 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Hybrid benzylidene thiazolidine-2,4-diones as potent apoptosis-inducing anticancer agents: design-driven optimization, cytotoxic profiling, and mechanistic validation in prostate cancer
Saad Shaaban†
 *a,
Samia S. Hawas†b,
Marwa Sharakyc,
Hussein Ba-Ghazala,
Ayman Abo Elmaatyde,
Khadra B. Alomarif,
Mohamed Alaasar
*a,
Samia S. Hawas†b,
Marwa Sharakyc,
Hussein Ba-Ghazala,
Ayman Abo Elmaatyde,
Khadra B. Alomarif,
Mohamed Alaasar g,
Asma M. Elsharifh,
Fatema S. Alatawii,
Mohamed Alaa Mohamedj,
Arwa Omar Al Khatibk and
Ahmed A. Al-Karmalawy
g,
Asma M. Elsharifh,
Fatema S. Alatawii,
Mohamed Alaa Mohamedj,
Arwa Omar Al Khatibk and
Ahmed A. Al-Karmalawy *bl
*bl
aDepartment of Chemistry, College of Science, King Faisal University, Al-Ahsa 31982, Saudi Arabia. E-mail: sibrahim@kfu.edu.sa
bDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy, Horus University-Egypt, New Damietta 34518, Egypt. E-mail: akarmalawy@horus.edu.eg
cCancer Biology Department, Pharmacology Unit, National Cancer Institute (NCI), Cairo University, Cairo, Egypt
dMedicinal Chemistry Department, Faculty of Pharmacy, Port Said University, Port Said 42526, Egypt
eMedicinal Chemistry Department, Clinical Pharmacy Program, East Port Said National University, Port Said 42526, Egypt
fJazan University, Department of Physical Sciences, Chemistry Division, 45142, Jazan, Kingdom of Saudi Arabia
gDepartment of Chemistry, Faculty of Science, Cairo University, Giza, Egypt
hDepartment of Chemistry, College of Science, Imam Abdulrahman Bin Faisal University, Dammam 31441, Saudi Arabia
iDepartment of Biochemistry, Faculty of Science, University of Tabuk, Tabuk, Saudi Arabia
jDepartment of Chemistry, Faculty of Science, Mansoura University, Mansoura, Egypt
kFaculty of Pharmacy, Al-Ahliyya Amman University, Amman, Jordan
lDepartment of Pharmaceutical Chemistry, College of Pharmacy, The University of Mashreq, Baghdad 10023, Iraq
First published on 12th April 2026
Abstract
The smart combination of fragment merging, bioisosteric replacement, and conformational rigidification allowed for the design of a new series of N-(phenylthiazolyl)acetamide hybrids based on benzylidene-thiazolidine-2,4-dione as possible anticancer leads. Compounds with significantly increased cytotoxic activity were developed by methodically altering the benzylidene C4 ring structure and specifically modifying the thiazole olefinic bond. Analogues HB161 and HB162, which outperformed doxorubicin (DOX) against a variety of cancer types, demonstrated the highest average growth inhibition (GI) among the synthesized derivatives (HB121–HB169). A thorough IC50 analysis showed that HB123 and HB161 were more effective against PC-3 prostate cancer cells (7.14 and 7.90 µM, respectively), whereas HB162 showed balanced activity across colorectal, breast, and prostate cancer models. Strong apoptosis induction was demonstrated by mechanistic investigation, which showed that BAX and Caspase-3/7/9 were significantly upregulated while BCL-2, MMP-2, and MMP-9 were suppressed. Moreover, flow cytometry demonstrated that HB162-induced G0/G1 cell-cycle arrest is a major contributor to its antiproliferative action. To support HB161 and HB162 as BCL-2 downregulators, a molecular docking study was conducted focusing on the BCL-2 receptor, an essential component of the pathway that triggers apoptosis. Positive physicochemical behavior, non-mutagenicity, and good drug-likeness were highlighted by in silico ADMET profiling. In conclusion, this hybrid scaffold offers a promising platform for next-generation anticancer drugs that target apoptosis, especially for the treatment of prostate cancer.
1. Introduction
Despite significant progress in cancer treatment, the disease continues to be a universal healthcare challenge.1–3 There were suggestions that global cancer cases could rise significantly, with estimates indicating an increase of more than 50% compared to current levels.4,5 Conventional anticancer therapies aim to kill rapidly dividing cells. However, this method also damages normal cells along with cancer cells. This can cause side effects like nausea, a weakened immune system, hair loss, and anemia.6–8Two protein families play a key role in controlling apoptosis, or controlled cell destruction: BCL-2 and caspases. The caspase cascade is initiated by the release of cytochrome c. This procedure gets the cell ready to die.9,10 When activated, BAX, a crucial protein in the BCL-2 family that regulates apoptosis, causes mitochondrial outer membrane permeabilization (MOMP).11,12 This pathway is essential as it facilitates the release of cytochrome c and other pro-apoptotic molecules, which start programmed cell death.9 Additionally, p53 is essential for limiting cell proliferation. The role of p53 as a central tumor suppressor is highlighted by the fact that it is found in the nucleus, where it controls gene expression and can trigger apoptosis when required.13,14 Cellular stressors like DNA damage can cause p53 to build up inside the nucleus. When activated as a tumor suppressor, it can exhibit two responses: stopping the cell cycle to allow for DNA repair and starting apoptosis to get rid of damaged cells when repair is not possible.13,15
A class of zinc-dependent endopeptidases known as matrix metalloproteinases (MMPs) is essential for breaking down elements of the basement membrane (BM) and extracellular matrix (ECM).16,17 Therefore, gelatin and collagens IV and V are the main targets of matrix metalloproteinases MMP-2 and MMP-9.18,19 Their increased activity in cancer breaks down these barriers, allowing tumor invasion and metastasis, and providing insight into how MMP-2 and MMP-9 affect the development of cancer.20,21
Defects in the apoptosis pathway are linked to decreased pro-apoptotic proteins and increased anti-apoptotic proteins, which encourage researchers to use these markers to assess the effectiveness and safety of anticancer medications.22–24
In addition to conventional chemotherapeutic strategies, 2,4-thiazolidinedione (2,4-TZD) derivatives have drawn increasing attention in recent years as promising scaffolds for anticancer drug discovery. Originally developed as insulin sensitizers targeting peroxisome proliferator-activated receptor gamma (PPARγ), structural diversification of the TZD core has revealed significant anticancer potential across multiple tumor models beyond its antidiabetic applications.25,26 Recent medicinal chemistry efforts have highlighted that 2,4-TZD derivatives can modulate key pathways involved in tumorigenesis, including cell proliferation, apoptosis, angiogenesis, and metastasis. Their structural flexibility—especially at the C-5 position—allows for strategic substitution and functionalization that can enhance target affinity, selectivity, and cytotoxic potency.25 Mechanistically, many 2,4-TZDs appear to influence cell cycle regulators (e.g., p21, p27), apoptotic proteins (e.g., BAD, p53), and oncogenic signaling pathways such as PI3K/AKT/mTOR and RAS/RAF/MEK/ERK, as well as downregulate growth factor receptors like EGFR and angiogenic factors such as VEGF.25 Several recent studies have demonstrated the effectiveness of TZD analogues against various cancer cell lines. For instance, novel 2,4-TZD derivatives have been shown to disrupt oncogenic protein–protein interactions, inhibit tubulin polymerization, and induce apoptosis and cell cycle arrest in colon cancer models.27 Other reports describe the synthesis of TZD-bearing hybrid molecules exhibiting potent growth inhibition that rivals or surpasses standard chemotherapeutics such as doxorubicin (DOX), underlining their potential as cytotoxic agents.28 In addition, TZD derivatives incorporating heterocyclic or arylidene moieties continue to show promising antiproliferative profiles in breast, lung, and hepatic cancer cell lines, further supporting the versatility of this scaffold in anticancer drug design.29 Collectively, these advancements underscore 2,4-thiazolidinediones as valuable pharmacophores in the ongoing search for new anticancer therapeutics, motivating continued exploration of their structure–activity relationships and molecular mechanisms of action.25
The antitumor potential of N-(phenylthiazole) acetamide and benzylidene thiazolidine-2,4-dione scaffolds has been brought to light by efforts to find effective chemotherapeutics. In their study, Laxmi et al. developed benzylidene thiazolidine-2,4-dione analogues (compounds I and II, Fig. 1), which showed notable anticancer effects on a panel of 60 different types of cancer cells.30
The developed derivatives demonstrated significant efficacy against various cancer cell lines, including non-small cell lung cancer (NCI-H522), breast cancer (MDA-MB-468), prostate cancer (PC-3), colon cancer (COLO 205), and others.30 In addition, a series of benzylidene thiazolidine-2,4-dione derivatives (e.g., compound III, Fig. 1) were synthesized by El-Kashef et al., who subsequently evaluated their antitumor potential for breast cancer treatment.31 All synthesized compounds incorporated a thiazolidine-2,4-dione core conjugated with a trimethoxy-benzylidene group. Their anti-breast cancer activity was assessed using both normal breast cells and human breast cancer cells (MCF-7 and MDA-MB-231).31
Moreover, Al-Wahaibi et al. have afforded N-(phenylthiazole) acetamide derivatives as tubulin polymerization inhibitors endowed with apoptotic antiproliferative activity (e.g., compounds IV and V).32 The MTT assay was used to examine the novel compounds' antiproliferative activity against HCT-116 colon cancer, PC-3 prostate cancer, and MCF-7 and MDA-MB-231 breast cancer cell lines. The results showed a significant cytotoxic effect when compared to reference medications, sorafenib, and DOX.32
Herein, a fragment merging approach was used to make benzylidene thiazolidine-2,4-dione and N-(phenylthiazole) acetamide containing hybrids to reveal their activity as new cytotoxic candidates. Moreover, the benzylidene ring was substituted at C4 with diverse bio-isosteric groups to pursue a change in cytotoxic activity. Additionally, the olefinic bond of the benzylidene moiety was replaced by a freely rotatable single bond for some derivatives to highlight the effect of conformation restriction of the designed compounds on cytotoxic activity.
2. Results and discussion
2.1. Chemistry
The thiazole ring is a significant pharmacophore responsible for a variety of biological activities.33–37 They present as key structural components in numerous natural products, such as vitamin B1, and in pharmaceutical drugs, including the antidiabetic agents troglitazone and pioglitazone.34,38,39 Schemes 1 and 2 show the synthetic procedures utilized to create thiazolidinedione hybrids.Thiazolidinedione hybrids were obtained via reaction of thiourea and phenacyl bromide, which furnished 4-phenylthiazol-2-amine (3) in 99% yield (Scheme 1). Subsequent reaction of 2-amino-thiazol 3 with phenyl isothiocyanate afforded the 1-phenyl-3-(4-phenylthiazol-2-yl)thiourea (HB169) in 81% yield. Furthermore, the acylation of 2-amino-thiazol 3 with chloroacetyl chloride in dichloromethane afforded the corresponding acetamide (4), which, upon reaction with the K salt of thiazolidine-2,4-dione, provided 2-(2,4-dioxothiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB045) in 90% yield (Scheme 1).
The FT-IR spectra of HB169 revealed significant bands at 1593–1493 cm-1 that corresponded to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N and aromatic C
N and aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching of the thiazole and phenyl moieties, along with a broad N–H stretching absorption at 3144 and 2936 cm−1, which is typical of the NH groups of thiourea. In the 1HNMR spectrum, two downfield singlets at δ 13.34 and 12.76 ppm correspond to the two hydrogen-bonded thiourea NH protons, confirming formation of the N,N′-disubstituted thiourea. The remaining signals appear between δ 8.68 and 7.15 ppm as multiplets and doublets integrating for eleven aromatic protons, in agreement with two phenyl rings and the thiazole proton. At δ 8.68 ppm, the most deshielded singlet can be attributed to the thiazole ring's H-5, which is typically shifted downfield by the adjacent heteroatoms, while the pattern of doublets, triplets, and multiplets between δ 8.04–7.15 ppm matches two monosubstituted phenyl groups. Signals at δ 178.26 and 163.15 ppm in the 13C NMR spectra can be referred to the thiocarbonyl (C
C stretching of the thiazole and phenyl moieties, along with a broad N–H stretching absorption at 3144 and 2936 cm−1, which is typical of the NH groups of thiourea. In the 1HNMR spectrum, two downfield singlets at δ 13.34 and 12.76 ppm correspond to the two hydrogen-bonded thiourea NH protons, confirming formation of the N,N′-disubstituted thiourea. The remaining signals appear between δ 8.68 and 7.15 ppm as multiplets and doublets integrating for eleven aromatic protons, in agreement with two phenyl rings and the thiazole proton. At δ 8.68 ppm, the most deshielded singlet can be attributed to the thiazole ring's H-5, which is typically shifted downfield by the adjacent heteroatoms, while the pattern of doublets, triplets, and multiplets between δ 8.04–7.15 ppm matches two monosubstituted phenyl groups. Signals at δ 178.26 and 163.15 ppm in the 13C NMR spectra can be referred to the thiocarbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S) and C-2 of the thiazole ring, respectively. Signals in the range δ 139.63–124.01 ppm correspond to the aromatic carbons of both phenyl rings and the remaining thiazole carbons, while the resonance at δ 105.89 ppm is consistent with the thiazole C-5 carbon.
S) and C-2 of the thiazole ring, respectively. Signals in the range δ 139.63–124.01 ppm correspond to the aromatic carbons of both phenyl rings and the remaining thiazole carbons, while the resonance at δ 105.89 ppm is consistent with the thiazole C-5 carbon.
