Open Access Article
Open Access ArticleFrom in silico and machine learning model to validation: discovery of novel amide-functionalized imidazopyridines as PIM-1 kinase inhibitors using an integrated ligand and structure-based hopping approach
Vinayak Walhekara,
Shubham Dhadila,
Ankit Ganeshpurkara,
Srilakshmi Singagarib,
Ayesha Begum Khadernaickc,
Amol Muthala,
Vaibhav Shindea,
Ashwin Malia,
Raghavendra Kulkarnid and
Ravindra Kulkarni *a
*a
aDepartment of Pharmaceutical Chemistry, BVDU’S Poona College of Pharmacy, Erandawane Pune-411038, Maharashtra, India. E-mail: ravigk2000@gmail.com
bGitam School of pharmacy, Gitam University, Rushikonda, Visakhapatnam- 530045, India
cShadan Women's College of Pharmacy, Khairabad, Hyderabad, India
dDepartment of Pharmaceutics, Sri Sanganabasava Mahaswamiji College of Pharmacy and Research Center, Vijayapura, Karnataka, India
First published on 20th April 2026
Abstract
Pro-viral integration sites for Moloney murine leukaemia virus (PIM) kinases are the members of serine/threonine kinase family, that elevate cell division and inhibit apoptosis, thus making them as an attractive anticancer target. In this study, a virtual screening strategy based on a pharmacophore model was used to unearth imidazo[1,2-a]pyridin-2-yl phenyl benzamides as potential PIM1 inhibitors. A Python-based design workflow generated 27 candidate molecules, whose inhibitory activities were predicted using an in-house linear regression machine learning model. Selected compounds were synthesized, characterized, and subjected to cytotoxicity studies across multiple cancer cell lines, alongside assessments of PIM1 inhibition and apoptosis induction. Among the tested molecules, compound 6h demonstrated the most promising antiproliferation activity in HT-29, PC3, and A549 cells, inducing apoptosis in HT-29 cells. It inhibited PIM1 kinase activity by 34.5% ± 3.4% at 10 µM and exhibited favorable in silico ADME and drug-likeness profiles, including high predicted absorption and absence of hepatotoxicity. These findings establish 6h as a promising lead scaffold for the development of PIM1-1 targeted anticancer agents. The integration of pharmacophore modeling, machine learning prediction, and experimental validation highlights a viable strategy for accelerating kinase inhibitor discovery.
1 Introduction
Cancer is identified as a menace to human beings among the plethora of diseases. In spite of huge investments, the fatal nature endures, with finite treatment outcomes. Despite huge efforts over decades on anticancer drug discovery, copious agents failed to eradicate tumor completely while maintaining minimal adverse effects and toxicity.1,2 The modern era of anticancer drug discovery is based on dysregulating the function of kinases such as non-programmed proliferation and angiogenesis towards cancer cells.3,4 Kinase drug discovery has led to a constellation of anticancer agents for the treatment of numerous cancers, among which serine/threonine and tyrosine classes of kinase are in the spotlight.5PIM kinases are part of the serine/threonine kinase family, playing a vital role in cell proliferation and survival, followed by factors such as growth-factor-based activation of NF-κB and JAK/STAT, cytokines and stress.6,7 PIM phosphorylates various pro-oncogenic proteins such as BAD, CDC25A/C, and 4EBP1 and thus is responsible for cell survival, progression, growth, oncogenesis and migration.8 Alterations in the basic architecture of PIM kinases lead to the development of cancer. PIM-1, PIM-2 and PIM-3 kinases are mostly detected in solid and haematological malignancies (prostate, hepatocellular, and pancreatic tumours, acute myeloid leukaemia and multiple myeloma).9–14
The crystal structure of PIM kinases constitutes a unique hinge area, which is a part of the ATP binding motif, consisting of proline, which is not observed in other kinases. This unique structure provides a handle for the development of new inhibitors targeting PIM kinases.15 All three isoforms of PIM kinases are structural analogues, but PIM-2 has a lower binding affinity (Km: approx. 4 µM) for ATP than PIM-1 and PIM-3 kinases.16 Multitudinous PIM kinase inhibitors have been documented in the past two decades, among which SGI-1776, AZD-1208, PIM447 and INCB053914 are potent pan-PIM kinase inhibitors17–19 that have been put through the clinical pipeline but could not pass the trials due to persistant QTc prolongation, as shown in Fig. 1. Recently, a maximum number of ATP-competitive (Type-2) PIM kinase inhibitors have been developed instead of ATP mimetics due to their clinical significance, exemplified by TP-3654.20
Despite the numerous PIM1 kinase inhibitors explored in clinical trials, not a single molecule has been in clinical use in the treatment of cancer. Hence, this poses an enormous challenge for the discovery of new potential lead molecules. Our group has been consistently involved in exploring the structural information of kinases, especially PIM1, P38, GSK3β and EGFR and in searching for new kinase inhibitors using QSAR and machine leaning approaches.21–27
Numerous studies have explored computational tools for understanding the development of new PIM1 kinase inhibitors, including newer approaches like machine learning techniques. Such approaches eventually unearthed newer PIM1 kinase inhibitors, including some from natural sources.28–30
Recently, we applied a combination of computational approaches, in which a new lead was identified by pharmacophore and derivatized using the python/RdKit enumerator tool for analogues. A ML-based model predicted the PIM1 inhibitory activity of the molecules.31 This research manuscript is in continuation of our efforts towards the development of kinase inhibitors, reporting one such series of molecules bearing the imidazo[1,2-a]pyridin-2-yl-phenyl benzamide scaffold, as potential PIM1 kinase inhibitors. The design strategy included a complex pharmacophore utility that identified the imidazopyridine scaffold, a Python script that generated twenty-seven new derivatives with appropriate substitution and a linear-regression-based ML model that predicted the activity, which was confirmed by molecular docking and dynamics simulation studies, as well as the in vitro PIM1 kinase inhibitory activity.31
2. Experimental section
2.1. Materials
All reagents and solvents were collected from commercial suppliers, including Sigma-Aldrich (Merck), Tokyo Chemical Industry (TCI), and Carbanio (India) and utilized as received without further purification. Synthetic procedures were conducted in carefully oven-dried borosilicate glassware. The reaction progress was checked via thin layer chromatography (TLC) using Silica Gel 60 GF254 plates, with visualization accomplished using UV light or iodine staining.Compound characterization was performed as follows: The melting points were determined using a Veego VMP-PM digital machine and are presented as uncorrected. Infrared (IR) spectra were captured on a PerkinElmer FT-IR (ATR) spectrometer (results in cm−1). Nuclear magnetic resonance (NMR) spectra (1H at 500 MHz and 13C) were acquired on a Bruker Avance III HD system using DMSO-d6. Chemical shifts (δ) are cited in parts per million (ppm) in comparison to the solvent signal, with multiplicities designated as s, d, t, q, m, and br. Mass spectrometry (LC-MS/MS) using an Agilent Triple Quadrupole instrument was employed to confirm the identity of all intermediate and target molecules. Molecular modeling calculation studies were accomplished exclusively on desktops with the Windows 10 operating system, and molecular dynamics simulations were carried out on an Ubuntu 22.04 LTS desktop.
2.2. Pharmacophore-based virtual screening and machine-learning-based PIM1 kinase activity predictions
2.3. Method of synthesis
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretch), 1558 (-CH bending), 1520 (-NO2 symmetric stretch), 1342 (-NO2 asymmetric stretch), 717; 1H NMR (500 MHz, DMSO) δ 8.51 (d, J = 7.8 Hz, 1H, amide NH), 8.23 (s, 1H ArH), 7.54 (d, J = 9.0 Hz, 1H, ArH), 7.28–7.18 (m, 2H, ArH), 7.13–7.02 (m, 2H, ArH), 6.87 (td, J = 6.7, 1.1 Hz, 1H, ArH), 6.56–6.49 (m, 1H, ArH); LC/MS: calculated C13H9N3O2 [M + 1]+: m/z 239.23; observed, 239.90 (deconvoluted).
N stretch), 1558 (-CH bending), 1520 (-NO2 symmetric stretch), 1342 (-NO2 asymmetric stretch), 717; 1H NMR (500 MHz, DMSO) δ 8.51 (d, J = 7.8 Hz, 1H, amide NH), 8.23 (s, 1H ArH), 7.54 (d, J = 9.0 Hz, 1H, ArH), 7.28–7.18 (m, 2H, ArH), 7.13–7.02 (m, 2H, ArH), 6.87 (td, J = 6.7, 1.1 Hz, 1H, ArH), 6.56–6.49 (m, 1H, ArH); LC/MS: calculated C13H9N3O2 [M + 1]+: m/z 239.23; observed, 239.90 (deconvoluted).![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretch), 1608, 1585 (-CH bending), 744; 1H NMR (500 MHz, DMSO) δ 8.51 (d, J = 7.8 Hz, 1H, ArH), 8.23 (s, 1H, ArH), 7.54 (d, J = 9.0 Hz, 1H, ArH), 7.28–7.18 (m, 2H, ArH), 7.13–7.02 (m, 2H, ArH), 6.87 (td, J = 6.7, 1.1 Hz, 1H, ArH), 6.56–6.49 (m, 1H, ArH), 5.13 (s, 2H, NH2); 13C NMR (500 MHz, DMSO) δ 149.35 (ArCNH2), 145.67 (ArC
N stretch), 1608, 1585 (-CH bending), 744; 1H NMR (500 MHz, DMSO) δ 8.51 (d, J = 7.8 Hz, 1H, ArH), 8.23 (s, 1H, ArH), 7.54 (d, J = 9.0 Hz, 1H, ArH), 7.28–7.18 (m, 2H, ArH), 7.13–7.02 (m, 2H, ArH), 6.87 (td, J = 6.7, 1.1 Hz, 1H, ArH), 6.56–6.49 (m, 1H, ArH), 5.13 (s, 2H, NH2); 13C NMR (500 MHz, DMSO) δ 149.35 (ArCNH2), 145.67 (ArC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 145.06 (ArC), 134.85 (ArC), 129.58 (ArC), 127.20 (ArC), 125.06 (ArC), 116.97 (ArC), 113.97 (ArC), 113.95 (ArC), 112.47 (ArC), 111.70 (ArC), 109.08 (ArC); LC/MS calculated C13H11N3 [M + 1]+: m/z 209.25; observed, 208.99 (deconvoluted).
N), 145.06 (ArC), 134.85 (ArC), 129.58 (ArC), 127.20 (ArC), 125.06 (ArC), 116.97 (ArC), 113.97 (ArC), 113.95 (ArC), 112.47 (ArC), 111.70 (ArC), 109.08 (ArC); LC/MS calculated C13H11N3 [M + 1]+: m/z 209.25; observed, 208.99 (deconvoluted).2.4. Biological assays
2.4.2.1. Antiproliferative activity. The antiproliferative capabilities of the synthesized compounds were gauged using the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide) assay. In our previous publication, we reported that PIM-1 kinase expression was noted at elevated levels in carcinomas, including prostate, colon and lung cancers. Due to this reason, human cancer cell lines such as PC3, HT29 and A549 were selected.31 PC3, A549 and HCT-29 prostate, lung, and colon cancer cells, respectively, were cultured in 96-well plates using DMEM supplemented with 10% FBS. When the cell confluence was sufficient (up to 70%), the cells were treated with different concentrations of compounds, including the control, which contained DMSO for 24 h. The cells were washed with phosphate buffered saline after treatment with the compounds, and media aspiration was completed. In order to examine the viability of the cells, MTT was added (0.5 mg mL−1) to each well and the MTT-treated cells were incubated at 37 °C for 4 h in a dark environment. Upon the completion of incubation, the MTT staining solution was removed and replaced with 200 µL of DMSO and the plates were shaken at 180 rpm for 10 minutes in order to make a clear solution, which facilitates the optical density measurement at 570 nm using a plate reader (Benesphera 21). To achieve better reliability of the activity data, this entire process was performed in triplicate before performing the statistical computations and visualization of the results.34,35
2.4.2.2. PIM-1 kinase inhibitory activity. The PIM-1 kinase inhibitory activity of the molecules was assessed using the luminescence-based ADP-Glo™ kinase Assay. The diluted enzyme, test compounds and ATP kinase buffer were transferred to a 384-well plate and incubated for 60 minutes. 5 µl of ADP-Glo™ reagent was also mixed into the wells and further incubated at RT for 40 minutes. Followed by the addition of ADP-Glo reagent, 5 µl of kinase detection reagent was added and incubated for 30 minutes, and the luminescence was recorded with a luminometer. Further, the % inhibition of the compounds was determined.36
2.4.2.3. Apoptosis assay employing Alexa Fluor® 488 Annexin V examination. Upon confluence of the HT-29 cells (70–80%), the cells were seeded in a sterile 12-well plate. The 24 h seeded cells were treated with the apoptosis-inducing agent (provided drugs) as per the provided drug concentrations, with half media change. A negative control was prepared by incubating the cells without the apoptosis-inducing agent. The treated cells were harvested by the trypsinization protocol and the harvested cells were properly washed with PBS. The washed cells were once again suspended in 1× Annexin binding buffer, further treated with Annexin V dye (2 µL) and subsequently incubated for 40 minutes at 2–8 °C. Post incubation, the test mixture with cells was centrifuged at 1500 rpm using 1X PBS for 5 minutes, and the cells were resuspended in PBS. Staining dye PI at 100 µg mL−1 was added just before sample acquisition on a flow cytometer and the samples were kept over ice. Apoptosis of the cells was assessed based on dye pigmentation on the flow cytometer. Annexin V detection was accomplished using an Attune Nxt flow cytometer with dual detection wavelengths at 488 nm with Ex/Em of 499/521. Ex/EM of 535/617 was employed for PI recognition. The green fluorescence of Annexin V-FITC/Alexa Fluor® 488 was recorded from the FITC channel (FL1 or BL1), whereas the red fluorescence was detected from the PI channel (FL2 or BL2 or sometimes FL3 or BL3).31,37
2.5. Molecular modelling analysis
3 Results and discussion
3.1. Molecule hybridization
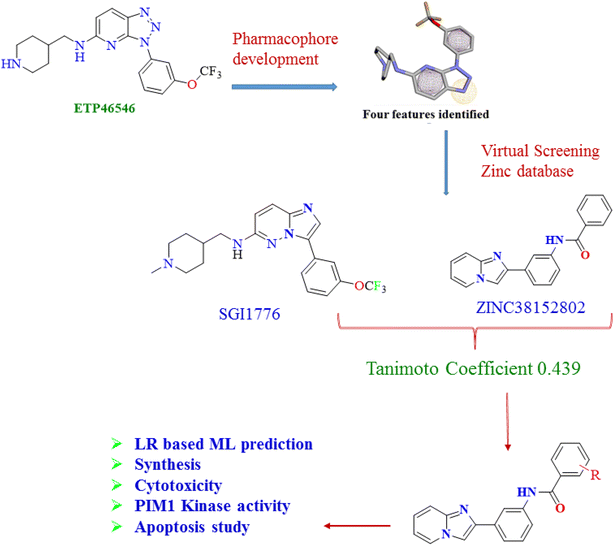
In the past few decades, plentiful PIM kinase inhibitors have been discovered, including SGI-1776 and AZD-1208, which were found to possess potent PIM kinase inhibitory strength but failed due to an unavoidable cardiac toxicity profile for SGI-1776 in the clinical trial phase-1. The design of imidazopyridine amides was based on multiple approaches involving pharmacophore and database search, ML activity prediction and evaluation of the scaffold hopping attributes using molecular docking and MD simulations.3.2. Pharmacophore postulate and database virtual screening
ETP46546 is a PIM-1 kinase inhibitor with an IC50 of 6 nM for type-2/ATP non-competitive inhibition. It constitutes the 1,2,3-triazolo[4,5-b]pyridine and piperidine rings vital for selective PIM-1 kinase inhibition, which was enhanced by substitutions of the alkyl functionalities affixed to the phenyl ring. We derived four featured pharmacophores, namely aromatic rings (A), two hydrophobic interactions, and a hydrogen bond acceptor (HA), as depicted in Fig. 2. Furthermore, the established pharmacophores were queried against the ZINC15 database using the Pharmit server to determine the virtual hits. Fifteen thousand hits were filtered from the database, with numerous aromatic and heterocyclic scaffolds, namely, tetrahydrobenzothiophene, imidazopyridine, benzothiophene and others. They were sorted based on the score, from which ZINC38152802 was identified, with a score of −9.21 kJ mol−1, which was encouraging as compared to the other hits.313.3. Similarity search
In order to verify the pharmacophore search and resultant molecule ZINC38152802, Tanimoto similarity and fingerprint search were performed by running an in-house Python script. Such similarity search has been successfully applied by us, which enhanced the confidence towards selecting these molecules.31 The comparison was accomplished against SGI1776, a known inhibitor with a PIM1 kinase inhibitory activity of 7 nM. The Tanimoto similarity determined by using MACCS keys with 166 bits had a moderate similarity coefficient of 0.439. This moderate similarity suggests approximately 44% substructural common features between ZINC38152802 and SGI1776.| Type | Iteration | R2 | Predicted R2 | Q2 |
|---|---|---|---|---|
| DFG in | 121 | 0.912![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 683 683 |
0.742![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 594 594 |
0.549![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 658 658 |
| DFG in | 1080 | 0.910![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 914 914 |
0.761![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 765 765 |
0.533![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 005 005 |
| DFG out | 871 | 0.929![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 642 642 |
0.740![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 439 439 |
0.63 |
| DFG out | 2018 | 0.908![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 237 237 |
0.812![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 169 169 |
0.61 |
Previously, we reported the development of QSAR models in support of structure-based docking and molecular dynamics simulation21 for predicting the activity of new molecules based on the ML model. In continuation of our previous research developments [unpublished data from our research], a Python script was executed by feeding SMILE strings of ZINC38152802, and monosubstitution was assigned on the 2nd, 3rd and 4th positions of the phenyl group of the amidic bridge, which generated twenty-six new analogues.31 The descriptors for the enumerated molecules were generated using the RdKit module, and the required descriptors were selected. The predicted activities of the enumerated molecules have were determined by employing the models listed in Table 1, and the ML-predicted activities are summarized in Table 2.
| Position | Substituent | DFG In | DFG out |
|---|---|---|---|
| 2 | C | 7.50 | 5.50 |
| 2 | OC | 7.14 | 5.62 |
| 2 | Cl | 7.48 | 5.38 |
| 2 | F | 7.30 | 5.37 |
| 2 | N | 6.69 | 5.40 |
| 2 | C(F)(F)F | 6.96 | 5.44 |
| 2 | OC(F)(F)F | 6.60 | 5.61 |
| 2 | c1ccc2ccccc2c1 | 8.28 | 5.37 |
| 2 | Br | 6.00 | 5.98 |
| 3 | C | 7.48 | 5.50 |
| 3 | OC | 7.14 | 5.65 |
| 3 | Cl | 7.48 | 5.43 |
| 3 | F | 7.30 | 5.40 |
| 3 | N | 6.67 | 5.43 |
| 3 | C(F)(F)F | 6.94 | 5.47 |
| 3 | OC(F)(F)F | 6.60 | 5.64 |
| 3 | c1ccc2ccccc2c1 | 8.28 | 5.37 |
| 3 | Br | 6.00 | 6.01 |
| 4 | C | 7.49 | 5.50 |
| 4 | OC | 7.14 | 5.65 |
| 4 | Cl | 7.48 | 5.43 |
| 4 | F | 7.30 | 5.40 |
| 4 | N | 6.67 | 5.43 |
| 4 | C(F)(F)F | 6.94 | 5.47 |
| 4 | OC(F)(F)F | 6.60 | 5.64 |
| 4 | c1ccc2ccccc2c1 | 8.28 | 5.37 |
| 4 | Br | 6.00 | 6.01 |
3.4. Discrimination of PIM-1 kinase inhibitors (Type-2) postulated by a structure-based molecule design strategy
Triazolopyridines constituting PIM-1 kinase inhibitor ETP46546 with an IC50 of 6 nM were the input for the development of the pharmacophore, which eventually identified ZINC38152802 as a new hit. ZINC38152802 was further derivatized and twenty-five such analogs were designed. These twenty-five molecules were docked within the crystal structure of PIM-1 kinase (PDB ID: 4A7C), from which eighteen molecules demonstrated affirmative binding interactions comparable to that of ETP46546.31 All eighteen molecules interacted through hydrogen interactions with Lsy67, Asp128 and Asp186 of the binding pocket of PIM-1 kinase, as depicted in Fig. 3.The PIM-1 kinase crystal structure holds a unique active site with Pro123 in the hinge region, which generates a bulge in the binding site and provides a handle for new PIM-1 kinase (Type-2) inhibitor development. Binding site exploration of PIM-1 kinases grounded on molecular docking of PIM-1 kinase inhibiting molecules (LGB321, ETP46546, AZD-1208 and SGI-9481) revealed that the type-II inhibitors interact with the binding area of the receptor, and its active pocket is divided into four regions: hinge, catalytic, DFG motif (hydrogen bond opulent area) and side chain region (hydrophobic area).
3.5. Chemistry
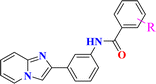
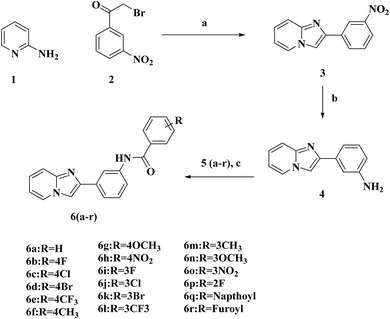
The construction of 3 (2-(3-nitrophenyl)imidazo[1,2-a]pyridine) was undertaken by the reaction between 2-amino pyridine 1 and 2-bromo-3′-nitroacetophenone 2 in the presence of NaHCO3 as a base in acetonitrile. Further, 3 was reduced with zinc and HCl, which selectively reduced the primary amine to the amino intermediate 4. 4 was further derivatized to numerous commensurable amides 6a–r with substituted benzoyl chlorides 5a–r in a Schotten–Baumann reaction, and the physical attributes of the target molecules are displayed in Table S1 (SI).Confirmation of the chemical structure of the synthesized molecules was in concordance with their corresponding FT-IR, 1H and 13C NMR, ESI-MS spectral details. FT-IR evaluation revealed –C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching in the range of 1600–1618 cm−1. The peak at 1640–1678 cm−1 was assigned to amide –C
N stretching in the range of 1600–1618 cm−1. The peak at 1640–1678 cm−1 was assigned to amide –C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, 3376–3239 cm−1 to amide –NH stretching and 3050–3084 cm−1 to aromatic –CH stretching. In the proton NMR analysis, amide –NH protons were identified as singlets at δ values of 10.26–10.69 ppm; aromatic and heteroaromatic hydrogen were observed as multiplets from 8.92–7.37 ppm. The signals of the methyl and methoxy functional-group protons were located at 2.41 and 3.86 ppm as singlets. Moreover, carbon NMR indicated a carbonyl amide signal in the range of 156.76–167.85 ppm and heteroaromatic and aromatic carbon signal at 145.23–109.60 ppm. The entire series of molecules was delineated based on the corresponding molecular ion peaks at the discrete molecular weights.
O, 3376–3239 cm−1 to amide –NH stretching and 3050–3084 cm−1 to aromatic –CH stretching. In the proton NMR analysis, amide –NH protons were identified as singlets at δ values of 10.26–10.69 ppm; aromatic and heteroaromatic hydrogen were observed as multiplets from 8.92–7.37 ppm. The signals of the methyl and methoxy functional-group protons were located at 2.41 and 3.86 ppm as singlets. Moreover, carbon NMR indicated a carbonyl amide signal in the range of 156.76–167.85 ppm and heteroaromatic and aromatic carbon signal at 145.23–109.60 ppm. The entire series of molecules was delineated based on the corresponding molecular ion peaks at the discrete molecular weights.
Reagents and conditions: (a) acetonitrile, NaHCO3, stirring at RT for 2 h; (b) Zn, HCl, deionized water, reflux; (c) substituted benzoyl chlorides, DCM, TEA, RT.
3.6. In vitro antiproliferative activity
The MTT assay stands is a standard, accurate and appropriate colorimetric method employed for the quantitative assessment of cell proliferation, viability, and activation. Concurrently, PIM kinases have emerged as pivotal players in the realm of tumor cell proliferation. In this context, an investigation into the cytotoxicity potential of the synthesized target compounds was undertaken, with the determination of the IC50 values across PC3, HT29, and A549 cell lines. Utilizing 5-fluorouracil (5FU) as a positive control, the results revealed that the target molecules exhibited notable potency against the PC3 cell line, with IC50 values ranging from 29.95 to 49.92 µM, while 5FU demonstrated comparable potency, possessing an IC50 value of 40.96 µM.Furthermore, the compounds were screened using the HT29 cell line, where they demonstrated varying degrees of activity within the range of 16 to 47 µM. In contrast, 5FU exhibited potent activity against this cell line, with an IC50 value of 43 µM. Similarly, the compounds were evaluated against the A549 cell line, revealing an admirable antiproliferative effect, with IC50 ranging from 25.49 to 57.38 µM. Notably, the positive control 5FU, exhibited an IC50 value of 42.08 µM (Table 3).
| Molecule Id | PC3 | A549 | HT29 | PCS100–024 | SI |
|---|---|---|---|---|---|
| IC50 (µM)b | |||||
| a Positive control.b IC50 (µM) ± SD of triplicates, $: IC50 of PCS100–024/IC50 of HT29. | |||||
| 6a | 47.92 ± 1.25 | 25.49 ± 1.59 | 35.22 ± 1.57 | n.d | n.d |
| 6b | 29.95 ± 1.5 | 65.86 ± 3.0 | 20.34 ± 0.87 | 57.67 ± 1.62 | 2.835 |
| 6c | 44.2 ± 1.56 | 31.42 ± 1.49 | 38.1 ± 1.86 | n.d | n.d |
| 6d | 44.66 ± 2.33 | 51.87 ± 1.71 | 41.98 ± 1.74 | n.d | n.d |
| 6e | 42.48 ± 0.57 | 46.63 ± 1.1 | 47.05 ± 0.99 | n.d | n.d |
| 6f | 48.45 ± 2.43 | 41.01 ± 1.1 | 39.05 ± 1.27 | n.d | n.d |
| 6g | 40.98 ± 2.27 | 34.96 ± 2.01 | 26.17 ± 2.1 | n.d | n.d |
| 6h | 30.89 ± 1.93 | 42.02 ± 3.22 | 18.1 ± 0.44 | 58.91 ± 2.04 | 3.254 |
| 6i | 48.52 ± 1.45 | 57.38 ± 2.84 | 42.16 ± 1.09 | n.d | n.d |
| 6j | 47.33 ± 1.54 | 57.38 ± 2.84 | 26.33 ± 1.75 | n.d | n.d |
| 6k | 38.89 ± 2.11 | 41.26 ± 1.93 | 45.3 ± 1.31 | n.d | n.d |
| 6l | 42.58 ± 1.36 | 39.88 ± 1.58 | 34.51 ± 1.62 | n.d | n.d |
| 6m | 33.73 ± 2.37 | 40.0 ± 1.9 | 25.22 ± 0.82 | n.d | n.d |
| 6n | 44.46 ± 1.65 | 37.54 ± 1.65 | 34.13 ± 2.02 | n.d | n.d |
| 6o | 43.4 ± 2.88 | 42.76 ± 3.08 | 31.37 ± 0.55 | n.d | n.d |
| 6p | 49.93 ± 1.32 | 37.33 ± 1.02 | 16.41 ± 1.5 | 54.13 ± 1.8 | 3.298 |
| 6q | 39.63 ± 3.55 | 39.77 ± 1.24 | 32.51 ± 2.53 | n.d | n.d |
| 6r | 49.33 ± 2.86 | 42.66 ± 1.67 | 24.05 ± 1.34 | n.d | n.d |
| 5FUa | 40.86 ± 1.56 | 42.08 ± 1.1 | 44.11 ± 1.56 | 43.53 ± 1.7 | 0.986 |
Among the target analogs, compound 6b exhibited notable potency, with an IC50 value of 29.25 µM, showcasing encouraging efficacy against the PC3 cell line. Conversely, compounds 6p (49.92 µM) and 6r (49.32 µM) demonstrated comparatively lower potency against PC3, in contrast to the positive control 5FU, which exhibited an IC50 value of 40.96 µM. Intriguingly, compounds 6p (18 µM) and 6h (16 µM) displayed excellent potential in inhibiting HT29 cell proliferation. On the other hand, compounds 6d (42 µM) 6e (74 µM), 6i (42 µM) and 6k (45 µM) exhibited lower activity against the HT29 cell line compared to 5FU (43 µM).
Besides, in the case of the A549 cell line, compounds 6a (25.49 µM) and 6c (31.41 µM) demonstrated improved activity over 5FU (42.08 µM). This underscores the promising antiproliferative effects of these specific analogs across different cell lines. The results suggest a nuanced spectrum of potency among the synthesized compounds, emphasizing their potential as candidates for further investigation and development in the context of cancer cell proliferation inhibition (Fig. 4).
In continuation of antiproliferative studies on three cancerous cell lines, three compounds 6b, 6h and 6p, along with the standard 5-fluorouracil, were screened against the normal cell line PCS1000-024 in order to explore the selectivity index. The three tested molecules demonstrated IC50 values of 57.67, 58.91 and 54.13 µM, compared to 43.53 µM for 5FU.31 All three exhibited greater IC50 in comparison to 5FU. The selectivity index is the ratio of IC50 for the normal cell line versus that of the HT29 cell line, and a SI value greater than 2.0 is essentially a favorable value. The SI values calculated for the three tested molecules were found to be 2.835, 3.254 and 3.298, which are favorable values, while 5FU showed the calculated SI value of 0.986.
3.7. In vitro PIM-1 kinase inhibitory activity
The PIM-1 kinase inhibitory assay is based on chemiluminescence, which was found to be the most precise method for evaluation. Herein, an evaluation of the enzyme inhibitory potential against PIM-1 kinase for the target molecules was performed to determine the % inhibition at a concentration of 10 µM by employing staurosporine as a standard drug. The results demonstrated inhibition of PIM-1 in the range of 9.8% ± 1.1% to 34.5% ± 3.4%, compared with the standard staurosporine (70.2% ± 1.1%).Furthermore, the synthesized compounds 6h and 6i demonstrated excellent inhibitory activity of 34.5% ± 3.4% and 32.7% ± 2.5%. Molecules 6k to 6p were spotted as moderately active, with % inhibitory potentials in the range of 22.9% ± 1.0% to 28.0% ± 2.3%, where 6n and 6o revealed a similar inhibitory activity of 26.5% ± 2.7% and 26.8% ± 2.1% with a minimal SE.
Compound 6f was noted with the lowest inhibitory effect of 9.8% ± 1.1% from the series of synthesized compounds, as compared to the positive control with a corresponding % inhibition of 70.2% ± 1.1%, as depicted in Table 4.
| Molecule Id | PIM-1 kinase % inhibition at 10 µM |
|---|---|
| 6a | 11.6 ± 2.1 |
| 6b | 13.0 ± 2.4 |
| 6c | 17.0 ± 1.9 |
| 6d | 14.1 ± 0.3 |
| 6e | 13.0 ± 1.3 |
| 6f | 9.8 ± 1.1 |
| 6g | 22.1 ± 2.8 |
| 6h | 34.5 ± 3.4 |
| 6i | 32.7 ± 2.5 |
| 6j | 10.0 ± 1.9 |
| 6k | 22.9 ± 1.0 |
| 6l | 21.9 ± 1.1 |
| 6m | 25.4 ± 2.6 |
| 6n | 26.5 ± 2.7 |
| 6o | 26.8 ± 2.1 |
| 6p | 28.0 ± 2.3 |
| 6q | 19.6 ± 1.0 |
| 6r | 18.2 ± 2.4 |
| Staurosporine | 70.2 ± 1.1 |
3.8. Apoptosis induction in HT-29 cells
Apoptosis is a regulated form of planned cell death that plays an indispensable role in normal development and tissue homeostasis. Any aberration in this process eventually leads to various pathological conditions, including Alzheimer's disease and malignancies like cancer. Apoptosis is characterized by distinct morphological and biochemical changes, in contrast to necrosis, which is a form of accidental cell death. Key features of apoptosis include chromatin condensation and fragmentation, cytoplasm shrinkage, and alterations in membrane asymmetry. These changes collectively facilitate the orderly dismantling and removal of cells without eliciting an inflammatory response.In order to study whether tumour cells treated with the control and target compounds show apoptosis, HT-29 cells were smeared using Annexin V and PI. FACS (Fluorescence Assisted Cell Sorting) can separate cells into four parts viz., Annexin V– PI- (live cells); Annexin V + PI- (early apoptotic cells); Annexin V + PI+ (late apoptotic cells) and Annexin V– PI + (necrotic cell). To evaluate the antiproliferative effects of the compounds on the HT-29 cell line, 6h and 6p were selected for the assay. 6h and 6p demonstrated IC50 of 9 and 16 µM, respectively, resulting in a high count of necrotic cells (28.59, 74.05%) compared to the untreated cells (<1%). Upon treatment with 6p, 74.05% necrotic cells and 7.23% late apoptotic cells were identified, indicating that 6p induces apoptosis of HT-29 cells, as illustrated in Fig. 5.
 | ||
| Fig. 5 Flow cytometry analysis of 6h and 6p for the determination of apoptosis in the HT-29 cell line, analysis from FACS with Annexin V vs. PI on HT-29 (a) control, (b) 6h and (c) 6p. | ||
3.9. Molecular docking analysis of target compounds
Molecular docking analysis was explored to identify the binding mode of the designed kinase inhibitors to the active site of PIM-1 kinase. To validate the docking protocol, the co-crystal ligand (ETP46546) was re-docked, and both the docked conformation and crystal ligand conformation were aligned, which displayed RMSD = 0.25 Å, as depicted in Fig. 6. All the tested compounds were docked in the active site of the PIM-1 kinase, and their corresponding docking scores are summarized in Table 5.| Comp. no. | Docking energy (kcal mol−1) | Hydrogen bond contacts | Electrostatic and hydrophobic interactions |
|---|---|---|---|
| ETP46546 (Co-crystal ligand) | −8.9 | Lys67 and Asp128 | Leu44, Val52, Phe49, Ala65, Ile104, Val126, Leu120, Glu171, and Ile185 |
| 6a | −9.8 | Lys67 and Asp186 | Leu44, Phe49, Val52, Ala65, Ile104, Leu120, Val126, Leu174 and Ile185 |
| 6b | −10.0 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Val126, Leu120, Glu171, and Ile185 |
| 6c | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Val126, Leu120, Glu171, and Ile185 | |
| 6d | −10.1 | Lys67 and Asp186 | Leu44, Phe49, Val52, Ala65, Ile104, Leu120, Val126, Glu171, and Ile185 |
| 6e | −10.4 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Val126, Leu120, Glu171, Ile185, Val126, Asp128, Glu171, Leu174 and Ile185 |
| 6f | −10.2 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Val126, Leu120, Glu171, and Ile185 |
| 6g | −9.8 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Arg122, Val126, Glu171, Asn172, Leu174 and Ile185 |
| 6h | −10.0 | Lys67, Glu171 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Glu171, and Ile185 |
| 6i | −10.1 | Lys67 and Asp186 | Leu44, Gly45, Phe49, Val52, Ala65, Leu120, Ile104, Arg122, Val126, Asn172, Glu171, Leu174 and Ile185 |
| 6j | Lys67 and Asp186 | Leu44, Gly45, Phe49, Val52, Ala65, Ile104, Leu120, Arg122, Val126, Glu171, Asn172, Leu174 and Ile185 | |
| 6k | −10.2 | Lys67 and Asp186 | Leu44, Gly45, Phe49, Val52, Ala65, Ile104, Leu120, Val126, Leu174 and Ile185 |
| 6l | −10.6 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Leu174 and Ile185 |
| 6m | −10.5 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Leu174 and Ile185 |
| 6n | −10.0 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Glu171, Leu174 and Ile185 |
| 6o | −9.7 | Lys67,Glu171, Asn172and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Glu171, Leu174 and Ile185 |
| 6p | −9.8 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Glu171, Leu174 and Ile185 |
| 6r | −9.1 | Lys67 and Asp186 | Leu44, Val52, Phe49, Ala65, Ile104, Leu120, Val126, Glu171, Leu174 and Ile185 |
The most potent molecule 6p, with an IC50 value of 16 µM for HT29, docked in the binding pocket of PIM-1 kinase, with docking energy = −9.8 kcal mol−1. The amide functional group formed polar hydrogen bonds with Lys67 (2.1 Å) and Asp186 (2 Å) in the catalytic region and with the DFG motif. The spacer phenyl ring established π-sigma interaction with Ile185 (3.9 Å). Moreover, the 2-fluoro phenyl ring generated π–π stacking hydrophobic interactions with Phe49 (4 Å) in the side pocket. In the lipophilic groove, the imidazopyridine ring was surrounded by Leu44 (3.90 Å), Ala65 (4.9 Å), Val126 (4.7 Å) and Leu174 (4.9 Å) and established π-alkyl contacts, as depicted in Fig. 7.
The active compound 6a with an IC50 value of 25.49 µM for A549 resided in the active pocket of the protein, with a docking energy of −9.8 kcal mol−1. The carbonyl amide functionality generated hydrogen bond contacts with Lys67 (2.09 Å) and Asp186 (2.2 Å) in the catalytic and DFG motifs. The imidazopyridine and the central phenyl ring resided in the hydrophobic pocket, surrounded by amino acids, namely, Leu44 (3.90, 4.88 Å), Ala65 (5.1, 4.8 Å), Val126 (4.7 Å) and Leu174 (4.9 Å), and established π-alkyl contacts. The phenyl ring of the amide group generated π–π stacking with Phe49 (4.1 Å), as shown in Fig. 8.
Compounds 6l and 6m, with docking energies of −10.6 and −10.5 kcal mol−1, respectively, well occupied the active cavity of PIM-1 kinase. The carbonyl oxygen of the amide functionality (6l and 6m) established two hydrogen bond interactions with the hydrogen on the terminal nitrogen of Lys67 (2.1 Å) and Asp186 (2.2 Å) in the catalytic region and DFG motif. The imidazopyridine and central phenyl rings were precisely placed in the hydrophobic cavity, forming hydrophobic π-alkyl contacts with Leu44 (4.91 Å), Val126 (4.7 Å) and Leu174 (5.9 Å), as well as electrostatic π-sigma interactions with Ile185 (5.16 Å). The 3-trifluoromethylphenyl and 3-methylphenyl moieties generated π–π stacking and π-alkyl interactions with Phe47 (5.8 Å) in the corner hydrophobic region of the kinase, as shown in Fig. 9.
Molecule 6e was perfectly positioned in the active-site pocket of PIM-1 kinase; the oxygen of the amide carbonyl formed two polar hydrogen bond interactions with Lys67 (2.1 Å) in the catalytic groove and Asp186 (2.9 Å) in the DFG motif. The imidazopyridine and central phenyl ring occupied the hydrophobic region of the protein, and the imidazopyridine fragment generated π-alkyl contacts with Leu44 (4.84, 4.87 Å), Val52 (6.77 Å), Val126 (4.72 Å) and Leu174 (5.99 Å). Moreover, the phenyl ring connected to the imidazopyridine nucleus established π-sigma and π-alkyl interactions with Ala65, Leu120, and Ile185. The trifluoromethyphenyl flanked with amide functional groups established π-sigma and π–π stacking contacts with Ile185 and Phe49 in the side hydrophobic pocket, with a corresponding docking energy of −10.4 kcal mol−1, as depicted in Fig. 10.
Molecules 6f and 6k were properly positioned in the binding area of PIM-1 kinase, with a docking energy of −10.2 kcal mol−1. The amide functionality of both compounds established hydrogen bond contacts with Lys67 (2.08, 2.10 Å) and Asp186 (3.2 Å) in the catalytic and DFG motifs. The imidazopyridine fragment of both compounds, located in the hydrophobic pocket, formed π-alkyl contacts with Leu44 (4.87 Å), Val52 (5.50 Å), Ala65 (5.20, 4.80 Å), Ile104 (5.38, 5.50 Å), Leu120 (5.11 Å), and Val126(5.23, 4.88 Å), as well as π-sigma interactions with Ile185(3.9 Å). Similar interactions were noticed for the central phenyl ring attached to the imidazoypridine. The 3-Br and 4-CH3 phenyl moieties established π–π stacking interactions with Phe49 (4.05, 4.09 Å) in the side hydrophobic cavity, as depicted in Fig. 11.
Compounds 6h and 6o having a –NO2 functionality at the para and meta positions are the isomeric forms of each other and reside in the active pocket of PIM-1 kinase, with corresponding docking energies of −10.0 and −9.7 kcal mol−1. Imidazoypridine and the spacer phenyl ring of the isomers developed electrostatic π-sigma interactions with Leu44 (3.98 Å) and Ile185 (3.92, 3.96 Å) but the 4-NO2 phenyl ring also established an additional π-sigma contact with Ile185 (3.99 Å) and π–π stacking interaction with Phe49 (3.98, 4.10 Å) in the corner side hydrophobic cavity. The amide functionality of the isomers formed polar hydrogen bond contacts with Lys67 (2.09 Å) and Asp186 (3.4 Å). Moreover, the –NO2functional group generated three polar hydrogen bond contacts with Glu171 (3.4 Å) and Asn172 (3.1 Å) in the side-chain region, as shown in Fig. 12.
 | ||
| Fig. 13 (a–c) RMSD graphical representations of PIM-1 kinase, 6h and the protein-6h complex; (d) RMSF plot; (e) Rg graph; and (f) hydrogen bond plot. | ||
The RMSD of macromolecule ligand (6h) was also analysed, indicating that in the inceptive phase, RMSD spikes were noticed from 3 and increased to 6 Å at 5 ns. A sudden downfall in the projections was observed at 2–3 Å in the timeframe of 6 to 25 ns; furthermore, a tangential increase in the RMSD at 3.5–4 Å was observed at 30 ns, indicating a moderate conformational change in 6h. From 35 to 45 ns, a sudden rise in the ligand RMSD projections to 6 Å was observed, followed by a quick downfall to 3 Å. From 46–100 ns, the ligand obtained conformational stability, with a continuous rise and fall in the RMSD spikes at 2.5–3 Å, as illustrated in Fig. 13b.
Evaluation of the 4A7C-6h complex showed an encouraging stability profile throughout the MD simulation, with RMSDs ranging from 2 to 2.5 Å, with minimal alterations in the RMSD projections. It was noticed that the nascent ligand displayed average stability features as compared to the ligand protein complex (Fig. 13c).
3.10. In silico ADME and drug-likeliness prediction
All the synthesized compounds were subjected to ADMET profiling and drug-likeness assessment employing the free online software https://www.swissadme.ch/index.php and https://biosig.unimelb.edu.au/pkcsm. All the screened molecules were tested for the in silico ADMET features by using SwissADME. The molecular weight calculated for all the molecules was in the range of 303.31–392.25 Da, with a maximum count of 5 HBA and 1 HBD. The TPSA of the compounds under scrutiny, 6a–f, 6i–m, 6q, were calculated with TPSA of 46.4 Å2 while 6g, 6n, showed 55.63 Å2 with 6r, was 59.54 Å2 and for 6h, and 6o, it was 92.22 Å2 demonstrated values of 46.4, 55.63, 59.54 and 92.22 Å2, respectively, with log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values ranging from 3.84 to 5.40. All molecules fall in the window of Lipinski's rule of 5 (Lipinski RO5) without any violation.
P values ranging from 3.84 to 5.40. All molecules fall in the window of Lipinski's rule of 5 (Lipinski RO5) without any violation.
Moreover, the compounds were predicted to have excellent intestinal absorption, from which the potent molecules 6a, 6b, 6c, 6h and 6p exhibited % absorptions of 92.92, 92.58, 91.52, 91.15 and 92.38%, respectively. The potent compounds are more readily transported through the BBB, among which 6h demonstrated poor distribution through the BBB. The most active compounds 6a, 6c, 6h, and 6p were safe and did not induce hepatotoxicity, whereas 6b induced hepatotoxicity. Overall, the synthesized molecules were drug-like according to the LRO5, and displayed excellent ADMET profiles. The detailed analysis is presented in Table 6.
| Molecule Id | MW | TPSA (Å2) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
HA | HD | Rotatable bonds | Lipinski rule | % Abs | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) BB BB |
CLtot(log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ml min−1 kg−1) ml min−1 kg−1) |
Hepatotoxicity |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 6a | 313.35 | 46.4 | 4.25 | 2 | 1 | 4 | Yes | 92.92 | 0.48 | 0.73 | No |
| 6b | 331.34 | 46.4 | 4.39 | 3 | 1 | 4 | 92.58 | 0.46 | 0.69 | Yes | |
| 6c | 347.8 | 46.4 | 4.90 | 2 | 1 | 4 | 91.52 | 0.46 | 0.72 | No | |
| 6d | 392.25 | 46.4 | 5.01 | 2 | 1 | 4 | 91.46 | 0.45 | 0.71 | ||
| 6e | 381.35 | 46.4 | 5.27 | 5 | 1 | 4 | 90.60 | 0.46 | 0.59 | ||
| 6f | 327.38 | 46.4 | 4.56 | 2 | 1 | 4 | 92.98 | 0.47 | 0.74 | ||
| 6g | 343.38 | 55.6 | 4.26 | 3 | 1 | 5 | 94.03 | 0.33 | 0.76 | Yes | |
| 6h | 358.35 | 92.2 | 4.16 | 4 | 1 | 5 | 91.15 | −0.69 | 0.50 | No | |
| 6i | 331.34 | 46.4 | 4.39 | 3 | 1 | 4 | 93.06 | 0.45 | 0.65 | Yes | |
| 6j | 347.8 | 46.4 | 4.90 | 2 | 1 | 4 | 91.95 | 0.45 | 0.66 | No | |
| 6k | 392.25 | 46.4 | 5.01 | 2 | 1 | 4 | 91.88 | 0.45 | 0.61 | ||
| 6l | 381.35 | 46.4 | 5.27 | 5 | 1 | 5 | 91.02 | 0.45 | 0.54 | ||
| 6m | 327.38 | 46.4 | 4.56 | 2 | 1 | 4 | 93.40 | 0.47 | 0.71 | ||
| 6n | 343.38 | 55.6 | 4.26 | 3 | 1 | 5 | 94.51 | 0.32 | 0.72 | Yes | |
| 6o | 358.35 | 92.2 | 4.16 | 4 | 1 | 5 | 91.93 | −0.68 | 0.46 | No | |
| 6p | 331.34 | 46.4 | 4.39 | 3 | 1 | 4 | 92.38 | 0.49 | 0.64 | ||
| 6q | 363.41 | 46.4 | 5.40 | 2 | 1 | 4 | 93.32 | 0.5 | 0.72 | ||
| 6r | 303.31 | 59.5 | 3.84 | 3 | 1 | 4 | 92.96 | 0.44 | 0.79 |
4. Conclusion
In conclusion, this research article revealed the comprehensive utility of various computation and machine learning tools in designing new compounds with different scaffolds for PIM1 inhibitory activity. The designed molecules were synthesized, characterized and subjected to numerous in vitro evaluations, including PIM1 inhibitory assay at 10 µM concentration. Among the synthesized compounds, 6h displayed superior antiproliferative activity against colon cancer cell lines, with an IC50 of 18.1 ± 0.44 µM. 6h could prompt apoptosis and prevent the growth of HT-29 cells due to its antiproliferative potential, thus augmenting its therapeutic potential against cancer cells. In order to confirm our hypothesis of PIM1 kinase inhibition, all compounds were tested at a fixed concentration for kinase inhibitory activity, which revealed moderate kinase inhibitory potentials of 34.5% ± 3.4%. Further, structure-based computational studies, including docking and molecular dynamics simulation, strengthened the approach, with favorable ADMET property predictions. Further, in vivo evaluation coupled with determination of the kinase inhibitory IC50 values confirmed the computational and machine learning approach for medicinal chemistry.Ethical statement
This research has been protected by filing a Patent (Indian Patent Application No. IN202421052696, 2024).Author contributions
All the authors contributed equally and approved the final version of the manuscript.Conflicts of interest
The authors declare no financial interest that could appear to affect the research work.Data availability
The data used for development of the ML models were obtained from open source published research articles. The codes for the ML were composed in-house on Jupiter note book and have not been taken from any other sources. The molecular docking results are included in the manuscript. The molecular dynamics simulation output files are very large in size and hence cannot be shared. The data derived from MDS such as RMSD, RG, RMSF and HB are included in the document.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6ra00514d.
Acknowledgements
VW extends gratitude to the Principal, Poona College of Pharmacy, Pune, for the constant encouragement. RK thanks the Management team of BLDEA University for providing the necessary help in cell line studies.References
- A. Upadhyay, Cancer: An unknown territory; rethinking before going ahead, Gene. Dis., 2021, 8, 655–661, DOI:10.1016/j.gendis.2020.09.002.
- S. Pandiyan and L. Wang, A comprehensive review on recent approaches for cancer drug discovery associated with artificial intelligence, Comput. Biol. Med., 2022, 150, 106140, DOI:10.1016/j.compbiomed.2022.106140.
- K. S. Bhullar, N. O. Lagarón, E. M. McGowan, I. Parmar, A. Jha, B. P. Hubbard and H. P. V. Rupasinghe, Kinase-targeted cancer therapies: progress, challenges and future directions, Mol. Cancer, 2018, 17, 1–20, DOI:10.1186/s12943-018-0804-2.
- A. Turdo, C. D'Accardo, A. Glaviano, G. Porcelli, C. Colarossi, L. Colarossi, M. Mare, N. Faldetta, C. Modica, G. Pistone, M. R. Bongiorno, M. Todaro and G. Stassi, Targeting phosphatases and kinases: how to checkmate cancer, Front. Cell Dev. Biol., 2021, 9, 690306, DOI:10.3389/fcell.2021.690306.
- R. R. Naik and A. K. Shakya, Exploring the chemotherapeutic potential of currently used kinase inhibitors: An update, Front. Pharmacol, 2023, 13, 1064472, DOI:10.3389/fphar.2022.1064472.
- M. Bellon and C. Nicot, Targeting Pim kinases in hematological cancers: molecular and clinical review, Mol. Cancer, 2023, 22, 18, DOI:10.1186/s12943-023-01721-1.
- S. Nandi, M. Salman, A. Sharma and M. Kumar, Signal transduction through JAK-STAT and NF-κB regulated Pim-1 Kinase: Novel target for anticancer leads, Curr. Signal Transduct. Ther., 2018, 13, 83–104, DOI:10.2174/1574888X13666180301124935.
- M. Bachmann, H. Hennemann, P. X. Xing, I. Hoffmann and T. Möröy, The Oncogenic Serine/Threonine Kinase Pim-1 phosphorylates and inhibits the activity of Cdc25C-associated kinase 1 (C-TAK1), J. Biol. Chem., 2004, 279, 48319–48328, DOI:10.1074/jbc.M404440200.
- Y. Peng, J. Li, F. Xie, J. Chen, Y. Yu, X. Ouyang and H. Liang, expression of pim-1 in tumors, tumor stroma and tumor-adjacent mucosa co-determines the prognosis of colon cancer patients, PLoS One, 2013, 8, e76693, DOI:10.1371/journal.pone.0076693.
- M. Zhang, T. Liu, H. Sun, W. Weng, Q. Zhang, C. Liu, Y. Han and W. Sheng, Pim1 supports human colorectal cancer growth during glucose deprivation by enhancing the Warburg effect, Cancer Sci., 2018, 109, 1468–1479, DOI:10.1111/cas.13562.
- S. S. Chauhan, R. K. Toth, C. C. Jensen, A. L. Casillas, D. F. Kashatus and N. A. Warfel, PIM kinases alter mitochondrial dynamics and chemosensitivity in lung cancer, Oncogene, 2020, 39, 2597–2611, DOI:10.1038/s41388-020-1168-9.
- J. H. Song and A. S. Kraft, Pim kinase inhibitors sensitize prostate cancer cells to apoptosis triggered by Bcl-2 family inhibitor ABT-737, Cancer Res., 2012, 72, 294–303, DOI:10.1158/0008-5472.CAN-11-3240.
- C. Blanco-Aparicio and A. Carnero, Pim kinases in cancer: Diagnostic, prognostic and treatment opportunities, Biochem. Pharmacol., 2013, 85, 629–643, DOI:10.1016/j.bcp.2012.09.018.
- X. Tang, T. Cao, Y. Zhu, L. Zhang, J. Chen, T. Liu, X. Ming, S. Fang, Y. Yuan, L. Jiang, J.-D. Huang and X.-Y. Guan, PIM2 promotes hepatocellular carcinoma tumorigenesis and progression through activating NF-κB signaling pathway, Cell Death Dis., 2020, 11, 510, DOI:10.1038/s41419-020-2700-0.
- V. Walhekar, C. Bagul, D. Kumar, A. Muthal, G. Achaiah and R. Kulkarni, Topical advances in PIM kinases and their inhibitors: Medicinal chemistry perspectives, Biochim. Biophys. Acta Rev. Cancer, 2022, 1877, 188725, DOI:10.1016/j.bbcan.2022.188725.
- J. Bogusz, K. Zrubek, K. P. Rembacz, P. Grudnik, P. Golik, M. Romanowska, B. Wladyka and G. Dubin, Structural analysis of PIM1 kinase complexes with ATP-competitive inhibitors, Sci. Rep., 2017, 7, 13399, DOI:10.1038/s41598-017-13557-z.
- L. S. Chen, S. Redkar, D. Bearss, W. G. Wierda and V. Gandhi, Pim kinase inhibitor, SGI-1776, induces apoptosis in chronic lymphocytic leukemia cells, Blood, 2009, 114, 4150–4157, DOI:10.1182/blood-2009-03-212852.
- E. K. Keeton, K. McEachern, K. S. Dillman, S. Palakurthi, Y. Cao, M. R. Grondine, S. Kaur, S. Wang, Y. Chen, A. Wu, M. Shen, F. D. Gibbons, M. L. Lamb, X. Zheng, R. M. Stone, D. J. DeAngelo, L. C. Platanias, L. A. Dakin, H. Chen, P. D. Lyne and D. Huszar, AZD1208, a potent and selective pan-Pim kinase inhibitor, demonstrates efficacy in preclinical models of acute myeloid leukemia, Blood, 2014, 123, 905–913, DOI:10.1182/blood-2013-04-495366.
- T. Paíno, A. Garcia-Gomez, L. González-Méndez, L. San-Segundo, S. Hernández-García, A.-A. López-Iglesias, E. M. Algarín, M. Martín-Sánchez, D. Corbacho, C. Ortiz-de-Solorzano, L. A. Corchete, N. C. Gutiérrez, M.-V. Maetos, M. Garayoa and E. M. Ocio, The Novel Pan-PIM kinase inhibitor, PIM447, displays dual antimyeloma and bone-protective effects, and potently synergizes with current standards of care, Clin. Cancer Res., 2017, 23, 225–238, DOI:10.1158/1078-0432.CCR-16-0230.
- H. Koblish, Y. Li, N. Shin, L. Hall, Q. Wang, K. Wang, M. Covington, C. Marando, K. Bowman, J. Boer, K. Burke, R. Wynn, A. Margulis, G. W. Reuther, Q. T. Lambert, V. Dostalik Roman, K. Zhang, H. Feng, C.-B. Xue, S. Diamond, G. Hollis, S. Yeleswaram, W. Yao, R. Huber, K. Vaddi and P. Scherle, Preclinical characterization of INCB053914, a novel pan-PIM kinase inhibitor, alone and in combination with anticancer agents, in models of hematologic malignancies, PLoS ONE, 2018, 13, e0199108, DOI:10.1371/journal.pone.0199108.
- V. S. Walhekar, C. Bagul, D. Kumar and R. Kulkarni, Computational modelling strategies in exploring triazolopyridazine PIM1 kinase inhibitors as anticancer agents, Anti-Cancer Agents Med. Chem., 2025, 25(13), 954–966, DOI:10.2174/1871520622666220820090353.
- A. Choudhary, M. Khandekar, C. Bagul, A. Muthal and R. Kulkarni, Investigation of structure activity relationship: in silico studies of [1,2,4]triazolo[4,3-a]pyridine ureas as P38 kinase inhibitors, Struct. Chem., 2023, 34, 915–929, DOI:10.1007/s11224-022-02046-3.
- G. K. Ravindra, G. Achaiah, S. Laufer and V. M. Chandrashekar, Synthesis, anti-inflammatory and p38 kinase inhibitory activity of N, N′-diaryl urea derivatives, Med. Chem., 2013, 9, 213–221, DOI:10.2174/1573406411309020006.
- G. K. Ravindra, G. Achaiah, S. Laufer and V. M. Chandrashekar, Synthesis, p38 kinase inhibitory and anti-inflammatory activity of new substituted benzimidazole derivatives, Med. Chem., 2013, 9, 90–99, DOI:10.2174/157340613804488440.
- R. Kulkarni, U. Mitkari, S. Divyani, G. Achaiah, S. Laufer, D. V. R. N. Bikshapti, V. M. Chandrashekar, P. Gurav and S. J. Joshi, Design, synthesis, anti-inflammatory, p38 kinase inhibitory, antimicrobial and antitubercular activities of substituted benzamides, Mini Rev. Med. Chem., 2018, 18(17), 1486–1497, DOI:10.2174/1389557517666170707105416.
- S. Kulkarni, S. Deshpande, M. Shetty, P. Sahare, V. Shinde, A. Patil, V. Walhekar, S. G. Alegaon, S. D. Ranade, S. patrudu, A. Ganeshpurkar and R. Kulkarni, 2-Substituted 3-oxindoles as glycogen synthase kinase 3β inhibitors: Insights from ML based QSAR, molecular docking, and dynamics simulations, Proc. Indian Natl. Sci. Acad., 2025 DOI:10.1007/s43538-025-00533-9.
- D. Anuradha, G. K. Ravindra, G. Achaiah and P. Radhakrishna, Pyrazole derivatives as potent inhibitors of c-Jun N-terminal kinase: synthesis and SAR studies, Bioorg. Med. Chem., 2014, 22, 6209–6219, DOI:10.1016/j.bmc.2014.08.028.
- G. N. D. Ion, G. M. Nitulescu and D. P. Mihai, A PIM-1 Kinase Inhibitor Docking Optimization Study Based on Logistic Regression Models and Interaction Analysis, Life, 2023, 13(8), 1635, DOI:10.3390/life13081635.
- H. Park, J. Jeon, K. Kim, S. Choi and S. Hong, Structure-Based Virtual Screening and De Novo Design of PIM1 Inhibitors with Anticancer Activity from Natural Products, Pharmaceuticals, 2021, 14(3), 275, DOI:10.3390/ph14030275.
- H. Almukadi, G. A. Jadkarim, A. Mohammed, M. Almansouri, N. Sultana, N. A. Shaik and B. Banaganapalli, Combining machine learning and structure-based approaches to develop oncogene PIM kinase inhibitors, Front. Chem., 2023, 11, 1137444, DOI:10.3389/fchem.2023.1137444.
- V. Walhekar, A. Muthal, M. Ghaisas, N. R. Mamidipalli, R. Kulkarni and R. Kulkarni, Reconnecting new tetrahydrobenzothieno-4-pyrimidine amides as potent PIM-1 kinase inhibitors via an integrated ligand and structure based scaffold hopping: In vitro anti-tumor investigations, Bioorg. Chem., 2026, 109487, DOI:10.1016/j.bioorg.2026.109487.
- M. S. Sunkara, V. Kuchana, J. P. Sree, R. Prabugari, A. Pilli, F. Irum, S. J. Tangeda and D. Bhowmik, Pharmacophore-based virtual screening & molecular docking studies on selected plant constituents of Plantago major, J. Applied Pharm. Sci., 2022, 157–167, DOI:10.7324/JAPS.2023.92723.
- D. Bajusz, A. Rácz and K. Héberger, Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations?, J. Cheminf., 2015, 7(20) DOI:10.1186/s13321-015-0069-3.
- J. van Meerloo, G. J. L. Kaspers and J. Cloos, in. Cell Sensitivity Assays: the MTT Assay, 2011, pp. 237–245, DOI:10.1007/978-1-61779-080-5_20.
- V. J. Sawant, S. R. Bamane, R. V. Shejwal and S. B. Patil, Comparison of drug delivery potentials of surface functionalized cobalt and zinc ferrite nanohybrids for curcumin in to MCF-7 breast cancer cells, J. Magn. Magn. Mater., 2016, 417, 222–229, DOI:10.1016/j.jmmm.2016.05.061.
- B. J. Alves, D. Ph, S. A. Goueli, D. Ph, H. Zegzouti, D. Ph, P. Corporation, PIM1 Kinase Assay, (n.d.) 1–2.
- I. Lakshmanan and S. Batra, Protocol for Apoptosis Assay by Flow Cytometry Using Annexin V Staining Method, Bio-protoc., 2013, 3, 374, DOI:10.21769/BioProtoc.374.
- S. Martínez-González, S. Rodríguez-Arístegui, C. A. Gómez de la Oliva, A. I. Hernández, E. González Cantalapiedra, C. Varela, A. B. García, O. Rabal, J. Oyarzabal, J. R. Bischoff, J. Klett, M. I. Albarrán, A. Cebriá, N. Ajenjo, B. García-Serelde, E. Gómez-Casero, M. Cuadrado-Urbano, D. Cebrián, C. Blanco-Aparicio and J. Pastor, Discovery of novel triazolo[4,3-b]pyridazin-3-yl-quinoline derivatives as PIM inhibitors, Eur. J. Med. Chem., 2019, 168, 87–109, DOI:10.1016/j.ejmech.2019.02.022.
- O. Trott and A. J. Olson, AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comb. Chem., 2010, 31, 455–461, DOI:10.1002/jcc.21334.
- R. Choudhary, V. Walhekar, A. Muthal, D. Kumar, C. Bagul and R. Kulkarni, Machine learning facilitated structural activity relationship approach for the discovery of novel inhibitors targeting EGFR, J. Biomol. Struct. Dyn., 2023, 12445–12463, DOI:10.1080/07391102.2023.2175263.
- K. Mangala, W. Vinayak, C. Aasiya, B. Chandrakant, M. Amol, D. Kumar and R. Kulkarni, Reconnoitering imidazopyridazines as anticancer agents based on virtual modelling approach: quantitative structure activity relationship, molecular docking and molecular dynamics, J. Biomol. Struct. Dyn., 2024, 42, 2392–2409, DOI:10.1080/07391102.2023.2204502.
- S. Jagannath, W. Vinayak, M. Amol, P. Sandhya, V. M. Chandrashekhar, S. Vaibhav and R. Kulkarni, Molecular dynamics directed neuroprotective activity of alcoholic extract of Garuga pinnata Roxb. in Experimental Rats, J. Ayurveda Integr. Med., 2025, 16, 101032, DOI:10.1016/j.jaim.2024.101032.
- D. E. V. Pires, T. L. Blundell and D. B. Ascher, pkCSM: Predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures, J. Med. Chem., 2015, 58, 4066–4072, DOI:10.1021/acs.jmedchem.5b00104.
- A. Daina, O. Michielin and V. Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Sci. Rep., 2017, 7, 42717, DOI:10.1038/srep42717.
| This journal is © The Royal Society of Chemistry 2026 |