 Open Access Article
Open Access ArticleRadiation-induced neurotoxicity: investigating human neuronal damage in MEA-integrated microfluidic platforms
Andie E. Padilla ,
Marion Jones
,
Marion Jones and
Binata Joddar
and
Binata Joddar *
*
Inspired Materials & Stem-Cell Based Tissue Engineering Laboratory, Chemical, Biological and Environmental Engineering, Oregon State University, 313 Gleeson Hall, 2115 SW Campus Way, Corvallis, OR 97331, USA. E-mail: binata.joddar@oregonstate.edu
First published on 19th March 2026
Abstract
In this study, we utilized a novel microelectrode array (MEA)-integrated 3D tissue-on-a-chip platform, developed by our team, to retain human glutamatergic neuronal networks and assess their functional response to 2.5 Gy and 5 Gy gamma radiation exposure. This platform enables real-time electrophysiology monitoring and mimics the human brain tissue 3D microenvironment more accurately than traditional 2D cultures. Cell viability analysis using live/dead staining in both the 3D microfluidic device and conventional 2D cultures revealed significant radiation-induced reductions in neuronal survival, with evidence of delayed onset of cell death following exposure. Longitudinal electrophysiological (EPHYS) measurements over a one week post-irradiation period demonstrated progressive deterioration of neuronal network performance, reflected by reduced firing frequency and decreased action potential amplitude. Biochemical assessments showed a marked downregulation of creatine kinase (CK), indicating impaired metabolic capacity in irradiated neurons. Additionally, DNA methylation analysis revealed radiation-associated alterations in epigenetic regulation, suggesting persistent molecular changes that may influence long-term neuronal function. Together, these findings highlight the susceptibility of human neuronal systems to ionizing radiation and demonstrate the value of microfluidic tissue-on-chip platforms for modeling extraterrestrial health risks. The combined decline in electrophysiological activity, metabolic integrity, and epigenetic stability provides key mechanistic insights into radiation-induced neurodegeneration and supports development of targeted countermeasures for astronaut health during deep-space missions.
1 Introduction
Human spaceflight beyond the low earth orbit (LEO) introduces astronauts to a wide array of stressors including microgravity and prolonged exposure to ionizing radiation. The complex environment of the solar system imparts an amalgamation of electromagnetic radiation and charged particles stemming from solar cosmic radiation, galactic cosmic rays, solar particle events, and trapped radiation belts.1 Although protons with energies less than 30 MeV are shielded by the spacecraft, the overall sum of charged particles dictates the dosage of radiation experienced and inflicted upon the vehicle and human body in relation.1,2 At the top of Earth's atmosphere, primary cosmic ray particles collide with atmospheric nuclei to form secondary particles, of which the primary mechanism of electromagnetic decay results in the emission of two photons, or gamma rays.3Gamma rays, characteristic of their ability to incite widespread cellular and molecular damage, are the highest form of electromagnetic radiation in the cosmos.4 Acute and chronic exposure to cosmic radiation is known to negatively impact neuronal and cardiac systems, prompting the formation of reactive oxygen species (ROS) and alterations to cell morphology and metabolic function.5,6 Gamma rays specifically have been shown to induce neural apoptosis and ROS formation, compounding the vulnerability of neuronal networks and facilitating amyloid-β accumulation, neuroinflammation, and cognitive decline.7–11
Current astronaut health protection strategies in space primarily focus on minimizing the duration of exposures, maximizing the distance between crew members and radiation sources, and shielding through strategic material selection for spacecraft and suits.2 However, the pervasive nature of space radiation and the high energy of certain particle types pose an extreme challenge towards fully managing and preventing exposure.2 As we progress toward long-duration missions, more robust health protection approaches will be necessary. The development of suitable and effective countermeasures, in turn, depends on a deeper understanding of the specific mechanistic changes space travel and radiation elicit within human physiological systems. Although significant efforts have been made to identify the physiological pathways influenced by spaceflight and radiation, the mechanisms through which ionizing radiation alters neuronal function remain poorly defined. This limitation stems partly from the ethical and practical challenges associated with studying live human tissues in space. Many investigations examining the effects of gamma radiation on neural cells rely on in vitro characterization using hippocampal brain slices derived from rats, mice, and other rodents.12–14 Such models, however, have limited relevance to true human physiology. To bridge this gap, engineered tissue systems that replicate the three-dimensional architecture and function of native human tissues present a promising alternative. These studies have shown that through utilization of 3D bioprinting techniques,15 self-organizing assembloids,16 and synthetic scaffold formation,17 we are able to construct a 3D environment which more closely mimics the behavior of neural tissues in comparison to 2D models. In doing so, we create a domain conducive to enhanced cell viability, differentiation, synaptic assembly, and network connectivity.
In this work, we employ a microfluidic device integrated with MEA technology18 to examine the effects of gamma radiation on the morphology and activity of human glutamatergic neurons. In prior studies, this platform was validated for its hardware functionality and its ability to sustain tissue viability throughout the study period included in this manuscript.18
This platform provides a physiologically relevant environment that more closely mimics native cellular functions while enabling non-invasive and real time electrophysiology (EPHYS) recordings from cultures exposed to ionizing radiation. By subjecting this model to gamma radiation, we aim to identify changes in neural network structure and function. These results may help determine the levels of ionizing radiation capable of causing significant neurotoxicity and support the use of this platform as an improved in vitro model for studying human neuronal responses to critical spaceflight-related conditions or other environmental stressors.
2 Results & discussion
Ionizing radiation has shown to increase oxidative stress, neuroinflammation, apoptosis, and amyloid-β accumulation in the human brain upon exposure.13,19–21 To gain a clearer understanding of these effects, specifically on the structural and functional properties of neuronal networks, we adopted a microfluidic platform which we previously developed and validated upon a suborbital flight (Fig. 1A).18 This device features a central hydrogel chamber interfaced with a Neuropixels MEA probe thereby enabling the real-time monitoring of spontaneous neuronal activity. Because a suborbital flight offers only minimal radiation exposure, we decided to conduct a rigorous follow-up study following our previously published work validating the device's hardware and its EPHYS recording capabilities.Here, we expand upon the capability of the device to capture EPHYS signals from glutamatergic neuronal networks using astrocyte co-culture and caffeine exposure treatments to further substantiate its use in assessing real time changes in human neuronal function. We then exposed these human glutamatergic neuron cultures in 2D to differing levels of gamma radiation (2.5 and 5 Gy) under Earth's gravity to study their dose responses from the cultures at 24 and 72 hours post radiation characterized by a live/dead viability assay and immunofluorescence staining. Then, neuronal networks cultured within the 3D microfluidic platform were exposed to varying doses of gamma radiation and subsequently evaluated using EPHYS, live/dead assays, creatine kinase (CK) assays, and DNA methylation analysis.
Due to the complexity of the study design, which involved multiple experimental steps and defining controls for each experiment, please refer to 3.1 to confirm the design and the corresponding controls for the experiments discussed below.
2.1 EPHYS validation
The capacity of the 3D microfluidic device to capture distinct changes in the firing activity of neurons was determined in the presence of human astrocytes (Fig. 1) and caffeine exposure (Fig. 2) respectively.Human glutamatergic neurons were co-cultured with human astrocytes (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) within a 3D microfluidic device for up to one week. In addition, human glutamatergic neurons alone and in the presence of astrocytes were cultured in a 2D MEA well-plate to serve as controls. Results showed an increase in total number of spikes, firing rate, and average spike amplitude in both co-cultures and in glutamatergic neurons alone (Fig. 1B, D and E) over the culture period in 3D. These findings are consistent with neuronal maturation and align with observations made in 2D MEA well-plate controls (Fig. 1C and F). Interestingly, the overall neural metrics recorded from the microfluidic device are much greater than what was observed in MEA well-plates (Fig. 1). That is, the total number of spikes in microfluidic cultures being 634, 1110, and 973 for astrocyte co-cultures and 153, 250, and 586 in glutamatergic-only cultures on days 1, 3, and 6 respectively. In comparison, MEA well-plates captured 0, 78, and 69 spikes in co-cultures and 0, 2, and 11 spikes in glutamatergic-only samples across the same time points. This validates the integration of electrodes within the 3D microfluidic device throughout the neuronal cultures, rather than at the surface alone, allowing for the capture of more robust activity deep within the neuronal networks. It is important to note, however, that the level of spontaneous activity of neuronal cultures can vary, even across biological replicates. SI Fig. S1 shows the difference in total number of spikes between at least two wells of an MEA well plate recorded simultaneously. This may account for variances observed in neural metrics across different experiments. Yet, our microfluidic platform still consistently captures EPHYS activity in greater magnitude compared to existing MEAs (Fig. 1 and 2). Additionally, EPHYS activity within both 3D and 2D MEA well-plate co-cultures was greater than glutamatergic neurons alone (Fig. 1). This suggests enhanced network formation in the presence of support cells such as astrocytes, in comparison to glutamatergic neurons alone.
1) within a 3D microfluidic device for up to one week. In addition, human glutamatergic neurons alone and in the presence of astrocytes were cultured in a 2D MEA well-plate to serve as controls. Results showed an increase in total number of spikes, firing rate, and average spike amplitude in both co-cultures and in glutamatergic neurons alone (Fig. 1B, D and E) over the culture period in 3D. These findings are consistent with neuronal maturation and align with observations made in 2D MEA well-plate controls (Fig. 1C and F). Interestingly, the overall neural metrics recorded from the microfluidic device are much greater than what was observed in MEA well-plates (Fig. 1). That is, the total number of spikes in microfluidic cultures being 634, 1110, and 973 for astrocyte co-cultures and 153, 250, and 586 in glutamatergic-only cultures on days 1, 3, and 6 respectively. In comparison, MEA well-plates captured 0, 78, and 69 spikes in co-cultures and 0, 2, and 11 spikes in glutamatergic-only samples across the same time points. This validates the integration of electrodes within the 3D microfluidic device throughout the neuronal cultures, rather than at the surface alone, allowing for the capture of more robust activity deep within the neuronal networks. It is important to note, however, that the level of spontaneous activity of neuronal cultures can vary, even across biological replicates. SI Fig. S1 shows the difference in total number of spikes between at least two wells of an MEA well plate recorded simultaneously. This may account for variances observed in neural metrics across different experiments. Yet, our microfluidic platform still consistently captures EPHYS activity in greater magnitude compared to existing MEAs (Fig. 1 and 2). Additionally, EPHYS activity within both 3D and 2D MEA well-plate co-cultures was greater than glutamatergic neurons alone (Fig. 1). This suggests enhanced network formation in the presence of support cells such as astrocytes, in comparison to glutamatergic neurons alone.
Caffeine has shown to lower the threshold for neuronal activation and increase spontaneous electrical activity in neurons.22 Therefore, a second validation study employing caffeine revealed the ability of the 3D microfluidic platform to capture EPHYS activity in direct response to external stimulation (Fig. 2). EPHYS activity was collected from 3D human glutamatergic neuron and astrocyte co-cultures (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) cultured within a microfluidic device for up to a week and dosed with a 100 µM caffeine solution on day 7. EPHYS acquisitions were performed prior to and immediately following the addition of caffeine. Results showed dramatic increases in all EPHYS metrics immediately upon exposure to caffeine in both microfluidic devices (Fig. 2A) and 2D MEA well-plate controls (Fig. 2B). In microfluidic devices, the number of spikes increased from 983 to 2815, mean spike amplitude from 240.16 µV to 323.27 µV, and frequency from 2.68 Hz to 11.49 Hz (Fig. 2A). In 2D MEA well-plates, the number of spikes rose from 79 to 131 and firing rate from 0.0069 Hz to 0.0114 Hz (Fig. 2B). Detection of these changes in both the microfluidic platform and 2D MEA well-plate controls confirms the capacity of the platform to accurately monitor real-time fluctuations in EPHYS activity.
1) cultured within a microfluidic device for up to a week and dosed with a 100 µM caffeine solution on day 7. EPHYS acquisitions were performed prior to and immediately following the addition of caffeine. Results showed dramatic increases in all EPHYS metrics immediately upon exposure to caffeine in both microfluidic devices (Fig. 2A) and 2D MEA well-plate controls (Fig. 2B). In microfluidic devices, the number of spikes increased from 983 to 2815, mean spike amplitude from 240.16 µV to 323.27 µV, and frequency from 2.68 Hz to 11.49 Hz (Fig. 2A). In 2D MEA well-plates, the number of spikes rose from 79 to 131 and firing rate from 0.0069 Hz to 0.0114 Hz (Fig. 2B). Detection of these changes in both the microfluidic platform and 2D MEA well-plate controls confirms the capacity of the platform to accurately monitor real-time fluctuations in EPHYS activity.
2.2 Gamma radiation dose response and time dependency
Following additional EPHYS validation of the 3D microfluidic device, we chose to study varying doses of gamma radiation applied to our neuronal tissue cultures in this project to determine dose dependent effects of radiation using a live/dead assay. In order to study the effects of space radiation on neuronal tissues in a ground-based experiment, gamma radiation exposures at doses of 2.5 Gy and 5 Gy were employed to model acute and chronic exposures and cultured for 24 and 72 hours, respectively, first in 2D cultures before optimizing those exposures for our 3D neuronal tissue chips. During space travel, it is estimated that astronauts are exposed to ionizing radiation doses of approximately 0.3–0.6 m Gy per day.23 Therefore, the total amount of radiation experienced is highly dependent upon mission duration and interaction with spacecraft materials. To understand the complete effects of radiation on central nervous system (CNS) function, however, more current studies focus on the effects of relatively low-dose exposure between 1–10 Gy.24 This study therefore focused on doses of 0, 2.5, and 5 Gy of gamma radiation.For this experiment, human glutamatergic neurons were mixed within Matrigel in a 35 mm Petri dish (100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–200
000–200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/12.5 µL Matrigel) and were referred to as 2D cultures or mixed within Matrigel at similar densities and introduced within the 3D microfluidic devices and were referred to as 3D cultures. Results showed similar viability across all groups at 24 hours following radiation exposure with 65.5%, 56.6%, and 59.2% in control, 2.5 Gy, and 5 Gy groups respectively (Fig. 3). After 72 hours, the number of viable cells in irradiated samples drastically decreased in the irradiated samples with 2.5 Gy showing 31.3% viability and 5 Gy showing 42.75% viability (Fig. 4). This delayed onset of cell death may suggest that the primary mechanisms driving apoptosis in response to radiation exposure involves more complex and time driven physiological changes within the cells. While longer study time points (∼72 hours) showed a significant reduction in cell viability for all radiation doses, the lasting impact of the 5 Gy dose for 24 hours, appeared most appropriate for studying the effects of gamma radiation on neuronal cultures in this study. Thus, we chose to further adopt the dose of 5 Gy for ∼24 hours for conducting other assays that are key to understanding the mechanisms involved in radiation-induced neuronal or non-neuronal tissue damage in human physiological systems prior to network-wide apoptosis. A viability assay performed on irradiated neuron cultures within a 3D microfluidic device (5 Gy gamma radiation, 24 hours) showed a decrease in viable cells (72.7%) compared to non-irradiated controls (85.54%) (Fig. 5), a difference that was statistically significant (p < 0.001, n = 3). Notably, within the irradiated group, 3D cultures exhibited slightly higher viability than 2D cultures at the same time point (Fig. 3), likely due to the enhanced network formation and protective microenvironment offered by 3D conditions.
000 cells/12.5 µL Matrigel) and were referred to as 2D cultures or mixed within Matrigel at similar densities and introduced within the 3D microfluidic devices and were referred to as 3D cultures. Results showed similar viability across all groups at 24 hours following radiation exposure with 65.5%, 56.6%, and 59.2% in control, 2.5 Gy, and 5 Gy groups respectively (Fig. 3). After 72 hours, the number of viable cells in irradiated samples drastically decreased in the irradiated samples with 2.5 Gy showing 31.3% viability and 5 Gy showing 42.75% viability (Fig. 4). This delayed onset of cell death may suggest that the primary mechanisms driving apoptosis in response to radiation exposure involves more complex and time driven physiological changes within the cells. While longer study time points (∼72 hours) showed a significant reduction in cell viability for all radiation doses, the lasting impact of the 5 Gy dose for 24 hours, appeared most appropriate for studying the effects of gamma radiation on neuronal cultures in this study. Thus, we chose to further adopt the dose of 5 Gy for ∼24 hours for conducting other assays that are key to understanding the mechanisms involved in radiation-induced neuronal or non-neuronal tissue damage in human physiological systems prior to network-wide apoptosis. A viability assay performed on irradiated neuron cultures within a 3D microfluidic device (5 Gy gamma radiation, 24 hours) showed a decrease in viable cells (72.7%) compared to non-irradiated controls (85.54%) (Fig. 5), a difference that was statistically significant (p < 0.001, n = 3). Notably, within the irradiated group, 3D cultures exhibited slightly higher viability than 2D cultures at the same time point (Fig. 3), likely due to the enhanced network formation and protective microenvironment offered by 3D conditions.
Although extended timepoints of 72 hours showed significant reductions in cell viability for all radiation doses (Fig. 4), the lasting impact of the 5 Gy dose for 24 hours demonstrated the most suitable balance for assessing the effects of gamma radiation in this study as it was enough to induce noticeable physiological changes in the cultures while still maintaining some network integrity needed to study functional changes. The apparent decrease in cell viability suggests that the selected radiation doses are adequate to trigger early metabolic responses which progress toward cell death at later timepoints. Since the goal of this work is to study functional changes of neuronal networks in response to radiation, subsequent analyses focused on earlier timepoints at these doses to examine the mechanistic consequences of radiation prior to extensive loss of viability. In doing so, we can observe radiation-induced stress response of the neuronal networks to gain a better perspective for potential countermeasure development.
Immunocytochemical analysis was performed on human GFP-glutamatergic neurons in order to assess changes in axon length in response to radiation exposure. These cells are already GFP-labelled and were additionally stained with microtubule associated protein 2 (MAP-2), a structural protein, expressed at the axons and dendrites of neurons25 and the nuclear stain DAPI. Changes to the expression of MAP-2 can help us visualize changes to network integrity and formation while GFP staining highlights overall cellular morphology, alongside DAPI, which pertains to the nucleus only. Immunostaining confirmed the presence of this marker in both control and irradiated samples in 2D wells and revealed physiological changes including evidence of axonal retraction in irradiated samples (Fig. 6), with a decrease of 20.05 ± 8.88 µM in axon length between radiation and control groups.
2.3 Gamma radiation response in human glutamatergic neurons
The functional responses of irradiated 3D neuronal networks were characterized using the EPHYS capabilities of the 3D microfluidic platform. EPHYS activity of the neuronal cultures was monitored for 7 days prior to 5 Gy gamma exposure. Spontaneous firing activity captured on day 4 (number of spikes = 1809, mean amplitude = 250.19 µV, and frequency = 1.97 Hz) suggests the presence of a healthy and functional network (Fig. 7). Increases in the number of spikes (n = 2579), mean amplitude (274.45 µV), and average frequency (3.52 Hz) were also observed with increasing time in culture which further implies appropriate maturation of the networks within the microfluidic platform. Following irradiation, on day 8, the cultures exhibit decreases across all neural metrics compared to pre-radiation acquisitions with number of spikes = 1930, mean amplitude = 258.66 µV, and average frequency = 2.86 Hz (Fig. 7), highlighting the deteriorative impact of gamma radiation on the EPHYS properties of human neuronal networks.Next, we assessed the Creatine Kinase (CK) metabolic activity levels of the neurons after being exposed to 5 Gy and cultured for up to ∼24 h. Although the assay kit we used can potentially detect all isoforms of the CK enzyme, the predominant form involved in this study was the CK-Brain-type isoenzyme or CK-BB, expressed by the neurons.26 In general, CK catalyzes the reversible transfer of a phosphate group from ATP to creatine to form phosphocreatine, which serves as an energy buffer ensuring a rapid ATP supply during high-demand processes such as synaptic transmission and neuronal firing. Thus, this system is crucial for maintaining neuronal energy metabolism, especially in cells with moderate metabolic activity such as those used in our study. On the other hand, CK is also expressed in fibroblasts, where it plays a role in energy metabolism however the isoform that could be involved would be distinctly different for the CK-BB isoform and may cross react with the CK-MB (cardiac) and the CK-MM (muscle) isoforms also present in the kit. Metabolic changes in response to gamma radiation were determined using a CK assay performed 24 hours following exposure in 2D cultures. Cardiac fibroblasts were employed as a positive control cell type. Decreases in the CK activity of both neurons (−1.70 ± 0.54 IU per µg protein) and fibroblast (−0.80 ± 0.21 IU per µg protein) cultures were observed in samples exposed to radiation (Fig. 8). Similarly, we observed slight decreases in the amount of methylated DNA in both neurons and fibroblasts (Fig. 9), with irradiated neurons exhibiting a mean of 0.009% 5 mC and irradiated fibroblasts exhibiting a mean of 0.0088% 5 mC. This suggests potential epigenetic alterations that could influence gene regulation and neuronal function, indicating that radiation exposure may disrupt DNA methylation homeostasis, a mechanism increasingly linked to neurodegeneration.
These findings align with published works which display declines in CK activity in samples exposed to radiation,27 and are confirmed by the disruptions in spontaneous EPHYS activity observed within our platform following radiation treatment. Downregulation in CK activity, associated with reduced protection against oxidative stress, has also shown to indicate central nervous system cell damage in neurodegenerative diseases such as Alzheimer's and Parkinson's disease.26,28 This further validates the use of this microfluidic device for studying functional changes in neuronal networks and suggests that the gamma radiation induces metabolic dysfunction in neurons which is tied to their signalling capacity and therefore may be linked to the development of neurodegenerative disorders downstream.
3 Materials and methods
3.1 3D and 2D EPHYS and other substrates used in the study
Data in this study were gathered from both 2D and 3D substrates. EPHYS data was collected from custom 3D microfluidic platforms, referred to as ‘microfluidic device’ or ‘3D cultures’, and Axion MEA 6-well plates, referred to as ‘2D MEA well-plate’. Networks cultured in 35 mm Petri dishes were employed for viability assays, CK assays, DNA methylation, and immunohistochemistry and are referred to as ‘2D cultures’.3.2 3D microfluidic device preparation
A microfluidic device previously described in Padilla et al.18 was adopted for the present study. Devices were cast in Ostemer 322 (Mercene Labs, Stockholm, Sweden) resin cast from a Sylgard 184 (Dow Chemical Company, Midland, MI, USA) mold and subsequently fitted with Microelectrode array (MEA) probes and bonded to a glass slide. Refer to next section for PEI coating procedures.3.3 Microelectrode array (MEA) preparation
All MEA substrates, 3D microfluidic devices and 2D MEA well-plates, used in this study were coated with polyethyleneimine (PEI) prior to use. This step has shown to promote attachment and of iPSC-derived cultures on MEA substrates for EPHYS characterization.29 Following sterilization, a 0.1% PEI solution was added to each culture vessel, microfluidic devices and Axion well plates, such that the solution was in contact with the entire culture surface. Vessels were incubated for 1 hour at 37 °C. The solution was then aspirated, and vessels were immediately washed 3 times with PBS. Vessels were then allowed to air dry overnight in a biosafety cabinet.3.4 Glutamatergic neuron cell culture
Human iCell Glutamatergic Neurons (Cellular Dynamics, Madison, WI, USA) or iXCell Glutamatergic Neurons (iXCells Biotechnologies, San Diego, California, USA) were thawed and resuspended in 9 mL complete maintenance medium prepared according to manufacturer protocol. The suspension was centrifuged at 400 rpm for 5 minutes and resuspended in 500 µL complete maintenance medium. A small volume of the suspension, totalling 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells, was transferred to a 1.5 mL Eppendorf tube and centrifuged at 400 rpm for 3 minutes. The supernatant was aspirated, and cells were resuspended in 5 µL complete maintenance medium. 12.5 µL Matrigel (Corning Inc., Corning, New York, USA) was added to the suspension and gently mixed by pipetting to achieve a density of 100
000 cells, was transferred to a 1.5 mL Eppendorf tube and centrifuged at 400 rpm for 3 minutes. The supernatant was aspirated, and cells were resuspended in 5 µL complete maintenance medium. 12.5 µL Matrigel (Corning Inc., Corning, New York, USA) was added to the suspension and gently mixed by pipetting to achieve a density of 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/12.5 µL Matrigel. For microfluidic devices, glutamatergic neurons were thawed as previously described and seeded at a density of 200
000 cells/12.5 µL Matrigel. For microfluidic devices, glutamatergic neurons were thawed as previously described and seeded at a density of 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/25 µL Matrigel to completely fill the hydrogel chamber. The Matrigel suspension was pipetted onto the culture dish and incubated at 37 °C for 30 minutes, after which, complete culture medium was added to each sample. iCell Glutamatergic Neurons were used to conduct EPHYS experimentation, including co-culture with astrocytes, caffeine exposure, radiation studies, and accompanying viability assays. iXCell Neurons were utilized in creatine kinase and DNA methylation assays, followed by immunocytochemical analysis.
000 cells/25 µL Matrigel to completely fill the hydrogel chamber. The Matrigel suspension was pipetted onto the culture dish and incubated at 37 °C for 30 minutes, after which, complete culture medium was added to each sample. iCell Glutamatergic Neurons were used to conduct EPHYS experimentation, including co-culture with astrocytes, caffeine exposure, radiation studies, and accompanying viability assays. iXCell Neurons were utilized in creatine kinase and DNA methylation assays, followed by immunocytochemical analysis.