A broad N–H stretching band at 3224–3090 cm−1 in the FT-IR spectrum of HB045 is suggestive of the amide/thiazolidinedione NH functionality. Strong absorptions at 1759 and 1654 cm−1 are characteristic of the two carbonyl groups of the thiazolidinedione ring (imide-type C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) and the amide C
O) and the amide C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, respectively. The amide and thiazolidinedione fragments' N–H bending and C–N/C–O stretching vibrations are represented by extra bands at 1559, 1325–1275, and 1182 cm−1. The amide/thiazolidinedione NH proton was identified by a singlet signal at δ 12.68 ppm in the 1H NMR spectrum. The aromatic region (δ 7.88–7.32 ppm) shows one singlet and three sets of doublets/triplets integrating for five protons, in agreement with a monosubstituted phenyl ring and the thiazole proton. The methylene group tying the amide nitrogen to the thiazolidinedione ring and the thiazole-attached acetyl methylene are represented by two singlets at δ 4.47 and 4.32 ppm (2H each), respectively. In the 13C NMR spectrum, two carbonyl carbons of the thiazolidinedione ring appear at δ 172.01 and 171.51 ppm, while the amide carbonyl and thiazole C-2 resonate at δ 164.79 and 158.38 ppm, respectively. Aromatic and thiazole carbons are observed at δ 128.83, 127.92, 126.26, and 108.22 ppm. The two methylene carbons appear at δ 43.79 and 34.50 ppm, supporting the presence of distinct CH2 groups adjacent to heteroatoms (N and C
O, respectively. The amide and thiazolidinedione fragments' N–H bending and C–N/C–O stretching vibrations are represented by extra bands at 1559, 1325–1275, and 1182 cm−1. The amide/thiazolidinedione NH proton was identified by a singlet signal at δ 12.68 ppm in the 1H NMR spectrum. The aromatic region (δ 7.88–7.32 ppm) shows one singlet and three sets of doublets/triplets integrating for five protons, in agreement with a monosubstituted phenyl ring and the thiazole proton. The methylene group tying the amide nitrogen to the thiazolidinedione ring and the thiazole-attached acetyl methylene are represented by two singlets at δ 4.47 and 4.32 ppm (2H each), respectively. In the 13C NMR spectrum, two carbonyl carbons of the thiazolidinedione ring appear at δ 172.01 and 171.51 ppm, while the amide carbonyl and thiazole C-2 resonate at δ 164.79 and 158.38 ppm, respectively. Aromatic and thiazole carbons are observed at δ 128.83, 127.92, 126.26, and 108.22 ppm. The two methylene carbons appear at δ 43.79 and 34.50 ppm, supporting the presence of distinct CH2 groups adjacent to heteroatoms (N and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O).
O).
The corresponding arylidene derivatives were then obtained by subjecting the thiazolidinedione–thiazole hybrid HB045 to the Knoevenagel condensation with a number of aromatic aldehydes, including benzaldehyde, 4-bromobenzaldehyde, 4-chlorobenzaldehyde, 4-fluorobenzaldehyde, and picolinaldehyde (Scheme 2). We assessed a variety of catalysts (AcOH, piperidine, and triethylamine) and solvents (polar protic, polar aprotic, and nonpolar) in our optimization investigations. These conditions, however, frequently resulted in a considerable recovery of the starting material but generally low conversions and product purity. However, utilizing ethanol as the solvent in conjunction with a catalytic mixture of piperidine and AcOH proved to be the most effective, allowing for complete conversion to the intended thiazolidinone-based arylidenes in fair to good yields (up to 42–73%).
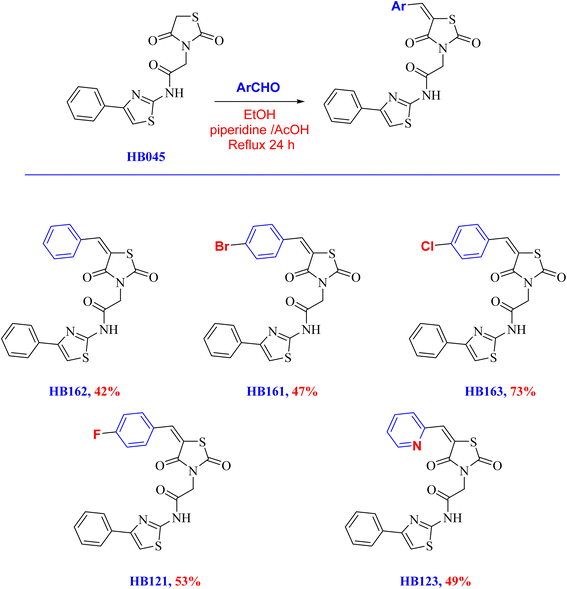 | ||
| Scheme 2 Synthesis of thiazolidinone-based arylidene derivatives via Knoevenagel condensation reactions. | ||
Broad bands in the 3250–3380 cm−1 range were visible in the FT-IR for HB162, HB161, HB163, HB121, and HB123, which is consistent with the amide/thiazolidinedione N–H functionality. Strong absorptions between 1758–1738 and 1692–1654 cm−1 are attributed to the amide carbonyl and the two imide C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups of the thiazolidinedione ring. Additional bands at 1540–1560 cm−1 and 1610–1619 cm−1 are indicative of the thiazole ring and arylidene fragments' C
O groups of the thiazolidinedione ring. Additional bands at 1540–1560 cm−1 and 1610–1619 cm−1 are indicative of the thiazole ring and arylidene fragments' C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C/C
C/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching. The amide/thiazolidinedione NH was identified by a downfield singlet signal in the 1HNMR at δ ≈ 12.7 ppm. At δ ≈ 8.0 ppm, the olefinic proton of the 5 arylidene group was detected as a singlet. The methylene group (CH2CO) linker that connects the thiazolidinedione core to the thiazole acetamide is responsible for the singlet signal that all compounds displayed at δ ≈ 4.64–4.66 ppm (2H). All compounds showed three downfield carbonyl signals in the 13CNMR around δ ≈ 171–169 and 167–165 ppm, which correspond to the amide C
N stretching. The amide/thiazolidinedione NH was identified by a downfield singlet signal in the 1HNMR at δ ≈ 12.7 ppm. At δ ≈ 8.0 ppm, the olefinic proton of the 5 arylidene group was detected as a singlet. The methylene group (CH2CO) linker that connects the thiazolidinedione core to the thiazole acetamide is responsible for the singlet signal that all compounds displayed at δ ≈ 4.64–4.66 ppm (2H). All compounds showed three downfield carbonyl signals in the 13CNMR around δ ≈ 171–169 and 167–165 ppm, which correspond to the amide C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O of the acetamide chain and the two thiazolidinedione carbonyls. Additionally, a methylene (CH2) carbon at δ ≈ 44–46 ppm was present in all derivatives, which corresponds to the methylene group of the acetamide linker.
O of the acetamide chain and the two thiazolidinedione carbonyls. Additionally, a methylene (CH2) carbon at δ ≈ 44–46 ppm was present in all derivatives, which corresponds to the methylene group of the acetamide linker.
2.2. Evaluation of biology
To further evaluate the safety of the synthesized compounds in non-malignant cells, the human skin fibroblast (HSF) line was incorporated into the study. The GI% for analogues HB121–HB169 are presented in Table 1, with their efficacy compared against the reference anticancer drug (DOX).
| HB121 | HB123 | HB161 | HB162 | HB163 | HB169 | DOX | |
|---|---|---|---|---|---|---|---|
| H460 | 81.99 | 68.07 | 83.86 | 90.34 | 80.03 | 79.48 | 73.58 |
| ±5.13 | ±2.73 | ±2.32 | ±6.66 | ±11.12 | ±1.97 | ±1.76 | |
| HCT-116 | 63.10 | 44.65 | 85.56 | 82.52 | 67.48 | 55.48 | 71.28 |
| ±3.55 | ±8.90 | ±2.81 | ±5.75 | ±4.18 | ±5.45 | ±5.07 | |
| MCF7 | 58.27 | 47.80 | 79.70 | 81.28 | 42.35 | 70.17 | 69.00 |
| ±9.17 | ±4.30 | ±6.57 | ±3.20 | ±4.92 | ±6.14 | ±0.16 | |
| HuH7 | 63.49 | 51.84 | 29.41 | 17.51 | 51.21 | 53.81 | 74.81 |
| ±4.73 | ±5.86 | ±10.04 | ±9.18 | ±6.10 | ±4.34 | ±0.69 | |
| PC-3 | 76.24 | 70.00 | 89.70 | 89.04 | 82.49 | 82.68 | 68.93 |
| ±3.95 | ±4.33 | ±5.14 | ±4.82 | ±4.94 | ±3.18 | ±5.42 | |
| A549 | 59.21 | 20.06 | 63.34 | 65.14 | 53.12 | 66.55 | 49.00 |
| ±2.48 | ±3.66 | ±3.22 | ±5.91 | ±6.76 | ±3.61 | ±1.46 | |
| MG63 | 31.07 | −11.31 | 76.84 | 78.42 | 32.84 | 27.76 | 69.87 |
| ±4.71 | ±22.51 | ±2.09 | ±3.89 | ±7.29 | ±2.65 | ±1.61 | |
| HSF | 66.66 | 21.54 | 65.30 | 62.05 | 59.29 | 66.72 | 26.79 |
| ±5.79 | ±6.83 | ±7.33 | ±6.77 | ±8.73 | ±5.31 | ±6.62 |
Interestingly, compounds HB161 and HB162 could display the highest mean GI% (72.63% and 72.04%, respectively), exceeding the DOX average GI% (68.07%). Furthermore, among the compounds under investigation, compounds HB169, HB121, and HB163 had an exceptional mean GI% (62.27, 61.91, and 58.50%, respectively). Among the compounds provided, compound HB123 may exhibit the lowest mean GI% (41.59%), as shown in Table 1.
Furthermore, compound HB123 demonstrated favorable safety and selectivity toward malignant cells, as reflected by its low GI% in normal HSF.
2.2.2.1 Structure–activity relationships (SAR) study. The bioisosteric replacement of the phenyl ring of the benzylidene thiazolidine-2,4-dione core with a pyridine ring and retaining the thiazole exocyclic olefinic bond (compound HB123), Fig. 3, resulted in the highest cytotoxic potency against prostate cancer with the lowest IC50 (7.14 µM), better than the reference cytotoxic drug DOX (8.41 µM). This enhanced activity can be attributed to the electron-deficient nature of the pyridine ring, which may improve hydrogen bonding and polar interactions within the biological target.
Besides, it was revealed that C-4 substitution of the phenyl ring of the benzylidene thiazolidine-2,4-dione core with bromide and giving up the thiazole exocyclic olefinic bond (compound HB161) can afford highly outstanding prostate cytotoxic potential (IC50 = 7.90 µM). The improved activity of this analogue may be associated with the strong electron-withdrawing effect and increased polarizability of the bromine atom, which can enhance hydrophobic and halogen-bonding interactions. Hence, we can argue that conformational restriction of our afforded compounds does not play a pivotal role in prostate cytotoxic biological activity, Fig. 3. The absence of benzylidene thiazolidine-2,4-dione core (compound HB169) can display the least prostate cytotoxic potential with an IC50 value of 12.93 µM, highlighting the critical role of this scaffold in maintaining cytotoxic potency.
Furthermore, the investigated compounds exhibited lower cytotoxic potential against breast cancer (MCF-7) cells compared to DOX. It was displayed that without C-4 substitution of the phenyl ring of the benzylidene thiazolidine-2,4-dione core and keeping the thiazole exocyclic olefinic bond (compound HB162), reasonable breast cytotoxic potential can be exhibited with IC50 = 15.44 µM (DOX IC50 = 10.74 µM). This suggests that the absence of substituents on the phenyl ring may reduce electronic modulation and binding affinity. Similarly, the absence of benzylidene thiazolidine-2,4-dione core (compound HB169) can display the lower breast cytotoxic potential with an IC50 value of 65.66 µM. Hence, we can argue that the benzylidene thiazolidine-2,4-dione core may be regarded as essential for prostate and breast cytotoxic potential.
In the case of colorectal cancer (HCT-116), it was demonstrated that, in contrast to DOX (IC50 = 10.46 µM), dependable colon cytotoxic potential could be achieved with IC50 values of 14.82 and 17.95 µM, respectively, without C-4 substitution of the phenyl ring of the benzylidene thiazolidine-2,4-dione core and maintaining the thiazole exocyclic olefinic bond (compound HB162) or C-4 substitution of the phenyl ring of the benzylidene thiazolidine-2,4-dione core with bromide and in the absence of the thiazole exocyclic olefinic bond (compound HB161), which may be attributed to a balance between steric bulk and electronic effects. With an IC50 value of 100.91 µM, compound HB123, which bioisosterically substitutes a pyridine ring for the phenyl ring of the benzylidene thiazolidine-2,4-dione core while keeping the thiazole exocyclic olefinic link, can exhibit the lowest colon cytotoxic potential, indicating that the pyridine substitution may not be favorable for interaction with targets relevant to colon cancer.
2.2.2.2 Apoptosis-related indicators' protein expression. The apoptotic activity of the two analogues, HB161 and HB162, in PC-3 prostate cancer cells was assessed using a protein expression study. In order to shed light on how each molecule alters cell death signaling pathways, the evaluation concentrated on important pro- and anti-apoptotic indicators.
The pro-apoptotic proteins Caspase-3, Caspase-7, and Caspase-9 (1.87-, 1.12-, and 2.53-fold, respectively) and BAX (2.81-fold) were clearly upregulated after HB161 treatment. The anti-apoptotic and metastasis-associated proteins MMP-2, MMP-9, and BCL-2, on the other hand, were downregulated, exhibiting 1.31-, 1.89-, and 2.23-fold lower expression levels, respectively.
A similar pattern was observed with HB162, which also enhanced pro-apoptotic signaling. BAX was increased to 2.70-fold, while Caspase-3, Caspase-7, and Caspase-9 rose to 2.04-, 1.41-, and 1.93-fold, respectively. The anti-apoptotic markers MMP-2, MMP-9, and BCL-2, on the other hand, were suppressed, decreasing to 1.39-, 1.75-, and 2.48-fold, respectively.
When everything is taken into account, HB162 has a particularly potent apoptotic effect on PC-3 cells, as evidenced by the marked upregulation of pro-apoptotic mediators and the obvious suppression of proteins linked to invasion and apoptosis. The potential of both analogues as viable apoptosis-inducing candidates is further supported by the significant apoptotic activity displayed by HB161. Fig. 4 shows apoptotic markers.
To ensure the reliability of these findings, the experiment was repeated three times, and representative histograms are provided in the SI (Supplementary Figures S1 and S2).
2.3. Studies conducted in silico
The binding scores for the analogues HB161 and HB162 were −7.31 and −7.34 kcal mol−1, respectively, with root mean square deviation (RMSD) values of 1.64 and 1.90 Å. Additionally, at an RMSD of 1.89 Å, the docked co-crystal of BCL-2 achieved a binding score of −9.94 kcal mol−1. The smaller sizes of the analogues HB161 and HB162 in comparison to the large co-crystal inhibitor (N-heteroaryl sulfonamide), which can establish more hydrophobic and hydrogen contacts, may be the reason for their lower scores.
