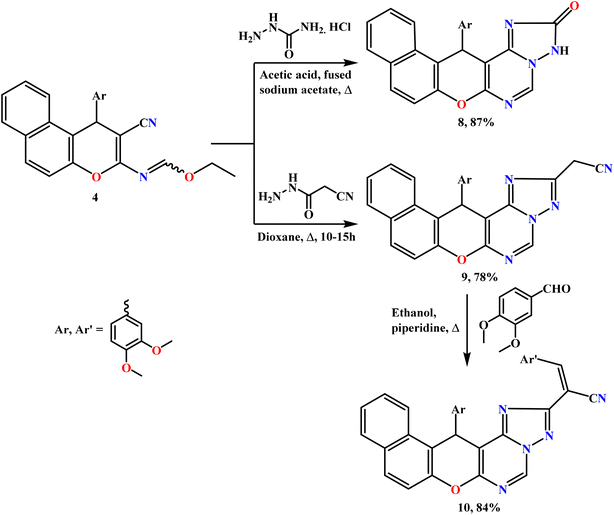
In this study, polyfunctionalized binary and fused triazole scaffolds were synthesized to facilitate the incorporation of functional groups that are known to enhance biological activity. The design of these scaffolds was guided by structure–activity relationship (SAR) insights, and earlier studies have emphasized the insecticidal properties of 1,2,4-triazoles and related heterocyclic compounds.34 The reactivity of compound 4 toward selected acid hydrazides, namely semicarbazide hydrochloride and 2-cyanoacetohydrazide, was subsequently investigated (Scheme 3). In this regard, the triheterocyclic compound 4 acted as a key precursor for the construction of tetra- and penta-heterocyclic systems. Thus, heating the ethyl formimidate derivative 4 with semicarbazide hydrochloride in acetic acid containing fused sodium acetate yielded the corresponding benzochromenotriazolopyrimidine derivative 8 as the sole product in 87% yield. The structure of compound 8 was determined from its spectroscopic data; notably, the IR spectrum lacked the characteristic C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N absorption band.
N absorption band.
Similarly, reaction of 4 with 2-cyanoacetohydrazide in refluxing 1,4-dioxane yielded acetonitrile derivative 9 as yellow crystals in 78% yield. Its structure was proposed from analytical and spectral data: the IR spectrum exhibited a characteristic ν(C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N) absorption at 2262 cm−1, and the 1H NMR spectrum displayed a singlet signal at δ 4.52 ppm, attributable to the –CH2CN protons, confirming the successful cyclization.
N) absorption at 2262 cm−1, and the 1H NMR spectrum displayed a singlet signal at δ 4.52 ppm, attributable to the –CH2CN protons, confirming the successful cyclization.
Importantly, the base-catalyzed condensation of active methylene candidates with aromatic aldehydes yielded the corresponding Knoevenagel products. In a similar manner, the reaction of compound 9 with 3,4-dimethoxybenzaldehyde in refluxing ethanol in the presence of a catalytic amount of piperidine led to the formation of the desired arylidene derivative 10 (84%), as illustrated in Scheme 3. The structure of compound 10 was verified through elemental analysis and spectroscopic data.
3. Antiproliferative activity
The synthesized compounds (2–10) were assessed for their antiproliferative activity on three human cancer cell lines, HCT-116, MCF-7, and HepG2, alongside the normal human lung fibroblast cell line WI-38. The cytotoxic potency was expressed as IC50 values (µM), while the selectivity index (SI) was calculated relative to the normal cell line. The results were compared with the reference anticancer drug doxorubicin.Among the tested compounds, derivatives 6, 7, and 10 demonstrated the most potent cytotoxic effects. In particular, compound 7 emerged as the most active candidate, displaying IC50 values of 5.98 ± 0.4 µM, 6.52 ± 0.4 µM, and 8.51 ± 0.6 µM against HCT-116, MCF-7, and HepG2 cells, respectively. These values approach those of doxorubicin, suggesting that compound 7 possesses strong anticancer potential. Importantly, compound 7 showed relatively low toxicity toward WI-38 cells (IC50 = 47.42 ± 3.8 µM), resulting in the highest selectivity index (SI = 6.7) among the tested compounds. This indicates that compound 7 preferentially targets cancer cells over normal cells, a desirable property for anticancer drug candidates.
Similarly, compounds 6 and 10 also demonstrated notable cytotoxic activities. Compound 6 exhibited IC50 values of 12.33 ± 1.4 µM, 10.66 ± 0.7 µM, and 7.26 ± 0.5 µM against HCT-116, MCF-7, and HepG2 cells, respectively, with an SI value of 5.4. Compound 10 also showed strong inhibition of cancer cell growth, particularly against MCF-7 (IC50 = 9.25 ± 0.6 µM) and HCT-116 (IC50 = 11.72 ± 1.2 µM), while maintaining relatively low toxicity toward WI-38 cells (IC50 = 67.32 ± 4.4 µM), leading to an SI of 5.6. These findings suggest that compounds 6 and 10 possess promising anticancer selectivity and may serve as valuable lead structures for further optimization (Table 1).
| Compounds | HCT-116 | MCF-7 | HepG2 | WI-38 | SIb |
|---|---|---|---|---|---|
| a IC50 values are presented as the mean ± standard deviation (SD) from three independent experiments.b SI: selectivity index, calculated as IC50 for WI-38 divided by the mean IC50 for the cancer cell lines. | |||||
| 2 | 65.17 ± 3.6 | 47.12 ± 2.8 | 71.24 ± 4.2 | 52.46 ± 4.2 | 0.9 |
| 3 | 56.88 ± 3.4 | 88.14 ± 4.6 | >100 | 36.16 ± 3.8 | 0.4 |
| 4 | 75.38 ± 4.7 | 62.64 ± 3.4 | 98.60 ± 4.8 | 44.54 ± 3.8 | 0.5 |
| 5 | 54.32 ± 3.2 | 71.44 ± 4.4 | 68.16 ± 3.6 | 54.84 ± 4.2 | 0.8 |
| 6 | 12.33 ± 1.4 | 10.66 ± 0.7 | 7.26 ± 0.5 | 55.23 ± 4.4 | 5.4 |
| 7 | 5.98 ± 0.4 | 6.52 ± 0.4 | 8.51 ± 0.6 | 47.42 ± 3.8 | 6.7 |
| 8 | 61.49 ± 3.3 | 55.21 ± 3.4 | 75.86 ± 4.2 | 28.61 ± 1.8 | 0.4 |
| 9 | 92.54 ± 4.9 | >100 | 81.63 ± 4.4 | 35.87 ± 2.2 | 0.3 |
| 10 | 11.72 ± 1.2 | 9.25 ± 0.6 | 15.14 ± 1.4 | 67.32 ± 4.4 | 5.6 |
| DOX | 5.23 ± 0.3 | 4.17 ± 0.2 | 4.50 ± 0.2 | 6.71 ± 0.5 | 1.4 |
In contrast, compounds 2, 3, 4, 5, 8, and 9 displayed moderate to weak cytotoxic activities, with IC50 values generally exceeding 50 µM against most tested cancer cell lines. Compound 3 showed relatively weak activity, particularly against HepG2 cells, where the IC50 exceeded 100 µM, indicating limited efficacy. Similarly, compounds 8 and 9 exhibited poor anticancer activity and low selectivity indices (0.4 and 0.3, respectively), suggesting that these derivatives lack sufficient potency and selectivity for further consideration.
4. ADME profiles
A computational ADME assessment was conducted using the SwissADME platform to evaluate the physicochemical properties, drug-likeness, and predicted oral bioavailability of compounds 2–10. Drug-likeness was assessed according to Lipinski's Rule of Five and Veber's criteria, widely recognized guidelines for evaluating the potential of compounds as orally active drug candidates.Most synthesized derivatives exhibited physicochemical properties compatible with orally active small molecules. Compounds 2–5 and 8–9 possess molecular weights ranging from 359.37 to 449.46 g mol−1, remaining within the recommended Lipinski threshold (<500 g mol−1). Compounds 6, 7, and 10 showed higher molecular weights (554.62–597.62 g mol−1), resulting in a single Lipinski violation. Nevertheless, their hydrogen-bonding capacities remained within acceptable limits (HBA = 5–9 and HBD = 0–2), indicating adequate polarity for intermolecular interactions without excessive hydrophilicity. The calculated lipophilicity values (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 3.09–5.27) indicate moderate hydrophobicity across the series, a property generally favorable for passive membrane diffusion. Most compounds fall within the recommended lipophilicity range (log
P = 3.09–5.27) indicate moderate hydrophobicity across the series, a property generally favorable for passive membrane diffusion. Most compounds fall within the recommended lipophilicity range (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≤ 5), with compound 10 slightly exceeding this value. All derivatives exhibited a predicted bioavailability score of 0.55, indicating a moderate probability of oral absorption.
P ≤ 5), with compound 10 slightly exceeding this value. All derivatives exhibited a predicted bioavailability score of 0.55, indicating a moderate probability of oral absorption.
According to Veber's criteria, TPSA values ranged from 71.71 to 123.91 Å2, well below the recommended threshold (≤140 Å2), supporting favorable intestinal permeability. The number of rotatable bonds (3–7) indicates moderate molecular flexibility, which may facilitate ligand–target interactions while preserving structural stability. Importantly, no Veber's rule violations were detected (Table 2, SI).
| Compound | MW | HBA | HBD | TPSA | Rotatable bonds | Bioavailability score | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≤ 5 P ≤ 5 |
Lipinski violations | Veber violations |
|---|---|---|---|---|---|---|---|---|---|
| 2 | 385.42 | 5 | 2 | 80.22 | 3 | 0.55 | 3.68 | 0 | 0 |
| 3 | 359.37 | 5 | 1 | 71.71 | 3 | 0.55 | 3.72 | 0 | 0 |
| 4 | 414.45 | 6 | 0 | 73.07 | 6 | 0.55 | 4.48 | 0 | 0 |
| 5 | 400.43 | 5 | 2 | 95.38 | 3 | 0.55 | 3.09 | 0 | 0 |
| 6 | 554.62 | 7 | 2 | 123.91 | 6 | 0.55 | 4.40 | 1 | 0 |
| 7 | 571.63 | 6 | 1 | 96.29 | 5 | 0.55 | 4.98 | 1 | 0 |
| 8 | 426.42 | 6 | 1 | 90.74 | 3 | 0.55 | 3.32 | 0 | 0 |
| 9 | 449.46 | 7 | 0 | 94.56 | 4 | 0.55 | 3.79 | 0 | 0 |
| 10 | 597.62 | 9 | 0 | 113.02 | 7 | 0.55 | 5.27 | 1 | 0 |
Among the evaluated derivatives, compound 4 displays one of the most balanced physicochemical profiles (MW = 414.45 g mol−1, log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 4.48, TPSA = 73.07 Å2, and six rotatable bonds), reflecting an optimal balance between lipophilicity, polarity, and conformational flexibility that may favor membrane permeability and pharmacokinetic performance.
P = 4.48, TPSA = 73.07 Å2, and six rotatable bonds), reflecting an optimal balance between lipophilicity, polarity, and conformational flexibility that may favor membrane permeability and pharmacokinetic performance.
To further evaluate absorption and brain penetration properties, the BOILED-Egg model was employed. This predictive model correlates lipophilicity (W![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) and TPSA to estimate passive GI absorption and blood–brain barrier permeability. In the BOILED-Egg diagram, the white area indicates compounds with a high probability of human intestinal absorption (HIA), while the yellow region (yolk) represents molecules predicted to cross the central nervous system (CNS).
P) and TPSA to estimate passive GI absorption and blood–brain barrier permeability. In the BOILED-Egg diagram, the white area indicates compounds with a high probability of human intestinal absorption (HIA), while the yellow region (yolk) represents molecules predicted to cross the central nervous system (CNS).
The BOILED-Egg model was additionally used to predict gastrointestinal absorption and blood–brain barrier (BBB) permeability based on lipophilicity and TPSA. As illustrated in Fig. 4, most compounds (2 and 4–10) fall within the white region of the diagram, indicating a high probability of human intestinal absorption. Interestingly, compound 3 appears in the yellow region (yolk), suggesting a potential ability to penetrate the blood–brain barrier. Such behavior may indicate possible central nervous system exposure, whereas the remaining derivatives are predicted to have limited BBB permeability, which may be advantageous for peripherally acting therapeutic agents by reducing the risk of neurological side effects.
 | ||
| Fig. 4 BOILED-Egg model illustrating the predicted gastrointestinal absorption and brain penetration potential of the synthesized candidates. | ||
The BOILED-Egg analysis also provides insight into interactions with P-glycoprotein, a key efflux transporter involved in drug absorption and distribution. Several derivatives are predicted to behave as non-substrates, indicating a reduced probability of efflux-mediated elimination and potentially improved intracellular retention following absorption.
Overall, the integrated ADME and BOILED-Egg analyses indicate that the synthesized compounds possess generally favorable pharmacokinetic characteristics, including acceptable drug-likeness and predicted oral absorption. Most derivatives exhibit limited blood–brain barrier (BBB) permeability, whereas compound 3 is predicted to penetrate the BBB. These findings suggest that careful optimization of molecular size, lipophilicity, and polarity within this scaffold may yield compounds with promising pharmacokinetic profiles suitable for further biological evaluation.
5. Molecular docking
Molecular docking simulations were conducted to evaluate the binding affinity and interaction patterns of the prepared compounds toward the epidermal growth factor receptor (EGFR) tyrosine kinase using the crystal structure PDB ID: 4HJO. The docking protocol was validated by re-docking the co-crystallized ligand into the active site of the receptor. The obtained RMSD value of 1.34 Å confirmed that the docking procedure was able to reproduce the experimentally observed binding mode with acceptable accuracy, indicating the reliability of the applied docking parameters (Fig. 5). | ||
| Fig. 5 Superimposition of the docked AQ4 ligand (green) with the original ligand (pink) showing a RMSD value of 1.34 Å and a binding score of −9.12 kcal mol−1. | ||
The anticancer drug gefitinib, a clinically approved EGFR inhibitor, was used as a reference ligand to compare the binding affinity and interaction behavior of the benzochromene derivatives. Gefitinib achieved a docking score of −8.249 kcal mol−1 and established key hydrogen bonds with LEU 694 (A) and MET 769 (A), along with hydrophobic π–H interactions within the ATP-binding pocket. These residues are essential for stabilizing inhibitors within the EGFR kinase domain.
The docking scores of the investigated compounds ranged from −5.941 to −9.067 kcal mol−1, suggesting variable but generally favorable binding affinities toward the EGFR active site. Notably, most compounds established interactions with essential residues located within the ATP-binding region, including LYS 692, LEU 694, MET 769, CYS 773, and ASP 776, which are crucial for ligand recognition and stabilization (Table 3, SI).
| Ligands | S (kcal mol−1) | rmsd_refine (Å) | Binding site | Interaction | Distance (Å) | E (kcal mol−1) | |
|---|---|---|---|---|---|---|---|
| Liganda | Receptor | ||||||
| a Repeated entries of “6-ring” represent the different ligand rings or functional groups interacting at distinct positions within the EGFR binding site. | |||||||
| 2 | −6.333 | 1.248 | N 31 | LYS 692 | H-Acceptor | 2.92 | −10.4 |
| 6-Ring | LEU 694 | Pi-H | 4.32, 4.07 | −1.0, −0.8 | |||
| 3 | −5.941 | 1.092 | O 1 | LYS 692 | H-Acceptor | 3.08 | −10.4 |
| N 5 | LEU 694 | H-Acceptor | 3.50 | −0.9 | |||
| O 1 | LYS 692 | Ionic | 3.08 | −3.9 | |||
| 6-Ring | LEU 694 | Pi-H | 4.11 | −0.6 | |||
| 4 | −6.731 | 1.686 | 6-Ring | LEU 694 | Pi-H | 4.02, 3.69 | −0.7, −1.0 |
| 6-Ring | GLY 772 | Pi-H | 4.00 | −0.7 | |||
| 5 | −6.031 | 1.850 | O 12 | LYS 692 | H-Acceptor | 2.94 | −0.8 |
| N 31 | ASP 776 | Ionic | 3.85 | −0.8 | |||
| 6-Ring | LEU 694 | Pi-H | 4.45, 3.98 | −0.6, −0.8 | |||
| 6 | −7.937 | 0.933 | O 49 | MET 769 | H-Acceptor | 3.51 | −0.6 |
| 6-Ring | LYS 692 | Pi-cation | 3.67, 4.31 | −1.1 | |||
| 6-Ring | LEU 694 | Pi-H | 3.97 | −0.9 | |||
| 7 | −9.067 | 1.747 | N 1 | ARG 817 | H-Donor | 2.74 | −3.4 |
| 5-Ring | THR 830 | Pi-H | 4.36 | −1.6 | |||
| 8 | −6.519 | 1.403 | 6-Ring | LEU 694 | Pi-H | 4.04, 3.75 | −1.0, −1.3 |
| 6-Ring | GLY 772 | Pi-H | 3.89 | −1.0 | |||
| 9 | −6.712 | 1.094 | N 30 | MET 769 | H-Acceptor | 3.55 | −1.6 |
| 6-Ring | CYS 773 | Pi-H | 3.82, 3.46 | −0.8 | |||
| 5-Ring | CYS 773 | Pi-H | 4.32 | −0.7 | |||
| 10 | −8.175 | 1.610 | N 28 | MET 769 | H-Acceptor | 3.17 | −3.7 |
| 6-Ring | LYS 692 | Pi-cation | 3.98 | −1.4 | |||
| Gefitinib reference | −8.249 | 1.383 | N 42 | LEU 694 | H-Donor | 3.20 | −5.9 |
| N 14 | MET 769 | H-Acceptor | 3.04 | −4.8 | |||
| 6-Ring | LEU 694 | Pi-H | 3.63 | −0.7 | |||
| Co-crystallized ligand | −8.674 | 2.039 | N 2 43 | MET 769 | H-Acceptor | 3.02 | −4.8 |
| 6-Ring | LEU 694 | Pi-H | 3.63 | −0.8 | |||
Among the evaluated compounds, compound 7 showed the most promising binding affinity, achieving a docking score of −9.067 kcal mol−1, surpassing that of the reference drug gefitinib. The ligand formed a strong hydrogen-bond donor interaction with ARG 817 at a distance of 2.74 Å, accompanied by a π–H interaction with THR 830. These interactions contribute significantly to stabilizing the ligand within the EGFR binding pocket and indicate a favorable binding orientation that may enhance inhibitory potential, as illustrated in Table 3 and Fig. 6. Compound 10 also demonstrated a promising binding profile, with a docking score of −8.175 kcal mol−1, comparable to that of gefitinib. The ligand established a hydrogen bond with MET 769 at 3.17 Å, together with a π–cation interaction with LYS 692 at 3.98 Å. Such interactions are frequently observed in potent EGFR inhibitors and suggest that compound 10 effectively occupies the ATP-binding site.
 | ||
| Fig. 6 Schematic of the benzochromenopyrimidine derivative 7 illustrating its 2D and 3D interactions within the active site of the EGFR tyrosine kinase. | ||
Similarly, compound 6 displayed a favorable docking score of −7.937 kcal mol−1 and a low RMSD value (0.933 Å), indicating a stable and reliable docking pose. The compound formed a hydrogen-bond interaction with MET 769 and multiple π–cation interactions with LYS 692, along with π–H interactions with LEU 694. These interactions emphasize the importance of aromatic systems in promoting hydrophobic and electrostatic contacts within the kinase active site.
Compounds 2, 3, and 5 exhibited moderate docking scores (−6.333, −5.941, and −6.031 kcal mol−1, respectively). Their binding modes were primarily stabilized through hydrogen-bond interactions with LYS 692 and π–H interactions with LEU 694. In addition, compound 3 formed an ionic interaction with LYS 692, while compound 5 showed an ionic interaction with ASP 776, suggesting that electrostatic contacts also contribute to ligand stabilization within the EGFR binding cavity.
Compounds 8 and 9 presented docking scores of −6.519 and −6.712 kcal mol−1, respectively. Their binding orientations were predominantly stabilized by hydrophobic π–H interactions with residues such as LEU 694, GLY 772, and CYS 773. Furthermore, compound 9 established a hydrogen-bond acceptor interaction with MET 769, which further enhanced the stabilization of the ligand within the active site. The interaction analysis also showed that several synthesized compounds mimic the key interactions observed for gefitinib, particularly those involving MET 769 and LEU 694, confirming that these residues are critical anchors within the EGFR ATP-binding pocket. The presence of aromatic rings and heteroatoms in the synthesized molecules appears to facilitate the formation of hydrogen bonds, π–H interactions, and π–cation contacts with these residues, thereby enhancing their binding affinity.
Importantly, while compound 7 exhibited a docking score numerically higher than that of gefitinib, the docking score alone is not sufficient to conclusively determine promising binding affinity. Therefore, additional parameters, such as RMSD values and detailed interaction profiles, were considered to support the reliability of the predicted binding modes. All reported RMSD values (<2 Å) indicate acceptable and stable docking conformations. Furthermore, several synthesized compounds mimic key interactions observed for gefitinib, particularly those involving MET 769 and LEU 694, which are critical anchors within the EGFR ATP-binding pocket. It should also be emphasized that molecular docking provides a static representation of ligand–protein interactions and does not fully account for solvent effects, protein flexibility, or binding kinetics. Consequently, further studies, such as molecular dynamics simulations and experimental kinetic validation, are required for a more comprehensive assessment of binding stability and inhibitory potential.
Overall, the docking results indicate that hydrogen bonding, hydrophobic π–interactions, and electrostatic contacts play fundamental roles in stabilizing ligand binding within the EGFR kinase domain. Among the tested compounds, 7, 10, and 6 demonstrated the most favorable binding profiles, with docking scores comparable to or better than that of the reference inhibitor gefitinib. These findings suggest that these compounds may represent promising scaffolds for further development as potential EGFR tyrosine kinase inhibitors.
6. Structure–activity relationship
The benzochromene derivatives 2–5 showed weak to moderate cytotoxic activity (IC50 > 50 µM), indicating that the basic benzochromene scaffold with simple substituents provides limited interactions within the EGFR binding pocket. Docking results suggest that these compounds mainly form weak hydrophobic π–H interactions with residues such as LEU 694.Introducing nitrogen-rich heterocyclic systems significantly enhanced activity. Compounds 6 and 7 displayed strong antiproliferative effects, likely owing to the presence of additional heteroatoms that facilitate hydrogen bonding and electrostatic interactions with key residues such as LYS 692 and MET 769. Among them, compound 7 showed the highest activity and promising binding affinity, attributed to its extended heteroaromatic system that improves both hydrogen bonding and hydrophobic stabilization within the EGFR active site. ADME predictions indicate that compounds 6 and 7 maintain moderate lipophilicity and acceptable oral bioavailability (bioavailability score = 0.55), supporting their potential as orally active leads.
In contrast, compounds 8 and 9 exhibited weak activity, possibly due to less favorable orientations of their heterocyclic moieties within the binding pocket. Compound 10 demonstrated notable activity, which may be related to the presence of the cyano group and additional heterocyclic features that enhance interactions with residues such as MET 769 and LYS 692. Despite a slightly higher log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (5.27) and molecular weight (597.62 g mol−1), its predicted oral absorption remains moderate, and limited BBB penetration may reduce potential CNS-related side effects. Overall, the results indicate that the incorporation of nitrogen-rich fused heterocycles, electron-withdrawing groups, and hydrogen-bonding functionalities enhances anticancer activity. Compounds 6, 7, and 10 emerge as the most promising candidates, balancing potent EGFR interactions with favorable pharmacokinetic profiles.
P (5.27) and molecular weight (597.62 g mol−1), its predicted oral absorption remains moderate, and limited BBB penetration may reduce potential CNS-related side effects. Overall, the results indicate that the incorporation of nitrogen-rich fused heterocycles, electron-withdrawing groups, and hydrogen-bonding functionalities enhances anticancer activity. Compounds 6, 7, and 10 emerge as the most promising candidates, balancing potent EGFR interactions with favorable pharmacokinetic profiles.
7. Conclusion
Benzochromene-based derivatives (2–10) were successfully synthesized and structurally characterized, and their antiproliferative activities were evaluated against HCT-116, MCF-7, and HepG2 cancer cell lines, as well as normal WI-38 fibroblasts. Several compounds demonstrated promising anticancer activity, with good selectivity toward cancer cells. Among them, compounds 6, 7, and 10 exhibited the most potent cytotoxic effects. Notably, compound 7 showed the highest activity, with IC50 values comparable to the reference drug doxorubicin and relatively low toxicity toward normal WI-38 cells, resulting in a favorable selectivity index. Compounds 6 and 10 also displayed significant antiproliferative activity and acceptable selectivity profiles.Molecular docking studies targeting the epidermal growth factor receptor (EGFR) tyrosine kinase suggested that the most active compounds (7, 10, and 6) may bind favorably within the EGFR ATP-binding pocket, showing docking scores comparable to that of gefitinib and forming key interactions with residues such as Lys692, Met769, and Leu694. However, these findings are based on computational predictions, and further experimental validation, such as enzymatic inhibition assays, is required to confirm their activity against EGFR. In addition, ADME and BOILED-Egg analyses indicated that these compounds possess generally favorable pharmacokinetic properties, including acceptable oral absorption, moderate lipophilicity, and limited blood–brain barrier penetration (with the exception of compound 3), supporting their potential for oral bioavailability.
Overall, the present study highlights benzochromene derivatives as promising scaffolds for anticancer drug development. Among the tested compounds, compound 7 emerges as a potential lead candidate for further optimization and biological investigation, particularly with respect to its mechanism of action and target validation.
8. Materials and methods
8.1. Chemistry
All melting points were determined using a Griffin and George melting-point apparatus (Griffin & Georgy Ltd, Wembley, Middlesex, UK). IR spectra were recorded with a Pye Unicam SP1200 spectrophotometer (Pye Unicam Ltd, Cambridge, UK) using the KBr pellet method. 1H NMR spectra were acquired with a 500 MHz JEOL NMR spectrometer with tetramethylsilane (TMS) as the internal standard (chemical shifts reported in δ), and 13C NMR spectra were recorded at 120 MHz. Elemental analyses were performed at the Microanalytical Unit, Faculty of Science, Ain Shams University, using a PerkinElmer 2400 CHN analyzer (Waltham, MA), yielding satisfactory results within ±0.4% for all compounds.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), 1648 (C
N), 1648 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). Anal. calcd. for C22H18N2O3 (358.13): C, 73.73; H, 5.06; N, 7.82. Found: C, 73.85; H, 5.22; N, 7.68.
N). Anal. calcd. for C22H18N2O3 (358.13): C, 73.73; H, 5.06; N, 7.82. Found: C, 73.85; H, 5.22; N, 7.68.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.58 (s, 3H, OCH3), 3.65 (s, 3H, OCH3), 5.94 (s, 1H, C4H-pyran), 6.75 (d, 1H, Ar–H, J = 8.5 Hz), 6.86, 6.87 (dd, 1H, Ar–H, J = 1.5, 1.5 Hz), 7.18 (brs, 2H, NH2, exchangeable with D2O), 7.24 (d, 1H, Ar–H, J = 1.5 Hz), 7.34 (s, 1H, Ar–H), 7.43 (t, 1H, Ar–H, J = 7.5 Hz), 7.50 (d, 1H, Ar–H, J = 9 Hz), 7.57 (t, 1H, Ar–H, J = 7.5 Hz), 7.91 (d, 2H, Ar–H, J = 9 Hz), 8.10 (s, 1H, Ar–H). 13C-NMR (125 MHz, DMSO-d6) δppm: 33.8, 55.3, 55.5, 97.2, 111.9, 117.6, 119.9, 123.2, 124.8, 127.0, 128.6, 129.2, 147.4, 147.7, 148.2, 156.2, 161.9, 162.4. Anal. calcd. for C23H19N3O3 (385.14): C, 71.68; H, 4.97; N, 10.90. Found: C, 71.73; H, 4.81; N, 10.79.
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.58 (s, 3H, OCH3), 3.65 (s, 3H, OCH3), 5.94 (s, 1H, C4H-pyran), 6.75 (d, 1H, Ar–H, J = 8.5 Hz), 6.86, 6.87 (dd, 1H, Ar–H, J = 1.5, 1.5 Hz), 7.18 (brs, 2H, NH2, exchangeable with D2O), 7.24 (d, 1H, Ar–H, J = 1.5 Hz), 7.34 (s, 1H, Ar–H), 7.43 (t, 1H, Ar–H, J = 7.5 Hz), 7.50 (d, 1H, Ar–H, J = 9 Hz), 7.57 (t, 1H, Ar–H, J = 7.5 Hz), 7.91 (d, 2H, Ar–H, J = 9 Hz), 8.10 (s, 1H, Ar–H). 13C-NMR (125 MHz, DMSO-d6) δppm: 33.8, 55.3, 55.5, 97.2, 111.9, 117.6, 119.9, 123.2, 124.8, 127.0, 128.6, 129.2, 147.4, 147.7, 148.2, 156.2, 161.9, 162.4. Anal. calcd. for C23H19N3O3 (385.14): C, 71.68; H, 4.97; N, 10.90. Found: C, 71.73; H, 4.81; N, 10.79.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), 1678 (C
N), 1678 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Olactone, keto form). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.66 (s, 3H, OCH3), 3.72 (s, 3H, OCH3), 5.53 (s, 1H, C4-Hpyran + 1H, OH), 6.57, 6.99 (dd, 1H, Ar–H, J = 1.5 Hz), 6.85 (d, 1H, Ar–H, J = 8 Hz), 7.08 (d, 1H, Ar–H, J = 2.5 Hz), 7.49–7.52 (m, 1H, Ar–H), 7.57 (t, 1H, Ar–H, J = 7 Hz), 7.98 (d, 1H, Ar–H, J = 9 Hz), 8.05 (d, 1H, Ar–H, J = 9 Hz). 13C-NMR (125 MHz, DMSO-d6) δppm: 40.8, 55.3, 55.4, 112.1, 117.0, 117.2, 119.3, 123.4, 125.7, 127.7, 128.6, 128.9, 129.8, 130.4, 130.8, 148.2, 148.7, 162.0. Anal. calcd. for C22H17NO4 (359.12): C, 73.53; H, 4.77; N, 3.90. Found: C, 73.44; H, 4.69; N, 3.98.
Olactone, keto form). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.66 (s, 3H, OCH3), 3.72 (s, 3H, OCH3), 5.53 (s, 1H, C4-Hpyran + 1H, OH), 6.57, 6.99 (dd, 1H, Ar–H, J = 1.5 Hz), 6.85 (d, 1H, Ar–H, J = 8 Hz), 7.08 (d, 1H, Ar–H, J = 2.5 Hz), 7.49–7.52 (m, 1H, Ar–H), 7.57 (t, 1H, Ar–H, J = 7 Hz), 7.98 (d, 1H, Ar–H, J = 9 Hz), 8.05 (d, 1H, Ar–H, J = 9 Hz). 13C-NMR (125 MHz, DMSO-d6) δppm: 40.8, 55.3, 55.4, 112.1, 117.0, 117.2, 119.3, 123.4, 125.7, 127.7, 128.6, 128.9, 129.8, 130.4, 130.8, 148.2, 148.7, 162.0. Anal. calcd. for C22H17NO4 (359.12): C, 73.53; H, 4.77; N, 3.90. Found: C, 73.44; H, 4.69; N, 3.98.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), 1652 (C
N), 1652 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 1.30 (t, 3H,
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 1.30 (t, 3H, ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif)
![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 3CH2, J = 7.5 Hz), 3.65 (s, 3H, OCH3), 3.67 (s, 3H, OCH3), 4.31 (q, 2H, CHE3
3CH2, J = 7.5 Hz), 3.65 (s, 3H, OCH3), 3.67 (s, 3H, OCH3), 4.31 (q, 2H, CHE3![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif)
![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2, J = 7 Hz), 5.50 (s, 1H, C4-Hpyran), 6.63, 6.65 (dd, 1H, Ar–H, J = 2, 1.5 Hz), 6.82 (d, 1H, Ar–H, J = 8 Hz), 6.96 (s, 1H, Ar–H), 7.41–7.46 (m, 3H, Ar–H), 7.86 (d, 1H, Ar–H, J = 8 Hz), 7.90–7.92 (m, 1H, Ar–H), 7.96 (d, 1H, Ar–H, J = 8.5 Hz), 8.70 (s, 1H, Ar–H). 13C-NMR (125 MHz, DMSO-d6) δppm: 13.9, 55.4, 55.5, 63.9, 81.3, 111.4, 112.0, 117.1, 119.7, 123.9, 125.1, 127.1, 128.5, 129.8, 131.1, 146.9, 147.8, 148.7, 156.6, 161.8. Anal. calcd. for C25H22N2O4 (414.16): C, 72.45; H, 5.35; N, 6.76. Found: C, 72.51; H, 5.29; N, 6.63.
2, J = 7 Hz), 5.50 (s, 1H, C4-Hpyran), 6.63, 6.65 (dd, 1H, Ar–H, J = 2, 1.5 Hz), 6.82 (d, 1H, Ar–H, J = 8 Hz), 6.96 (s, 1H, Ar–H), 7.41–7.46 (m, 3H, Ar–H), 7.86 (d, 1H, Ar–H, J = 8 Hz), 7.90–7.92 (m, 1H, Ar–H), 7.96 (d, 1H, Ar–H, J = 8.5 Hz), 8.70 (s, 1H, Ar–H). 13C-NMR (125 MHz, DMSO-d6) δppm: 13.9, 55.4, 55.5, 63.9, 81.3, 111.4, 112.0, 117.1, 119.7, 123.9, 125.1, 127.1, 128.5, 129.8, 131.1, 146.9, 147.8, 148.7, 156.6, 161.8. Anal. calcd. for C25H22N2O4 (414.16): C, 72.45; H, 5.35; N, 6.76. Found: C, 72.51; H, 5.29; N, 6.63.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.59 (s, 3H, OCH3), 3.67 (s, 3H, OCH3), 5.72 (brs, 2H, NH2, exchangeable with D2O), 5.58 (s, 1H, C4-Hpyran), 6.72 (d, 1H, Ar–H, J = 8 Hz), 6.84 (s, 1H, Ar–H), 7.29 (s, 1H, Ar–H), 7.34 (s, 1H, Ar–H), 7.41 (t, 1H, Ar–H, J = 7 Hz), 7.45–7.47 (m, 1H, Ar–H), 7.52 (t, 1H, Ar–H, J = 7 Hz), 7.88 (d, 2H, J = 8.5 Hz), 8.09 (s, 1H, N
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.59 (s, 3H, OCH3), 3.67 (s, 3H, OCH3), 5.72 (brs, 2H, NH2, exchangeable with D2O), 5.58 (s, 1H, C4-Hpyran), 6.72 (d, 1H, Ar–H, J = 8 Hz), 6.84 (s, 1H, Ar–H), 7.29 (s, 1H, Ar–H), 7.34 (s, 1H, Ar–H), 7.41 (t, 1H, Ar–H, J = 7 Hz), 7.45–7.47 (m, 1H, Ar–H), 7.52 (t, 1H, Ar–H, J = 7 Hz), 7.88 (d, 2H, J = 8.5 Hz), 8.09 (s, 1H, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 8.25 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 35.4, 55.3, 55.5, 99.7, 117.4, 120.4, 124.8, 126.9, 128.5, 129.1, 130.5, 130.9, 147.3, 147.4, 148.1, 150.1, 155.1. Anal. calcd. for C23H20N4O3 (400.15): C, 68.99; H, 5.03; N, 13.99. Found: C, 68.84; H, 5.17; N, 13.78.
CH), 8.25 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 35.4, 55.3, 55.5, 99.7, 117.4, 120.4, 124.8, 126.9, 128.5, 129.1, 130.5, 130.9, 147.3, 147.4, 148.1, 150.1, 155.1. Anal. calcd. for C23H20N4O3 (400.15): C, 68.99; H, 5.03; N, 13.99. Found: C, 68.84; H, 5.17; N, 13.78.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 2.18 (s, 3H, CH3), 3.63 (s, 3H, OCH3), 3.64 (s, 3H, OCH3), 5.99 (s, 1H, C4-Hpyran), 6.72–6.77 (m, 2H, Ar–H), 6.85 (s, 1H, Ar–H), 7.21 (s, 1H, Ar–H), 7.25 (d, 2H, Ar–H), 7.46 (t, 1H, Ar–H, J = 7 Hz), 7.54–7.91 (m, 2H, Ar–H), 7.94 (t, 2H, Ar–H, J = 7 Hz), 8.10 (d, 1H, ArH), 8.21 (brs, 1H, NH, exchangeable with D2O), 8.90 (s, 1H, N
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 2.18 (s, 3H, CH3), 3.63 (s, 3H, OCH3), 3.64 (s, 3H, OCH3), 5.99 (s, 1H, C4-Hpyran), 6.72–6.77 (m, 2H, Ar–H), 6.85 (s, 1H, Ar–H), 7.21 (s, 1H, Ar–H), 7.25 (d, 2H, Ar–H), 7.46 (t, 1H, Ar–H, J = 7 Hz), 7.54–7.91 (m, 2H, Ar–H), 7.94 (t, 2H, Ar–H, J = 7 Hz), 8.10 (d, 1H, ArH), 8.21 (brs, 1H, NH, exchangeable with D2O), 8.90 (s, 1H, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 8.99 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 20.6, 33.8, 55.3, 55.4, 98.5, 111.5, 111.8, 116.5, 117.3, 119.8, 123.1, 125.4, 125.8, 127.4, 128.7, 129.6, 129.9, 134.3, 140.9, 146.7, 152.4, 158.4. Anal. calcd. for C30H26N4O5S (554.16): C, 64.97; H, 4.73; N, 10.10; S, 5.78. Found: C, 64.86; H, 4.61; N, 10.23; S, 5.59.
CH), 8.99 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 20.6, 33.8, 55.3, 55.4, 98.5, 111.5, 111.8, 116.5, 117.3, 119.8, 123.1, 125.4, 125.8, 127.4, 128.7, 129.6, 129.9, 134.3, 140.9, 146.7, 152.4, 158.4. Anal. calcd. for C30H26N4O5S (554.16): C, 64.97; H, 4.73; N, 10.10; S, 5.78. Found: C, 64.86; H, 4.61; N, 10.23; S, 5.59.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 1.86 (s, 3H, CH3), 3.08 (s, 3H, NCH3), 3.58 (s, 3H, OCH3), 3.66 (s, 3H, OCH3), 6.18 (s, 1H, C4-Hpyran), 6.73 (s, 2H, Ar–H), 7.34 (t, 1H, Ar–H, J = 7 Hz), 7.42–7.45 (m, 4H, Ar–H), 7.51–7.55 (m, 4H, Ar–H), 7.93 (t, 2H, Ar–H, J = 8 Hz), 8.14 (d, 1H, ArH, J = 6.5 Hz), 8.68 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 10.2, 33.4, 55.3, 55.5, 98.8, 112.1, 117.6, 119.7, 123.7, 129.1, 129.2, 130.6, 130.8, 147.3, 147.7, 148.1, 153.3, 155.8, 160.6, 161.7, 162.5. Anal. calcd. for C34H29N5O4 (571.22): C, 71.44; H, 5.11; N, 12.25. Found: C, 71.66; H, 5.07; N, 12.11.
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 1.86 (s, 3H, CH3), 3.08 (s, 3H, NCH3), 3.58 (s, 3H, OCH3), 3.66 (s, 3H, OCH3), 6.18 (s, 1H, C4-Hpyran), 6.73 (s, 2H, Ar–H), 7.34 (t, 1H, Ar–H, J = 7 Hz), 7.42–7.45 (m, 4H, Ar–H), 7.51–7.55 (m, 4H, Ar–H), 7.93 (t, 2H, Ar–H, J = 8 Hz), 8.14 (d, 1H, ArH, J = 6.5 Hz), 8.68 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 10.2, 33.4, 55.3, 55.5, 98.8, 112.1, 117.6, 119.7, 123.7, 129.1, 129.2, 130.6, 130.8, 147.3, 147.7, 148.1, 153.3, 155.8, 160.6, 161.7, 162.5. Anal. calcd. for C34H29N5O4 (571.22): C, 71.44; H, 5.11; N, 12.25. Found: C, 71.66; H, 5.07; N, 12.11.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1636 (C
O), 1636 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.57 (s, 3H, OCH3), 3.66 (s, 3H, OCH3), 6.07 (s, 1H, C4-Hpyran), 6.71 (d, 2H, Ar–H, J = 8 Hz), 7.19 (s, 1H, Ar–H), 7.42 (t, 1H, Ar–H, J = 7 Hz), 7.48–7.51 (m, 1H, Ar–H), 7.55 (d, 1H, Ar–H, J = 9 Hz), 7.92 (d, 1H, ArH, J = 8 Hz), 7.97 (d, 1H, ArH, J = 9 Hz), 8.01 (d, 1H, ArH, J = 8 Hz), 9.25 (s, 1H, ArH), 12.64 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 36.3, 55.3, 55.5, 100.8, 111.7, 112.2, 117.5, 120.0, 125.0, 127.3, 128.6, 129.8, 147.7, 147.9, 148.4, 153.1, 139.9. Anal. calcd. for C24H18N4O4 (426.13): C, 67.60; H, 4.25; N, 13.14. Found: C, 67.88; H, 4.11; N, 13.29.
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.57 (s, 3H, OCH3), 3.66 (s, 3H, OCH3), 6.07 (s, 1H, C4-Hpyran), 6.71 (d, 2H, Ar–H, J = 8 Hz), 7.19 (s, 1H, Ar–H), 7.42 (t, 1H, Ar–H, J = 7 Hz), 7.48–7.51 (m, 1H, Ar–H), 7.55 (d, 1H, Ar–H, J = 9 Hz), 7.92 (d, 1H, ArH, J = 8 Hz), 7.97 (d, 1H, ArH, J = 9 Hz), 8.01 (d, 1H, ArH, J = 8 Hz), 9.25 (s, 1H, ArH), 12.64 (brs, 1H, NH, exchangeable with D2O). 13C-NMR (120 MHz, DMSO-d6) δppm: 36.3, 55.3, 55.5, 100.8, 111.7, 112.2, 117.5, 120.0, 125.0, 127.3, 128.6, 129.8, 147.7, 147.9, 148.4, 153.1, 139.9. Anal. calcd. for C24H18N4O4 (426.13): C, 67.60; H, 4.25; N, 13.14. Found: C, 67.88; H, 4.11; N, 13.29.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), 1635 (C
N), 1635 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.54 (s, 3H, OCH3), 3.55 (s, 3H, OCH3), 4.52 (s, 2H,
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.54 (s, 3H, OCH3), 3.55 (s, 3H, OCH3), 4.52 (s, 2H, ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif)
![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2CN), 6.20 (s, 1H, C4-Hpyran), 6.48, 6.50 (dd, 1H, Ar–H, J = 2 Hz), 6.66 (d, 1H, Ar–H, J = 8 Hz), 7.40–7.48 (m, 3H, Ar–H), 7.57 (d, 1H, Ar–H, J = 9 Hz), 7.92 (d, 1H, ArH, J = 8 Hz), 7.98 (d, 1H, ArH, J = 9 Hz), 8.02 (d, 1H, ArH, J = 8 Hz), 9.60 (s, 1H, ArH). 13C-NMR (125 MHz, DMSO-d6) δppm: 18.0, 36.4, 55.2, 55.4, 102.6, 111.7, 112.4, 114.7, 116.6, 117.4, 119.6, 123.7, 125.1, 127.3, 128.5, 129.9, 131.1, 135.3, 140.0, 147.5, 147.9, 148.3, 160.7. Anal. calcd. for C26H19N5O3 (449.15): C, 69.48; H, 4.26; N, 15.58. Found: C, 69.55; H, 4.37; N, 15.41.
2CN), 6.20 (s, 1H, C4-Hpyran), 6.48, 6.50 (dd, 1H, Ar–H, J = 2 Hz), 6.66 (d, 1H, Ar–H, J = 8 Hz), 7.40–7.48 (m, 3H, Ar–H), 7.57 (d, 1H, Ar–H, J = 9 Hz), 7.92 (d, 1H, ArH, J = 8 Hz), 7.98 (d, 1H, ArH, J = 9 Hz), 8.02 (d, 1H, ArH, J = 8 Hz), 9.60 (s, 1H, ArH). 13C-NMR (125 MHz, DMSO-d6) δppm: 18.0, 36.4, 55.2, 55.4, 102.6, 111.7, 112.4, 114.7, 116.6, 117.4, 119.6, 123.7, 125.1, 127.3, 128.5, 129.9, 131.1, 135.3, 140.0, 147.5, 147.9, 148.3, 160.7. Anal. calcd. for C26H19N5O3 (449.15): C, 69.48; H, 4.26; N, 15.58. Found: C, 69.55; H, 4.37; N, 15.41.![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), 1635 (C
N), 1635 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.35 (s, 3H, OCH3), 3.53 (s, 3H, OCH3), 3.81 (s, 3H, OCH3), 3.82 (s, 3H, OCH3), 6.20 (s, 1H, C4-Hpyran), 6.41, 6.42 (dd, 1H, Ar–H, J = 1.5, 1.5 Hz), 6.65 (d, 1H, Ar–H, J = 9 Hz), 7.06 (d, 1H, Ar–H, J = 9 Hz), 7.38–7.41 (m, 1H, Ar–H), 7.46 (t, 1H, ArH, J = 7 Hz), 7.56 (d, 1H, ArH, J = 10 Hz), 7.64 (d, 1H, ArH, J = 2.5 Hz), 7.73 (d, 1H, ArH, J = 2 Hz), 7.90–7.93 (m, 2H, ArH), 7.98 (d, 1H, ArH, J = 9.5 Hz), 8.47 (s, 1H,
N). 1H-NMR (500 MHz, DMSO-d6) δppm: 3.35 (s, 3H, OCH3), 3.53 (s, 3H, OCH3), 3.81 (s, 3H, OCH3), 3.82 (s, 3H, OCH3), 6.20 (s, 1H, C4-Hpyran), 6.41, 6.42 (dd, 1H, Ar–H, J = 1.5, 1.5 Hz), 6.65 (d, 1H, Ar–H, J = 9 Hz), 7.06 (d, 1H, Ar–H, J = 9 Hz), 7.38–7.41 (m, 1H, Ar–H), 7.46 (t, 1H, ArH, J = 7 Hz), 7.56 (d, 1H, ArH, J = 10 Hz), 7.64 (d, 1H, ArH, J = 2.5 Hz), 7.73 (d, 1H, ArH, J = 2 Hz), 7.90–7.93 (m, 2H, ArH), 7.98 (d, 1H, ArH, J = 9.5 Hz), 8.47 (s, 1H, ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NCH), 9.53 (s, 1H,
NCH), 9.53 (s, 1H, ![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH). 13C-NMR (120 MHz, DMSO-d6) δppm: 36.5, 55.2, 55.3, 55.4, 55.7, 97.1, 102.3, 111.8, 117.4, 125.1, 125.5, 130.4, 131.1, 135.2, 140.1, 148.1, 148.2, 148.7, 162.6. Anal. calcd. for C35H27N5O5 (597.20): C, 70.34; H, 4.55; N, 11.72. Found: C, 70.47; H, 4.33; N, 11.68.
CH). 13C-NMR (120 MHz, DMSO-d6) δppm: 36.5, 55.2, 55.3, 55.4, 55.7, 97.1, 102.3, 111.8, 117.4, 125.1, 125.5, 130.4, 131.1, 135.2, 140.1, 148.1, 148.2, 148.7, 162.6. Anal. calcd. for C35H27N5O5 (597.20): C, 70.34; H, 4.55; N, 11.72. Found: C, 70.47; H, 4.33; N, 11.68.8.2. In Vitro cytotoxic activity
Cytotoxic effects of the synthesized compounds were evaluated using three human cancer cell lines, breast adenocarcinoma (MCF-7), colon carcinoma (HCT-116), and hepatocellular carcinoma (HepG2), along with normal human fibroblasts (WI-38). All cell lines were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in RPMI-1640 medium supplemented with 10% heat-inactivated fetal bovine serum, 100 U mL−1 penicillin, and 100 µg mL−1 streptomycin, maintained at 37 °C in a humidified atmosphere containing 5% CO2.Exponentially growing cells were detached using trypsin, counted, and seeded into 96-well plates at a density of approximately 2000 cells per well. Following a 24 hours incubation to ensure proper attachment, cells were treated with the synthesized compounds at concentrations ranging from 0.5 to 100 µM for 48 hours, while untreated cells served as controls.
Cell viability was determined using the MTT assay with full procedural details to ensure reproducibility. After treatment, the medium was removed, and 200 µL of MTT solution (5 mg mL−1) was added to each well, followed by a 4 hours incubation at 37 °C to allow formazan crystal formation. The MTT solution was then discarded, and the crystals were solubilized in DMSO for 30 minutes with continuous shaking at room temperature, protected from light, using a MaxQ 2000 plate shaker (Thermo Fisher Scientific, MA, USA). Absorbance was recorded at 570 nm using an ELISA plate reader. Cell viability was expressed as a percentage relative to the control group, and IC50 values (the concentration required to inhibit 50% of cell proliferation) were calculated.
8.3. In silico studies
Author contributions
Data curation: MHH, FSMA and YMA. Formal analysis: MHH, YMA and AYAZ. Investigation: MHH, FSMA, AYAZ and SA. Writing-original draft: MHH and FSMA. Software: MHH, AYAZ and SA. Methodology: MHH, FSMA and YMA. Validation: MHH, AYAZ and SA. Writing-review and editing: MHH. All authors read and approved the final version of the manuscript.Conflicts of interest
The authors declare no conflicts of interest.Data availability
All data generated or analyzed during this study are included in the published article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra02423h.References
- B. Katta, C. Vijayakumar, S. Dutta, B. Dubashi and V. Prasad, The incidence and severity of patient-reported side effects of chemotherapy in routine clinical care: a prospective observational study, Cureus, 2023, 15(4), e38301, DOI:10.7759/cureus.38301.
- A. Silva, S. Martins and F. Pinto-Ribeiro, Chromene-based compounds as drug candidates for renal and bladder cancer therapy: a systematic review, Bioorg. Chem., 2024, 153, 107865 CrossRef PubMed.
- G. Mancuso, A. Midiri, E. Gerace and C. Biondo, Bacterial antibiotic resistance: the most critical pathogens, Pathogens, 2021, 10(10), 1310, DOI:10.3390/pathogens10101310.
- D. G. J. Larsson, Antibiotic resistance in the environment, Nat. Rev. Microbiol., 2022, 20, 257–269 CrossRef CAS PubMed.
- O. Gustavo, et al., Evidence that the anti-inflammatory effect of 4-aryl-4H-chromenes is linked to macrophage repolarization, Fundam. Clin. Pharmacol., 2022, 6, 1020–1030, DOI:10.1111/fcp.12809.
- H. M. Mohamed, A comprehensive review on chromene derivatives: potent anticancer drug development and therapeutic potential, Orient. J. Chem., 2025, 584–608 CAS.
- D. Kumar, V. B. Reddy, S. Sharad, U. Dube and S. Kapur, A facile one-pot green synthesis and antibacterial activity of tetrahydro-4H-chromenes, Eur. J. Med. Chem., 2009, 44, 3805–3809 Search PubMed.
- T. A. Johnson, et al., Synthesis, antioxidant activity, DNA interaction, electrochemical and spectroscopic properties of chromene-based Schiff bases: experimental and theoretical approach, J. Mol. Struct., 2024, 1307, 138020 CrossRef CAS.
- O. J. Jesumoroti, D. Mnkandhla, M. Isaacs, H. C. Hoppe and R. Klein, Evaluation of novel N′-(3-hydroxybenzoyl)-2-oxo-2H-chromene-3-carbohydrazide derivatives as potential HIV-1 integrase inhibitors, Med. Chem. Commun., 2019, 10, 80–88, 10.1039/c8md00328a.
- S. Gwon, et al., Anti-allergic effect of 5,7-dihydroxy-4-methylcoumarin in IgE-mediated RBL-2H3 cells and PCA murine model, Appl. Biol. Chem., 2025, 68(4) DOI:10.1186/s13765-025-00980-4.
- S. Cardoso, et al., Synthesis and anti-leishmanial activities of uniflorol analogues, Med. Chem. Res., 2024, 33, 1657–1670 Search PubMed.
- O. Udomsin, T. Juengwatanatrakul, G. Yusakul and W. Putalun, Chromene stability: the most potent estrogenic compounds in White Kwao Krua (Pueraria candollei var. mirifica) crude extract, J. Funct. Foods, 2015, 19, 269–277 CrossRef CAS.
- M. W. Pertino, et al., Exploring benzo[h]chromene derivatives as agents against protozoal and mycobacterial infections, Pharmaceuticals, 2024, 17(10), 1375, DOI:10.3390/ph17101375.
- Y. Hou, Ageing as a risk factor for neurological disease, Nat. Rev. Neurol., 2019, 10, 565–581, DOI:10.1038/s41582-019-0244-7.
- M. S. Fernandopulle, J. Lippincott-Schwartz and M. E. Ward, RNA transport and local translation in neurodevelopmental and neurodegenerative disease, Nat. Neurosci., 2021, 24, 622–632 CrossRef CAS PubMed.
- H. Madhav and E. Jameel, Recent advancements in chromone as a privileged scaffold toward the development of small molecules, MedChemComm, 2022, 13, 258–279 RSC.
- Q. T. L. He, Z. H. W. Liu, S. Z. Lin, L. H. Yang and L. Guan, Design, synthesis, and antidepressant/anticonvulsant activities of 3H-benzo[f]chromen chalcone derivatives, Med. Chem. Res., 2021, 30, 217–224 CrossRef.
- N. R. Kamdar, D. D. Haveliwala, P. T. Mistry and S. K. Patel, Design, synthesis and in vitro evaluation of antitubercular and antimicrobial activity of novel pyranopyrimidines, Eur. J. Med. Chem., 2010, 45, 5056–5063 Search PubMed.
- W. Kemnitzer, et al., Discovery of 4-aryl-4H-chromenes as a new series of apoptosis inducers using a cell- and caspase-based high-throughput screening assay. Structure–activity relationships of the 4-aryl group, J. Med. Chem., 2004, 47, 6299–6310 CrossRef CAS PubMed.
- M. Hanifeh, et al., Synthesis, characterization, antiproliferative properties and DNA binding of benzochromene derivatives: increased Bax/Bcl-2 ratio and caspase-dependent apoptosis in colorectal cancer cell line, Bioorg. Chem., 2019, 93, 103329 Search PubMed.
- J. M. Doshi, D. Tian and C. Xing, Structure–activity relationship studies of ethyl antagonists for antiapoptotic Bcl-2 proteins to overcome drug resistance in cancer, J. Med. Chem., 2006, 49, 7731–7739 CrossRef CAS PubMed.
- J. Wang, et al., Structure-based discovery of an organic compound that binds Bcl-2 protein and induces apoptosis of tumor cells, Proc. Natl. Acad. Sci. U. S. A., 2000, 97, 7124–7129, DOI:10.1073/pnas.97.13.7124.
- N. Gao, Y. Dai, M. Rahmani, P. Dent and S. Grant, Contribution of disruption of the NF-κB pathway to induction of apoptosis in human leukemia cells by histone deacetylase inhibitors and flavopiridol, Mol. Pharmacol., 2014, 66, 956–963 CrossRef PubMed.
- A. M. Fouda, et al., Microwave synthesis and biological evaluation of halogenated 3-amino-1H-benzo[f]chromene derivatives targeting dual inhibition of topoisomerase II and microtubules, Bioorg. Chem., 2020, 95, 103549 CrossRef CAS PubMed.
- M. El-Sayed, et al., Synthesis and evaluation of antitumor activity of 9-methoxy-1H-benzo[f]chromene derivatives, Bioorg. Chem., 2021, 116, 105402 CrossRef PubMed.
- V. Raj and J. Lee, 2H/4H-Chromenes: a versatile biologically attractive scaffold, Front. Chem., 2020, 8, 623, DOI:10.3389/fchem.2020.00623.
- W. Kemnitzer, et al., Discovery of 4-aryl-4H-chromenes as apoptosis inducers: structure–activity relationships of the 5-, 6-, 7- and 8-positions, Bioorg. Med. Chem. Lett., 2005, 15, 4745–4751, DOI:10.1016/j.bmcl.2005.07.066.
- D. D. Miller, Chromenes: potential new chemotherapeutic agents for cancer, Future Med. Chem., 2013, 14, 1647–1660 Search PubMed.
- A. Kulshrestha, G. K. Katara, S. A. Ibrahim and R. Patil, Microtubule inhibitor SP-6-27 inhibits angiogenesis and induces apoptosis in ovarian cancer cells, Oncotarget, 2017, 8, 67017–67028 CrossRef PubMed.
- S. Wang, et al., FDA-approved pyrimidine-fused bicyclic heterocycles for cancer therapy: synthesis and clinical application, Eur. J. Med. Chem., 2021, 214, 113218 CrossRef CAS PubMed.
- O. Afzal, M. Yusuf, M. J. Ahsan and A. S. A. Altamimi, Chemical modification of curcumin into semi-synthetic analogs bearing pyrimidinone moiety as anticancer agents, Plants, 2022, 11(20), 2737, DOI:10.3390/plants11202737.
- K. Byrne, et al., Development of novel anticancer pyrazolopyrimidinones targeting glioblastoma, ChemMedChem, 2025, 20, e202500337, DOI:10.1002/cmdc.202500337.
- P. Seboletswe, P. Awolade and P. Singh, Recent developments on the synthesis and biological activities of fused pyrimidinone derivatives, ChemMedChem, 2021, 16, 2050–2067, DOI:10.1002/cmdc.202100083.
- A. T. Ali and M. H. Hekal, Convenient synthesis and anti-proliferative activity of some benzochromenes and chromenotriazolopyrimidines under classical methods and phase transfer catalysis, Synth. Commun., 2019, 49, 3498–3509 CrossRef CAS.
- M. R. Mahmoud, F. S. M. Abu El-Azm, M. F. Ismail, M. H. Hekal and Y. M. Ali, Synthesis and antitumor evaluation of novel tetrahydrobenzo[4′,5′]thieno[3′,2′:5,6]pyrimi-do[1,2-b] isoquinoline derivatives, Synth. Commun., 2018, 48, 428–438 CrossRef CAS.
- H. Ghonim, et al., Synthesis, SAR, and in silico studies of benzochromene derivatives as insecticidal agents against Culex pipiens, Sci. Rep., 2025, 15(1), 43485, DOI:10.1038/s41598-025-30027-z.
- M. R. Mahmoud, W. S. I. Abou-Elmagd, H. A. Derbala and M. H. Hekal, Synthesis and spectral characterization of phthalazinone derivatives, J. Chem. Res., 2012, 36, 75, DOI:10.3184/174751912X13274297624330.
- M. Kamal, et al., Synthesis and in silico studies of thiophene-isoquinolinone hybrids as potential larvicides against Culex pipiens, Sci. Rep., 2025, 15, 28031, DOI:10.1038/s41598-025-13063-7.
- M. R. Mahmoud, W. S. I. Abou-Elmagd, M. M. El-Shahawi and M. H. Hekal, Novel Fused and Spiro Heterocyclic Compounds derived from 4-(4-amino-5-mercapto-4H-1,2,4-triazol-3-yl)phthalazin-1(2H)-one, Eur. Chem. Bull., 2014, 3(7), 723–728 Search PubMed.
- M. R. Mahmoud, M. M. El-Shahawi, W. S. I. Abou-Elmagd and M. H. Hekal, Novel synthesis of isoquinoline derivatives derived from (Z)-4-(1,3-diphenylpyrazol-4-yl)isochromene-1,3-dione, Synth. Commun., 2015, 45, 1632–1641, DOI:10.1080/00397911.2015.1030760.
- A. A. Abdalha and M. H. Hekal, An efficient synthesis and evaluation of some novel quinazolinone-pyrazole hybrids as potential antiproliferative agents, Synth. Commun., 2021, 51, 2498–2509 CrossRef CAS.
- N. A. Hamed, M. I. Marzouk, M. F. Ismail and M. H. Hekal, N′-(1-([1,1′-Biphenyl]-4-yl)ethylidene)-2-cyanoacetohydrazide as scaffold for heterocyclic compounds with antitumor and antimicrobial activity, Synth. Commun., 2019, 49, 3017–3029 CAS.
- M. H. Hekal, S. S. Samir, Y. M. Ali and W. M. El-Sayed, New Benzochromeno[2,3-d]Pyrimidines and Benzochromenotriazolo[1,5-c]Pyrimidines as Potential Inhibitors of Topoisomerase II, Polycycl. Aromat. Compd., 2022, 42, 7644–7660 CrossRef CAS.
- M. H. Hekal, et al., Rational design and synthesis of new pyrrolone candidates as prospective insecticidal agents against Culex pipiens L. Larvae, Sci. Rep., 2024, 14, 24467 CrossRef CAS PubMed.
- M. H. Hekal, F. S. M. Abu El-Azm and H. A. Sallam, Synthesis, Spectral Characterization, and In Vitro Biological Evaluation of Some Novel Isoquinolinone-based Heterocycles as Potential Antitumor Agents, J. Heterocycl. Chem., 2019, 56, 795–803 CrossRef CAS.
- M. H. Hekal, et al., New 1,3,4-thiadiazoles as potential anticancer agents: pro-apoptotic, cell cycle arrest, molecular modelling, and ADMET profile, RSC Adv., 2023, 13, 15810–15825 RSC.
- M. H. Hekal, F. S. M. Abu El-Azm and S. S. Samir, An efficient approach for the synthesis and antimicrobial evaluation of some new benzocoumarins and related compounds, Synth. Commun., 2021, 51, 2175–2186 CrossRef CAS.
- M. H. Hekal, A. M. El-Naggar, F. S. M. Abu El-Azm and W. M. El-Sayed, Synthesis of new oxadiazol-phthalazinone derivatives with anti-proliferative activity; molecular docking, pro-apoptotic, and enzyme inhibition profile, RSC Adv., 2020, 10, 3675–3688, 10.1039/C9RA09016A.
- M. H. Hekal and F. S. M. Abu El-Azm, Efficient MW-Assisted Synthesis of Some New Isoquinolinone Derivatives with In Vitro Antitumor Activity, J. Heterocycl. Chem., 2017, 54, 3056–3064, DOI:10.1002/JHET.2916.
- D. S. A. Haneen, M. H. Hekal, W. S. I. Abou-Elmagd and W. M. El-Sayed, Novel pyrano[2,3-c]pyrazolopyrimidines as promising anticancer agents: design, synthesis, and cell cycle arrest of HepG2 cells at S phase, Synth. Commun., 2024, 54, 655–671, DOI:10.1080/00397911.2024.2327047.
- M. H. Hekal, Y. M. Ali, D. R. Abdel-Haleem and F. S. M. Abu El-Azm, Diversity oriented synthesis and SAR studies of new quinazolinones and related compounds as insecticidal agents against Culex pipiens L. Larvae and associated predator, Bioorg. Chem., 2023, 133, 106436, DOI:10.1016/j.bioorg.2023.106436.
- M. H. Hekal, F. S. M. Abu El-Azm and S. R. Atta-Allah, Ecofriendly and highly efficient microwave-induced synthesis of novel quinazolinone-undecyl hybrids with in vitro antitumor activity, Synth. Commun., 2019, 49, 2630–2641 CrossRef CAS.
- N. Abdelgawad, M. F. Ismail, M. H. Hekal and M. I. Marzouk, Design, Synthesis, and Evaluation of Some Novel Heterocycles Bearing Pyrazole Moiety as Potential Anticancer Agents, J. Heterocycl. Chem., 2019, 56, 1771–1779 CrossRef CAS.
- M. H. Hekal, P. S. Farag, M. M. Hemdan and W. M. El-Sayed, New N-(1,3,4-thiadiazol-2-yl)furan-2-carboxamide derivatives as potential inhibitors of the VEGFR-2, Bioorg. Chem., 2021, 115, 105176, DOI:10.1016/j.bioorg.2021.105176.
- M. R. Mahmoud, W. S. I. Abou-Elmagd, H. A. Derbala and M. H. Hekal, Novel Synthesis of Some Phthalazinone Derivatives, Chin. J. Chem., 2011, 29, 1446–1450 CrossRef CAS.
- F. S. M. Abu El-Azm, A. T. Ali and M. H. Hekal, Facile Synthesis and Anticancer Activity of Novel 4-Aminothieno[2,3-d]pyrimidines and Triazolothienopyrimidines, Org. Prep. Proced. Int., 2019, 51, 507–520, DOI:10.1080/00304948.2019.1666635.
- M. H. Hekal and F. S. M. Abu El-Azm, New potential antitumor quinazolinones derived from dynamic 2-undecyl benzoxazinone: synthesis and cytotoxic evaluation, Synth. Commun., 2018, 48, 2391–2402 CrossRef CAS.
- M. H. Hekal, A. A. Abdalha, H. Farag and A. T. Ali, Synthesis, DFT calculations, and biological studies of new 2-cyano-3-(naphthalene-1-yl)acryloyl amide analogues as anticancer agents, Chem. Biodivers., 2024, e202401023, DOI:10.1002/cbdv.202401023.
- A. I. Hassaballah, et al., New pyrazolo[3,4-d]pyrimidine derivatives as EGFR-TK inhibitors: design, green synthesis, potential anti-proliferative activity and P-glycoprotein inhibition, RSC Adv., 2024, 14, 1995–2015 RSC.
- M. H. Hekal, F. S. M. Abu El-Azm, S. S. Samir, A. Y. Alzahrani and Y. M. Ali, Synthesis, biological evaluation, and in silico studies of new pyrrolone derivatives as potent antiproliferative agents, J. Mol. Struct., 2026, 1355, 144970 CrossRef CAS.
- M. H. Hekal, Y. M. Ali and F. S. M. Abu El-Azm, Utilization of cyanoacetohydrazide and 2-(1,3-dioxoisoindolin-2-yl) acetyl chloride in the synthesis of some novel anti-proliferative heterocyclic compounds, Synth. Commun., 2020, 50, 2839–2852 CrossRef CAS.
- F. S. Abu El-Azm, M. R. Mahmoud and M. H. Hekal, Recent developments in chemistry of phthalazines, Org. Chem. Curr. Res., 2014, 4(1), 1–12 Search PubMed.
- H. Sharma, M. Mourya, L. K. Soni, D. Guin, Y. C. Joshi, M. P. Dobhal and A. K. Basak, Iodine mediated synthesis of coumarins from chromenes, Tetrahedron Lett., 2015, 56, 7100–7104 CrossRef CAS.
- M. D. Hanwell, et al., Avogadro: an advanced semantic chemical editor, visualization, and analysis platform, J. Cheminf., 2012, 4, 1–17 Search PubMed.
- R. Huey, G. M. Morris and S. Forli, et al., Using AutoDock 4 and AutoDock Vina with autodocktools: a tutorial, Scripps Res. Inst. Mol. Graph Lab., 2012, 10550, 1000 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |