Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
New isoxazole-based heterocyclic hybrids with dual antimicrobial and antioxidant bioactivity: integrated synthesis, in vitro assessment, and computational exploration
Aziz Arzinea,
Soumia Ait Assoub,
Lamiae El Bouamri c,
Mohammed Chalkha
c,
Mohammed Chalkha *ad,
Asmae Nakkabiad,
Samir Chtita
*ad,
Asmae Nakkabiad,
Samir Chtita c,
Reem M. Aljowaiee,
Mourad A. M. Aboul-Soud
c,
Reem M. Aljowaiee,
Mourad A. M. Aboul-Soud *f,
Mohammed El Hassouni
*f,
Mohammed El Hassouni b,
John P. Giesy
b,
John P. Giesy ghi and
Mohamed El Yazidi
ghi and
Mohamed El Yazidi a
a
aEngineering Laboratory of Organometallic, Molecular Materials and Environment, Faculty of Sciences Dhar EL Mahraz, Sidi Mohamed Ben Abdellah University, P.O. Box 1796 (Atlas), 30000 Fez, Morocco. E-mail: arzineaziz@gmail.com; mohammed.chalkha1@usmba.ac.ma
bBiotechnology, Environment, Agri-Food and Health Laboratory, Faculty of Sciences Dhar El Mahraz, Sidi Mohamed Ben Abdellah University, BP. 1796, Atlas, Fez, Morocco
cLaboratory of Analytical and Molecular Chemistry, Faculty of Sciences Ben M'Sik, Hassan II University of Casablanca, Casablanca B.P 7955, Morocco
dLaboratory of Materials Engineering for the Environment and Natural Resources, Faculty of Sciences and Techniques, University of Moulay Ismail of Meknes, B.P 509, Boutalamine, 52000, Errachidia, Morocco
eDepartment of Botany and Microbiology, College of Science, King Saud University, P.O. Box 2455, Riyadh 11451, Saudi Arabia
fCenter of Excellence in Biotechnology Research, College of Applied Medical Sciences, King Saud University, Riyadh 11433, Saudi Arabia. E-mail: maboulsoud@ksu.edu.sa
gDepartment of Veterinary Biomedical Sciences, Toxicology Centre, Western College of Veterinary Medicine, University of Saskatchewan, Saskatoon, SK S7N 5B4, Canada
hDepartment of Integrative Biology, Center for Integrative Toxicology, Michigan State University, East Lansing, MI 48824, USA
iDepartment of Environmental Sciences, Baylor University, Waco, 76706, USA
First published on 10th March 2026
Abstract
In the present study, novel isoxazole-based hybrid compounds 5a–h were synthesized with satisfactory yields. Their structure was confirmed by FT-IR, NMR (1H, 13C, 2D), and HRMS. The antimicrobial properties of this class of compounds were thoroughly investigated in vitro against a variety of Gram-positive and Gram-negative bacteria as well as different fungi, both yeast and molds. Their antioxidant ability was also assessed by molybdate reduction assay. The compounds showed excellent antifungal activity, particularly against Aspergillus niger and Fusarium oxysporum, equivalent to that of fluconazole. Among the series, 5d was the most potent antibacterial agent against Escherichia coli (minimal inhibitory concentration, MIC = 2.582 µmol mL−1), whereas the highest potency against Bacillus subtilis was found for 5h (MIC = 0.083 µmol mL−1) which was comparable to the efficacy of ampicillin. With respect to the antifungal activity, 5h showed the lowest MIC value against Candida albicans (MIC = 0.083 µmol mL−1) and A. niger (MIC = 0.083 µmol mL−1), and 5g was the most effective against Aspergillus flavus (MIC = 0.044 µmol mL−1). All hybrids were more effective as compared to fluconazole against F. oxysporum (MIC range: 0.43–0.94 µmol mL−1). Furthermore, assessment of their antioxidant potential shows that compounds 5e and 5g exhibit excellent reducing potency. Molecular docking and dynamic simulations demonstrated that compounds 5a–h, especially 5d, interact strongly and stably with key bacterial and fungal proteins, forming hydrogen bonds, hydrophobic interactions, and water bridges, with minimal root-mean-square fluctuations (RMSF), confirming the structural integrity of the complexes. Additionally, in silico predictions of Absorption, Distribution, Metabolism, Excretion, and Toxicity (ADME-Tox) analysis predicted 5d to possess an optimal profile, with high gastrointestinal absorption and minimal toxicity, highlighting it as the lead candidate for the development of broad-spectrum antimicrobial drugs.
1. Introduction
Nitrogen-containing heterocycles represent a significant class of bioactive compounds that play a fundamental role in the medicinal chemistry field.1–3 Among these, azoles constitute a subclass of five-membered aromatic heterocycles, comprising at least one nitrogen atom and often one or more additional heteroatoms, such as oxygen or sulfur.4–7The isoxazole pharmacophore, a five membered heterocyclic ring containing two adjacent heteroatoms (oxygen and nitrogen), plays a central role in many pharmaceutical agents.8–10 The importance of this structural element is highlighted in its role as a key mediator in the synthesis of several biologically active molecules.11–18 At the molecular level, the isoxazole residue can establish various non-covalent interactions, including nitrogen and oxygen hydrogen bonds, pi–pi, and hydrophobic interactions (cLogP = 0.334 at pH = 7.4).19 These properties improve isoxazole's extensive pharmacological applications, as shown by its incorporation into various therapeutic drugs, including β-lactam antibiotics (cloxacillin and dicloxacillin) and antibacterial sulfonamides (sulfisoxazole and sulfamethoxazole), as well as anti-inflammatory agents such as valdecoxib (a selective COX-II inhibitor) and leflunomide (a disease-modifying immunosuppressant (DMARD)).6,20–24 The isoxazole heterocycle has been found in numerous biologically active natural and synthetic substances, including the drugs cycloserine, acivicine, and muscimol. These compounds were originally obtained from microorganisms, higher plants and marine sponges.25–27 In addition, a wide range of therapeutic potential of isoxazole derivatives has also been reported, including antifungal,28,29 antiviral,30,31 antihistamine,32 antimicrobial,11,13–15 antioxidant,16,33 anticancer,17,34 and anti-inflammatory35,36 effects. Apart from their importance in medicine, these compounds are widely used in agriculture and industry,8 as insecticides,37 herbicides,38 fungicides,39 and as anti-corrosion coating agents.40 This broad-spectrum of potential applications highlights the isoxazole nucleus as a highly promising component at the interface between organic chemistry, biology and materials science.
Molecule hybridization is, currently, considered as one of the most innovative and rational approaches for designing new chemical entities exhibiting multiple therapeutic applications.41–43 This hybridization strategy involves designing a substance that contains two or more pharmacophores from different compounds, with each pharmacophore contributing to specific biological effects or physicochemical proprieties.44–46 This kind of combination has been found to enhance the biological activity of these molecules, as well as their selectivity and pharmacokinetic profile. Such combination has also been demonstrated to attenuate side effects and drug resistance.44,47,48 Given these considerations, the isoxazole moiety was considered as a crucial bioactive scaffold in developing new hybrid molecules. To increase their activity and to the investigate the formation of new synergistic effects.12,14 In the last decades, many hybrids have been synthesized and investigated, in which a variety of bioactive moieties were linked including isoxazole–(iso)oxazole types standing out as a significant example of heterocyclic hybrids. The molecules obtained through these hybridizations have demonstrated various pharmacological activities, including anticancer, antibacterial, anti-inflammatory, immunosuppressive, antitubercular, as well as antiviral and antifungal activities.14,41,49–51
In the present study, an efficient synthesis is described for developing new hybrids that contain di- and trisubstituted isoxazole moieties. This approach is based on the idea that the pharmacophoric elements act synergistically leading to molecules that display improved biologically effect. The newly synthesized hybrids were subjected to a combined assessment of their antioxidant and antimicrobial activity, in vitro, and the results indicated encouraging findings. Concurrently, a series of in silico studies, including molecular docking, ADME-Tox and drug-likeness prediction, and molecular dynamics simulation (MDS) were conducted to support the results obtained from the, in vitro, experiments tests. Structure-based approaches have been used to study binding interactions, evaluate pharmacokinetic features, and verify the stability of the subsequent ligand–protein complexes. Consequently, the experimental and theoretical investigations provide deeper understanding into the modification of new isoxazole–isoxazole hybrids.
2. Results and discussion
2.1. Chemistry
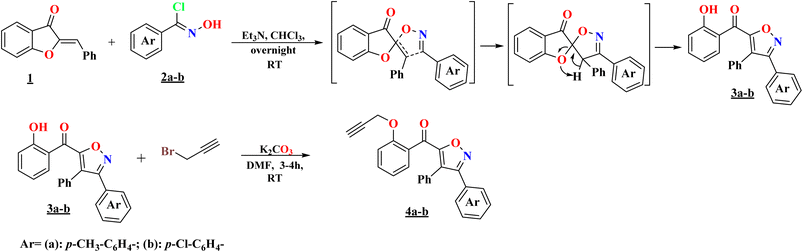
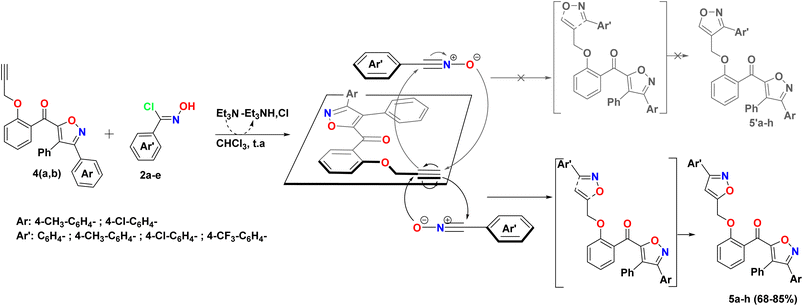
The o-propargylated isoxazoles 4a–b have been prepared from aroylisoxazoles 3a–b, which were obtained by a 1,3-dipolar cycloaddition reaction involving 2-benzylidene-3(2H)-benzofuranone 1 and 1,3-arylnitriloxide dipoles 2a–b. These 1,3-dipoles are generated in situ by the action of triethylamine on the corresponding chlorinated aldoximes. The reaction, due to a prototropic-1,3 rearrangement, does not lead to the expected spiroisoxazolines, the result is instead aroylisoxazoles. The high reactivity of the hydroxyl group of the substrate was then exploited in an o-alkylation reaction with propargyl bromide, in the presence of K2CO3 as a base and DMF as a solvent at room temperature, leading to the formation of the corresponding o-propargylated isoxazoles 4a–b (Scheme 1).11,16The previously synthesized o-propargylated isoxazoles 4a–b, were used as dipolarophiles in 1,3-dipolar cycloaddition reactions with arylnitriloxide dipoles 2a–d, which were generated in situ from the corresponding chlorinated aldoximes under the action of triethylamine in chloroform at room temperature. The course of these reactions was marked by a high degree of regioselectivity, culminating in the exclusive formation of 3,5-disubstituted isoxazole rings. Consequently, a novel series of hybrid systems 5a–h was obtained, comprising two 3,5-di- and 3,4,5-trisubstituted isoxazole rings, linked by a methoxybenzoyl motif serving as a structural linker, with satisfactory yields (Scheme 2). This remarkable regioselectivity underscores the controlled reactivity of arylnitriloxide dipoles toward o-propargylated dipolarophiles, thereby providing an efficient synthetic route for the construction of complex heterocyclic structures with high biological potential.
A series of in-depth spectroscopic analyses were conducted to confirm the structures of the newly synthesized hybrid compounds 5a–h, including FT-IR, NMR (1H, 13C, and 2D), and HRMS. The combined evaluation of the results confirmed the structures of the obtained compounds unambiguously.
The hybrid compound 5b has been selected as a representative example from the series 5a–h. An examination of its IR spectrum (Fig. S9) revealed a band at 3115 cm−1 associated with the absorption of the C4–H bond of the isoxazole ring. A second absorption band of the aromatic C–H stretching vibration occurred at approximately 3112 cm−1. Two bands at around 2925 and 2871 cm−1 are weak (low-intensity) and are assigned to aliphatic C–H stretching. A strong band at approximately 1661 cm−1 corresponds to the stretching vibration of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O carbonyl function. The 1H NMR spectrum of the compound (Fig. S6) displays two singlet peaks at 2.31 and 2.41 ppm, indicating the presence of protons from two methyl groups (–CH3). A singlet resonance at 5.07 ppm was assigned to the methylene protons (–O–CH2–). It is evident that an additional singlet signal at 6.39 ppm is consistent with the isoxazole proton (–C4H–). The 13C NMR spectrum (Fig. S7) of the sample exhibited two signals at 21.33 and 21.44 ppm, attributed to methyl carbons (–CH3). A signal at 61.90 ppm corresponds to the methylene carbon atom (–O–CH2–). A signal at 101.22 ppm is attributed to the disubstituted isoxazole carbon (C4). Two additional signals emerge at 167.07 and 156.59 ppm, corresponding respectively to the carbons (C5) of the di- and trisubstituted isoxazole nuclei. The signal at 183.50 ppm is attributed to the carbonyl function carbon (C
O carbonyl function. The 1H NMR spectrum of the compound (Fig. S6) displays two singlet peaks at 2.31 and 2.41 ppm, indicating the presence of protons from two methyl groups (–CH3). A singlet resonance at 5.07 ppm was assigned to the methylene protons (–O–CH2–). It is evident that an additional singlet signal at 6.39 ppm is consistent with the isoxazole proton (–C4H–). The 13C NMR spectrum (Fig. S7) of the sample exhibited two signals at 21.33 and 21.44 ppm, attributed to methyl carbons (–CH3). A signal at 61.90 ppm corresponds to the methylene carbon atom (–O–CH2–). A signal at 101.22 ppm is attributed to the disubstituted isoxazole carbon (C4). Two additional signals emerge at 167.07 and 156.59 ppm, corresponding respectively to the carbons (C5) of the di- and trisubstituted isoxazole nuclei. The signal at 183.50 ppm is attributed to the carbonyl function carbon (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). The 2D-HMBC (1H–13C) NMR spectrum (Fig. S8) of compound 5b revealed long-distance correlations (2J and 3J). The obtained correlations confirm the regioselectivity of the cycloaddition, thereby establishing unequivocally that the isolated compound corresponds to the 3,5-disubstituted regioisomer and not to the 3,4-disubstituted isomer. The methyl protons Ha (5.07 ppm) exhibited two cross-correlations with two characteristic carbons of the isoxazole ring: C4 (101.22 ppm) and C5 (167.07 ppm). The aforementioned protons also exhibit a correlation with carbon C1′ (156.59 ppm), thereby suggesting a direct spatial proximity. In addition, the Hb proton of the isoxazole (6.04 ppm) showed two significant correlations with C3 (162.44 ppm) and C5 (167.09 ppm). The correlations confirmed their significance in the isoxazole ring, particularly between C5 and the Ha and Hb protons. The correlations obtained were further confirmed by the literature, which accurately confirmed that the cycloadduct obtained is a 3,5-disubstituted regioisomer.12,52–55 The structures of the synthesized hybrid compounds were confirmed by HRMS. For compounds 5b, taken as a representative example, its spectrum (Fig. S10) shows a molecular ion peak at m/z = 549.1456, which corresponds exactly to the sodium ion mass [M + Na]+ at m/z = 549.1903. A second peak, at m/z = 527.19299, is attributed to the protonated molecular ion, [M + H]+, and matches the calculated value well (m/z = 527.19708). Additionally, it demonstrates typical fragments of the molecular ion; with several examples illustrated in Scheme S1.
O). The 2D-HMBC (1H–13C) NMR spectrum (Fig. S8) of compound 5b revealed long-distance correlations (2J and 3J). The obtained correlations confirm the regioselectivity of the cycloaddition, thereby establishing unequivocally that the isolated compound corresponds to the 3,5-disubstituted regioisomer and not to the 3,4-disubstituted isomer. The methyl protons Ha (5.07 ppm) exhibited two cross-correlations with two characteristic carbons of the isoxazole ring: C4 (101.22 ppm) and C5 (167.07 ppm). The aforementioned protons also exhibit a correlation with carbon C1′ (156.59 ppm), thereby suggesting a direct spatial proximity. In addition, the Hb proton of the isoxazole (6.04 ppm) showed two significant correlations with C3 (162.44 ppm) and C5 (167.09 ppm). The correlations confirmed their significance in the isoxazole ring, particularly between C5 and the Ha and Hb protons. The correlations obtained were further confirmed by the literature, which accurately confirmed that the cycloadduct obtained is a 3,5-disubstituted regioisomer.12,52–55 The structures of the synthesized hybrid compounds were confirmed by HRMS. For compounds 5b, taken as a representative example, its spectrum (Fig. S10) shows a molecular ion peak at m/z = 549.1456, which corresponds exactly to the sodium ion mass [M + Na]+ at m/z = 549.1903. A second peak, at m/z = 527.19299, is attributed to the protonated molecular ion, [M + H]+, and matches the calculated value well (m/z = 527.19708). Additionally, it demonstrates typical fragments of the molecular ion; with several examples illustrated in Scheme S1.
2.2. Antimicrobial activity
The newly synthesized 5a–h hybrids were subjected to, in vitro, antimicrobial assay. This study aimed to evaluate the potential antibacterial activity of these hybrids against three strains: two Gram-positive bacteria; Staphylococcus aureus ATCC 29213 and Bacillus subtilis ATCC 6633 and one Gram-negative bacterium Escherichia coli K12. Concurrently, they were evaluated as antifungal agents against four fungal strains: the yeast Candida albicans ATCC 10231 and the molds Aspergillus niger MTCC 282, Aspergillus flavus MTCC 9606, and Fusarium oxysporum MTCC 9913. The experimental tests were carried out in accordance with the MIC assay method, with previous protocols being adapted through certain modifications to optimize the testing conditions.3,56–58 Ampicillin was used as a positive control for both Gram-positive and Gram-negative bacteria, while fluconazole and amphotericin B served as positive controls for antifungal activity. DMSO was employed as a negative control to verify the absence of solvent effect.As shown in Table 1, the MICs (in µmol mL−1) vary depending on the nature of the aromatic substituents, Ar (4-CH3–C6H4–; 4-Cl–C6H4–) and Ar′ (C6H5–; 4-CH3–C6H4–; 4-Cl–C6H4–; 4-CF3–C6H4–), as well as the susceptibility of the different microbial strains examined.
| Pdts | Ar | Ar′ | Antibacterial activity | Antifungal activity | |||||
|---|---|---|---|---|---|---|---|---|---|
| MIC (µmol mL−1) | |||||||||
| E. coli | B. subtilis | S. aureus | C. albicans | A. niger | A. flavus | F. oxysporum | |||
| a MIC values represent two independent experiments performed in duplicate. Identical results were obtained in both replicates for all tested compounds; therefore, standard deviation was not applicable. | |||||||||
| 5a | 4-CH3–C6H4– | C6H5– | 4.878 | 0.097 | NA | 0.097 | 0.156 | 0.048 | 0.048 |
| 5b | 4-CH3–C6H4– | 4–CH3–C6H4– | NA | 0.094 | NA | 0.094 | 0.094 | 0.094 | 0.094 |
| 5c | 4-CH3–C6H4– | 4-Cl–C6H4– | 3.289 | 0.146 | NA | 0.091 | 0.091 | 0.091 | 0.091 |
| 5d | 4-CH3–C6H4– | 4-CF3–C6H4– | 2.582 | 0.430 | NA | 0.086 | 0.086 | 0.086 | 0.043 |
| 5e | 4-Cl–C6H4– | C6H5– | 4.690 | 0.938 | NA | 0.093 | 0.150 | 0.093 | 0.093 |
| 5f | 4-Cl–C6H4– | 4-CH3–C6H4– | NA | 1.828 | NA | 0.091 | 0.091 | 0.091 | 0.045 |
| 5g | 4-Cl–C6H4– | 4-Cl–C6H4– | 4.406 | 1.409 | NA | 0.088 | 0.088 | 0.044 | 0.088 |
| 5h | 4-Cl–C6H4– | 4-CF3–C6H4– | 4.161 | 0.083 | NA | 0.083 | 0.083 | 0.083 | 0.083 |
| Amphotericin B | N.D. | N.D. | N.D. | 0.002 | N.D. | N.D. | N.D. | ||
| Ampicillin | 0.0286 | 0.002 | 0.108 | N.D. | N.D. | N.D. | N.D. | ||
| Fluconazole | N.D. | N.D. | N.D. | N.D. | 0.130 | 0.032 | 0.130 | ||
The analysis of the MICs against E. coli reveals notable differences among compounds 5a–h regarding their antibacterial effect. Particularly, compound 5d stands out with an MIC of 2.582 µmol mL−1, indicating significant inhibitory activity. Compounds 5c, 5h, and 5g demonstrate slightly enhanced activity, whereas derivatives 5a and 5e, which bear an unsubstituted Ar′ ring, are among the least effective. These trends align with the enhanced activity against E. coli when electron-withdrawing groups, like –Cl and –CF3, were introduced. The combination of an Ar (4-CH3–C6H4) and a strong electron-withdrawing group in Ar′ (4-CF3–C6H4) is considered to be the most effective, and this combination produced smallest MIC value for 5d. In the case of B. subtilis, effective MICs were observed for hybrids 5a–h, where 5h showed high efficacy with an MIC of 0.083 µmol mL−1, which is almost equivalent to ampicillin's MIC (0.002 µmol mL−1). Compounds 5a–c presented high MIC values of 0.097 and 0.146 µmol mL−1, respectively, but compounds 5f and 5g showed loss of efficacy.
Collectively, these results show that the electron-withdrawing groups (–Cl, –CF3) enhance the activity of the compounds, particularly against B. subtilis, as manifested by compound 5h. None of the prepared compounds were active against S. aureus, showing its resistance to this variety of drugs. Although both bacteria are Gram-positive, the compounds showed more activity against B. subtilis than S. aureus due to the stronger binding of the peptidoglycan layer to teichoic and lipoteichoic acids, with high concentration, in S. aureus, thereby decreasing the cell wall permeability of compounds.59–61 On the other hand, B. subtilis has less compact cell wall architecture, allowing easy access for the compounds, hence a high sensitivity was observed.62
Regarding antifungal activity, the compounds 5a–h showed strong activity against A. niger and F. oxysporum. For C. albicans, the compounds 5h, 5g, and 5h had the lowest MICs, and the results were significantly different from the other compounds (p < 0.05). Compound 5h showed the strongest activity against C. albicans, with an MIC of 0.083 µmol mL−1, which was nearly close to that of amphotericin B (MIC = 0.002 µmol mL−1). Compounds 5a–c and 5e–f showed MIC values of 0.091 to 0.097 µmol mL−1, respectively, indicating strong anti-C. albicans activity compared to the reference drug. Analysis of the results shows that the Ar and Ar′ with –CF3 and –Cl have a strong to moderate activity against C. albicans when substituents are –CH3. However, the parent compounds had the lowest antifungal activity. Concerning the antifungal activity against the molds A. niger, A. flavus, and F. oxysporum, all compounds showed potent activity equivalent to that of fluconazole, especially against F. oxysporum. For the antifungal activity against A. niger, all the compounds with Ar′ having –CF3 showed stronger inhibition than fluconazole, specifically compounds 5h, Ar′ (–CF3) and 5e, Ar′ (–CF3), with MICs of 0.083 µmol mL−1, and 0.086 µmol mL−1, respectively. Thus, compounds 5b–c and 5f–g showed MICs of 0.088 to 0.094 µmol mL−1 lower than that of fluconazole, indicating potent activity.
These results confirm that the substituents Cl and –CF3 enhance the antifungal activity. In contrast, fluconazole was more effective than most the compounds against A. flavus with exception of 5a and 5g, which had lower MICs of 0.048 and 0.044 µmol mL−1, respectively. Against F. oxysporum, MIC values of the hybrid series 5a–h ranged from 0.43 to 0.94 µmol mL−1 well below fluconazole at 0.130 µmol mL−1. It therefore follows that a structural factor of a double isoxazole ring provides importantly high activity against F. oxysporum to the tested compounds. Moreover, the substituent groups (–Cl and –CF3) make the developed compounds more potent as represented by 5d and 5f.
2.3. Antioxidant activity
The antioxidant capacity of the synthesized hybrids (5a–h) was determined by using the phosphomolybdenum test, as reported in the method of Zengin et al.63 This test is based on the reduction of Mo(VI) to Mo(V), where a green Mo(V) complex forms, quantifiable by its measurement through a spectrophotometer at a wavelength of 630 nm. This was subjected to calculation, where the result was presented as a milligram of ascorbic acid equivalent to a gram of compound (mg AAE per g). DMSO was used as a negative control, while ascorbic acid was used as a positive control. Replication of the result was done by completing all of the mentioned tasks thrice (Fig. 1).The results, as shown in Fig. 3, indicate compounds 5e and 5g as the most effective compounds with maximum values (231.20 ± 2.15 and 197.90 ± 3.01 mg AAE per g, respectively), suggesting their high antioxidant capability. Compounds 5e and 5g were significantly different (p < 0.05) from the other derivatives. For compounds 5c, 5f and 5h, the reducing activity ranged from 124.51 to 175.06 mg of active principles (mg AAE per g). The lowest activity has been exhibited by compounds 5b (60.83 ± 0.86 mg AAE per g) and 5d (67.93 ± 2.37 mg AAE per g). This significant difference in activity can be explained by the type and position of substituents at the isoxazole ring, which modulate electron density and electron transfer ability. This fully functionalized isoxazole–isoxazole hybrid core structure, in particular with substituent electron donors, appears to be crucial for enhancing the reducing capacity of these compounds.
2.4. Structure–activity relationship (SAR)
Structure activity relationship analysis indicates that the substituents present in the aromatic rings (Ar and Ar′) play a critical part in the activity of the 5a–h series of compounds against microorganisms. Generally, the presence of strong electronegative substituents such as –CF3 and –Cl groups showed a marked improvement in activity, while the –CH3 group showed moderate activity and the least active were the unsubstituted groups. This trend was observed specifically for E. coli, among which the most active compound 5d (Ar = 4-CH3–C6H4 and Ar′ = 4-CF3–C6H4) had the lowest MIC value. This suggests that reaching a balance between the electronic and lipophilic properties is critical for the penetration of these molecules into the outer membrane of the Gram-negative bacteria.64,65 This trend was also observed with the activity level of the series against B. subtilis, where the strongest activity was exhibited by the electron–pull substituents, particulary 5h. This substituent could develop an affinity for the target enzymes for the Gram-positive bacteria.66,67 On the other hand, S. aureus was resistant to the series of compounds given its thick peptidoglycan cell wall and the presence of permeability barriers and efflux pump.68,69 For C. albicans, the strongest activity was observed with the 5d, 5g, and 5h series of compounds, thus indicating that electroattractive groups have the ability to improve the binding affinities for the fungal membranes and the biochemical targets.70,71 Minimally substituted molecules indicated that higher hydrophobicity and altered electron densities were required for improving the antifungal activity.72,73 This trend observed for C. albicans was reflected for the mold's series: the derivatives with –CF3 and –Cl substituents (5e, 5h, 5d, and 5f) were the least active in terms of MIC and were even more active than fluconazole against F. oxysporum. This activity could be due to the two isoxazole nuclei present in the molecule.41 This arrangement may be improving the interaction with the cell wall components. This trend indicates the importance of enhancing the antimicrobial activity through the creation of more attractive substituents for this class of compounds. The arrangement for the 5d and 5h series is worth noting for enhancing their potential for other properties.Analyses based on antioxidant activity show that the level of antioxidant activity potential could be influenced by the nature of the substituent group present in the isoxazole ring structure. Compounds 5e and 5g, containing the –Cl substituent (an electron withdrawing group but also participating in the enhancement of electron density due to delocalization), display high antioxidant activity. However, compounds 5b and 5d with the lowest activity show the significant impact of mesomerism and inductive effects on the relative stabilization of the generated radicals. Moderate activity was noted for compounds 5c, 5f, and 5h, which display less favorable structure characteristics. These observations warrant that the nature and characteristics of the particular substituent group, specifically electron-donating groups, have significant effects in increasing the reducing power of the compounds and can be utilized in designing more active isoxazole derivatives.74–76
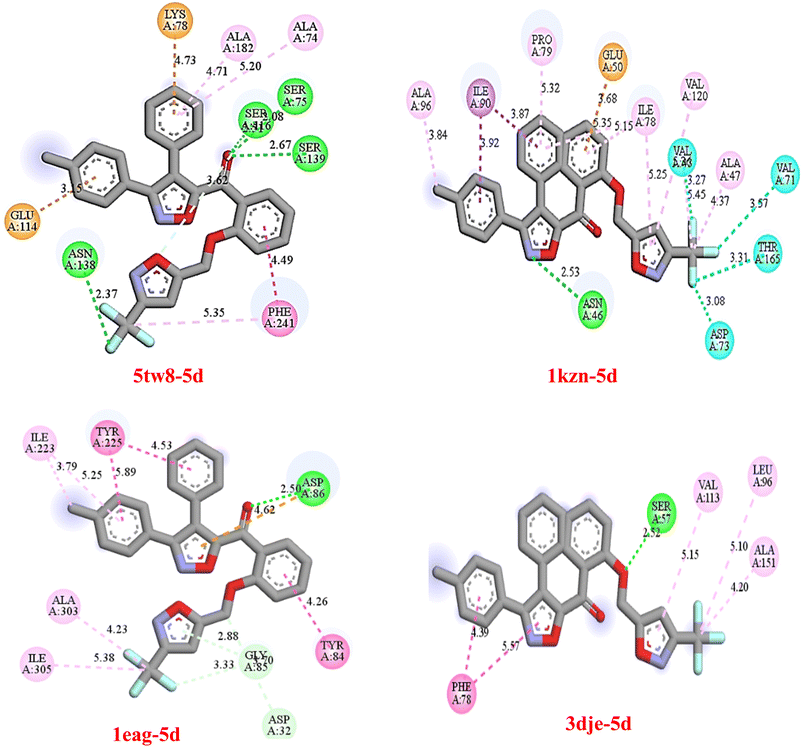
2.5. Molecular docking analysis
To investigate the binding affinity of compounds 5a–h with the desired bacterial (1KZN and 5TW8) and fungal (1EAG and 3DJE) protein targets that are of prime importance in the pathogenicity of microbes. The docking study revealed a high binding affinity with considerable dock scores of ΔG (Gibbs free energy) in the range of −8.3 kcal mol−1 to −11.8 kcal mol−1 (Table 2). Among the tested compounds 5a, 5b and 5c proved to have elite dual antibacterial and antifungal properties with high binding energies of up to −11.7 kcal mol−1 for the fungal protein 3DJE and greater than −9.0 kcal mol−1 for the bacterial protein targets. Compound 5g proved to have the highest binding energy of −11.8 kcal mol−1 for the fungal protein and also retains high antibacterial properties. The compound possesses the ability to have broad-spectrum activity. Also, the binding energy of the newly designed compounds proved to be much higher than the standard antibacterial and antifungal compounds ampicillin (−7.3 kcal mol−1), fluconazole (−6.6 kcal mol−1 to −7.9 kcal mol−1), and amphotericin B (−8.1 kcal mol−1). The result indicated better binding of the newly synthesized compounds with the active site of the microbial protein. It appears that the enhanced docking ability of these compounds provides a robust basis for being capable of blocking the crucial enzymatic function necessary for survival, and hence, they are promising leads for the development of new drugs targeting the pathogen. However, further validation and study are necessary to establish biological activity.| Compounds | Antibacterial target | Antifungal target | ||
|---|---|---|---|---|
| 1kzn | 5tw8 | 1eag | 3dje | |
| 5a | −9.8 | −9.9 | −10.1 | −11.6 |
| 5b | −9.1 | −9.4 | −8.5 | −11.7 |
| 5c | −9.0 | −9.3 | −8.5 | −11.7 |
| 5d | −9.6 | −9.3 | −9.5 | −8.3 |
| 5e | −9.4 | −9.6 | −9.4 | −11.4 |
| 5f | −8.7 | −10.2 | −10.5 | −8.5 |
| 5g | −9.0 | −10.3 | −10.0 | −11.8 |
| 5h | −9.2 | −10.2 | −10.6 | −8.8 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Scoring for the reference drug (kcal mol−1) | ||||
| Amphotericin B | *** | −8.1 | *** | *** |
| Ampicillin | −7.3 | *** | *** | *** |
| Fluconazole | *** | *** | −6.6 | −7.9 |
2.6. Computational ADME-Tox and drug-likeness analyses
For the assessment of the pharmacokinetic behavior and drug likeness of the identified molecules, an in-depth ADME-Tox study was conducted. The Swiss ADME and pkCSM online servers were employed to predict the parameters of interest, including oral bioavailability, solubility, lipophilicity, and toxicity. Table 3 represents the bioavailability characteristics of the eight most active molecules selected based on the data obtained from the docking simulation studies.| Molecule | 5a | 5b | 5c | 5d | 5e | 5f | 5g | 5h |
|---|---|---|---|---|---|---|---|---|
| MW (size) | 512.55 | 450.49 | 470.90 | 504.46 | 532.97 | 547.00 | 567.42 | 600.97 |
| GI absorption | Low | High | Low | Low | Low | Low | Low | Low |
| Fraction Csp3 (saturation index) | 0.06 | 0.11 | 0.07 | 0.11 | 0.03 | 0.06 | 0.03 | 0.06 |
| #Rotatable bonds (flexibility) | 8 | 7 | 7 | 8 | 8 | 8 | 8 | 9 |
| #H-bond acceptors | 6 | 6 | 6 | 9 | 6 | 6 | 6 | 9 |
| #H-bond donors | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| TPSA (polarity) | 78.36 | 78.36 | 78.36 | 78.36 | 78.36 | 78.36 | 78.36 | 78.36 |
| XLOGP3 (lipophilicity) | 7.31 | 6.05 | 6.61 | 6.57 | 7.57 | 7.93 | 8.20 | 8.45 |
| MLOGP | 3.76 | 3.01 | 3.28 | 3.57 | 4.03 | 4.21 | 4.48 | 4.75 |
ESOL log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S (insolubility) S (insolubility) |
−7.74 | −6.59 | −7.07 | −7.14 | −8.03 | −8.33 | −8.62 | −8.88 |
| Lipinski #violations | 1 | 0 | 0 | 1 | 1 | 2 | 2 | 2 |
| Bioavailability score | 0.55 | 0.55 | 0.55 | 0.55 | 0.55 | 0.17 | 0.17 | 0.17 |
| PAINS #alerts | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Synthetic accessibility | 4.41 | 4.13 | 4.03 | 4.18 | 4.29 | 4.41 | 4.29 | 4.40 |
Compounds 5a–h were subjected to in silico ADME-Tox analyses utilizing the SwissADME and pkCSM platforms to provide preliminary insights into their pharmacokinetic and drug-like characteristics. Table 3 summarizes the expected physicochemical properties and bioavailability characteristics, while Table 4 presents the full ADME-Tox profiles. The results show that numerous compounds surpass the acceptable molecular weight threshold of 500 g mol−1 and have high lipophilicity values (XLOGP3 > 5), resulting in Lipinski rule breaches. These physicochemical properties may have a deleterious impact on water solubility and oral bioavailability. The estimated ESOL log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S values (−6.59 to −8.88) indicate low water solubility across the sequence, especially for compounds 5f–5h (Table 3). Among the compounds tested, compound 5d had a substantially better projected safety profile, owing to the absence of hepatotoxicity alerts, AMES toxicity, hERG I inhibition, and skin sensitization, as shown in Table 4. It also had a high projected intestinal absorption rate (95.49%) and acceptable Caco-2 permeability (1.089) (Table 4). However, its molecular weight (504.46 g mol−1) and lipophilicity (XLOGP3 = 6.57) are slightly higher than the ideal range suggested by Lipinski's rule of five (Table 3), indicating that structural optimization may be required to improve its physicochemical balance. The remaining analogues (5a, 5c, 5e–5h) had moderate to low predicted GI absorption and greater lipophilicity values, which could account for their lower bioavailability ratings (Table 3). Furthermore, some substances were predicted to inhibit CYP2C9 and CYP3A4 (Table 4), highlighting the possibility of drug–drug interactions. The total clearance and distribution parameters presented in Table 4 indicate moderate elimination rates and little central nervous system penetration throughout the series. It is important to emphasize that these findings are based exclusively on computational predictions and should be interpreted as preliminary screening data rather than definitive evidence of drug potential. Experimental validation through in vitro and in vivo pharmacokinetic and toxicity studies is required to confirm these theoretical assessments.
S values (−6.59 to −8.88) indicate low water solubility across the sequence, especially for compounds 5f–5h (Table 3). Among the compounds tested, compound 5d had a substantially better projected safety profile, owing to the absence of hepatotoxicity alerts, AMES toxicity, hERG I inhibition, and skin sensitization, as shown in Table 4. It also had a high projected intestinal absorption rate (95.49%) and acceptable Caco-2 permeability (1.089) (Table 4). However, its molecular weight (504.46 g mol−1) and lipophilicity (XLOGP3 = 6.57) are slightly higher than the ideal range suggested by Lipinski's rule of five (Table 3), indicating that structural optimization may be required to improve its physicochemical balance. The remaining analogues (5a, 5c, 5e–5h) had moderate to low predicted GI absorption and greater lipophilicity values, which could account for their lower bioavailability ratings (Table 3). Furthermore, some substances were predicted to inhibit CYP2C9 and CYP3A4 (Table 4), highlighting the possibility of drug–drug interactions. The total clearance and distribution parameters presented in Table 4 indicate moderate elimination rates and little central nervous system penetration throughout the series. It is important to emphasize that these findings are based exclusively on computational predictions and should be interpreted as preliminary screening data rather than definitive evidence of drug potential. Experimental validation through in vitro and in vivo pharmacokinetic and toxicity studies is required to confirm these theoretical assessments.
| ADMET properties | 5a | 5b | 5c | 5d | 5e | 5f | 5g | 5h | |
|---|---|---|---|---|---|---|---|---|---|
| Absorption | Water solubility (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mol L−1) mol L−1) |
−2.933 | −3.276 | −3.525 | −3.460 | −2.933 | −3.072 | −3.069 | −3.063 |
Caco2 permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Papp) Papp) |
1.05 | 1.10 | 1.02 | 1.09 | 1.04 | 1.07 | 1.06 | 1.11 | |
| Intestinal absorption (%) | 96.85 | 100.0 | 97.17 | 95.49 | 95.39 | 97.73 | 96.27 | 94.89 | |
Skin permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp) Kp) |
−2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | −2.735 | |
| Distribution | VDss (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) L kg−1) L kg−1) |
−0.487 | −0.335 | −0.284 | −0.214 | −0.484 | −0.395 | −0.390 | −0.390 |
| Fraction unbound | 0.383 | 0.363 | 0.331 | 0.333 | 0.383 | 0.365 | 0.365 | 0.369 | |
BBB permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BB) BB) |
−0.868 | −0.866 | −0.984 | −1.245 | −1.043 | −1.098 | −1.274 | −1.546 | |
CNS permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PS) PS) |
−1.596 | −1.882 | −1.810 | −1.760 | −1.556 | −1.430 | −1.390 | −1.349 | |
| Metabolism | CYP2D6 substrate | No | No | No | No | No | No | No | No |
| CYP3A4 substrate | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | |
| CYP1A2 inhibitor | No | No | No | No | No | No | No | No | |
| CYP2C19 inhibitor | No | No | No | No | No | No | No | No | |
| CYP2C9 inhibitor | Yes | Yes | Yes | Yes | Yes | Yes | Yes | Yes | |
| CYP2D6 inhibitor | No | No | No | No | No | No | No | No | |
| CYP3A4 inhibitor | Yes | Yes | Yes | Yes | Yes | No | No | No | |
| Excretion | Total clearance (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mL min−1 kg−1) mL min−1 kg−1) |
0.483 | 0.388 | 0.238 | 0.132 | 0.259 | 0.201 | 0.127 | 0.180 |
| Renal OCT2 substrate | No | No | No | No | No | No | No | No | |
| Toxicity | AMES toxicity | No | No | No | No | No | No | No | No |
| hERG I inhibitor | No | No | No | No | No | No | No | No | |
| Hepatotoxicity | Yes | Yes | Yes | No | Yes | Yes | Yes | Yes | |
| Skin sensitization | No | No | No | No | No | No | No | No | |
2.7. Molecular dynamic simulation analysis
3. Materials and methods
3.1. Chemistry
The supplementary file includes a detailed description of reagents and solvents for the synthesis of target isoxazoles, instrumentation for analysis of characterization spectra, and syntheses.3.2. Antimicrobial activity
The supplementary file also holds information about the methodological approach in the bacterial and fungal strains that will be used to measure the antimicrobial potential of compounds 5a–h.3.3. Antioxidant activity
The procedure used for screening the antioxidant activity of compounds 5a–h is given in SI.3.4. Molecular docking simulation
For the exploration of the antimicrobial properties of selected natural bioactive compounds, molecular docking simulation studies were also performed with the help of AutoDock Vina. The major aim was to assess and study the binding affinity of these natural bioactive molecules with selected microbial target proteins.77 For this purpose, two antibacterial target proteins – DNA gyrase subunit B (PDB ID: 1KZN), which takes part in bacterial DNA synthesis, and penicillin-binding protein 2a (PDB ID: 5TW8), which plays an imperative function in cell wall synthesis, as well as two antifungal target proteins 1EAG, linked with Candida albicans, and 3DJE, obtained from Aspergillus fumigatus, which performs a pivotal function in fungal pathogenicity and life,77 were selected.783.5. Protein structure preparation
High-resolution crystal structures of the target microbial proteins were sourced from the RCSB Protein Data Bank (https://www.rcsb.org/) in PDB format. These proteins consisted of both antibacterial and antifungal targets that are associated with microbial pathogenesis. Structure visualization was done using BIOVIA Discovery Studio, with structure preparation using AutoDock Tools version 4.2.6. The preparation steps included the removal of water molecules and heteroatoms, the addition of polar hydrogen atoms, and the assignment of partial charges, which included Kollman charges in the case of proteins and Gasteiger charges in the case of ligands. The optimized protein structures were then saved in PDBQT format to make them compatible with the AutoDock Vina tool and to increase the accuracy of the docking simulations, as pointed out in ref. 79. To accurately target the active sites of the proteins and ensure that the complete active site was covered in simulating protein–ligand interactions, the docking boxes were placed around the coordinates and sizes of the 3D grid boxes as presented in Table 5.| PDB ID | Target | Centers (Å) | Size (Å) | Exhaustiveness | ||
|---|---|---|---|---|---|---|
| X | Y | Z | X, Y, Z | = | ||
| 1kzn | Antimicrobial | 19.1 | 30.3 | 34.7 | 20 | 9 |
| 5tw8 | Antimicrobial | 21.3 | 62.2 | 36.1 | 40 | 9 |
| 1eag | Antifungal | 41.6 | 24.9 | 13.6 | 40 | 9 |
| 3dje | Antifungal | 43.2 | 7.7 | 39.1 | 40 | 9 |
3.6. Ligand preparation
The identified bioactive molecules were initially drawn and optimized with the aid of Chem3D software in addition to energy minimization with Avogadro software. The molecules' geometry optimization was achieved through the steepest descent algorithm with the MMFF94 energetic term to obtain conformers with less energy as well as optimized conformers ready for docking experiments.80 Energy-optimized molecules' structures were then transformed to PDBQT files ready to be evaluated by AutoDock Vina to predict binding affinity to targeted molecules in agreement with previous docking experiments.79 Finally, a comparative study between the binding affinity scores from docking experiments and those from known available medications with well-known effectiveness as antibacterial and antifungal agents provided a relative effectiveness level of all test molecules in vitro.3.7. Computational ADME and drug-likeness analysis
In order to make an assessment about the pharmacokinetic optimality and potency of these compounds, an extensive in silico screening for their ADME-Tox profiles was carried out. Based on their SMILES codes, the https://www.swissadme.ch server, available as an open-access WEB resource, was used for these compounds to calculate their physicochemical and pharmacokinetic characteristics such as lipophilicity, water solubility, gastrointestinal absorption, and their aptness for the rule of five, which are essential criteria for analyzing their oral bioavailability and drug likeness.81 At the same time, another program named pkCSM was used for an extensive analysis relating to their ADME-Tox, particularly for their hepatotoxic, skin sensibility, blood–brain barrier permeability, and cytochrome P450 binding properties, as an attempt to get an insight into their pharmacokinetic optimality. In this way, an extensive pharmacokinetic screening was performed, aiming at the early selection of these compounds with potentially favorable pharmacokinetic profiles and low toxicity. These profiles work as filters at the early stages prior to their further laboratory tests.823.8. Molecular dynamics simulation (MDS)
Molecular dynamics simulation (MDS) was employed to explore the dynamic behavior and stability of the most promising ligand–protein complex at the atomic level. All simulations were conducted using Desmond, a component of the Schrödinger suite, over a 100 ns-time frame. The protein structure was first prepared using Schrödinger's Protein Preparation Wizard at a physiological pH of 7.4 to ensure structural accuracy and stability.83 The solvated system was constructed via the System Builder module using the TIP3P water model, with system neutrality achieved by the addition of Na+/Cl− counterions at a 0.15 M salt concentration. The OPLS3e force field was applied to define atomic interactions. Equilibration began under the NVT ensemble with gradual temperature increase, followed by the NPT ensemble regulated by the Nose–Hoover thermostat (300 K) and the Martyna–Tobias–Klein barostat (1.01325 bar). During the 100 ns production run, key parameters such as root mean square deviation (RMSD), root mean square fluctuation (RMSF), and protein–ligand interaction profiles were monitored to assess the conformational stability and binding performance of the complex.84Molecular dynamics simulation (MDS) was used to investigate the dynamic properties and stability of the most favorable ligand–protein complex at the atomic level. All MDS runs were carried out by the Desmond module of the Schrödinger software suite over a 100 ns simulation time course. Prior to the simulation, the protein was prepared using the Protein Preparation Wizard module of the Schrödinger suite at physiological pH 7.4 to guarantee structural correctness and conformational stability.83 The solvent system was set up using the System Builder module with the TIP3P water force field; neutrality was ensured by the addition of Na+/Cl− counter-ions at 0.15 M concentration 83. The OPLS3e force filed was used to describe the atomic interactions. The equilibration procedure was performed separately at first in the NVT ensemble with linear temperature scaling, followed by the NPT ensemble with control by the Nose–Hoover temperature regulator at 300 K and the Martyna–Tobias–Klein pressure regulator at 1.01325 bar 83. During the 100 ns simulation run, respect parameters like the root mean squared deviation (RMSD), the root mean squared fluctuation (RMSF), or protein–ligand interaction analyses were applied to examine the conformational and stabilizing properties of the ligand–protein complex.84
4. Conclusions
The current research works highlight the effective synthesis of a new series of isoxazole–isoxazole hybrids (5a–h) successfully isolated with substantial yields, where a comprehensive characterization is presented employing FT-IR, NMR (1H, 13C, 2D), and HR-MS. Evaluation of the numerous biological properties taken into account revealed that the hybrids presented potent and variable activities. They showed inhibitory effect towards different bacteria, including Gram-positive, Gram-negative, in addition to some fungal, yeast, and mold strains. Compounds 5d and 5h demonstrated substantial activity against E. coli, in addition to B. subtilis, while 5h, 5g presented pronounced activity towards C. albicans, A. niger, A. flavus, in addition to F. oxysporum. Notably, the hybrids presented exhibited improved activity compared to fluconazole, the reference drug. Also, in conjunction, evaluation of the antioxidant activity demonstrates that 5e, 5g presented substantial reducing capability. Importantly, the current works highlight the pivotal impact of the synthesized substituents in modifying the biological activities of the new isoxazole hybrids.In silico validation was done to confirm these findings. Using molecular docking and molecular dynamics simulations, it was proven that the isoxazole–isoxazole compounds, especially 5d, have stable and various interactions with bacterial (1KZN, 5TW8) and fungal (1EAG, 3DJE) protein structures using mechanisms such as hydrogen bonding, hydrophobic interactions, and water bridging. Within this group of compounds, the binding affinity of 5d stands out as the strongest. By the root mean square fluctuation (RMSF) calculation, these interactions help in decreasing the fluctuation of residues and thus can enhance the binding affinity of the complex. The ADME-Tox profile of the in silico analysis suggests that the most promising candidate with high gastrointestinal absorption, drug-likeness, and balanced pharmacokinetic properties suitable for development as an antimicrobial agent is 5d.
It is pertinent to highlight that the significance of the isoxazole–isoxazole ring structure is made explicit in this research work. It is clear from the above-stated observations that the outcomes also verify the efficiency and effectiveness associated with the isoxazole–isoxazole ring structure. It can thus be added that the feasibility associated with the experimentally designed synthesis contributes effectively towards validating the efficiency associated with the isoxazole–isoxazole framework.
Conflicts of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.Data availability
All data generated or analysed during this study are included in the article and its supplementary information (SI). Supplementary information: spectroscopic data and copies of IR, NMR and HRMS spectra of the newly synthesized compounds, and all additional details. See DOI: https://doi.org/10.1039/d6ra01005a.Acknowledgements
This work was funded by the Ongoing Research Funding Program, (ORF-2026-418), King Saud University, Riyadh, Saudi Arabia. The authors are very much thankful to the staff members of the “Cité de l'Innovation” of the Sidi Mohamed Ben Abdellah University, Fez, Morocco.References
- X. Wang, Q. Hu, H. Tang and X. Pan, Pharmaceuticals, 2023, 16, 228 CrossRef CAS PubMed.
- V. V. Pothuri, P. V. S. Machiraju and V. S. S. Rao, Russ. J. Gen. Chem., 2020, 90, 889–894 CrossRef CAS.
- M. Chalkha, H. Nour, K. Chebbac, A. Nakkabi, L. Bahsis, M. Bakhouch, M. Akhazzane, M. Bourass, S. Chtita, Y. A. Bin Jardan, M. Augustyniak, M. Bourhia, M. A. M. Aboul-Soud and M. El Yazidi, ACS Omega, 2022, 7, 46731–46744 CrossRef CAS PubMed.
- B. C. Monk and M. V. Keniya, J. Fungi, 2021, 7, 1–36 Search PubMed.
- J. Zhang, L. Li, Q. Lv, L. Yan, Y. Wang and Y. Jiang, Front. Microbiol., 2019, 10, 69 CrossRef PubMed.
- K. V. Chikkula and S. Raja, Int. J. Pharm. Pharm. Sci., 2017, 9, 13 CrossRef CAS.
- A. M. Fuentefria, B. Pippi, D. F. Dalla Lana, K. K. Donato and S. F. de Andrade, Lett. Appl. Microbiol., 2018, 66, 2–13 CrossRef CAS PubMed.
- N. Agrawal and P. Mishra, Med. Chem. Res., 2018, 27, 1309–1344 CrossRef CAS PubMed.
- G. J. Martis and S. L. Gaonkar, RSC Adv., 2025, 15, 8213–8243 RSC.
- U. Malik and D. Pal, Eur. J. Med. Chem., 2024, 279, 116898 CrossRef CAS PubMed.
- A. Arzine, S. A. Assou, F. Z. Guerguer, K. Boujdi, Y. Rhazi, N. Barghady, M. Chalkha, A. Nakkabi, K. Chkirate, J. T. Mague, M. Bakhouch, S. Chtita, H. Mabrouk, M. El Hassouni and M. El, ChemistrySelect, 2025, 10, e05455 CrossRef CAS.
- Y. Rhazi, R. Sghyar, N. Deak, B. Es-Sounni, B. Rossafi, A. Soran, M. Laghmari, A. Arzine, A. Nakkabi, K. Hammani, S. Chtita, M. M. Alanazi, G. Nemes and M. E. Yazidi, Pharmaceuticals, 2024, 17, 1390 CrossRef CAS PubMed.
- A. Arzine, H. Hadni, K. Boujdi, K. Chebbac, N. Barghady, Y. Rhazi, M. Chalkha, A. Nakkabi, K. Chkirate, J. T. Mague, S. M. A. Kawsar, G. Al Houari, M. M. Alanazi and M. El Yazidi, Molecules, 2024, 29, 3366 CrossRef CAS PubMed.
- A. Arzine, S. Ait Assou, F. Z. Guerguer, K. Boujdi, Y. Rhazi, M. Chalkha, A. Nakkabi, M. Akhazzane, J. T. Mague, S. Chtita, M. El Hassouni and M. El Yazidi, Chem. Biodiversity, 2025, e00600 CrossRef CAS PubMed.
- N. Barghady, S. A. Assou, M. Er-rajy, K. Boujdi, A. Arzine, Y. Rhazi, B. Tüzün, A. Nakkabi, M. Chalkha, M. El Hassouni and A. Kabra, Open Chem., 2024, 22, 202401 Search PubMed.
- A. Arzine, O. Abchir, M. Chalkha, K. Chebbac, Y. Rhazi, N. Barghady, I. Yamari, A. E. L. Moussaoui, A. Nakkabi, M. Akhazzane, M. Bakhouch, S. Chtita and M. E. L. Yazidi, Comput. Biol. Chem., 2023, 107993 Search PubMed.
- G. C. Arya, K. Kaur and V. Jaitak, Eur. J. Med. Chem., 2021, 221, 113511 CrossRef CAS PubMed.
- G. C. Arya, M. Khalid, S. Mehla and V. Jakhmola, J. Biomol. Struct. Dyn., 2024, 42, 4909–4935 CrossRef CAS PubMed.
- J. Zhu, J. Mo, H. zhi Lin, Y. Chen and H. peng Sun, Bioorg. Med. Chem., 2018, 26, 3065–3075 CrossRef CAS PubMed.
- M. A. Barmade, P. R. Murumkar, M. K. Sharma and M. R. Yadav, Curr. Top. Med. Chem., 2016, 16, 2863–2883 CrossRef CAS PubMed.
- R. G. Marks, L. Ravich and S. F. Wilhelm, J. Urol., 1955, 73, 179–181 CrossRef CAS PubMed.
- J. O. Straub, Environ. Toxicol. Chem., 2016, 35, 767–779 CrossRef CAS PubMed.
- A. Egorova, E. Kazakova, B. Jahn, S. Ekins, V. Makarov and M. Schmidtke, Eur. J. Med. Chem., 2020, 188, 112007 CrossRef CAS PubMed.
- R. I. Fox, J. Rheumatol., 1998, 25, 20–26 Search PubMed.
- Y. Shinde, B. Khairnar and S. Bangale, ChemistrySelect, 2024, 9, e202401423 CrossRef CAS.
- M. Majirská, M. B. Pilátová, Z. Kudličková, M. Vojtek and C. Diniz, Drug Discovery Today, 2024, 29, 104059 CrossRef PubMed.
- J. Wang, D. B. Wang, L. L. Sui and T. Luan, Arabian J. Chem., 2024, 17, 105794 CrossRef CAS.
- T. Ni, X. Chi, F. Xie, L. Li, H. Wu, Y. Hao, X. Wang, D. Zhang and Y. Jiang, Eur. J. Med. Chem., 2023, 246, 115007 CrossRef CAS PubMed.
- K. Tripathi, P. Kaushik, D. K. Yadav, R. Kumar, S. R. Misra, R. Godara, B. M. Bashyal, V. S. Rana, R. Kumar, J. Yadav and N. A. Shakil, Pest Manage. Sci., 2024, 81, 2579 CrossRef PubMed.
- V. A. Makarov, O. B. Riabova, V. G. Granik, P. Wutzler and M. Schmidtke, J. Antimicrob. Chemother., 2005, 55, 483–488 CrossRef CAS PubMed.
- Y. S. Lee and H. K. Byeang, Bioorg. Med. Chem. Lett., 2002, 12, 1395–1397 CrossRef CAS PubMed.
- R. Díaz-Trelles, A. Novelli and M. T. Fernández-Sánchez, Neurosci. Lett., 2003, 345, 136–140 CrossRef PubMed.
- N. Ganji, S. Daravath, A. Rambabu, K. Venkateswarlu, D. Shiva Shankar and Shivaraj, Inorg. Chem. Commun., 2020, 121, 108247 CrossRef CAS.
- A. Sysak and B. Obmińska-Mrukowicz, Eur. J. Med. Chem., 2017, 137, 292–309 CrossRef CAS PubMed.
- W. Alam, H. Khan, M. Saeed Jan, U. Rashid, A. Abusharha and M. Daglia, Front. Chem., 2023, 11, 1–19 Search PubMed.
- M. Zimecki, U. Bachor and M. Maczyński, Molecules, 2018, 23, 2724 CrossRef PubMed.
- Z. Yang, Y. Zhao, P. Li and Y. He, J. Heterocycl. Chem., 2019, 56, 3042–3047 CrossRef CAS.
- F. Ye, Y. Zhai, T. Kang, S. L. Wu, J. J. Li, S. Gao, L. X. Zhao and Y. Fu, Pestic. Biochem. Physiol., 2019, 157, 60–68 CrossRef CAS PubMed.
- X. Lin, Y. Li, W. Zhong, T. Hong, L. Li, S. Song and D. He, J. Agric. Food Chem., 2021, 69, 9520–9528 CrossRef CAS PubMed.
- M. Lasri, A. Bimoussa, A. Ait-karra, Y. Laamari, O. Zakir, R. Idouhli, M. Maatallah, K. M. Eddine, A. Auhmani, M. Y. A. Itto and A. Abouelfida, Colloids Surf., A, 2024, 695, 134227 CrossRef CAS.
- U. Bąchor, M. Mączyński and A. Sochacka-Ćwikła, Int. J. Mol. Sci., 2025, 26, 1–26 Search PubMed.
- M. Szumilak, A. Wiktorowska-Owczarek and A. Stanczak, Molecules, 2021, 26, 1–31 CrossRef PubMed.
- A. K. Singh, A. Kumar, H. Singh, P. Sonawane, H. Paliwal, S. Thareja, P. Pathak, M. Grishina, M. Jaremko, A. H. Emwas, J. P. Yadav, A. Verma, H. Khalilullah and P. Kumar, Pharmaceuticals, 2022, 15, 1071 CrossRef CAS PubMed.
- I. K. P. de Albuquerque, D. L. de Santana, F. de Assis Graciano dos Santos, F. N. Coutinho, V. M. de Almeida, A. R. de Faria, D. P. C. Macêdo and R. P. Neves, Braz. J. Microbiol., 2024, 55, 1811–1816 CrossRef CAS PubMed.
- A. Oubella, M. Fawzi, A. Bimoussa, A. N'Ait Ousidi, A. Auhmani, A. Riahi, A. Robert, L. El Firdoussi, H. Morjani and M. Y. Ait Itto, Chem. Pap., 2022, 76, 2935–2946 CrossRef CAS.
- A. Dubba and S. K. Koppula, Chem. Biol. Lett., 2024, 11, 1–6 Search PubMed.
- M. Hawash, N. Jaradat, A. M. Eid, A. Abubaker, O. Mufleh, Q. Al-Hroub and S. Sobuh, BMC Chem., 2022, 16, 1–12 Search PubMed.
- A. Adegoke, G. Azeez, A. Lawal and M. Imran, Adv. J. Chem., Sect. B, 2021, 3, 148 CAS.
- J. Magalhães, N. Franko, S. Raboni, G. Annunziato, P. Tammela, A. Bruno, S. Bettati, S. Armao, C. Spadini, C. S. Cabassi, A. Mozzarelli, M. Pieroni, B. Campanini and G. Costantino, Pharmaceuticals, 2021, 14, 1–16 CrossRef PubMed.
- A. Sochacka-Ćwikła, M. Mączyński, Ż. Czyżnikowska, B. Wiatrak, I. Jęśkowiak, A. Czerski and A. Regiec, Int. J. Mol. Sci., 2022, 13, 11694 CrossRef PubMed.
- A. E. Taishev, E. E. Galenko, M. S. Novikov and A. F. Khlebnikov, Russ. J. Gen. Chem., 2023, 93, 1246–1260 CrossRef CAS.
- D. C. B. Da Silva-Alves, J. V. Dos Anjos, N. N. M. Cavalcante, G. K. N. Santos, D. M. do A. F. Navarro and R. M. Srivastava, Bioorg. Med. Chem., 2013, 21, 940–947 CrossRef CAS PubMed.
- A. Naouri, A. Djemoui, M. R. Ouahrani, M. B. Lahrech, N. Lemouari, D. H. A. Rocha, H. Albuquerque, R. F. Mendes, F. A. Almeida Paz, L. A. Helguero, K. Bachari, O. Talhi and A. M. S. Silva, J. Mol. Struct., 2022, 1217, 128325 CrossRef.
- L. Phongphane, S. N. Mohd Radzuan, M. H. Abu Bakar, M. T. Che Omar, U. Supratman, D. Harneti, H. A. Wahab and M. N. Azmi, Comput. Biol. Chem., 2023, 106, 107938 CrossRef CAS PubMed.
- R. A. Al-Qawasmeh, L. A. Al-Nazer, S. A. Dawlat-Kari, L. Abu-Qatouseh, S. S. Sabri, M. A. Aldamen and M. Sinnokrot, Open Chem., 2020, 18, 138–148 CAS.
- S. Ait Assou, J. Anissi, K. Sendide and M. El Hassouni, Sci. World J., 2023, 2023, 6106673 Search PubMed.
- A. El Barnossi, F. Moussaid and A. Iraqi Housseini, E3S Web Conf., 2020, 150, 02003 CrossRef CAS.
- K. Chebbac, Z. Benziane Ouaritini, A. El Moussaoui, M. Chalkha, S. Lafraxo, Y. A. Bin Jardan, H. A. Nafidi, M. Bourhia and R. Guemmouh, Life, 2023, 13, 807 CrossRef CAS PubMed.
- C. Hansch, A. Leo and R. W. Taft, Chem. Rev., 1991, 91, 165–195 CrossRef CAS.
- E. Pontiki, A. Peperidou, I. Fotopoulos and D. Hadjipavlou-Litina, Curr. Pharm. Biotechnol., 2019, 19, 1019–1048 Search PubMed.
- K. Vashisht, P. Sethi, A. Bansal and P. Bansal, Vietnam J. Chem., 2025, 63, 195–213 CrossRef CAS.
- C. Weidenmaier and A. Peschel, Nat. Rev. Microbiol., 2008, 6, 276–287 CrossRef CAS PubMed.
- G. Zengin, T. Arkan, A. Aktumsek, G. O. Guler and Y. S. Cakmak, J. Food Biochem., 2013, 37, 646–653 CrossRef CAS.
- A. Rusu, I. M. Moga, L. Uncu and G. Hancu, Pharmaceutics, 2023, 15, 2554 CrossRef CAS PubMed.
- M. Mardirossian, M. Rubini, M. F. A. Adamo, M. Scocchi, M. Saviano, A. Tossi, R. Gennaro and A. Caporale, Molecules, 2021, 26, 7401 CrossRef CAS PubMed.
- J. Jays and S. Janardhanan, Discover Chem., 2025, 2, 139 CrossRef.
- Y. Liu, R. Qin, S. A. J. Zaat, E. Breukink and M. Heger, J. Clin. Transl. Res., 2015, 1, 140–167 Search PubMed.
- P. Nikolic and P. Mudgil, Microorganisms, 2023, 11, 259 CrossRef CAS PubMed.
- G. Kumar and K. Engle, Nat. Prod. Rep., 2023, 40, 1608–1646 RSC.
- N. A. Hamed, M. I. Marzouk, M. F. Ismail and M. H. Hekal, Synth. Commun., 2019, 49, 3017–3029 CAS.
- V. Tomar, G. Bhattacharjee, Kamaluddin and A. Kumar, Bioorg. Med. Chem. Lett., 2007, 17, 5321–5324 CrossRef CAS PubMed.
- S. Kagan, A. Jabbour, E. Sionov, A. A. Alquntar, D. Steinberg, M. Srebnik, R. Nir-Paz, A. Weiss and I. Polacheck, J. Antimicrob. Chemother., 2014, 69, 416–427 CrossRef CAS PubMed.
- D. Batovska, S. Parushev, A. Slavova, V. Bankova, I. Tsvetkova, M. Ninova and H. Najdenski, Eur. J. Med. Chem., 2007, 42, 87–92 CrossRef CAS PubMed.
- M. Adardour, M. Ait Lahcen, M. Oubahmane, W. Ettahiri, I. Hdoufane, H. Bouamama, M. M. Alanazi, D. Cherqaoui, M. Taleb, E. Z. Garcia and A. Baouid, Pharmaceuticals, 2023, 16, 1648 CrossRef CAS PubMed.
- W. S. Shehab, N. K. R. Elhoseni, W. M. Aboulthana, M. G. Assy, S. M. Mousa and G. T. El-Bassyouni, Sci. Rep., 2025, 15, 1–17 Search PubMed.
- K. Kaur, A. K. Al-Khazaleh, D. J. Bhuyan, F. Li and C. G. Li, Antioxidants, 2024, 13, 1092 CrossRef CAS PubMed.
- Schrödinger, Software, 2021, https://www.schrodinger.com/release-download/.
- P. Mark and L. Nilsson, J. Phys. Chem. A, 2001, 105, 9954–9960 CrossRef CAS.
- Q. Ke, X. Gong, S. Liao, C. Duan and L. Li, J. Mol. Liq., 2022, 365, 120116 CrossRef CAS.
- H. Nour, O. Daoui, O. Abchir, S. ElKhattabi, S. Belaidi and S. Chtita, Heliyon, 2022, 8, e11991 CrossRef CAS PubMed.
- O. Abchir, O. Daoui, S. Belaidi, M. Ouassaf, F. A. Qais, S. ElKhattabi, S. Belaaouad and S. Chtita, J. Mol. Model., 2022, 28, 106 CrossRef CAS PubMed.
- M. Khedraoui, H. Nour, I. Yamari, O. Abchir, A. Errougui and S. Chtita, Chem. Phys. Impact, 2023, 7, 100361 CrossRef.
- I. Yamari, A. Mouhib, B. Es-Sounni, R. Nejjari, N. Mazoir, M. Bakhouch, A. Mouzdahir, A. Benharref, M. El Kouali and S. Chtita, Chem. Phys. Impact, 2023, 7, 100372 CrossRef.
- I. Yamari, O. Abchir, F. Siddique, H. Zaki, A. Errougui, M. Talbi, M. Bouachrine, M. ElKouali and S. Chtita, Sci. Afr., 2024, 23, e01986 CAS.
| This journal is © The Royal Society of Chemistry 2026 |