Open Access Article
Open Access ArticleDesign, synthesis, enzymatic and in vivo antihyperglycemic activity evaluation of 1,3,4-oxadiazole derivatives targeting multiple pathways in type 2 diabetes
Shereena,
Ayesha Tahirb,
Awais Ahmedb,
Muhammad Saeed Jan c,
Nouman Aslamb,
Sohail Hassan*a and
Umer Rashid
c,
Nouman Aslamb,
Sohail Hassan*a and
Umer Rashid *b
*b
aDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy and Pharmaceutical Sciences, University of Karachi, Karachi-75270, Pakistan. E-mail: shassan@uok.edu.pk
bDepartment of Chemistry, COMSATS University Islamabad, Abbottabad Campus, 22060 Abbottabad, Pakistan. E-mail: umerrashid@cuiatd.edu.pk
cDepartment of Pharmacy, University of Swabi, Anbar, 23430, Khyber Pakhtunkhwa, Pakistan
First published on 29th April 2026
Abstract
The multifactorial and complex pathophysiology of type 2 diabetes mellitus (T2DM) requires therapeutic interventions that regulate key enzymes involved in glucose homeostasis. The study described the design of a new family of oxadiazole–tetrahydropyrimidinone hybrid compounds (18–23 and 30–36), their synthesis, and the testing of these compounds as multitarget antidiabetic agents. The in vitro enzyme inhibition was performed on four key targets: α-amylase, α-glucosidase, dipeptidyl peptidase-IV, and protein tyrosine phosphatase 1B. Among the synthesized derivatives, compound 23 exhibited excellent inhibitory activity across all four tested targets, demonstrating a balanced multitarget profile, while compounds 18 and 19 showed good to moderate activity. The in vivo antihyperglycemic activity of the chosen compounds, particularly compound 23, was proven using oral glucose tolerance tests (OGTT) and chronic studies in diabetic rat models, resulting in a significant decrease in fasting blood glucose. The docking simulations revealed favorable binding affinities with all four targets, supporting the observed multitarget behavior. Together, these results demonstrate oxadiazole–tetrahydropyrimidinone hybrids as promising scaffolds for the development of balanced multitarget antidiabetic agents with potential for synergistic glycemic control through coordinated modulation of complementary metabolic pathways, offering improved therapeutic prospects in T2DM management.
Introduction
Diabetes mellitus is a multi-etiological endocrine glucose, lipid and protein metabolic dysfunction that is currently prevalent in 830 million individuals across the globe and is characterized by severe cardiovascular, renal, visual and neurological adverse effects.1,2 As a result, patients with diabetes must always maintain normal levels of blood glucose. Diabetes mellitus is categorized into a number of different types: diabetes mellitus type 1 (the autoimmune destruction of pancreatic beta cells), diabetes mellitus type 2 (insulin resistance, progressive beta-cell dysfunction), gestational diabetes (appearing during pregnancy), and prediabetes (pre-diagnostic hyperglycemia that occurs).3Type 2 diabetes mellitus (T2DM) is a multifactorial metabolic disease where some enzyme targets are of central importance to the process of glucose homeostasis and are valuable targets of intervention during the development of therapeutic interventions. The most notable among them are α-amylase and α-glucosidase that stimulate the breakdown of complex carbohydrates into absorbable monosaccharides, which are major contributors to postprandial glucose spikes. The inhibition of these enzymes slows down the digestion of carbohydrates and glucose absorption and, therefore, is an effective approach in preventing postprandial hyperglycemia.4 Acarbose5 (Fig. 1), voglibose,6 and miglitol7 are the clinically approved oral antidiabetic drugs that are used to control postprandial hyperglycemia in individuals with T2DM. The drugs act as carbohydrate analogs. Acarbose uniquely inhibits both α-glucosidase and α-amylase, disrupting carbohydrate digestion at various steps in the gastrointestinal tract. Despite their efficacy, they are limited by the side effects of the gastrointestinal tract (flatulence, abdominal discomfort, and diarrhea), which are mainly caused by the effect of undigested carbohydrates on the colon.8,9
Dipeptidyl peptidase-4 (DPP-4) is a serine protease that has been shown to degrade incretin hormones like GLP-1 and GIP, both of which are necessary to stimulate glucose-stimulated insulin secretion and inhibit glucagon secretion.10 Thus, DPP-4 inhibition increases incretin action, thus enhancing glycemic regulation and does not cause insulin secretion stimulation directly.11 Some of the DPP-4 inhibitors (gliptins) are currently marketed, such as sitagliptin, linagliptin, and anagliptin12 (Fig. 1), and are extensively used as monotherapy or with metformin or SGLT 2 inhibitors in managing T2D. The cardiovascular and renal effects of these agents have proven useful in addition to glycemic control, and other DPP-4 inhibitors (gosogliptin, gemigliptin, trelagliptin, anagliptin, and dutogliptin) are under clinical development.13
Beyond glucose absorption and incretin regulation, protein tyrosine phosphatase 1B (PTP1B) serves as a critical negative regulator of insulin and leptin signalling by dephosphorylating their respective receptors, leading to reduced insulin sensitivity and metabolic dysfunction. Inhibiting PTP1B thus holds therapeutic potential in enhancing insulin action and addressing associated metabolic syndromes.14 Structure–activity relationship studies have revealed that compounds containing hydrophobic aromatic or alkyl moieties combined with hydrophilic heterocyclic rings demonstrate enhanced binding affinity and selectivity for PTP1B. Although synthetic inhibitors, including ertiprotafib (Fig. 1), trodusquemine, and JTT-551, have advanced to clinical trials, these candidates were ultimately discontinued due to poor bioavailability and inadequate pharmacokinetic profiles, highlighting the ongoing need for rational structural modifications to develop clinically viable PTP1B inhibitors with improved drug-like properties.15
Given the multifactorial nature of diabetes mellitus and the limitations of current therapies, there is growing interest in multi-target antidiabetic strategies that can simultaneously modulate several pathological pathways.16 Numerous inhibitors targeting each of these enzymes have been synthesized and reported in the literature, along with several dual and multi-targeted inhibitors, all demonstrating promising therapeutic potential.17,18 However, the development of such agents remains an evolving field, with considerable scope for further research and optimization within this modern and promising therapeutic strategy.
Our team has recently synthesized and reported thiazolidinedione and dihydropyrimidine-based inhibitors targeting these key enzymes, offering a multitargeted approach for glycemic management by simultaneously modulating carbohydrate digestion, incretin signaling, and insulin sensitivity.19,20
The 1,3,4-oxadiazole ring is a privileged medicinal chemistry scaffold, recognized for its favorable pharmacokinetics, diverse bioactivity, and structural versatility.21,22 Its proven therapeutic relevance is evidenced by its presence in clinically approved drugs such as raltegravir, furamizole, zibotentan, tiodazosin, and nesapidil (Fig. 2). It is based on these properties that we chose the oxadiazole core as the core scaffold on which we would synthesize new derivatives.23 In this context, we report the design and synthesis of new oxadiazole-based compounds evaluated as multi-target inhibitors of α-amylase, α-glucosidase, DPP-4, and PTP1B, offering a comprehensive approach to glycemic control.
Materials and methods
General
Chemicals and solvents such as ionized solvents, used in this research were acquired through commercial stores, namely Sigma-Aldrich and Alfa Aesar, and were utilized without any further purification. Bruker 400 MHz spectrometer was used to record both the 1H and the related 13C nuclear magnetic resonance (NMR) with the use of appropriate deuterated solvents. A Stuart SMP10 melting point apparatus was used to measure melting points in open capillary tubes. Thin-layer chromatography (TLC) on silica gel 60 F254 pre-coated plates was used to monitor the progress of the reactions. A Vario EL III CHN analyzer was used to determine the elemental composition. An Agilent Technologies 1200 series high-performance liquid chromatography (HPLC) system with a C18 reversed-phase column (particle size 3.5 µm; 100 mm length × 4.6 mm internal diameter; Agilent Technologies) was used to perform liquid chromatography-mass spectrometry (LC-MS) analysis. LC-MS analysis was performed using acetonitrile and water (0.1% formic acid, 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10, v/v) under isocratic conditions.
10, v/v) under isocratic conditions.
General method for synthesis of (4-substituted) ethyl-2-oxo-6-(trifluoromethyl)-1,2,3,4-tetrahydropyrimidine-5-carboxylates (6–8)
A mixture of the aldehyde (formaldehyde 1, benzaldehyde 2, or p-methoxybenzaldehyde 3) (25 mmol), urea 4 (25 mmol), and ethyl 4,4,4-trifluoro-3-oxobutanoate 5 (30 mmol) was added to a round-bottom flask containing 60 mL of acetonitrile. Tin(II) chloride dihydrate (SnCl2·2H2O, 10 mol%) was introduced as a Lewis acid activator, and the reaction mixture was refluxed for approximately 12 hours. The reaction progress was monitored by TLC. Upon completion, the mixture was poured into ice-cold water and stirred for 10 minutes. The resulting precipitate was filtered, dried, and recrystallized from ethanol to afford the pure products 6–8.General procedure for synthesis of (4-substituted) 2-oxo-6-(trifluoromethyl)-1,2,3,4-tetrahydropyrimidine-5-carbohydrazides (10–12)
Equimolar amounts (20 mmol) of substituted tetrahydropyrimidine carboxylates (6–8) and hydrazine hydrate 9 were added to a round-bottom flask containing ethanol. The reaction mixture was refluxed for 5–6 hours until completion, as monitored by TLC. Upon completion, the solid product precipitated out, which was then filtered, dried, and recrystallized from ethanol to afford the corresponding pure hydrazide derivatives (10–12).General procedure for synthesis of 1,3,4-oxadiazole-based derivatives (14–16)
Methyl 4-formylbenzoate (13, 15 mmol) and the synthesized carbohydrazides (10–12, 15 mmol) were dissolved in ethanol (40 mL) and refluxed for 8–10 hours to allow in situ formation of the corresponding hydrazone intermediates. After the completion of the condensation, as monitored by TLC, the solvent was removed under reduced pressure using a rotary evaporator, and the resulting crude concentrate was dissolved in DMSO (15 mL). To the resulting solution, potassium carbonate (K2CO3, 30 mmol) and iodine (I2, 12 mmol) were added sequentially. The reaction mixture was stirred at 100 °C for 3–4 hours to promote oxidative cyclization. After cooling to room temperature, the mixture was diluted with a sodium thiosulfate solution (Na2S2O3, 30 mL) to quench excess iodine, then extracted with ethyl acetate (3 × 15 mL). The combined organic layers were washed with brine (15 mL), dried over anhydrous sodium sulfate and magnesium sulfate, and concentrated using a rotary evaporator. The crude product was purified by silica gel column chromatography (ethyl acetate/petroleum ether, 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 7) to afford the corresponding 1,3,4-oxadiazole derivatives (14–16) in pure form.
7) to afford the corresponding 1,3,4-oxadiazole derivatives (14–16) in pure form.
General procedure for synthesis of oxadiazole-based benzohydrazides (18–23)
To a round-bottom flask containing ethanol (30 mL), equimolar amounts (10 mmol) of the previously synthesized oxadiazole-based esters (14–16) were added along with either hydrazine hydrate (9, 10 mmol) or phenylhydrazine (17, 10 mmol). The mixture was refluxed for 5–6 hours under constant stirring, and the reaction progress was monitored by TLC. After the reaction was completed, the resulting solid product was allowed to cool, filtered, washed with cold ethanol, and dried. Recrystallization from ethanol afforded the corresponding pure hydrazide derivatives (18–23).Using this procedure, a total of six hydrazide compounds (18–23) were synthesized; two from each oxadiazole ester (one using hydrazine hydrate and the other using phenylhydrazine).
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 8.98 (s, 1H, NH), 8.24 (d, J = 8.5 Hz, 2H, ArH), 8.07 (d, J = 8.5 Hz, 2H, ArH), 5.01 (br-s, 2H, NH2), 3.94–3.88 (m, 1H, CH–CH3), 1.44 (d, J = 6.8 Hz, 3H, CH3). 13C NMR (100 MHz, CDCl3) δ 167.5, 164.5, 154.5, 152.9, 133.2 (q, J = 29.4 Hz), 131.7, 129.4, 128.7, 127.9 (q, J = 247 Hz, CF3), 124.9, 123.0, 120.5, 97.9, 43.1, 23.4. LCMS: m/z = 383.2 [M + H]+.
O), 8.98 (s, 1H, NH), 8.24 (d, J = 8.5 Hz, 2H, ArH), 8.07 (d, J = 8.5 Hz, 2H, ArH), 5.01 (br-s, 2H, NH2), 3.94–3.88 (m, 1H, CH–CH3), 1.44 (d, J = 6.8 Hz, 3H, CH3). 13C NMR (100 MHz, CDCl3) δ 167.5, 164.5, 154.5, 152.9, 133.2 (q, J = 29.4 Hz), 131.7, 129.4, 128.7, 127.9 (q, J = 247 Hz, CF3), 124.9, 123.0, 120.5, 97.9, 43.1, 23.4. LCMS: m/z = 383.2 [M + H]+.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 9.60 (s, 1H, NH), 9.44 (s, 1H, NH), 8.24 (d, J = 8.4 Hz, 2H, ArH), 8.07 (d, J = 8.4 Hz, 1H, NH), 7.24 (s, 2H, ArH), 7.17–7.08 (m, 2H, ArH), 3.94–3.88 (m, 1H, CH), 1.44 (d, J = 6.7 Hz, 3H, CH3). 13C NMR (100 MHz, CDCl3) δ 167.8, 164.7, 154.2, 152.5, 147.2, 133.2 (q, J = 28.9 Hz, C–CF3), 131.5, 128.4 (q, J = 248 Hz, CF3), 125.3 (2C), 122.8 (2C), 120.3 (2C), 119.2, 115.2, 97.8, 43.0, 23.1. LCMS: m/z = 459.1 [M + H]+.
O), 9.60 (s, 1H, NH), 9.44 (s, 1H, NH), 8.24 (d, J = 8.4 Hz, 2H, ArH), 8.07 (d, J = 8.4 Hz, 1H, NH), 7.24 (s, 2H, ArH), 7.17–7.08 (m, 2H, ArH), 3.94–3.88 (m, 1H, CH), 1.44 (d, J = 6.7 Hz, 3H, CH3). 13C NMR (100 MHz, CDCl3) δ 167.8, 164.7, 154.2, 152.5, 147.2, 133.2 (q, J = 28.9 Hz, C–CF3), 131.5, 128.4 (q, J = 248 Hz, CF3), 125.3 (2C), 122.8 (2C), 120.3 (2C), 119.2, 115.2, 97.8, 43.0, 23.1. LCMS: m/z = 459.1 [M + H]+.General procedure for synthesis of oxadiazole-based benzoic acids (24–26)
The previously synthesized oxadiazole-based ester derivatives (14–16, 10 mmol) were hydrolyzed under basic conditions to obtain the corresponding carboxylic acids. Each ester was dissolved in ethanol (25 mL), followed by the addition of 1 N potassium hydroxide solution (15 mL). The reaction mixture was refluxed for approximately 3 hours, and the progress was monitored by TLC. After the reaction completion, the mixture was allowed to cool at room temperature and neutralized by the dropwise addition of dilute hydrochloric acid. Stirring the neutralized mixture for 10 minutes led to the formation of solid precipitates, which were collected by filtration, washed with distilled water (2 × 10 mL), and dried to afford the purified oxadiazole-based benzoic acids (24–26).General procedure for synthesis of compounds (30–36)
Under a nitrogen atmosphere, the oxadiazole-based carboxylic acid derivatives (24–26, 6 mmol) were dissolved in dry dichloromethane (DCM, 10 mL). While stirring the solution at 0 °C, 1-hydroxybenzotriazole (HOBt, 5.0 mmol) was added, followed sequentially by the corresponding secondary amines (27–29, 5.0 mmol), 1-(3-dimethylaminopropyl)-3-ethylcarbodiimide hydro-chloride (EDC·HCl, 5.0 mmol), and 4-dimethylaminopyridine (DMAP, 5.0 mmol). The reaction mixture was stirred at room temperature for 24 hours, and TLC confirmed completion. After completion of the reaction, ethyl acetate was added and rinsed with aqueous 1 N HCl solution (20 mL). Multiple solvent extractions were performed using 30 mL of ethyl acetate each. At the end, the organic layers (ethyl acetate) were thoroughly mixed with the brine and sodium bicarbonate solution (20 mL each). The combined organic layers were dried over anhydrous sodium sulfate and concentrated under reduced pressure using a rotary evaporator. Purification of crude products 30–36 was performed by column chromatography using n-hexane/ethyl acetate (70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30–60
30–60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40, v/v) as the eluent.
40, v/v) as the eluent.
In vitro enzyme inhibition assay
Alpha amylase (α-amylase from porcine pancreas, Macklin), alpha glucosidase (from Saccharomyces cerevisiae, Macklin), recombinant human recombinant DPP-4 enzyme (Enzo Life Sciences), and human recombinant protein tyrosine phosphatase-1B (PTP1B, Enzo Life Sciences), which are commercially available, were used to evaluate the synthesized compounds' inhibitory potential. The substrates used were soluble starch for α-amylase, p-nitrophenyl-β-D-glucopyranoside (p-NPG) for α-glucosidase, and p-nitrophenyl phosphate (p-NPP, Macklin) for PTP1B. Substrate (Gly-Pro-NA) for DPP-4 was purchased from Sigma-Aldrich. Tris–HCl (pH 7.6) ethylenediaminetetraacetic acid (EDTA), NaCl, and dithiothreitol (DTT) were purchased from Macklin. Standard drugs sitagliptin, acarbose and ursolic acid were also purchased from Macklin. The procedure was performed in a 96-well plate using Multiplate Reader, and the reported method was used to accomplish these assays.19In vivo antidiabetic assays
OGTT
After fasting overnight, the normal rats were divided into several groups, each containing eight rats, and an oral glucose tolerance test was conducted. Group 2 received glibenclamide, whereas group 1 received 2% (w/v) Tween-80. The tested samples were administered in separate doses to the other groups. All of the rats were given an oral dosage of glucose (3 g kg−1) after ½ hours of delivery. Blood glucose levels were assessed via an SD glucometer, Active Blood Glucose Meter (ACCU-CHECK, Korea) at 0, 30, 60, 90, and 120 minutes after glucose injection.Diabetes induction
After acclimatization, rats that had fasted for the whole night were administered nicotinamide (110 mg per kg body weight, i.p.) dissolved in normal saline. After 15 minutes, an intraperitoneal (i.p.) injection of citrate buffer (0.1 M) solution containing streptozotocin (50 mg kg−1) was given. Rats were given a 10% glucose solution for three days in a row to keep them from suffering from streptozotocin-induced hypoglycemia shock and to guarantee their survival. After 72 hours, an SD glucometer from the tail vein and touch-sensitive glucometer strips were used to assess blood glucose levels. Rats were identified as diabetics and selected for additional research if their fasting blood glucose levels were greater than 250 mg dL−1.Antidiabetic activity
Each group, comprising eight subjects, was randomly assigned to the animals in the experiment. These groups included a normal control group, a diabetic control group, and a diabetic group that received varying dosages of compounds 23 and 18 at 10 and 20 mg per kg b.w. Furthermore, glibenclamide was given to a positive control group at a dosage of 500 µg per kg, p.o. The only vehicle used as controls was administered to the streptozotocin (STZ)-induced diabetic rat group and the normal control group. For a total of four weeks (28 days), compounds 18, 19, 21, 23, and 31 were administered orally every day. Following the dosage, their levels of blood sugar and weight were recorded on the first day and then weekly until the experiment's end.Statistical evaluation
The information is available as ±standard error of the mean. GraphPad Prism software version 8.0 was utilized to use statistical techniques, such as Dunnett's test and analysis of variance (ANOVA), to look at significant differences between the groups.Biochemical estimation
Serum biochemistry
The serum biochemical parameters were quantified by enzymatic colorimetric methods using a commercially available diagnostic kit (Human Gesellschaft für Biochemica und Diagnostica MBH, Germany). The following parameters were measured: creatinine, phosphorus, LDL cholesterol, HDL cholesterol, total cholesterol, triglycerides, alanine aminotransferase (ALT), aspartate aminotransferase (AST), alkaline phosphatase (ALP), albumin (ALB), total bilirubin (T. BIL), glucose (Glu), and total protein (T. PROT).Analysis was performed via automatic biochemical analyzer (Cobas Mira, Thermo Scientific Konelab 20/20XT), following the manufacturer's standard protocols.
In vivo acute toxicity of compound 23
The in vivo acute toxicity study was performed using a previously described method.24 The median lethal dose (LD50) of compound 23 after oral administration was estimated using the up-and-down method as outlined in the Organization for Economic Cooperation and Development (OECD) guidelines.25 The number of rats used at each dose level was progressively increased: three animals at the first level, five at the second, seven at the third, nine at the fourth, and ten at the fifth level. Compound 23 was given orally at doses of 500, 1000, 1500, 2000, 2500, 3000, and 3500 mg per kg b.w. The animals were carefully and continuously observed for signs of toxicity immediately after dosing and throughout the experimental period.Molecular docking
The molecular docking studies were performed using the Molecular Operating Environment (MOE, version 2016.0802). All synthesized compounds were docked into the active sites of four antidiabetic targets, namely α-amylase, α-glucosidase, dipeptidyl peptidase-4 (DPP-4), and protein tyrosine phosphatase 1B (PTP-1B). The three-dimensional crystal structures of α-amylase, DPP-4, and PTP-1B were obtained from the Protein Data Bank (PDB) using the accession codes 4W93, 1X70, and 1NNY, respectively. The structural model of α-glucosidase was adopted from a previously established and reported homology model.19All docking procedures were conducted following a previously validated and published protocol that had been optimized. Therefore, the same docking parameters, scoring functions, and refinement methods were utilized without modification. Briefly, the synthesized compounds were first protonated and energy-minimized using the default parameters in MOE. Ligand placement within the binding pockets was performed using the Triangle Matcher algorithm, and initial scoring was carried out using the London dG scoring function. Subsequently, the top-ranked poses were subjected to further refinement using the GBVI/WSA dG scoring function under a rigid receptor approach to obtain the most favorable binding conformations.
The docked ligand–protein complexes were evaluated based on their docking scores, binding orientations, and interactions with key active site residues. The profiles of all the best-docked conformations with the ligand were visually investigated and analysed through the ligand interaction module of MOE. The Discovery Studio Visualizer (version 2020) was utilized to create two-dimensional interaction diagrams illustrating hydrogen bonding, hydrophobic interactions, and other contacts between the ligands and their respective enzyme active sites.
Results and discussion
Design rational
The present study was designed on a rational molecular hybridization approach, integrating two privileged heterocyclic scaffolds, pyrimidine and 1,3,4-oxadiazole, to generate multi-target directed ligands (MTDLs) capable of regulating key enzymatic pathways involved in Type 2 Diabetes Mellitus (T2DM). The design concept was inspired by structural features of reported alpha-glucosidase,26,27 alpha-amylase,26,28 DPP-4,29,30 and PTP-1B inhibitors, as illustrated in Fig. 3. 1,3,4-Oxadiazole ring present in our design strategy originates from cyclization of the DHPM-5-ethyl ester through the corresponding hydrazide intermediate and serves as a bioisosteric replacement of the ester/hydrazide functionality. This transformation is well recognized in medicinal chemistry for: (i) improving metabolic stability by reducing hydrolytic liability, (ii) enhancing lipophilicity and membrane permeability, and (iii) preserving key hydrogen-bonding interactions due to the heteroatom-rich oxadiazole ring. In our rationale (Fig. 3), the 1,3,4-oxadiazole motif (highlighted in blue) represents the optimized bioisosteric unit, while the DHPM core (highlighted in green) was deliberately retained as a conserved pharmacophore. This decision was based on literature evidence indicating that DHPM derivatives possess significant biological activity, including relevance to metabolic disorders. Therefore, maintaining the DHPM scaffold ensures preservation of essential target-binding interactions, while the oxadiazole moiety modulates physicochemical and pharmacokinetic properties. | ||
| Fig. 3 Design rationale for the synthesis of targeted compounds.26–30 | ||
Introduction of electron-withdrawing groups (e.g., CF3) and aromatic substituents was meant to enhance hydrophobic interactions within the active sites of α-glucosidase and α-amylase, whereas amide/heteroatom functionalities were included to facilitate key hydrogen bonding with DPP-4 and PTP-1B active sites. This design was postulated to generate compounds capable of exerting immediate postprandial glucose control (via α-glucosidase/α-amylase inhibition), intermediate incretin stabilization (via DPP-4 inhibition), and long-term insulin sensitization (via modulation of insulin signaling pathways). Thus, the pyrimidine–oxadiazole hybrid scaffold was visualized as a structurally consistent, pharmacologically integrated platform for multi-pathway intervention in T2DM, which may provide improved therapeutic activity without necessarily involving combination therapy.
Chemistry
Synthetic access to the target compounds was planned strategically with the aim of building a hybrid framework that includes a 1,3,4-oxadiazole ring connected to a tetrahydropyrimidine core. The key tetrahydropyrimidine intermediates 6–8 was synthesized via a one-pot three-component reaction involving aldehydes 1–3, urea 5, and ethyl 4,4,4-trifluoro-3-oxobutanoate (4) in refluxing conditions in acetonitrile (ACN), which proceeded smoothly to afford the desired products in good yields. Subsequent treatment of these intermediates with hydrazine hydrate 9 under reflux conditions in absolute ethanol led to the formation of the corresponding carbohydrazides 10–12. The central 1,3,4-oxadiazole-containing scaffold was then obtained by condensing the carbohydrazides with methyl 4-formylbenzoate 13 in ethanol. The reaction proceeds via in situ hydrazone formation followed by oxidative cyclization using molecular iodine and potassium carbonate, furnishing the 1,3,4-oxadiazole derivatives 14–16. The synthesized compounds possess a chiral center at the C-4 position and were obtained as racemic mixtures, which are represented by wavy bonds at C-4 in the structures (Scheme 1).In Scheme 2, oxadiazole intermediates 14–16 were further reacted with hydrazine hydrate 9 and phenylhydrazine 17, employing the same hydrazinolysis conditions as described earlier, to yield the corresponding hydrazide derivatives 18–23. Each oxadiazole intermediate provided two final products via nucleophilic substitution, expanding the molecular diversity of the series.
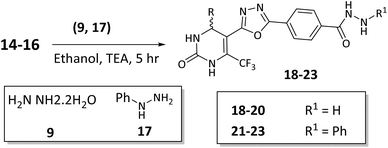 | ||
| Scheme 2 Synthesis of hydrazide derivatives 18–23 via hydrazinolysis of oxadiazole intermediates 14–16. | ||
In Scheme 3, the methyl ester functionality of compounds 14–16 was hydrolyzed under basic conditions using potassium hydroxide in ethanol under reflux, to obtain the corresponding carboxylic acids 24–26.
In Scheme 4, synthesized acid derivatives 24–26 were then finally subjected to amide bond formation using EDC·HCl, HOBt, and DMAP in dichloromethane, with three structurally distinct secondary amines (tert-butyl piperidin-3-ylcarbamate (27), pyrrolidine-2-carbonitrile (28), and 3-(trifluoromethyl)-3,5,6,7,8,8a-hexahydro-[1,2,4]triazolo[4,3-a]pyrazine (29)). This coupling strategy afforded a series of final amide derivatives 30–36, all bearing the key oxadiazole–tetrahydropyrimidine framework with appended amide moieties.
In vitro enzyme inhibition assay results
The synthesized compounds were studied for their enzyme inhibitory effects via in vitro assays on α-amylase, α-glucosidase, DPP-IV, and PTP1B enzyme assays, using standard protocols that have been followed in previous studies. The results in terms of IC50 values are presented in Table 1. The designed compounds were classified into two series (18–23 and 30–36) with modifications at both sides of the oxadiazole–tetrahydropyrimidine core. This study demonstrated the crucial role of substituents for increasing the inhibitory efficacy against α-amylase, α-glucosidase, DPP-4, and PTP1B.| S. no. | R | R1 | R2 | IC50 (µM) ± SEM | |||
|---|---|---|---|---|---|---|---|
| Alpha amylase | Alpha glucosidase | DPP-4 | PTP1B | ||||
| a SEM = ±standard error of mean of three experiments (i.e. n = 3), n.a. = no activity. * = compounds marked with an asterisk (*) did not achieve complete inhibition at the highest tested concentration (100 µM) and are therefore classified as partial inhibitors. IC50 values for these compounds correspond to 50% of their maximal inhibitory response. | |||||||
| 18 | CH3 | H | — | 5.27 ± 0.37 | 3.94 ± 0.16 | 0.42 ± 0.04 | 0.94 ± 0.06 |
| 19 | Ph | H | — | 1.04 ± 0.06 | 0.86 ± 0.06* | 7.05 ± 0.28 | 3.61 ± 0.22 |
| 20 | 4-OCH3·C6H4 | H | — | 2.49 ± 0.15 | 0.74 ± 0.06* | 30.09 ± 1.20* | 4.07 ± 0.20 |
| 21 | CH3 | Ph | — | 0.63 ± 0.05* | 2.73 ± 0.16 | 3.02 ± 0.15 | >50 |
| 22 | Ph | Ph | — | 1.38 ± 0.08 | >50 | 6.71 ± 0.47 | 19.05 ± 0.76 |
| 23 | 4-OCH3·C6H4 | Ph | — | 0.75 ± 0.05 | 2.21 ± 0.04 | 0.55 ± 0.03 | 1.01 ± 0.15 |
| 30 | CH3 | — | 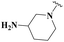 |
21.90 ± 1.75 | 27.58 ± 1.38 | 1.61 ± 0.11 | 15.03 ± 0.75 |
| 31 | CH3 | — | 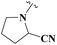 |
33.47 ± 1.34 | 1.81 ± 0.11 | 1.08 ± 0.11 | 17.84 ± 0.71 |
| 32 | CH3 | — | 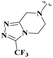 |
6.52 ± 0.39 | n.a. | 3.37 ± 0.24 | 15.94 ± 0.80* |
| 33 | Ph | — | 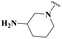 |
0.96 ± 0.06 | 35.07 ± 1.40 | 13.29 ± 0.93 | 13.72 ± 0.69 |
| 34 | Ph | — | 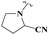 |
0.59 ± 0.03* | >50 | 13.61 ± 0.95* | 2.73 ± 0.12* |
| 35 | 4-OCH3·C6H4 | — | 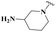 |
0.73 ± 0.04 | >50 | 7.04 ± 0.56 | 6.03 ± 0.30 |
| 36 | 4-OCH3·C6H4 | — | 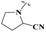 |
0.27 ± 0.06* | >50 | 17.28 ± 0.86 | 9.71 ± 0.18 |
| Sitagliptin | — | — | 0.009 ± 0.001 | — | |||
| Ursolic acid | — | — | — | 2.63 ± 0.16 | |||
| Acarbose | 11.36 ± 0.93 | 10.72 ± 0.99 | — | — | |||
α-Amylase inhibition assay results
In this assay, our synthesized 3,4-oxadiazole-based tetrahydropyrimidinone hybrids showed a strong inhibitory effect against the α-amylase enzyme. Having IC50 values in the micromolar range, all of the compounds exhibited good to moderate inhibition (Table 1). Acarbose was used as the standard compound in this test. All compounds of both series displayed superior inhibitory activity compared to the standard drug acarbose (IC50 = 11.36 ± 0.93 µM), except for compounds 30 and 31. In series 1, compound 21 (IC50 = 0.63 ± 0.03 µM), with a methyl substitution on tetrahydro-pyrimidinone, showed the best potency. N′-Phenyl hydrazide moiety generally increased activity, as in compound 21 compared to compound 18 and compound 23 compared to compound 20. But it reduced the inhibitory activity in compound 22 as compared to compound 19, which may indicate that the nature of substitution on the pyrimidine ring impacts the influence of N′-phenyl substitution. Further, electron-donating groups like methoxy or methyl on the aryl ring of the THP core further enhanced potency. While in series 2, the effect of substitution on the THP ring played a crucial role; compound 36 (IC50 = 0.27 ± 0.02 µM) with a methoxy-phenyl moiety and compound 34 (IC50 = 0.59 ± 0.02 µM) with a pyrrolidine-carbonitrile fragment exhibited the most potent activity overall. Compounds containing aminopiperidine and methyl-substituted THP moieties (compounds 30 and 31) presented poor activity.α-Glucosidase inhibition assay results
The outcomes of the α-glucosidase inhibition assay demonstrated that in series 1, compounds 19 (IC50 = 0.86 ± 0.06 µM) and 20 (IC50 = 0.74 ± 0.06 µM) showed strong inhibition, overtaking standard drug acarbose (IC50 = 10.72 ± 0.99 µM), signifying the importance of unsubstituted hydrazide and an electron-donating group (methoxy). Compound 22, on the other hand, has shown an IC50 above 50 µM, indicating that the N′-phenyl hydrazide moiety might hinder activity. In series 2, only compound 31 (IC50 = 1.81 ± 0.09 µM) showed good inhibition, while others, like compounds 30, 32–34, 35, and 36, showed weak or no inhibition (Table 1), demonstrating that aminopiperidine and bulky aryl substituents reduce potency. This implies that the hydrazide moiety is critical for α-glucosidase inhibition activity, as unsubstituted hydrazide and N′-phenyl hydrazide moiety on the east side of the molecules are well-tolerated (18–21 and 23), but replacement with amide-containing pyrrolidine/piperidine substituents (30–36) resulted in reduced or complete loss of activity.DPP-4 inhibition assay results
The inhibitory activity of the synthesized compounds against the DPP-4 enzyme is listed in Table 1. Compared to the positive control sitagliptin (IC50 = 0.009 ± 0.001 µM), several compounds presented good to moderate inhibition. Compound 18 (IC50 = 0.42 ± 0.03 µM) demonstrated the most potent activity in series 1, indicating the absence of N′-phenyl and the advantageous role of the methyl group on the THP ring. Compound 23 (IC50 = 0.55 ± 0.02 µM) also displayed good inhibition, revealing that a methoxyphenyl group at position 4 of the THP when combined with N′-phenyl hydrazide moiety contributes positively. Compound 20, which had the same substitution but no N′-phenyl, was less active (IC50 = 30.09 ± 1.20 µM), indicating that the N′-phenyl hydrazide moiety improves DPP-4 inhibition when combined with certain aryl moieties in this case. Compounds 30 and 31 in series 2 demonstrated encouraging inhibition, particularly when aminopiperidine or pyrrolidine-carbonitrile were included at the terminal position (IC50 = 1.61 ± 0.11 µM and 1.08 ± 0.08 µM, respectively). Overall, in the second series (30–36), irrespective of the amide substituent, bulky groups like phenyl or methoxyphenyl (33–36) when replaced with methyl groups (30 and 31) led to reduced activity.PTP1B inhibition assay results
The inhibitory activity of synthesized compounds (18–23 and 30–36) against the PTP1B enzyme was evaluated, and ursolic acid was taken as a reference drug. In the first series (18–20), compound 18 (IC50 = 0.94 ± 0.04 µM) exhibited the most potent inhibition, followed by compound 23 (IC50 = 1.01 ± 0.17 µM), both surpassing the standard ursolic acid (IC50 = 2.63 ± 0.86 µM). The PTP1B inhibition values indicate that small substituents (methyl) on the THP ring may be favorable. Still, size alone does not explain any difference, as illustrated by the comparable values of 19 (phenyl substituent) and 20 (methoxy-phenyl substituent), as well as the compounds containing N′-phenyl hydrazide (21–23) exhibiting a different pattern. Compound 23 emerged as potent, followed by compounds 22 and loss of activity in 21 indicated that the effect of the substituent on THP ring is dependent on the moieties of the eastern side (N′-phenylhydrazide or hydrazide). While in series 2, only compound 34 (IC50 = 2.73 ± 0.11 µM) showed activity comparable to the standard, suggesting that the phenyl-substituted THP ring and pyrrolidine-carbonitrile moiety contribute positively. Moderate inhibition was demonstrated by other compounds in this series with IC50 values in the range of 6–17 µM.It is noteworthy that several compounds (Table 1) exhibited low IC50 values but did not achieve complete inhibition, indicating a partial inhibition profile despite high potency. This behavior may suggest a non-competitive or allosteric mode of action, thereby limiting their maximal inhibitory effect. The dose response curves (Fig. 4) demonstrated that compounds 21, 23, and 36 against α-amylase, 19 and 20 against α-glucosidase, 31 against DPP-4, and 34 against PTP-1B showed partial inhibition as they do not completely suppress the enzyme activity even at increased concentration.
Oral glucose tolerance
This assay evaluated how different doses (10 mg kg−1 and 20 mg kg−1) of test compounds affect blood glucose levels over time after glucose intake, compared to a control group and a standard antidiabetic drug. Following the oral administration (3 g kg−1) to the test animals, the group's serum glucose levels peaked at 30 minutes and subsequently decreased to 128.85 ± 4.71 mg dL−1 after two hours.Pre-administration of compounds 18, 19, 21, 23, and 31 (at 10 and 20 mg kg−1 doses) along with glibenclamide caused a notable reduction in the concentrations of serum glucose levels compared to the control group, as illustrated in Table 2. The outcomes of the oral glucose tolerance test (OGTT) showed that all selected compounds exhibited varying degrees of antihyperglycemic activity compared to the control, as shown in Fig. 5. However, among them, compound 23 revealed the most potent effect, closely approaching the reference drug. According to these findings, the most potent compounds show potential for more in vivo studies.
| Groups | Dose (mg kg−1) | Blood glucose level in mg dL−1 | ||||
|---|---|---|---|---|---|---|
| 0 | 30 | 60 | 90 | 120 | ||
| a The sample size is n = 8, and the data are stated as means ± SEM. Dunnett's post hoc multiple comparison test and a one-way ANOVA were used to evaluate the significance of the values (as shown by P). There were significant differences between the normal control group and the diabetes control group (###P < 0.001, ##P < 0.01). Additionally, using a one-way ANOVA followed by Dunnett's post hoc test, comparisons between the test samples and the diabetes control group, as well as the groups treated with glibenclamide, revealed significant differences, as indicated by *P < 0.05, **P < 0.01, and ***P < 0.001. | ||||||
| Control group (sugar-treated) | — | 106.22 | 175.24 ± 4.12### | 151.74 ± 3.96### | 135.12 ± 4.01## | 128.85 ± 4.71## |
| 18 | 10 | 101.87 ± 4.75 | 121.23 ± 3.95* | 123.45 ± 4.62* | 115.63 ± 4.12** | 113.33 ± 4.11* |
| 20 | 103.53 ± 4.23 | 116.53 ± 4.09** | 115.36 ± 4.71* | 111.21 ± 4.35* | 109.73 ± 3.99** | |
| 19 | 10 | 109.22 ± 4.26 | 134.33 ± 4.75* | 145.63 ± 4.14* | 134.52 ± 4.58** | 130.23 ± 4.31* |
| 20 | 104.16 ± 3.86 | 119.83 ± 4.81* | 123.32 ± 4.36** | 120.82 ± 4.82*** | 121.42 ± 4.13* | |
| 21 | 10 | 104.52 ± 4.69 | 125.36 ± 4.15** | 118.53 ± 4.63** | 114.96 ± 4.03** | 116.65 ± 4.25* |
| 20 | 103.46 ± 4.52 | 118.26 ± 4.35* | 123.23 ± 4.59* | 111.87 ± 4.43* | 110.35 ± 4.22* | |
| 23 | 10 | 101.62 ± 4.37 | 115.74 ± 4.13** | 131.33 ± 4.66* | 115.23 ± 3.97* | 110.18 ± 4.55* |
| 20 | 102.32 ± 4.07 | 120.24 ± 4.21* | 116.28 ± 4.33*** | 108.45 ± 4.51** | 109.43 ± 4.56** | |
| 31 | 10 | 104.56 ± 4.13 | 128.70 ± 4.25** | 117.73 ± 4.31* | 116.78 ± 4.76* | 115.89 ± 4.54* |
| 20 | 100.22 ± 4.77 | 118.23 ± 4.06* | 116.76 ± 3.89** | 109.61 ± 4.16** | 110.13 ± 4.33* | |
| Standard | — | 103.03 ± 4.26 | 106.98 ± 4.75** | 110.36 ± 4.53*** | 103.66 ± 4.21*** | 104.15 ± 4.33* |
 | ||
| Fig. 5 Time course graph of the oral glucose tolerance test (OGTT) curves of rats treated with the synthesized compounds (18, 19, 21, 23, and 31) at the doses of 10 mg kg−1 and 20 mg kg−1. | ||
Anti-diabetic activity assessment using a chronic model
Rats were used in an induced chronic paradigm to assess the hypoglycemic effects. We chose the most optimal multitarget inhibitors (compounds 23 and 18) for this purpose. Throughout the trial, the normal control group's evaluation revealed 97–103 mg dL−1. Blood glucose levels (BGL) in the diabetic control group ranged from 247 to 283 from day 1 to day 28. Rats in the conventional control group (glibenclamide) had blood glucose levels of 291 on day 1 and 102 on day 28. Likewise, the tested compound 23 at 20 mg kg−1 presented notably lower fasting blood glucose (FBG) levels from day 1 to day 28 in rats, with 293 ± 3.95, 185 ± 4.19, 158 ± 4.66, 125 ± 4.75, and 115 ± 4.46 mg dL−1. Similarly, the BGL dropped from 290 to 129 mg dL−1 in the compound 18 test group, which was less active than compound 23. As seen in Fig. 6, compound 23 was most effective in this assay. As illustrated in Fig. 6, a detailed statistical comparison was made between the treatment groups. While both compounds significantly reduced BGL compared to the diabetic control (***P < 0.001). Similarly, when compared the compound 18 and 23 the compound 23 demonstrated superior efficacy over compound 18. This superiority became statistically significant at day 21 and 28 (###P < 0.001, ##P < 0.01).Furthermore, when comparing the test compounds to the standard drug, compound 23 showed no significant difference (n.s.) from glibenclamide at day 21, while significant at the end of the study at 28 day (!!P < 0.01) suggesting that its hypoglycemic potency is comparable to the reference drug. In contrast, compound 18 remained significantly less effective than both glibenclamide (!!!P < 0.001) at the final time points. These results highlight compound 23 as the most potent lead candidate in this chronic model.
Changes in serum biochemistry in experimental rats
Administration of STZ induced a noticeable variation in the serum biochemical parameters when compared to the normal control group. Diabetic rats displayed significant elevation in the serum creatinine (0.8 ± 0.2 mg dL−1) and urea (43.8 ± 3.1 mg dL−1) relative to the normal control group animals (0.3 ± 0.1 and 27.2 ± 1.1 mg dL−1), respectively. Treatment with glibenclamide or the tested synthesized compounds significantly ameliorated these alterations. Compound 23 at a dose of 20 mg kg−1 caused a greater reduction in creatinine (0.46 ± 0.2 mg dL−1, p < 0.01) and urea (34.6 ± 2.5 mg dL−1, p < 0.01) compared to compound 18 (0.55 ± 0.3 and 37.1 ± 2.8 mg dL−1, p < 0.05), indicating better nephroprotective effects. Additionally, lipid profile analysis showed that diabetic rats had significantly lower HDL cholesterol (19.5 ± 3.2 mg dL−1) and higher LDL (155.2 ± 5.1 mg dL−1), total cholesterol (109.6 ± 4.1 mg dL−1), and triglycerides (101.8 ± 6.2 mg dL−1) compared to the normal control group. Both the standard drug glibenclamide and the compound 23 improved these parameters. Compound 23 notably increased HDL (39.6 ± 2.9 mg dL−1, p < 0.01) and decreased LDL (99.8 ± 3.6 mg dL−1, p < 0.01), cholesterol (76.1 ± 3.2 mg dL−1, p < 0.01), and triglycerides (82.3 ± 4.4 mg dL−1, p < 0.01). While compound 18 also improved lipid levels, the effects were less pronounced, with HDL at 33.4 ± 3.2 mg dL−1 and higher LDL, cholesterol, and triglycerides compared to compound 23. Regarding the hepatic biomarkers, the diabetic animals displayed elevated AST (95.1 ± 5.2 U L−1), ALT (33.8 ± 4.3 U L−1), and ALP (160.7 ± 5.6 U L−1) relative to the control group. Treatment with glibenclamide and the tested compounds reduced these enzyme levels. Compound 23 produced greater decreases in AST (75.2 ± 3.2 U L−1, p < 0.01), ALT (28.1 ± 3.0 U L−1, p < 0.01), and ALP (137.1 ± 4.9 U L−1, p < 0.01) compared to compound 18 (80.5 ± 3.9, 29.8 ± 3.3, and 143.3 ± 5.3 U L−1), respectively. Similarly, the serum bilirubin was significantly higher in the diabetic control group (1.4 ± 0.7 mg dL−1) compared with normal rats (0.6 ± 0.2 mg dL−1). Both the compound 23 (0.92 ± 0.5 mg dL−1, p < 0.01) and compound 18 (1.03 ± 0.6 mg dL−1, p < 0.05) reduced bilirubin levels, with compound 23 displaying a more pronounced effect. Likewise, total protein levels, which were reduced in diabetic rats (5.1 ± 1.5 g dL−1), were significantly improved by both compounds, with compound 23 (6.0 ± 0.5 g dL−1, p < 0.01) restoring values nearly to those of glibenclamide, while compound 18 displayed a moderate increase (5.7 ± 0.7 g dL−1, p < 0.05). Overall, the biochemical analysis demonstrated that both compounds 23 and 18 exerted beneficial effects against the diabetic induced alterations, with compound 23 consistently displaying superior efficacy over 18 in normalizing renal, hepatic, and lipid parameters, as shown in Table 3.| Biochemical parameters | Normal control | Diabetic control | Glibenclamide-0.5 mg kg−1 | Compound 18 (20 mg kg−1) | Compound 23 (20 mg kg−1) |
|---|---|---|---|---|---|
| a AST: (aspartate aminotransferase), ALT: (alanine aminotransferase), ALP: (alkaline phosphatase), T. BIL: (total bilirubin), T. PROT: (total proteins), HDL: (high-density lipoprotein), LDL: (low-density lipoprotein). Values are expressed as mean ± SEM (n = 6). Statistical significance: *p < 0.05, **p < 0.01 compared with diabetic control. Compound 23 exhibited greater potency than compound 18 in restoring biochemical alterations toward normal levels. | |||||
| Creatinine (mg dL−1) | 0.3 ± 0.1 | 0.8 ± 0.2 | 0.5 ± 0.2** | 0.55 ± 0.3* | 0.46 ± 0.2** |
| Urea (mg dL−1) | 27.2 ± 1.1 | 43.8 ± 3.1 | 33.5 ± 2.4** | 37.1 ± 2.8* | 34.6 ± 2.5** |
| HDL cholesterol (mg dL−1) | 56.1 ± 1.6 | 19.5 ± 3.2 | 45.2 ± 2.1** | 33.4 ± 3.2* | 39.6 ± 2.9** |
| LDL cholesterol (mg dL−1) | 87.5 ± 1.2 | 155.2 ± 5.1 | 91.4 ± 4.8** | 111.2 ± 4.5* | 99.8 ± 3.6** |
| Cholesterol (mg dL−1) | 60.5 ± 2.1 | 109.6 ± 4.1 | 70.2 ± 3.1** | 83.4 ± 3.5* | 76.1 ± 3.2** |
| Triglyceride (mg dL−1) | 65.1 ± 3.7 | 101.8 ± 6.2 | 77.5 ± 3.8* | 89.2 ± 5.0* | 82.3 ± 4.4** |
| AST (U L−1) | 45.6 ± 2.5 | 95.1 ± 5.2 | 70.8 ± 3.6** | 80.5 ± 3.9* | 75.2 ± 3.2** |
| ALT (U L−1) | 22.1 ± 1.7 | 33.8 ± 4.3 | 26.6 ± 2.9** | 29.8 ± 3.3* | 28.1 ± 3.0** |
| ALP (U L−1) | 90.5 ± 3.0 | 160.7 ± 5.6 | 132.2 ± 6.1** | 143.3 ± 5.3* | 137.1 ± 4.9** |
| T. bilirubin (mg dL−1) | 0.6 ± 0.2 | 1.4 ± 0.7 | 0.9 ± 0.4* | 1.03 ± 0.6* | 0.92 ± 0.5** |
| T. protein (g dL−1) | 6.7 ± 0.4 | 5.1 ± 1.5 | 6.1 ± 0.6** | 5.7 ± 0.7* | 6.0 ± 0.5** |
In vivo acute toxicity of compound 23
Healthy adult Wistar rats weighing 180–220 g were used to assess acute toxicity. We assessed acute toxicity using the methodology and recommendations as described in the earlier study. Using the up and down procedure (UDP), the doses are given to the animals one at a time, beginning with an estimated LD50 dose. The subsequent animal receives a higher dosage if the preceding one lives. On the other hand, if the prior animal dies, a lower dosage is administered. Depending on the results of the preceding animal, the dosage is changed either up or down for each subsequent animal. Estimating the oral toxicity of compound 23 given by oral gavage to animals at varied doses (100–3500 mg per kg body weight) and with varying numbers of rats for each concentration was the aim of this study. In the first phase, three rats that received a dose of chemical 23 at 3500 mg per kg b.w. died. Four of the five rats perished as a result of the second phase dosage of 3000 mg per kg b.w. In the third phase, 2500 mg kg−1 was given to seven rats. Throughout the 96-hour observation period, three rats perished. The dosage was reduced to 1500 mg kg−1 in the subsequent phase after two of the nine rats died at the 2000 mg kg−1 dosage in the fourth phase. All of the rats survived. Rats were also administered dosages of Compound 23 at 1000 and 500 mg kg−1 in order to measure the No Observed Adverse Effect Level (NOAEL). None of those animals displayed any adverse symptoms, and all survived the 96-hour observations Table 4.| Doses (mg kg−1) | No. of animals tested | Death no. | Survival time | % mortality |
|---|---|---|---|---|
| 3500 | 3 | 3 | 20 min; 32 min; 45 min | 100% |
| 3000 | 5 | 4 | 26 min; 55 min; 2 h | 80% |
| 2500 | 7 | 3 | 2 h; 3.5 h; 7 h | 42.80% |
| 2000 | 9 | 2 | 16 h, 28 h | 22.22% |
| 1500 | 10 | 0 | >96 h | 0 |
| 1000 | 10 | 0 | >96 h | 0 |
| 500 | 10 | 0 | >96 h | 0 |
Based on the data, a nonlinear regression fitting method calculated compound 23's oral LD100 to be 3500 mg per kg b.w. (GraphPad Prism 8) and its estimated LD50 to be 2515 mg per kg b.w. (95% CI: 2273–2745 mg kg−1; R2 = 0.9776) (Fig. 7).
Molecular docking studies results
To gain insights into the molecular basis of the observed in vitro potency, molecular docking studies were carried out on the most active compounds, 18 and 23, which exhibited the strongest inhibitory activity across the enzymatic assays. Docking simulations were performed using the Molecular Operating Environment (MOE 08.02) software to evaluate its binding interactions with four key antidiabetic targets: α-amylase, α-glucosidase, DPP-4, and PTP1B.The high-resolution 3D crystal structures of α-amylase, DPP-4, and PTP1B were retrieved from the Protein Data Bank (PDB) using their respective IDs (4W93,31 1X70,32 and 1NNY33), while a validated homology model was utilized for α-glucosidase, which we have previously developed and reported.19 This computational study aimed to predict the binding orientation, key interactions within the active sites, and the possible molecular rationale behind the compound's multitarget inhibitory potential. Binding energy values (kcal mol−1) and interacting residues along with their types and bond distances (Å) for synthesized compounds 18 and 23 against α-amylase, α-glucosidase, DPP4, and PTP-1B are presented in Table S-5 in SI.
Docking studies on α-amylase
Compound 18 had a good binding orientation in the active site of α-amylase, and it has polar and non-covalent interactions with the important catalytic and structural residues. Three hydrogen bonds were seen: two were between the terminal hydrazide moiety, which binds with the side chains of Glu240 and Lys200, but another hydrogen bond was observed between the pyrimidine nitrogen and the side chain of Asp300, a residue that is related to substrate binding.The 2D interaction diagram generated by Discovery Studio Visualizer (Fig. 8a) indicated contacts involving the fluorine atoms of the CF3 group, which are labeled as halogen (fluorine) interactions. However, from a chemical perspective, fluorine in a trifluoromethyl group is a weak halogen bond donor due to its low polarizability. Therefore, these contacts are more appropriately interpreted as weak fluorine-mediated interactions, such as C–F⋯H interactions or van der Waals contacts, rather than classical halogen bonding.
The CF3 group on the pyrimidine ring participates in weak fluorine-mediated contacts (e.g., C–F⋯H or van der Waals interactions) with the backbone of residue Tyr62 located within the hydrophobic cleft. Moreover, the oxadiazole ring of compound 18 interacts with the side chain of Glu233, a key catalytic residue of α-amylase, through a π–anion interaction, contributing to the stabilization of the ligand within the active site.
Compound 23 demonstrated a clear profile of interaction in the active site of α-amylase, which shows its possible important inhibitory activity. Six hydrogen bonds were realized. The hydrazide terminal element provided two hydrogen bonds with the NH protons of Tyr151 and Ile235, and the O group of the moiety also provided a hydrogen bond with the backbone of Ile235, indicating that there were strong anchors at the peripheral part of the binding cleft. The CF3 group of the compound is involved in a weak fluorine-mediated interaction (C–F⋯H) with the –NH group of Asp197, rather than forming a classical hydrogen bond (Fig. 8b). In addition, the NH proton of the pyrimidine ring was hydrogen bonded to the backbone of Tyr62 (2.51 Å), and the oxygen atom of its carbonyl group was hydrogen bonded to the backbone's NH of Gln63 (2.41 Å), which are also located in the known catalytic environment of α-amylase (Fig. 8b).
Also, the CF3 group exhibited two interactions with halogen-fluorine, one of them with backbone of Tyr62 (3.47 Å), and another one with the side chain of Asp197 (3.28 Å), which further stabilized in the active pocket. The aromatic methoxyphenyl ring of the compound also developed a π–π T-shaped interaction with the aromatic moiety of Trp59, which is a stacking interaction-forming residue that is important in substrate recognition.
Docking studies on α-glucosidase
Compound 18 showed a positive binding capacity in the active site of α-glucosidase, where it was found to interact with several catalytic and structural residues in the site. Five hydrogen bonds were found in total, with one of them being a hydrogen bond with the nitrogen atom of the tetrahydropyrimidine ring with the side chains of Asp349 (2.54 Å) and Glu276, which are two of the members of the catalytic triad in the enzyme. The oxadiazole nitrogen group in the middle was used to bind the protein through a hydrogen bond with the side chain's amino group of His279, whereas the terminal oxadiazole was engaged in a hydrophobic interaction with the backbone of Phe310. Moreover, the two hydrogen bonding interactions involved the backbones of Arg312 (2.03 Å, 2.86 Å), one with the carbonyl group of the hydrazide moiety and the other with a fluorine atom in the CF3 group. To further stabilize the binding, the oxadiazole core and the related aromatic ring were involved in the π–π stacking and π–alkyl interaction with both the side chain of Arg312 and aromatic moiety of Phe300, which are located in the active site of the enzyme (Fig. 9a). Compound 23 showed a wide spectrum of stabilizing interactions in the active site of α-glucosidase, and this compound can be a good inhibitor of the enzyme. The carbonyl oxygen of the pyrimidine ring was hydrogen-bonded to the side chain's amino group of Arg439, and the nitrogen atom in the central oxadiazole moiety joined with the hydrogen bond to Arg312's side chain. Further, the hydrogen of the terminal moiety phenyl hydrazide formed a hydrogen bond with the backbone of Pro309, which signified anchoring in the substrate-binding pocket. Indole moiety of Trp242 also had a π–sigma interaction with the same phenyl hydrazide ring as well as formed a π–π T-shaped interaction with the aromatic moieties of Trp242 and His239, residues that are known to be involved in the aromatic pocket architecture of the enzyme (Fig. 9b).In addition, the π–π T-shaped interaction of the central oxadiazole ring with the phenyl ring of Phe300 was also observed, which additionally stabilized the ligand in the catalytic cleft. The methoxyphenyl ring, which was bound to the pyrimidine moiety, had a π–π stacked interaction with the phenyl ring of Phe157 (5.12 Å), enriching the hydrophobic interaction in the binding site. Notably, the interactions between the CF3 group and the side chains of Asp408 (2.71 Å, 2.89 Å) and Phe157 (2.81 Å, 3.30 Å), which were described as halogen-fluorine bondings, but cannot be classified as true halogen bonds. Classical halogen bonding requires a halogen atom with a positively polarized σ-hole interacting with an electron-rich nucleophile (Cavallo, et al.34), which is not present for the fluorines in CF3. Although MOE identifies these contacts as halogen interactions based on geometric criteria, chemically they correspond to a dipole interactions rather than classical halogen bonding, which strengthened the binding force through the non-covalent halogen bonding.
Docking studies on DPP4
Compound 18 showed a high binding affinity to the active site of DPP-4 and interacted with multiple important ones of the residues in subsites, including S1, S2, and S2-extensive. The hydrazide moiety of the compound was hydrogen-bonding with the backbone of Val207 and side chain of Glu206, which are residues of the S2 and S2-extensive regions. Moreover, the pyrimidine core was also engaged in a hydrogen bond with the catalytic site Ser630 by both its carbonyl and nitrogen group indicating stabilization in position S1. Interestingly, CF3 group played an important role in binding, they made hydrogen bonds with the side chain's amino groups of Asn710 (2.91 Å) and Arg125 (6.62 Å), and formed halogen bonds with the imidazole's nitrogen atom of His740 (2.93 Å) and Glu205's side chain (2.91 Å), a feature that has frequently been linked to increased binding specificity. Moreover, the aromatic ring on the central oxadiazole moiety also interacted with the phenyl ring of the Phe357, which is a major residue of the S2-extensive subsite, by π–π stacking interactions (Fig. 10a). Compound 23 has shown a good binding orientation in the active site of DPP-4 enzyme, which is involved in multiple and significant non-covalent interactions leading to its possible inhibitory effect. The carbonyl oxygen atom of the hydrazide moiety was connected by two strong hydrogen bonds to peptide S2-extensive subsite residues, i.e. terminal amino group of Arg358 and the hydroxyl group of Ser209, which points to anchoring in an area of significance in potent DPP-4 inhibition. Moreover, the central oxadiazole ring, as well as the aromatic phenyl substituent that was attached to it, both formed pi–pi stacked interactions with the phenyl ring of Phe357, a well-established key residue in the S2-extensive subsite. The methoxyphenyl ring of the pyrimidine backbone was shown to have additional aromatic interactions with the phenyl moiety of Tyr662, a π–π stacked interaction, and Tyr666 (phenyl moiety) a π–π T-shaped interaction, both of which are located in the S1 subsite. These are the same interactions as observed with the reference drug sitagliptin, indicating that compound 23 is an effective occupant of the S1 and S2-extensive sub-pockets of the DPP-4 active site, which supports the potential of this compound 23 being a promising DPP-4 inhibitor (Fig. 10b).Docking studies on PTP1B
The PTP-1B active site is made up of two primary components: the catalytic-binding pocket (A site) and the secondary aryl phosphorylation-binding pockets (B, C, and D sites). The A site, or catalytic-binding pocket, is the most polar location. PTP1B's enzymatic activity is determined by the amino acids Asp181 and Gln262, whereas hydrophobic residues (Tyr46, Val49, Phe182, Ala217, Cys215, Arg221, and Ile219) play an important role in the efficacy of PTP1B inhibitors. The residues Arg24, Arg254, Tyr20, Ala27, Phe52, and Met258–Gly259 make up the majority of the noncatalytic binding region known as the B site. PTP1B's potency and selectivity are owing to a charged area produced by the C site, Arg47, Tyr46, and Asp48. The polar and charged residues that border the D site (Lys120, Asp181, Tyr46, Glu115, and Ser216).19Compound 18 showed a high binding affinity to the PTP1B active site and established a interactions with both the PTP1B active site and non-catalytic regions of the enzyme. Six typical hydrogen bonds were seen. The terminal hydrazide functional group formed important conventional hydrogen bonding patterns with the side chains of Asp29 (2.69 Å) and Phe30 (2.22 Å, 3.05 Å), with the nitrogen atom of the oxadiazole ring forming hydrogen bonds with the side chain's amino group of Gly259 (2.49 Å) and Arg254 (3.00 Å), residues that were near the B-site. The oxadiazole ring also interacted through π–sigma with the backbone of Met258 (2.56 Å), and another π–sulfur interaction was also found between the sulfur atom of Met258 and the neighboring aromatic ring (4.17 Å). The two oxygen atoms of the carbonyl group in the molecule also added additional stability due to the hydrogen bonding with the side chain's NH2 of Lys36 and Gln262, which was also a critical residue that played an important role in catalytic activity. Also, the alkyl interaction with Met258 and halogen bonding between the CF3 group (on the pyrimidine ring) and the side chain of Asp48 were also identified (Fig. 11a). Compound 23 had a high binding affinity in the PTP1B catalytic domain, in which several interactions with major active site residues were stabilizing. It is noteworthy that the terminal phenyl hydrazide moiety formed a hydrogen bond with Asp48's side chain hydroxyl group (1.48 Å), which is one of the key residues with which the enzyme catalyzes the reaction. Also, the NH proton of the pyrimidine ring was grouped with the side chain of Gly183, which is a signature WPD loop region necessary to make the activity of PTP1B catalytic. The CF3 substituent also contributed to specificity by forming a polar interaction with the side chains of Gln266 and Gly183 additional stabilizing interactions (e.g., C–F⋯H or dipolar contacts) which indicated a well-linked binding with the active cleft of the enzyme (Fig. 11b).
Furthermore, the methoxy group on the methoxyphenyl ring formed two additional hydrogen bonds with the side chain's amino group of Ala217 and Ser216, residues located near the peripheral binding pocket. The aromatic methoxyphenyl ring also interacted with the –SH of Cys215 and the nitrogen of Arg221, respectively, by π–cation and π–sulfur interactions, both of which belong to the catalytic and phosphate binding loops.
Conclusions
In summary, a series of oxadiazole–tetrahydropyrimidinone hybrids was designed rationally. These hybrid compounds were assessed for their effectiveness as multitarget agents in treating diabetes. The compounds demonstrated promising inhibitory activity against α-amylase, α-glucosidase, DPP-4, and PTP1B, with compound 23 exhibiting a balanced multitarget profile. In vivo studies further confirmed its significant antihyperglycemic efficacy and favorable safety profile. Molecular docking supported the experimental findings by revealing stable binding interactions within the active sites of all four enzymes. Overall, these hybrids represent promising lead scaffolds for the development of multitarget-directed therapies for type 2 diabetes mellitus.Author contributions
Umer Rashid and Sohail Hassan conceived, designed, and supervised the study and were involved in all phases of the work, including synthesis, pharmacological evaluation, and manuscript writing and editing. Shereen carried out the synthesis of the compounds, with assistance from Ayesha Tahir. The in vitro biological evaluations were performed by Shereen and Ayesha Tahir. Molecular docking studies were conducted by Awais Ahmed and Nouman Aslam, while the docking results were analyzed and interpreted by Umer Rashid. The in vivo studies were performed by Muhammad Saeed Jan. The manuscript was initially drafted by Shereen and Ayesha Tahir and subsequently reviewed and edited by Umer Rashid and Sohail Hassan. All authors have read and approved the final version of the manuscript and agree to its submission for publication.Conflicts of interest
The authors declare no conflict of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: dose–response curve of sitagliptin (standard drug), molecular docking analysis, and spectroscopic characterization data (NMR and LC-MS chromatograms) for the synthesized compounds. See DOI: https://doi.org/10.1039/d6ra01810f.Acknowledgements
Sohail Hassan expresses his sincere gratitude to the Department of Pharmaceutical Chemistry, Faculty of Pharmacy and Pharmaceutical Sciences, University of Karachi, Karachi, Pakistan, for providing the necessary laboratory infrastructure and technical support for this work. Umer Rashid would like to thank the Higher Education Commission, Pakistan, for supporting the research under the National Research Program for Universities (NRPU) (20-14513/NRPU/R&D/HEC/2021 2021).References
- A. D. Deshpande, M. Harris-Hayes and M. Schootman, Phys. Ther., 2008, 88, 1254–1264 CrossRef PubMed.
- C. Tudoran, Biomedicines, 2025, 13(8), 1980 CrossRef PubMed.
- American Diabetes Association, Diabetes Care, 2021, 44, S15–S33 CrossRef PubMed.
- V. G. Klochkov, E. N. Bezsonova, M. Dubar, D. D. Melekhina, V. V. Temnov, E. V. Zaryanova, N. A. Lozinskaya, D. A. Babkov and A. A. Spasov, Bioorg. Med. Chem. Lett., 2022, 55, 128449 CrossRef CAS PubMed.
- J. Tian, C. Li, Z. Dong, Y. Yang, J. Xing, P. Yu, Y. Xin, F. Xu, L. Wang and Y. Mu, Nat. Metab., 2023, 5, 896–909 CrossRef CAS PubMed.
- L. K. Tijare, L. T. Deshmukh, A. A. Mohod, V. M. Bhandakkar, S. Hore and N. N. Padole, Asian J. Pharm. Res. Dev., 2022, 10, 90–110 CrossRef CAS.
- H.-M. Kim and C.-G. Hyun, Molecules, 2022, 28, 115 CrossRef PubMed.
- H. S. Yee and N. T. Fong, Pharmacotherapy, 1996, 16, 792–805 CrossRef CAS PubMed.
- D. Ismail and D. S. A. Deshmukh, Diabetes, 2012, 7, 8 Search PubMed.
- C. F. Deacon, Peptides, 2018, 100, 150–157 CrossRef CAS PubMed.
- R. Yin, Y. Xu, X. Wang, L. Yang and D. Zhao, Molecules, 2022, 27, 3055 CrossRef CAS PubMed.
- E. Z. Fisman and A. Tenenbaum, Cardiovasc. Diabetol., 2015, 14, 129 CrossRef PubMed.
- K. Saini, S. Sharma and Y. Khan, Front. Mol. Biosci., 2023, 10, 1130625 CrossRef CAS PubMed.
- R. Liu, C. Mathieu, J. Berthelet, W. Zhang, J.-M. Dupret and F. Rodrigues Lima, Int. J. Mol. Sci., 2022, 23, 7027 CrossRef CAS PubMed.
- Z. Liu, H. Gao, Z. Zhao, M. Huang, S. Wang and J. Zhan, Biomed. Pharmacother., 2023, 157, 113990 CrossRef PubMed.
- X. H. Makhoba, C. Viegas Jr, R. A. Mosa, F. P. Viegas and O. J. Pooe, Drug Des., Dev. Ther., 2020, 3235–3249 CrossRef CAS PubMed.
- M. Nadeem, M. W. Mumtaz, M. Danish, U. Rashid, H. Mukhtar and A. Irfan, Ind. Crops Prod., 2020, 152, 112445 CrossRef CAS.
- A. Ahmad, F. Ullah, A. Sadiq, M. Ayaz, M. Saeed Jan, M. Shahid, A. Wadood, F. Mahmood, U. Rashid and R. Ullah, Drug Des., Dev. Ther., 2020, 2165–2178 CrossRef CAS PubMed.
- M. Shah, M. S. Jan, A. Sadiq, S. Khan and U. Rashid, Eur. J. Med. Chem., 2023, 258, 115591 CrossRef CAS PubMed.
- B. Mobeen, M. Shah, H. M. Rehman, M. S. Jan and U. Rashid, Eur. J. Med. Chem., 2024, 279, 116834 CrossRef CAS PubMed.
- S. Lata, L. Choudhary, A. Bharwal, A. Pandit and V. Abbot, Med. Chem., 2025, 21(10), 1051–1071 CrossRef CAS PubMed.
- O. M. Hendawy, Arch. Pharm., 2022, 355, 2200045 CrossRef CAS PubMed.
- A. Bukhari, H. Nadeem, M. Imran and S. A. Muhammad, Iran. J. Basic Med. Sci., 2021, 24, 1632 Search PubMed.
- F. Hussain, A. Tahir, H. M. Rehman, Y. Wu, M. Shah and U. Rashid, Eur. J. Med. Chem., 2025, 117327 CrossRef CAS PubMed.
- Organisation for Economic Co-operation and Development, Test No. 425: Acute Oral Toxicity: Up-And-Down Procedure, OECD Publishing, 2008 Search PubMed.
- U. Acar Çevik, I. Celik, L. Paşayeva, H. Fatullayev, H. E. Bostancı, Y. Özkay and Z. A. Kaplancıklı, Arch. Pharm., 2023, 356, 2200663 CrossRef PubMed.
- V. Kamat, D. A. Barretto, B. Poojary, A. Kumar, V. B. Patil and S. Hamzad, Bioorg. Chem., 2024, 143, 107085 CrossRef CAS PubMed.
- U. Arshad, S. Ahmed, N. Shafiq, Z. Ahmad, A. Hassan, N. Akhtar, S. Parveen and T. Mehmood, Molecules, 2021, 26, 4424 CrossRef CAS PubMed.
- S. Nordhoff, S. Bulat, S. Cerezo-Gálvez, O. Hill, B. Hoffmann-Enger, M. López-Canet, C. Rosenbaum, C. Rummey, M. Thiemann and V. G. Matassa, Bioorg. Med. Chem. Lett., 2009, 19, 6340–6345 CrossRef CAS PubMed.
- P. B. Jadhav, S. B. Jadhav, M. Zehravi, M. S. Mubarak, F. Islam, P. Jeandet, S. L. Khan, N. Hossain, S. Rashid and L. C. Ming, Molecules, 2022, 28, 149 CrossRef PubMed.
- S. M. Sireesha and B. D. Prasad, J. Mol. Struct., 2025, 1343, 142771 CrossRef CAS.
- J. Upadhyay and A. Gajjar, ChemistrySelect, 2024, 9, e202403537 CrossRef CAS.
- M. H. Mahnashi, Y. S. Alqahtani, A. O. Alqarni, B. A. Alyami, M. S. Jan, M. Ayaz, F. Ullah, U. Rashid and A. Sadiq, BMC Complementary Med. Ther., 2021, 21, 270 CrossRef PubMed.
- G. Cavallo, P. Metrangolo, R. Milani, T. Pilati, A. Priimagi, G. Resnati and G. Terraneo, Chem. Rev., 2016, 116(4), 2478–2601 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |