Open Access Article
Open Access ArticleExploring an azo-uracil based nickel(II) complex for anticancer and phosphatase like activities
Subhasis Ghosha,
Samrat Daripab,
Tandrim Shyama,
Sumit Kumar Hira *b and
Debasis Das
*b and
Debasis Das *a
*a
aDepartment of Chemistry, The University of Burdwan, Golapbag, Burdwan, India. E-mail: ddas100in@yahoo.com; subhasisg456@gmail.com; tandrimshyam@gmail.com
bCellular Immunology Laboratory, Department of Zoology, The University of Burdwan, Burdwan, 713104, W.B., India. E-mail: sdaripa40@gmail.com; sumit.hira2008@gmail.com
First published on 2nd April 2026
Abstract
An azo-uracil based Ni2+ complex (N1) has been synthesized and structurally characterized by spectroscopic techniques, namely FTIR, NMR, ESI-MS and, authenticated by single-crystal X-ray diffraction analyses as a triclinic mononuclear system (CCDC 2003428). The N1 exhibits concentration-dependent cytotoxicity against murine colon carcinoma cell lines (CT26, MC-38) and human colorectal cancer cells (HCT-15) while displaying minimal toxicity toward normal fibroblast NIH-3T3 cells indicating good selectivity. Flow cytometric and fluorescence analyses reveal significant induction of apoptosis in CT26 cells, at sub-micromolar concentration of N1, as evidenced by Annexin V-FITC/PI staining and a dose-dependent increase in early and late apoptotic populations. Mechanistic investigations indicate that N1 induces ROS-mediated, caspase-dependent apoptosis, supported by enhanced intracellular ROS generation and attenuation of cytotoxicity upon ROS scavenging and caspase inhibition. The N1 also interacts efficiently with ct-DNA, exhibiting a binding constant, 2.168 × 105 M−1. In addition, the N1 demonstrates phosphatase-like catalytic activity, by promoting the hydrolysis of p-nitrophenyl phosphate (p-NPP) under pseudo-first-order conditions with an apparent binding constant (Kapp), 1.18 × 105 M−1. Density functional theoretical (DFT) studies at the TD-SCF/B3LYP/3-21G level support the proposed N1–p-NPP interaction and provide insights into the optimized geometries and associated electronic (HOMO–LUMO) properties. This work bridges catalytic chemistry and cancer biology, positioning the azo-uracil derived Ni(II) complex as an emerging multifunctional therapeutic candidate.
Introduction
Development of anticancer drugs1–3 that selectively act on particular cancer cells is challenging and hence being exhaustively studied. Many anticancer drugs impart toxicity towards normal human cells, resulting in various side effects.The safety and utility of metal-based drugs depend upon ligand design that governs interactions with biomolecular targets, controls the release of metal ions, and offers steric protection to the metal core. Although Ni2+ is essential at trace level, unbound Ni2+ have harmful effects on biological systems. However, rational ligand coordination stabilizes the metal center under physiological settings, enhances biocompatibility, controls lipophilicity, and target selectivity, restricts cancer cell proliferation,4–6 lowers the availability of free Ni2+, regulates the interaction with biomolecules and, drastically modifies its toxicity.7,8
Recent computational and experimental research indicate that suitably designed Ni2+ complexes may impart desired biological activity9 while displaying acceptable toxicity profile. Biological investigations, in association with theoretical ADMET calculations reveal that Ni2+ complexes may exhibit better pharmacokinetic behavior over corresponding uncoordinated species.10–14
Thus, there exists ample space to contribute towards the application of new Ni2+ complexes as potential antibacterial/anticancer agents15–17 with regulated toxicity.18 The application of nickel complexes as metallo-drugs and therapeutic agents remains relatively underexplored.19–23 Relatively inexpensive nature of nickel complexes provides additional dimension and inspiration in this context.
On the other hand, phosphate metabolism24,25 plays an important role in humans. Appropriate phosphate level is essential in human as phosphorus plays vital role towards formation of bones and teeth. Moreover, hydrolysis of ATP provides energy for cellular processes. Hydrolysis of phosphodiesters26,27 is catalyzed by phosphatase enzymes. Understanding the mechanism of such bond-cleavage reactions, both enzymatic and non-enzymatic have long been an interesting research area.
Interestingly, hydrolysis of phosphodiesters is also a key focus in various industrial processes. In humans, energy transfer and phosphorylation28 are closely related to information storage and signal conduction. Due to extreme low reactivity, studies on phosphate monoester dianions are challenging. Hence, aryl phosphate monoester dianion with excellent leaving group, specially, p-nitrophenyl phosphate (p-NPP)29 have been widely used to investigate the reaction mechanism.
Complexes containing bio-relevant ligands such as uracil derivatives have demonstrated enhanced pharmacological profiles as they interact with nucleic acids and enzymes, involved in nucleotide metabolism.30
The background to choose the uracil derivative as a ligand for preparation of new Ni2+ complex has a reasoning. Uracil, a natural pyrimidine base, forms the structural core of several chemotherapeutic agents31 and participates in essential biological processes such as DNA/RNA synthesis, base-pair fidelity, and cellular energy metabolism. Chemical modification of uracil through azo-linkage and its coordination to Ni2+ may significantly alter its electronic and steric environment, leading to improved membrane permeability, DNA-binding ability32 and enzyme-like catalytic potential. As nitrogen and oxygen donors readily coordinate to Ni2+, the azo-uracil derivative33 readily forms the new Ni2+ complex (N1) as planned. Moreover, as per our speculation that the new Ni2+ complex (N1) may interact with phosphate and nucleobase-rich regions34 of bio-macromolecules, and thereby disrupt key proliferative35 and metabolic pathways of cancer cells.
To evaluate this hypothesis, the anticancer potential of the N1 is tested against murine and human colon carcinoma cell lines (CT-26, MC-38, and HCT-15), along with normal fibroblast NIH-3T3 cells to assess cytocompatibility.36 The mechanistic basis of cytotoxicity is further examined using Annexin V-FITC and propidium iodide (PI) staining, followed by fluorescence microscopy and flow cytometry37 to quantitatively assess apoptosis.
In addition to its anti-proliferative properties,38 this study also explores the enzyme-mimetic behavior of the N1, specifically towards phosphate metabolism. The ability of N1 to catalyze hydrolysis of p-nitrophenyl phosphate (p-NPP) has been investigated to assess its phosphatase-like activity.39,40 Thus, investigation of dual functionality, viz. anticancer and enzyme-mimetic properties of N1 may unveil some valuable insights about its therapeutic41 and enzymatic potential.42
Experimental
Materials
All chemicals and reagents were of analytical grade and used without further purification unless otherwise stated. Nickel(II) nitrate hexahydrate (Ni(NO3)2·6H2O), 1,3-dimethyl-6-aminouracil, p-aminobenzoic acid, p-nitrophenyl phosphate (p-NPP), N-acetyl cysteine (NAC), Z-VAD-FMK, and PhosSTOP™ phosphatase inhibitor cocktail were purchased from Sigma-Aldrich (USA). Organic solvents including methanol and acetonitrile were of spectroscopic grade.Cell culture reagents, including Dulbecco's Modified Eagle Medium (DMEM), fetal bovine serum (FBS), penicillin–streptomycin, and trypsin-EDTA, were obtained from Gibco (Thermo Fisher Scientific, USA). The murine colon carcinoma cell lines CT26, human colorectal cancer cell line HCT-15, and normal murine fibroblast cell line NIH-3T3 were procured from ATCC (USA).
For biological assays, Annexin V-FITC/propidium iodide (PI) apoptosis detection kit, CellROX™ Green reagent, MitoSOX™ Red mitochondrial superoxide indicator, JC-1 mitochondrial membrane potential probe, and LDH cytotoxicity assay kit were obtained from Invitrogen (USA) and used according to the manufacturer's instructions.
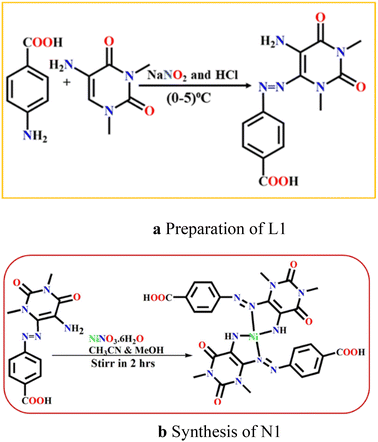
Synthesis of azauracil derivative (Scheme 1a)43
1,3-Dimethyl-6-aminouracil (224.6 mg, 1.458 mmol) and p-amino benzoic acid (200 mg, 1.458 mmol) are used to prepare the azo dye (L1) following conventional diazotization procedure (Scheme 1a).Preparation of Ni(II) complex (Scheme 1b)
The solutions of Ni(NO3)2·6H2O (40 mg, 0.138 mmol) and L1 (83.6 mg, 0.276 mmol) in acetonitrile–methanol mixture (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) was stirred for 2 h. Slow evaporation of the solvent resulted red colour X-ray diffraction quality single crystals of N1 in 72% yield. The N1 was also characterized by mass (Fig. S1, SI), FTIR (Fig. S2, SI) and NMR (Fig. S3, SI) spectra.
1, v/v) was stirred for 2 h. Slow evaporation of the solvent resulted red colour X-ray diffraction quality single crystals of N1 in 72% yield. The N1 was also characterized by mass (Fig. S1, SI), FTIR (Fig. S2, SI) and NMR (Fig. S3, SI) spectra.
Characterization of L1 and N1
Anal. calcd. for N1, C26 H24 N10 NiO8, 14 (H2O) (%): C, 49.17; H, 4.70 and N, 19.77; found: C, 49.19; H, 4.8 and N, 19.87. The absorption spectra of L1 and N1 has main characteristics peaks at 517 nm and 600 nm respectively (Fig. 1). Since, L1 is previously reported,43 some of its spectra are presented as supporting information that may be useful to note the changes after complexation with Ni2+. Hence, 1H NMR of L1 is excluded here. The FTIR spectra of N1 has characteristics ν(Ni–N), 544 cm−1; ν(N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1395; ν(sp3 C–H), 2975, ν(–C
N), 1395; ν(sp3 C–H), 2975, ν(–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) 1708, 1654, ν(–C–N), 1370, ν(–C
O) 1708, 1654, ν(–C–N), 1370, ν(–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 1510, ν(–C–N) 1299 (Fig. S2, SI).
C), 1510, ν(–C–N) 1299 (Fig. S2, SI).
QTOF-MS ES(+) (Fig. S1, SI): [N1 + Li + 3H]+ 718.79. 1H NMR of N1 (400 MHz, DMSOd6, δ, ppm) (Fig. S3, SI): 7.00–8.00 (d, 2H, aromatic H), 7.71–7.69 (d, 2H, aromatic H), 3.42–3.24 (m, 6H, N–CH3). 13C NMR spectra of L1 and N1 are presented in Fig. S4 and S5 (SI) respectively.
Cell lines and culture conditions
Murine colon carcinoma cell lines CT26 and MC-38, human colorectal carcinoma cell line HCT-15, and normal murine fibroblast cell line NIH-3T3 were procured from the ATCC, USA. All cell lines were cultured in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS), 1% penicillin–streptomycin, and maintained in a humidified incubator at 37 °C with 5% CO2 atmosphere. Cells were sub-cultured every 2–3 days using 0.25% trypsin-EDTA and used during the logarithmic growth phase for experiments.44Cytotoxicity assay (MTT assay)
The cytotoxic effect of the Ni2+ complex (N1) was evaluated by standard MTT assay. Briefly, cells were seeded at a density of 10 × 103 cells per well in 96-well plates and allowed to adhere for 24 h. Cells were then treated with increasing concentrations of N1 (ranging from 0.01 µg mL−1 to 1 µg mL−1) for 48 h. After incubation, 20 µL of MTT reagent (5 mg mL−1) was added to each well and further incubated for 4 h. The resulting formazan crystals were dissolved in 150 µL DMSO, and absorbance was measured at 570 nm using a microplate reader (Bio-Tek, USA). Cell viability was expressed as a percentage relative to untreated control, and the IC50 values for each cell line (CT26, MC-38, HCT-15, and NIH-3T3) were calculated by nonlinear regression using GraphPad Prism 10 software.45Apoptosis detection by Annexin V-FITC and propidium iodide (PI) staining
To determine the apoptotic potential of the N1, CT26 cells were seeded in six-well plates (2 × 105 cells per well) and treated with 0.04 µg mL−1 and 0.08 µg mL−1 of N1 for 12 h. Doxorubicin (0.25 µM) was used as a positive control. Following treatment, cells were harvested, washed twice with cold PBS, and resuspended in 1× Annexin V binding buffer. Subsequently, 5 µL Annexin V-FITC and 10 µL PI (50 µg mL−1) were added, followed by 15 min incubation at room temperature at dark. Samples were analyzed using a BD C6accuri flow cytometer, and data were processed using FlowJo v10.8 software. The distribution of viable, early apoptotic, late apoptotic, and necrotic cell populations was quantified. Representative images were captured by fluorescence microscopy (Leica DMi8, 200×) to visualize Annexin-FITC and PI-positive cell.46,47Intracellular ROS measurement
CT26 cells were seeded in 24-well plates (2 × 105 cells per well) and treated with N1 (0.04 and 0.08 µg mL−1) for 12 h. Cells were then incubated with CellROX™ Green (5 µM) for total ROS and MitoSOX™ Red (5 µM) for mitochondrial superoxide at 37 °C for 30 min in the dark. After washing with PBS, fluorescence was analysed by flow cytometry (BD Accuri C6), and data were processed using FlowJo software.Mitochondrial membrane potential (ΔΨm)
Changes in mitochondrial membrane potential were assessed using JC-1 dye. Following treatment with N1 for 12 h, cells were incubated with JC-1 (5 µg mL−1) at 37 °C for 20 min, washed with PBS, and analysed by flow cytometry. The ratio of red (aggregates) to green (monomers) fluorescence was used to determine ΔΨm.LDH release assay and ROS scavenging
Cellular cytotoxicity was evaluated using an LDH release assay in CT26 and HCT-15 cells. Cells were pre-treated with N-acetyl cysteine (NAC, 5 mM, 1 h) followed by N1 treatment for 12 h. LDH activity in the culture supernatant was measured according to the manufacturer's protocol, and cytotoxicity was expressed relative to untreated controls.Phosphatase inhibition assay
To assess the role of phosphatase-related signaling, cells are pre-treated with PhosSTOP™ phosphatase inhibitor cocktail (1 × 1 h) prior to N1 exposure. Cytotoxicity is subsequently evaluated using LDH assay as described above.Caspase inhibition assay
For apoptosis validation, CT26 cells were pre-treated with the pan-caspase inhibitor Z-VAD-FMK (20 µM, 1 h) followed by N1 treatment for 12 h. Apoptosis was quantified by Annexin V-FITC/PI staining and analysed by flow cytometry.Statistical analysis
All experiments have been performed in triplicate and data expressed as mean ± standard deviation (SD). Statistical comparisons are carried out using one-way ANOVA followed by Tukey's multiple comparison test in GraphPad Prism 9. A p-value < 0.05 is considered statistically significant.Results and discussion
Synthesis and structure
Conventional diazotization process was followed for preparation of uracil based azo dye, namely 1,3-dimethyl-5-arylazo-6-amino uracil. For this, the diazotized amine was subjected to coupling with aqueous solution of 1,3-dimethyl-5-amino uracil followed by neutralization with NaHCO3 solution to obtain the red solid azo dye. To understand the structure of dye, the spectroscopic data like UV-vis, fluorescence, FTIR, ESI-MS and 1H NMR spectra were recorded. In addition, the structure of L1 and N1 were authenticated by single crystal X-ray diffraction analysis which indicate existence of three tautomeric forms of the L1, namely azo-enamine-keto (HLAZO), hydrazone-amine-keto (HLhydrazo) and azo-imine-enol (HLazo-enol).Single crystal X-ray diffraction analysis of N1
The stereo-chemical structure of N1 is confirmed by single crystal X-ray diffraction analysis. The molecular stereochemical view is shown in Fig. 2 while its packing pattern is shown in Fig. S6 (SI). The crystallographic data and refinement parameters are presented in Table S1 (SI). Selected bond lengths and angles are listed in Table S2 (SI). N1 is triclinic having space group P![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) (CCDC no. 2003428): a, 7.89(2) Å; b, 8.49(2) Å; c, 15.86(4) Å; α (°), 80.77(4); β (°), 79.09(5); γ(°), 89.38(4); volume (Å3), 103 0(4); Z = 1. The bond lengths Ni2–N5, 1.828(12) Å and Ni2–N4, 1.855(14) Å indicate single bond character of Ni–N whereas N3–N4, 1.329(16) Å indicate double bond character of two N–N bonds azo group. The bond lengths N3–C5, 1.314(18) Å and C12–N4, 1.484(17) Å indicate single bond character of C–N, adjacent to Ni–N bond. The bond angles C5 N3 N4 120.0 (14)° and C12 N4 Ni2 122.4 (9)° indicate sp2 nature of the N3 and N4 centres.
(CCDC no. 2003428): a, 7.89(2) Å; b, 8.49(2) Å; c, 15.86(4) Å; α (°), 80.77(4); β (°), 79.09(5); γ(°), 89.38(4); volume (Å3), 103 0(4); Z = 1. The bond lengths Ni2–N5, 1.828(12) Å and Ni2–N4, 1.855(14) Å indicate single bond character of Ni–N whereas N3–N4, 1.329(16) Å indicate double bond character of two N–N bonds azo group. The bond lengths N3–C5, 1.314(18) Å and C12–N4, 1.484(17) Å indicate single bond character of C–N, adjacent to Ni–N bond. The bond angles C5 N3 N4 120.0 (14)° and C12 N4 Ni2 122.4 (9)° indicate sp2 nature of the N3 and N4 centres.
1H NMR spectral studies
The 1H NMR spectra reflects the mode of binding of L1 with Ni2+. Upon addition of Ni(NO3)2·6H2O (1.0 equiv.) to 2.0 equiv. of L1, the –NH protons of the imine moiety (c, 11.280 ppm) and benzoic acid (d, 12.02 ppm) moiety have been observed. The phenyl ring protons (d, 7.731 ppm), close to the coordination sphere shifted downfield to 7.742 ppm (b). On the other hand, the amine proton (3.0657 ppm) disappears with the appearance of imine proton (c, 11.280 ppm) (Fig. S3, SI). 13C NMR spectra of L1 and N1 (Fig. S4 and S5, SI) support their formation and the peak positions are as usual as expected and no anomalous peaks have been observed.Density functional theoretical (DFT) studies
DFT studies have been performed using the TDSCF/DFT/B3LYP/3-21G level of theory. For L1, the HOMO–LUMO energy gap is 0.08063 eV (Table S3, SI and Fig. 3). The S0/S3 transition (HOMO to LUMO, HOMO−1 to LUMO, HOMO−2 to LUMO, HOMO to LUMO+1, HOMO to LUMO+9) at 690.46 nm has maximum value of fb, 0.0198, that indicates higher possibility of intra-molecular charge transfer (ICT) in the visible region over other transitions. The LUMO of N1 is more stable than the LUMO of L1 but the HOMO of N1 is less destabilized than the HOMO of L1, which indicates that N1 is more stable than L1. The S0/S1 transition at 552.66 nm of L1 justifies its yellow colour. Other possible transitions associated with N1 and L1 are furnished in the Tables S4–S6 (SI).Absorption spectroscopic studies on the interaction between N1 and ct-DNA
The binding interaction of the N1 with DNA is monitored by UV-vis spectroscopy. The concentration of the N1 is kept unaltered (6 µM in DMSO) while that of DNA is varied from 1–20 µM. The ionic strength of the media is maintained using NaCl (60 mM) while the pH is maintained at 7.2 using HEPES buffer (10 mM). It is observed that the absorbance at 425 nm increases with increasing concentration of ct-DNA. After mixing ct-DNA with N1, the temperature is maintained in between 0–5 °C for 15 minutes, followed by the absorbance is measured. The binding parameters have been evaluated considering the following equilibrium: N1 + DNA ⇌ N1-DNA where Kd = [N1-DNA]/[N1] [DNA] where [N1], [DNA] and [N1-DNA] are equilibrium concentrations of N1, DNA and N1-DNA adduct respectively, using the equation, 1/A = 1/ΔAmax + Kd/ΔAmax (CD − CL) where ΔA and ΔAmax are changes in absorbance with increasing DNA concentration and maximum change of absorbance respectively. The CD and CL are concentrations of DNA and N1 respectively.The plot of (CD − CL)−1 vs. (ΔA)−1 is linear and allows to determine Kd, the inverse of the apparent binding constant (Kapp) and ΔAmax. The (ΔA/ΔAmax)−1 vs. (CD)−1 plot gives the linear fit analysis that generates, Kapp as 2.168 × 105 M−1 (Fig. 4).
Although, it is inconclusive whether the interaction follows purely electrostatic interaction/groove binding or traditional intercalation. However, literature indicates that increase in absorbance (hyperchromism) upon gradual addition of ct-DNA to the N1, probably due to electrostatic interaction/groove binding rather than traditional intercalation (Fig. 5). The hyperchromism and binding constant48 hints to groove binding49 with partial intercalation.50 The fluorescence competitive experiment also supports the binding interaction as detailed below.
Competitive binding fluorescence measurements
The interaction between N1 and ct-DNA is monitored using competitive binding fluorescence measurements.51 Ethidium bromide (EB) bound ct-DNA system is treated with the solution of N1 in DMSO to assess the binding interaction. The pH of the media is maintained at 7.2 using 10 mM HEPES buffer, containing 60 mM NaCl at room temperature.The concentration of the ct-DNA–EB (8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) is varied between 1 to 18 µM, while that of N1 is kept unaltered at 6 µM. Before measuring fluorescence (λex, 500 nm, λem, 540 nm), the mixture is incubated for 15 minutes at room temperature to allow the reaction to reach equilibrium. Upon addition of N1, the fluorescence intensity of the system drops due to reduced concentration of the DNA–EB (8
1) is varied between 1 to 18 µM, while that of N1 is kept unaltered at 6 µM. Before measuring fluorescence (λex, 500 nm, λem, 540 nm), the mixture is incubated for 15 minutes at room temperature to allow the reaction to reach equilibrium. Upon addition of N1, the fluorescence intensity of the system drops due to reduced concentration of the DNA–EB (8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) system, indicating interaction of N1 with EB–DNA system, resulting its disruption to cause fluorescence quenching (Fig. 6).
1) system, indicating interaction of N1 with EB–DNA system, resulting its disruption to cause fluorescence quenching (Fig. 6).
From literature, it seems that the nature of interaction of N1 with ct-DNA is predominantly groove binding with a discernible contribution of partial intercalation. The DNA binding efficiency of N1 is compared with few pioneering Ni2+ complexes in Table 1.
| Ni(II) complex | Donors | DNA binding constant (Kb, M−1) | Binding mode | Ref. |
|---|---|---|---|---|
| Schiff base | N, O | 1.5 × 105 | Groove | 52 |
| [Ni(5-Cl-sal-phe)(phen)(H2O)] | N, O | 1.6 × 104 | Groove | 53 |
| [Ni(5-Cl-sal-phe)(bpy)(H2O)] | N, O | 2.30 × 104 | Groove | 53 |
| Ni(II) bis(thiosemicarbazone) | N, S | 3.50 × 105 | Intercalation | 54 |
| Azo-uracil | N, O | 2.168 × 105 | Possibly groove binding with partial intercalation | Present |
Studies on p-nitrophenyl phosphatase (PNPP) activity
The rate of phosphoester bond hydrolysis is generally slow,55 however it enhances in presence of suitable catalyst.56–62 Herein, different concentrations of p-NPP is reacted with catalytic quantity of the N1 (0.001 mmol) in MeOH–H2O (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, v/v), maintaining the pH at 7.4 using PBS buffer at room temperature. The generation of yellow p-nitrophenol as the hydrolysis product is reflected in increase of absorbance at 428 nm (Fig. 7). The rate constant, 9.11 × 10−3 s−1 is determined following initial rate method by reacting N1 (0.001 mmol) with PNPP of different concentrations and, maintaining pseudo-first-order condition (Fig. S7, SI). The value of binding constant is 0.1182 × 106 M−1 (Fig. 8).
4, v/v), maintaining the pH at 7.4 using PBS buffer at room temperature. The generation of yellow p-nitrophenol as the hydrolysis product is reflected in increase of absorbance at 428 nm (Fig. 7). The rate constant, 9.11 × 10−3 s−1 is determined following initial rate method by reacting N1 (0.001 mmol) with PNPP of different concentrations and, maintaining pseudo-first-order condition (Fig. S7, SI). The value of binding constant is 0.1182 × 106 M−1 (Fig. 8).
The hydrolysis reaction is further supported by the ESI-MS spectrum, where the peak at m/z, 139.0932 (calcd. m/z 139.11; Fig. S8, SI) indicates the formation of p-nitrophenol.
To explore the mode of interaction between N1 and p-NPP and to unlock the associated energy parameters that facilitate the interaction, density functional theoretical (DFT) studies have been performed. The optimized structures at the initial (step I) and final (step F) stages, along with the associated orbital (HOMO and LUMO) energies for the interaction between the N1 and p-NPP are presented in Fig. 9. Relevant interaction parameters are presented in detail in Table S7 (SI).
 | ||
| Fig. 9 Optimized structures and HOMO–LUMO energies of the initial (Step I) and final (Step F) stages of interaction between N1 and p-NPP. | ||
Cytotoxic activity of the N1
The N1 is first screened for anti-proliferative potential against two murine and one human colon carcinoma cell lines, viz. CT26, MC-38 and HCT-15 by MTT assay. Fig. 10A reveals that N1 exhibits dose-dependent inhibition of cancer cell proliferation, indicating a strong cytostatic effect across all tested lines. The inhibitory response is most prominent in CT26 cells, followed by MC-38 and HCT-15, suggesting differential cellular susceptibility, possibly linked to intrinsic variations in their metabolic and redox environments.Importantly, when tested on normal murine fibroblast cells (NIH-3T3), the N1 shows negligible cytotoxicity even at higher concentrations (Fig. 10B), confirming its selectivity toward malignant cells and reasonable biocompatibility. The calculated half-maximal inhibitory concentrations (IC50) are ∼0.13 µg mL−1 (CT26), 0.2 µg mL−1 (MC-38) and 0.12 µg mL−1 (HCT-15) (Fig. 10C–E). The results highlight that the N1 has potential anti-cancer activity at sub-microgram concentration, competitive to several reported Ni2+ based cytotoxic agents, while it is safe towards non-tumorigenic fibroblasts.
The studies on anti-cancer activities of the N1 against two murine colon cancer cell lines (CT26 and MC-38) and one human colon cancer cell line (HCT-15) reveals a concentration-dependent inhibition of cell growth (A). In contrast, the N1 shows minimal cytotoxicity toward normal murine fibroblast cells (NIH-3T3), indicating reasonable biocompatibility (B). The half-maximal inhibitory concentration (IC50) values for CT26 (C), MC-38 (D) and HCT-15 (E) cells, mentioned in the figure are satisfactory.
N1 induced caspase-dependent apoptosis in colon cancer cells
To elucidate the underlying mechanism of the observed cytotoxicity, apoptotic analysis has been performed on CT26 cells using dual Annexin V-FITC/PI staining. As shown in Fig. 11A, a distinct increase in green (Annexin-FITC) and red (PI) fluorescence has been observed upon treatment with the N1, indicating externalization of phosphatidylserine and loss of membrane integrity characteristics of apoptotic cells.Flow cytometric quantification (Fig. 11B–E) further confirmed a dose-dependent elevation of both early and late apoptotic populations upon exposure to 0.04 µg mL−1 and 0.08 µg mL−1 of N1 for 12 h, compared to untreated controls. The apoptotic response is comparable to that induced by doxorubicin (0.25 µM), used as a positive control. The histogram of FITC fluorescence (Fig. 11F) demonstrated a rightward shift, corroborating enhanced Annexin binding, while the bar diagram (Fig. 11G) quantifies significant apoptosis induction (p < 0.01).
To validate, whether the growth inhibition induced by the N1 is due to apoptosis, CT26 cells are treated with 0.04 µg mL−1 and 0.08 µg mL−1 of the N1 for 12 h and subjected to Annexin V-FITC and propidium iodide (PI) stain. Representative fluorescence micrographs (Leica DMi8, 200×) show Annexin V-FITC- and PI-positive apoptotic cells (A). Flow-cytometric dot plots illustrate apoptotic populations in untreated control cells (B), doxorubicin-treated cells (0.25 µM, positive control) (C) and cells treated with the indicated concentrations of the N1 (D–E). Histogram shows fluorescence shifts in the FITC channel (F) and corresponding bar diagram (G) quantitatively represents the percentage of apoptotic cells.
To verify whether the observed cytotoxicity arises from the coordinated complex rather than the metal ion alone, control experiments were performed using free Ni2+ under identical conditions. As shown in Fig. S9A, free Ni2+ exhibits significantly lower cytotoxicity compared to N1 at corresponding 0.04 µg mL−1 concentrations. Consistently, Annexin V-FITC/PI analysis (Fig. S9B) reveals minimal induction of apoptosis in cells treated with free Ni2+, whereas N1 induces a pronounced increase in apoptotic cell populations. These findings confirm that the enhanced cytotoxic and pro-apoptotic effects are predominantly associated with the ligand-coordinated Ni2+ complex, highlighting the critical role of the azo-uracil framework in modulating biological activity.
To further validate the involvement of caspase signalling, cells were pre-treated with the pan-caspase inhibitor Z-VAD-FMK, which resulted in a marked reduction in Annexin V-positive cells, confirming that N1 induces caspase-dependent apoptosis (Fig. 12A and B). Collectively, these findings suggest that the N1 suppresses tumour cell growth primarily through the induction of programmed cell death rather than nonspecific necrosis. The structural design of the N1, a Ni2+ complex of an uracil based azo derivative facilitates the interactions with ct-DNA and intracellular phosphate moieties, thereby triggering intrinsic apoptotic pathways.
ROS-mediated cytotoxicity and mitochondrial involvement
Given the known redox activity of azo-metal complexes, intracellular ROS generation was evaluated using flow cytometry. Treatment with N1 led to a pronounced increase in total intracellular ROS (CellROX Green) whereas mitochondrial superoxide levels (MitoSOX Red) showed a comparatively modest increase (Fig. 12C), indicating that ROS generation is predominantly non-mitochondrial in origin. To assess mitochondrial involvement, JC-1 staining was performed. N1 treatment resulted in a moderate decrease in mitochondrial membrane potential (ΔΨm), suggesting partial mitochondrial perturbation but not extensive mitochondrial damage (Fig. 12D).The functional role of ROS in cytotoxicity is further confirmed using N-acetyl cysteine (NAC), a ROS scavenger. NAC pre-treatment significantly reduces LDH release in both CT26 and HCT-15 cells, demonstrating that ROS generation is a key driver of N1-induced cytotoxicity (Fig. 13).
Contribution of phosphatase-related processes
Considering the phosphatase-like activity of N1, its potential involvement in cytotoxicity is preliminarily examined using a phosphatase inhibitor (PhosSTOP). Inhibition of phosphatase activity results in a partial reduction in cytotoxicity in both CT26 and HCT-15 cells (Fig. 11). While this observation suggests a possible contribution of phosphate-related cellular processes, the current data are not sufficient to establish a direct causal relationship. Accordingly, this aspect has been interpreted cautiously.Biological implications and mechanistic insight
Collectively, the results demonstrate that N1 exerts selective anticancer activity primarily through ROS-mediated, caspase-dependent apoptosis. While mitochondrial depolarization is observed, it appears to play a secondary role. The biological observations may well be explained from the structural features of the uracil-azo-Ni2+ coordination framework that provides redox-active azo centre, capable of generating localized oxidative stress within tumour cells.Concurrently, the phosphatase like activity of the N1, demonstrated by its catalytic hydrolysis of p-nitrophenyl phosphate (p-NPP), may interfere with cellular phosphate signalling and nucleic acid metabolism, further enhancing its anti-proliferative efficacy. The phosphatase-like activity of N1 may contribute to the overall biological response, but its direct involvement in cytotoxicity remains to be fully established.
Thus, the multifunctional nature of the azo-uracil-based Ni2+ complex, combining redox activity with enzyme-mimetic behaviour, provides a promising framework for the development of next-generation metal-based anticancer agents. However, further detailed biochemical studies will be required to fully delineate the interplay between catalytic activity and cellular toxicity.
Conclusion
In summary, a smart uracil-derived azo based Ni2+ complex (N1), structurally characterized by spectroscopic techniques and authenticated by single-crystal X-ray diffraction analysis shows phosphatase-like catalytic behaviour, efficiently hydrolysing p-nitrophenyl phosphate (p-NPP) under mild conditions. DFT studies validate the experimental findings, optimizes the geometries of the reactants and products along with associated HOMO–LUMO energy gap, supporting the driving force behind the catalytic behaviour. DFT studies also allows to unveil the underlying mechanism of phosphatase-like catalytic activity.Biologically, N1 demonstrates potent and selective cytotoxicity against murine and human colon carcinoma cell lines (CT26, MC-38 and HCT-15), while exhibiting minimal toxicity toward normal fibroblast cells (NIH-3T3). Mechanistic investigations reveal that the cytotoxic effect is primarily mediated through ROS-dependent, caspase-mediated apoptosis, as evidenced by intracellular ROS generation, attenuation of cytotoxicity upon ROS scavenging, and reduction of apoptosis in the presence of a pan-caspase inhibitor. Moderate mitochondrial membrane depolarization suggests partial involvement of mitochondrial pathways.
Although the phosphatase-like activity of N1 may contribute to its biological effects, the current data indicate that this relationship remains indirect and requires further investigation.
Overall, the present study highlights the potential of azo-uracil-based Ni2+ complex as a multifunctional system combining catalytic activity with selective anticancer properties. These findings provide a foundation for the rational design of metal-based therapeutic agents, while emphasizing the need for further studies to fully elucidate structure–activity relationships and underlying biochemical mechanisms.
Author contributions
SG has performed synthesis, characterization and interaction studies with DNA and p-NPP while SD has performed biological studies. TS helped with MS drawings and plots. SKH and DD are supervisors.Conflicts of interest
There are no conflicts to declare.Data availability
Please note that any data related to the above mentioned manuscript are available to us and will be shared if the situation demands. Although all related data are available in supplementary information (SI), in addition to the text. Supplementary information: NMR, mass, SC-XRD, FTIR, DFT data, plot for determination of rate constant for N1 catalyzed hydrolysis of p-NPP, anti-cancer activities etc. See DOI: https://doi.org/10.1039/d6ra01587e.CCDC 2003428 (N1) contains the supplementary crystallographic data for this paper.63
Acknowledgements
The present work is partially supported by the grant from ICMR, GOI Smal grant (IIRPSG-2024-01-00993) to SKH. We are grateful to WB-DSTBT and CSIR for fellowship to SG and SD respectively. We are grateful to The University of Burdwan Instrumentation Center (USIC) for XRD facility.References
- Y. Gilad, M. Firer and G. Gellerman, Biomedicines, 2016, 4, 11 CrossRef PubMed.
- A. Wojtyla, M. Gładysz and B. Rubiś, Mol. Biol. Rep., 2011, 38, 3339–3349 CrossRef CAS PubMed.
- A. Kumar Singh, A. Kumar, H. Singh, P. Sonawane, P. Pathak, M. Grishina, J. Pal Yadav, A. Verma and P. Kumar, Chem. Biodiversity, 2023, 20, e202300061 CrossRef CAS.
- C. Zhao, X. Chen, D. Zang, X. Lan, S. Liao, C. Yang, P. Zhang, J. Wu, X. Li, N. Liu and Y. Liao, Oncogene, 2016, 35, 5916–5927 CrossRef CAS PubMed.
- M. Nejat Dehkordi and B. Åkerman, Chem.-Biol. Interact., 2018, 282, 55–62 CrossRef CAS PubMed.
- S. Diyali, A. K. Mohanty, S. Sarkar, M. F. Hossain, B. Bhowmik and B. Biswas, Adv. Chemicobiol. Res., 2021, 1, 22–30 CrossRef.
- G. Genchi, A. Carocci, G. Lauria, M. S. Sinicropi and A. Catalano, Int. J. Environ. Res. Public Health, 2020, 17, 679 CrossRef CAS PubMed.
- R. R. Kumar, M. K. M. Subarkhan and R. Ramesh, RSC Adv., 2015, 5, 46760–46773 RSC.
- N. V. Loginova, H. I. Harbatsevich, N. P. Osipovich, G. A. Ksendzova, T. V. Koval’chuk and G. I. Polozov, Curr. Med. Chem., 2020, 27, 5213–5249 CrossRef CAS PubMed.
- A. A. Aly, E. M. Abdallah, S. A. Ahmed, M. M. Rabee and S. Bräse, Molecules, 2023, 28, 1808 CrossRef CAS PubMed.
- A. Rambabu, N. Ganji, S. Daravath, K. Venkateswarlu and K. Rangan, J. Mol. Struct., 2020, 1199, 127006 CrossRef CAS.
- S. K. Singh, A. Bagul, A. Tufail, M. S. Ali, A. Ragusa, S. W. Kulkarni, S. Sharma and A. Dubey, Appl. Organomet. Chem., 2025, 39, e70286 CrossRef CAS.
- Y. Gou, X. Jia, L. X. Hou, J. G. Deng, G. J. Huang, H. W. Jiang and F. Yang, J. Med. Chem., 2022, 65, 6677–6689 CrossRef CAS PubMed.
- R. Ahmad and M. Choudhary, New J. Chem., 2025, 49, 12401–12422 RSC.
- H. M. Abd El-Lateef, M. M. Khalaf, M. Gouda, A. A. Amer, A. A. Abdelhamid and A. Abdou, Appl. Organomet. Chem., 2025, 39, e7843 CrossRef.
- G. Genchi, A. Carocci, G. Lauria, M. S. Sinicropi and A. Catalano, Int. J. Environ. Res. Public Health, 2020, 17, 679 CrossRef CAS PubMed.
- R. R. Kumar, M. K. M. Subarkhan and R. Ramesh, RSC Adv., 2015, 5, 46760–46773 RSC.
- J. Haribabu, K. Jeyalakshmi, Y. Arun, N. S. P. Bhuvanesh, P. T. Perumal and R. Karvembu, RSC Adv., 2015, 5, 46031–46049 RSC.
- D. Tanwar, T. Kaur, R. Kumar, D. Ahluwalia, D. Sharma and U. Kumar, ACS Appl. Bio Mater., 2023, 6, 134–145 CrossRef CAS PubMed.
- Y. Hoshimoto, M. Ohashi and S. Ogoshi, Acc. Chem. Res., 2015, 48, 1746–1755 CrossRef CAS PubMed.
- E. Kuah, S. Toh, J. Yee, Q. Ma and Z. Gao, Chem.–Eur. J., 2016, 22, 8404–8430 CrossRef CAS PubMed.
- J. K. Fathima, V. Lavanya, S. Jamal and N. Ahmed, Curr. Pharmacol. Rep., 2022, 8, 236–252 CrossRef.
- K. Banerjee, M. K. Biswas and S. K. Choudhuri, J. Biol. Inorg. Chem., 2017, 22, 1223–1249 CrossRef CAS PubMed.
- A. G. Cassano, V. E. Anderson and M. E. Harris, J. Am. Chem. Soc., 2002, 124, 10964–10965 CrossRef CAS PubMed.
- V. Singh, M. Ram, R. Kumar, R. Prasad, B. K. Roy and K. K. Singh, Protein J., 2017, 36, 1–6 CrossRef CAS PubMed.
- M. A. Chagas, E. S. Pereira, J. C. S. Da Silva and W. R. Rocha, J. Mol. Model., 2018, 24, 259 CrossRef PubMed.
- J. Rawlings, W. W. Cleland and A. C. Hengge, J. Inorg. Biochem., 2003, 93, 61–65 CrossRef CAS PubMed.
- S. Nath and J. Villadsen, Biotechnol. Bioeng., 2015, 112, 429–437 CrossRef CAS PubMed.
- X.-Y. Yi, T. C. H. Lam, I. D. Williams and W.-H. Leung, Inorg. Chem., 2010, 49, 2232–2238 CrossRef CAS PubMed.
- S. M. Hacker, A. Buntz, A. Zumbusch and A. Marx, ACS Chem. Biol., 2015, 10, 2544–2552 CrossRef CAS.
- J. K. Fathima, V. Lavanya, S. Jamal and N. Ahmed, Curr. Pharmacol. Rep., 2022, 8, 236–252 CrossRef.
- G. Barone, A. Terenzi, A. Lauria, A. M. Almerico, J. M. Leal, N. Busto and B. García, J. Inorg. Biochem., 2013, 257, 2848–2862 CAS.
- V. Yadav, R. Kumar, Rishu and V. K. Bhardwaj, J. Chem. Sci., 2024, 136, 9 CrossRef CAS.
- Z. -J. Yang, M.-G. Concilio, V. Ramesh and L.-H. Zhang, Epitranscriptomics: Methods and Protocols, 2018, pp. 151–163 Search PubMed.
- J. Zhu and C. B. Thompson, Nat. Rev. Mol. Cell Biol., 2019, 20, 436–450 CrossRef CAS PubMed.
- M. Mikulewicz and K. Chojnacka, Biol. Trace Elem. Res., 2011, 142, 865–889 CrossRef CAS PubMed.
- A. Adan, G. Alizada, Y. Kiraz, Y. Baran and A. Nalbant, Crit. Rev. Biotechnol., 2017, 37, 163–176 CrossRef CAS PubMed.
- H. Regassa, A. Sourirajan, V. Kumar, S. Pandey, D. Kumar and K. Dev, Cancers, 2022, 14, 3898 CrossRef CAS PubMed.
- N. Dutta, S. Haldar, G. Vijaykumar, S. Paul, A. P. Chattopadhyay, L. Carrella and M. Bera, Inorg. Chem., 2018, 57, 10802–10820 CrossRef CAS PubMed.
- J. W. Mao, H. Zhou, Y. F. Chen, G. Z. Cheng and Z. Q. Pan, Transition Met. Chem., 2012, 37, 385–391 CrossRef CAS.
- T. Chattopadhyay, M. Mukherjee, A. Mondal, P. Maiti, A. Banerjee, K. S. Banu, S. Bhattacharya, B. Roy, D. J. Chattopadhyay, T. K. Mondal, M. Nethaji, E. Zangrando and D. Das, Inorg. Chem., 2010, 49, 3121–3129 CrossRef CAS PubMed.
- D.-L. Ma, C. Wu, G. Li, T.-L. Yung and C.-H. Leung, J. Mater. Chem. B, 2020, 8, 4715–4725 RSC.
- M. Banerjee, M. Ghosh, S. Pradhan, J. S. Matalobos, A. Rej, S. K. Hira and D. Das, ACS Appl. Bio Mater., 2019, 2, 1184–1196 CrossRef CAS PubMed.
- A. Paladhi, S. Daripa, A. Nath and S. K. Hira, J. Immunol., 2024, 213, 1255–1263 CrossRef CAS PubMed.
- A. Paladhi, S. Daripa, I. Mondal and S. K. Hira, Front. Immunol., 2022, 13, 988071 CrossRef CAS PubMed.
- S. K. Hira, K. Mitra, P. Srivastava, S. Singh, S. Vishwakarma, R. Singh, B. Ray and P. P. Manna, Nanomedicine, 2020, 24, 102128 CrossRef CAS PubMed.
- P. Srivastava, S. K. Hira, A. Paladhi, R. Singh, U. Gupta, D. N. Srivastava, R. A. Singh and P. P. Manna, J. Mater. Chem. B, 2020, 8, 1411–1421 RSC.
- M. Das, R. Nasani, M. Saha, S. M. Mobin and S. Mukhopadhyay, Dalton Trans., 2015, 44, 2299–2310 RSC.
- X.-B. Fu, D.-D. Liu, Y. Lin, W. Hu, Z.-W. Mao and X.-Y. Le, Dalton Trans., 2014, 43, 8721–8731 RSC.
- P. Kumar, S. Gorai, M. K. Santra, B. Mondal and D. Manna, Dalton Trans., 2012, 41, 7573–7581 RSC.
- Y. Li, Z. Yang, M. Zhou, Y. Li, J. He, X. Wang and Z. Lin, RSC Adv., 2017, 7, 41527–41539 RSC.
- K. Karami, Z. M. Lighvan, S. A. Barzani, A. Y. Faal, M. Poshteh-Shirani, T. Khayamian, V. Eigner and M. Dušek, New J. Chem., 2015, 39, 8708–8719 RSC.
- D. İ. Özbağcı, Appl. Organomet. Chem., 2024, 38, e7697 CrossRef.
- R. R. Kumar and R. Ramesh, RSC Adv., 2015, 5, 101932–101948 RSC.
- A. K. De, T. Shaym, S. Ghosh, S. Dogra, A. R. Choudhury and D. Das, RSC Adv., 2026, 16, 11130–11136 RSC.
- J. L. Vanhooke, M. M. Benning, F. M. Raushel and H. M. Holden, Biochemistry, 1996, 35, 6020–6025 CrossRef CAS PubMed.
- J. Chen, X. Wang, Y. Zhu, J. Lin, X. Yang and Y. Li, Inorg. Chem., 2005, 44, 3430–3437 Search PubMed.
- I. Nikolic-Hughes, P. J. O'Brien and D. Herschlag, J. Am. Chem. Soc., 2005, 127, 9314–9315 CrossRef CAS PubMed.
- R. Sanyal, A. Guha, T. Ghosh, T. K. Mondal, E. Zangrando and D. Das, Inorg. Chem., 2014, 53, 855–865 CrossRef PubMed.
- M. Garai, D. Dey, H. R. Yadav, M. Maji, A. R. Choudhury and B. Biswas, J Chem. Sci., 2017, 129, 1513–1521 CrossRef CAS.
- M. Subat, K. Woinaroschy, C. Gerstl, B. Sarkar, W. Kaim and B. König, Inorg. Chem., 2008, 47, 4661–4670 CrossRef CAS PubMed.
- S. Anbu, S. Kamalraj, B. Varghese, J. Muthumary and M. Kandaswamy, Dalton Trans., 2012, 41, 13353–13361 Search PubMed.
- CCDC 2003428: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc257qqw.
| This journal is © The Royal Society of Chemistry 2026 |