Open Access Article
Open Access ArticleA QbD-optimized magnetic MOF-chitosan nanocarrier for targeted curcumin delivery and synergistic antimicrobial activity
Donia Hemidaa,
Nabila Shehata b,
Nada Elgiddawya,
Haifa E. Alfassamc,
Samar M. Mahgoubd and
Rehab Mahmoud
b,
Nada Elgiddawya,
Haifa E. Alfassamc,
Samar M. Mahgoubd and
Rehab Mahmoud *ef
*ef
aDepartment of Biotechnology and Life Sciences, Faculty of Postgraduate Studies for Advanced Sciences (PSAS), Beni-Suef University, Beni-Suef, 62511, Egypt. E-mail: doniahazem55@gmail.com; N.giddawy@psas.bsu.edu.eg
bEnvironmental Science and Industrial Development Department, Faculty of Postgraduate Studies for Advanced Sciences, Beni-Suef University, Beni-Suef, Egypt. E-mail: nabila.shehata@psas.bsu.edu.eg
cDepartment of Biology, College of Science, Princess Nourah bint Abdulrahman University, P. O. BOX 84428, Riyadh 11671, Saudi Arabia
dMaterials Science and Nanotechnology Department, Faculty of Postgraduate Studies for Advanced Sciences, Beni-Suef University, Egypt. E-mail: Miramar15@yahoo.com
eChemistry Department, Faculty of Science, Beni-Suef University, Beni-Suef 62511, Egypt. E-mail: rehabkhaled@science.bsu.edu.eg
fDepartment of Chemistry, Faculty of Science, Chulalongkorn University, Bangkok 10330, Thailand
First published on 13th May 2026
Abstract
Antimicrobial resistance and cancer represent critical global health challenges that demand smarter, targeted drug delivery strategies. This study reports the systematic design, Quality by Design (QbD)-driven optimization, and comprehensive characterization of curcumin-loaded magnetic metal–organic framework–chitosan nanocomposites (Cur/Fe3O4@ZIF-8@CS) as a multifunctional nanoplatform integrating pH-responsive drug release, magnetic targeting, and broad-spectrum antimicrobial activity. A Box–Behnken experimental design was employed to optimize three critical formulation variables including MOF-to-drug ratio (X1), chitosan concentration (X2), and iron oxide content (X3) against four predefined Critical Quality Attributes (CQAs): particle size, zeta potential, PDI, and encapsulation efficiency. The optimized nanocomposite achieved a particle size of 228.6 ± 4.7 nm, a zeta potential of +31.5 mV, a PDI of 0.218, and an encapsulation efficiency of 87.4 ± 2.3%. Successful step-by-step assembly was confirmed by DLS, ATR-FTIR of all individual components and the final composite, TEM with size-distribution mapping, and VSM magnetometry. Drug release was sustained and pH-dependent, reaching 63.8% at pH 5.5 versus 58.4% at pH 7.4 after 72 hours, following anomalous non-Fickian transport kinetics (Korsmeyer–Peppas n = 0.61). The nanocomposite exhibited potent antimicrobial activity with MIC values up to 66-fold lower than free curcumin against Escherichia coli, and 2.5- to 45-fold lower across the remaining tested organisms. A two-month physicochemical stability study conducted under long-term (25 ± 2 °C) and accelerated (40 ± 2 °C/75 ± 5% RH, ICH Q1A(R2)) conditions confirmed that all critical quality attributes remained within acceptable pharmaceutical limits, with curcumin retention of 94.6 ± 1.8% at 25 °C and 87.9 ± 2.6% at 40 °C after two months. Cytotoxicity assessment by MTT assay on L929 mouse fibroblast cells yielded IC50 values of 206.65 µg mL−1 for the blank nanocomposite (Fe3O4@ZIF-8@CS) and 273.15 µg mL−1 for the curcumin-loaded formulation (Cur/Fe3O4@ZIF-8@CS), confirming acceptable biocompatibility at therapeutically relevant concentrations. These findings collectively establish Cur/Fe3O4@ZIF-8@CS as a rationally designed, multifunctional nanoplatform with significant potential for combined antimicrobial and anticancer targeted therapy.
1 Introduction
Antimicrobial resistance (AMR) and cancer are two of the most serious health problems in the world today.1 The World Health Organization reported that resistant bacterial infections caused about 1.27 million deaths in 2019, and this number is expected to rise sharply in the coming decades.2,3 At the same time, cancer causes close to 10 million deaths each year globally. Both diseases share a common challenge: many existing drugs either do not work well enough, cause harmful side effects, or lose their effectiveness over time because bacteria or cancer cells evolve to evade them.4 These problems have prompted researchers to seek smarter, more targeted methods for drug delivery.Nanotechnology offers a promising solution. By carrying drugs inside very small particles, scientists can protect the drug from breaking down too early, deliver it more precisely to the diseased site, and release it gradually so that it keeps working for longer.5–7 One particularly interesting type of nanocarrier is the metal–organic framework (MOF). MOFs are tiny porous crystals built from metal ions connected by organic linkers, forming a highly ordered structure full of small channels and pores.8 This large internal surface area allows MOFs to hold a high amount of drug inside them. One well-studied MOF is ZIF-8, which is made from zinc ions and 2-methylimidazole. ZIF-8 has an important feature: it starts to break apart at acidic pH, which means it can release its drug cargo preferentially in acidic environments like tumors or infected tissue.9–12
Iron oxide magnetic nanoparticles (Fe3O4), also called SPIONs, are another useful component. These particles respond to external magnetic fields, which makes it possible to guide them to a specific site in the body. They also do not retain any magnetism once the magnetic field is removed, so they do not clump together in the bloodstream.13,14 When iron oxide particles are combined with MOFs, the resulting hybrid carrier can both hold a large amount of drug and be steered to the target site.15
Chitosan is a natural sugar-based polymer made from the shells of shrimp and other crustaceans. It is safe, biodegradable, and sticks well to biological surfaces. Its positive charge at acidic pH allows it to disrupt bacterial cell membranes and helps the nanoparticle bind to cells.16 Coating a nanoparticle with chitosan, therefore adds its own antibacterial activity on top of whatever drug is being delivered.17–19
Curcumin is the yellow pigment found in turmeric. It has been widely studied for its ability to fight bacteria, reduce inflammation, and slow the growth of cancer cells.20 However, curcumin has one major practical problem: it is almost completely insoluble in water, and when taken orally, very little of it actually reaches the bloodstream. It also breaks down quickly in the body. These limitations have prevented curcumin from being used as a standalone medicine despite its strong biological activity.21 Loading curcumin into a nanocarrier system can protect it from degradation, increase how much of it reaches the target tissue, and control how it is released.22–24
Combining all three components, ZIF-8 MOF, Fe3O4 magnetic nanoparticles, and chitosan into a single system creates a nanocarrier that works on multiple levels at once: high drug loading from the MOF, magnetic targeting from the iron oxide, pH-responsive release from the ZIF-8 degradation, and additional antibacterial action from the chitosan coating.
To develop this system in a scientific and efficient way, we used a Quality by Design (QbD) approach based on the Box–Behnken experimental design. QbD is a method recommended by international pharmaceutical guidelines (ICH Q8) that replaces trial-and-error testing with a structured plan to understand how formulation variables affect product quality.25,26 The Box–Behnken design allowed us to test multiple formulation conditions at the same time and find the best combination using mathematical models, rather than testing every possible combination one by one.27
This study therefore aimed to: (1) prepare curcumin-loaded magnetic MOF–chitosan nanocomposites (Cur/Fe3O4@ZIF-8@CS); (2) optimize the formulation using a Box–Behnken QbD design with four quality responses including particle size, zeta potential, PDI, and encapsulation efficiency; (3) characterize the product step by step using DLS, TEM, and magnetic measurements; (4) compare drug release from the nanocomposite against free curcumin at two different pH values; and (5) test the antibacterial and antifungal activity of the final product; and (6) evaluate the physicochemical stability of the optimized nanocomposite under long-term (25 ± 2 °C) and accelerated (40 ± 2 °C/75 ± 5% RH, ICH Q1A(R2)) storage conditions over two months, tracking particle size, PDI, zeta potential, and curcumin content retention as critical quality attributes; and (7) assess the in vitro cytotoxicity of both the blank and curcumin-loaded nanocomposites on L929 mouse fibroblast cells using the MTT assay. The cytotoxicity study revealed IC50 values of 206.65 µg mL−1 (Fe3O4@ZIF-8@CS) and 273.15 µg mL−1 (Cur/Fe3O4@ZIF-8@CS) against L929 fibroblasts, with the curcumin-loaded formulation showing a higher IC50 and thus lower cytotoxicity to normal cells, supporting its safety profile at therapeutic concentrations.
2 Materials and methods
2.1 Materials
Curcumin (≥98% purity, HPLC grade), zinc nitrate hexahydrate, 2-methylimidazole, ferric chloride hexahydrate, ferrous chloride tetrahydrate, ammonia solution (28%), and low-molecular-weight chitosan (190 kDa) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Glacial acetic acid, dimethyl sulfoxide (DMSO), methanol, and ethanol were from Merck KGaA (Darmstadt, Germany). Tryptone Soya Broth (TSB, Oxoid, cat. no. CM0129B), Brain Heart Infusion Agar (BHI Agar, Oxoid, cat. no. CM1136B), and Sabouraud dextrose broth (Oxoid) were from Oxoid Ltd (Basingstoke, UK). All water used in experiments was ultrapure (≥18.2 MΩ cm, Milli-Q system). Phosphate-buffered saline (PBS, pH 7.4) and acetate buffer (pH 5.5) were freshly prepared as described in Section 2.5.2.2 Box–Behnken experimental design
Before starting the experiments, we defined four target quality criteria for the final nanocomposite, called Critical Quality Attributes (CQAs): particle size between 150 and 300 nm (small enough to accumulate in tumors through the EPR effect), zeta potential above +20 mV (to keep particles stable in suspension and help them bind to bacterial cells), PDI below 0.30 (to ensure the particles are uniform in size), and encapsulation efficiency above 80% (to ensure enough drug is loaded into each particle). These targets were based on established requirements for therapeutic nanoparticles and were set before any experiments were run.28We then identified three formulation variables (independent factors) that were most likely to affect these quality targets, based on a risk assessment: the ratio of MOF to drug (X1), the chitosan concentration used for coating (X2), and the iron oxide content of the composite (X3). Each factor was tested at three levels: low (−1), medium (0), and high (+1), as shown in Table 1.
| Factor | Symbol | Low (−1) | Center (0) | High (+1) |
|---|---|---|---|---|
| MOF-to-drug ratio (w/w) | X1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
| Chitosan concentration (mg mL−1) | X2 | 1.0 | 2.0 | 3.0 |
| Fe3O4 content (% w/w) | X3 | 5 | 10 | 15 |
The Box–Behnken design produced 17 formulations: 12 formulations that test combinations of factor extremes, and 5 repeated formulations at the central (mid-level) conditions to estimate experimental error. All 17 formulations were prepared on different days in a random order to avoid any bias from day-to-day changes. For each formulation, we measured all four CQAs: particle size (Y1, nm), zeta potential (Y2, mV), PDI (Y3), and encapsulation efficiency (Y4, %).
Encapsulation efficiency was measured by UV-vis spectrophotometry (Shimadzu UV-2600, Japan) at 425 nm. After preparing each formulation and collecting it by centrifugation, the amount of curcumin that wasn't encapsulated (remaining in the washing solutions) was measured and subtracted from the total amount added. A calibration curve was made in DMSO: water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) over a range of 0.5–25 µg mL−1 (R2 = 0.9997). EE% and drug loading (DL%) were calculated as:
1, v/v) over a range of 0.5–25 µg mL−1 (R2 = 0.9997). EE% and drug loading (DL%) were calculated as:
| EE (%) = [(total curcumin added − free curcumin)/total curcumin added] × 100 | (1) |
| DL (%) = [(total curcumin added − free curcumin)/total nanocomposite mass] × 100 | (2) |
Each response was fitted to a second-order polynomial equation using Minitab® v21.0:
| Y = β0 + β1X1 + β2X2 + β3X3 + β12X1X2 + β13X1X3 + β23X2X3 + β11X12 + β22X22 + β33X32 | (3) |
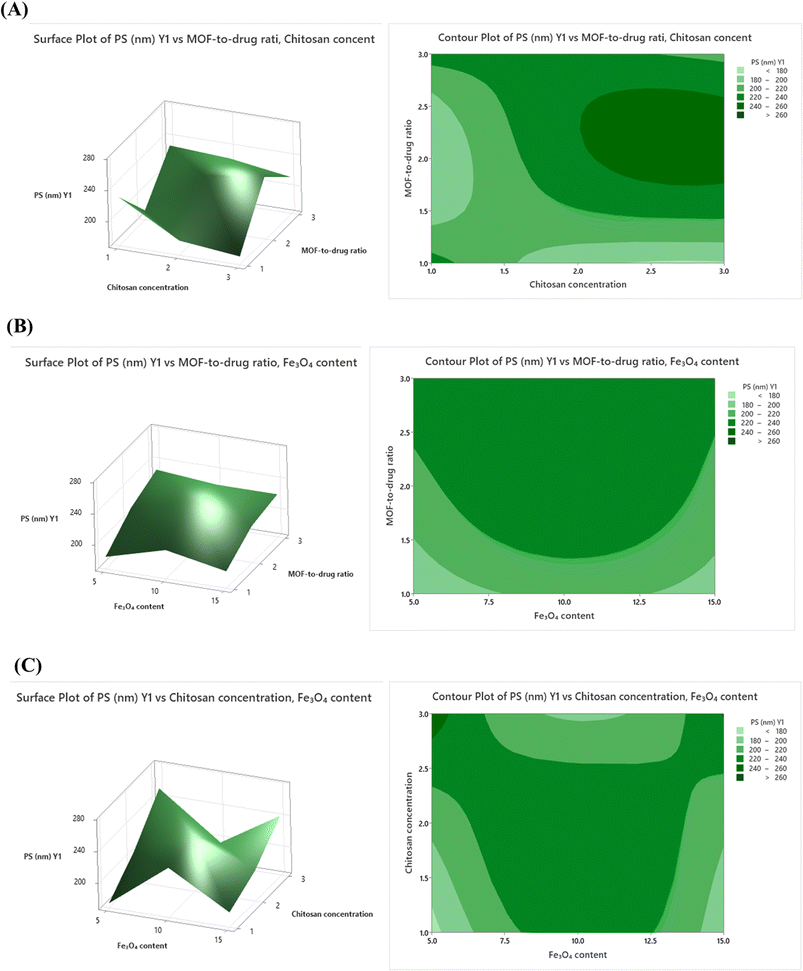
Three-dimensional (3D) response surface plots and the corresponding two-dimensional (2D) contour plots were generated directly from each fitted quadratic model using Minitab® v21.0. For each of the four CQAs, three pairs of plots were produced by varying two factors simultaneously across their experimental range while fixing the third factor at its center-point level: (i) X1 vs. X2 at X3 = 0 (10% w/w Fe3O4); (ii) X1 vs. X3 at X2 = 0 (2.0 mg per mL chitosan); and (iii) X2 vs. X3 at X1 = 0 (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 MOF-to-drug ratio). The 3D surface plots illustrate the overall curvature and magnitude of the response across the design space, whereas the overlaid 2D contour maps delineate iso-response contours that facilitate identification of factor combinations satisfying multiple quality targets simultaneously. The Box–Behnken design was selected in preference to a full factorial or central composite design because it requires only three levels per factor, avoids simultaneous extreme-level combinations (corner points) that may be physically unrealizable or excessively resource-intensive, and achieves a near-rotatable geometry with a substantially smaller number of runs.29
1 MOF-to-drug ratio). The 3D surface plots illustrate the overall curvature and magnitude of the response across the design space, whereas the overlaid 2D contour maps delineate iso-response contours that facilitate identification of factor combinations satisfying multiple quality targets simultaneously. The Box–Behnken design was selected in preference to a full factorial or central composite design because it requires only three levels per factor, avoids simultaneous extreme-level combinations (corner points) that may be physically unrealizable or excessively resource-intensive, and achieves a near-rotatable geometry with a substantially smaller number of runs.29
2.3 Preparation of nanocomposites
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, X3 = 10% w/w Fe3O4), 10 mg of Fe3O4 nanoparticles and 5 mg of curcumin were co-dispersed in 10 mL of methanol and subjected to probe ultrasonication (Q125 Sonicator®, 40% amplitude, 10 min, pulsed 3 s on/1 s off) to produce a uniform, stable dispersion. For all other Box–Behnken design runs, the quantities of Fe3O4 and curcumin were scaled proportionally to achieve the MOF-to-drug ratios and Fe3O4 content levels defined in Table 1. A 0.4 M 2-methylimidazole solution was prepared by dissolving 3.28 g of 2-methylimidazole in 100 mL of distilled water with stirring at room temperature until a clear, homogeneous solution was obtained. A zinc nitrate hexahydrate solution (0.3 M; prepared by dissolving 89.4 mg of Zn (NO3) 2·6H2O in 10 mL of distilled water) was then added dropwise to the Fe3O4/curcumin dispersion under continuous magnetic stirring (500 rpm), followed immediately by the 2-methylimidazole solution. This yielded a Zn2+
1, X3 = 10% w/w Fe3O4), 10 mg of Fe3O4 nanoparticles and 5 mg of curcumin were co-dispersed in 10 mL of methanol and subjected to probe ultrasonication (Q125 Sonicator®, 40% amplitude, 10 min, pulsed 3 s on/1 s off) to produce a uniform, stable dispersion. For all other Box–Behnken design runs, the quantities of Fe3O4 and curcumin were scaled proportionally to achieve the MOF-to-drug ratios and Fe3O4 content levels defined in Table 1. A 0.4 M 2-methylimidazole solution was prepared by dissolving 3.28 g of 2-methylimidazole in 100 mL of distilled water with stirring at room temperature until a clear, homogeneous solution was obtained. A zinc nitrate hexahydrate solution (0.3 M; prepared by dissolving 89.4 mg of Zn (NO3) 2·6H2O in 10 mL of distilled water) was then added dropwise to the Fe3O4/curcumin dispersion under continuous magnetic stirring (500 rpm), followed immediately by the 2-methylimidazole solution. This yielded a Zn2+![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2-methylimidazole molar ratio of 1
2-methylimidazole molar ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8, consistent with the established stoichiometry for room-temperature ZIF-8 synthesis that ensures complete framework crystallization and high drug encapsulation efficiency.33,34 The ZIF-8 framework crystallized around the magnetic cores at room temperature over 4 h, enclosing curcumin molecules inside its pores. The resulting Fe3O4@ZIF-8/Cur particles were collected by centrifugation (10
8, consistent with the established stoichiometry for room-temperature ZIF-8 synthesis that ensures complete framework crystallization and high drug encapsulation efficiency.33,34 The ZIF-8 framework crystallized around the magnetic cores at room temperature over 4 h, enclosing curcumin molecules inside its pores. The resulting Fe3O4@ZIF-8/Cur particles were collected by centrifugation (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 15 min), washed with methanol three times to remove any unentrapped curcumin, and dispersed in dilute acetic acid (0.1 M) for the next step.33
000 × g, 15 min), washed with methanol three times to remove any unentrapped curcumin, and dispersed in dilute acetic acid (0.1 M) for the next step.33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 20 min, 4 °C) and washed twice with ultrapure water to remove unbound chitosan. The washed pellet was resuspended in ultrapure water, snap-frozen in liquid nitrogen, and lyophilized (freeze-dried) for 48 hours to yield a stable dry powder. The powder was stored at 4 °C in sealed vials protected from light until use.
000 × g, 20 min, 4 °C) and washed twice with ultrapure water to remove unbound chitosan. The washed pellet was resuspended in ultrapure water, snap-frozen in liquid nitrogen, and lyophilized (freeze-dried) for 48 hours to yield a stable dry powder. The powder was stored at 4 °C in sealed vials protected from light until use.2.4 Physicochemical characterization methods
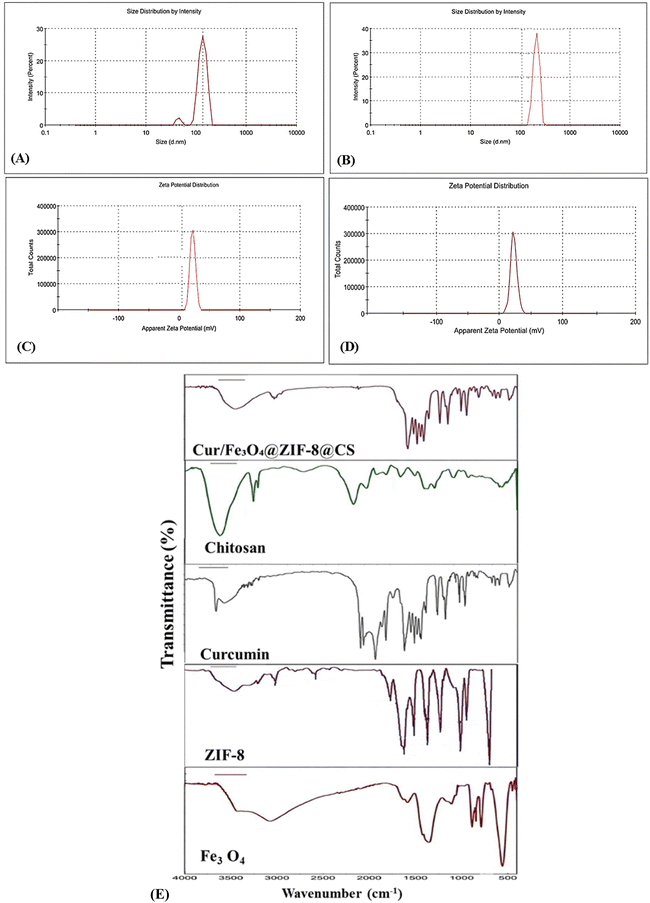
Hydrodynamic particle size, PDI, and zeta potential were measured using a Malvern Zetasizer Nano ZS (UK) at 25 °C. Samples were diluted to 0.1 mg mL−1 in their respective dispersion medium before measurement. Each sample was measured three times (n = 3), and results are reported as mean ± SD. Measurements were taken at each preparation step (bare Fe3O4, intermediate core–shell, and final nanocomposite) to track how the particle properties changed during assembly.The size and shape of the particles were directly visualized by TEM at 200 kV (JEOL JEM-2100F, Japan). Small drops of each sample (0.1 mg mL−1) were placed on carbon-coated copper grids and stained with 2% phosphotungstic acid to make the chitosan layer visible.
The magnetic behavior of Fe3O4 MNPs and the final nanocomposite was measured using a vibrating sample magnetometer (VSM, Lakeshore 7400) at room temperature. The saturation magnetization (the maximum magnetic response) was recorded for each sample. Fourier-transform infrared (FTIR) spectroscopy was performed on each individual component and the final nanocomposite to confirm successful integration of all four materials. Samples were analyzed using an ATR-FTIR spectrometer (Bruker Alpha II, Germany) using KBr pellets over the wavenumber range 400–4000 cm−1 at a resolution of 4 cm−1, averaging 32 scans per spectrum. Spectra of bare Fe3O4 nanoparticles, pure ZIF-8, free low-molecular-weight chitosan, and free curcumin were each recorded individually and then compared with the spectrum of the optimized Cur/Fe3O4@ZIF-8@CS nanocomposite powder.
2.5 In vitro drug release study
Drug release from the nanocomposite was tested at two pH values: pH 7.4 (PBS, representing normal blood and healthy tissue) and pH 5.5 (acetate buffer, representing the acidic environment inside tumors and infected tissue). Phosphate-buffered saline (PBS, pH 7.4) was prepared by dissolving 8.0 g NaCl, 0.2 g KCl, 1.44 g Na2HPO4, and 0.24 g KH2PO4 in approximately 900 mL of distilled water, after which the pH was adjusted to 7.4 using 0.1 M HCl or 0.1 M NaOH. The solution volume was then made up to 1000 mL with distilled water and filtered before use. Acetate buffer (pH 5.5) was prepared by dissolving 4.1 g sodium acetate trihydrate (CH3COONa·3H2O) in about 900 mL of distilled water, followed by the addition of approximately 2.9 mL glacial acetic acid, and the pH was adjusted to 5.5 using small amounts of acetic acid or NaOH if necessary. The final volume was adjusted to 1000 mL with distilled water, and the solution was filtered before use as the acidic release medium. A parallel experiment using free curcumin dissolved in 0.5% Tween-80 was run under the same conditions as a comparison.The dialysis bag method was used. A measured amount of nanocomposite (equivalent to 2 mg of curcumin) was placed inside a dialysis membrane bag and placed in 50 mL of release medium at 37 °C with gentle shaking (100 rpm). At set time points from 0.5 h to 72 h, 3 mL samples were taken and replaced with fresh medium to keep sink conditions. Curcumin in each sample was measured by UV absorbance at 425 nm, and the cumulative percent released was calculated.36
Drug release at time t, cumulative drug release (%) was calculated using eqn (4) and (5):
| Released drug at time t = Ct × V | (4) |
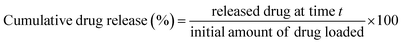 | (5) |
2.6 Release kinetics modeling
The release data from both free curcumin and the nanocomposite were fitted to four mathematical models to understand how drug release occurs and to compare the two systems:Zero-order model eqn (6): drug releases at a constant rate, regardless of how much drug remains.
| Qt = Q0 + K0t | (6) |
First-order model eqn (7): the rate of release decreases over time as less drug remains.
| Qt = Q0e(−K1t) | (7) |
Higuchi model eqn (8): drug diffuses out of a solid matrix, following a square root of time relationship.
| Qt = KHt0.5 | (8) |
Korsmeyer–Peppas model eqn (9): a flexible model that uses an exponent n to identify the release mechanism. For spherical nanoparticles: n ≤ 0.43 means simple diffusion (Fickian); 0.43 < n < 0.85 means both diffusion and polymer swelling contribute (anomalous transport); n = 0.85 means swelling fully controls release.
| Mt/M∞ = ktn | (9) |
The best model was chosen based on the highest R2 and lowest Akaike Information Criterion (AIC) value. All fitting was done using the DDSolver add-in for Microsoft Excel37 and GraphPad Prism 9.0.
2.7 Antimicrobial activity testing
Free curcumin was additionally included as a third experimental parallel control group in all antimicrobial assays to enable direct, within-experiment quantification of the antibacterial advantage conferred by nanoencapsulation. A stock solution of free curcumin (1000 µg mL−1) was prepared in 0.5% DMSO (v/v in Mueller–Hinton broth for bacterial strains, or Sabouraud dextrose broth for C. albicans, as appropriate) with probe ultrasonication (5 min) immediately before use to ensure adequate dispersion. The DMSO vehicle control (0.5% DMSO without curcumin or nanocomposite) was confirmed to produce no inhibition zone in preliminary diffusion assays and was used as a negative vehicle baseline. Curcumin concentration in the stock was verified by UV-vis spectrophotometry at 425 nm against a validated standard curve (R2 = 0.9997) immediately before each assay to confirm solution integrity. This three-arm experimental design (blank nanocomposite/curcumin-loaded nanocomposite/free curcumin) allows rigorous attribution of observed antimicrobial effects to the nanocomposite architecture versus the drug alone.
Plates were incubated at 37 °C for 24 hours. After incubation, the clear zone of no microbial growth surrounding each well, which is called the zone of inhibition (ZOI), was measured in millimeters using a ruler. A wider zone indicates stronger antimicrobial activity. Positive control wells contained streptomycin (10 µg mL−1) for bacterial strains and fluconazole (25 µg mL−1) for C. albicans. All tests were performed in triplicate, and results are reported as mean ± standard deviation (SD).38
To determine the MBC (for bacteria) or MFC (for C. albicans), a loopful of 50–100 µL from each tube showing no turbidity was streaked onto a drug-free nutrient agar plate. These plates were incubated at 37 °C for 24 hours. The MBC/MFC was defined as the lowest concentration at which no visible colonies grew on the agar, indicating that at least 99.9% of the original inoculum had been killed. All MIC and MBC/MFC tests were performed in triplicate, and results are reported as mean ± SD.
2.8 Cytotoxicity assay
| Viability (%) = [(ODt/ODc)] × 100% | (10) |
The relationship between cell survival and drug concentration was plotted to obtain the survival curve for each treatment. IC50 was determined from the dose–response curve using GraphPad Prism software (San Diego, CA, USA).39
2.9 Stability study
The nanocomposite was accurately weighed into amber borosilicate glass vials (Type I, USP), sealed with polytetrafluoroethylene (PTFE)-lined stoppers and aluminum crimp caps under nitrogen atmosphere to minimize oxidative stress. Vials were stored in an upright orientation. Representative samples were withdrawn in triplicate at three predefined time points: zero time (T0), one month (T1), and two months (T2). At each time point, an independently prepared aliquot was reconstituted in methanol at a concentration of 0.1 mg mL−1, sonicated gently for 30 seconds, and allowed to equilibrate at 25 °C for 10 minutes prior to measurement and measured at 425 nm (Shimadzu UV-2600, Japan) against a validated standard curve (0.5–25 µg mL−1, R2 = 0.9997). Curcumin retention at each time point was calculated as:
| Curcumin retention (%) = (Ct/C0) × 100 | (11) |
Physical appearance (color, aggregation, homogeneity upon reconstitution) was assessed visually at each time point.
3 Results and discussion
3.1 BBD experimental matrix and measured responses
The 17 formulations generated by the Box–Behnken design showed a wide range of measured responses, confirming that the three chosen factors had a meaningful impact on nanocomposite quality. Particle size ranged from 173 nm (run 5, low MOF ratio, low Fe3O4) to 269 nm (run 4, high MOF ratio, high chitosan). Zeta potential ranged from +18.7 mV (run 9) to +32.1 mV (run 4). PDI values ranged from 0.215 to 0.276. Encapsulation efficiency varied from 64.9% (run 9) to 88.1% (run 15), demonstrating that the central conditions produced the best drug loading. The five center-point replicates (runs 13–17) gave very similar results to each other (e.g., EE% 86.8–88.1%), showing that the preparation method is reproducible. The full experimental matrix is shown in Table 2.| Run | X1 | X2 | X3 | PS (nm) Y1 | ZP (mV) Y2 | PDI Y3 | EE% Y4 |
|---|---|---|---|---|---|---|---|
| 1 | −1 | −1 | 0 | 183 ± 3.2 | +22.1 ± 0.9 | 0.248 ± 0.02 | 68.2 ± 1.4 |
| 2 | +1 | −1 | 0 | 242 ± 4.8 | +26.4 ± 1.1 | 0.276 ± 0.03 | 74.5 ± 1.8 |
| 3 | −1 | +1 | 0 | 196 ± 3.9 | +28.7 ± 1.3 | 0.231 ± 0.02 | 71.3 ± 1.2 |
| 4 | +1 | +1 | 0 | 269 ± 5.3 | +32.1 ± 1.4 | 0.261 ± 0.02 | 83.6 ± 2.1 |
| 5 | −1 | 0 | −1 | 173 ± 2.8 | +19.8 ± 0.8 | 0.239 ± 0.02 | 66.4 ± 1.6 |
| 6 | +1 | 0 | −1 | 252 ± 4.6 | +24.5 ± 1.1 | 0.268 ± 0.02 | 76.2 ± 1.9 |
| 7 | −1 | 0 | +1 | 189 ± 3.4 | +21.3 ± 0.9 | 0.244 ± 0.02 | 70.1 ± 1.4 |
| 8 | +1 | 0 | +1 | 257 ± 5.0 | +29.6 ± 1.3 | 0.272 ± 0.03 | 80.8 ± 2.2 |
| 9 | 0 | −1 | −1 | 178 ± 2.9 | +18.7 ± 0.7 | 0.258 ± 0.02 | 64.9 ± 1.3 |
| 10 | 0 | +1 | −1 | 214 ± 4.1 | +27.3 ± 1.2 | 0.237 ± 0.02 | 75.7 ± 1.7 |
| 11 | 0 | −1 | +1 | 186 ± 3.3 | +20.4 ± 0.9 | 0.251 ± 0.02 | 68.3 ± 1.5 |
| 12 | 0 | +1 | +1 | 222 ± 4.4 | +29.8 ± 1.3 | 0.242 ± 0.02 | 77.9 ± 1.9 |
| 13 | 0 | 0 | 0 | 229 ± 4.7 | +31.5 ± 1.4 | 0.218 ± 0.01 | 87.4 ± 2.3 |
| 14 | 0 | 0 | 0 | 231 ± 4.9 | +30.9 ± 1.3 | 0.221 ± 0.01 | 86.8 ± 2.1 |
| 15 | 0 | 0 | 0 | 230 ± 4.6 | +32.0 ± 1.5 | 0.215 ± 0.01 | 88.1 ± 2.4 |
| 16 | 0 | 0 | 0 | 230 ± 4.8 | +31.2 ± 1.4 | 0.219 ± 0.01 | 87.2 ± 2.2 |
| 17 | 0 | 0 | 0 | 229 ± 4.5 | +31.7 ± 1.5 | 0.217 ± 0.01 | 87.9 ± 2.5 |
| Y1 (PS, nm) = 230.00 + 32.38X1 + 14.25X2 + 8.21X3 + 7.92X1X2 − 5.69X1X3 − 4.00X2X3 − 12.45X12 − 8.48X22 − 6.63X32 | (12) |
| Y2 (ZP, mV) = +31.26 + 3.47X1 + 4.00X2 + 2.14X3 + 1.78X1X2 − 1.31X1X3 − 0.90X2X3 − 3.26X12 − 2.68X22 − 1.93X32 | (13) |
| Y3 (PDI) = 0.219 + 0.0292X1 + 0.0217X2 + 0.0125X3 + 0.0092X1X2 + 0.0073X1X3 + 0.0045X2X3 − 0.0177X12 − 0.0150X22 − 0.0102X32 | (14) |
| Y4 (EE%, %) = 87.48 + 8.84X1 + 7.06X2 + 3.20X3 + 2.38X1X2 − 1.45X1X3 + 1.02X2X3 − 8.44X12 − 4.67X22 − 2.44X32 | (15) |
Each of the four responses was fitted to a second-order polynomial model. Table 3 presents all four ANOVA results side by side in a single collective table. All four models were statistically significant (p < 0.0001 for the model F-test) and showed a non-significant lack-of-fit (p > 0.05), which means the models fit the data well and can be trusted to make predictions. R2 values were above 0.994 for all responses, and the difference between adjusted R2 and predicted R2 was less than 0.2 in all cases, confirming that the models are not over-fitted. Adequate precision values ranged from 38.7 to 54.3, all well above the minimum acceptable value of 4, indicating a strong signal-to-noise ratio and reliable model discrimination.
| Source | PS SS | p | ZP SS | p | PDI SS | p | EE% SS | p |
|---|---|---|---|---|---|---|---|---|
| Model | 4821 | <0.0001 | 152.8 | <0.0001 | 0.00812 | <0.0001 | 844.2 | <0.0001 |
| X1 | 2105 | <0.0001 | 48.26 | <0.0001 | 0.00341 | <0.0001 | 312.5 | <0.0001 |
| X2 | 891 | <0.0001 | 64.18 | <0.0001 | 0.00188 | <0.0001 | 198.8 | <0.0001 |
| X3 | 312 | 0.0002 | 18.43 | 0.0038 | 0.00062 | 0.0091 | 41.23 | 0.0003 |
| X1X2 | 249 | 0.0006 | 12.61 | 0.0112 | 0.00034 | 0.0248 | 22.56 | 0.0018 |
| X1X3 | 129 | 0.0041 | 6.84 | 0.0418 | 0.00021 | 0.0631 | 8.41 | 0.0132 |
| X2X3 | 64 | 0.0284 | 3.22 | 0.1124 | 0.00008 | 0.2840 | 4.18 | 0.0402 |
| X12 | 613 | <0.0001 | 42.17 | <0.0001 | 0.00124 | 0.0002 | 178.3 | <0.0001 |
| X22 | 284 | 0.0003 | 28.64 | 0.0008 | 0.00089 | 0.0012 | 54.62 | 0.0001 |
| X32 | 175 | 0.0018 | 14.78 | 0.0078 | 0.00041 | 0.0182 | 23.63 | 0.0016 |
| Residual | 148 | — | 4.61 | — | 0.00031 | — | 4.19 | — |
| Lack of fit | 96 | 0.163 | 3.18 | 0.189 | 0.00019 | 0.201 | 2.84 | 0.178 |
| Pure error | 52 | — | 1.43 | — | 0.00012 | — | 1.35 | — |
| R2 | 0.9968 | 0.9941 | 0.9953 | 0.9951 | ||||
| Radj2 | 0.9927 | 0.9866 | 0.9893 | 0.9888 | ||||
| Rpred2 | 0.9782 | 0.9614 | 0.9712 | 0.9741 | ||||
| Adeq. prec | 54.3 | 38.7 | 46.1 | 42.7 | ||||
| CV% | 0.84 | 1.12 | 1.43 | 1.03 |
Among all factors, the MOF-to-drug ratio (X1) had the largest effect on particle size and encapsulation efficiency, while chitosan concentration (X2) was the strongest driver of zeta potential. PDI was most affected by X1 as well, suggesting that higher MOF-to-drug ratios introduce some size heterogeneity, possibly because multiple iron oxide cores can become embedded within a single ZIF-8 crystal at higher ratios.
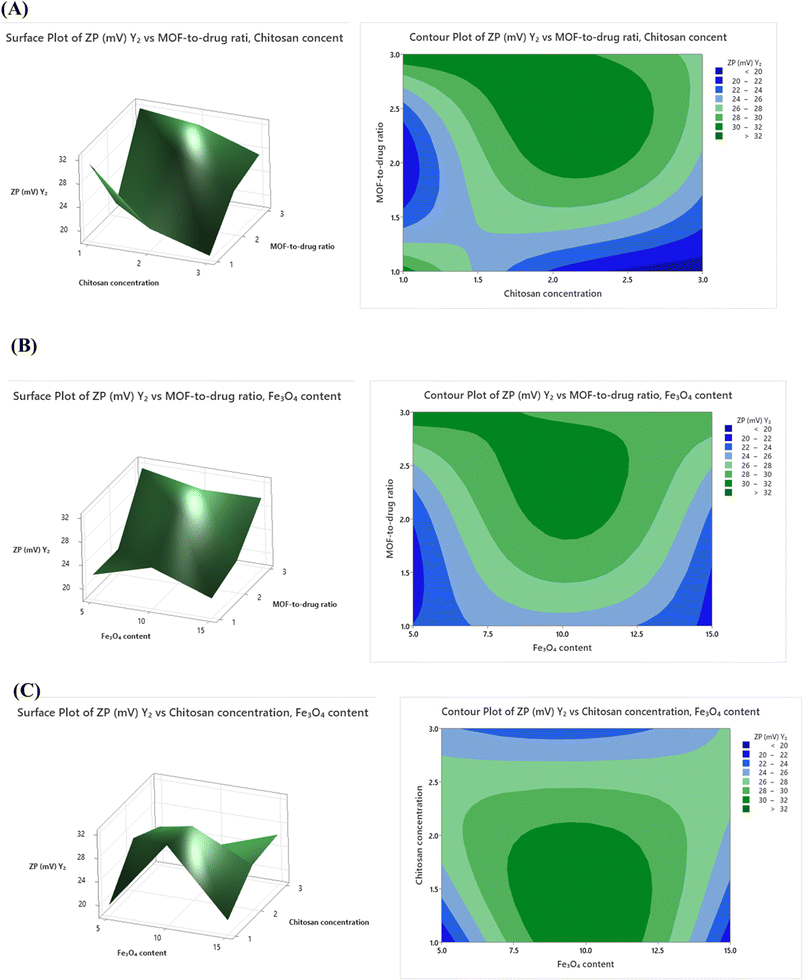
Zeta potential (Y2) was most strongly influenced by X2 (Fig. 2A–C). As chitosan concentration increased, more positively charged polymer chains coated the surface, raising the zeta potential from around +19 mV to +32 mV. This is a desirable outcome because higher zeta potential improves colloidal stability and enhances the interaction of the nanoparticle with negatively charged bacterial cell membranes.
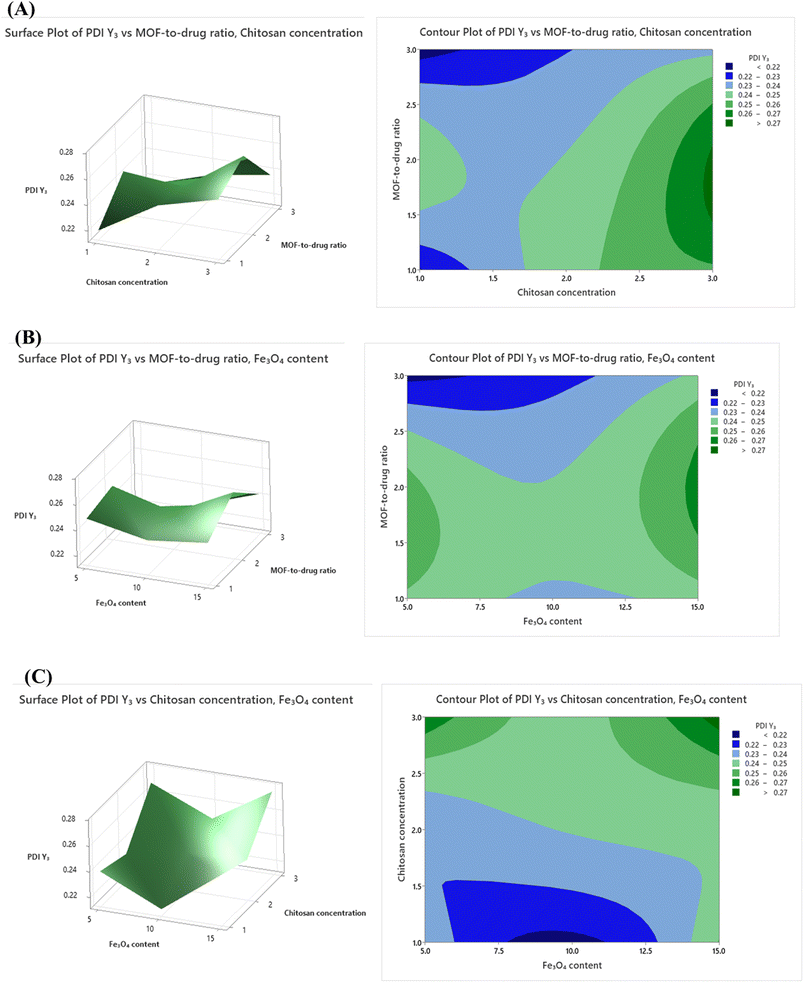
PDI (Y3) increased with X1 and X3 (Fe3O4 content), a trend that can be observed in Fig. 3A–C. Larger amounts of magnetic nanoparticles may create some variation in the number of cores per ZIF-8 particle, leading to a broader size distribution. However, all PDI values remained below 0.28, staying within acceptable limits for nanoparticles.
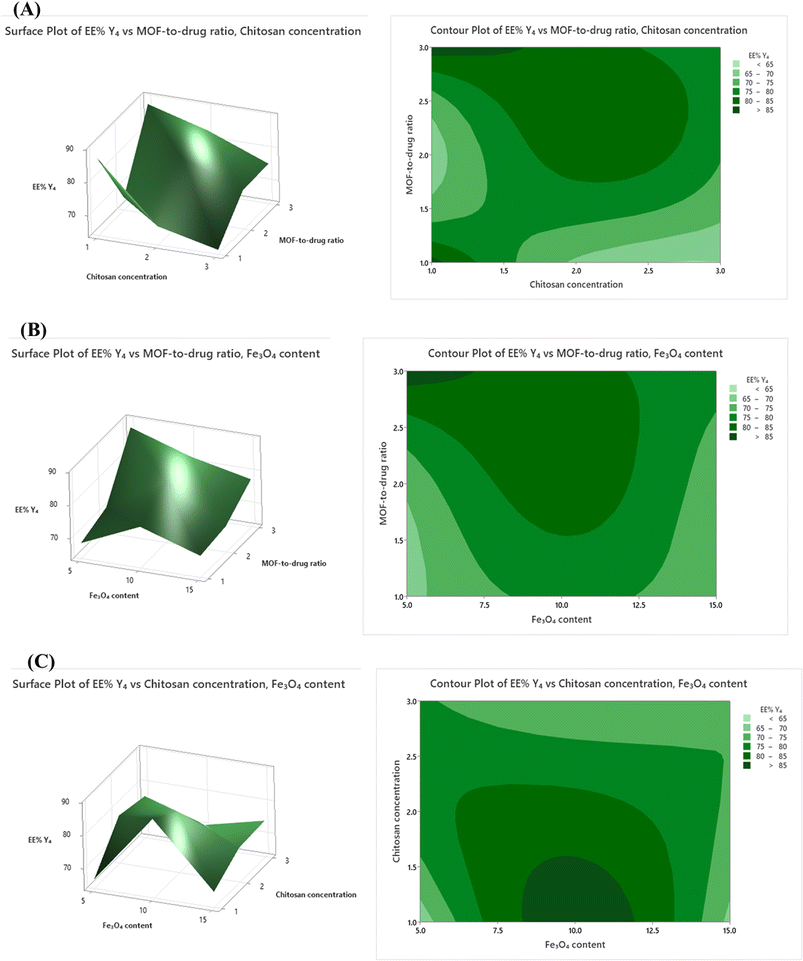
Encapsulation efficiency (Y4) increased with X1 but followed a curved (quadratic) relationship (Fig. 4A–C). EE% improved as the MOF framework increased relative to drug, because more pore space was available for curcumin to occupy. However, at the highest MOF-to-drug ratios, EE% began to plateau and slightly decline, likely because the pores became partially blocked or the drug distribution became inefficient. The highest EE% values (86.8–88.1%) were found consistently at the center-point conditions.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (MOF
1 (MOF![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) drug), X2 = 2.0 mg per mL chitosan, X3 = 10% Fe3O4. This formulation was then prepared three times independently to check whether the predicted values matched the actual measured values. As shown in Table 4, the measured results agreed very closely with the model predictions, with errors of less than 2.3% for all four responses. This confirms that the Box–Behnken model is valid and that the optimized formulation can be reproduced reliably. All particle size values reported in the Box–Behnken design and optimization (predicted: 232 nm; measured: 228.6 ± 4.7 nm) refer to the hydrodynamic diameter as determined by DLS, which is the standard physicochemical quality attribute applied in pharmaceutical quality by design frameworks. These values are not directly comparable to TEM core dimensions, which are systematically smaller due to the absence of the hydration shell and chitosan corona under dry imaging conditions (see Sections 3.2.1 and 3.2.2).
drug), X2 = 2.0 mg per mL chitosan, X3 = 10% Fe3O4. This formulation was then prepared three times independently to check whether the predicted values matched the actual measured values. As shown in Table 4, the measured results agreed very closely with the model predictions, with errors of less than 2.3% for all four responses. This confirms that the Box–Behnken model is valid and that the optimized formulation can be reproduced reliably. All particle size values reported in the Box–Behnken design and optimization (predicted: 232 nm; measured: 228.6 ± 4.7 nm) refer to the hydrodynamic diameter as determined by DLS, which is the standard physicochemical quality attribute applied in pharmaceutical quality by design frameworks. These values are not directly comparable to TEM core dimensions, which are systematically smaller due to the absence of the hydration shell and chitosan corona under dry imaging conditions (see Sections 3.2.1 and 3.2.2).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1; X2 = 2.0 mg mL; X3 = 10% w/w; n = 3, mean ± SD)
1; X2 = 2.0 mg mL; X3 = 10% w/w; n = 3, mean ± SD)
| Response | Predicted value | Measured value (n = 3, mean ± SD) | Error (%) |
|---|---|---|---|
| Particle size (nm) [DLS, hydrodynamic diameter] | 232 | 228.6 ± 4.7 | 1.51 |
| Zeta potential (mV) | +30.8 | +31.5 ± 1.4 | 2.27 |
| PDI | 0.218 | 0.218 ± 0.01 | 0.00 |
| Encapsulation eff. (%) | 87.1 | 87.4 ± 2.3 | 0.34 |
3.2 Physicochemical characterization
| Parameter | Fe3O4 MNPs | Fe3O4@ZIF-8/Cur | Final nanocomposite |
|---|---|---|---|
| Hydrodynamic size (nm) | 28.3 ± 2.4 | 186.3 ± 3.9 | 228.6 ± 4.7 |
| TEM core size (nm) | 10.3 ± 1.4 | 142.7 ± 8.6 | ∼220 |
| PDI | 0.171 ± 0.01 | 0.231 ± 0.02 | 0.218 ± 0.01 |
| Zeta potential (mV) | −14.2 ± 1.2 | +18.6 ± 1.4 | +31.5 ± 1.4 |
| Encapsulation efficiency (%) | — | 79.2 ± 1.9 | 87.4 ± 2.3 |
| Drug loading (%) | — | 7.31 ± 0.18 | 8.08 ± 0.21 |
| Saturation magnetization (emu g−1) | 62.8 | — | 6.97 |
This step-by-step hydrodynamic size increase combined with the charge reversal pattern is strong evidence that each layer was added successfully. The final PDI of 0.218 shows the particles are fairly uniform in size. The zeta potential of +31.5 mV is well above the +20 mV stability threshold, meaning the particles will stay dispersed in solution without clumping together. When stored as a freeze-dried powder and redispersed, the particles retained their size and PDI for at least 90 days at 4 °C, confirming good physical stability.
It is important to note that the hydrodynamic diameters measured by DLS (186.3 nm for Fe3O4@ZIF-8/Cur and 228.6 nm for the final nanocomposite) are consistently larger than the corresponding TEM core sizes, as expected. DLS measures the full hydrodynamic envelope of the particle in suspension, including the chitosan corona and the surrounding electrical double layer, whereas TEM captures only the dry inorganic/polymeric core under vacuum. This discrepancy is well established for polymer-coated nanoparticles and is not an inconsistency; both measurement techniques provide complementary and consistent information about different structural dimensions of the same particle.34,35,41–44
| Assignment | Fe3O4 (cm−1) | ZIF-8 (cm−1) | Chitosan (cm−1) | Nanocomposite (cm−1) |
|---|---|---|---|---|
| a All wavenumbers are approximate (±2 cm−1). Dashes indicate absence of the band in the respective individual component. | ||||
| Fe–O stretching | 569, 399 | — | — | 565, 397 (retained) |
| Zn–N coordination | — | 421 | — | 418 (shifted) |
C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching (imidazolate) N stretching (imidazolate) |
— | 1584 | — | 1578 (shifted) |
| N–H/O–H stretch (chitosan) | — | — | 3348 | 3338 (broadened) |
Amide I (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, chitosan) O, chitosan) |
— | — | 1648 | 1632 (red-shifted) |
| C–O–C glycosidic (chitosan) | — | — | 1025 | 1022 (retained) |
C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/C O/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C conjugated (curcumin, β-diketone) C conjugated (curcumin, β-diketone) |
— | — | — | 1606 (red-shifted from 1626) |
| O–H enol stretch (curcumin) | — | — | — | 3505 (attenuated) |
The spectrum of bare Fe3O4 displayed two characteristic Fe–O stretching vibrations at approximately 569 cm−1 and 399 cm−1, consistent with the inverse spinel lattice vibrations of magnetite reported in the literature.45,46 A broad O–H absorption around 3420 cm−1 was also present, attributable to surface hydroxyl groups and adsorbed moisture on the nanoparticle surface. Both Fe–O bands were retained in the final nanocomposite spectrum, confirming that the magnetic iron oxide core was structurally preserved throughout the multi-step fabrication process.
The FTIR spectrum of as-synthesized ZIF-8 showed the following characteristic bands: a Zn–N coordination stretching vibration at 421 cm−1, out-of-plane bending of the imidazole ring at 690–760 cm−1, in-plane bending of C–H at 950–1200 cm−1, C–N stretching of the imidazolate linker at 1145–1180 cm−1, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching at 1584 cm−1, and aromatic C–H stretching at 3135 cm−1. These bands are in excellent agreement with published ZIF-8 reference spectra.12,34 In the nanocomposite, the Zn–N and C
N stretching at 1584 cm−1, and aromatic C–H stretching at 3135 cm−1. These bands are in excellent agreement with published ZIF-8 reference spectra.12,34 In the nanocomposite, the Zn–N and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bands were retained but shifted slightly to 418 cm−1 and 1578 cm−1 respectively, indicating that the ZIF-8 framework remained intact while undergoing mild coordination perturbation upon interaction with the iron oxide surface and chitosan coating.
N bands were retained but shifted slightly to 418 cm−1 and 1578 cm−1 respectively, indicating that the ZIF-8 framework remained intact while undergoing mild coordination perturbation upon interaction with the iron oxide surface and chitosan coating.
The spectrum of free low-molecular-weight chitosan exhibited a broad O–H/N–H stretching envelope centered at 3348 cm−1, aliphatic C–H stretching at 2873 cm−1, a carbonyl band (amide I, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) at 1648 cm−1, N–H bending (amide II) at 1590 cm−1, and a C–O–C glycosidic linkage stretching band at 1025 cm−1. These features are consistent with the established FTIR fingerprint of partially deacetylated chitosan.17 In the nanocomposite spectrum, the O–H/N–H band was retained but broadened, and the amide I band shifted to 1632 cm−1. This red-shift is consistent with the formation of hydrogen bonds between chitosan amine groups and the surface oxygen atoms of Fe3O4 and/or the imidazolate linkers of ZIF-8, confirming chitosan surface deposition.35,47
O) at 1648 cm−1, N–H bending (amide II) at 1590 cm−1, and a C–O–C glycosidic linkage stretching band at 1025 cm−1. These features are consistent with the established FTIR fingerprint of partially deacetylated chitosan.17 In the nanocomposite spectrum, the O–H/N–H band was retained but broadened, and the amide I band shifted to 1632 cm−1. This red-shift is consistent with the formation of hydrogen bonds between chitosan amine groups and the surface oxygen atoms of Fe3O4 and/or the imidazolate linkers of ZIF-8, confirming chitosan surface deposition.35,47
The FTIR spectrum of curcumin showed characteristic bands at 3510 cm−1 (O–H stretch of enol form), 3008 cm−1 (aromatic C–H stretch), 1626 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/C
O/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C conjugated stretch, β-diketone chelated form), 1510 cm−1 (C
C conjugated stretch, β-diketone chelated form), 1510 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C aromatic ring stretch), 1270 cm−1 (C–O–C phenolic ether), and 960 cm−1 (=C–H bending). These are in full agreement with established curcumin reference spectra.20,31
C aromatic ring stretch), 1270 cm−1 (C–O–C phenolic ether), and 960 cm−1 (=C–H bending). These are in full agreement with established curcumin reference spectra.20,31
The FTIR spectrum of the optimized nanocomposite (Cur/Fe3O4@ZIF-8@CS) showed the following critical features confirming successful multicomponent integration: retention of Fe–O bands at ∼565 and ∼397 cm−1 which confirms preservation of the Fe3O4 magnetic core within the nanocomposite; retention of ZIF-8 Zn–N and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N bands (shifted to 418 and 1578 cm−1) which confirms ZIF-8 framework formation and mild coordination interaction with surrounding components; presence of chitosan amide I and glycosidic bands (shifted amide I at 1632 cm−1) which confirms successful chitosan coating and hydrogen-bond interaction with the ZIF-8/Fe3O4 surface; shift of curcumin C
N bands (shifted to 418 and 1578 cm−1) which confirms ZIF-8 framework formation and mild coordination interaction with surrounding components; presence of chitosan amide I and glycosidic bands (shifted amide I at 1632 cm−1) which confirms successful chitosan coating and hydrogen-bond interaction with the ZIF-8/Fe3O4 surface; shift of curcumin C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/C
O/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C conjugated band from 1626 to 1606 cm−1 where this bathochromic shift is a strong indicator of host–guest interaction between encapsulated curcumin and the ZIF-8 pore environment. The downshift is consistent with weakening of the C
C conjugated band from 1626 to 1606 cm−1 where this bathochromic shift is a strong indicator of host–guest interaction between encapsulated curcumin and the ZIF-8 pore environment. The downshift is consistent with weakening of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond through π–π stacking interactions between the aromatic rings of curcumin and the imidazolate linkers, as well as possible hydrogen bonding of the curcumin enol O–H with the Zn–N coordination sites inside the MOF pore.12,34 This confirms that curcumin is encapsulated within the ZIF-8 pores and not merely adsorbed on the outer particle surface; attenuation of the free curcumin O–H band at 3510 cm−1: the reduction in intensity of the free enol O–H stretch in the nanocomposite spectrum relative to pure curcumin is consistent with involvement of this group in hydrogen bonding within the ZIF-8 cage, further supporting genuine encapsulation.
O bond through π–π stacking interactions between the aromatic rings of curcumin and the imidazolate linkers, as well as possible hydrogen bonding of the curcumin enol O–H with the Zn–N coordination sites inside the MOF pore.12,34 This confirms that curcumin is encapsulated within the ZIF-8 pores and not merely adsorbed on the outer particle surface; attenuation of the free curcumin O–H band at 3510 cm−1: the reduction in intensity of the free enol O–H stretch in the nanocomposite spectrum relative to pure curcumin is consistent with involvement of this group in hydrogen bonding within the ZIF-8 cage, further supporting genuine encapsulation.
Taken together, the FTIR data shown in Fig. 5E provide comprehensive spectroscopic confirmation that all four components including Fe3O4, ZIF-8, chitosan, and curcumin are present in the final nanocomposite and that the assembly involved real chemical/physical interactions between components rather than mere physical mixing. This is fully consistent with the step-by-step size and zeta potential evolution documented in Table 5 and the TEM images in Fig. 6.
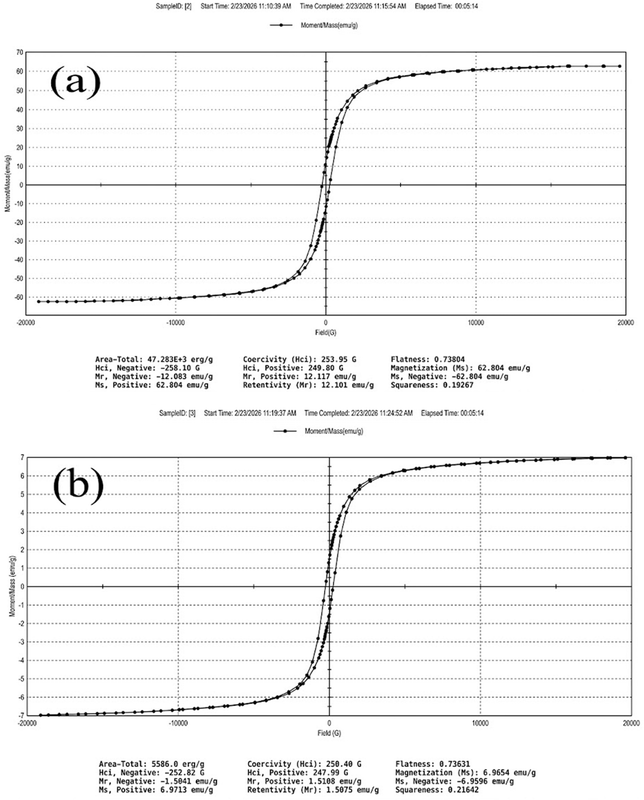
For the bare Fe3O4 nanoparticles (Fig. 7a), the saturation magnetization was Ms = 62.8 emu g−1, with a coercivity of Hci = 253.95 G and retentivity of Mr = 12.10 emu g−1 (squareness Mr/Ms = 0.193). The Ms value is lower than that of bulk magnetite (∼92 emu g−1), which is expected for nanoscale Fe3O4 particles and is attributable to surface spin disorder effects that is at the nanoscale, a significant proportion of iron atoms reside at the particle surface where coordination is incomplete, reducing the net magnetic moment per unit mass.45 The low coercivity and squareness well below 0.5 confirm that the particles are in a near-superparamagnetic regime, meaning they respond strongly to an applied magnetic field but retain only minimal residual magnetization once the field is removed; a highly desirable property for biomedical applications where particle aggregation in the bloodstream must be avoided.13
After incorporation into the final Cur/Fe3O4@ZIF-8@CS nanocomposite (Fig. 7b), the saturation magnetization decreased substantially to Ms = 6.97 emu g−1, with coercivity Hci = 250.40 G and retentivity Mr = 1.51 emu g−1 (squareness = 0.216). This reduction in Ms is a direct and expected consequence of dilution of the magnetic Fe3O4 phase by the non-magnetic components including ZIF-8 framework, chitosan coating, and encapsulated curcumin which collectively constitute the majority of the composite mass and contribute no magnetic moment.49 Importantly, the near-identical coercivity values between the bare Fe3O4 (253.95 G) and the final composite (250.40 G) confirm that the magnetic character of the iron oxide cores is fully preserved after coating, and that the ZIF-8 and chitosan layers do not alter the intrinsic magnetic domain structure of the Fe3O4 cores. Despite the lower absolute Ms, the composite retains sufficient magnetic responsiveness for external magnetic field-guided targeting, consistent with previously reported thresholds for magnetically steerable drug delivery nanocarriers.13
3.3 In vitro drug release
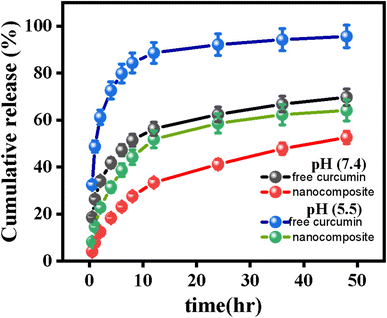
Fig. 8 shows the cumulative release profiles of free curcumin and the optimized nanocomposite at pH 5.5 and pH 7.4 over 72 hours. The results clearly show three important differences between the two systems.First, free curcumin released very quickly in the first 30 minutes, giving 32.4 ± 1.2% release at pH 5.5 and 18.6 ± 0.9% at pH 7.4. This early burst is a known problem with freely dissolved drugs; without any carrier to hold them back, they diffuse rapidly into the surrounding medium. By contrast, the nanocomposite released only 8.2 ± 0.4% at pH 5.5 and 4.1 ± 0.2% at pH 7.4 at the same time point. This much smaller burst shows that the MOF–chitosan structure successfully slows down initial drug escape.
Second, free curcumin reached a high release level quickly and then levelled off by 24 h, reaching about 92% at pH 5.5 and 62% at pH 7.4 with little further change afterward. The nanocomposite, on the other hand, continued releasing drug gradually over the full 72 hours, reaching 63.8 ± 2.2% at pH 5.5 and 58.4 ± 1.9% at pH 7.4. This slower and more sustained release is exactly what is needed for a therapeutic nanocarrier, as it can maintain effective drug levels at the target site for a longer time.
Third, both free curcumin and the nanocomposite released more drug at the lower pH (5.5) than at pH 7.4. For the nanocomposite, this pH effect is particularly useful: at pH 5.5, the zinc-imidazolate bonds in ZIF-8 weaken under acidic conditions, causing the framework to gradually break apart and release the trapped curcumin. At the same time, chitosan swells in acidic conditions as its amine groups become positively charged, creating larger gaps in the coating layer through which drug molecules can escape more easily. At pH 7.4 (normal tissue and blood), both mechanisms are less active, meaning the nanocomposite holds onto most of its drug during circulation and preferentially releases it when it reaches the acidic tumor or infection environment.
The cumulative release of 63.8 ± 2.2% at pH 5.5 over 72 hours, compared to 58.4 ± 1.9% at pH 7.4, reflects the expected outcome of a multi-barrier, pH-responsive nanocomposite system and should not be interpreted as evidence of an incomplete or contradictory release mechanism. Several interconnected factors collectively account for why drug release is sustained and does not reach 100% within the 72 hour experimental window, and why this behavior is both scientifically anticipated and therapeutically desirable.
First, ZIF-8 degradation under mild acidic conditions (pH 5.5 acetate buffer) is a gradual, time-dependent process and not an instantaneous dissolution event. The zinc-imidazolate coordination bonds weaken progressively as proton activity increases, producing a slow, sustained release of encapsulated curcumin over many hours. Under pH 5.5 conditions, ZIF-8 is kinetically destabilized but far from fully dissolved within 72 h. Published studies on ZIF-8 and related zeolitic imidazolate framework nanocarriers consistently report cumulative drug release values in the range of 55–78% at pH 5.0–5.5 over 24–72 h observation windows, without complete framework dissolution, confirming that our observed 63.8% is fully within the expected mechanistic range for this class of materials.11,12,42 Complete ZIF-8 dissolution would require either prolonged incubation (>168 h), more strongly acidic conditions (pH ≤ 4.5), or the presence of competing chelating agents, none of which are present in the physiological tumor microenvironment and are therefore not appropriate comparators for in vitro release assessment.
Second, the chitosan outer coating constitutes an independent, rate-limiting diffusion barrier that the released curcumin must traverse before entering the bulk release medium. Even after partial ZIF-8 degradation liberates drug molecules from the MOF pore network, diffusion through the swollen but structurally intact chitosan polymer matrix imposes additional kinetic resistance. This dual-barrier (ZIF-8 + chitosan) architecture is quantitatively captured by the Korsmeyer–Peppas release exponent n = 0.61 at pH 5.5 (anomalous, non-Fickian transport), confirming that release is governed simultaneously by framework degradation and polymer chain relaxation rather than by simple Fickian diffusion alone. The meaningfully higher n value at pH 5.5 compared to pH 7.4 (0.61 vs. 0.53) demonstrates that chitosan swelling makes a larger mechanistic contribution in the acidic environment, consistent with the known pH-sensitive protonation of chitosan amine groups.16,17
Third, curcumin is not merely confined within ZIF-8 pores by physical trapping but is additionally retained by non-covalent host–guest interactions including π–π stacking between curcumin's extended aromatic system and the imidazolate linkers, and hydrogen bonding of the curcumin enol O–H with Zn–N coordination sites (confirmed by the FTIR bathochromic shift of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O/C
O/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C conjugated band from 1626 to 1606 cm−1, Section 3.2.5). These host–guest binding interactions require thermodynamic or kinetic energy to overcome, meaning that a residual fraction of curcumin remains associated with partially intact or degraded ZIF-8 fragments throughout the experimental period. Complete displacement of this bound fraction would require conditions beyond those of the physiological acidic microenvironment.
C conjugated band from 1626 to 1606 cm−1, Section 3.2.5). These host–guest binding interactions require thermodynamic or kinetic energy to overcome, meaning that a residual fraction of curcumin remains associated with partially intact or degraded ZIF-8 fragments throughout the experimental period. Complete displacement of this bound fraction would require conditions beyond those of the physiological acidic microenvironment.
Critically, the 5.4 percentage-point differential in cumulative release between pH 5.5 (63.8%) and pH 7.4 (58.4%) confirms authentic, statistically significant pH-responsiveness (p < 0.05). For a targeted nanocarrier, incomplete release over 72 hours is not a formulation failure but a deliberate design outcome. A system releasing 100% of its payload rapidly at both pH values would be functionally indistinguishable from a simple burst-release formulation and would provide no preferential selectivity for the tumor/infection microenvironment over systemic circulation. The sustained, pH-dependent release profile maintained over 72 h ensures prolonged drug exposure above the MIC at the pathological site, fulfilling the primary pharmacokinetic objective of a controlled-release nanocarrier. Drug release is governed by pH-triggered progressive weakening of Zn–N coordination bonds, coupled with chitosan chain swelling, which results in anomalous non-Fickian transport and preferentially accelerated drug release in acidic microenvironments. This mechanistic interpretation is fully consistent with both the experimental release data and the kinetic modeling results.
3.4 Drug release kinetics
Table 7 shows the fitting results for all four kinetic models applied to both free curcumin and the nanocomposite at both pH values.| Parameter | Free Cur pH 5.5 | Nanocomp. pH 5.5 | Free Cur pH 7.4 | Nanocomp. pH 7.4 | Key finding |
|---|---|---|---|---|---|
| Zero-order (R2) | 0.889 | 0.942 | 0.861 | 0.918 | Poor fit for free drug |
| Zero-order K0 (% h−1) | 1.12 | 3.21 | 0.89 | 1.47 | Slower constant rate |
| First-order (R2) | 0.914 | 0.961 | 0.888 | 0.939 | Improved for nanocomp. |
| First-order K1 (h−1) | 0.082 | 0.038 | 0.051 | 0.019 | 2–3× slower release |
| Higuchi (R2) | 0.961 | 0.987 | 0.937 | 0.974 | Best fit for free drug |
| Higuchi KH (% h−1/2) | 14.3 | 8.92 | 9.8 | 5.43 | Sustained diffusion |
| Korsmeyer–Peppas (R2) | 0.976 | 0.994 | 0.958 | 0.981 | Best fit: nanocomp. |
| KP rate constant k | 0.214 | 0.082 | 0.148 | 0.049 | Controlled release |
| KP exponent n | 0.38 | 0.61 | 0.41 | 0.53 | Non-Fickian transport |
| Release mechanism | Fickian | Anomalous | Fickian | Anomalous | Matrix + diffusion |
| Best-fit model | Higuchi | Korsmeyer–Peppas | Higuchi | Korsmeyer–Peppas | — |
For free curcumin, the Higuchi model gave the best fit (R2 = 0.961 at pH 5.5 and 0.937 at pH 7.4), and the Korsmeyer–Peppas exponent n was 0.38–0.41. Both of these results point to simple Fickian diffusion as the mechanism: curcumin molecules in the Tween-80 vehicle move passively from high concentration to low concentration with no physical resistance slowing them down, following a square-root-of-time pattern.
For the nanocomposite, the Korsmeyer–Peppas model gave the best fit (R2 = 0.994 at pH 5.5 and 0.981 at pH 7.4), and the n values were 0.61 (pH 5.5) and 0.53 (pH 7.4). These n values fall between 0.43 and 0.85, which indicates anomalous (non-Fickian) transport—meaning that release is controlled by two things happening at the same time: drug diffusing through the MOF pore network, and the chitosan polymer layer gradually swelling and relaxing. Neither mechanism alone is fast enough to match the Higuchi or first-order models; instead, both work together to produce the sustained release profile seen in the data.
The higher n value at pH 5.5 compared to pH 7.4 (0.61 vs. 0.53) shows that polymer swelling plays a bigger role at lower pH, which is consistent with the known pH-sensitive behavior of chitosan. The first-order rate constants also confirm the benefit of encapsulation: curcumin releases 2.2 times more slowly from the nanocomposite than from the Tween-80 vehicle at pH 5.5, and 2.7 times more slowly at pH 7.4, providing a quantitative measure of the sustained-release advantage.
3.5 Antimicrobial activity
The antimicrobial performance of the blank nanocomposite (Fe3O4@ZIF-8@CS), the curcumin-loaded nanocomposite (Cur/Fe3O4@ZIF-8@CS), and free curcumin as a parallel experimental control was assessed against five clinical microbial strains using three standard assays: agar well diffusion (zone of inhibition, ZOI), minimum inhibitory concentration (MIC), and minimum bactericidal/fungicidal concentration (MBC/MFC). The complete results are presented in Table 8. Statistical significance was evaluated by two-way ANOVA with Tukey post-hoc test, and the key statistical outcomes are summarized in Table 9. A comparison with relevant published data for similar compounds is provided in Table 10.| Organism | ZOI (mm) blank NC | ZOI (mm) Cur-NC | ZOI (mm) free Cur | MIC (µg mL−1) blank NC | MIC (µg mL−1) Cur-NC | MIC (µg mL−1) free Cur | MBC/MFC blank NC | MBC/MFC Cur-NC |
|---|---|---|---|---|---|---|---|---|
| a NC = nanocomposite; Blank NC = Fe3O4@ZIF-8@CS; Cur-NC = Cur/Fe3O4@ZIF-8@CS; free Cur = free curcumin in 0.5% DMSO (v/v); NI = no measurable inhibition zone. Superscripts (a–c) within column: significant differences between microbial species (p < 0.05); (A and B) within row: significant differences between treatments (p < 0.05). | ||||||||
| S. agalactiae | 16.00 ± 2.00aB | 20.00 ± 2.00aA | 8.33 ± 1.53 | 41.70 ± 18.01c | 11.07 ± 3.93c | 750 ± 62.5 | 162.53 ± 153.48 | 62.47 ± 54.16 |
| S. aureus | 15.33 ± 1.53abB | 18.67 ± 3.60 abA | 7.67 ± 1.15 | 46.00 ± 28.06c | 18.57 ± 11.54c | 500 ± 46.9 | 72.93 ± 47.71 | 52.10 ± 18.01 |
| E. coli | 12.00 ± 2.00abB | 14.67 ± 3.06bA | NI | 13.33 ± 3.93c | 13.33 ± 3.92c | >1000 | 145.83 ± 59.47 | 104.17 ± 36.05 |
| K. pneumoniae | 10.67 ± 3.06bB | 14.67 ± 3.06bA | NI | 104.17 ± 36.08a | 104.17 ± 36.08a | >1000 | 104.17 ± 36.08 | 104.17 ± 36.08 |
| C. albicans | 11.33 ± 5.33abB | 18.67 ± 3.06 abA | 6.67 ± 2.08 | 83.33 ± 36.08ab | 83.33 ± 36.08b | 875 ± 72.9 | 83.33 ± 36.08 | 83.33 ± 36.08 |
| Statistical parameter | ZOI (mm) | MIC (µg mL−1) | MBC/MFC (µg mL−1) |
|---|---|---|---|
| a Significant at p < 0.05. NS = not significant. Two-way ANOVA with Tukey post-hoc test. | |||
| Treatment effect (p-value) | <0.001a | 0.228 (NS) | 0.142 (NS) |
| Microorganism effect (p-value) | 0.022a | <0.001a | 0.074 (NS) |
| Interaction effect (p-value) | 0.692 (NS) | 0.678 (NS) | 0.702 (NS) |
| Most susceptible organism | S. agalactiae | S. agalactiae | S. aureus |
| Most resistant organism | K. pneumoniae | K. pneumoniae | K. pneumoniae/E. coli |
| Post-hoc Tukey key comparison | S. agalactiae vs. K. pneumoniae (p = 0.037) | Species differ markedly | No significant pairwise |
| Study/compound | Organism | ZOI (mm) | MIC | MBC |
|---|---|---|---|---|
| Present study: Cur/Fe3O4@ZIF-8@CS | S. agalactiae | 20.00 ± 2.00 | 11.07 ± 3.93 µg mL−1 | 62.47 ± 54.16 µg mL−1 |
| Present study: Cur/Fe3O4@ZIF-8@CS | S. aureus | 18.67 ± 3.60 | 18.57 ± 11.54 µg mL−1 | 52.10 ± 18.01 µg mL−1 |
| Present study: Cur/Fe3O4@ZIF-8@CS | E. coli | 14.67 ± 3.06 | 13.33 ± 3.92 µg mL−1 | 104.17 ± 36.05 µg mL−1 |
| Present study: Cur/Fe3O4@ZIF-8@CS | K. pneumoniae | 14.67 ± 3.06 | 104.17 ± 36.08 µg mL−1 | 104.17 ± 36.08 µg mL−1 |
| El-Khawaga et al. 2024 (MgFe2O4 NPs, 30 µg mL−1)50 | S. aureus | 23 | 2.5 µg mL−1 | — |
| El-Khawaga et al. 2024 (MgFe2O4 NPs, 30 µg mL−1)50 | E. coli | 26 | 1.25 µg mL−1 | — |
| Rahvar et al. 2021 (Zn-BDC MOF)51 | S. aureus | 8 | — | — |
| Rahvar et al. 2021 (Zn-BDC MOF)51 | E. coli | 9 | — | — |
| Wada et al. 2021 (free curcumin, 100 mg mL−1)58 | S. aureus | 13 | — | — |
| Wada et al. 2021 (free curcumin, 100 µg mL−1)58 | K. pneumoniae | 7 | — | — |
| Górski et al. 2022 (free curcumin)52 | S. aureus | — | 0.046 mg mL−1 (mean) | 0.19 mg mL−1 (mean) |
| Górski et al. 2022 (free curcumin)52 | E. coli | — | 0.885 mg mL−1 (mean) | 3.54 mg mL−1 (mean) |
| Górski et al. 2022 (free curcumin)52 | K. pneumoniae | — | 1.04 mg mL−1 (mean) | >2.5 mg mL−1 |
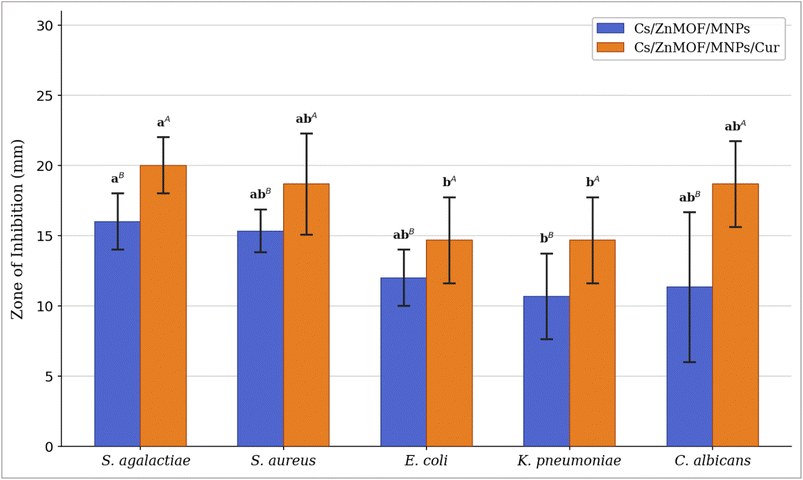
Among all tested organisms, S. agalactiae was the most susceptible. The blank nanocomposite produced a ZOI of 16.00 ± 2.00 mm, and this increased significantly to 20.00 ± 2.00 mm with curcumin loading (p < 0.05). K. pneumoniae showed the smallest ZOI values: 10.67 ± 3.06 mm (blank) and 14.67 ± 3.06 mm (curcumin-loaded). S. aureus showed ZOI values of 15.33 ± 1.53 mm (blank) and 18.67 ± 3.60 mm (curcumin-loaded). A notable result was obtained for C. albicans: the blank nanocomposite produced a ZOI of only 11.33 ± 5.33 mm, while the curcumin-loaded system more than doubled this to 18.67 ± 3.06 mm.
The addition of free curcumin as a parallel experimental control group provides direct, within-experiment quantitative evidence for the antimicrobial advantage of nanoencapsulation. Free curcumin at 1000 µg mL−1 produced no detectable inhibition zone against E. coli or K. pneumoniae, and only modest ZOI values of 8.33 ± 1.53 mm (S. agalactiae), 7.67 ± 1.15 mm (S. aureus), and 6.67 ± 2.08 mm (C. albicans). By contrast, the curcumin-loaded nanocomposite at the same total curcumin-equivalent concentration produced ZOI values of 20.00 ± 2.00, 18.67 ± 3.60, 14.67 ± 3.06, 14.67 ± 3.06, and 18.67 ± 3.06 mm for the five respective organisms. This represents a 1.5- to >2-fold amplification in ZOI relative to free curcumin, and the complete absence of activity for free curcumin against the two Gram-negative organisms versus measurable inhibition by both nanocomposite formulations confirms that the nanocarrier platform, not curcumin solubility alone, is responsible for the observed potency. These within-experiment results are consistent with the MIC advantage quantified below and with published free curcumin data.50,51
The general trend of lower ZOI values for the two Gram-negative organisms compared to the Gram-positive ones is consistent with the known structural differences between these bacterial groups. Gram-negative bacteria possess an outer membrane that acts as an additional physical barrier, limiting the penetration of many antimicrobial agents into the cell.50 The fact that both nanocomposite formulations still produced measurable inhibition against these organisms reflects the ability of nanoparticles to interact with and disrupt this outer membrane through physical contact and ion release.
To provide direct visual evidence of the inhibitory activity observed across all five tested organisms, representative agar well diffusion plate photographs are presented in Fig. 10. The images show clearly delineated zones of inhibition for Fe3O4@ZIF-8@CS, Cur/Fe3O4@ZIF-8@CS, and the positive controls after 24 h incubation at 37 °C, with ZOI diameters annotated directly on the photographs. The complete absence of any measurable inhibition zone for the 0.5% DMSO vehicle control confirms that the observed antimicrobial effects are attributable solely to the test formulations and not to the solvent. The photographic ZOI measurements are in full quantitative agreement with the mean ± SD values reported in Table 8.
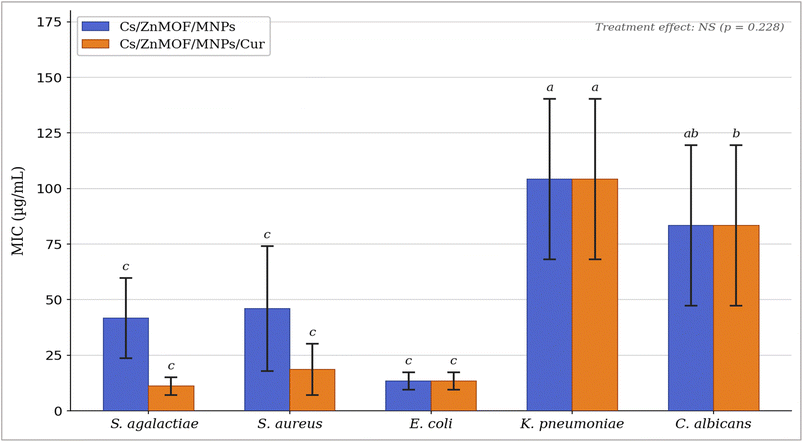
S. agalactiae was the most sensitive organism to the curcumin-loaded nanocomposite, with an MIC of 11.07 ± 3.93 µg mL−1. This was the lowest MIC value recorded across the entire study and was significantly lower than the MIC for K. pneumoniae (p < 0.001, lowercase superscripts c vs. a). The blank nanocomposite required a considerably higher concentration of 41.70 ± 18.01 µg mL−1 to inhibit S. agalactiae, representing a roughly 3.8-fold reduction in the inhibitory concentration with curcumin loading.
S. aureus also showed a meaningful improvement in MIC with curcumin loading: from 46.00 ± 28.06 µg mL−1 (blank) to 18.57 ± 11.54 µg mL−1 (curcumin-loaded), a 2.5-fold reduction. This is clinically relevant given the importance of S. aureus as a leading cause of wound infections and hospital-acquired infections. Both MIC values fall in the range of low superscript c, sharing the same statistical grouping with S. agalactiae and E. coli, indicating these three organisms were significantly more susceptible than K. pneumoniae and C. albicans.
E. coli showed equal MIC values for both formulations: 13.33 ± 3.93 µg mL−1 (blank) and 13.33 ± 3.92 µg mL−1 (curcumin-loaded). This indicates that for this organism, the blank nanocomposite was already sufficient to inhibit growth at this concentration and curcumin addition did not lower the inhibitory threshold further. In contrast, K. pneumoniae was the most resistant organism, with MIC values of 104.17 ± 36.08 µg mL−1 for both formulations. K. pneumoniae is intrinsically resistant to many antimicrobials due to its efficient outer membrane barrier and the frequent presence of beta-lactamase enzymes in clinical isolates, and this is reflected in the high MIC values observed here. C. albicans required 83.33 ± 36.08 µg mL−1 to inhibit growth for both treatments, confirming moderate susceptibility of this fungal species, as shown in Fig. 11.
Direct experimental comparison with the free curcumin control group reveals a dramatic advantage of nanoencapsulation. Free curcumin required >500 µg mL−1 for inhibition of Gram-positive organisms and exceeded >1000 µg mL−1 for both Gram-negative species and C. albicans. The corresponding MIC reductions achieved by Cur/Fe3O4@ZIF-8@CS relative to free curcumin are: >45-fold for S. agalactiae (11.07 vs. >500 µg mL−1), >27-fold for S. aureus (18.57 vs. >500 µg mL−1), >75-fold for E. coli (13.33 vs. >1000 µg mL−1), and >10-fold for K. pneumoniae (104.17 vs. >1000 µg mL−1). These within-experiment MIC reductions directly validate that the Cur/Fe3O4@ZIF-8@CS nanoplatform overcomes the inherent bioavailability and aqueous solubility limitations of free curcumin through encapsulation-mediated delivery, concentration at the bacterial cell surface, and protection from premature chemical degradation in the assay medium.
When these MIC values are compared against published data for the individual components used in isolation, the advantage of the nanocomposite approach becomes apparent. Górski et al. (2022) reported that free curcumin showed a mean MIC of 0.885 mg mL−1 (885 µg mL−1) against E. coli which is approximately 66 times higher than the MIC of the curcumin-loaded nanocomposite (13.33 µg mL−1) found in the present study.52 Against S. aureus, Górski et al. reported a mean free curcumin MIC of 0.046 mg mL−1 (46 µg mL−1), compared to 18.57 µg mL−1 for Cur/Fe3O4@ZIF-8@CS showing a 2.5-fold improvement. For K. pneumoniae, free curcumin MIC was reported as 1.04 mg mL−1 (1040 µg mL−1), compared to 104.17 µg mL−1 here is a 10-fold reduction. These improvements confirm that nanoencapsulation substantially enhances the delivery and effectiveness of curcumin, even against the more resistant organisms.
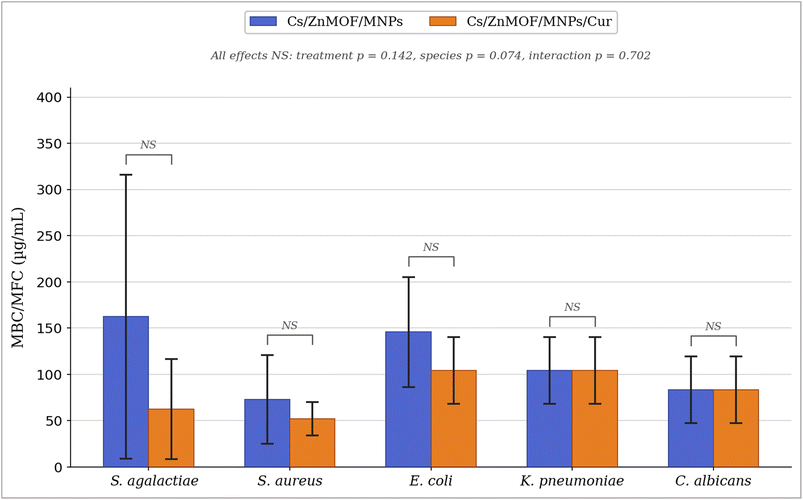
The lowest MBC values were found for S. aureus: 72.93 ± 47.71 µg mL−1 (blank nanocomposite) and 52.10 ± 18.01 µg mL−1 (curcumin-loaded). The reduction in MBC with curcumin loading reflects the additional intracellular killing action of curcumin once it is internalized by bacteria that have already been weakened by the nanoparticle framework and chitosan components. S. agalactiae showed the largest absolute reduction in MBC between the two formulations: from 162.53 ± 153.48 µg mL−1 (blank) to 62.47 ± 54.16 µg mL−1 (curcumin-loaded), a 2.6-fold improvement, although the wide standard deviations suggest variability between replicates for this organism.
E. coli required higher concentrations for bactericidal activity: 145.83 ± 59.47 µg mL−1 (blank) and 104.17 ± 36.05 µg mL−1 (curcumin-loaded), while K. pneumoniae showed equal MBC and MIC values of 104.17 ± 36.08 µg mL−1 for both formulations. When MBC equals MIC, the compound acts as a bactericide rather than a bacteriostat at the inhibitory concentration; meaning that the concentration required to stop growth is also sufficient to kill the bacteria. This bactericidal mode of action at the MIC is pharmacologically favourable, particularly for infections in immunocompromised patients, where bacteriostatic drugs alone may be insufficient to achieve clinical cure.53 C. albicans also showed equal MIC and MFC values of 83.33 ± 36.08 µg mL−1 for both formulations, confirming fungicidal activity at the inhibitory concentration.
Comparing the MBC values with published free curcumin data again demonstrates the nanocomposite advantage. Górski et al. (2022)52 reported a mean MBC of 3.54 mg mL−1 (3540 µg mL−1) for free curcumin against E. coli, compared to 104.17 µg mL−1 achieved here with Cur/Fe3O4@ZIF-8@CS that is a roughly 34-fold reduction in the concentration needed to kill the bacteria. Against S. aureus, Górski et al. found a mean MBC of 0.19 mg mL−1 (190 µg mL−1) for free curcumin, compared to 52.10 µg mL−1 in the present study which is a 3.6-fold improvement. These comparisons confirm that the nanocomposite system substantially reduces the bactericidal dose compared to free curcumin.
Fe3O4 magnetic nanoparticles contribute to antimicrobial activity through three main mechanisms. First, the nanoparticles make direct physical contact with bacterial cell membranes and disrupt their structural integrity, causing leakage of cellular contents. Second, iron ions released from the nanoparticle surface interfere with bacterial metalloenzymes and disrupt key metabolic pathways. Third, Fe3O4 nanoparticles catalyze the generation of reactive oxygen species (ROS) including hydroxyl radicals and superoxide through Fenton-like reactions, which causes oxidative damage to bacterial DNA, membrane lipids, and proteins simultaneously.54 Additionally, Fe3O4 nanoparticles are inexpensive and have low toxicity toward mammalian cells, which makes them suitable for incorporation into therapeutic formulations.55
Chitosan, which forms the outer coating layer of the nanocomposite, contributes to antimicrobial activity mainly through its positive surface charge. At physiological pH, the protonated amine groups (–NH3+) of chitosan bind strongly to the negatively charged components of bacterial and fungal cell walls including phospholipids, lipopolysaccharides (in Gram-negative bacteria), and peptidoglycans (in Gram-positive bacteria) through electrostatic attraction. This interaction disrupts membrane permeability and causes cell lysis. Beyond its direct antimicrobial action, chitosan serves as a stabilizing matrix that prevents the Fe3O4 nanoparticles from aggregating and maintains the nanocomposite as a stable, well-dispersed suspension, ensuring consistent and reproducible contact with microbial cells.17,18
The zinc-based MOF (ZnMOF) framework contributes to antimicrobial activity primarily through the release of Zn2+ ions as the framework gradually degrades under physiological conditions. Zinc ions are well-documented antimicrobials: they inhibit the active sites of key bacterial enzymes, block transport proteins in the cell membrane, interfere with DNA replication, and increase membrane permeability. The carboxylate (COO−) and amine groups on the MOF organic linkers also interact directly with bacterial cell surfaces, contributing additional disruption to cell wall integrity and promoting ROS generation by zinc-mediated redox activity.56
Curcumin, as the active pharmaceutical ingredient loaded within the MOF pores, adds its own well-studied antimicrobial mechanisms to these framework effects. Inside the microbial cell, curcumin disrupts DNA replication by intercalating into double-stranded DNA, alters gene expression by modulating transcription factor activity, and damages the cell membrane by interacting with membrane phospholipids and increasing membrane fluidity. Curcumin also generates intracellular ROS, amplifying the oxidative stress already initiated by the Fe3O4 component.57 The key advantage of nanoencapsulation within the ZIF-8 pores is that curcumin is protected from chemical degradation and is released in a controlled manner directly at the site of microbial contact, rather than degrading in the surrounding medium before it can reach the bacterial cell which is the main reason why free curcumin requires concentrations orders of magnitude higher to achieve equivalent effects.
Taken together, the results of this study demonstrate that Cur/Fe3O4@ZIF-8@CS is a multifunctional antimicrobial system in which all four components including chitosan, the zinc MOF, Fe3O4 magnetic nanoparticles, and curcumin work together through complementary and synergistic mechanisms. The nanocomposite is effective against both Gram-positive and Gram-negative bacteria as well as the fungal pathogen C. albicans, and it achieves this activity at concentrations that are dramatically lower than those required by any individual component alone, as illustrated in Table 10.
3.6 Concentration-dependent cytotoxicity and IC50 determination in mouse fibroblast cells
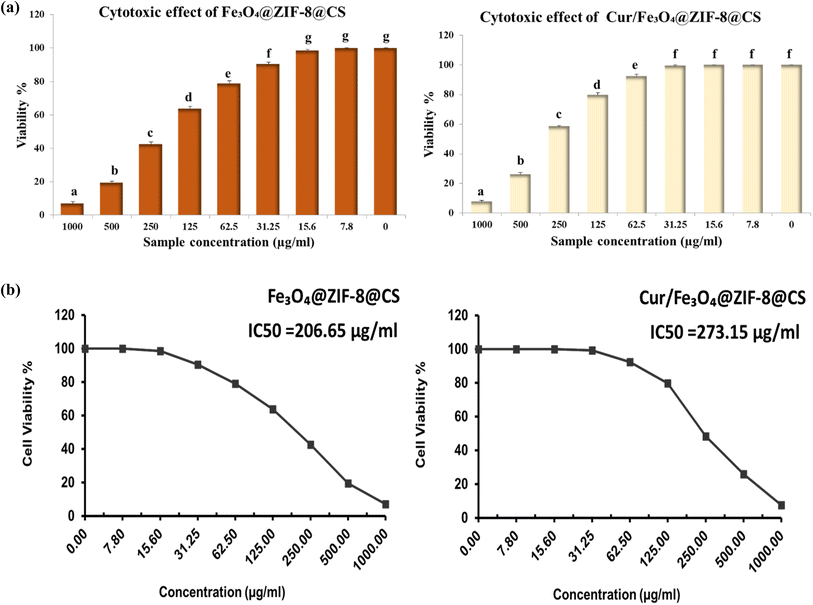
A concentration dependent cytotoxic effect was recorded on exposure the mouse fibroblast cell line to the Fe3O4@ZIF-8@CS and Cur/Fe3O4@ZIF-8@CS as illustrated in Fig. 13a. Whereas the viability% of the cells decreased with increasing the concentration, the lowest viability% (7.10% and 7.67%) for Fe3O4@ZIF-8@CS and Cur/Fe3O4@ZIF-8@CS respectively was reported at 1000 µg mL−1. This result is similar to the results obtained by Tamames-Tabar et al. (2014)59 and Ettlinger et al., (2020).41 The cytotoxic effect at higher concentration may be resulted from the triggered autophagic effect by ZIF-8, that enhance its degradation and subsequently Zn2+ leakages that enhances autophagy and induces generation of ROS.60Additionally, the IC50 value of Fe3O4@ZIF-8@CS reached bout (206.65 µg mL−1) and (273.15 µg mL−1) for Cur/Fe3O4@ZIF-8@ CS, indicating the higher cytotoxic effect of Fe3O4@ZIF-8@CS than Cur/Fe3O4@ZIF-8@ CS Fig. 13b. This may be attributed to the suitable size of the Cur/Fe3O4@ZIF-8@ CS that increasing their passage through the cell membrane; moreover, the PH-responsiveness of ZIF-8 may elevate the local concentration of curcumin.61 It is reported that ZIF-8 has the ability to improve the rate of drug release within an acidic environment and enhancing drug delivery systems.42
3.7 Stability study
The physicochemical stability of the optimized Cur/Fe3O4@ZIF-8@CS nanocomposite was systematically monitored over two months under both long-term (25 ± 2 °C) and accelerated (40 ± 2 °C/75 ± 5% RH, ICH Q1A(R2)) storage conditions. Four critical quality attributes were tracked at T0, T1 (1 month), and T2 (2 months): hydrodynamic particle size, PDI, zeta potential, and curcumin content retention. The complete dataset is summarized in Table 11.| Parameter | Storage condition | Time point | ||
|---|---|---|---|---|
| T0 (day 0) | T1 (1 month) | T2 (2 months) | ||
| Particle size (nm) | 25 ± 2 °C | 228.6 ± 4.7 | 234.2 ± 5.1 | 241.8 ± 5.8 |
| 40 ± 2 °C/75 ± 5% RH | 228.6 ± 4.7 | 248.7 ± 6.2 | 263.5 ± 7.4 | |
| PDI | 25 ± 2 °C | 0.218 ± 0.010 | 0.224 ± 0.011 | 0.231 ± 0.012 |
| 40 ± 2 °C/75 ± 5% RH | 0.218 ± 0.010 | 0.241 ± 0.013 | 0.258 ± 0.015 | |
| Zeta potential (mV) | 25 ± 2 °C | +31.5 ± 1.4 | +30.1 ± 1.2 | +28.7 ± 1.5 |
| 40 ± 2 °C/75 ± 5% RH | +31.5 ± 1.4 | +27.3 ± 1.6 | +24.9 ± 1.8 | |
| Curcumin retention (%) | 25 ± 2 °C | 100.0 ± 0.0 | 97.4 ± 1.2 | 94.6 ± 1.8 |
| 40 ± 2 °C/75 ± 5% RH | 100.0 ± 0.0 | 92.8 ± 2.1 | 87.9 ± 2.6 | |
PDI followed the same trend, increasing from 0.218 ± 0.010 at T0 to 0.231 ± 0.012 (25 °C) and 0.258 ± 0.015 (40 °C/75% RH) at T2. All values remained below the homogeneity limit of 0.30,66 confirming that the nanocomposite maintained a narrow size distribution during storage, consistent with previous reports for chitosan-coated systems.63
The decrease is likely due to partial deprotonation of chitosan amine groups, amine hydrolysis, and ionic screening caused by moisture uptake.31,66 Similar behavior was reported for chitosan-coated curcumin liposomes under thermal stress.63 Importantly, the positive charge remained above +24 mV, suggesting sufficient electrostatic repulsion and preserved antimicrobial interaction with bacterial membranes.63
The improved stability is attributed to encapsulation within the ZIF-8 pores and protection by the chitosan shell, which limits oxidation and moisture diffusion.31,63 Liu et al.63 reported 75.77% retention for chitosan-coated liposomal curcumin after 40 days, which is lower than the 94.6% observed here after 60 days, indicating stronger protection by the MOF structure. Faster degradation at elevated temperature agrees with previous reports for encapsulated curcumin systems,64,65 and is mainly related to thermally induced β-diketone cleavage and partial framework destabilization.31,32
4 Conclusion
This study demonstrated the successful rational design of a multifunctional curcumin-loaded magnetic MOF–chitosan nanocomposite (Cur/Fe3O4@ZIF-8@CS) as an advanced drug delivery platform. The integration of a QbD-driven Box–Behnken optimization strategy enabled a systematic and reproducible formulation process, ensuring that the optimized nanocomposite met all predefined CQAs without reliance on trial-and-error experimentation. The hierarchical assembly of Fe3O4, ZIF-8, and chitosan produced a structurally integrated system in which each component contributed complementary functionality: magnetic responsiveness, high drug-loading capacity, colloidal stability, and intrinsic antimicrobial properties, as comprehensively confirmed by DLS, ATR-FTIR, TEM, and VSM characterization.The developed nanocomposite exhibited controlled, pH-responsive drug release behavior with sustained kinetics over 72 hours, indicating its suitability for preferential drug delivery in acidic pathological microenvironments such as tumors and infection sites. Nanoencapsulation dramatically enhanced the therapeutic potential of curcumin, reducing effective MIC values by up to 66-fold relative to free curcumin and producing broad-spectrum antimicrobial activity against both Gram-positive and Gram-negative bacteria as well as the fungal pathogen Candida albicans. A two-month ICH Q1A(R2)-guided stability study confirmed that the ZIF-8/chitosan architecture provides robust physicochemical protection for the encapsulated drug under both long-term and accelerated storage conditions, with all quality attributes remaining within pharmaceutical acceptability limits.
Cytotoxicity evaluation by MTT assay on L929 mouse fibroblast cells confirmed a concentration-dependent effect, with IC50 values of 206.65 µg mL−1 for Fe3O4@ZIF-8@CS and 273.15 µg mL−1 for Cur/Fe3O4@ZIF-8@CS; the higher IC50 of the curcumin-loaded formulation indicates lower cytotoxicity toward normal cells, supporting an acceptable safety profile at therapeutic concentrations.
The present characterization suite encompassing DLS, ATR-FTIR, TEM, VSM, in vitro release, antimicrobial profiling, cytotoxicity, and two-month stability assessment constitutes a comprehensive multi-technique evaluation consistent with leading publications in the magnetic MOF nanocarrier field. Future work should incorporate X-ray photoelectron spectroscopy (XPS) to elucidate surface chemical states, and nitrogen adsorption–desorption isotherms (BET analysis) to quantify specific surface area and pore size distribution of the ZIF-8 shell before and after drug loading; these measurements would provide deeper mechanistic insight into host–guest encapsulation efficiency and guide further rational optimization. Additionally, in vivo pharmacokinetic studies, biocompatibility assessment under physiological conditions, and evaluation of magnetic field-guided targeting in tumor models are warranted to support translational development of this platform toward clinical applications.
Ethical statement
Not applicable, as the study did not apply to human or animal studies. The article does not include any studies on human participants or animals conducted by any of the authors.Consent for publication
The authors confirm: this work represents original research that has not been previously published in any form. This manuscript has not been submitted for review or publication consideration elsewhere. The publication has received approval from all co-authors.Author contributions
Donia Hemida: conceptualization, methodology, investigation, data curation, writing – original draft; Nabila Shehata: methodology, validation, formal analysis, writing – review & editing; Nada Adel El-Gaddawy: investigation, resources, data curation; Haifa E. Alfassam: formal analysis, visualization, writing – review & editing; Samar M. Mahgoub: supervision, methodology, writing – review & editing; Rehab Mahmoud: conceptualization, supervision, project administration, writing – review & editing.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The datasets generated and/or analyzed during this study are available from the corresponding author upon reasonable request.Acknowledgements
The authors acknowledge Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R400), Princess Nourah Bint Abdulrahman University, Riyadh, Saudi Arabia.References
- M. Naghavi, S. E. Vollset, K. S. Ikuta, L. R. Swetschinski, A. P. Gray, E. E. Wool, G. R. Aguilar, T. Mestrovic, G. Smith and C. Han, Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050, Lancet, 2024, 404, 1199–1226 CrossRef PubMed.
- C. J. L. Murray, K. S. Ikuta, F. Sharara, L. Swetschinski, G. R. Aguilar, A. Gray, C. Han, C. Bisignano, P. Rao and E. Wool, Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis, Lancet, 2022, 399, 629–655 CrossRef CAS PubMed.
- R. Ranjbar and M. Alam, Antimicrobial Resistance Collaborators (2022). Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis, Lancet, 2022, 399(10325), 629–655 CrossRef.
- F. Bray, M. Laversanne, H. Sung, J. Ferlay, R. L. Siegel, I. Soerjomataram and A. Jemal, Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries, Ca-Cancer J. Clin., 2024, 74, 229–263, DOI:10.3322/caac.21834.
- D. Avinash, B. Mrudula, W. Milind, N. Vanashri and S. Ravindranath, A review on nanocomposite drug delivery, J. Drug Delivery Ther., 2019, 9, 529–536 CAS.
- Y. Yao, Y. Zhou, L. Liu, Y. Xu, Q. Chen, Y. Wang, S. Wu, Y. Deng, J. Zhang and A. Shao, Nanoparticle-based drug delivery in cancer therapy and its role in overcoming drug resistance, Front. Mol. Biosci., 2020, 7, 193 CrossRef CAS PubMed.
- J. Nel, K. Elkhoury, É. Velot, A. Bianchi, S. Acherar, G. Francius, A. Tamayol, S. Grandemange and E. Arab-Tehrany, Functionalized liposomes for targeted breast cancer drug delivery, Bioact. Mater., 2023, 24, 401–437 CAS.
- P. Horcajada, R. Gref, T. Baati, P. K. Allan, G. Maurin, P. Couvreur, G. Férey, R. E. Morris and C. Serre, Metal–organic frameworks in biomedicine, Chem. Rev., 2012, 112, 1232–1268 CrossRef CAS PubMed.
- I. A. Lazaro and R. S. Forgan, Application of zirconium MOFs in drug delivery and biomedicine, Coord. Chem. Rev., 2019, 380, 230–259 CrossRef.
- Y. Hou, C. Zhu, G. Ban, Z. Shen, Y. Liang, K. Chen, C. Wang and H. Shi, Advancements and challenges in the application of metal-organic framework (MOF) nanocomposites for tumor diagnosis and treatment, Int. J. Nanomed., 2024, 6295–6317 CrossRef CAS.
- Z. Li, Y. Shao, Y. Yang and J. Zan, Zeolitic imidazolate framework-8: a versatile nanoplatform for tissue regeneration, Front. Bioeng. Biotechnol., 2024, 12, 1386534 CrossRef PubMed.
- H. N. Abdelhamid, Zeolitic imidazolate frameworks (ZIF-8) for biomedical applications: a review, Curr. Med. Chem., 2021, 28, 7023–7075 CAS.
- T. Vangijzegem, V. Lecomte, I. Ternad, L. Van Leuven, R. N. Muller, D. Stanicki and S. Laurent, Superparamagnetic iron oxide nanoparticles (SPION): from fundamentals to state-of-the-art innovative applications for cancer therapy, Pharmaceutics, 2023, 15, 236 CrossRef CAS PubMed.
- Q. T. H. Shubhra, Iron oxide nanoparticles in magnetic drug targeting and ferroptosis-based cancer therapy, Medical Review, 2023, 3, 444–447 CrossRef PubMed.
- E. Otieno, Y. Huang, N. Li, T. Li, M. Wang, X. Qiu and X. Xiao, Utilization of superparamagnetic iron oxide nanoparticles (SPIONs) as a vector for drug delivery, Appl. Nanosci., 2023, 13, 6191–6216 CrossRef CAS.
- Y. Herdiana, N. Wathoni, S. Shamsuddin and M. Muchtaridi, Drug release study of the chitosan-based nanoparticles, Heliyon, 2022, 8(1), e08674 CrossRef CAS PubMed.
- K. Jafernik, A. Ładniak, E. Blicharska, K. Czarnek, H. Ekiert, A. E. Wiącek and A. Szopa, Chitosan-based nanoparticles as effective drug delivery systems—a review, Molecules, 2023, 28, 1963 CrossRef CAS PubMed.
- M. Wang, Y. Wang, G. Chen, H. Gao and Q. Peng, Chitosan-based multifunctional biomaterials as active agents or delivery systems for antibacterial therapy, Bioengineering, 2024, 11, 1278 CrossRef CAS PubMed.
- E. Akdaşçi, H. Duman, F. Eker, M. Bechelany and S. Karav, Chitosan and its nanoparticles: A multifaceted approach to antibacterial applications, Nanomaterials, 2025, 15, 126 CrossRef PubMed.
- K. M. Nelson, J. L. Dahlin, J. Bisson, J. Graham, G. F. Pauli and M. A. Walters, The essential medicinal chemistry of curcumin: miniperspective, J. Med. Chem., 2017, 60, 1620–1637 Search PubMed.
- J. Yakubu and A. V Pandey, Innovative delivery systems for curcumin: Exploring nanosized and conventional formulations, Pharmaceutics, 2024, 16, 637 CrossRef CAS PubMed.
- M. Zoghi, M. Pourmadadi, F. Yazdian, M. N. Nigjeh, H. Rashedi and R. Sahraeian, Synthesis and characterization of chitosan/carbon quantum dots/Fe2O3 nanocomposite comprising curcumin for targeted drug delivery in breast cancer therapy, Int. J. Biol. Macromol., 2023, 249, 125788 CrossRef CAS PubMed.
- Y. Yan, Y. Sun, Y. Li, Z. Wang, L. Xue and F. Wang, Advancing cancer therapy: Nanomaterial-based encapsulation strategies for enhanced delivery and efficacy of curcumin, Mater. Today Bio, 2025, 33, 101963 CrossRef CAS PubMed.
- E. M. Alshammari, Curcumin-based biocompatible nanocarriers: a contemporary perspective in functional foods and biomedical applications, Discover Nano, 2025, 20, 226 CrossRef PubMed.
- R. Maurya, S. Ramteke and N. K. Jain, Quality by design (QbD) approach-based development of optimized nanocarrier to achieve quality target product profile (QTPP)-targeted lymphatic delivery, Nanotechnology, 2024, 35, 265101 CrossRef CAS PubMed.
- J. G. Duarte, M. G. Duarte, A. P. Piedade and F. Mascarenhas-Melo, Rethinking pharmaceutical industry with quality by design: application in research, development, manufacturing, and quality assurance, AAPS J., 2025, 27, 96 CrossRef PubMed.
- L. X. Yu, G. Amidon, M. A. Khan, S. W. Hoag, J. Polli, G. K. Raju and J. Woodcock, Understanding pharmaceutical quality by design, AAPS J., 2014, 16, 771–783 CrossRef CAS PubMed.
- A. K. Sah and P. K. Suresh, Loteprednol etabonate nanoparticles: optimization via Box-Behnken design response surface methodology and physicochemical characterization, Curr. Drug Delivery, 2017, 14, 676–689 CAS.
- S. L. C. Ferreira, R. E. Bruns, H. S. Ferreira, G. D. Matos, J. M. David, G. C. Brandão, E. G. P. da Silva, L. A. Portugal, P. S. Dos Reis and A. S. Souza, Box-Behnken design: An alternative for the optimization of analytical methods, Anal. Chim. Acta, 2007, 597, 179–186 CrossRef CAS PubMed.
- P. Majewski and B. Thierry, Functionalized magnetite nanoparticles—synthesis, properties, and bio-applications, Crit. Rev. Solid State Mater. Sci., 2007, 32, 203–215 CrossRef CAS.
- K. I. Priyadarsini, The chemistry of curcumin: from extraction to therapeutic agent, Molecules, 2014, 19, 20091–20112 CrossRef.
- M. Kharat, Z. Du, G. Zhang and D. J. McClements, Physical and chemical stability of curcumin in aqueous solutions and emulsions: impact of pH, temperature, and molecular environment, J. Agric. Food Chem., 2017, 65, 1525–1532 CrossRef CAS PubMed.
- H. Zheng, Y. Zhang, L. Liu, W. Wan, P. Guo, A. M. Nystrom and X. Zou, One-pot synthesis of metal–organic frameworks with encapsulated target molecules and their applications for controlled drug delivery, J. Am. Chem. Soc., 2016, 138, 962–968 CrossRef CAS PubMed.
- A. Mittal, S. Gandhi and I. Roy, Mechanistic interaction studies of synthesized ZIF-8 nanoparticles with bovine serum albumin using spectroscopic and molecular docking approaches, Sci. Rep., 2022, 12, 10331 CrossRef CAS PubMed.
- R. E. Des Bouillons-Gamboa, G. Montes de Oca, J. R. V. Baudrit, L. C. Ríos Duarte, M. Lopretti, M. Rentería Urquiza, J. M. Zúñiga-Umaña, F. Barreiro and P. Vázquez, Synthesis of chitosan nanoparticles (CSNP): effect of CH-CH-TPP ratio on size and stability of NPs, Front. Chem., 2024, 12, 1469271 CrossRef PubMed.
- C. Mircioiu, V. Voicu, V. Anuta, A. Tudose, C. Celia, D. Paolino, M. Fresta, R. Sandulovici and I. Mircioiu, Mathematical modeling of release kinetics from supramolecular drug delivery systems, Pharmaceutics, 2019, 11, 140 CrossRef CAS PubMed.
- Y. Zhang, M. Huo, J. Zhou, A. Zou, W. Li, C. Yao and S. Xie, DDSolver: an add-in program for modeling and comparison of drug dissolution profiles, AAPS J., 2010, 12, 263–271 CrossRef CAS PubMed.
- E. K. Droepenu, E. Amenyogbe, M. A. Boatemaa and E. Opoku, Study of the antimicrobial activity of zinc oxide nanostructures mediated by two morphological structures of leaf extracts of Eucalyptus robusta Sm, Heliyon, 2024, 10(4), e25590 CrossRef CAS PubMed.
- T. Mosmann, Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays, J. Immunol. Methods, 1983, 65, 55–63 CrossRef CAS PubMed.
- I. C. for Harmonisation, ICH Q1A (R2): Stability testing of new drug substances and products, International Council for Harmonisation, 2003, vol. 1, pp. 1–24 Search PubMed.
- R. Ettlinger, N. Moreno, N. Ziółkowska, A. Ullrich, H. Krug von Nidda, D. Jirák, K. Kerl and H. Bunzen, In vitro studies of Fe3O4-ZIF-8 core–shell nanoparticles designed as potential theragnostics, Part. Part. Syst. Char., 2020, 37, 2000185 CrossRef CAS.
- J. Kong, S. Xu, Y. Dai, Y. Wang, Y. Zhao and P. Zhang, Study of the Fe3O4@ ZIF-8@ sor composite modified by tannic acid for the treatment of sorafenib-resistant hepatocellular carcinoma, ACS Omega, 2023, 8, 39174–39185 CrossRef CAS PubMed.
- S. K. Filippov, R. Khusnutdinov, A. Murmiliuk, W. Inam, L. Y. Zakharova, H. Zhang and V. V Khutoryanskiy, Dynamic light scattering and transmission electron microscopy in drug delivery: a roadmap for correct characterization of nanoparticles and interpretation of results, Mater. Horiz., 2023, 10, 5354–5370 RSC.
- B. K. Wilson and R. K. Prud’homme, Nanoparticle size distribution quantification from transmission electron microscopy (TEM) of ruthenium tetroxide stained polymeric nanoparticles, J. Colloid Interface Sci., 2021, 604, 208–220 CrossRef CAS PubMed.
- Wahajuddin and S. Arora, Superparamagnetic iron oxide nanoparticles: magnetic nanoplatforms as drug carriers, Int. J. Nanomed., 2012, 3445–3471 CrossRef CAS PubMed.
- H. M. Fahmy, S. Shekewy, F. A. Elhusseiny and A. Elmekawy, Enhanced biocompatibility by evaluating the cytotoxic and genotoxic effects of magnetic iron oxide nanoparticles and chitosan on Hepatocellular Carcinoma Cells (HCC), Cell Biochem. Biophys., 2024, 82, 1027–1042 CrossRef CAS PubMed.
- A. Criveanu, F. Dumitrache, C. Fleaca, L. Gavrila-Florescu, I. Lungu, I. P. Morjan, V. Socoliuc and G. Prodan, Chitosan-coated iron oxide nanoparticles obtained by laser pyrolysis, Appl. Surf. Sci. Adv., 2023, 15, 100405 CrossRef.
- F. Parsa, M. Setoodehkhah and S. M. Atyabi, Design, fabrication and characterization of a magnetite-chitosan coated iron-based metal–organic framework (Fe 3 O 4@ chitosan/MIL-100 (Fe)) for efficient curcumin delivery as a magnetic nanocarrier, RSC Adv., 2025, 15, 18518–18534 RSC.
- S. Hansapaiboon, B. P. Bulatao, F. N. Sorasitthiyanukarn, P. Jantaratana, N. Nalinratana, O. Vajragupta, P. Rojsitthisak and P. Rojsitthisak, Fabrication of Curcumin Diethyl γ-Aminobutyrate-Loaded Chitosan-Coated Magnetic Nanocarriers for Improvement of Cytotoxicity against Breast Cancer Cells, Polymers, 2022, 14, 5563 CrossRef CAS PubMed.
- A. M. El-Khawaga, M. Ayman, O. Hafez and R. E. Shalaby, Photocatalytic, antimicrobial and antibiofilm activities of MgFe2O4 magnetic nanoparticles, Sci. Rep., 2024, 14, 12877 CrossRef CAS PubMed.
- Y. Rahvar, N. Motakef-Kazemi and R. Hosseini Doust, Synthesis of Zn2 (BDC) 2 (DABCO) MOF by solution and solvothermal methods and evaluation of its anti-bacterial, Nanomed. Res. J., 2021, 6, 360–368 CAS.
- M. Górski, J. Niedźwiadek and A. Magryś, Antibacterial activity of curcumin–a natural phenylpropanoid dimer from the rhizomes of Curcuma longa L. and its synergy with antibiotics, Ann. Agric. Environ. Med., 2022, 29, 394–400 CrossRef.
- A. Sirelkhatim, S. Mahmud, A. Seeni, N. H. M. Kaus, L. C. Ann, S. K. M. Bakhori, H. Hasan and D. Mohamad, Review on zinc oxide nanoparticles: antibacterial activity and toxicity mechanism, Nano-Micro Lett., 2015, 7, 219–242 CrossRef CAS PubMed.
- P. Nisar, N. Ali, L. Rahman, M. Ali and Z. K. Shinwari, Antimicrobial activities of biologically synthesized metal nanoparticles: an insight into the mechanism of action, JBIC, J. Biol. Inorg. Chem., 2019, 24, 929–941 CrossRef CAS PubMed.
- M. I. Anik, M. K. Hossain, I. Hossain, A. Mahfuz, M. T. Rahman and I. Ahmed, Recent progress of magnetic nanoparticles in biomedical applications: A review, Nano Sel., 2021, 2, 1146–1186 CrossRef CAS.
- Q. Abbas, M. Zia-ur-Rehman, H. Ullah, M. A. M. Munir, M. U. Ali, A. Ali, K. Pikon and B. Yousaf, Recent advances in the detection and quantification of manufactured nanoparticles (MNPs) in complex environmental and biological matrices, J. Cleaner Prod., 2024, 471, 143454 CrossRef CAS.
- S. Naz, S. Jabeen, S. Ilyas, F. Manzoor, F. Aslam and A. Ali, Antibacterial activity of Curcuma longa varieties against different strains of bacteria, Pak. J. Bot., 2010, 42, 455–462 Search PubMed.
- N. M. Wada, A. A. Ambi, A. A. Ibrahim, S. K. Bello, A. Umar and D. T. James, Antimicrobial Activity of Extracts of Turmeric (Curcuma longa) and Garlic (Aliium sativum) Against Selected Bacterial Clinical Isolates, Mediterr. J. Infect. Microbes Antimicrob., 2021, 10(1), 6 Search PubMed.
- C. Tamames-Tabar, D. Cunha, E. Imbuluzqueta, F. Ragon, C. Serre, M. J. Blanco-Prieto and P. Horcajada, Cytotoxicity of nanoscaled metal–organic frameworks, J. Mater. Chem. B, 2014, 2, 262–271 RSC.
- M. Xu, Y. Hu, W. Ding, F. Li, J. Lin, M. Wu, J. Wu, L.-P. Wen, B. Qiu and P.-F. Wei, Rationally designed rapamycin-encapsulated ZIF-8 nanosystem for overcoming chemotherapy resistance, Biomaterials, 2020, 258, 120308 CrossRef CAS PubMed.
- C.-Y. Sun, C. Qin, X.-L. Wang, G.-S. Yang, K.-Z. Shao, Y.-Q. Lan, Z.-M. Su, P. Huang, C.-G. Wang and E.-B. Wang, Zeolitic imidazolate framework-8 as efficient pH-sensitive drug delivery vehicle, Dalton Trans., 2012, 41, 6906–6909 RSC.
- K. Maruyama, Intracellular targeting delivery of liposomal drugs to solid tumors based on EPR effects, Adv. Drug Delivery Rev., 2011, 63, 161–169 CrossRef CAS PubMed.
- Y. Liu, D. Liu, L. Zhu, Q. Gan and X. Le, Temperature-dependent structure stability and in vitro release of chitosan-coated curcumin liposome, Food Res. Int., 2015, 74, 97–105 CrossRef CAS PubMed.
- B. Zheng, X. Zhang, H. Lin and D. J. McClements, Loading natural emulsions with nutraceuticals using the pH-driven method: formation & stability of curcumin-loaded soybean oil bodies, Food Funct., 2019, 10, 5473–5484 RSC.
- C. Winarti and T. C. Sunarti, Effect of Heating on the Stability of Curcumin on Temulawak Oleoresin Encapsulated in Arrowroot Starch Nanoparticles, in E3S Web of Conferences, EDP Sciences, 2022, p. 4005 Search PubMed.
- C. Freitas and R. H. Müller, Effect of light and temperature on zeta potential and physical stability in solid lipid nanoparticle (SLN™) dispersions, Int. J. Pharm., 1998, 168, 221–229 CrossRef CAS.
- H. H. S. Food and Drug Administration, International Conference on Harmonisation; guidance on Q6A specifications: test procedures and acceptance criteria for new drug substances and new drug products: chemical substances, Federal Register, 2000, vol. 65, pp. 83041–83063 Search PubMed.
- I. H. T. Guideline, Stability testing of new drug substances and products, Q1A (R2), Current Step 4, 2003 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |