 Open Access Article
Open Access ArticleDNA/BSA binding, antidiabetic and antioxidant studies of copper(II) complexes derived from ONO-tridentate Schiff bases and diimines as auxiliary ligands
Segun D. Oladipo*a,
Adesola A. Adelekeb,
Kolawole A. Olofisanc,
Bernard Omondib and
Robert C. Luckay *a
*a
aDepartment of Chemistry and Polymer Science, Stellenbosch University, Private Bag X1, Matieland 7602, South Africa. E-mail: segun.oladipo@oouagoiwoye.edu.ng; rcluckay@sun.ac.za
bSchool of Agriculture and Science, Discipline of Chemistry, University of Kwazulu-Natal, Private Bag X54001, Durban, South Africa
cDepartment of Pharmacology, University of Free State, Bloemfontein 9300, South Africa
First published on 10th April 2026
Abstract
Six new copper(II) complexes with general molecular formula [CuL1(Phen)] (1), [CuL2(Phen)] (2), [CuL3(Phen)] (3), [CuL1(DM-Phen)] (4), [CuL2(DM-Phen)] (5) and [CuL3(DM-Phen)] (6) (L = (E)-1-(((4-chloro-2-hydroxyphenyl)imino)methyl)naphthalen-2-ol (L1), (E)-1-(((2-hydroxy-5-methylphenyl)imino)methyl)naphthalen-2-ol (L2), (E)-1-(((2-hydroxy-4-nitrophenyl)imino)methyl)naphthalen-2-ol (L3), Phen = 1,10-phenanthroline and DM-Phen = 2,9-dimethyl-1,10-phenanthroline) have been synthesized and characterized by spectroscopic as well as crystallographic techniques. Molecular structures of 2, 4, 5 and 6 revealed that they are mononuclear species where the copper(II) center is five-coordinated to a pair of oxygen atoms as well as one nitrogen atom from the ligands (L1–L3) and to a pair of nitrogen atoms from the auxiliary ligand (Phen or DM-Phen) conforming to a distorted square pyramid. The binding affinities of ligands L1–L3 and copper(II) complexes 1–6 with calf thymus DNA (ctDNA) and bovine serum albumin (BSA) were evaluated using UV-visible absorption spectroscopy. For ctDNA, all compounds showed hypochromism and bathochromic shifts suggesting intercalative binding, with Kb values ranging from 1.49 × 104 to 2.74 × 105 M−1. The complexes exhibited 3–18-fold higher affinity than ligands, especially those with 2,9-dimethyl-1,10-phenanthroline coligands. BSA titrations of complexes 1–6 revealed hyperchromic shifts at ∼280 nm, with Kb values of 9.59 × 104 to 2.41 × 105 M−1. These moderate-to-strong bindings were enhanced by lipophilic substituents that promote hydrophobic and π-stacking interactions in protein pockets. All of the compounds inhibit α-amylase better than acarbose with complex 5 (IC50 = 0.274 mM) showing the highest activity. For the α-glucosidase assay, only complexes 2 (IC50 = 0.055 mM) and 4 (IC50 = 0.054 mM) outshined acarbose (IC50 = 0.059 mM) while other compounds showed moderate to good α-glucosidase inhibition activity. Antioxidant activities of the free ligands showed better activity than that of the metal complexes with none of the compounds performing better than quercetin in all of the assays explored for this study.
1 Introduction
The research on the design and development of metallodrugs with multifunctional therapeutic activities continues to gain significant interest in the field of bioinorganic and medicinal chemistry.1,2 Among metal ions explored in metal-based drugs, copper has been unique due to its important role in biological systems. Copper exists in crucial enzymes (i.e., ceruloplasmin, laccase, superoxide dismutase, ascorbate oxidase, among others).3,4 These enzymes are essential in energy metabolism and nervous system development.5 It is found in humans either as Cu2+ (oxidized form) or Cu+ (reduced form). While high doses of copper are toxic, copper(II) ions are less toxic than copper(I) ions at appropriate concentrations. This is because, in aqueous solutions, copper(II) is the most stable oxidation state, while copper(I) is highly prone to oxidation (less stable) and not easy to develop into reliable oral medication. In pharmaceuticals, copper-based drugs or supplements are mainly copper(II) salts or complexes, i.e., copper(II) glycinate and copper(II) citrate, both common in supplements.6,7 Copper(II) participates in diverse physiological processes, including enzymatic catalysis, electron transfer, and oxidative stress regulation, making copper-based complexes potential candidates for therapeutic application.8Schiff base ligands have been widely explored as chelate ligands in coordination chemistry.9,10 They are well known for their structural versatility, robust metal-binding ability, and tunable electronic properties. The bio-active azomethine functional group, which allows nucleophilic and electrophilic reactions at the electron-rich nitrogen and electron-deficient carbon atoms, respectively, has been linked to the inherent therapeutic qualities of Schiff bases.11,12 Copper(II) complexes derived from Schiff bases have been reported to display a myriad of medicinal properties, which include anticancer, antioxidant, antimicrobial, and antifungal with enhanced medicinal activity compared to the free ligands.13,14 However, the incorporation of planar nitrogen heterocyclic (N-heterocyclic) i.e., 2,2-bipyridine, 1,10-phenanthroline, and 2,9-dimethyl-1,10-phenanthroline, is known to further enhance the biological activities of copper(II) complexes. These N-heterocyclic ligands are well known for their strong π–π stacking interaction with nucleic acid–base pairs of DNA, hence facilitating DNA intercalation.15
DNA binding studies play an important role in developing DNA molecule probes and chemotherapeutics.16,17 It also helps in understanding the mechanisms of action of metal-based drugs, as interaction with DNA can influence replication and transcription processes.18 Strong binding affinities for DNA via intercalative, groove-binding, or electrostatic modes have been shown by copper(II) complexes, especially those with Schiff bases and N-heterocyclic ligands and these affinities do corroborate with biological activities.19–23 Hence, the need to evaluate the DNA binding potential of newly synthesized compounds is crucial to elucidate possible mechanisms of biological activity. Similarly, BSA binding studies are equally vital, as bovine serum albumin (BSA), a model plasma protein, transports metal complexes in blood, influencing their bioavailability, distribution, and toxicity. Copper(II) Schiff base complexes often exhibit strong BSA affinity through hydrophobic interactions or hydrogen bonding with its Sudlow sites, which correlates with enhanced therapeutic efficacy.24 These studies provide insights into drug–protein interactions, complementing DNA binding data for multifunctional agents.
In addition to the DNA and BSA binding potential of copper(II) complexes, they have also attracted interest as antidiabetic agents, through the inhibition of carbohydrate-hydrolyzing enzymes such as α-glucosidase and α-amylase enzymes.25–27 Inhibiting these enzymes can reduce postprandial hyperglycemia by retarding the rate at which dietary starch breaks down to glucose in the body and this has been validated as an effective strategy to manage diabetes.28 It is important to note that, drugs like acarbose and miglitol have been successfully used to treat diabetes by acting as α-glucosidase and α-amylase inhibitors. However, adverse effects associated with their usage i.e., stomach discomfort, meteorism, and diarrhea due to undigested carbohydrate fermenting in the bowel, have significantly decreased their use as antidiabetic medication.29,30 Hence, the need for new α-glucosidase and α-amylase inhibitors with little or no side effects.
Interestingly, oxidative stress has been associated with the progression and complications of diabetes, highlighting the relevance of antioxidant activity in potential antidiabetic drugs.31–33 Aside from that free radicals are closely linked to the cause of diabetes, they are also associated to deadly diseases such as liver cirrhosis, cancer, atherosclerosis among others.34 Hence, an antioxidant drug could play a crucial chemoprotective role in the treatment of diabetes and other diseases. Copper(II) complexes do display radical scavenging activity due to facile redox cyclic between Cu(II) and Cu(I) states, contributing to their therapeutic relevance.14 Copper(II) complexes derived from Schiff bases have demonstrated significant antioxidant activities in multiple radical scavenging assays.9,14,35
In this study, the ligand design was based on introducing controlled variations in electronic, steric and physicochemical properties. Chloro and nitro substituents were selected as electron-withdrawing groups, while the methyl group was included as an electron-donating substituent to facilitate comparison of electronic effects on the resulting copper(II) complexes. Furthermore, 1,10-phenanthroline and 2,9-dimethyl-1,10-phenanthroline were used as co-ligands to probe steric and lipophilic influences, as methyl substitution at the 2 and 9 positions may increase steric hindrances and hydrophobicity. These variations were considered in relation to their potential impact on biological interactions, including DNA binding affinity and inhibitory activity against α-amylase and α-glucosidase enzymes. Herein, we report the synthesis and characterization of copper(II) complexes derived from ONO-donor Schiff base ligands and N-heterocyclic (1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline) as coligands. Their DNA/BSA binding, antidiabetic, and antioxidant potential are investigated to evaluate their suitability as multifunctional bioactive agents.
2 Experimental
2.1 Reagents
The reagents used for the preparation of the metal complexes are 1,10-phenanthroline (≥99%), 2,9-dimethyl-1,10-phenanthroline (≥98%), 2-hydroxy-4-nitroaniline (98%), 2-amino-5-chlorophenol (97%), 2-hydroxy-5-methylaniline (97%), acetic acid, and copper(II) acetate monohydrate (≥98%). Solvents such as dimethyl sulfoxide (DMSO), methanol and diethyl ether are A.C.S. grade and used as obtained. All reagents are purchased from Merck.2.2 Instrumentation
The FT-IR spectra for complexes 1–6 were obtained using a PerkinElmer Universal A.T.R. spectrum 100 FT-IR spectrometer, and the elemental/combustion analysis for the complexes was carried out using a Vario elemental E.L. cube CHNS analyser. Shimadzu UV-Vis-NIR spectrophotometer was used to obtain the UV-Visible spectra of all the complexes and Waters Synapt G2 coupled to a Waters UPLC., ESI probe, ESI Positive, Cone Voltage 15 V was used to analyze the mass spectra for the complexes. The JEOL FA200 spectrometer was used to record the electron paramagnetic resonance (EPR) spectra for compounds 1–6.2.3 Preparation of complexes 1–6
We have previously reported the synthetic procedures as well as fully characterized the Schiff base ligands L1–L3 used in the preparation of complexes 1–6.36 For the preparation of complexes 1–6, we followed the procedure reported by Kiran et al.37 with slight modification. 0.751 mmol (0.150 g) of copper(II) acetate monohydrate was dissolved in 20 mL of methanol in 100 mL round bottom flask and stirred until complete dissolution. Thereafter, 0.751 mmol of methanolic solution of 1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline was added to the copper(II) salt solution and heated to around 40–50 °C while stirring vigorously for 1 h. For the 1,10-phenanthroline, the solution turned deep blue, while the one for 2,9-dimethyl-1,10-phenanthroline, it turned light green. 0.751 mmol of L1–L3 dissolved in 20 mL of dimethyl sulfoxide is added to the reaction mixture in a round bottom flask and further stirred for another 1 h. After 10 min of stirring, a reddish-brown precipitate crashed out of the reaction mixture, which was later washed with methanol and rinsed four times with diethyl ether. The obtained products were dried in an oven at 60–70 °C.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1600), (νC
N = 1600), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1576), (νC–N = 1395), (νC–O = 1291), and (νCu–N = 557). ES-TOF MS: m/z(%): [M + H]+, 539.0454, [M + Na-naph]+, 436.9912 MS calc. analysis: 538.04 (100.0%), 540.04 (44.6%). Anal. calc. for C29H18ClCuN3O20.5C2H6OS: C, 62.28, H, 3.66, N, 7.26. Found: C, 62.30, H, 3.89, N, 7.46.
C = 1576), (νC–N = 1395), (νC–O = 1291), and (νCu–N = 557). ES-TOF MS: m/z(%): [M + H]+, 539.0454, [M + Na-naph]+, 436.9912 MS calc. analysis: 538.04 (100.0%), 540.04 (44.6%). Anal. calc. for C29H18ClCuN3O20.5C2H6OS: C, 62.28, H, 3.66, N, 7.26. Found: C, 62.30, H, 3.89, N, 7.46.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1600), (νC
N = 1600), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1583), (νC–N = 1388), (νC–O = 1299), and (νCu–N = 557). ES-TOF MS: m/z(%): [M + H]+, 519.100, [M + Na-naph]+, 417.0460 MS calc. analysis: 518.09 (100.0%), 520.09 (44.6%), 519.10 (32.4%). Anal. calc. for C30H21CuN3O2: C, 69.42, H, 4.08, N, 8.10. Found: C, 69.30, H, 3.42, N, 8.04.
C = 1583), (νC–N = 1388), (νC–O = 1299), and (νCu–N = 557). ES-TOF MS: m/z(%): [M + H]+, 519.100, [M + Na-naph]+, 417.0460 MS calc. analysis: 518.09 (100.0%), 520.09 (44.6%), 519.10 (32.4%). Anal. calc. for C30H21CuN3O2: C, 69.42, H, 4.08, N, 8.10. Found: C, 69.30, H, 3.42, N, 8.04.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1598), (νC
N = 1598), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1584), (νC–N = 1365), (νC–O = 1284), and (νCu–N = 572). ES-TOF MS: m/z(%): [M + H]+, 550.0701, [M + Na-naph]+, 448.0151 MS calc. analysis: 549.06 (100.0%), 551.06 (44.6%), 550.07 (31.4%). Anal. calc. for C29H18CuN4O40.5C2H6OS: C, 61.17, H, 3.59, N, 9.51. Found: C, 61.29, H, 3.87, N, 9.82.
C = 1584), (νC–N = 1365), (νC–O = 1284), and (νCu–N = 572). ES-TOF MS: m/z(%): [M + H]+, 550.0701, [M + Na-naph]+, 448.0151 MS calc. analysis: 549.06 (100.0%), 551.06 (44.6%), 550.07 (31.4%). Anal. calc. for C29H18CuN4O40.5C2H6OS: C, 61.17, H, 3.59, N, 9.51. Found: C, 61.29, H, 3.87, N, 9.82.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1601), (νC
N = 1601), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1587), (νC–N = 1394), (νC–O = 1290), and (νCu–N = 569). ES-TOF MS: m/z(%): [M + H]+, 567.0773, [M + Na-ArCl]+, 479.1292 MS calc. analysis: 566.07 (100.0%), 568.07 (44.6%), 567.07 (33.5%). Anal. calc. for C29H18CuN4O40.5C2H6OS: C, 64.22, H, 4.06, N, 7.11. Found: C, 64.44, H, 4.26, N, 7.24.
C = 1587), (νC–N = 1394), (νC–O = 1290), and (νCu–N = 569). ES-TOF MS: m/z(%): [M + H]+, 567.0773, [M + Na-ArCl]+, 479.1292 MS calc. analysis: 566.07 (100.0%), 568.07 (44.6%), 567.07 (33.5%). Anal. calc. for C29H18CuN4O40.5C2H6OS: C, 64.22, H, 4.06, N, 7.11. Found: C, 64.44, H, 4.26, N, 7.24.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1601), (νC
N = 1601), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1585), (νC–N = 1389), (νC–O = 1275), and (νCu–N = 555). ES-TOF MS: m/z(%): [M + Na-ArCH3]+, 479.1297 [M + H]+, 547.12 MS calc. analysis: 546.1 (100.0%), 548.12 (44.6%), 547.13 (34.6%). Anal. calc. for C32H25CuN3O20.3C2H6OS: C, 68.63, H, 4.73, N, 7.38. Found: C, 68.30, H, 4.78, N, 7.24.
C = 1585), (νC–N = 1389), (νC–O = 1275), and (νCu–N = 555). ES-TOF MS: m/z(%): [M + Na-ArCH3]+, 479.1297 [M + H]+, 547.12 MS calc. analysis: 546.1 (100.0%), 548.12 (44.6%), 547.13 (34.6%). Anal. calc. for C32H25CuN3O20.3C2H6OS: C, 68.63, H, 4.73, N, 7.38. Found: C, 68.30, H, 4.78, N, 7.24.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N = 1600), (νC
N = 1600), (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C = 1587), (νC–N = 1367), (νC–O = 1277), and (νCu–N = 571). ES-TOF MS: m/z(%): [M + Na-ArNO2]+, 479.1288 MS calc. analysis: 577.09 (100.0%), 579.09 (44.6%), 578.10 (33.5%). Anal. calc. for C31H22CuN4O40.5C2H6OS: C, 62.88, H, 4.08, N, 9.08. Found: C, 62.74, H, 3.92, N, 9.21.
C = 1587), (νC–N = 1367), (νC–O = 1277), and (νCu–N = 571). ES-TOF MS: m/z(%): [M + Na-ArNO2]+, 479.1288 MS calc. analysis: 577.09 (100.0%), 579.09 (44.6%), 578.10 (33.5%). Anal. calc. for C31H22CuN4O40.5C2H6OS: C, 62.88, H, 4.08, N, 9.08. Found: C, 62.74, H, 3.92, N, 9.21.2.4 Single crystal X-ray crystallography
Crystals with dimensions of 0.258 × 0.075 × 0.032 mm3 (for 2), 0.231 × 0.117 × 0.107 mm3 (for 4), 0.483 × 0.221 × 0.074 mm3 (for 5), and 0.153 × 0.140 × 0.039 mm3 (for 6) were mounted on a Bruker DUO Apex II X-ray diffractometer equipped with an INCOATEC IµS HB microfocus tube (MoKα radiation λ = 0.71073 Å) fitted with a multilayer monochromator. Data were captured with a CCD (charge-coupled device) area detector. Data collection was carried out at 100 K using an Oxford Cryosystems cryostat (700 series Cryostream Plus) attached to the diffractometer. Data collection and reduction were carried out using the Bruker software package APEX4,38 using standard procedures. All structures were solved and refined using SHELX-201639 employed within the X-Seed40 environment. Hydrogen atoms were placed in calculated positions using riding models. Diagrams for the crystal structures were obtained by Mercury software.41 Table 1 details the structural data together with the refinement parameters.| 2 | 4 | 5 | 6 | |
|---|---|---|---|---|
| Empirical formula | C31H23Cl2CuN3O2 | C32H23Cl4CuN3O2 | C33H29CuN3O3 | C32H24Cl2CuN4O4 |
| Formula weight | 603.96 | 686.87 | 579.13 | 662.99 |
| Crystal system | Monoclinic | Monoclinic | Monoclinic | Triclinic |
| Space group | P21/c | P21/c | P21/n | P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) |
| a/Å | 10.1131(6) | 10.3155(3) | 9.6459(6) | 7.418(4) |
| b/Å | 18.2779(10) | 22.5459(6) | 20.2675(13) | 13.351(7) |
| c/Å | 14.6964(8) | 13.4635(4) | 14.4479(9) | 15.437(8) |
| α/° | 90 | 90 | 90 | 93.220(16) |
| β/° | 105.1330(10) | 107.6710(10) | 108.352(2) | 103.676(15) |
| γ/° | 90 | 90 | 90 | 103.705(16) |
| Volume/Å3 | 2622.4(3) | 2983.49(15) | 2680.9(3) | 1433.1(14) |
| Z | 4 | 4 | 4 | 2 |
| ρcalc/g cm−3 | 1.530 | 1.529 | 1.435 | 1.536 |
| µ/mm−1 | 1.072 | 1.126 | 0.856 | 0.995 |
| F(000) | 1236 | 1396 | 1204 | 678 |
| Crystal size/mm3 | 0.258 × 0.075 × 0.032 | 0.231 × 0.117 × 0.107 | 0.483 × 0.221 × 0.074 | 0.153 × 0.140 × 0.039 |
| 2Θ range for data collection/° | 1.817 to 26.410 | 1.806 to 26.386 | 1.793 to 26.420 | 1.581 to 26.582 |
| Index ranges | −12 ≤ h ≤ 12 | −12 ≤ h ≤ 12 | −12 ≤ h ≤ 11 | −9 ≤ h ≤ 9 |
| −22 ≤ k ≤ 22 | −28 ≤ k ≤ 28 | −25 ≤ k ≤ 25 | −16 ≤ k ≤ 16 | |
| −18 ≤ l ≤ 18 | −16 ≤ l ≤ 16 | −18 ≤ l ≤ 17 | −19 ≤ l ≤ 19 | |
| Reflections collected | 53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 446 446 |
67![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 118 118 |
64![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 228 228 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 441 441 |
| Independent reflections | 5383 [Rint = 0.0653] | 6105 [Rint = 0.0410] | 5477 [Rint = 0.0477] | 5874 [Rint = 0.0949] |
| Data/restraints/parameters | 5383/0/353 | 6105/0/381 | 5477/0/366 | 5874/718/418 |
| Goodness-of-fit on F2 | 1.056 | 1.031 | 1.040 | 0.992 |
| Final R indexes [I ≥ 2σ(I)] | R1 = 0.0391, wR2 = 0.0910 | R1 = 0.0424, wR2 = 0.1077 | R1 = 0.0479, wR2 = 0.1139 | R1 = 0.0590, wR2 = 0.1492 |
| Final R indexes [all data] | R1 = 0.0602, wR2 = 0.1008 | R1 = 0.0610, wR2 = 0.1216 | R1 = 0.0614, wR2 = 0.1210 | R1 = 0.0951, wR2 = 0.1714 |
| Largest diff. peak and hole (e Å−3) | 0.453 and −0.399 | 0.516 and −0.370 | 1.063 and −0.442 | 0.399 and −0.949 |
2.5 DNA binding study
The DNA binding study was done using modified literature procedures.42,43 UV-visible absorption spectroscopy was employed to investigate the binding affinity of ligands L1–L3 and complexes 1–6 with calf thymus DNA (ctDNA). The ctDNA stock solution was prepared by dissolving an appropriate amount in phosphate buffer saline (pH 7.2), with its concentration quantified spectrophotometrically using the molar absorptivity coefficient ε260 = 6600 M−1 cm−1. The purity of the ctDNA solution was confirmed by a UV absorbance ratio A260/A280 > 1.8, verifying the absence of protein contamination. Binding interactions were evaluated by maintaining a fixed compound concentration of 40 µM while incrementally adding ctDNA (0–25 µM). The compound-DNA mixtures were incubated for 5 min at 25 °C prior to recording absorption spectra. Intrinsic binding constants Kb were determined by nonlinear fitting of the data to the Wolfe–Shimmer equation (eqn (1)). The experiments were performed in triplicate to confirm the precision and accuracy of the procedure and in all the experiments on the interaction of the compounds with ctDNA, the final concentration of DMSO in the incubation mixture was ∼2% (v/v).
 | (1) |
2.6 Protein binding studies
The interactions of complexes 1–6 were performed using the UV-vis absorption titration method.44 BSA stock solutions were prepared by dissolving the protein in phosphate-buffered saline (pH 7.2) at 25 °C under constant stirring, with concentrations determined spectrophotometrically using the molar absorptivity coefficient ε280 = 44![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 M−1 cm−1. The complexes' stock solutions (1 × 103 mM) were dissolved in DMSO. Titrations were done by adding varying complex concentrations (0–10 µM) to a fixed BSA concentration (6 µM), followed by 5 min incubation at 25 °C. The experiments were performed in triplicate to confirm the precision and accuracy of the procedure, and in all the experiments on the interaction of the compounds with ctDNA, the final concentration of DMSO in the incubation mixture was ∼1% (v/v). Absorption at λmax = 280 nm was measured, and the binding constant Kb was calculated using the Benesi–Hildebrand equation (eqn (2)) from the intercept to the slope ratio of 1/[A − A0] vs. 1/[complex].
300 M−1 cm−1. The complexes' stock solutions (1 × 103 mM) were dissolved in DMSO. Titrations were done by adding varying complex concentrations (0–10 µM) to a fixed BSA concentration (6 µM), followed by 5 min incubation at 25 °C. The experiments were performed in triplicate to confirm the precision and accuracy of the procedure, and in all the experiments on the interaction of the compounds with ctDNA, the final concentration of DMSO in the incubation mixture was ∼1% (v/v). Absorption at λmax = 280 nm was measured, and the binding constant Kb was calculated using the Benesi–Hildebrand equation (eqn (2)) from the intercept to the slope ratio of 1/[A − A0] vs. 1/[complex].
 | (2) |
2.7 Antidiabetes studies
2.8 Antioxidant studies
3 Results and discussion
3.1 Preparation of complexes 1–6
The free ligands L1–L3 explored in synthesizing complexes 1–6 were prepared by the condensation reaction between 2-hydroxy-1-napthaldehyde and respective primary amines viz; 2-amino-5-chlorophenol (L1), 2-hydroxy-5-methylaniline (L2) and 2-hydroxy-4-nitroaniline (L3) in ratio 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 at room temperature. The reaction between ligand (L1/L2/L3), copper(II) salt (Cu(CH3COO)2.H2O) and N,N′-adducts ligands (1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline) at 40–50 °C in 1
1 at room temperature. The reaction between ligand (L1/L2/L3), copper(II) salt (Cu(CH3COO)2.H2O) and N,N′-adducts ligands (1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline) at 40–50 °C in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 respectively affords complexes 1–6 (Scheme 1) in good yield. The resulting copper(II) complexes are reddish brown in colour and very stable at room temperature. Generally, the complexes dissolve readily in DMSO, dichloromethane as well as chloromethane, while they do not dissolve readily in tetrahydrofuran and protic solvents i.e., ethanol and methanol. They are completely insoluble in hexane.
1 respectively affords complexes 1–6 (Scheme 1) in good yield. The resulting copper(II) complexes are reddish brown in colour and very stable at room temperature. Generally, the complexes dissolve readily in DMSO, dichloromethane as well as chloromethane, while they do not dissolve readily in tetrahydrofuran and protic solvents i.e., ethanol and methanol. They are completely insoluble in hexane.
3.2 Spectroscopic elucidation of complexes 1–6
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N)str which is the diagnostic peak for Schiff base derivatives was observed around 1598–1601 cm−1 in 1–6 and when compared to the ones for the free ligands, L1–L3 (1612–1616 cm−1) previously reported, it appeared at a lower vibrational frequency, and this confirmed the involvement of the azomethine nitrogen atom (CH
N)str which is the diagnostic peak for Schiff base derivatives was observed around 1598–1601 cm−1 in 1–6 and when compared to the ones for the free ligands, L1–L3 (1612–1616 cm−1) previously reported, it appeared at a lower vibrational frequency, and this confirmed the involvement of the azomethine nitrogen atom (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) in coordination to the copper(II) metal center. Furthermore, the absence of a pronounced vibrational broad band within the region of 3200–3500 cm−1 unravel the deprotonation of the hydroxyl group and coordination of the resulting phenolate as well as naphtholate oxygen to the copper(II) center. The vibrational bands which were observed around 500–650 cm−1 in the spectra of all the complexes might be due to the formation of Cu–O and Cu–N bonds.
N) in coordination to the copper(II) metal center. Furthermore, the absence of a pronounced vibrational broad band within the region of 3200–3500 cm−1 unravel the deprotonation of the hydroxyl group and coordination of the resulting phenolate as well as naphtholate oxygen to the copper(II) center. The vibrational bands which were observed around 500–650 cm−1 in the spectra of all the complexes might be due to the formation of Cu–O and Cu–N bonds.The electronic absorption spectra of the compounds were recorded within the range of 200–800 nm using the dichloromethane solution of the respective compound (Fig. 1). In the ultraviolet region, one intense band as well as one weak shoulder band were observed around 265–270 nm and 360–365 nm and these bands can be attributed to π → π* and n → π*, respectively due to the intra-ligand transition within the complexes. In the visible region, an intense band can also be observed around 400–600 nm in the spectra of complexes 1–6, and this is due to metal-to-ligand charge transfer (LMCT). It could be observed that the absorption band for complexes 3 and 6 with nitro (–NO2) moiety appeared at a longer wavelength. This is because the nitro group is a strong electron-withdrawing group which acts as a strong chromophore that elongates the absorption range into longer wavelengths by lowering the electronic energy gap. A very weak band due to d–d transitions were also observed around 600–650 nm in the electronic spectra of the complexes.
3.3 Single crystal X-ray structures
We used the single crystal X-ray crystallography technique to determine the structures of complexes 2, 4, 5 and 6. Single crystals suitable for this experiment were obtained by the slow evaporation of 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 dichloromethane: methanol or chloromethane: methanol solution. Fig. 3 shows the crystal structures of the complexes using an ORTEP diagram with 50% probability thermal ellipsoids, while Table 2 details selected bond angles as well as bond lengths. Complexes 2, 4 and 5 crystallize as monoclinic with a space group of P21/c for complexes 2 and 4 and P21/n for 5 while complex 6 crystallizes as triclinic with a space group of P
1 dichloromethane: methanol or chloromethane: methanol solution. Fig. 3 shows the crystal structures of the complexes using an ORTEP diagram with 50% probability thermal ellipsoids, while Table 2 details selected bond angles as well as bond lengths. Complexes 2, 4 and 5 crystallize as monoclinic with a space group of P21/c for complexes 2 and 4 and P21/n for 5 while complex 6 crystallizes as triclinic with a space group of P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) . Generally, the asymmetric unit of the complexes contain a whole molecule with one molecule of solvent i.e. one molecule of dichloromethane for 2 and 6, one molecule of chloromethane for 4, and one molecule of methanol for 6. All the crystal structures are mononuclear species in which the copper(II) ion is five-coordinated to a pair of oxygen atoms as well as one nitrogen atom from the Schiff base ligand (L1–L3) and to a pair of nitrogen atoms from the auxiliary ligand (1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline) conforming to a distorted square pyramid with O1N7O2N1 atoms forming the basal plane for Cu1. The apical Cu1–N2 bond length for 2, 5 and 6 and Cu1–N1 for 3 are longer than other two Cu–N bond lengths. For example, the Cu1–N2 bond length is 2.272(2)° in complex 2, while the ones for Cu1–N1 as well as Cu1–N7 are 2.049(2)° and 1.944(2)° respectively. We used the structural parameter index (eqn (3)) for five coordinated complexes51 to affirm the geometry of 2, 4, 5 and 6.
. Generally, the asymmetric unit of the complexes contain a whole molecule with one molecule of solvent i.e. one molecule of dichloromethane for 2 and 6, one molecule of chloromethane for 4, and one molecule of methanol for 6. All the crystal structures are mononuclear species in which the copper(II) ion is five-coordinated to a pair of oxygen atoms as well as one nitrogen atom from the Schiff base ligand (L1–L3) and to a pair of nitrogen atoms from the auxiliary ligand (1,10-phenanthroline or 2,9-dimethyl-1,10-phenanthroline) conforming to a distorted square pyramid with O1N7O2N1 atoms forming the basal plane for Cu1. The apical Cu1–N2 bond length for 2, 5 and 6 and Cu1–N1 for 3 are longer than other two Cu–N bond lengths. For example, the Cu1–N2 bond length is 2.272(2)° in complex 2, while the ones for Cu1–N1 as well as Cu1–N7 are 2.049(2)° and 1.944(2)° respectively. We used the structural parameter index (eqn (3)) for five coordinated complexes51 to affirm the geometry of 2, 4, 5 and 6.
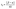 | (3) |
 | ||
| Fig. 3 Crystal structure of copper(II) complexes 2, 4, 5, and 6 with thermal ellipsoids drawn at 50% probability. Hydrogen atoms were removed for clarity. | ||
| Parameters | 2 | 4 | 5 | 6 |
|---|---|---|---|---|
| C8–N7 | 1.296(4) | 1.294(4) | 1.301(4) | 1.311(4) |
| C10–O2 | 1.295(4) | 1.302(4) | 1.302(3) | 1.298(5) |
| C1–O1 | 1.328(3) | 1.321(4) | 1.330(3) | 1.316(4) |
| Cu1–O1 | 1.933(2) | 1.977(2) | 1.954(2) | 1.942(3) |
| Cu1–O2 | 1.918(2) | 1.930(2) | 1.918(2) | 1.911(3) |
| Cu1–N7 | 1.944(2) | 1.945(3) | 1.955(2) | 1.955(3) |
| Cu1–N1 | 2.049(2) | 2.303(2) | 2.026(2) | 2.052(3) |
| Cu1–N2 | 2.272(2) | 2.045(3) | 2.288(3) | 2.323(4) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Bond angles | ||||
| O2–Cu1–O1 | 171.20(8) | 168.09(9) | 166.63(9) | 171.4(1) |
| N2–Cu1–N1 | 77.75(8) | 76.9(1) | 78.0(1) | 76.4(1) |
| N7–Cu1–O1 | 85.32(8) | 83.68(9) | 84.4(1) | 84.6(1) |
| N7–Cu1–O2 | 92.23(8) | 90.92(9) | 92.4(1) | 91.8(1) |
| O2–Cu1–N2 | 91.64(8) | 91.2(1) | 98.18(9) | 92.0(1) |
| O1–Cu1–N1 | 90.18(8) | 94.24(9) | 91.6(1) | 89.0(1) |
3.4 DNA binding study of ligands L1–L3 and complexes 1–6
The UV-visible absorption spectroscopy technique is commonly employed to study the noncovalent DNA binding modes for potential metallodrugs, particularly those targeting DNA replication in anticancer therapies. In this study, we evaluate the binding affinities of ligands L1–L3 and complexes 1–6 to calf thymus DNA (ctDNA) by monitoring their spectral changes upon incremental ctDNA addition. All the compounds displayed absorption bands between 447 and 508 nm, showing hypochromism of 9.44–40.69% and bathochromic shifts of 2–9 nm with increasing ctDNA concentration (Fig. 4 and S15–S22). These observations suggest a potential intercalative binding, wherein the aromatic π-systems of the ligands stack with DNA base pairs, reducing transition probabilities. The pronounced hypochromism and red shifts signify strong π–π interactions, common in phenanthroline-containing copper complexes. Using the Wolfe–Shimmer equation (eqn (1)), the intrinsic binding constant (Kb) was calculated to quantify the binding interactions of the compounds. | ||
| Fig. 4 Electronic absorption spectrum of complex 5 in the absence (dashed line) and the presence of different concentrations of ctDNA. | ||
The intrinsic binding constants (Kb) of L1–L3 and complexes 1–6 (Table 3) revealed moderate to strong DNA affinities. The Kb values for L1, L2 and L3 were 1.49 × 104 M−1, 2.46 × 104 M−1, and 1.56 × 104 M−1, respectively (L2 > L3 > L1; Table 3), reflecting the ligands' moderate DNA affinities.
| Cpd. | DNA | BSA | ||||||
|---|---|---|---|---|---|---|---|---|
| Kb (×104 M−1) | λmax (nm) | Spec. Shift (nm) | %Hypo | Kb (×105 M−1) | λmax (nm) | Spec. Shift (nm) | %Hyper | |
| a Kb = intrinsic binding constant; the Kb values shown are mean ± standard deviation. | ||||||||
| L1 | 1.49 ± 0.01 | 471 | 3(red) | 40.69 | — | — | — | — |
| L2 | 2.46 ± 0.06 | 446 | 3(red) | 36.96 | — | — | — | — |
| L3 | 1.56 ± 0.01 | 508 | 9(red) | 9.44 | — | — | — | — |
| 1 | 4.73 ± 0.08 | 504 | 3(red) | 30.63 | 1.81 ± 0.03 | 280 | 7(blue) | 64.42 |
| 2 | 7.15 ± 0.01 | 483 | 2(red) | 32.14 | 2.41 ± 0.06 | 280 | 4(blue) | 51.52 |
| 3 | 9.31 ± 0.11 | 507 | 3(red) | 22.83 | 1.53 ± 0.01 | 280 | 8(blue) | 56.01 |
| 4 | 17.5 ± 0.20 | 462 | 2(red) | 29.73 | 1.57 ± 0.07 | 280 | 3(blue) | 61.55 |
| 5 | 27.4 ± 0.20 | 459 | 5(red) | 36.65 | 1.46 ± 0.02 | 280 | 8(blue) | 60.53 |
| 6 | 24.8 ± 0.05 | 490 | 3(red) | 26.03 | 0.96 ± 0.03 | 280 | 4(blue) | 54.28 |
Ligand L2, bearing a methyl substituent, exhibited the highest affinity due to enhanced π-electron density and hydrophobic interactions with DNA base pairs. In contrast, L1 (chloro-substituted) and L3 (nitro-substituted) showed lower affinities, likely due to the electron-withdrawing groups reducing π-stacking efficiency. Complexes 1–6 showed enhanced binding affinity, with Kb values ranging between 4.73 × 104 and 2.74 × 105 M−1 in the order 5 > 6 > 4 > 3 > 2 > 1. Complex 5 exhibited the highest affinity (Fig. 4), attributable to the synergistic effect of the methyl group on its Schiff base ligand and the 2,9-dimethyl-1,10-phenanthroline coligand, which enhances hydrophobicity and π-stacking relative to unsubstituted phenanthroline.
The obtained Kb values of 1–6 show that the coordination of L1–L3 to the copper(II) centre increased their binding affinities by 3 to 18-fold compared to the free ligands, suggesting metal-assisted stabilization of the potential intercalative mode. Notably, complexes 4–6 (with 2,9-dimethyl-1,10-phenanthroline as co-ligand) show 2–3 times higher Kb values than 1–3, due to methyl-induced hydrophobicity, as such encouraging π-stacking with DNA bases. Overall, the Kb values within 104–105 M−1 recorded for the compounds suggest their non-covalent intercalative binding modes, aligning with literature reports for mixed-ligand copper-phen Schiff base complexes (Kb = ∼104–106 M−1),53,54 positioning these compounds as classical intercalators suitable for biomedical applications.
3.5 Bovine serum albumin interaction study
Drug molecule binding to plasma proteins offers useful information about drug distribution, transportation, and excretion, which is crucial in drug discovery and design. Bovine serum albumin (BSA), the most abundant plasma protein, serves as a cost-effective and structurally similar model to human serum albumin for drug–protein interaction studies. In this study, UV-visible absorption spectroscopy was employed to evaluate the binding affinity of copper(II) complexes 1–6 to BSA. The binding study was monitored by the conformational changes in BSA characterized by its intense absorbance at 280 nm, attributed to the polarity of tyrosine and tryptophan residues. Ligands L1–L3 were excluded, as their ctDNA binding affinities were lower than those of the complexes.Upon titration of increasing complexes 1–6 concentrations into fixed BSA solutions, continuous hyperchromic shifts were observed between 260 and 300 nm, with hyperchromism of 51.52–64.42% and hypsochromic shifts of 3–8 nm, indicating alterations in the protein microenvironment (Fig. 5 and S23–S27).
 | ||
| Fig. 5 Electronic absorption spectra of BSA in the absence (dashed line) and the presence of different concentrations of complex 2. | ||
Binding constants (Kb) were calculated from the intercept-to-slope ratio of linear plots of 1/(A − A0) vs. 1/[complex], where A and A0 are the absorbances with and without complex, respectively. The Kb values for complexes 1–6 ranged from 9.59 × 104 M−1 to 2.41 × 105 M−1 (Table 3), falling within the typical 104 to 106 M−1 range for drug–protein interactions. This suggests their moderate to strong binding to protein, as such, suitable for pharmacological applications.
Complex 2 exhibited the highest binding affinity (Kb = 2.41 × 105 M−1, Fig. 5), followed by complex 1 (1.81 × 105 M−1, Fig. S23) and complex 4 (1.57 × 105 M−1, Fig. S24), highlighting the influence of ligand substituents and co-ligands. In complex 2 (bearing L2 with an electron-donating methyl group on the aniline), the methyl group enhances lipophilicity, thus promoting stronger hydrophobic interactions and π–π stacking with BSA's aromatic residues in hydrophobic pockets like subdomain IIA. For complex 1, the planarity of the unsubstituted 1,10-phenanthroline co-ligand could have influenced the insertion into the binding pocket. Similarly, complex 4's affinity can be related to the steric bulk and hydrophobic effects of the 2,9-dimethyl substituents on 1,10-phenanthroline, which promote stable interactions within the BSA subdomain. These trends correspond with literature reports on copper(II) Schiff base complexes, where substituents modulate binding through steric and electronic effects.55,56 Overall, the Kb values position complexes 1–6, especially 1, 2, and 4, as viable drug carriers with protein, strong enough for transport and allowing free complex release for bioactivity.
3.6 In vitro antidiabetes activity
| Compounds | IC50 (mM) | |
|---|---|---|
| α-Amylase | α-Glucosidase | |
| L1 | 0.780 | 0.062 |
| L2 | 0.415 | 0.071 |
| L3 | 0.604 | 0.081 |
| 1 | 0.329 | 0.093 |
| 2 | 0.361 | 0.055 |
| 3 | 0.335 | 0.121 |
| 4 | 0.290 | 0.054 |
| 5 | 0.274 | 0.076 |
| 6 | 0.275 | 0.065 |
| Acarbose | 0.912 | 0.059 |
3.7 In vitro antioxidant activity
| Compounds | IC50 (mM) | ||
|---|---|---|---|
| NO˙ | DPPH | FRAP | |
| L1 | 0.050 | 0.087 | >5.00 |
| L2 | 0.053 | 0.261 | 4.375 |
| L3 | 0.050 | 0.531 | 3.414 |
| 1 | 0.066 | 0.627 | >5.00 |
| 2 | 0.049 | 1.222 | >5.00 |
| 3 | 0.079 | 1.323 | >5.00 |
| 4 | 0.074 | 2.788 | >5.00 |
| 5 | 0.065 | 1.001 | >5.00 |
| 6 | 0.064 | 3.951 | >5.00 |
| Quercetin | 0.060 | 0.126 | 0.034 |
4 Conclusion
In conclusion, six new copper(II) complexes derived from mixed ligands (Schiff bases and diimines) with general molecular formula [CuL1(Phen/DM-Phen)] have been synthesized and characterized by core spectroscopic techniques. The crystallography analysis of complexes 2, 4, 5 and 6 unraveled the geometry around the CuO2N3 of the complexes to be distorted square pyramidal. UV-visible absorption spectroscopy revealed the interactions of ligands L1–L3 and complexes 1–6 with ctDNA and BSA. Spectral analysis showed hypochromism with slight bathochromic shifts, suggesting an intercalation binding mode with ctDNA via classical non-covalent π-stacking. Complexes 1–6 displayed moderate-to-strong binding affinities to ctDNA (Kb = 4.73 × 104 to 2.74 × 105 M−1), better than those of the free ligands L1–L3 (Kb = 1.49 × 104 to 2.46 × 104 M−1). This positions the complexes as promising DNA-targeting agents for anticancer applications. For BSA, complexes 1–6 exhibited strong binding (Kb = 9.59 × 104 to 2.41 × 105 M−1), accompanied by hyperchromic shifts that signal microenvironment alterations and hydrophobic/π–π interactions, particularly driven by lipophilic methyl substituents (highest in complex 2). These affinities lie within optimal pharmacological ranges, facilitating effective protein-potential drug transport and release for bioactivity, especially for complexes 1, 2, and 4. All the complexes displayed good α-amylase and α-glucosidase inhibition potential with complexes 2 and 4 having lower IC50 values than the ones for acarbose in both assays, suggesting them to have potential as antidiabetic therapeutics. Compounds with methyl group substituents displayed better DNA and BSA binding affinities when compared to those with chloro and nitro groups. They also inhibit α-amylase and α-glucosidase more than their counterparts. The ligands exhibited good antioxidant activities compared to copper(II) complexes and also scavenge nitric oxide and DPPH (only L1) radical better than quercetin, the standard used for this study.Author contributions
Conceptualization, S. D. O. and R. C. L.; methodology; S. D. O., A. A. A., B. O. R. C. L. and K. A. O.; software, S. D. O., K. A. O., and A. A. A., validation, S. D. O., A. A. A., K. A. O., B. O., and R. C. L.; formal analysis; S. D. O., K. A. O., and A. A. A.; resources' R. C. L., B. O., data curation, S. D. O, A. A. A. and K. A. O.; writing original draft preparation; S. D. O., A. A. and K. A. O.; writing review and editing, S. D. O., A. A. A., K. A. O., B. O. and R. C. L., visualization, S. D. O., A. A. A., K. A. O., B. O., and R. C. L., supervision, R. C. L., project administration, R. C. L., funding acquisition, S. D. O and R. C. L.Conflicts of interest
The authors declare no conflicts of interest.Data availability
CCDC 2530540 (2), 2530541 (4), 2530542 (5) and 2530543 (6) contain the supplementary crystallographic data for this paper.63a–dAll data generated or analyzed during this study are included in this published article and its supplementary information (SI). Supplementary information (with charaterization data) is available. See DOI: https://doi.org/10.1039/d6ra01650b.
Acknowledgements
The authors acknowledge the funding from the National Research Foundation (grant number: CPRR23041794158), Stellenbosch University, Matieland, 7602, South Africa. The first author is a recipient of a postdoctoral fellowship award of the NRF at Stellenbosch University, Matieland, South Africa.References
- A. A. Ali, M. Shebl, O. M. Adly, M. S. Mahfouz and R. Fouad, J. Mol. Struct., 1359, 206, 145313 Search PubMed.
- S. D. Oladipo, B. Omondi and C. Mocktar, Polyhedron, 2019, 170, 712–722 CrossRef CAS.
- L. Esmaeili, M. G. Perez, M. Jafari, J. Paquin, P. Ispas-Szabo, V. Pop, M. Andruh, J. Byers and M. A. Mateescu, J. Inorg. Biochem., 2019, 192, 87–97 CrossRef CAS PubMed.
- I. Bento, C. Peixoto, V. N. Zaitsev and P. F. Lindley, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2007, 63, 240–248 CrossRef CAS PubMed.
- F. Tisato, C. Marzano, M. Porchia, M. Pellei and C. Santini, Med. Res. Rev., 2010, 30, 708–749 CrossRef CAS PubMed.
- X. Wu, Y. Zhou, Z. Lu, Y. Zhang, T. Zhang and Q. Jiang, Anim. Biosci., 2024, 38, 530 CrossRef PubMed.
- M. Azeem, K. Li, Y. Qin, L. Dong and W. Li, CrystEngComm, 2021, 23, 1815–1820 RSC.
- I. Iakovidis, I. Delimaris and S. M. Piperakis, Mol. Biol. Int., 2011, 2011, 594529 CrossRef PubMed.
- S. D. Oladipo, R. C. Luckay, K. A. Olofinsan, A. A. Badeji and S. Mokoena, Inorg. Chim. Acta, 2025, 575, 122447 CrossRef CAS.
- T. L. Yusuf, I. Waziri, S. D. Oladipo, M. S. Abd El-Maksoud, A. J. Muller and B. Vatsha, ACS Omega, 2025, 10, 50795–50805 CAS.
- S. D. Oladipo, R. C. Luckay and K. A. Olofinsan, Results Chem., 2025, 16, 102471 CrossRef CAS.
- S. Thakur, A. Jaryal and A. Bhalla, Results Chem., 2024, 7, 101350 CrossRef CAS.
- M. E. Moreno-Narváez, L. González-Sebastián, R. Colorado-Peralta, V. Reyes-Márquez, L. O. Franco-Sandoval, A. Romo-Pérez, J. A. Cruz-Navarro, I. V. Mañozca-Dosman, A. Aragón-Muriel and D. Morales-Morales, Inorganics, 2025, 13, 38 CrossRef.
- R. D. Mohan, S. Sarukrishna, A. S. Babu, D. Santhosh, S. A. Fathima and B. Parvathy, Inorg. Chim. Acta, 2026, 550, 122980 CrossRef.
- A. Gil, M. Melle-Franco, V. Branchadell and M. J. Calhorda, J. Chem. Theor. Comput., 2015, 11, 2714–2728 CrossRef CAS PubMed.
- M. A. Alossaimi, T. Aldakhil, H. Elmansi, F. Belal and G. Magdy, RSC Adv., 2026, 16, 5437–5454 RSC.
- X.-B. Fu, D.-D. Liu, Y. Lin, W. Hu, Z.-W. Mao and X.-Y. Le, Dalton Trans., 2014, 43, 8721–8737 RSC.
- J. C. García-Ramos, R. Galindo-Murillo, F. Cortés-Guzmán and L. Ruiz-Azuara, J. Mex. Chem. Soc., 2013, 57, 245–259 Search PubMed.
- D. Ray, A. Manna, M. F. Ansari, F. Arjmand, Z. Jaglicic and R. Herchel, Dalton Trans., 2026, 55, 3750 RSC.
- R. Ghanta, S. Jhulki, T. Chowdhury, P. Giri, R. Mondal, S. Kundu, S. Dasgupta, U. K. Das, M. Jana and A. Pal, Appl. Organomet. Chem., 2026, 40, e70511 CrossRef CAS.
- S. V. Saravanaselvam, X. Arulanantham, M. Tamilelakkiya, R. Sakthivadivel, S. Ponnusamy, P. Pandi, P. Velusamy, S. E. Muthu and K. A. Kumar, J. Mol. Struct., 2025, 1325, 141000 CrossRef CAS.
- M. Lištiak, J. Valentová, E. Zahradníková, R. Varga, E. Samoľová, M. Litecká, K. Lušpai, M. Puchoňová, M. Malček Šimunková and J. Moncoľ, Eur. J. Inorg. Chem., 2025, 28, e202500143 CrossRef.
- V. Vinayagam, D. Chandren, P. Arthur, S. Theerthagiri and W. Yong, Inorg. Chem. Commun., 2024, 161, 112116 CrossRef CAS.
- M. Maity, U. Pramanik, V. R. Hathwar, P. Brandao, S. Mukherjee, S. Maity, R. Maity, T. Maity and B. C. Samanta, Heliyon, 2022, 8, e11345 CrossRef CAS PubMed.
- N. Yasumatsu, Y. Yoshikawa, Y. Adachi and H. Sakurai, Bioorg. Med. Chem., 2007, 15, 4917–4922 CrossRef CAS PubMed.
- S. D. Oladipo and R. C. Luckay, New J. Chem., 2024, 48, 13276–13288 RSC.
- S. S. Lakshmi, K. Geetha, M. Gayathri and G. Shanmugam, J. Chem. Sci., 2016, 128, 1095–1102 CrossRef CAS.
- T. S. Chaithanya and V. Sabareesh, Pept. Sci., 2026, 118, e70017 CrossRef CAS.
- A. Reuser and H. Wisselaar, Eur. J. Clin. Invest., 1994, 24, 19–24 CrossRef CAS PubMed.
- S. R. Pathak, K. R. Senwar and K. N. Sharma, in Diabetes Mellitus, Elsevier, 2025, pp. 63–78 Search PubMed.
- M. Sabu and R. Kuttan, J. Ethnopharmacol., 2002, 81, 155–160 CrossRef CAS PubMed.
- P. Thakur, A. Kumar and A. Kumar, J. Drug Targeting, 2018, 26, 766–776 CrossRef CAS PubMed.
- A. Ceriello, Diabetes Care, 2003, 26, 1589–1596 CrossRef CAS PubMed.
- M. Valko, C. Rhodes, J. Moncol, M. Izakovic and M. Mazur, Chem.-Biol. Interact., 2006, 160, 1–40 CrossRef CAS PubMed.
- T. Dhanya, M. P. Kurup, K. Rajimon, G. A. Krishna, J. K. Varughese, K. Raghu, S. Philip, K. Divya, M. Augustine and P. Mohanan, Dalton Trans., 2025, 54, 3216–3234 RSC.
- S. D. Oladipo, R. C. Luckay, A. A. Badeji, V. A. Obakachi, K. K. Govender and A. D. Osinubi, Chem. Biodiversity, 2026, 23, e02739 CrossRef CAS PubMed.
- T. Kiran, V. G. Prasanth, M. Balamurali, C. Vasavi, P. Munusami, K. I. Sathiyanarayanan and M. Pathak, Inorg. Chim. Acta, 2015, 433, 26–34 CrossRef CAS.
- APEX4, SAINT, and SADABS, Bruker AXS Inc., Madison, WI, 2022 Search PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Adv., 2015, 71, 3–8 CrossRef PubMed.
- L. J. Barbour, J. Appl. Crystallogr., 2020, 53, 1141–1146 CrossRef CAS.
- C. F. Macrae, I. Sovago, S. J. Cottrell, P. T. Galek, P. McCabe, E. Pidcock, M. Platings, G. P. Shields, J. S. Stevens and M. Towler, J. Appl. Crystallogr., 2020, 53, 226–235 CrossRef CAS PubMed.
- A. A. Adeleke, Md. S. Islam, K. Olofinsan, V. F. Salau, C. Mocktar and B. Omondi, New J. Chem., 2021, 45, 17827–17846 RSC.
- T. L. Yusuf, D. C. Akintayo, S. D. Oladipo, A. A. Adeleke, K. Olofinsan, B. Vatsha and N. Mabuba, New J. Chem., 2022, 46, 12968–12980 RSC.
- S. J. Zamisa, A. A. Adeleke, N. Devnarain, M. A. Rhman, P. M. O. Owira and B. Omondi, RSC Adv., 2023, 13, 21820–21837 RSC.
- O. Ibitoye, K. Olofinsan, K. Teralı, U. Ghali and T. Ajiboye, J. Food Biochem., 2018, 42, e12627 CrossRef.
- K. A. Olofinsan, O. L. Erukainure, N. Z. Msomi and M. S. Islam, Asian Pac. J. Trop. Biomed., 2022, 12, 300–311 CrossRef CAS.
- B. K. Beseni, K. A. Olofinsan, V. F. Salau, O. L. Erukainure and M. S. Islam, Asian Pac. J. Trop. Biomed., 2022, 12, 453–465 CrossRef CAS.
- P. Changlian, C. Shaowei, L. Zhifang and L. Guizhu, Europe PMC, 2000, 27, 658–661 Search PubMed.
- M. K. Koley, S. U. Parsekar, N. Duraipandy, M. S. Kiran, B. Varghese, P. T. Manoharan and A. P. Koley, Inorg. Chim. Acta, 2018, 478, 211–221 CrossRef CAS.
- W. A. Munzeiwa, S. D. Oladipo, C. U. Ibeji, C. Mocktar and B. Omondi, J. Inorg. Biochem., 2021, 225, 111600 CrossRef CAS PubMed.
- A. W. Addison, T. N. Rao, J. Reedijk, J. van Rijn and G. C. Verschoor, J. Chem. Soc., Dalton Trans., 1984, 1349–1356 RSC.
- B. Mohan and M. Choudhary, J. Mol. Struct., 2021, 1246, 131246 CrossRef CAS PubMed.
- K. Paliwal, P. Haldar, P. K. S. Antharjanam and M. Kumar, ACS Omega, 2022, 7, 21961–21977 CrossRef CAS PubMed.
- L. Li, Q. Guo, J. Dong, T. Xu and J. Li, J. Photochem. Photobiol., B, 2013, 125, 56–62 CrossRef CAS PubMed.
- S. Mali, S. D. Vatagude, M. Horitani and P. M. Gurubasavaraj, J. Mol. Struct., 2026, 1360, 145599 CrossRef CAS.
- S. K. Tarai, S. Mandal, A. Tarai, I. Som, A. Pan, A. Bagchi, A. Biswas and S. C. Moi, Appl. Organomet. Chem., 2023, 37, e7164 CrossRef CAS.
- H. O. Omoregie, T. L. Yusuf, S. D. Oladipo, K. A. Olofinsan, M. B. Kassim and S. Yousuf, Polyhedron, 2022, 217, 115738 CrossRef CAS.
- A. d. S. Portas, D. C. Miguel, J. K. Yokoyama-Yasunaka, S. R. B. Uliana and B. P. Espósito, J. Biol. Inorg. Chem., 2012, 17, 107–112 CrossRef CAS PubMed.
- S. D. Oladipo, R. C. Luckay, K. A. Olofinsan, V. A. Obakachi, S. J. Zamisa, A. A. Adeleke, A. A. Badeji, S. A. Ogundare and B. P. George, Heliyon, 2024, 10, e23174 CrossRef CAS PubMed.
- U. Rao, B. A. Ahmad and K. S. Mohd, Malays. J. Anal. Sci., 2016, 20, 1191–1202 CrossRef.
- E. Rodríguez-Arce and M. Saldías, Biomed. Pharmacother., 2021, 143, 112236 CrossRef PubMed.
- İ. Gulcin and S. H. Alwasel, Processes, 2025, 13, 1296 CrossRef.
- (a) CCDC 2530540: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qy7b8; (b) CCDC 2530541: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qy7c9; (c) CCDC 2530542: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qy7db; (d) CCDC 2530543: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qy7fc.
| This journal is © The Royal Society of Chemistry 2026 |