3.5 Glutamatergic neuron–astrocyte co-culture
Human glutamatergic neurons (iCell®) and human astrocytes (iCell® Astrocytes 2.0) derived from iPSCs were purchased from Cellular Dynamics International (Madison, WI, USA). Human glutamatergic neurons were thawed at 37 °C for 2 minutes and resuspended in 9 mL complete BrainPhys maintenance medium (BrainPhys Neuronal Medium, iCell Neural Supplement B, iCell Nervous System Supplement, N-2 Supplement, Laminin, and Penicillin–Streptomycin). Cells were centrifuged at 400g for 5 minutes and resuspended in 1 mL complete BrainPhys Medium and Laminin [0.1 mg mL−1] in a labelled 15 mL falcon tube to achieve a density of 1.8 × 106 cells per mL. Cell counts are based on the vial certificate of analysis. Astrocytes were thawed and resuspended following the same procedure and placed in a separate 15 mL falcon tube to achieve a density of 1.5 × 106 cells per mL. Cells were seeded at a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 (astrocytes
6 (astrocytes![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) neurons) following manufacturer's recommendation. 2D MEA studies of astrocyte and neuron co-culture at this ratio showed to promote network formation.30 This was accomplished by transferring 111.11 µL of the neuron suspension and 22.22 µL suspension to a 1.5 mL Eppendorf tube. Cells were centrifuged at 400g for 3 minutes and resuspended in 25 µL Matrigel to achieve a final density of 200
neurons) following manufacturer's recommendation. 2D MEA studies of astrocyte and neuron co-culture at this ratio showed to promote network formation.30 This was accomplished by transferring 111.11 µL of the neuron suspension and 22.22 µL suspension to a 1.5 mL Eppendorf tube. Cells were centrifuged at 400g for 3 minutes and resuspended in 25 µL Matrigel to achieve a final density of 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/25 µL Matrigel at a ratio of 1
000 cells/25 µL Matrigel at a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6. The gel was pipetted into the hydrogel chamber of a PEI coated microfluidic device incubated for 30 minutes at 37 °C, 5% CO2. Control samples were pipetted onto the electrodes of a PEI coated Axion MEA 6-well plate (Axion Biosystems, Atlanta, GA, USA). A microfluidic device and 3 MEA wells were seeded with glutamatergic neurons only at a density of 200
6. The gel was pipetted into the hydrogel chamber of a PEI coated microfluidic device incubated for 30 minutes at 37 °C, 5% CO2. Control samples were pipetted onto the electrodes of a PEI coated Axion MEA 6-well plate (Axion Biosystems, Atlanta, GA, USA). A microfluidic device and 3 MEA wells were seeded with glutamatergic neurons only at a density of 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/25 µL Matrigel. A complete BrainPhys maintenance medium was then added to all devices. A 50% medium change was performed the next day and exchanged every 1–2 days thereafter.
000 cells/25 µL Matrigel. A complete BrainPhys maintenance medium was then added to all devices. A 50% medium change was performed the next day and exchanged every 1–2 days thereafter.
3.6 Fibroblasts
Human cardiac fibroblasts (HCF) obtained from Cell Applications Inc. (San Diego, CA, USA) from passage 10 were detached from a T-75 flask using Trypsin/EDTA for 3 minutes at 25 °C. The suspension was transferred to a 15 mL falcon tube and centrifuged for 5 minutes at 220×g. Cells were resuspended in 5 mL culture medium and plated at a density of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well of a 96 well plate. Samples were maintained at 37 °C for 7 days, with medium exchanges every 1–2 days.
000 cells per well of a 96 well plate. Samples were maintained at 37 °C for 7 days, with medium exchanges every 1–2 days.
3.7 Cell viability assay
A live/dead kit was purchased from Invitrogen (Gibco, CA, USA) and used according to the manufacturer's protocol. The assay was performed at 24 h and 72 h post-irradiation. Images were obtained using an Echo Revolve Fluorescence microscope (Discover Echo, USA) at 10× and 20× magnifications and were processed using ImageJ to quantify the number of live and dead cells in the image. Individual cells were selected by changing the image type to 8-bit, then using the ‘Threshold’ tool and adjusting the parameters until the areas of interest, or cells, were selected. These selected areas were then counted using the ‘Analyze Particles’ tool. % Cell viability was calculated by counting the number of live and dead cells in images captured from each sample respectively, then dividing this number by the total number of cells in the image and multiplied by 100.3.8 Electrophysiology from microfluidic device
Electrophysiology measurements from 3D microfluidic devices were obtained using a Neuropixels 1.0 probe with a metal cap (imec, Leuven, Belgium) integrated into the device. Data acquisition was performed using a OneBox (imec, Leuven, Belgium) operated by a Python-based code. A headstage and cable were connected to each probe at the time of recording. A wire connected to a grounded outlet was also inserted into the grounding ports of each device. Acquisitions were collected from banks 0 and 1, using a gain of 500 and an external reference. The external reference refers to grounding wires which are soldered on to allocated points on the probe prior to the fabrication step. These are connected to the same ground as the culture medium and OneBox during EPHYS collection. Data acquisitions began 24 hours after the seeding of cell-laden hydrogels into the microfluidic devices. Acquisitions were facilitated via a headstage, cable, and data collection system designed and developed by imec. These components are connected to microfluidic biodevices and provide the Neuropixels probe with a power supply, bi-directional communication capabilities, and data transmission to a PC.183.9 Electrophysiology from 2D MEA well plate
2D Electrophysiology measurements were captured using an (Axion Maestro Edge Axion Biosystems, Atlanta, GA, USA). iCell human glutamatergic neurons were seeded onto an Axion MEA 6-well plate (Axion Biosystems, Atlanta, GA, USA) following the protocol described in Section 3.5. After 24 hours in culture, the well plates were placed into the Maestro Edge and data was collected using the Axis Navigator software. Spontaneous neural recordings were captured for 60 seconds using the default program settings. The resulting data was then exported and processed using the Neural Metrics software developed by Axion.3.10 Caffeine exposure
Human glutamatergic neurons and astrocytes were seeded in a microfluidic device following the protocol described in Section 3.5. Human glutamatergic neurons alone were seeded within a separate microfluidic to serve as controls. Samples were maintained for 7 days at 37 °C and 5% CO2, with medium exchanges performed every other day. On day 7, EPHYS acquisitions were performed on each sample for 60 seconds. A 100 µM caffeine solution was prepared from Caffeine powder (99%) (ThermoFisher Scientific, Waltham, MA USA) diluted in MiliQ water. Culture medium was removed from samples immediately following EPHYS acquisitions and replaced with the caffeine solution. A second EPHYS acquisition was then performed immediately to assess functional changes.3.11 Gamma radiation exposure
Samples were subjected to 2.5 Gy and 5 Gy gamma radiation using a Gammacell 220 Cobalt-60 gamma irradiator (Nordion, Ottawa, Ontario, Canada) at Oregon State University. The exposure rate of this machine is 508![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 455 R h−1 (4459 Gy h−1). To achieve radiation levels of 2.5 Gy and 5 Gy, samples were exposed for roughly 2.02 seconds and 4.04 seconds respectively. The samples were transported to our laboratory for analysis immediately following radiation exposure.
455 R h−1 (4459 Gy h−1). To achieve radiation levels of 2.5 Gy and 5 Gy, samples were exposed for roughly 2.02 seconds and 4.04 seconds respectively. The samples were transported to our laboratory for analysis immediately following radiation exposure.
3.12 Immunocytochemistry (IHC)
Human glutamatergic neurons (iXCell) in Matrigel were seeded and maintained at a density of 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells/25 µL for 7 days. Cells were fixed using 4% paraformaldehyde (ThermoFisher Scientific, Waltham, MA) for 30 min. Samples were washed and blocked using 1× PBS, 5% BSA, and 0.3% Triton-X solution for 1 h. Immunocytochemistry was performed using MAP-2 Monoclonal Antibody (M13) as a primary antibody (ThermoFisher Scientific, Waltham, MA) diluted in 1× PBS, 1% BSA, and 0.3% Triton-X at a ratio of 1
000 cells/25 µL for 7 days. Cells were fixed using 4% paraformaldehyde (ThermoFisher Scientific, Waltham, MA) for 30 min. Samples were washed and blocked using 1× PBS, 5% BSA, and 0.3% Triton-X solution for 1 h. Immunocytochemistry was performed using MAP-2 Monoclonal Antibody (M13) as a primary antibody (ThermoFisher Scientific, Waltham, MA) diluted in 1× PBS, 1% BSA, and 0.3% Triton-X at a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400. Samples were incubated overnight at 4 °C. Alexa Fluor Plus 647 (Invitrogen, USA) was used as a secondary antibody diluted to 1
400. Samples were incubated overnight at 4 °C. Alexa Fluor Plus 647 (Invitrogen, USA) was used as a secondary antibody diluted to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400. Secondary antibodies were incubated for 1 h at 25 °C, protected from light. Samples were washed, and DAPI Fluoromount G (Southern Biotech, Birmingham, AL, USA) was applied. Images were acquired using an Echo Revolve Fluorescence microscope (Discover Echo, USA).
400. Secondary antibodies were incubated for 1 h at 25 °C, protected from light. Samples were washed, and DAPI Fluoromount G (Southern Biotech, Birmingham, AL, USA) was applied. Images were acquired using an Echo Revolve Fluorescence microscope (Discover Echo, USA).
3.13 Creatine kinase assay
A CK Activity Assay Kit was purchased from Millipore Sigma (Burlington, MA, USA). Cells were collected from samples following exposure to 2.5 or 5 Gy radiation and homogenized in cold 50 mM potassium phosphate buffer. Samples were centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 15 minutes at 4 °C and stored at −80 °C until the time of assay. Samples were brought to 25 °C at the time of assay. The CK assay was performed in a 96-well plate according to manufacturer protocol. CK activity was calculated for each sample by dividing the average 5 minutes and initial absorbance values by the difference between calibrator and blank absorbance values and multiplying the dividend by the equivalent activity of the blank and calibrator. Data were then normalized to average protein expression from each sample, measured using a Quick Start Bradford Protein Assay (Bio-Rad Laboratories, Fort Worth, TX, USA). The equation used (eqn (1)) for this calculation is as follows:
000g for 15 minutes at 4 °C and stored at −80 °C until the time of assay. Samples were brought to 25 °C at the time of assay. The CK assay was performed in a 96-well plate according to manufacturer protocol. CK activity was calculated for each sample by dividing the average 5 minutes and initial absorbance values by the difference between calibrator and blank absorbance values and multiplying the dividend by the equivalent activity of the blank and calibrator. Data were then normalized to average protein expression from each sample, measured using a Quick Start Bradford Protein Assay (Bio-Rad Laboratories, Fort Worth, TX, USA). The equation used (eqn (1)) for this calculation is as follows:
 | (1) |
3.14 DNA methylation
DNA was extracted from cell cultures 24 hours following irradiation using the FitAmp Blood and Cultured Cell DNA Extraction Kit (EpiGentek, Farmingdale, NY, USA). DNA methylation was then performed using the MethylFlash Global DNA Methylation (5 mC) ELISA Easy Kit (Colorimetric) (EpiGentek, Farmingdale, NY, USA). Input and control DNA were normalized to 25 ng. 5 mC (%) was calculated following a generated logarithmic regression equation based on the positive control standard curve obtained at the time of assay. The linear regression model equation sets (eqn (2) and (3)) used for this calculation were obtained from the manufacturer protocol and are as follows:
Y = 0.422![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(x) + 2.6699 ln(x) + 2.6699
| (2) |
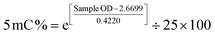 | (3) |
3.15 Data analysis
Data are expressed as mean ± SD and were analyzed using a Welch's t-test or one-way analysis of variance (ANOVA) using GraphPad Prism 10 software. Statistical significance was set at p ≤ 0.05 for all analyses.4 Conclusions
Long-duration space travel, particularly missions beyond low Earth orbit (LEO), exposes astronauts to a complex array of environmental stressors, including microgravity and ionizing radiation. Among these, exposure to cosmic radiation including galactic cosmic rays (GCRs), solar particle events (SPEs), and secondary radiation poses significant risks to human physiology, especially the central nervous system.31 Ionizing radiation has been shown to adversely affect sensitive neuronal structures, leading to the generation of reactive oxygen species (ROS), accumulation of amyloid-β peptides, neuroinflammation, and alterations in cellular morphology and metabolic activity. These changes are associated with cognitive impairments and increased risk of neurodegenerative conditions, raising concerns for astronaut health during extended space missions.32,33In our previously published works, we developed a neuronal tissue-on-a-chip electrophysiology sensing platform that was tested and validated for its robustness and function aboard a suborbital space flight. Since space vehicles have a sufficient amount of anti-radiation hardware and protection built in, this system may not have captured the effects of sustained LEO radiation on our tissue culture microfluidic chip system. Thus, a follow-up investigation and study were initiated wherein human glutamatergic neurons were encapsulated within the same neuronal tissue-on-a-chip reported by our group before.18
In this study, we again subjected the platform to a few other functional studies, such as the addition of caffeine to explore its effects on neuronal firing and the addition of astrocytes to modulate the amplitude and frequency of firing of the neurons. After being successfully validated, the system was now ready to be exposed for radiation studies.
Cell viability was not found to be significantly different among cultured neurons in the 3D microfluidic device and in 3D well plates after being exposed to 5 Gy and cultured for up to ∼24 hours. This highlights the use of this device for culturing healthy neuronal networks and further strengthened the premise in the utilization of the neuronal tissue chip platform for the remainder of this study. Similarly, microtubule-associated protein 2 (MAP-2) expression was significantly reduced in this set of neurons after being exposed to 5 Gy and cultured for up to ∼24 hours. Since MAP-2 plays a critical role in stabilizing microtubules and maintaining dendritic architecture, its absence or reduction in expression correlated with unhealthy or distressed neurons.34 Other studies have indicated that MAP-2 disruption may be a marker of excitotoxicity.34
Interestingly, the exposure of neurons and fibroblasts to 5 Gy and cultured for up to ∼24 hours, both showed a slight reduction in CK enzyme expression levels. This confirmed the changes in both the activity and the biosynthesis of the CK enzyme, which serves as the basis for the use of analysis of the activity of enzymes for evaluation of the effect of ionizing radiation and various radioprotectors, as also disseminated by others.35,36 It is well known that exposures to ionizing radiation such as Gamma has a direct effect on cells, causing breaks in chemical bonds in macromolecules, including enzyme proteins such as CK, and indirect effect, which can be further studies and determined by the formation of free radicals that aggressively interact with macromolecules.27 Prior studies have also shown that CK is the most radiosensitive enzyme,27 and that in similar reports an initial reduction was witnessed by others followed by radiation exposures which matches and validates our findings as well.
In parallel we also assessed DNA methylation which is an epigenetic modification involving the addition of a methyl group to the 5′ position of cytosine residues, typically within CpG dinucleotides in neurons after being exposed to 5 Gy and cultured for up to ∼24 h. A methylation level of 5 mC (%) indicates that approximately 5% of cytosines in the analyzed region are methylated.37 Thus, quantifying methylation percentages was critical for understanding epigenetic regulation under different radiation exposure conditions. Our findings indicated that, under the experimental conditions applied, the neuronal cultures maintained a permissive epigenetic landscape rather than undergoing widespread transcriptional silencing. Such low methylation levels may reflect the inherent plasticity of neuronal cells or a stress-adaptive response to gamma radiation exposure. These results underscore the importance of monitoring methylation dynamics as a sensitive indicator of cellular state and highlight the potential role of epigenetic regulation in mediating neuronal resilience or vulnerability under space radiation analog conditions.
Radiation exposure triggers profound mechanistic changes in neuronal and non-neuronal cell types, initiating a cascade of pathological events. In neurons, early disruptions include excessive calcium influx,38 mitochondrial dysfunction, and oxidative stress, which impair ATP production and destabilize cellular homeostasis as evidenced by the altered CK and 5 mC (%) expression. These changes activate apoptotic and necrotic pathways, leading to progressive neuronal death.27,39 Other non-neuronal cell types also undergo inflammatory activation, releasing cytokines and reactive oxygen species that exacerbate tissue damage, as was the case with fibroblasts used as a control cell type in our study. Collectively, these processes amplify neurotoxicity and impair synaptic connectivity, resulting in widespread functional deficits. Critically, a therapeutic window exists within the first 24 hours post-exposure, as shown by our study, during which timely administration of countermeasures can reverse mitochondrial dysfunction, reduce oxidative stress, and suppress inflammatory cascades, thereby promoting cellular recovery. However, if intervention is delayed beyond 72 hours, the damage becomes irreversible due to extensive cell death and systemic failure, ultimately leading to fatal outcomes.
Author contributions
Andie Padilla: conceptualization, methodology, software, data curation, writing – original draft preparation, visualization, investigation, formal data analysis Marion Jones: methodology (CK assay), investigation (CK assay) Binata Joddar: supervision, funding acquisition, project administration, writing – reviewing and editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
All data generated in this study are included as figures, graphs, and tables in Fig. 1–9 and supplementary information (SI) S1. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra01084a.Acknowledgements
The authors would like to acknowledge Hazel Beatty-Witt for her contribution to immunohistochemistry image processing and characterization. This research was supported by grants from the National Science Foundation (# 2515123, # 2511985, # 2515120) awarded to BJ. We gratefully acknowledge the start-up package offered to BJ from OSU for this work. BJ also acknowledges the Valley Health Foundation fellowship for this effort.References
- G. A. Nelson, Space Radiation and Human Exposures, A Primer, Radiat. Res., 2016, 185, 349–358 CrossRef CAS PubMed.
- K. D. Held, Space Radiation: An Overview, in Handbook of Bioastronautics, Springer International Publishing, Cham, 2021, pp. 257–262, DOI:10.1007/978-3-319-12191-8_131.
- C. Navia and M. N. D. Oliveira, Gamma Rays from Space, in New Insights on Gamma Rays, InTech, 2017, DOI:10.5772/67176.
- R. C. Hartman, et al., Galactic plane gamma-radiation, Astrophys. J., 1979, 230, 597 CrossRef CAS.
- K. Kandarpa, V. Schneider and K. Ganapathy, Human health during space travel: An overview, Neurol. India, 2019, 67, 176 CrossRef PubMed.
- G. Pani, et al., Combined Exposure to Simulated Microgravity and Acute or Chronic Radiation Reduces Neuronal Network Integrity and Survival, PLoS One, 2016, 11, e0155260 CrossRef PubMed.
- M.-S. Song, et al., Gamma-Ray Susceptibility of Immature and Mature Hippocampal Cultured Cells, J. Vet. Med. Sci., 2010, 72, 605–609 CrossRef CAS PubMed.
- W. Sudprasert, P. Navasumrit and M. Ruchirawat, Effects of low-dose gamma radiation on DNA damage, chromosomal aberration and expression of repair genes in human blood cells, Int. J. Hyg Environ. Health, 2006, 209, 503–511 CrossRef CAS PubMed.
- A. Graupner, et al., Gamma radiation at a human relevant low dose rate is genotoxic in mice, Sci. Rep., 2016, 6, 32977 CrossRef CAS PubMed.
- M. Yang, et al., Cytotoxicity of gamma-ray in rat immature hippocampal neurons, J. Vet. Sci., 2011, 12, 203 CrossRef PubMed.
- M. Roy-O’Reilly, A. Mulavara and T. Williams, A review of alterations to the brain during spaceflight and the potential relevance to crew in long-duration space exploration, npj Microgravity, 2021, 7, 5 CrossRef PubMed.
- P. H. Wu, et al., Radiation Induces Acute Alterations in Neuronal Function, PLoS One, 2012, 7, e37677 CrossRef CAS PubMed.
- T. C. Pellmar and D. L. Lepinski, Gamma radiation (5-10 Gy) impairs neuronal function in the guinea pig hippocampus, Radiat. Res., 1993, 136, 255–261 CrossRef CAS PubMed.
- B. P. Tseng, et al., Characterizing low dose and dose rate effects in rodent and human neural stem cells exposed to proton and gamma irradiation, Redox Biol., 2013, 1, 153–162 CrossRef CAS PubMed.
- Y. Yan, et al., 3D bioprinting of human neural tissues with functional connectivity, Cell Stem Cell, 2024, 31, 260–274 CrossRef CAS PubMed.
- Y. Miura, et al., Engineering brain assembloids to interrogate human neural circuits, Nat. Protoc., 2022, 17, 15–35 CrossRef CAS PubMed.
- J. M. Galindo, et al., Optimization of 3D Synthetic Scaffolds for Neuronal Tissue Engineering Applications, Chem.–Eur. J., 2024, 30, e202302481 CrossRef CAS PubMed.
- A. E. Padilla, et al., Adoption of microfluidic MEA technology for electrophysiology of 3D neuronal networks exposed to suborbital conditions, npj Microgravity, 2025, 11, 20 CrossRef CAS PubMed.
- F. R. Algeda, N. A. Eltahawy, S. M. Shedid and H. N. Saada, The impact of gamma-radiation on the cerebral- and cerebellar- cortex of male rats' brain, Brain Res. Bull., 2022, 186, 136–142 CrossRef CAS PubMed.
- A. Nakkazi, D. Forster, G. A. Whitfield, D. P. Dyer and B. R. Dickie, A systematic review of normal tissue neurovascular unit damage following brain irradiation—Factors affecting damage severity and timing of effects, Neuro-Oncol. Adv., 2024, 6, vdae098 CrossRef PubMed.
- R. Soffietti, A. Pellerino, F. Bruno, A. Mauro and R. Rudà, Neurotoxicity from Old and New Radiation Treatments for Brain Tumors, Int. J. Mol. Sci., 2023, 24, 10669 CrossRef CAS PubMed.
- C. Walton, J. M. Kalmar and E. Cafarelli, Effect of caffeine on self-sustained firing in human motor units, J. Physiol., 2002, 545, 671–679 CrossRef CAS PubMed.
- L. Strigari, S. Strolin, A. G. Morganti and A. Bartoloni, Dose-Effects Models for Space Radiobiology: An Overview on Dose-Effect Relationships, Front. Public Health, 2021, 9, 733337 CrossRef PubMed.
- B. P. Tseng, et al., Characterizing low dose and dose rate effects in rodent and human neural stem cells exposed to proton and gamma irradiation, Redox Biol., 2013, 1, 153–162 CrossRef CAS PubMed.
- R. F. Halliwell, H. Salmanzadeh, L. Coyne and W. S. Cao, An Electrophysiological and Pharmacological Study of the Properties of Human iPSC-Derived Neurons for Drug Discovery, Cells, 2021, 10, 1953 CrossRef CAS PubMed.
- T. S. Bürklen, et al., The Creatine Kinase/Creatine Connection to Alzheimer′s Disease: CK Inactivation, APP-CK Complexes and Focal Creatine Deposits, BioMed Res. Int., 2006, 035936 CrossRef PubMed.
- M. S. Petrosyan, L. S. Nersesova, N. A. Adamyan, M. G. Gazaryants and Z. I. Akopyan, The Effect of Ionizing Radiation on the Creatine–Creatine Kinase System in the Rat Brain and the Radioprotective Effect of Creatine, Neurochem. J., 2019, 13, 295–301 CrossRef CAS.
- M. Aksenov, M. Aksenova, D. A. Butterfield and W. R. Markesbery, Oxidative Modification of Creatine Kinase BB in Alzheimer's Disease Brain, J. Neurochem., 2000, 74, 2520–2527 CrossRef CAS PubMed.
- M. Yang, et al., Polyethyleneimine facilitates the growth and electrophysiological characterization of iPSC-derived motor neurons, Sci. Rep., 2024, 14, 26106 CrossRef CAS PubMed.
- Fujifilm, ICell ® Astrocytes 2.0 User's Guide, 2023, https://www.fujifilmcdi.com/wp/wp-content/uploads/2023/09/FCDI_iCellAstrocytes-2.0_UG_31Aug2023_FINAL.pdf.
- V. K. Parihar, et al., Cosmic radiation exposure and persistent cognitive dysfunction, Sci. Rep., 2016, 6, 34774 CrossRef CAS PubMed.
- N. Begum, B. Wang, M. Mori and G. Vares, Does ionizing radiation influence Alzheimer's disease risk?, J. Radiat. Res., 2012, 53, 815–822 CrossRef CAS PubMed.
- K. Krukowski, et al., The impact of deep space radiation on cognitive performance: From biological sex to biomarkers to countermeasures, Sci. Adv., 2021, 7, eabg6702 CrossRef CAS PubMed.
- C. Arias, I. Arrieta, L. Massieu and R. Tapia, Neuronal damage and MAP2 changes induced by the glutamate transport inhibitor dihydrokainate and by kainate in rat hippocampus in vivo, Exp. Brain Res., 1997, 116, 467–476 CrossRef CAS PubMed.
- G. N. Catravas and C. G. McHale, Changed activities of brain enzymes involved in neurotransmitter metabolism in rats exposed to different qualities of ionizing radiation, J. Neurochem., 1975, 24, 673–676 CrossRef CAS PubMed.
- Y. S. T, et al., Radioprotective effects of diallyl disulphide and Carica papaya (L.) leaf extract in electron beam radiation induced hematopoietic suppression, Cogent Biol., 2016, 2, 1247607 CrossRef.
- EpigenTek, MethylFlash™ Global DNA Methylation (5-mC) ELISA Easy Kit (Colorimetric) User Guide Search PubMed.
- N. Samari, et al., Non-conventional apoptotic response to ionising radiation mediated by N-methyl D-aspartate receptors in immature neuronal cells, Int. J. Mol. Med., 2013, 31, 516–524 CrossRef CAS PubMed.
- D. Averbeck and C. Rodriguez-Lafrasse, Role of Mitochondria in Radiation Responses: Epigenetic, Metabolic, and Signaling Impacts, Int. J. Mol. Sci., 2021, 22, 11047 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |









