Open Access Article
Open Access ArticleTransformation of 3-(4-methoxybenzylidine)-5-(p-tolyl)-2(3H)-furanone into new nitrogen heterocyclic candidates as insecticidal agents with in silico studies
Nourhan M. Gada,
David S. A. Haneen *a,
Mahmoud Kamal
*a,
Mahmoud Kamal b,
Eslam M. Hosni
b,
Eslam M. Hosni b,
Mohamed H. Hekal
b,
Mohamed H. Hekal a,
Abdelaal A. Abdalhaa,
Wael S. I. Abou-Elmagd
a,
Abdelaal A. Abdalhaa,
Wael S. I. Abou-Elmagd a and
Sayed K. Ramadan
a and
Sayed K. Ramadan *a
*a
aChemistry Department, Faculty of Science, Ain Shams University, Cairo, 11566, Egypt. E-mail: david.shoukry@sci.asu.edu.eg; sayed.karam2008@sci.asu.edu.eg
bEntomology Department, Faculty of Science, Ain Shams University, Cairo, 11566, Egypt
First published on 19th March 2026
Abstract
A series of new nitrogen-containing heterocyclic compounds derived from a 2(3H)-furanone precursor was synthesized via reactions with mono- and bidentate nucleophiles, affording pyrrolone, benzoxazinone, quinazolinone, and pyridazine frameworks. The synthesized candidates were characterized spectroscopically and evaluated for larvicidal activity against third-instar Culex pipiens larvae, revealing a wide potency range (LC50 = 0.1–458.6 µg mL−1). Among them, compound 4 exhibited the highest toxicity (LC50 = 0.1 µg mL−1), while several others showed moderate activity (LC50 = 34–95 µg mL−1). Molecular docking against Culex acetylcholinesterase (AChE) and the methoprene-tolerant (Met) receptor indicated that the tested compounds engage catalytically relevant residues such as HIS567 and TRP212 in AChE and HIS23 in Met, though binding energies did not consistently parallel larvicidal potency. Molecular-dynamics simulations (100 ns) supported the stability of the ligand–protein complexes and provided structural evidence for the potential accommodation of the compounds within both targets. In silico ADME predictions revealed high gastrointestinal absorption, balanced lipophilicity, and generally good BBB permeability, with only minor differences, such as the non-BBB-permeant compounds 7, 8, and 14, that may influence access to neural targets. Overall, the combined experimental and computational findings suggest that larvicidal activity arises from a multifactorial interplay between molecular recognition, transport properties, and physicochemical balance rather than docking affinity alone. These results indicate that 2(3H)-furanone-derived heterocycles may represent promising scaffolds for further investigation as dual-target larvicidal candidates. The present findings may contribute to the rational design of next-generation mosquito control agents and support ongoing efforts toward developing more effective and sustainable strategies for vector-borne disease management.
1. Introduction
2(3H)-Furanones are highly adaptable building blocks for synthesizing a wide range of valuable heterocyclic compounds with significant synthetic and biological relevance.1–4 Our research group has focused on exploring the reactivity of these furanones with various nitrogen-based nucleophiles, aiming to develop diverse nitrogen-holding heterocyclic systems with notable pharmacological properties, such as insecticidal, antiviral, antitumor, and antimicrobial activities.5–10 These furanones are particularly reactive due to their tendency to undergo facile ring opening upon nucleophilic attack.11,12 Nitrogen-containing heterocycles are prevalent in nature, and they are significant structural motifs in the advancement of new drugs and play essential roles in both agrochemicals and pharmaceuticals.13,14Among them, chlorfenapyr, a pyrrole derivative, was used as a pesticide (Fig. 1). Also, pyrrole, benzoxazinone, quinazolinone, and pyridazine candidates were found to exhibit promising biological effects, including insecticidal, antimicrobial, antiviral, and antitumor activities depending on their structural features and substitution patterns.4,15–23 Their nitrogen, oxygen, and/or sulfur atoms are important for their biological efficacy through formation of distinct non-covalent interactions such as hydrophobic contacts, hydrogen bonding, dipole–dipole interactions and van der Waals forces with protein targets. In many ligand–protein systems, hydrogen bonding and hydrophobic contacts are among the most frequently observed interactions and play a central role in stabilizing ligand binding within the active site.24
 | ||
| Fig. 1 Structures of chlorfenapyr, and some reported pyrrolones I–III and pyridazinones IV, V with insecticidal activity. | ||
In 2025, two synthesized pyrrolones I and II (Fig. 1) revealed strong larvicidal activity with LC50 values of 0.4 and 4.9 µg mL−1, respectively, through which pyrrolone I outperformed the reference insecticide, chlorpyrifos.6 Also, three pyrrolone derivatives IIIa–c were highly toxic to Culex pipiens with LC50 of 21.05, 22.81, and 38.07 µg mL−1 compared to the reference, chlorfenapyr with LC50 of 25.43 µg mL−118 (cf. Fig. 1). In 2022, two pyridazinones IV and V (Fig. 1) were described as undertaking agents for the control of insect pests, where they were evaluated towards the third larval instar of lab and field strains of C. pipiens, which showed high toxicity, making them suitable for controlling C. pipiens at larval stage.5 Therefore, widespread research is still needed to advance and enhance their properties as insecticidal agents and to reduce their adverse effects.
Mosquito-borne diseases continue to represent a serious threat to global public health, with Culex pipiens serving as a major vector of pathogens such as West Nile virus, Rift Valley fever, and various filarial nematodes.25–29 The widespread and persisted use of chemical insecticides has led to rapid development of resistance within mosquito populations, undermining the effectiveness of conventional control programs. This resistance is often associated with genetic mutations in target enzymes and enhanced metabolic detoxification mechanisms that reduce insecticide sensitivity.30–33 Most traditional insecticides, including organophosphates and carbamates, act through the inhibition of acetylcholinesterase (AChE), resulting in the accumulation of acetylcholine at synapses and subsequent neuronal hyperexcitation.34–38 Although highly effective, the repeated use of these neurotoxic compounds has promoted the selection of resistant alleles and enzymatic variants of AChE, particularly those harboring mutations at the catalytic serine or oxyanion hole residues.39
To address this challenge, alternative molecular targets have gained attention, including the methoprene-tolerant (Met) receptor, a member of the basic helix–loop–helix Per-Arnt-Sim (bHLH-PAS) family that functions as the receptor for the juvenile hormone (JH). The Met receptor regulates critical developmental processes including metamorphosis, reproduction, and cuticle formation in insects.40–42 Juvenile hormone analogs (JHAs) such as pyriproxyfen and methoprene exploit this pathway by interfering with hormonal signaling, thereby preventing the pupal–adult transition and causing delayed mortality in larvae. Several compounds targeting these pathways have been approved and widely used in mosquito control programs, highlighting the practical importance of these molecular targets in vector management. Although JH-based insect growth regulators are safer for non-target organisms, their slow action makes them less efficient in short-term larval management. Consequently, combining fast-acting neurotoxicity with hormone disruption through dual-target insecticidal agents could offer an effective strategy to achieve potent larvicidal effects while minimizing the risk of resistance.40,41,43,44
Accordingly, our present study was designed not only to synthesize and evaluate new nitrogen-containing heterocyclic compounds derived from 2(3H)-furanone scaffolds, but also to explore their potential to act through different molecular mechanisms and targets. We hypothesized that such structural diversification could yield molecules capable of interacting with both the classical neurotoxic target (AChE) and the hormonal regulatory site (Met receptor), thereby providing a multitarget mode of action that may help mitigate resistance arising from single-target insecticides.
The synthesized compounds were screened for acute toxicity in larval bioassays alongside the reference insecticide chlorpyrifos. To rationalize their biological activity and explore potential mechanisms of action, an integrated computational approach was employed, incorporating molecular docking, molecular dynamics simulations, and ADME predictions. The selected molecular targets, acetylcholinesterase (AChE) and methoprene-tolerant (Met) receptor, are key to insecticidal action. The heterocyclic rings in the synthesized compounds, including both five-membered and six-membered systems, contain polar and aromatic features commonly associated with bioactive insecticidal scaffolds. Rather than implying direct scaffold similarity to reported co-crystallized ligands, the present docking strategy focused on evaluating whether the designed molecules can geometrically accommodate the same binding cavities and engage conserved catalytic or hormone-binding residues within the active sites. Such spatial compatibility and interaction patterns, even in the absence of structural resemblance, may indicate the feasibility of target recognition at both neural (AChE) and endocrine (Met) levels.
This combined experimental and computational approach provides a mechanistic framework for understanding the observed larvicidal activities and highlights the potential of 2(3H)-furanone-derived heterocycles as multitarget insecticidal candidates capable of simultaneously modulating neural and endocrine pathways. Such an approach could contribute to the advance of next-generation insecticides with enhanced selectivity, reduced resistance risk, and enhanced sustainability in mosquito vector control. It should be emphasized that the present investigation represents an initial exploratory stage primarily focused on identifying the most promising molecular scaffolds through an integrated in silico and bioassay evaluation. This early-stage screening enables the rational selection of one or two potent candidates for subsequent mechanistic, environmental, and toxicological assessments. Among the synthesized derivatives, compound 4 demonstrated the highest larvicidal potency against Culex pipiens, highlighting the potential of this scaffold for further development. Accordingly, comprehensive toxicity studies on non-target organisms will be conducted in future stages once the most active and selective compounds have been prioritized, ensuring that experimental efforts are directed toward the most viable leads. Compared with previously reported heterocyclic insecticidal candidates, the present study introduces new furanone-derived scaffolds and combines experimental larvicidal evaluation with integrated molecular docking and dynamics analyses to explore their potential mechanisms of action, which may support future research toward the development of improved mosquito control agents.
2. Results and discussion
2.1 Synthesis and characterization of target compounds
2(3H)-Furanones are considered as useful synthons for an inclusive diversity of other heterocycles of synthetic and biological importance. Thus, the key synthon, E/Z-2(3H)-furanone 1 (ref. 45) was used in this work to construct derivatives of 1,3-dihydro-2H-pyrrol-2-one, benzoxazinone, and pyridazine. Indeed, it reacted with ammonium acetate in refluxing ethanol to produce the respective 1,3-dihydro-2H-pyrrol-2-one derivative 2 (ref. 46) (Scheme 1). In its IR spectrum, lactone C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption disappeared while the amide NH and C
O absorption disappeared while the amide NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption bands appeared. Also, its 1H NMR spectrum indicated a singlet signal in a downfield region related to amide NH proton, in addition to four singlet signals at δ 7.11, 6.80, 3.83, 2.35 ppm corresponding to olefinic (CH
O absorption bands appeared. Also, its 1H NMR spectrum indicated a singlet signal in a downfield region related to amide NH proton, in addition to four singlet signals at δ 7.11, 6.80, 3.83, 2.35 ppm corresponding to olefinic (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), C4–H pyrrole, OCH3, and CH3 (tolyl) protons, respectively. Also, its 13C NMR spectrum strongly reinforced the assigned structure.
), C4–H pyrrole, OCH3, and CH3 (tolyl) protons, respectively. Also, its 13C NMR spectrum strongly reinforced the assigned structure.
 | ||
| Scheme 1 Synthesis of 1,3-dihydro-2H-pyrrol-2-one derivatives 2, 4, 5, and N-benzylamide 3 from 2(3H)-furanone 1. | ||
Reacting furanone 1 with benzylamine in refluxing ethanol for 6 h furnished N-benzylamide 3 as a sole product, through initial nucleophilic attack of primary amino group of benzylamine on the lactone-carbonyl carbon to affect ring opening. Its IR spectrum showed absorption bands for NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups of ketonic and amide. Also, its 1H NMR spectrum presented a broad singlet signal for NH proton and singlet signals for –COCH2– protons. Noteworthy, two doublet signals appeared for the magnetically non-equivalent protons of benzyl-methylene group (one of them was next to carbonyl oxygen and the other was next to phenyl moiety), with appropriate J coupling constant, that supported the open-structure.
O groups of ketonic and amide. Also, its 1H NMR spectrum presented a broad singlet signal for NH proton and singlet signals for –COCH2– protons. Noteworthy, two doublet signals appeared for the magnetically non-equivalent protons of benzyl-methylene group (one of them was next to carbonyl oxygen and the other was next to phenyl moiety), with appropriate J coupling constant, that supported the open-structure.
In turn, treating the furanone 1 with 4-methoxyaniline in refluxing ethanol for a longer time (12 h) produced 1,3-dihydro-2H-pyrrol-2-one derivative 4, through initial lactone ring opening by primary amino group of 4-methoxyaniline followed by 5-exo-trig intramolecular ring closure. The 1H NMR spectrum of 1,3-dihydro-2H-pyrrol-2-one derivative 4 displayed its being as a mixture of E- and Z-stereoisomers, possibly occurred in situ, in a proportion of 78![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22%, respectively. Presumably, the preferred E-configuration was inferred from the steric hindrance between anisyl ring and carbonyl group in the counter Z-form. This was also suggested from the study of space models and MM2 calculations11,47 (cf. Table S1) which revealed the more thermodynamic stability of E-isomer (total E = 60.75 kcal mol−1) than the counter Z-form (total E = 66.03 kcal mol−1) with different 1,4-van-der Waals (VDW) interactions.
22%, respectively. Presumably, the preferred E-configuration was inferred from the steric hindrance between anisyl ring and carbonyl group in the counter Z-form. This was also suggested from the study of space models and MM2 calculations11,47 (cf. Table S1) which revealed the more thermodynamic stability of E-isomer (total E = 60.75 kcal mol−1) than the counter Z-form (total E = 66.03 kcal mol−1) with different 1,4-van-der Waals (VDW) interactions.
On the other hand, reaction of the furanone 1 with methyl 4-aminobenzoate in refluxing ethanol remains unchanged for long time. So, the reaction condition was changed to fusion of the furanone 1 with that reagent in an oil bath at 150–160 °C for 4 h, which afforded the 1,3-dihydro-2H-pyrrol-2-one derivative 5 (cf. Scheme 1). The IR spectra of 1,3-dihydro-2H-pyrrol-2-ones 4 and 5 lacked lactone absorptions. In turn, 1H NMR spectrum of 1,3-dihydro-2H-pyrrol-2-one derivative 5 afforded singlet signal at δ 3.84 ppm, integrated for six protons corresponding to two methoxy sets.
On the other hand, benzoxazinone 7 was obtained upon treating furanone 1 with 2-aminobenzoic acid in neat conditions through the dehydration of the non-isolable intermediate 6 (Scheme 2). Spectroscopically, its IR spectrum offered carbonyl absorption bands related to lactone and ketone carbonyl groups. It 1H NMR spectrum revealed its presence as a mixture of Z- and E-isomers in a proportion of 78![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22%, respectively, which may be attributed to the absence of repulsion between two benzene rings in Z-configuration. The preferred Z-configuration was also proven from the study of space models and MM2 calculations11,47 (cf. Table S1), which revealed the more thermodynamic stability of Z-isomer (total E = 47.31 kcal mol−1) than the counter E-form (total E = 1092.68 kcal mol−1).
22%, respectively, which may be attributed to the absence of repulsion between two benzene rings in Z-configuration. The preferred Z-configuration was also proven from the study of space models and MM2 calculations11,47 (cf. Table S1), which revealed the more thermodynamic stability of Z-isomer (total E = 47.31 kcal mol−1) than the counter E-form (total E = 1092.68 kcal mol−1).
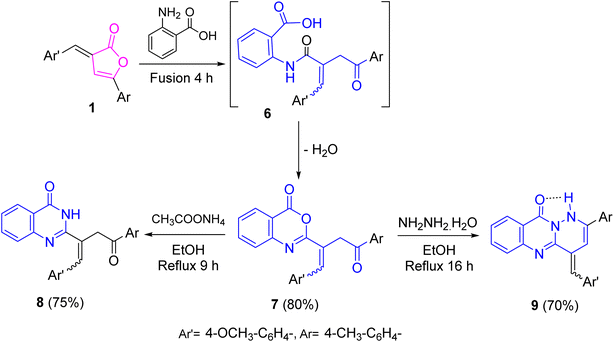 | ||
| Scheme 2 Synthesis of benzoxazinone 7 and its reactions with ammonium acetate and hydrazine hydrate. | ||
Refluxing of benzoxazinone 7 with ammonium acetate in ethanol produced quinazolinone 8 (Scheme 2). Its IR spectrum revealed absorption bands for amide NH and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, in addition to ketonic carbonyl group. Its 1H NMR spectrum displayed singlet signal in the downfield region related to NH proton, along with a singlet signal integrated to two protons corresponding to methylene protons and it was found as Z- and E-isomers in a proportion of 66
O, in addition to ketonic carbonyl group. Its 1H NMR spectrum displayed singlet signal in the downfield region related to NH proton, along with a singlet signal integrated to two protons corresponding to methylene protons and it was found as Z- and E-isomers in a proportion of 66![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
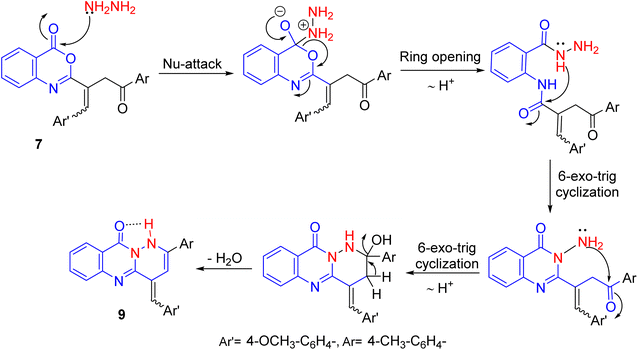
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 33%, respectively, with conserved the preferred Z-configuration (cf. Table S1). In contrast, hydrazinolysis of benzoxazinone 7 using hydrazine hydrate in refluxing ethanol produced pyridazinoquinazoline candidate 9 (cf. Scheme 2). Its IR spectrum exposed NH, C
33%, respectively, with conserved the preferred Z-configuration (cf. Table S1). In contrast, hydrazinolysis of benzoxazinone 7 using hydrazine hydrate in refluxing ethanol produced pyridazinoquinazoline candidate 9 (cf. Scheme 2). Its IR spectrum exposed NH, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, and C
O, and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N absorption bands. The broad band of NH absorption was attributed to intramolecular hydrogen bond with the neighboring carbonyl group. Its 1H NMR spectrum revealed that E- and Z-isomers in a ratio of 60
N absorption bands. The broad band of NH absorption was attributed to intramolecular hydrogen bond with the neighboring carbonyl group. Its 1H NMR spectrum revealed that E- and Z-isomers in a ratio of 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40%, respectively, with preferred E-configuration (cf. Table S1). A probable pathway for this reaction was displayed in Scheme 3.
40%, respectively, with preferred E-configuration (cf. Table S1). A probable pathway for this reaction was displayed in Scheme 3.
Aiming to obtain pyridazine derivatives, furanone 1 underwent hydrazinolysis reaction using hydrazine hydrate in ethanol at room temperature or under reflux to produce pyridazinone derivative 10 (ref. 48) (Scheme 4). Its IR spectrum lacked lactone carbonyl absorption and presented an absorption band for amide carbonyl group. Also, its 1H NMR spectrum offered a singlet signal for methylene protons (δ = 3.78 ppm), in addition to a singlet signal for NH proton in a downfield region (δ = 13.03 ppm). Presumably, this reaction was viewed via initial lactone ring opening by hydrazine (by attack of amino group on lactone carbonyl group) followed by 6-exo-trig intramolecular cyclization. Thiation of pyridazinone 10 using phosphorus pentasulfide in refluxing toluene offered thione derivative 11 (Scheme 4).
The carbonyl absorption disappeared in its IR spectrum, and thioxo absorption band was detected. Also, its 1H NMR spectrum revealed singlet signals for NHCS proton in a downfield region (δ = 14.85 ppm) and methylene protons at δ 4.09 ppm. Further, its 13C NMR showed signals for thioxo-carbon at δ 179.1 ppm and methylene carbon at δ 37.72 ppm. Chlorination of pyridazinone 10 using phosphorus oxychloride furnished 3-chloropyridazine derivative 12 (Scheme 4). Its IR spectrum exposed the absence of carbonyl absorption and showed band for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N group at ν 1612 cm−1. Furthermore, its 1H NMR spectrum presented singlet signal for methylene protons at δ 4.06 ppm, affected by the anisotropic effect of pyridazine aromatic ring and the electron-withdrawing effect of chlorine atom.
N group at ν 1612 cm−1. Furthermore, its 1H NMR spectrum presented singlet signal for methylene protons at δ 4.06 ppm, affected by the anisotropic effect of pyridazine aromatic ring and the electron-withdrawing effect of chlorine atom.
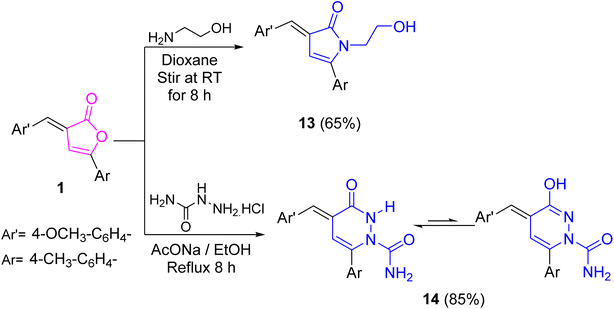
On the other hand, treating furanone 1 with 2-aminoethanol upon stirring in 1,4-dioxane at ambient temperature offered the N-hydroxyethyl-1,3-dihydro-2H-pyrrol-2-one derivative 13 (Scheme 5). Carbonyl absorption band of pyrrolone ring appeared in its IR spectrum. Its 1H NMR spectrum was reliable with the proposed structure. Finally, treatment of furanone 1 with semicarbazide hydrochloride in refluxing ethanol and sodium acetate anhydrous led to the construction of pyridazinone 14 (Scheme 5). Spectroscopically, its IR spectrum was devoid of the lactone carbonyl absorption and showed absorption bands for carbonyl, NH, and NH2 groups. In consistency, its 1H NMR spectrum offered singlet signals for amide (NHCO, 70%) and isoamide (N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–OH, 30%) protons at δ 8.55 and 5.83 ppm, respectively, as well as a singlet signal for NH2 protons at δ 6.13 ppm.
C–OH, 30%) protons at δ 8.55 and 5.83 ppm, respectively, as well as a singlet signal for NH2 protons at δ 6.13 ppm.
2.2 Insecticidal activity evaluation
The larvicidal bioassay results of the produced heterocyclic compounds towards third-instar Culex pipiens larvae are summarized in Table 1 and Fig. 2. All bioassays were conducted in 3 replicates under the WHO-recommended protocol to ensure the reliability of the obtained mortality data. The tested compounds exhibited a wide range of larvicidal activity after 24 h of exposure, with LC50 values varying from 0.1 to 458.6 µg mL−1. Among the series, compound 4 showed the strongest activity (LC50 = 0.1 µg mL−1) and was therefore designated as the internal reference for calculating the toxicity index (100%). Compounds 9 (34.1 µg mL−1), 8 (66.3 µg mL−1), 5 (84.7 µg mL−1), 10 (83.4 µg mL−1), 14 (85.3 µg mL−1), 11 (93.8 µg mL−1), and 7 (94.8 µg mL−1) also demonstrated considerable toxicity compared with the remaining derivatives, which showed LC50 values exceeding 120 µg mL−1. The least active compound in the series was furanone 1 (LC50 = 458.6 µg mL−1).| Compound | LC50/µg mL−1 | χ2cal./tab.(7.8) | r2 | P value | Toxicity index |
|---|---|---|---|---|---|
| 1 | 458.6 | 7.70 | 0.93 | 0.000 | 0.021 |
| 2 | 379.2 | 7.60 | 0.91 | 0.000 | 0.026 |
| 3 | 362.6 | 7.40 | 0.92 | 0.000 | 0.027 |
| 4 | 0.1 | 0.01 | 0.2 | 0.900 | 100 |
| 5 | 84.7 | 3.60 | 0.82 | 0.200 | 0.118 |
| 7 | 94.8 | 4.40 | 0.82 | 0.200 | 0.100 |
| 8 | 66.3 | 5.30 | 0.83 | 0.100 | 0.150 |
| 9 | 34.1 | 1.70 | 0.88 | 0.600 | 0.290 |
| 10 | 83.4 | 3.60 | 0.83 | 0.300 | 0.119 |
| 11 | 93.8 | 7.00 | 0.82 | 0.020 | 0.100 |
| 12 | 127.7 | 6.90 | 0.81 | 0.004 | 0.078 |
| 13 | 343.9 | 6.60 | 0.94 | 0.005 | 0.029 |
| 14 | 85.3 | 6.00 | 0.87 | 0.100 | 0.117 |
| Chlorpyrifos | 420 | 5.20 | 0.96 | 0.300 | 0.023 |
For comparison, the reference insecticide chlorpyrifos exhibited an LC50 value of 420 µg mL−1 under the same bioassay conditions. Chlorpyrifos was included as a standardized toxicological benchmark widely used in WHO-guided larval bioassays to allow contextual evaluation of potency under identical experimental conditions, rather than as a chemotype-related structural comparator. Under these conditions, all compounds except compound 1 displayed higher toxicity, with compound 4 being markedly more active. The calculated toxicity indices, expressed relative to compound 4 (100%), ranged between 0.021 and 0.299 for the remaining derivatives, reflecting their lower relative potency within the series.
The regression analysis demonstrated generally high determination coefficients for most compounds (R2 = 0.82–0.96) together with acceptable χ2 values (≤7.8), supporting the statistical reliability of the dose–response data. An exception was compound 4, whose extremely low LC50 value produced a steeper dose–response slope and consequently a lower correlation coefficient, despite its clear biological potency. Overall, these findings indicate that the larvicidal activity of the tested heterocyclic derivatives is highly dependent on structural variation and that compound 4 represents the most promising lead for further investigation.
2.3 Molecular docking studies
Molecular docking was performed to elucidate the possible molecular mechanisms underlying the observed larvicidal activity and to rationalize the structure–activity trends among the synthesized compounds. Two key insecticidal targets were selected: acetylcholinesterase (AChE), representing the classical neurotoxic site, and the methoprene-tolerant (Met) receptor, the intracellular receptor for the juvenile hormone (JH). The choice of these targets was guided by the distinct structural features of the synthesized heterocycles, which contain polar and heteroaromatic moieties capable of interacting with the catalytic residues of AChE and resembling functional motifs of JH analogs such as pyriproxyfen. Thus, docking into both proteins was intended to determine whether the most active compounds, particularly 1,3-dihydro-2H-pyrrol-2-one derivative 4 and its analogs, might act through one or both molecular pathways, providing a mechanistic explanation for their enhanced larvicidal potency and potential dual-target mode of action.Homology models of Culex pipiens acetylcholinesterase (AChE) and the methoprene-tolerant (Met) receptor were successfully generated to provide accurate protein structures for docking simulations (Fig. 3 and 4). The AChE model was built using the Anopheles gambiae AChE crystal structure (PDB ID: 5X61, chain A) as a template, while the Met PAS-B domain was modeled using the human HIF-2α PAS-B domain (PDB ID: 6E3U, chain B) as a structural analogue owing to the conserved PAS-fold architecture among bHLH-PAS receptors. Both models showed high structural quality, as confirmed by their Ramachandran plots, in which more than 92% of residues occupied favored regions with no residues in disallowed areas. The QMEAN Z-scores (−0.47 for AChE and −0.62 for Met) and global QMEAN scores (∼0.7–0.8) indicated geometries consistent with experimentally determined structures, reflecting good stereochemical reliability.
To ensure biologically meaningful docking, the native co-crystal ligands from the template structures, BT7 (from PDB 6ARY) for AChE and HNJ (from PDB 6E3U) for Met, were transferred to the corresponding modeled cavities. These ligands defined the catalytic gorge of AChE and the hormone-binding pocket of Met with high spatial precision, allowing accurate grid generation and docking in the correct active sites. Collectively, the modeling and validation results confirm that the obtained protein structures are geometrically sound, energetically stable, and appropriately configured for subsequent docking and dynamic analyses.
| Compound | AChE | MET | ||
|---|---|---|---|---|
| S | RMSD | S | RMSD | |
| 1 | −6.01 | 1.16 | −6.70 | 0.87 |
| 2 | −5.85 | 1.23 | −6.65 | 1.26 |
| 3 | −8.06 | 1.40 | −8.74 | 1.27 |
| 4 | −7.38 | 1.02 | −5.06 | 1.04 |
| 5 | −8.08 | 1.59 | −6.51 | 1.18 |
| 7 | −7.34 | 1.64 | −8.06 | 1.46 |
| 8 | −7.38 | 1.25 | −7.16 | 1.40 |
| 9 | −6.97 | 1.55 | −7.05 | 1.91 |
| 10 | −5.18 | 1.94 | −6.82 | 0.83 |
| 11 | −7.15 | 1.52 | −6.34 | 1.42 |
| 12 | −7.09 | 1.02 | −6.46 | 1.49 |
| 13 | −6.42 | 0.78 | −5.44 | 0.99 |
| 14 | −4.45 | 0.94 | −5.98 | 1.59 |
| Chlorpyrifos | −6.72 | 1.94 | — | — |
| Redocked CCL (AChE) | −6.31 | 0.65 | — | — |
| Pyriproxyfen | — | — | −7.36 | 1.54 |
| Redocked CCL (MET) | — | — | −5.90 | 1.40 |
All selected docking poses for the synthesized compounds exhibited RMSD values below 2.0 Å, indicating stable and well-converged conformations. The reference insecticide chlorpyrifos yielded a docking score of −6.72 kcal mol−1, forming hydrogen bonds with HIS567, GLY247, and SER327, together with a π–H interaction involving TRP212. These residues belong to the catalytic triad and peripheral aromatic gorge, consistent with its established inhibitory behavior.
The synthesized derivatives displayed docking scores ranging from −5.18 to −8.08 kcal mol−1. Several ligands showed recurrent engagement of HIS567 and TRP212, with frequent π–π or π–H interactions anchoring the heteroaromatic systems within the aromatic gorge. Compound 4, the most potent derivative in the biological assay, achieved a docking score of −7.38 kcal mol−1 and adopted a stable pose (RMSD = 1.02 Å), exhibiting spatial overlap within the catalytic gorge similar to that observed for the reference ligands at HIS567 and TRP212 (Fig. 5). Although none of the synthesized ligands simultaneously engaged all members of the catalytic triad, the observed interaction patterns confirm that they can be accommodated within the AChE active gorge. Docking scores were not interpreted as quantitative predictors of larvicidal potency. Rather, the docking analysis was used to assess structural accommodation and conserved residue engagement within the catalytic pocket.6–8,49–51
Docking simulations were similarly conducted on the homology-modeled PAS-B domain of the Culex pipiens Met receptor. The co-crystallized ligand HNJ (PDB ID: 6E3U) was superposed to define the ligand-binding cavity prior to docking. Redocking reproduced its orientation with an RMSD of 1.40 Å and a docking score of −5.90 kcal mol−1. Although no strong interactions were reported in the native crystal structure, the redocked pose established hydrogen bonds with MET56 and HIS23, supporting correct cavity definition.
All synthesized compounds exhibited RMSD values below 2.0 Å within the PAS-B domain, indicating stable binding poses. The reference juvenile hormone analog pyriproxyfen yielded a docking score of −7.36 kcal mol−1, forming hydrophobic and π–H contacts with LEU68 and HIS23, consistent with its reported orientation within the hormone-binding cavity. The synthesized ligands displayed docking scores ranging from −5.06 to −8.74 kcal mol−1. Hydrogen bonding with HIS23 represented the most recurrent interaction across the series, often accompanied by contacts involving MET56, CYS115, or THR21 within the PAS-B cavity. Compound 4 exhibited a docking score of −5.06 kcal mol−1 and an RMSD of 1.04 Å, forming a hydrogen bond with HIS23 and showing partial spatial overlap with both HNJ and pyriproxyfen (Fig. 6). Although its docking energy was less favorable than that of pyriproxyfen or compound 3, it maintained a stable and well-converged orientation within the hormone-binding pocket.
Collectively, the docking findings indicate that the synthesized ligands can be accommodated within both the AChE catalytic gorge and the Met PAS-B cavity, reproducing selected interactions with conserved residues such as HIS567 and HIS23. The computational analysis is therefore presented as structural, hypothesis-generating evidence supporting the geometric feasibility of dual-site engagement, without implying direct proportionality between docking energy and biological potency.
2.4 Molecular dynamics simulation analysis
Molecular dynamics (MD) simulations were performed to validate and complement the docking findings by assessing the stability and behavior of the most active compound (compound 4) within the binding pockets of both targets (AChE and Met receptors). This technique was selected because it provides a dynamic view of ligand–protein interactions over time, accounting for solvent effects and conformational flexibility, factors that are not fully captured in static docking studies.6,7,52 Compound 4 was chosen for this detailed analysis based on its highest larvicidal activity and favorable docking affinity values against both targets.Root Mean Square Deviation (RMSD) and Root Mean Square Fluctuation (RMSF) profiles were computed over a 100 ns simulation to evaluate the overall system stability and local residue flexibility. For AChE, the RMSD plots (Fig. 7a–f) indicated that compound 4 maintained stable binding within the active pocket with minimal fluctuations, comparable to the reference insecticide chlorpyrifos and the redocked co-crystallized ligand (BT7). The RMSF curves also confirmed that the residues surrounding the catalytic gorge remained stable throughout the simulation, suggesting that no major conformational rearrangements occurred upon ligand binding.
Similarly, for the methoprene-tolerant (Met) receptor (Fig. S1a–f), compound 4 exhibited a consistent RMSD trajectory, aligning closely with the reference juvenile-hormone analog pyriproxyfen and the re-docked native ligand (HNJ). Local residue fluctuations (RMSF) were limited to flexible loop regions, while the key binding residues displayed minimal deviation, confirming stable accommodation of the ligand within the PAS-B domain pocket.
Protein–ligand contact histograms (Fig. 8 and S2) provided insight into the persistence and nature of the interactions. For both targets, compound 4 maintained several hydrogen bonds and hydrophobic contacts with residues previously identified from the docking analysis, confirming the reliability of the docking predictions. Notably, in AChE, the interactions involving catalytic residues (such as HIS567, TRP212, and TRP559) persisted for most of the simulation time, whereas in the Met receptor, compound 4 preserved strong hydrophobic and hydrogen-bond interactions within the hormone-binding cavity. In comparison, the reference insecticides exhibited similar but sometimes less persistent contact frequencies, and the co-crystallized ligands followed expected binding behavior, serving as internal validation controls.
The MD trajectories corroborate the docking results, demonstrating that compound 4 forms stable and persistent interactions within both AChE and Met active pockets. The consistency between docking and MD findings supports the internal coherence of the computational workflow and indicates that the designed molecules can maintain stable interactions within the modeled systems under simulated physiological conditions.
Although docking and MD analyses provide strong predictive evidence, further biochemical assays on enzyme inhibition and hormonal pathway disruption are necessary to confirm the proposed mechanisms of action. Additional studies such as enzyme inhibition assays, gene expression analysis of hormonal pathways, and extended toxicity evaluations may further clarify the biological mechanisms underlying the observed larvicidal activity. Together, these in silico insights suggest that the studied compounds may exert their larvicidal effects primarily through AChE inhibition and/or juvenile-hormone mimicry, consistent with the observed mortality patterns in bioassay results. Hence, molecular dynamics not only strengthened the docking conclusions but also underscored the practical value of combining these computational methods as efficient screening tools that reduce experimental workload by focusing on the most promising candidates for future validation.
2.5 Structure–activity relationship (SAR) analysis
A comparative evaluation of the structural features of the synthesized heterocyclic compounds and their corresponding larvicidal activities revealed distinct structure–activity relationships, where variations in heterocyclic ring systems and substituent patterns influenced both potency and target affinity. Although compound 4 displayed the lowest LC50 value (0.1 µg mL−1), its docking scores against AChE and Met were within the moderate range rather than the most favorable across the series. A closer comparison with structurally related analogues provides additional insight. For instance, compound 3, which bears a bulkier benzylamide moiety, showed stronger predicted binding energy in AChE yet markedly lower larvicidal potency. This suggests that increased steric bulk and higher polarity (TPSA = 55.40 Å2) may partially restrict membrane penetration or intracellular accessibility despite favorable in silico interactions. Similarly, compound 5, incorporating an ester functionality and slightly higher lipophilicity (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 4.49), achieved strong docking scores but did not translate this into comparable biological activity, possibly due to altered conformational behavior or distribution properties in the larval system.
P = 4.49), achieved strong docking scores but did not translate this into comparable biological activity, possibly due to altered conformational behavior or distribution properties in the larval system.
In contrast, compound 4 exhibits a balanced physicochemical profile (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 4.36; TPSA = 38.77 Å2) combined with the absence of predicted efflux liability and preserved interaction geometry within both targets. This combination of moderate rigidity, controlled polarity, and adequate lipophilicity may facilitate efficient penetration through larval barriers and sustained target engagement. Therefore, its superior potency likely reflects an optimized interplay between structural compatibility and physicochemical accessibility rather than maximal docking affinity alone.49,51
P = 4.36; TPSA = 38.77 Å2) combined with the absence of predicted efflux liability and preserved interaction geometry within both targets. This combination of moderate rigidity, controlled polarity, and adequate lipophilicity may facilitate efficient penetration through larval barriers and sustained target engagement. Therefore, its superior potency likely reflects an optimized interplay between structural compatibility and physicochemical accessibility rather than maximal docking affinity alone.49,51
The docking analyses nonetheless indicate that compound 4 maintains relevant stabilizing interactions with residues critical for target recognition, HIS567 and TRP212 in AChE, and HIS23 in Met, supporting the likelihood of simultaneous engagement of both neurotoxic and hormonal pathways. The combination of an electronically balanced aromatic scaffold and favorable physicochemical parameters may therefore facilitate dual-site compatibility, even without exceptionally strong docking energies, helping to rationalize its overall biological performance.
Compounds 5, 7, 8, 9, 10, and 11, which exhibited moderate larvicidal activity (LC50 = 34–95 µg mL−1), shared comparable docking energies but displayed variations in substituent polarity and spatial orientation. These structural differences may have reduced their overall bioavailability or target accessibility despite satisfactory binding scores. For instance, compound 5 showed a strong docking score in AChE (−8.08 kcal mol−1) yet produced lower larvicidal activity, suggesting that binding affinity alone is not the sole determinant of biological efficacy. Such discrepancies can arise from differences in lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), molecular size, or hydrogen-bonding capacity, all of which affect the ability of the compound to penetrate larval cuticle barriers and reach internal target sites.
P), molecular size, or hydrogen-bonding capacity, all of which affect the ability of the compound to penetrate larval cuticle barriers and reach internal target sites.
In contrast, compounds 1, 2, 3, and 13 exhibited weak activity (LC50 > 340 µg mL−1) despite reasonable docking scores. Their reduced efficacy may stem from unfavorable pharmacokinetic characteristics, such as excessive polarity or steric bulk, limiting their diffusion through the larval membrane or their stability in the aquatic medium. These observations emphasize that a high docking score does not necessarily correlate with strong biological activity when physicochemical factors restrict compound-target encounters.
Conversely, certain compounds (e.g., 3 and 5) showed stronger binding energies than compound 4 but lower in vivo potency, indicating that their active conformations, although energetically favorable in silico, may not be accessible under physiological conditions due to rigidity, solubility constraints, or metabolic instability. Therefore, while docking and MD simulations provide valuable insights into potential target engagement, the overall larvicidal outcome reflects a complex interplay between molecular recognition, physicochemical behavior, and biological transport processes.
Taken together, the SAR findings suggest that the presence of electron-donating heteroatoms (N, O, S) within a moderately rigid aromatic framework enhances the dual-target potential of these furanone-derived heterocycles. Subtle changes in polarity or substituent positioning can drastically shift the biological outcome by modulating both target affinity and physicochemical accessibility. This highlights the importance of optimizing molecular balance between hydrophobicity and polarity to achieve both efficient penetration and stable binding. Future structure refinement of compound 4 and its analogues could therefore focus on maintaining its favorable interaction profile while improving aqueous stability and metabolic persistence to yield more effective, broad-spectrum larvicidal candidates. These observations also suggest that maintaining a balanced combination of aromatic scaffolds, heteroatom-containing functionalities, and moderate lipophilicity may represent useful structural considerations for the design of future insecticidal agents.
2.6 In silico ADME and drug-likeness evaluation
Given the partial inconsistencies between docking predictions and experimental bioassay outcomes, an in silico pharmacokinetic (ADME) analysis was conducted to determine whether variations in absorption, distribution, metabolism, or permeability among the synthesized compounds could account for their differing larvicidal efficacies. This complementary approach links molecular recognition with physicochemical and biological transport properties. The SwissADME platform was used to predict the key pharmacokinetic and physicochemical parameters of the heterocyclic derivatives, providing early insight into how these molecules may behave within insect systems, particularly their potential for cuticular or gut absorption and subsequent access to neural and hormonal targets.All synthesized compounds exhibited molecular weights and topological polar surface areas (TPSA) within the optimal range for passive membrane diffusion, together with moderate lipophilicity (Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≈ 2.9–4.5), supporting a balanced hydrophilic–lipophilic character (Table 3). The uniformly high gastrointestinal (GI) absorption predicted for the series indicates good potential for penetration through the insect gut or cuticular membranes, an essential prerequisite for bioavailability in larval systems.
P ≈ 2.9–4.5), supporting a balanced hydrophilic–lipophilic character (Table 3). The uniformly high gastrointestinal (GI) absorption predicted for the series indicates good potential for penetration through the insect gut or cuticular membranes, an essential prerequisite for bioavailability in larval systems.
| Compound | Molecular weight (g mol−1) | TPSA (Å2) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (iLOGP) P (iLOGP) |
GI absorption | BBB permeability | P-gp substrate | Bioavailability score |
|---|---|---|---|---|---|---|---|
| 1 | 292.33 | 35.53 | 3.36 | High | Yes | No | 0.55 |
| 2 | 291.34 | 38.33 | 3.20 | High | Yes | No | 0.55 |
| 3 | 399.48 | 55.40 | 3.50 | High | Yes | Yes | 0.55 |
| 4 | 397.47 | 38.77 | 4.36 | High | Yes | No | 0.55 |
| 5 | 425.48 | 55.84 | 4.49 | High | Yes | No | 0.55 |
| 7 | 411.45 | 69.40 | 3.45 | High | No | No | 0.55 |
| 8 | 410.46 | 72.05 | 3.33 | High | No | No | 0.55 |
| 9 | 407.46 | 59.39 | 4.16 | High | Yes | No | 0.55 |
| 10 | 306.36 | 54.98 | 2.99 | High | Yes | No | 0.55 |
| 11 | 322.42 | 70.00 | 3.21 | High | Yes | No | 0.55 |
| 12 | 324.81 | 56.15 | 3.54 | High | Yes | No | 0.55 |
| 13 | 335.40 | 62.09 | 3.45 | High | Yes | No | 0.55 |
| 14 | 349.39 | 72.65 | 2.88 | High | No | No | 0.55 |
Minor pharmacokinetic differences were nonetheless observed. Most compounds were predicted to cross the blood–brain barrier (BBB), consistent with possible access to neural targets such as AChE, whereas compounds 7, 8, and 14 were non-BBB-permeant, suggesting that their activity could be mediated primarily through peripheral or endocrine routes rather than direct neurotoxicity. Furthermore, only compound 3 was identified as a P-glycoprotein (P-gp) substrate, implying a higher likelihood of efflux and reduced intracellular retention relative to the rest of the series (Fig. 9).
The identical bioavailability score (0.55) across all derivatives reflects a consistent physicochemical design with generally favorable absorption and distribution potential (Table 3). Taken together, these data indicate that while no compound exhibits a pronounced pharmacokinetic advantage, subtle differences in BBB permeability, polarity, and efflux susceptibility may help explain part of the mismatch between docking affinities and larvicidal potencies (Fig. 9). Hence, the overall biological performance likely results from the combined effects of molecular recognition, membrane transport, and physicochemical stability rather than binding affinity alone.
Although these computational pharmacokinetic results complement the docking and bioassay findings, experimental validation at the enzyme and tissue levels remain essential to confirm actual absorption and target engagement. Nonetheless, the in silico screening provides a useful, resource-efficient framework for prioritizing insecticidal candidates with balanced permeability and stability profiles for future optimization.
3. Conclusion
This study successfully transformed the 2(3H)-furanone synthon into a diverse series of nitrogen-containing heterocycles including 1,3-dihydro-2H-pyrrol-2-ones, benzoxazinones, quinazolinones, and pyridazines, that were evaluated for their insecticidal potential. The bioassay results revealed wide variations in potency, with compound 4 emerging as the most active (LC50 = 0.1 µg mL−1), while several analogues displayed moderate activity. Molecular docking and dynamics simulations showed that all compounds can occupy the AChE catalytic gorge and the Met hormone-binding cavity, reproducing selected key contacts without a direct correlation between docking scores and biological activity. These findings indicate that larvicidal efficacy is governed not solely by binding strength but also by physicochemical and pharmacokinetic factors affecting membrane permeability and target accessibility.The ADME predictions confirmed favorable absorption and lipophilicity across the series, with minor differences, particularly in BBB permeability and efflux potential, likely contributing to variations in observed activity. Taken together, the integration of bioassay data, molecular modeling, dynamics simulation, and pharmacokinetic profiling provides a coherent mechanistic framework that links structural features with biological performance. The results suggest that 1,3-dihydro-2H-pyrrol-2-one-based candidates may represent promising scaffolds for further optimization toward efficient and selective larvicidal agents, with possible interaction across neurotoxic and hormonal targets, providing a basis for future research aimed at developing improved mosquito control agents.
4. Materials and methods
4.1 Synthesis and characterization
Solvents, reagents, and chemicals obtained from Sigma-Aldrich, Merck, Fluka, and El-Nasr pharmaceutical chemicals companies were purified and dried by standard procedures. The reactions and purity of products were tracked by thin-layer chromatography (TLC) using TLC aluminum sheets silica gel 60 F254 (Merck, Whitehouse Station, NJ). Melting points were measured on GALLENKAMP electrothermal melting point apparatus. The IR (infrared) spectra (ν, cm−1) were documented using the KBr wafer technique on Fourier Transform Infrared Thermo Electron Nicolet iS10 Spectrometer (Thermo Fisher Scientific Inc., Waltham, MA) at Faculty of Science, Ain Shams University. The 1H and 13C NMR spectra (δ, ppm) were recorded on a Varian 300 and 75 MHz GEMINI (GEMINI, Manufacturing & Engineering Inc., Anaheim, CA, USA) and on a Bruker AV 400 and 100 MHz with tetramethyl silane (TMS) as an internal standard, in DMSO-d6 as a solvent, at Faculty of Science, Cairo University and at the Microanalytical Unit – FOPCU – NMR laboratory, Faculty of pharmacy Ain Sham University, respectively. The EI-MS (electron impact mass spectra) were measured on a Shimadzu GC-MS-QP-1000EX mass spectrometer working at 70 eV (Shimadzu Scientific Instruments, Inc., MD, USA). The elemental analyses were executed on a PerkinElmer 2400 CHN elemental analyzer (PerkinElmer, Waltham, MA) at Faculty of Science, Ain Shams University. The synthon 2(3H)-furanone 1 was synthesized as a mixture of E- and Z-isomers (as confirmed by TLC under UV light using diethyl ether as a mobile phase) using the literature.444.2 3-(4-Methoxybenzylidene)-5-(p-tolyl)-1,3-dihydro-2H-pyrrol-2-one (2)45
A mixture of furanone 1 (1.46 g, 5 mmol) and dry ammonium acetate (0.42 g, 5.5 mmol) was refluxed in ethanol (20 mL) for 9 h. The residue was triturated with ethanol and the solid obtained was collected and recrystallized by ethanol to get green crystals, mp 200–202 °C, yield 90%. IR (KBr, ν, cm−1): 3175 (NH), 2957, 2918 (CH aliphatic), 1692 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O pyrrolone). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 10.40 (br.s, 1H, NH, exchangeable), 7.79–7.74 (m, 4H, Ar–H), 7.27 (d, 2H, Ar–H, J = 8.1 Hz), 7.11 (s, 1H, CH
O pyrrolone). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 10.40 (br.s, 1H, NH, exchangeable), 7.79–7.74 (m, 4H, Ar–H), 7.27 (d, 2H, Ar–H, J = 8.1 Hz), 7.11 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.03 (d, 2H, Ar–H, J = 8.7 Hz), 6.80 (s, 1H, C4–H pyrrole), 3.83 (s, 3H, OCH3), 2.35 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 171.3, 160.5, 144.8, 139.5, 132.4, 129.6 (2), 128.9, 128.3 (2), 127.1, 125.4, 121.0, 120.9, 114.8 (2), 96.8, 55.6, 21.2. Anal. calcd for C19H17NO2 (291.35): C, 78.33; H, 5.88; N, 4.81%; found: C, 78.29; H, 5.80; N, 4.87%.
), 7.03 (d, 2H, Ar–H, J = 8.7 Hz), 6.80 (s, 1H, C4–H pyrrole), 3.83 (s, 3H, OCH3), 2.35 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 171.3, 160.5, 144.8, 139.5, 132.4, 129.6 (2), 128.9, 128.3 (2), 127.1, 125.4, 121.0, 120.9, 114.8 (2), 96.8, 55.6, 21.2. Anal. calcd for C19H17NO2 (291.35): C, 78.33; H, 5.88; N, 4.81%; found: C, 78.29; H, 5.80; N, 4.87%.
4.3 N-Benzyl-2-(4-methoxybenzylidene)-4-oxo-4-(p-tolyl)butanamide (3)
A solution of furanone 1 (1.46 g, 5 mmol) and benzylamine (0.55 mL, 5 mmol) in ethanol (20 mL) was refluxed for 6 h. The solid obtained after cooling was collected and recrystallized by ethyl alcohol to produce white crystals, mp 260–262 °C, yield 60%. IR (KBr, ν, cm−1): 3263 (NH), 3062, 3031 (CH arom.), 2961, 2923 (CH aliph.), 1670 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1636 (C
O ketone), 1636 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.85 (br.s, 1H, NH), 7.46–6.97 (m, 14H, Ar–H + CH
O amide). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.85 (br.s, 1H, NH), 7.46–6.97 (m, 14H, Ar–H + CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 4.32 (d, 1H, 1H of N–CH2, J = 14.7 Hz), 4.04 (d, 1H, 1H of N–CH2, J = 15.0 Hz), 3.76 (s, 3H, OCH3), 3.40 (s, 2H, CH2CO), 2.34 (s, 3H, CH3). EI-MS (70 eV, m/z, %): 399.24 (M+, 48), 344.50 (77), 266.77 (95), 175.15 (100), 148.67 (66), 112.76 (26), 84.82 (36). Anal. calcd for C26H25NO3 (399.49): C, 78.17; H, 6.31; N, 3.51; found: C, 78.02; H, 6.25; N, 3.54%.
), 4.32 (d, 1H, 1H of N–CH2, J = 14.7 Hz), 4.04 (d, 1H, 1H of N–CH2, J = 15.0 Hz), 3.76 (s, 3H, OCH3), 3.40 (s, 2H, CH2CO), 2.34 (s, 3H, CH3). EI-MS (70 eV, m/z, %): 399.24 (M+, 48), 344.50 (77), 266.77 (95), 175.15 (100), 148.67 (66), 112.76 (26), 84.82 (36). Anal. calcd for C26H25NO3 (399.49): C, 78.17; H, 6.31; N, 3.51; found: C, 78.02; H, 6.25; N, 3.54%.
4.4 (E,Z)-3-(4-Methoxybenzylidene)-1-(4-methoxyphenyl)-5-(p-tolyl)-1,3-dihydro-2H-pyrrol-2-one (4)
A solution of furanone 1 (1.46 g, 5 mmol) and 4-methoxyaniline (0.62 g, 5 mmol) in ethanol (20 mL) was refluxed for 12 h, respectively. The solid obtained after cooling was collected and recrystallized by ethanol to produce yellow crystals, mp 190–192 °C, yield 58%. IR (KBr, ν, cm−1): 2955, 2934 (CH aliphatic), 1677 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O pyrrolone). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (E- & Z-isomers, 78
O pyrrolone). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (E- & Z-isomers, 78![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22%): 7.84 & 7.51 (d, 2H, Ar–H, J = 8.1 & 9 Hz), 7.36 & 7.31 (s, 1H, CH
22%): 7.84 & 7.51 (d, 2H, Ar–H, J = 8.1 & 9 Hz), 7.36 & 7.31 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.30–7.18 (m, 5H, Ar–H + C4–H pyrrolone), 7.15 & 7.06 (d, 2H, Ar–H, J = 8.7 & 8.1 Hz), 6.99 & 6.77 (d, 2H, Ar–H, J = 8.7 & 9 Hz), 6.63 & 6.50 (d, 2H, Ar–H, J = 6.6 Hz), 3.78 & 3.72 (s, 3H, OCH3), 3.67 & 3.61 (s, 3H, OCH3), 2.36 & 2.22 (s, 3H, CH3). Anal. calcd for C26H23NO3 (397.47): C, 78.57; H, 5.83; N, 3.52%; found: C, 78.63; H, 5.88; N, 3.55%.
), 7.30–7.18 (m, 5H, Ar–H + C4–H pyrrolone), 7.15 & 7.06 (d, 2H, Ar–H, J = 8.7 & 8.1 Hz), 6.99 & 6.77 (d, 2H, Ar–H, J = 8.7 & 9 Hz), 6.63 & 6.50 (d, 2H, Ar–H, J = 6.6 Hz), 3.78 & 3.72 (s, 3H, OCH3), 3.67 & 3.61 (s, 3H, OCH3), 2.36 & 2.22 (s, 3H, CH3). Anal. calcd for C26H23NO3 (397.47): C, 78.57; H, 5.83; N, 3.52%; found: C, 78.63; H, 5.88; N, 3.55%.
4.5 Methyl-4-(3-(4-methoxybenzylidene)-2-oxo-5-(p-tolyl)-2,3-dihydro-1H-pyrrol-1-yl)benzoate (5)
A mixture of furanone 1 (1.46 g, 5 mmol) and methyl 4-aminobenzoate (0.76 g, 5 mmol) was heated in oil bath at 150–160 °C for 4 h. The residue was triturated with methanol and the solid obtained was collected and recrystallized by methanol to give orange crystals, mp 160–162 °C, yield 78%. IR (KBr, ν, cm−1): 2952 (CH aliphatic), 1711 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.93 (d, 2H, Ar–H, J = 8.4 Hz), 7.86 (d, 2H, Ar–H, J = 8.4 Hz), 7.36 (s, 1H, CH
O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.93 (d, 2H, Ar–H, J = 8.4 Hz), 7.86 (d, 2H, Ar–H, J = 8.4 Hz), 7.36 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.23 (d, 2H, Ar–H, J = 8.4 Hz), 7.18–7.11 (m, 4H, Ar–H), 7.05 (d, 2H, Ar–H, J = 8.7 Hz), 6.82 (s, 1H, C4–H pyrrole), 3.84 (s, 6H, 2 –OCH3), 2.27 (s, 3H, CH3). 13C NMR (100 MHz, DMSO-d6): 166.2, 161.4, 139.4, 133.2 (2), 130.1 (2), 129.5 (2), 128.2, 127.9 (2), 127.5, 126.7, 115.2 (2), 103.1, 63.5, 55.9, 52.7, 21.3. EI-MS (70 eV, m/z, %): 425.20 (M+, 18), 327.41 (39), 277.44 (37), 239.47 (41), 190.42 (47), 164.09 (75), 84.48 (62), 70.82 (58), 44.85 (100). Anal. calcd for C27H23NO4 (425.48): C, 76.22; H, 5.45; N, 3.29%; found: C, 76.29; H, 5.50; N, 3.20%.
), 7.23 (d, 2H, Ar–H, J = 8.4 Hz), 7.18–7.11 (m, 4H, Ar–H), 7.05 (d, 2H, Ar–H, J = 8.7 Hz), 6.82 (s, 1H, C4–H pyrrole), 3.84 (s, 6H, 2 –OCH3), 2.27 (s, 3H, CH3). 13C NMR (100 MHz, DMSO-d6): 166.2, 161.4, 139.4, 133.2 (2), 130.1 (2), 129.5 (2), 128.2, 127.9 (2), 127.5, 126.7, 115.2 (2), 103.1, 63.5, 55.9, 52.7, 21.3. EI-MS (70 eV, m/z, %): 425.20 (M+, 18), 327.41 (39), 277.44 (37), 239.47 (41), 190.42 (47), 164.09 (75), 84.48 (62), 70.82 (58), 44.85 (100). Anal. calcd for C27H23NO4 (425.48): C, 76.22; H, 5.45; N, 3.29%; found: C, 76.29; H, 5.50; N, 3.20%.
4.6 (E,Z)-2-(1-(4-Methoxyphenyl)-4-oxo-4-(p-tolyl)but-1-en-2-yl)-4H-benzo[d][1,3]oxazin-4-one (7)
A mixture of furanone 1 (1.46 g, 5 mmol) and 2-aminobenzoic acid (0.69 g, 5 mmol) was heated in a sand bath at 140–150 °C for 4 h. Then, the residue was triturated with ethanol, and the solid obtained was filtered and recrystallized by methanol to get yellow crystals, mp 230–232 °C, yield 80%. IR (KBr, ν, cm−1): 2968, 2930 (CH aliphatic), 1745 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O oxazinone), 1702 (C
O oxazinone), 1702 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1648 (C
O ketone), 1648 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (Z- & E-isomers, 78
N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (Z- & E-isomers, 78![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22%): 8.16 (d, 1H, Ar–H, J = 8.1 Hz), 7.89–7.75 (m, 3H, Ar–H), 7.60–7.53 (m, 3H, Ar–H + CH
22%): 8.16 (d, 1H, Ar–H, J = 8.1 Hz), 7.89–7.75 (m, 3H, Ar–H), 7.60–7.53 (m, 3H, Ar–H + CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.34–7.04 (m, 4H, Ar–H), 7.0 (d, 2H, Ar–H, J = 8.7 Hz), 3.79 & 3.85 (s, 3H, OCH3), 3.44 & 3.17 (s, 2H, CH2), 2.20 & 2.37 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 166.6, 161.6, 160.7, 138.9, 137.8, 137.0, 136.1, 134.4, 132.4, 129.8 (2), 128.9, 128.4, 126.8 (2), 124.9 (2), 122.9, 121.4, 117.0, 114.9, 114.60 (2), 94.6, 55.4, 41.7, 20.6. EI-MS (70 eV, m/z, %): 411.46 (M+, 40), 410.29 (44), 373.41 (97), 312.86 (77), 285.11 (100), 253.97 (99), 175.45 (62), 105.04 (89), 91.12 (94). Anal. calcd for C26H21NO4 (411.46): C, 75.90; H, 5.14; N, 3.40%; found: C, 75.85; H, 5.19; N, 3.46%.
), 7.34–7.04 (m, 4H, Ar–H), 7.0 (d, 2H, Ar–H, J = 8.7 Hz), 3.79 & 3.85 (s, 3H, OCH3), 3.44 & 3.17 (s, 2H, CH2), 2.20 & 2.37 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 166.6, 161.6, 160.7, 138.9, 137.8, 137.0, 136.1, 134.4, 132.4, 129.8 (2), 128.9, 128.4, 126.8 (2), 124.9 (2), 122.9, 121.4, 117.0, 114.9, 114.60 (2), 94.6, 55.4, 41.7, 20.6. EI-MS (70 eV, m/z, %): 411.46 (M+, 40), 410.29 (44), 373.41 (97), 312.86 (77), 285.11 (100), 253.97 (99), 175.45 (62), 105.04 (89), 91.12 (94). Anal. calcd for C26H21NO4 (411.46): C, 75.90; H, 5.14; N, 3.40%; found: C, 75.85; H, 5.19; N, 3.46%.
4.7 (E,Z)-2-(1-(4-Methoxyphenyl)-4-oxo-4-(p-tolyl)but-1-en-2-yl)quinazolin-4(3H)-one (8)
A solution of benzoxazinone 7 (1.23 g, 3 mmol) and ammonium acetate (0.27 g, 3.5 mmol) in ethanol (20 mL) was refluxed for 9 h. The solid obtained after cooling was filtered and recrystallized by ethanol/dioxane mixture (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to produce green crystals, mp 220–222 °C, yield 75%. IR (KBr, ν, cm−1): 3286 (NH), 2954, 2933 (CH aliphatic), 1679 (C
1) to produce green crystals, mp 220–222 °C, yield 75%. IR (KBr, ν, cm−1): 3286 (NH), 2954, 2933 (CH aliphatic), 1679 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O ketone), 1641 (C
O ketone), 1641 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide), 1607 (C
O amide), 1607 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (Z- & E-isomers, 66
N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (Z- & E-isomers, 66![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 33%): 10.41 (br.s, 1H, NH, exchangeable), 7.83–7.74 (m, 2H, Ar–H), 7.57 & 7.40 (s, 1H, CH
33%): 10.41 (br.s, 1H, NH, exchangeable), 7.83–7.74 (m, 2H, Ar–H), 7.57 & 7.40 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.27–6.67 (m, 10H, Ar–H), 3.82 & 3.78 (s, 3H, OCH3), 3.50 & 3.45 (s, 2H, CH2), 2.33 & 2.23 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6 δ, ppm): (Z- and E-isomers, 66
), 7.27–6.67 (m, 10H, Ar–H), 3.82 & 3.78 (s, 3H, OCH3), 3.50 & 3.45 (s, 2H, CH2), 2.33 & 2.23 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6 δ, ppm): (Z- and E-isomers, 66![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 33%): 171.3, 169.2, 166.9, 160.8, 160.5, 147.4, 144.8, 139.4, 138.8, 137.8, 136.5, 135.3, 133.9, 133.7, 132.4, 131.6, 130.3 (2), 130.0, 129.7, 129.5, 129.1, 128.9, 128.3 (2), 128.0 (2), 127.8, 127.6, 127.1, 126.7, 125.6, 124.7, 114.8, 114.6 (2), 114.0, 75.0, 55.5, 29.1, 21.1. Anal. calcd for C26H22N2O3 (410.47): C, 76.08; H, 5.40; N, 6.82%. found: C, 76.18; H, 5.50; N, 6.78%.
33%): 171.3, 169.2, 166.9, 160.8, 160.5, 147.4, 144.8, 139.4, 138.8, 137.8, 136.5, 135.3, 133.9, 133.7, 132.4, 131.6, 130.3 (2), 130.0, 129.7, 129.5, 129.1, 128.9, 128.3 (2), 128.0 (2), 127.8, 127.6, 127.1, 126.7, 125.6, 124.7, 114.8, 114.6 (2), 114.0, 75.0, 55.5, 29.1, 21.1. Anal. calcd for C26H22N2O3 (410.47): C, 76.08; H, 5.40; N, 6.82%. found: C, 76.18; H, 5.50; N, 6.78%.
4.8 (E,Z)-4-(4-Methoxybenzylidene)-2-(p-tolyl)-1,4-dihydro-10H-pyridazino[6,1-b]quinazolin-10-one (9)
A solution of benzoxazinone 7 (1.23 g, 3 mmol) and hydrazine hydrate (0.2 mL, 3.2 mmol, 80%) in ethyl alcohol (20 mL) was refluxed for 16 h. The solid obtained after cooling was collected and recrystallized by ethanol to produce white crystals, mp 170–172 °C, yield 70%. IR (KBr, ν, cm−1): 3406 (br.NH, hydrogen bonding), 3037, 3004 (CH arom.), 2959, 2921 (CH aliph.), 1670 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1611 (C
O), 1611 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (E- & Z-isomers, 60
N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): (E- & Z-isomers, 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40%): 12.85 (br.s, 1H, NH), 8.18 (d, 1H, Ar–H, J = 7.8 Hz), 7.88–7.79 (m, 3H, Ar–H), 7.61–7.27 (m, 6H, Ar–H + CH
40%): 12.85 (br.s, 1H, NH), 8.18 (d, 1H, Ar–H, J = 7.8 Hz), 7.88–7.79 (m, 3H, Ar–H), 7.61–7.27 (m, 6H, Ar–H + CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.09–6.98 (m, 3H, Ar–H), 6.74 (s, 1H, C4–H pyridazine), 3.83 & 3.79 (s, 3H, OCH3), 2.18 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 168.9, 166.8, 161.3, 160.7, 147.1, 140.6, 136.9, 135.2, 134.3, 132.4, 132.2, 130.5, 129.8 (2), 128.9 (2), 127.7 (2), 126.9, 126.7, 125.9, 124.9, 122.7, 121.3, 116.9, 114.6 (2), 101.2, 94.4, 55.3, 41.5. EI-MS (70 eV, m/z, %): 407.90 (M+, 23), 376.16 (83), 257.94 (46), 227.25 (86), 202.19 (100), 160.39 (69), 110.75 (75), 80.15 (87), 76.86 (97). Anal. calcd for C26H21N3O2 (407.47): C, 76.64; H, 5.19; N, 10.31; found: C, 76.50; H, 5.11; N, 10.26%.
), 7.09–6.98 (m, 3H, Ar–H), 6.74 (s, 1H, C4–H pyridazine), 3.83 & 3.79 (s, 3H, OCH3), 2.18 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 168.9, 166.8, 161.3, 160.7, 147.1, 140.6, 136.9, 135.2, 134.3, 132.4, 132.2, 130.5, 129.8 (2), 128.9 (2), 127.7 (2), 126.9, 126.7, 125.9, 124.9, 122.7, 121.3, 116.9, 114.6 (2), 101.2, 94.4, 55.3, 41.5. EI-MS (70 eV, m/z, %): 407.90 (M+, 23), 376.16 (83), 257.94 (46), 227.25 (86), 202.19 (100), 160.39 (69), 110.75 (75), 80.15 (87), 76.86 (97). Anal. calcd for C26H21N3O2 (407.47): C, 76.64; H, 5.19; N, 10.31; found: C, 76.50; H, 5.11; N, 10.26%.
4.9 4-(4-Methoxybenzyl)-6-(p-tolyl)pyridazin-3(2H)-one (10)44
A solution of the furanone 1 (1.46 g, 5 mmol) and hydrazine hydrate (0.3 mL, 5.2 mmol, 80%) in absolute ethanol (20 mL) was refluxed for 6 h. After cooling, the solid obtained was filtered and recrystallized by ethanol to afford white crystals, mp 208–210 °C (decomp.), yield 80%. IR (KBr, ν, cm−1): 3207 (NH), 2963, 2919 (CH aliphatic), 1650 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1605 (C
O), 1605 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 13.03 (s, 1H, NH, exchangeable), 7.78 (s, 1H, C5–H pyridazine), 7.68 (d, 2H, Ar–H, J = 8.1 Hz), 7.29–6.25 (m, 4H, Ar–H), 6.86 (d, 2H, Ar–H, J = 9 Hz), 3.78 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.34 (s, 3H, CH3). Anal. calcd for C19H18N2O2 (306.36): C, 74.49; H, 5.92; N, 9.14%; found: C, 74.55; H, 5.86; N, 9.11%.
N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 13.03 (s, 1H, NH, exchangeable), 7.78 (s, 1H, C5–H pyridazine), 7.68 (d, 2H, Ar–H, J = 8.1 Hz), 7.29–6.25 (m, 4H, Ar–H), 6.86 (d, 2H, Ar–H, J = 9 Hz), 3.78 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.34 (s, 3H, CH3). Anal. calcd for C19H18N2O2 (306.36): C, 74.49; H, 5.92; N, 9.14%; found: C, 74.55; H, 5.86; N, 9.11%.
4.10 4-(4-Methoxybenzyl)-6-(p-tolyl)pyridazine-3(2H)-thione (11)
A suspension of pyridazinone 10 (0.92 g, 3 mmol) and phosphorus pentasulfide (0.67 g, 3 mmol) in dry toluene (20 mL) was refluxed for 6 h. The solid obtained after cooling was filtered and recrystallized by ethanol to produce grey crystals, mp 185–187 °C, yield 40%. IR (KBr, ν, cm−1): 3226 (NH), 3060, 3002 (CH arom.), 2952, 2923 (CH aliph.), 1609 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1247 (C
N), 1247 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 14.85 (br.s, 1H, NH), 7.81–7.70 (m, 3H, Ar–H + C5–H pyridazine), 7.32–7.21 (m, 4H, Ar–H), 6.86 (d, 2H, Ar–H, J = 8.4 Hz), 4.09 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.33 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 179.1, 157.9, 152.0, 149.4, 139.9, 131.2, 130.3 (2), 129.7 (2), 129.6 (2), 125.9, 123.0, 113.8 (2), 55.0, 37.7, 20.9. Anal. calcd for C19H18N2OS (322.43): C, 70.78; H, 5.63; N, 8.69%. found: C, 70.72; H, 5.67; N, 8.61%.
S). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 14.85 (br.s, 1H, NH), 7.81–7.70 (m, 3H, Ar–H + C5–H pyridazine), 7.32–7.21 (m, 4H, Ar–H), 6.86 (d, 2H, Ar–H, J = 8.4 Hz), 4.09 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.33 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 179.1, 157.9, 152.0, 149.4, 139.9, 131.2, 130.3 (2), 129.7 (2), 129.6 (2), 125.9, 123.0, 113.8 (2), 55.0, 37.7, 20.9. Anal. calcd for C19H18N2OS (322.43): C, 70.78; H, 5.63; N, 8.69%. found: C, 70.72; H, 5.67; N, 8.61%.
4.11 3-Chloro-4-(4-methoxybenzyl)-6-(p-tolyl)pyridazine (12)
A suspension of pyridazinone 10 (0.92 g, 3 mmol) in phosphorus oxychloride (5 mL) was heated in water bath at 80–90 °C for 8 h. After cooling, the residue was triturated with ice-cold water. The solid obtained was filtered and recrystallized by ethanol–dioxane (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to give grey crystals, mp 210–212 °C, yield 45%. IR (KBr, ν, cm−1): 3036, 3006 (CH arom.), 2926, 2908 (CH aliph.), 1612 (C
1) to give grey crystals, mp 210–212 °C, yield 45%. IR (KBr, ν, cm−1): 3036, 3006 (CH arom.), 2926, 2908 (CH aliph.), 1612 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 8.18 (s, 1H, C5–H pyridazine), 7.99 (d, 2H, Ar–H, J = 8.1 Hz), 7.35 (d, 2H, Ar–H, J = 8.1 Hz), 7.23 (d, 2H, Ar–H, J = 8.4 Hz), 6.88 (d, 2H, Ar–H, J = 8.7 Hz), 4.06 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.37 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 164.8, 158.4, 155.8, 144.4, 138.4, 132.8, 130.1 (2), 129.8 (2), 128.4 (2), 126.8, 124.3, 123.1, 114.1 (2), 55.1, 36.7, 21.0. Anal. calcd for C19H17ClN2O (324.81): C, 70.26; H, 5.28; N, 8.62%; found: C, 70.32; H, 5.35; N, 8.52%.
N). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 8.18 (s, 1H, C5–H pyridazine), 7.99 (d, 2H, Ar–H, J = 8.1 Hz), 7.35 (d, 2H, Ar–H, J = 8.1 Hz), 7.23 (d, 2H, Ar–H, J = 8.4 Hz), 6.88 (d, 2H, Ar–H, J = 8.7 Hz), 4.06 (s, 2H, CH2), 3.71 (s, 3H, OCH3), 2.37 (s, 3H, CH3). 13C NMR (75 MHz, DMSO-d6, δ, ppm): 164.8, 158.4, 155.8, 144.4, 138.4, 132.8, 130.1 (2), 129.8 (2), 128.4 (2), 126.8, 124.3, 123.1, 114.1 (2), 55.1, 36.7, 21.0. Anal. calcd for C19H17ClN2O (324.81): C, 70.26; H, 5.28; N, 8.62%; found: C, 70.32; H, 5.35; N, 8.52%.
4.12 1-(2-Hydroxyethyl)-3-(4-methoxybenzylidene)-5-(p-tolyl)-1,3-dihydro-2H-pyrrol-2-one (13)
2-Aminoethanol (0.2 mL, 3 mmol, 98%) was added dropwise to a stirred solution of furanone 1 (0.88 g, 3 mmol) in dioxane (20 mL) at room temperature, and stirring stayed for 8 h. The orange precipitate obtained was collected and recrystallized by ethanol to afford orange crystals, mp 170–172 °C, yield 65%. IR (KBr, ν, cm−1): 3422 (br. OH), 3105, 3073 (CH arom.), 2954, 2930 (CH aliph.), 1667 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.75 (d, 2H, Ar–H, J = 8.7 Hz), 7.54 (d, 2H, Ar–H, J = 7.8 Hz), 7.31 (d, 2H, Ar–H, J = 7.8 Hz), 7.22 (s, 1H, CH
O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 7.75 (d, 2H, Ar–H, J = 8.7 Hz), 7.54 (d, 2H, Ar–H, J = 7.8 Hz), 7.31 (d, 2H, Ar–H, J = 7.8 Hz), 7.22 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.01 (d, 2H, Ar–H, J = 8.7 Hz), 6.34 (s, 1H, C4–H pyrrolone), 3.81 (s, 4H, OCH3 + OH), 3.67–3.63 (m, 4H, 2 CH2), 2.37 (s, 3H, CH3). Anal. calcd for C21H21NO3 (335.40): C, 75.20; H, 6.31; N, 4.18%. found: C, 75.02; H, 6.25; N, 4.30%.
), 7.01 (d, 2H, Ar–H, J = 8.7 Hz), 6.34 (s, 1H, C4–H pyrrolone), 3.81 (s, 4H, OCH3 + OH), 3.67–3.63 (m, 4H, 2 CH2), 2.37 (s, 3H, CH3). Anal. calcd for C21H21NO3 (335.40): C, 75.20; H, 6.31; N, 4.18%. found: C, 75.02; H, 6.25; N, 4.30%.
4.13 4-(4-Methoxybenzylidene)-3-oxo-6-(p-tolyl)-3,4-dihydropyridazine-1(2H)-carboxamide (14)
A solution of furanone 1 (0.88 g, 3 mmol) and semicarbazide hydrochloride (0.35 g, 3 mmol) in ethanol (20 mL) containing sodium acetate anhydrous (0.25 g, 3 mmol) was refluxed for 8 h. The solid obtained after cooling was collected and recrystallized by ethanol to produce yellow crystals, mp 240–242 °C, yield 85%. IR (KBr, ν, cm−1): 3424, 3320, 3290 (NH, NH2), 1709, 1675 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 8.55 (br.s, 1H, NHCO, lactam form, 70%), 7.80 (d, 2H, Ar–H, J = 8.7 Hz), 7.60 (d, 2H, Ar–H, J = 7.8 Hz), 7.25 (d, 2H, Ar–H, J = 7.7 Hz), 7.24 (s, 1H, CH
O). 1H NMR (300 MHz, DMSO-d6, δ, ppm): 8.55 (br.s, 1H, NHCO, lactam form, 70%), 7.80 (d, 2H, Ar–H, J = 8.7 Hz), 7.60 (d, 2H, Ar–H, J = 7.8 Hz), 7.25 (d, 2H, Ar–H, J = 7.7 Hz), 7.24 (s, 1H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 7.03 (d, 2H, Ar–H, J = 8.4 Hz), 6.56 (s, 1H, C5–H pyridazine), 6.13 (br.s, 2H, NH2), 5.83 (br.s, 1H, OH lactim form, 30%), 3.83 (s, 3H, OCH3), 2.35 (s, 3H, CH3). 13C NMR (100 MHz, DMSO-d6): 170.0, 161.1, 139.6, 132.9 (2), 131.8, 129.5 (2), 128.3, 127.8 (2), 127.4, 125.5, 115.2 (2), 55.8, 21.4. EI-MS (70 eV, m/z, %): 349.87 (M+, 24), 284.25 (32), 256.01 (81), 222.91 (67), 166.42 (29), 130.21 (50), 120.08 (57), 102.89 (47), 91.95 (52), 77.17 (100). Anal. calcd for C20H19N3O3 (349.39): C, 68.75; H, 5.48; N, 12.03; found: C, 68.64; H, 5.42; N, 12.00%.
), 7.03 (d, 2H, Ar–H, J = 8.4 Hz), 6.56 (s, 1H, C5–H pyridazine), 6.13 (br.s, 2H, NH2), 5.83 (br.s, 1H, OH lactim form, 30%), 3.83 (s, 3H, OCH3), 2.35 (s, 3H, CH3). 13C NMR (100 MHz, DMSO-d6): 170.0, 161.1, 139.6, 132.9 (2), 131.8, 129.5 (2), 128.3, 127.8 (2), 127.4, 125.5, 115.2 (2), 55.8, 21.4. EI-MS (70 eV, m/z, %): 349.87 (M+, 24), 284.25 (32), 256.01 (81), 222.91 (67), 166.42 (29), 130.21 (50), 120.08 (57), 102.89 (47), 91.95 (52), 77.17 (100). Anal. calcd for C20H19N3O3 (349.39): C, 68.75; H, 5.48; N, 12.03; found: C, 68.64; H, 5.42; N, 12.00%.
4.14 Biological assay
The Culex pipiens strain used in this study originated from a long-established laboratory population maintained for nearly forty consecutive generations at the Entomology Department, Faculty of Science, Ain Shams University. Mosquitoes were reared under consistently regulated insectary conditions, including a temperature close to 27 °C (±2 °C), relative humidity around 75%, and a photoperiod of 12 h light followed by 12 h darkness, following routine colony management practices.6–8,25,28,49–51,53Larval toxicity tests were performed following the standard procedures outlined by the World Health Organization.54 Third-instar Culex pipiens larvae (20 individuals per replicate) were subjected to a gradient of test concentrations 250, 500, 1000, 1500, and 2000 µg mL−1, to determine the larvicidal potency of each compound. Stock solutions were prepared in dimethylformamide (DMF) and subsequently diluted with distilled water to achieve the desired concentrations. For each treatment, three independent replicates were conducted alongside a solvent control containing only DMF and water. After 24 h of exposure, larval mortality was recorded, with death confirmed by the lack of movement upon gentle stimulation.6–8,25,49–51,53,55
The recorded larval mortality data were analyzed statistically using the LDP Line software. Median lethal concentrations (LC50) and their corresponding 95% confidence limits were estimated to quantify larvicidal potency. Mortality observed in control groups was adjusted through Abbott's correction formula. The reliability and precision of the fitted probit regression lines were verified using Finney's statistical approach, the chi-square goodness-of-fit test, and the coefficient of determination (r2).56,57 In addition, the relative toxicity (toxicity index, TI) of each tested compound was computed following the equation proposed by Sun and colleagues.58
4.15 Molecular docking and dynamics simulations
Molecular docking and dynamics studies were conducted to rationalize the larvicidal activity of the tested heterocyclic compounds against Culex pipiens and to elucidate their possible binding mechanisms toward two molecular targets: acetylcholinesterase (AChE) and the methoprene-tolerant receptor (Met). These proteins represent the principal neurotoxic and endocrine targets in mosquitoes, respectively. The dual-target in silico strategy was designed to determine whether the compounds act as AChE inhibitors similar to organophosphate or carbamate insecticides, as juvenile-hormone agonists resembling pyriproxyfen, or as multitarget agents capable of interacting with both sites, thereby providing a promising route for resistance management and the development of broad-spectrum larvicides.Because no crystallographic structures are available for Culex pipiens AChE or Met, homology models of both proteins were generated. The amino acid sequences were retrieved from UniProt under accession numbers Q86GC8 (AChE) and Q4ZH01 (Met). For AChE, the model was built using PDB 5X61 (chain A) as a template, while for Met, the PAS-B domain was modeled based on PDB 6E3U (chain B) containing the co-crystallized ligand HNJ.
To ensure accurate definition of the binding cavities, the reference ligands BT7 (from PDB 6ARY) and HNJ (from PDB 6E3U) were transferred into the respective Culex models by structural superposition and alignment with their template proteins. This procedure preserved the biologically relevant orientation of the ligands and allowed precise mapping of the active and hormonal binding sites for subsequent docking. The accuracy of each docking protocol was further validated by redocking the transferred ligands into their corresponding binding sites, confirming that the reproduced poses closely matched the experimental orientations (RMSD < 2.0 Å), thereby demonstrating proper grid definition and parameter calibration.
Docking simulations were carried out for all synthesized compounds, along with the co-crystallized ligands and the benchmark insecticides chlorpyrifos (AChE reference) and pyriproxyfen (Met reference), using the Molecular Operating Environment (MOE 2024.06) software. Protein structures were prepared by adding polar hydrogens, assigning Gasteiger charges, and performing energy minimization to optimize the geometry of the active sites. For each target, a total of 100 docking poses were generated per ligand, and the best-ranked conformations were selected based on multiple criteria, including the lowest binding energy, an RMSD ≤ 2.0 Å, and consistent alignment of key interacting residues with those of the co-crystallized and reference ligands. The binding interactions were then analyzed to identify the most relevant amino acid residues involved in hydrogen bonding, hydrophobic contacts, and π–π stacking within the defined active-site pocket.6–8,25,49–53,59
To validate the molecular docking outcomes and explore the conformational stability of the ligand–protein assemblies over time, molecular dynamics (MD) simulations were conducted using the Desmond package integrated within the Schrödinger Suite (Schrödinger, LLC, New York, NY, USA). Four representative complexes were selected for simulation: the most active compound (compound 4), the co-crystallized ligands of AChE and Met, and the benchmark insecticides chlorpyrifos and pyriproxyfen. Each molecular system was immersed in an explicit TIP3P water model confined within an orthorhombic box extending 10 Å beyond the solute in all directions. The systems were neutralized by adding appropriate numbers of Na+ and Cl− ions and adjusted to a physiological salt concentration of 0.15 M NaCl.
All simulations employed the OPLS4 force field and periodic boundary conditions. Prior to production, the solvated complexes were energy-minimized through 2000 iterations of steepest descent followed by conjugate-gradient refinement. The equilibrated configurations were then propagated for 100 ns under an NPT ensemble at 300 K and 1 atm, controlled by the Nose–Hoover chain thermostat and Martyna–Tobias–Klein barostat algorithms. Long-range electrostatic interactions were computed using the Particle-Mesh Ewald (PME) approach with a 9.0 Å cutoff for short-range forces. To maintain bond rigidity involving hydrogen atoms, the SHAKE constraint was applied, enabling a stable integration time step of 2 fs.6,7,49,52
Trajectory analyses were performed using the Simulation Interaction Diagram (SID) module implemented in Desmond to investigate the temporal behavior and structural integrity of each complex. The Root Mean Square Deviation (RMSD) profile was employed to track overall conformational stability, whereas Root Mean Square Fluctuation (RMSF) values characterized the mobility of individual amino acid residues, particularly those located in the binding cavity. To further examine the interaction dynamics, contact histograms and frequency plots were generated, illustrating the persistence of hydrogen bonds, hydrophobic interactions, and π–π stacking events throughout the 100 ns simulation period.
Complementary parameters, including the radius of gyration (Rg), solvent-accessible surface area (SASA), and total potential energy, were also monitored to provide a comprehensive assessment of system equilibration and compactness (File S2). Comparative trajectory profiles of compound 4, the co-crystallized ligands, and the reference insecticides corroborated the docking results, delineating key stabilizing residues and revealing distinct dynamic patterns associated with neurotoxic and endocrine-modulating mechanisms of action.6,7,49,52
4.16 Rationale for in silico validation
The computational validation was performed to bridge the observed larvicidal bioassay results with the possible molecular mechanisms underlying the biological activity of the tested compounds. Because conventional insecticides targeting a single site such as acetylcholinesterase have frequently led to the development of resistance in mosquito populations, understanding whether newly synthesized molecules can simultaneously interact with multiple molecular targets has become a critical step in rational insecticide design. By combining docking, redocking, and molecular dynamics simulations, this study sought to identify the molecular determinants of ligand recognition and to evaluate whether the tested compounds could behave as neurotoxic inhibitors, juvenile-hormone mimics, or dual-action agents.The redocking of the co-crystallized ligands served as an essential internal validation step that confirmed the reliability of the docking protocol, the accuracy of grid box definition, and the consistency of scoring functions. Comparison of binding residues between the co-crystallized ligands, reference insecticides, and tested compounds enabled the identification of conserved amino acids that are likely to mediate stable and biologically meaningful interactions. Furthermore, molecular dynamics simulations provided an energetic and conformational validation of docking predictions, revealing the stability of each complex under near-physiological conditions and offering insight into the flexibility of active-site residues and the persistence of key noncovalent contacts.
Together, the integrated bioassay and in silico framework employed in this study provides a mechanistic basis for interpreting larvicidal potency in terms of molecular binding behavior. This approach also supports the broader objective of discovering multitarget larvicides capable of simultaneously disrupting both the neural and endocrine systems of Culex pipiens, thereby offering a promising strategy to overcome resistance and enhance vector control efficacy.
4.17 In silico ADME and drug-likeness evaluation
The pharmacokinetic and physicochemical properties of the synthesized compounds were predicted in silico using the SwissADME web tool (https://www.swissadme.ch/, Swiss Institute of Bioinformatics). Canonical SMILES of the optimized structures were submitted to evaluate lipophilicity, solubility, molecular descriptors, and major pharmacokinetic parameters including GI absorption, BBB permeability, P-gp interaction, and CYP450 inhibition potential. Drug-likeness was assessed according to the Lipinski, Veber, Egan, Ghose, and Muegge filters, and visualized through bioavailability radar and BOILED-Egg plots. All calculations were performed using default SwissADME settings, and the results were used to support the ADME-based interpretation of the biological and docking data.6,7,52Author contributions
N. M. Gad was responsible for the synthesis and characterization of compounds obtained. D. S. A. Haneen, M. H. Hekal, A. A. Abdalha, W. S. I. Abou-Elmagd, and S. K. Ramadan supervised and conceptualized the chemical part of the study and provided critical revisions to the manuscript. E. M. Hosni conducted the larvicidal bioassays and analyzed the insecticidal activity data. M. Kamal conceptualized and conducted in silico studies, including molecular docking and molecular dynamics simulations. All authors contributed to the writing and revision of the manuscript and approved the final version.Conflicts of interest
No potential conflict of interest was reported by the author(s).Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information (SI) files. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00490c.References
- A. Husain, A. Ahmad, M. M. Alam, M. Ajmal and P. Ahuja, Fenbufen based 3-[5-(substituted aryl)-1,3,4-oxadiazol-2-yl]-1-(biphenyl-4-yl)propan-1-ones as safer antiinflammatory and analgesic agents, Eur. J. Med. Chem., 2009, 44, 3798–3804, DOI:10.1016/J.EJMECH.2009.04.009.
- M. M. Alam, A. Husain, S. M. Hasan, Suruchi and T. Anwer, Synthesis and pharmacological evaluation of 2(3H)-furanones and 2(3H)-pyrrolones, combining analgesic and anti-inflammatory properties with reduced gastrointestinal toxicity and lipid peroxidation, Eur. J. Med. Chem., 2009, 44(6), 2636–2642, DOI:10.1016/j.ejmech.2008.10.030.
- A. K. El-Ziaty, W. S. I. Abou-Elmagd, S. K. Ramadan and A. I. Hashem, Synthesis and biological screening of some chromonyl-substituted heterocycles derived from 2(3H)-furanone derivative, Synth. Commun., 2017, 47, 471–480, DOI:10.1080/00397911.2016.1271896.
- A. Husain, M. M. Alam, S. M. Hasan and M. S. Yar, 2(3H)-furanones and 2(3H)-pyrrolones: synthesis and antimycobacterial evaluation, Acta Pol. Pharm., 2009, 66, 173–180 CAS . https://pubmed.ncbi.nlm.nih.gov/19719052/.
- S. K. Ramadan, D. R. Abdel Haleem, H. S. M. Abd-Rabboh, N. M. Gad, W. S. I. Abou-Elmagd and D. S. A. Haneen, Synthesis, SAR studies, and insecticidal activities of certain N-heterocycles derived from 3-((2-chloroquinolin-3-yl)methylene)-5-phenylfuran-2(3H)-one against Culex pipiens L. larvae, RSC Adv., 2022, 12, 13628–13638, 10.1039/D2RA02388A.
- E. A. E. El-Helw, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and S. K. Ramadan, Synthesis of Benzo[h]quinoline Derivatives and Evaluation of Their Insecticidal Activity Against Culex pipiens L. Larvae, Eur. J. Med. Chem., 2025, 117565, DOI:10.1016/J.EJMECH.2025.117565.
- S. K. Ramadan, W. S. I. Abou-Elmagd, E. M. Hosni, M. Kamal, A. I. Hashem and E. A. E. El-Helw, Synthesis, in vivo evaluation, and in silico molecular docking of benzo[h]quinoline derivatives as potential Culex pipiens L. larvicides, Bioorg. Chem., 2025, 154, 108090, DOI:10.1016/J.BIOORG.2024.108090.
- E. A. E. El-Helw, E. M. Hosni, M. Kamal, A. I. Hashem and S. K. Ramadan, Synthesis, insecticidal Activity, and molecular docking analysis of some benzo[h]quinoline derivatives against Culex pipiens L. Larvae, Bioorg. Chem., 2024, 107591, DOI:10.1016/J.BIOORG.2024.107591.
- W. S. I. Abou-Elmagd and A. I. Hashem, Conversion of Some 2(3H)-Furanones into Pyrrolinotriazine and Oxazolopyrimidine Derivatives, J. Heterocycl. Chem., 2012, 49, 947–950, DOI:10.1002/CHIN.201306131.
- S. K. Ramadan, W. S. I. Abou-Elmagd and A. I. Hashem, Reactions of 2(3H)-furanones, Synth. Commun., 2019, 49, 3031–3057, DOI:10.1080/00397911.2019.1647441.
- N. M. Gad, W. S. I. Abou-Elmagd, D. S. A. Haneen and S. K. Ramadan, Reactivity of 5-phenyl-3-[(2-chloroquinolin-3-yl)methylene] furan-2(3H)-one towards hydrazine and benzylamine: A comparative study, Synth. Commun., 2021, 51, 1384–1397, DOI:10.1080/00397911.2021.1882498.
- A. I. Hashem, W. S. I. Abou-Elmagd, A. K. El-Ziaty and S. K. Ramadan, Ring Transformation of a 2(3H)-furanone Derivative into Oxazinone and Pyrimidinone Heterocycles, J. Heterocycl. Chem., 2017, 54, 3711–3715, DOI:10.1002/JHET.2937.
- Y. Xu and Q.-X. Guo, Syntheses of Heterocyclic Compounds under Microwave Irradiation, Heterocycles, 2004, 63, 903–974 CrossRef CAS.
- S. R. Atta-Allah, R. S. Gouhar, M. M. Hemdan, W. S. I. Abou-Elmagd, D. S. A. Haneen, K. A. A. Kandeel and A. S. A. Youssef, Synthesis and antitumor activity evaluation of some novel pyrazolotriazine derivatives, Synth. Commun., 2017, 47, 299–309, DOI:10.1080/00397911.2016.1262422.
- A. A. El-Barbary, D. R. Imam, M. M. T. El-Tahawy, S. M. El-Hallouty, N. A. Kheder and A. I. Khodair, Unexpected synthesis, characterization, biological evaluations, and computational details of novel nucleosides containing triazine-pyrrole hybrid, J. Mol. Struct., 2023, 1272, 134182, DOI:10.1016/J.MOLSTRUC.2022.134182.
- K. S. Mohamed and A. A. Fadda, Synthesis, characterization and cytotoxicity evaluation of some novel pyrazole and pyrrole derivatives containing benzothiazole moiety, Heterocycles, 2015, 91, 1937–1954, DOI:10.3987/COM-15-13304.
- S. K. Ramadan, H. S. M. Abd-Rabboh, N. M. Gad, W. S. I. Abou-Elmagd and D. S. A. Haneen, Synthesis and Characterization of Some Chitosan-Quinoline Nanocomposites as Potential Insecticidal Agents, Polycyclic Aromat. Compd., 2023, 43, 7013–7026, DOI:10.1080/10406638.2022.2128831.
- M. H. Hekal, A. I. Hashem, F. S. M. A. El-Azm, D. R. Abdel-Haleem, E. H. Rafat and Y. M. Ali, Rational design and synthesis of new pyrrolone candidates as prospective insecticidal agents against Culex pipiens L. Larvae, Sci. Rep., 2024, 14, 1–17, DOI:10.1038/S41598-024-74011-5.
- A. A. Fadda, R. Rabie, S. Bondock and H. A. Etman, Derivational, Structural, and Biological Studies of Some New Pyrazolyl, Isoxazolyl, Pyrimidinyl, Pyridazinyl, and Pyridopyridazinyl from 4-Substituted Antipyrine, J. Heterocycl. Chem., 2017, 54, 1304–1310, DOI:10.1002/JHET.2707.
- A. I. Khodair, M. A. Alsafi and M. S. Nafie, Synthesis, molecular modeling and anti-cancer evaluation of a series of quinazoline derivatives, Carbohydr. Res., 2019, 486, 107832, DOI:10.1016/j.carres.2019.107832.
- H. E. Hashem, D. S. A. Haneen, K. F. Saied and A. S. A. Youssef, Synthesis of new annulated pyridazine derivatives and studying their antioxidant and antimicrobial activities, Synth. Commun., 2019, 49, 3169–3180, DOI:10.1080/00397911.2019.1658786.
- R. S. Gouhar, D. S. A. Haneen and S. M. El-Hallouty, Synthesis and Anticancer Evaluation of Some Novel Quinazolin-4(3H)-one Derivatives, J. Heterocycl. Chem., 2019, 56, 1651–1660, DOI:10.1002/JHET.3559.
- D. S. A. Haneen, R. S. Gouhar, H. E. Hashem and A. S. A. Youssef, Synthesis and reactions of 4H-3,1-benzoxazin-4-one derivative bearing pyrazolyl moiety as antimicrobial and antioxidant agents, Synth. Commun., 2019, 49, 2840–2855, DOI:10.1080/00397911.2019.1646288.
- X. Du, Y. Li, Y. L. Xia, S. M. Ai, J. Liang, P. Sang, X. L. Ji and S. Q. Liu, Insights into Protein-Ligand Interactions: Mechanisms, Models, and Methods, Int. J. Mol. Sci., 2016, 17(2), 144, DOI:10.3390/ijms17020144.
- A. A. Rashad, S. M. Gad Allaha, R. M. Badawy, I. I. Ahmed, M. G. Shehata and M. Kamal, Aquatic Larvicidal Efficacy of Periplaneta americana Chitin and Essential Oils Against Culex pipiens: A Comparative Assessment, Egypt. J. Aquat. Biol., 2025, 29, 2911–2938, DOI:10.21608/EJABF.2025.434231.
- E. B. Vinogradova, Culex pipiens Pipiens Mosquitoes: Taxonomy, Distribution, Ecology, Physiology, Genetics, Applied Importance and Control, Pensoft Publishers, Sofia, Bulgaria, 2000 Search PubMed.
- M. Liu, Y. Zhang, Q. Li, X. Zhou, T. Yan, J. Li, H. Zhang, L. Wang, G. Wang, R. Li, Y. Tong and X. Zeng, Spatial distribution and environmental correlations of Culex pipiens pallens (Diptera: Culicidae) in Haidian district, Beijing, J. Med. Entomol., 2024, 61, 948–958, DOI:10.1093/JME/TJAE063.
- E. Adly, A. A. Hegazy, M. Kamal and S. H. Abu-Hussien, Midguts of Culex pipiens L. (Diptera: Culicidae) as a potential source of raw milk contamination with pathogens, Sci. Rep., 2022, 12, 13183, DOI:10.1038/s41598-022-16992-9.
- A. Farajollahi, D. M. Fonseca, L. D. Kramer and A. Marm Kilpatrick, “Bird biting” mosquitoes and human disease: A review of the role of Culex pipiens complex mosquitoes in epidemiology, Infect., Genet. Evol., 2011, 11, 1577–1585, DOI:10.1016/J.MEEGID.2011.08.013.
- H. Alout, P. Labbé, A. Berthomieu, P. Makoundou, P. Fort, N. Pasteur and M. Weill, High chlorpyrifos resistance in Culex pipiens mosquitoes: strong synergy between resistance genes, Heredity, 2016, 116, 224–231, DOI:10.1038/HDY.2015.92.
- P. Labbé, J.-P. David, H. Alout, P. Milesi, N. Pasteur, M. Weill, P. Labbé, J.-P. David, H. Alout, P. Milesi, L. Djogbénou, N. Pasteur and M. Weill, Evolution of Resistance to Insecticide in Disease Vectors Evolution of Resistance to Insecticide in Disease Vectors 14, 2017, DOI:10.1016/B978-0-12-799942-5.00014-7.
- N. Liu, Insecticide resistance in mosquitoes: impact, mechanisms, and research directions, Annu. Rev. Entomol., 2015, 60, 537–559, DOI:10.1146/ANNUREV-ENTO-010814-020828.
- Y. Sato, S. Jang, K. Takeshita, H. Itoh, H. Koike, K. Tago, M. Hayatsu, T. Hori and Y. Kikuchi, Insecticide resistance by a host-symbiont reciprocal detoxification, Nat. Commun., 2021, 12(1), 1–8, DOI:10.1038/s41467-021-26649-2.
- Y. P. Pang, Insect Acetylcholinesterase as a Target for Effective and Environmentally Safe Insecticides, Adv. Insect Physiol., 2014, 46, 435–494, DOI:10.1016/B978-0-12-417010-0.00006-9.
- S. Sakthivel, H. S. Mohideen, C. Raman and S. Bin Mohamad, Potential Acetylcholinesterase Inhibitor Acting on the Pesticide Resistant and Susceptible Cotton Pests, ACS Omega, 2022, 7, 20515–20527, DOI:10.1021/ACSOMEGA.1C07359.
- Y. Li, W. Wu, R. Jian, X. Ren, X. Chen, W. D. Hong, M. Wu, J. Cai, C. Lao, X. Xu and Z. Sheng, Larvicidal, acetylcholinesterase inhibitory activities of four essential oils and their constituents against Aedes albopictus, and nanoemulsion preparation, J. Pest Sci., 2023, 96, 961–971, DOI:10.1007/S10340-022-01555-8.
- A. K. Ray and M. C. Ghosh, Aquatic Toxicity of Carbamates and Organophosphates, Toxicology of Organophosphate & Carbamate Compounds, 2006, pp. 657–672, DOI:10.1016/B978-012088523-7/50046-6.
- B. C. Geyer, T. Evron, H. Soreq and T. S. Mor, Organophosphate intoxication: Molecular consequences, mechanisms and solutions, Handbook of Toxicology of Chemical Warfare Agents, 2009, pp. 691–717, DOI:10.1016/B978-012374484-5.00046-8.
- World Health Organization, Manual for monitoring insecticide resistance in mosquito vectors and selecting appropriate interventions, Organização Mundial Da Saúde, 2022, pp. 1–65. https://www.who.int/publications/i/item/9789240051089, accessed March 29, 2024.
- M. Jindra and L. Bittova, The juvenile hormone receptor as a target of juvenoid “insect growth regulators,”, Arch. Insect Biochem. Physiol., 2020, 103, e21615, DOI:10.1002/ARCH.21615.
- J. P. Charles, T. Iwema, V. C. Epa, K. Takaki, J. Rynes and M. Jindra, Ligand-binding properties of a juvenile hormone receptor, Methoprene-tolerant, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 21128–21133, DOI:10.1073/PNAS.1116123109.
- Y. Cui, Z.-L. Liu, C.-C. Li, X.-M. Wei, Y.-J. Lin, L. You, Z.-D. Zhu, H.-M. Deng, Q.-L. Feng, Y.-P. Huang, H. Xiang, Y. Cui, Z.-L. Liu, C.-C. Li, X.-M. Wei, Y.-J. Lin, L. You, Z.-D. Zhu, H.-M. Deng, Q.-L. Feng, Y.-P. Huang and H. Xiang, Role of juvenile hormone receptor Methoprene-tolerant 1 in silkworm larval brain development and domestication, Zool. Res., 2021, 42(5), 637–649, DOI:10.24272/J.ISSN.2095-8137.2021.126.
- J. L. Wang, T. T. Saha, Y. Zhang, C. Zhang and A. S. Raikhel, Juvenile hormone and its receptor methoprene-tolerant promote ribosomal biogenesis and vitellogenesis in the Aedes aegypti mosquito, J. Biol. Chem., 2017, 292, 10306–10315, DOI:10.1074/jbc.M116.761387.
- J. E. Casida and K. A. Durkin, Neuroactive insecticides: Targets, selectivity, resistance, and secondary effects, Annu. Rev. Entomol., 2013, 58, 99–117, DOI:10.1146/ANNUREV-ENTO-120811-153645.
- A. I. Hashem, S. M. El-Kousy, A. El-Torgoman and G. M. Salama, Conversion of Some AM-Butenolides and S-Oxo-2-pyrrolines into Thiono Derivatives, Indian J. Chem., 1986, 1985, 876, DOI:10.1002/CHIN.198603113.
- A. P. Lill, C. B. Rödl, D. Steinhilber, H. Stark and B. Hofmann, Development and evaluation of ST-1829 based on 5-benzylidene-2-phenylthiazolones as promising agent for anti-leukotriene therapy, Eur. J. Med. Chem., 2015, 89, 503–523, DOI:10.1016/j.ejmech.2014.10.054.
- S. K. Ramadan, H. S. M. Abd-Rabboh, A. A. Abdel Hafez and W. S. I. Abou-Elmagd, Some pyrimidohexahydroquinoline candidates: synthesis, DFT, cytotoxic activity evaluation, molecular docking, and in silico studies, RSC Adv., 2024, 14, 16584–16599, 10.1039/D4RA02271H.
- A. Husain, A. Ahmad, A. Bhandari and V. Ram, Synthesis and antitubercular activity of pyridazinone derivatives, J. Chil. Chem. Soc., 2011, 56, 778–780, DOI:10.4067/S0717-97072011000300013.
- M. Kamal, M. H. Hekal, F. S. M. A. El-Azm, E. M. Hosni, Y. M. Ali, A. Yahya Abdullah Alzahrani and E. H. Rafat, Synthesis and in silico studies of new thiophene-isoquinolinone hybrids as potential larvicides against Culex pipiens, Sci. Rep., 2025, 15, 1–24, DOI:10.1038/S41598-025-13063-7.
- M. S. Khalil, M. F. Ismail, M. Kamal, E. M. Hosni, G. A. Elsayed and A. M. A. Hassan, Synthesis and insecticidal assessment of nitrogenous heterocycles derived from 2-pyridone derivative against Culex pipiens L. Larvae, J. Mol. Struct., 2025, 1322, 140405, DOI:10.1016/J.MOLSTRUC.2024.140405.
- M. K. F. El-Sayed, M. M. Elshahawi, F. S. M. Abu El-Azm, E. M. Hosni, M. Kamal and Y. M. Ali, Synthesis, molecular modelling and evaluation of larvicidal efficacy of annulated Benzo[h]chromenes against Culex pipiens L. Larvae, Sci. Rep., 2024, 14(1), 1–19, DOI:10.1038/s41598-024-68035-0.
- D. S. A. Haneen, A. A. Abdalha, M. M. Alkhatib, M. Kamal, A. S. A. Youssef, W. S. I. Abou-Elmagd and S. S. Samir, Synthesis, comprehensive in silico studies, and cytotoxicity evaluation of novel quinazolinone derivatives as potential anticancer agents, Sci. Rep., 2025, 15, 1–25, DOI:10.1038/S41598-025-08062-7.
- A. Haikal, M. Kamal, E. M. Hosni and Y. Amen, Evaluation of hesperidin as a potential larvicide against Culex pipiens with computational prediction of its mode of action via molecular docking, Sci. Rep., 2025, 15(1), 1–16, DOI:10.1038/s41598-025-85760-2.
- World Health Organization, Guidelines for laboratory and field testing of mosquito larvicides, 2005, WHO/CDS/WHOPES/GCDPP/2005.13.
- M. F. Ismail, A. A. El-Sayed, E. M. Hosni and A. I. Hassaballah, Synthesis and evaluation of larvicidal efficacy against C. pipiens of some new heterocyclic compounds emanated from 2-cyano-N’-(2-(2,4-dichlorophenoxy)acetyl)acetohydrazide, Chem. Biodivers., 2024, 21, e202301560, DOI:10.1002/CBDV.202301560.
- W. S. Abbott, A Method of Computing the Effectiveness of an Insecticide, J. Econ. Entomol., 1925, 18, 265–267, DOI:10.1093/JEE/18.2.265A.
- D. J. Finney, Probit Analysis, Cambridge University Press, Cambridge, 3rd edn, 1971, DOI:10.1002/JPS.2600600940.
- Y.-P. Sun, Toxicity Index-An Improved Method of Comparing the Relative Toxicity of Insecticides1, J. Econ. Entomol., 1950, 43, 45–53, DOI:10.1093/JEE/43.1.45.
- A. I. Hassaballah, A. K. El-ziaty, M. M. Gado, H. A. E. Sayed, M. Kamal and R. S. Ali, Design synthesis, characterization, molecular docking and antimicrobial evaluation of novel heterocycles with acrylonitrile and anthracene moieties, Sci. Rep., 2025, 15(1), 1–18, DOI:10.1038/s41598-025-03272-5.
| This journal is © The Royal Society of Chemistry 2026 |