Three pi-hydrogen interactions with Asp70, Phe112, and Gly104 fixed the small analogue HB161 inside the BCL-2's active pocket. However, in addition to forming a hydrogen bond with Glu95, the small homologue HB162 demonstrated the creation of two pi-hydrogen interactions with Asp70 and Phe112. Additionally, four hydrogen bonds were established between the docked BCL-2 co-crystal inhibitor and Arg66, Gly104, and Asp70 (2). Additionally, it created a single pi–hydrogen bond with Leu96 (Fig. 6).
 | ||
| Fig. 6 3D binding interactions between the co-crystal inhibitor and the analogues HB161 and HB162 in the BCL-2 receptor's active site (PDB ID: 4IEH). | ||
As a result, it is strongly advised that the small analogues (HB161 and HB162) be considered as prospective BCL-2 downregulators. They may also be further optimized by using an extension strategy to reach the additional empty pocket of the BCL-2 active site.
 | ||
| Fig. 7 The Boiled-egg diagram for the newly synthesized analogues (HB121–HB169), along with DOX as a reference control. | ||
Interestingly, none of the synthesized hybrids were found to violate Lipinski's rule of five.43 SI Fig. S3 presents bioavailability radar plots for the investigated compounds, providing insights into their bioavailability characteristics.
Furthermore, none of the synthesized compounds exhibited Ames toxicity, suggesting a reduced likelihood of mutagenicity.44 Additionally, the synthesized compounds are predicted to be safe with respect to cardiotoxicity, as they do not inhibit hERG I channels involved in cardiac electrical activity.45 However, inhibition of hERG II by all synthesized compounds may pose a potential risk of cardiac arrhythmia.46 All examined derivatives appear to possess hepatotoxic potential, as shown in Table 2.
| HB121 | HB123 | HB161 | HB162 | HB163 | HB169 | DOX | ||
|---|---|---|---|---|---|---|---|---|
| a MS: moderately soluble; S: soluble. | ||||||||
| Molecular properties | Molar refractivity | 118.93 | 116.77 | 126.35 | 118.97 | 123.66 | 93.47 | 132.66 |
| TPSA (Az) | 132.91 | 145.80 | 132.91 | 132.91 | 132.91 | 97.28 | 206.07 | |
| Log P o/w (WLOGP) | 4.36 | 3.20 | 4.25 | 3.80 | 4.14 | 4.24 | −0.32 | |
| Consensus log P o/w | 3.68 | 2.77 | 3.97 | 3.43 | 3.89 | 3.77 | 0.44 | |
| Water solubility | MS | MS | MS | MS | MS | MS | S | |
| Pharmacokinetics parameters and drug/lead likeness | GI absorption | Low | Low | Low | Low | Low | High | Low |
| BBB permeant | No | No | No | No | No | No | No | |
| P-gp substrate | No | No | No | No | No | No | Yes | |
| CYP1A2 inhibitor | Yes | No | Yes | Yes | Yes | Yes | No | |
| CYP2C19 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| CYP2C9 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| CYP2D6 inhibitor | No | No | No | No | No | Yes | No | |
| CYP3A4 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| Drug likeness (lipinski) | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| Lead likeness | No | No | No | No | No | No | No | |
| Toxicity parameters | Ames toxicity | No | No | No | No | No | No | Yes |
| Max. Tolerated dose (log mg per kg per day) | 0.63 | 0.363 | 0.638 | 0.062 | 0.033 | 0.008 | 0.405 | |
| hERG I inhibitor | No | No | No | No | No | No | No | |
| hERG II inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| Oral rat acute toxicity (LD50) (mol kg−1) | 2.892 | 2.996 | 2.932 | 2.723 | 2.767 | 2.604 | 3.934 | |
| Oral rat chronic toxicity (log mg per kg_bw per day) | 1.69 | 1.166 | 1.609 | 0.886 | 0.954 | 1.113 | 3.435 | |
| Hepatotoxicity | Yes | Yes | Yes | Yes | Yes | Yes | No | |
| Minnow toxicity (log mM) | 0.298 | 1.443 | −0.144 | 1.129 | 0.071 | 1.889 | 8.145 | |
3. Conclusion
In this study, a rational design strategy integrating fragment merging, bioisosteric replacement, and conformational control successfully yielded a new class of benzylidene thiazolidine-2,4-dione/N-(phenylthiazole) acetamide hybrids with promising anticancer potential. With cytotoxic profiles that were either better than or comparable to DOX across a variety of cancer cell lines, HB161, HB162, and HB123 were found to be the most active analogues via systematic structural diversification, which also led to distinct SAR trends. Notably, HB162 showed broad-spectrum antiproliferative effects and caused a noticeable G0/G1 cell-cycle arrest, while HB123 and HB161 showed strong activity against PC-3 prostate cancer cells. By upregulating BAX and Caspases-3/7/9 and suppressing the anti-apoptotic mediators BCL-2, MMP-2, and MMP-9, mechanistic validation showed that HB161 and HB162 activate apoptotic pathways to produce their anticancer effects. The small analogues (HB161 and HB162) were suggested by molecular docking to be promising BCL-2 downregulators. They may be further optimized by using an extension strategy to reach the additional empty pocket of the BCL-2 active site. Although future optimization is necessary to improve oral bioavailability and reduce potential cardiotoxic and hepatotoxic liabilities, complementary ADMET predictions indicated favorable drug-likeness, lack of mutagenicity, and acceptable pharmacokinetic behavior. Taken together, these results present a flexible hybrid scaffold with proven biological activity and a well-defined mechanistic basis, establishing these compounds—specifically HB161 and HB162—as viable lead candidates for additional preclinical research in solid tumors, including prostate cancer. These hybrids may progress toward clinically useful anticancer treatments with further structural improvement and thorough pharmacological analysis.4. Materials and methods
4.1. Chemistry
Compounds 3,47 4,48 and thiazolidine-2,4-dione49 and its potassium salt were synthesized using the reported literature approach (see also SI for the preparation methods).39![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DMF (1
DMF (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1).
1).
4.1.3.1 Synthesis of 2-(5-benzylidene-2,4-dioxothiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB162). After being synthesized from HB045 (15 mmol) and benzaldehyde (18 mol), compound HB162 was separated as a white powder; yield = 42%; MP = 235–236 °C; FT-IR (ν, cm−1): 3286, 1741, 1692, 1670, 1610, 1560, 1483, 1376, 1148, 725, 665, 515; 1H NMR (500 MHz, DMSO-d6) δ 12.77 (s, 1H, NH), 8.00 (s, 1H, Ar–H), 7.89 (d, J = 7.5 Hz, 2H, Ar–H), 7.67 (s, 2H, Ar–H), 7.57 (t, J = 7.3 Hz, 2H, Ar–H), 7.53 (d, J = 7.1 Hz, 1H, Ar–H), 7.43 (t, J = 7.6 Hz, 2H, Ar–H), 7.33 (t, J = 7.3 Hz, 1H, Ar–H), 4.66 (s, 2H, CH2); 13C NMR (151 MHz, pyridine-d5) δ 167.54, 165.70, 164.76, 158.40, 134.27, 133.33, 130.70, 130.32, 129.32, 128.85, 127.94, 126.28, 122.98, 121.60, 108.29, 43.96; MS (ESI): m/z = calcd. For C21H15N3O3S2 [M+]: 421.4, found 419.6 [M+-2], 421.7 [M+-1]; HRMS calcd. For C21H15N3O3S2 [M+]: 421.4890, found 422.0625 [M+].
4.1.3.2 Synthesis of 2-(5-(4-bromobenzylidene)-2,4-dioxothiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB161). HB045 (15 mmol) and 4-bromo-benzaldehyde (18 mol) were combined to synthesize compound HB161, which was then separated as a white powder; yield = 47%; MP = 263–264 °C; FT-IR (ν, cm−1): 3363, 3118, 2980, 1756, 1686, 1618, 1838, 1483, 1275, 1071, 1005, 814; 1H NMR (500 MHz, DMSO-d6) δ 12.77 (s, 1H, NH), 7.98 (s, 1H, Ar–H), 7.89 (d, J = 7.6 Hz, 2H, Ar–H), 7.77 (d, J = 8.4 Hz, 2H, Ar–H), 7.67 (s, 1H, Ar–H), 7.61 (d, J = 8.4 Hz, 2H, Ar–H), 7.43 (t, J = 7.6 Hz, 2H, Ar–H), 7.32 (t, J = 7.3 Hz, 1H, Ar–H), 4.65 (s, 2H, CH2); 13C NMR (151 MHz, pyridine-d5) δ 174.74, 171.78, 166.94, 158.09, 137.84, 134.37, 133.99, 133.42, 133.20, 131.33, 130.29, 129.40, 127.78, 124.95, 109.73, 45.52; HRMS calcd. For C21H14BrN3O3S2 [M+]: 500.3850, found 501.9712 [M+ + 1], 502.9741 [M+ + 2].
4.1.3.3 Synthesis of 2-(5-(4-chlorobenzylidene)-2,4-dioxothiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB163). HB045 (15 mmol) and 4-chloro-benzaldehyde (18 mol) were combined to synthesize compound HB163, which was then separated as a white powder; yield = 73%; MP = 258–259 °C; FT-IR (ν, cm−1): 3364, 3117, 2977, 1758, 1686, 1677, 1654, 1611, 1542, 1483, 1542, 1370, 1324, 1275, 1073, 1061, 828; 1H NMR (500 MHz, DMSO-d6) δ 12.77 (s, 1H, NH), 8.00 (s, 1H, Ar–H), 7.87 (t, J = 14.6 Hz, 2H, Ar–H), 7.69 (d, J = 4.4 Hz, 1H, Ar–H), 7.68 (d, J = 5.2 Hz, 2H, Ar–H), 7.63 (d, J = 8.5 Hz, 2H, Ar–H), 7.43 (t, J = 7.6 Hz, 2H, Ar–H), 7.32 (t, J = 7.3 Hz, 1H, Ar–H), 4.65 (s, 2H, CH2); 13C NMR (151 MHz, pyridine-d5) δ 168.74, 167.07, 166.21, 159.89, 137.80, 134.34, 133.46, 133.14, 131.01, 130.35, 129.45, 127.79, 123.77, 109.80, 45.51; HRMS calcd. For C21H14ClN3O3S2 [M+]: 455.93100, found 456.0234 [M+ + 1], 457.0262 [M+ + 2], 458.0226 [M+ + 3], 459.0134 [M+ + 4], 460.0134 [M+ + 5].
4.1.3.4 Synthesis of 2-(5-(4-fluorobenzylidene)-2,4-dioxothiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB121). HB045 (15 mmol) and 4-flouro-benzaldehyde (18 mol) were combined to synthesize compound HB121, which was then separated as a green powder; yield = 53%; MP = 227–228 °C; FT-IR (ν, cm−1): 3363, 3252, 3208, 2948, 2922, 1687, 1619, 1546, 1517, 1479, 1436, 1197, 1013, 992; 1H NMR (500 MHz, DMSO-d6) δ 12.77 (s, 1H, NH), 8.01 (s, 1H, Ar–H), 7.89 (d, J = 7.5 Hz, 2H, Ar–H), 7.74 (dd, J = 8.1, 5.6 Hz, 2H, Ar–H), 7.67 (s, 1H, Ar–H), 7.45–7.37 (m, 4H, Ar–H), 7.32 (t, J = 7.3 Hz, 1H, Ar–H), 4.65 (s, 2H, CH2); 13C NMR (151 MHz, DMSO-d6) δ 166.76, 160.27, 154.28, 149.02, 134.87, 129.11, 128.03, 126.01, 107.01, 45.30; HRMS calcd. For C21H14FN3O3S2 [M+]: 439.4794, found 440.0522 [M+ + 1], 441.0554 [M+ + 2], 442.0463 [M+ + 3], 443.0502 [M+ + 4].
4.1.3.5 Synthesis of 2-(2,4-dioxo-5-(pyridin-2-ylmethylene)thiazolidin-3-yl)-N-(4-phenylthiazol-2-yl)acetamide (HB123). HB045 (15 mmol) and pyridine-2-carbaldehyde (18 mol) were combined to create compound HB123, which was then separated as a grey powder; yield = 49%; MP = 274–275 °C; FT-IR (ν, cm−1): 3254, 3203, 3050, 3003, 2936, 1738, 1682, 1656, 1616, 1582, 1408, 1386, 1276, 1249, 1148, 779; 1H NMR (500 MHz, DMSO-d6) δ 12.76 (s, 1H, NH), 8.79 (d, J = 4.2 Hz, 1H, Ar–H), 8.02 (s, 1H, Ar–H), 7.97 (t, J = 7.6 Hz, 1H, Ar–H), 7.93–7.86 (m, 3H, Ar–H), 7.67 (s, 1H, Ar–H), 7.50–7.38 (m, 3H, Ar–H), 7.32 (t, J = 7.3 Hz, 1H, Ar–H), 4.64 (s, 2H, CH2); 13C NMR (151 MHz, pyridine-d5) δ 171.63, 166.09, 164.94, 151.63, 137.12, 129.77, 128.84, 128.80, 127.92, 127.70, 127.52, 126.28, 123.81, 108.25, 43.36; MS (ESI): m/z = calcd. For C20H14N4O3S2 [M+]: 422.4, found 420.7 [M+ − 2], 421.7 [M+ − 1], 422.7 [M+]; HRMS calcd. For C20H14N4O3S2 [M+]: 422.4770, found 440.0522 [M+ + 1], 441.0554 [M+ + 2], 442.0463 [M+ + 3], 443.0502 [M+ + 4].
4.2. Evaluation of biology
4.3. Studies conducted in silico
Author contributions
Conceptualization and supervision: Saad Shaaban and Ahmed A. Al-Karmalawy; data curation, visualization, methodology, and writing–review & editing: Saad Shaaban, Samia S. Hawas, Marwa Sharaky, Hussein Ba-Ghazal, Ayman Abo Elmaaty, Khadra B. Alomari, Mohamed Alaasar, Asma M. Elsharif, Fatema S. Alatawi, Mohamed Alaa Mohamed, Arwa Omar Al Khatib, Ahmed A. Al-Karmalawy. Finally, all authors revised and approved the final submitted version of the manuscript.Conflicts of interest
The authors declared no conflict of interest.Data availability
The data supporting this article have been included in the manuscript and the Supporting Information (SI). Supplementary information: chemistry (methods and materials); spectroscopic data; biological data; ADMET and physicochemical studies. See DOI: https://doi.org/10.1039/d6ra01323f.Funding
This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia [Grant No. KFU261378].References
- Nat. Cancer, 2020, 1, 1–2 Search PubMed.
- F. R. Weth, G. B. Hoggarth, A. F. Weth, E. Paterson, M. P. J. White, S. T. Tan, L. Peng and C. Gray, Br. J. Cancer, 2024, 130, 703–715 Search PubMed.
- R. Kaur, A. Bhardwaj and S. Gupta, Mol. Biol. Rep., 2023, 50, 9663–9676 Search PubMed.
- F. Martínez-Jiménez, F. Muiños, I. Sentís, J. Deu-Pons, I. Reyes-Salazar, C. Arnedo-Pac, L. Mularoni, O. Pich, J. Bonet and H. Kranas, Nat. Rev. Cancer, 2020, 20, 555–572 CrossRef PubMed.
- T. Al-Warhi, A. A. Al-Karmalawy, A. A. Elmaaty, M. A. Alshubramy, M. Abdel-Motaal, T. A. Majrashi, M. Asem, A. Nabil, W. M. Eldehna and M. Sharaky, J. Enzyme Inhib. Med. Chem., 2023, 38, 176–191 CrossRef CAS PubMed.
- M. Demaria, M. N. O'Leary, J. Chang, L. Shao, S. Liu, F. Alimirah, K. Koenig, C. Le, N. Mitin and A. M. Deal, Cancer Discov., 2017, 7, 165–176 CrossRef CAS PubMed.
- A. A. Al-Karmalawy, M. S. Nafie, M. A. Shaldam, A. A. Elmaaty, S. A. Antar, A. A. El-Hamaky, M. A. Saleh, A. Elkamhawy and H. O. Tawfik, J. Med. Chem., 2023, 66, 777–792 CrossRef CAS PubMed.
- R. R. Bhandare, C. S. Munikrishnappa, G. V. Suresh Kumar, S. K. Konidala, D. K. Sigalapalli, Y. Vaishnav, S. Chinnam, H. Yasin, A. A. Al-karmalawy and A. B. Shaik, J. Saudi Chem. Soc., 2022, 26, 101447 CrossRef CAS.
- M. Brentnall, L. Rodriguez-Menocal, R. L. De Guevara, E. Cepero and L. H. Boise, BMC Cell Biol., 2013, 14, 1–9 CrossRef PubMed.
- Z. Zhou, T. Arroum, X. Luo, R. Kang, Y. J. Lee, D. Tang, M. Hüttemann and X. Song, Cell Death Differ., 2024, 31, 387–404 CrossRef CAS PubMed.
- P. E. Czabotar and A. J. Garcia-Saez, Nat. Rev. Mol. Cell Biol., 2023, 24, 732–748 CrossRef CAS PubMed.
- L. Lalier, F. Vallette and S. Manon, Biomolecules, 2022, 12, 162 CrossRef CAS PubMed.
- T. Ozaki and A. Nakagawara, Cancers, 2011, 3, 994–1013 CrossRef CAS PubMed.
- Y. Liu, Z. Su, O. Tavana and W. Gu, Cancer Cell, 2024, 42, 946–967 CrossRef CAS PubMed.
- M. Capuozzo, M. Santorsola, M. Bocchetti, F. Perri, M. Cascella, V. Granata, V. Celotto, O. Gualillo, A. M. Cossu, G. Nasti, M. Caraglia and A. Ottaiano, Biology, 2022, 11, 1325 CrossRef CAS PubMed.
- M. Wolosowicz, S. Prokopiuk and T. W. Kaminski, Int. J. Mol. Sci., 2024, 25, 13691 CrossRef CAS PubMed.
- D. Radzki, A. Negri, A. Kusiak and M. Obuchowski, Int. J. Mol. Sci., 2024, 25, 2763 CrossRef CAS PubMed.
- A. Nikolov and N. Popovski, Diagnostics, 2021, 11, 480 CrossRef CAS PubMed.
- I. Luchian, A. Goriuc, D. Sandu and M. Covasa, Int. J. Mol. Sci., 2022, 23, 1806 Search PubMed.
- H. Jiang and H. Li, BMC Cancer, 2021, 21, 149 CrossRef CAS PubMed.
- A. Shoari, A. Ashja Ardalan, A. M. Dimesa and M. A. Coban, Biomolecules, 2025, 15, 35 CrossRef CAS PubMed.
- J. Plati, O. Bucur and R. Khosravi-Far, J. Cell. Biochem., 2008, 104, 1124–1149 CrossRef CAS PubMed.
- C. M. Neophytou, I. P. Trougakos, N. Erin and P. Papageorgis, Cancers, 2021, 13, 4363 CrossRef CAS PubMed.
- V. Shoshan-Barmatz, T. Arif and A. Shteinfer-Kuzmine, Apoptosis, 2023, 28, 730–753 CrossRef PubMed.
- N. Malik and R. K. Singh, RSC Med. Chem., 2025, 16, 3344–3361 RSC.
- G. Latambale and K. Juvale, Mol. Divers., 2025, 29, 5185–5217 Search PubMed.
- Y. Guo, M. Cao, Z. Li, H. Zhou, Z. Chen and Q. J. A. d. P. Li, Arch. Pharm., 2025, 358 Search PubMed.
- S. Shaaban, S. S. Hawas, A. A. Elmaaty, H. Ba-Ghazal, M. Sharaky, M. Alaasar, A. M. Elsharif, A. O. Al Khatib, F. S. Alatawi, M. E. Eissa, H. M. Abd El-Lateef and A. A. Al-Karmalawy, Bioorg. Med. Chem., 2026, 136, 118610 CrossRef PubMed.
- N. S. Sethi, D. N. Prasad and R. K. Singh, Med. Chem. (Sharjah, United Arab Emirates), 2021, 17, 369–379 CAS.
- S. V. Laxmi, P. Anil, G. Rajitha, A. J. Rao, P. A. Crooks and B. Rajitha, J. Chem. Biol., 2016, 9, 97–106 CrossRef PubMed.
- H. El-Kashef, G. Badr, N. Abo El-Maali, D. Sayed, P. Melnyk, N. Lebegue and R. Abd El-Khalek, Bioorg. Chem., 2020, 96, 103569 CrossRef CAS PubMed.
- L. H. Al-Wahaibi, A. M. Elshamsy, T. F. Ali, B. G. Youssif, S. Bräse, M. Abdel-Aziz and N. A. El-Koussi, Front. Chem., 2025, 13, 1565699 CrossRef CAS PubMed.
- W. S. Hamama, G. G. El-Bana, S. Shaaban, O. Habib and H. H. Zoorob, RSC Adv., 2016, 6, 24010–24049 RSC.
- W. S. Hamama, M. A. Ismail, S. Shaaban and H. H. Zoorob, Med. Chem. Res., 2012, 21, 2615–2623 CrossRef CAS.
- S. Shaaban, M. A. Arafat and W. S. Hamama, Arkivoc, 2014, 2014, 470–505 Search PubMed.
- S. Shaaban, A. A. Nazeer, F. A. Azeez, H. S. R. Mohamed, T. A. Yousef and K. Shalabi, Colloids Surf., A, 2026, 732, 139218 CrossRef CAS.
- S. Shaaban, A. Y. M. Alabdali, M. H. Mousa, H. Ba-Ghazal, Y. S. Al-Faiyz, I. Elghamry, H. A. Althikrallah, A. O. A. Khatib, M. Alaasar and A. A. Al-Karmalawy, Drug Dev. Res., 2025, 86, e70075 CrossRef CAS PubMed.
- A. S. M. Al-Janabi, R. Zaky, T. A. Yousef, B. S. Nomi and S. Shaaban, J. Chin. Chem. Soc., 2020, 67(6), 1032–1044 CrossRef CAS.
- S. Shaaban, K. Shalabi, T. A. Yousef, M. Abou-Krisha, A. A. Alanazi, H. A. Althikrallah, M. Alaasar, A. M. Abu-Dief and A. S. Al-Janabi, J. Taiwan Inst. Chem. Eng., 2024, 165, 105766 CrossRef CAS.
- P. Skehan, R. Storeng, D. Scudiero, A. Monks, J. McMahon, D. Vistica, J. T. Warren, H. Bokesch, S. Kenney and M. R. Boyd, JNCI, J. Natl. Cancer Inst., 1990, 82, 1107–1112 CrossRef CAS PubMed.
- A. Scalbert, C. Morand, C. Manach and C. Rémésy, Biomed. Pharmacother., 2002, 56, 276–282 CrossRef CAS PubMed.
- M. Thanou, J. Verhoef and H. Junginger, Adv. Drug Deliv. Rev., 2001, 52, 117–126 CrossRef CAS PubMed.
- C. A. Lipinski, F. Lombardo, B. W. Dominy and P. J. Feeney, Adv. Drug Deliv. Rev., 1997, 23, 3–25 CrossRef CAS.
- D. D. Levy, E. Zeiger, P. A. Escobar, A. Hakura, M. Bas-jan, M. Kato, M. M. Moore and K.-i. Sugiyama, Mutat. Res., Genet. Toxicol. Environ. Mutagen., 2019, 848, 403074 CrossRef CAS PubMed.
- S. Roy and M. Mathew, J. Biol. Chem., 2018, 293, 4289–4303 CrossRef CAS PubMed.
- M. C. Sanguinetti, Curr. Opin. Pharmacol., 2014, 15, 22–27 CrossRef CAS PubMed.
- E. O. Olawode, R. Tandlich, E. Prinsloo, M. Isaacs, H. Hoppe, R. Seldon, D. F. Warner, V. Steenkamp and P. T. Kaye, ARKIVOC, 2018, 2018, 110–118 Search PubMed.
- G. Yan, L. Hao, Y. Niu, W. Huang, W. Wang, F. Xu, L. Liang, C. Wang, H. Jin and P. Xu, Eur. J. Med. Chem., 2017, 137, 462–475 CrossRef CAS PubMed.
- C. Nitsche, V. N. Schreier, M. A. Behnam, A. Kumar, R. Bartenschlager and C. D. Klein, J. Med. Chem., 2013, 56, 8389–8403 CrossRef CAS PubMed.
- M. M. Hammoud, M. Khattab, M. Abdel-Motaal, J. Van der Eycken, R. Alnajjar, H. S. Abulkhair and A. A. Al-Karmalawy, J. Biomol. Struct. Dynam., 2023, 41, 5199–5216 CAS.
- P. Skehan, R. Storeng, D. Scudiero, A. Monks, J. McMahon, D. Vistica, J. T. Warren, H. Bokesch, S. Kenney and M. R. Boyd, JNCI, J. Natl. Cancer Inst., 1990, 82, 1107–1112 CrossRef CAS PubMed.
- N. El-Gohary, S. Hawas, M. Gabr, M. Shaaban and M. El-Ashmawy, Bioorg. Chem., 2019, 92, 103109 CrossRef CAS PubMed.
- S. Shaaban, S. S. Hawas, A. M. Elsharif, M. Sharaky, H. Ba-Ghazal, M. Alaasar, F. S. Alatawi, K. B. Alomari, M. E. Eissa, A. O. Al Khatib, R. Alnajjar and A. A. Al-Karmalawy, RSC Adv., 2026, 16, 6539–6561 RSC.
- K. R. Senwar, T. S. Reddy, D. Thummuri, P. Sharma, V. G. M. Naidu, G. Srinivasulu and N. Shankaraiah, Eur. J. Med. Chem., 2016, 118, 34–46 CrossRef CAS PubMed.
- C. Brustolin Braga, J. C. Milan, M. Andrade Meirelles, B. Zavan, G. Á. Ferreira-Silva, E. S. Caixeta, M. Ionta and R. A. Pilli, RSC Med. Chem., 2024, 15, 3778–3794 RSC.
- V. P. de Souza, I. N. F. Gomes, S. J. Santos, C. B. Braga, A. J. A. Bezerra, E. A. Philot, C. R. N. Ramos, S. Q. Pantaleão, L. S. da Silva, R. A. Pilli, R. M. Reis, R. J. S. Oliveira and D. Russowsky, RSC Med. Chem., 2025, 16, 5989–6009 RSC.
- M. Wang, X. Zeng, S. Li, Z. Sun, J. Yu, C. Chen, X. Shen, W. Pan and H. Luo, Int. J. Mol. Sci., 2019, 20, 4459 CrossRef CAS PubMed.
- M. N. Sallam, A. A. Al-Karmalawy, E. M. Abbass, S. S. Hawas, A. M. El-Naggar and A. M. A. Hassan, RSC Adv., 2025, 15, 40078–40092 RSC.
- R. Huey, G. M. Morris and S. Forli, The Scripps Research Institute Molecular Graphics Laboratory, 2012, 10550, 1000 Search PubMed.
- S. Yuan, H. S. Chan and Z. Hu, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2017, 7, e1298 Search PubMed.
- T. Al-Warhi, M. M. Elbadawi, A. Bonardi, A. Nocentini, A. A. Al-Karmalawy, N. Aljaeed, O. J. Alotaibi, H. A. Abdel-Aziz, C. T. Supuran and W. M. Eldehna, J. Enzyme Inhib. Med. Chem., 2022, 37, 2635–2643 CrossRef CAS PubMed.
- R. M. Hazem, S. A. Antar, Y. K. Nafea, A. A. Al-Karmalawy, M. A. Saleh and M. F. El-Azab, Life Sci., 2022, 288, 120185 CrossRef CAS PubMed.
- A. Mahmoud, E. Kotb, A. I. Alqosaibi, A. A. Al-Karmalawy, I. S. Al-Dhuayan and H. Alabkari, Heliyon, 2021, 7, e08148 CrossRef CAS PubMed.
- A. Belal, M. A. Elanany, E. Y. Santali, A. A. Al-Karmalawy, M. O. Aboelez, A. H. Amin, M. H. Abdellattif, A. B. M. Mehany and H. Elkady, Molecules, 2022, 27, 3584 CrossRef CAS PubMed.
- D. B. Mahmoud, M. M. Bakr, A. A. Al-Karmalawy, Y. Moatasim, A. El Taweel and A. Mostafa, AAPS PharmSciTech, 2022, 23, 1–12 CrossRef PubMed.
- A. Daina, O. Michielin and V. Zoete, Sci. Rep., 2017, 7, 1–13 CrossRef PubMed.
- D. E. Pires, T. L. Blundell and D. B. Ascher, J. Med. Chem., 2015, 58, 4066–4072 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |






