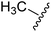Open Access Article
Open Access ArticleRational design and synthesis of non-competitive transcription inhibitors targeting a conserved RNA polymerase-σA interface
Mukesh Tandia,
Nilima Hatib,
Mukul Korea,
Debashree Behera b,
Twinkal Patelb,
Eebac,
Nisheeth Agarwalc,
Balasubramanian Gopal
b,
Twinkal Patelb,
Eebac,
Nisheeth Agarwalc,
Balasubramanian Gopal *bd and
Sandeep Sundriyal
*bd and
Sandeep Sundriyal *a
*a
aDepartment of Pharmacy, Birla Institute of Technology and Science Pilani, Pilani Campus, Rajasthan 333031, India. E-mail: sandeep.sundriyal@pilani.bits-pilani.ac.in
bMolecular Biophysics Unit, Indian Institute of Science, Bangalore, 560012, India. E-mail: bgopal@iisc.ac.in
cBRIC-Translational Health Science and Technology Institute, NCR Biotech Science Cluster, 3rd Mile Stone, Gurugram-Faridabad Expressway, Faridabad 121001, Haryana, India
dInstitute of Bioinformatics and Applied Biotechnology, Bangalore 560100, India
First published on 16th March 2026
Abstract
RNA polymerase (RNAP) inhibitors that block RNA chain elongation or disrupt DNA unwinding have long served as frontline therapeutic agents. However, a major limitation of competitive inhibitors like rifampicin is the rapid emergence of resistance due to point mutations at the RNAP active site. Transcription initiation, the first step in mRNA synthesis, depends on the interaction between RNAP and a sigma (σ) factor. We targeted the conserved region of the RNAP-σA interaction interface to inhibit transcription initiation. Mycobacterium tuberculosis σA was used as a template for inhibitor design due to the extensive experimental structural data on Mycobacterial RNAP in the apo and holo forms, with a promoter DNA fragment, as well as complexes with rifampicin and Nα-aroyl-N-aryl-phenylalaninamide inhibitors. Sequence comparison and subsequent mutational analysis validated Asp319, Glu322, and Gln323 in M. tuberculosis σA as critical residues for interaction with the RNAP β′ subunit. The interface encompassing this polypeptide segment served as a target for inhibitor design. Structure-based virtual screening (SBVS) of a Ugi reaction derived library (URDL) led to the identification of hydroxamate 2a that could bind to M. tuberculosis σA (KD 2.8 µM). Systematic exploration of SAR via 22 analogues led to non-hydroxamate 2u as a superior binder (KD 0.9 µM). Both 2a and 2u also displayed almost complete inhibition of transcription at 1 mM. In a whole-cell bactericidal assay, 2u exhibited clear bactericidal activity against Staphylococcus aureus (MIC 50 µM) and M. tuberculosis (MIC 250 µM). Overall, this study showcases the rational design of non-competitive inhibitors of M. tuberculosis transcription through a systematic application of in silico screening and multicomponent reaction chemistry.
1. Introduction
The transcription mechanism is essential for bacterial survival. The multi-subunit RNA polymerase enzyme driving this mechanism is widely conserved across bacteria. The structure of bacterial RNAPs is widely conserved despite differences in amino acid sequences across RNAP homologues. Transcription initiation, the first step of transcription, involves the recruitment of the RNAP enzyme to the promoter DNA. This step, which primarily serves as the checkpoint regulating gene expression, can vary across bacterial species.1 For example, in the case of M. tuberculosis, when compared to the well-characterized Escherichia coli RNAP, the open promoter RNAP-σA (holoenzyme) complex is inherently unstable and requires accessory proteins like the RNAP- binding protein A (RbpA) and Caspase recruitment domain (CarD) for transcript formation.2 Several other protein factors, like WhiB, modulate RNAP activity under specific environmental stress conditions. Indeed, M. tuberculosis RNAP has been better explored for development of inhibitors than RNAP from other bacterial pathogens.3 These insights have been compiled in a recent review that also provides a summary of all the antimycobacterial compounds developed against the RNAP enzyme, specifically targeting the transcription initiation step.4 To date, peptides and small molecules have been designed to target the base clamp region in the β subunit, the switch region and bridge helix in the β′ subunit, and the RNAP active site. However, a major drawback of rifampicin and other competitive modulators against transcription or other essential cellular processes is the emergence of drug-resistant mutations.5,6 It is worth noting in this context that the RNAP-σ factor interface, located away from the active site, has not been extensively explored for inhibitor design despite significant structural conservation across bacterial homologues.Targeting protein–protein interfaces is an emerging avenue for the design of non-competitive ligands in a variety of disease contexts despite several challenges.7–9 The interaction interface between the RNAP and a σ factor is well conserved across different bacteria and its disruption may result in transcription inhibition.10 Ma et al. utilized a homology model of the Bacillus subtilis RNAP β′–σ factor interface to develop a pharmacophore model to obstruct RNAP-σ factor binding. The pharmacophore model was used to screen an in-house library of peptidomimetic compounds to obtain inhibitors of RNAP β′–σ factor interactions. One of the bisindole derivatives, GKL003, was found to bind RNAP β′ and inhibit the initiation of complex formation. The compound was also found to inhibit transcription and bacterial growth in vitro, thus validating the RNAP β′–σ factor interface as a druggable target.11 Based on the new structural information, the authors further modified this pharmacophore model and discovered compound C5 from a commercial library that could inhibit transcription initiation in vitro and showed antibacterial activity.12 Recently, the same group developed triaryl derivatives binding to the RNAP β′ clamp helix region-σ interface and successfully demonstrated their activity against Streptococcus pneumoniae.13 These results demonstrate the validity of computational design for developing small molecule inhibitors of bacterial RNAP β′–σ factor interactions and subsequent disruption of transcription. Here we describe an approach towards structure-guided protein–protein interface inhibitors focussing on the rigid stretch of σ70 that is occluded upon binding the RNAP β′ subunit. We hypothesize that these inhibitors would obstruct recruitment of the RNAP apo-enzyme to the promoter DNA itself, thereby preventing transcription initiation.
The Pribnow box interacting domain (σ2) of σ factors belonging to the σ70 family share significant commonality in the RNAP interaction interface. We utilized the M. tuberculosis β′ and σ2 interface (in particular with the primary, essential σ factor σA) for the ligand strategy thus building upon the extensive structural information on the Mycobacterial RNAP complexes with multiple experimentally determined structural models.14 A structure-based high throughput-virtual screening (HTVS) strategy was employed using the Ugi reaction-derived library (URDL) with high synthetic tractability that was reported by our laboratory earlier.15 The synthesis of hit molecules and SAR studies resulted in the identification of an experimentally validated scaffold that could be further developed for the design of non-competitive RNAP inhibitors.
2. Results and discussion
2.1 Identification and experimental validation of critical σA residues in RNAP interactions
Across multiple structural complexes, interactions between residues in the coiled-coil region of the β′ domain and domain 2 of a σ factor were noted (Fig. 1A and B). For example, in M. tuberculosis σ2A, a cluster of four residues (Asp319, Glu322, Gln323 and Leu326) interact with β′ residues (Arg350, Arg353, Leu357, Leu360, Asn369 and Glu370) (Fig. 1A). From this interface, conserved interacting residues among M. tuberculosis σ factors were identified by performing a structure-based sequence alignment using all M. tuberculosis σ factor sequences (Fig. 1B and C). To evaluate the significance of the target site of σA binding to the RNAP, residues that were considered crucial were evaluated by mutational analysis. The mutants of σA, including D319K, Q322K, E323K, and KKK (a triple mutant where all three residues- D319, Q322, and E323 were replaced by a positively charged residue, K) were examined. An in vitro transcription assay was performed with these mutants using the M. tuberculosis rRNA AP3 promoter. We note that the D319K mutant exhibited a relatively minor effect on transcription activity, whereas the Q322K and E323K mutants showed no significant alteration in transcription. However, the triple mutant (KKK), encompassing all three mutations, demonstrated a notable and significant impact on transcriptional activity. These observations revealed that for M. tuberculosis σA mediated transcription, the interaction between σA residues Asp319, Glu322, and Gln323 with the β′ domain is crucial. Thus, we hypothesized that small molecules binding to this region of M. tuberculosis σA could prevent RNAP-σA interaction leading to the inhibition of M. tuberculosis transcription.2.2 Structure-based virtual screening (SBVS) against the identified σA site
Based on mutational analysis we selected a region around the residues Asp319, Glu322, and Gln323 of σA as the target site for the SBVS (PDB ID 5UH5, Fig. 2A). Earlier we reported a manually curated library of synthetically tractable molecules derived from the Ugi reaction (URDL) for virtual screening.15,16 The URDL molecules occupy property space of protein–protein inhibitors and provide an opportunity for the systematic exploration of multicomponent reactions in medicinal chemistry.17,18 These molecules are close to protein–protein inhibitors in chemical space and thus suitable for the given target. Moreover, most of the URDL molecules are drug-like and can be obtained in 1–2 synthetic steps under mild conditions.15 Thus, URDL is a suitable library for quick hit validation and SAR studies in the given scenario. The 5773 unique ligands in URDL were screened against M. tuberculosis σA structure (PDB ID 5UH5) employing the HTVS mode of the Schrodinger Suite to perform in silico screening (see SI). The hits were identified based on the docking scores and interactions with the conserved residues (Asp319, Glu322, and Gln323) found important for fostering interactions with RNAP. The hits were subsequently subjected to further analysis using the MMGBSA scoring.This in silico screening resulted in structurally close hits among which compound 2a was found to be most promising (SI, Fig. S1) with a docking score of −4.01 kcal mol−1 and a molecular mechanics with generalized born and surface area (MMGBSA) score of −64.28 kcal mol−1 (Table 1). While the hydroxamate group of 2a displayed hydrogen bonding and non-classical hydrogen bond interactions with Gln322, Glu323, and Gln362 (Fig. 2C and D), the phenyl ring and tert-butyl groups of 2a remain solvent-exposed (Fig. 2C). The compound 2a was originally designed as a selective inhibitor of human histone deacetylase-6 (HDAC-6). However, it displayed weak inhibitory activity against HDAC-6 and low cytotoxicity.19 Compound 2a fulfilled the criteria to be included in URDL owing to its convenient two step synthesis through Ugi reaction. For the experimental validation, we synthesized 2a (over 95% purity by HPLC) and evaluated it for binding by Micro Scale Thermophoresis (MST). Pleasingly, 2a displayed a strong binding with σA with a dissociation constant (KD) of 2.8 ± 0.8 µM (Fig. 3), thus validating the computational design. These results encouraged us to carry out a SAR study for this series of compounds.
2.3 Chemistry
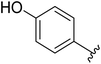
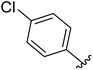
One of the reasons for using URDL was that the basic scaffold of these molecules could be assembled in a single-pot multicomponent reaction. Moreover, a series of analogues can be quickly generated by simply using a different set of building blocks.15–18 Accordingly, for the SAR study we planned to explore different regions P1–P4 of 2a scaffold arising from the four building blocks of the Ugi reaction (Fig. 4). | ||
| Fig. 4 SAR plan around 2a. Ugi MCR allows the exploration of regions P1–P4 by employing diverse aldehydes, acid, isonitrile, and amine components. | ||
Initially, we followed the synthetic procedure reported in the literature report to obtain the intermediate 1a. Thus, a mixture of 4-aminomethyl benzoic acid methyl ester, formaldehyde, benzoic acid, and tert-butyl isocyanide was stirred at 15–25 °C, providing 1a with satisfactory yield (Scheme 1).19 The same method was used to obtain Ugi adducts 1b–1r from other aldehydes, acids, and isocyanide components. The reported sodium methoxide-mediated hydroxyaminolysis, however, yielded unsatisfactory results with isolated yields up to 30% for the hydroxamates. Nonetheless, the usage of KOH base at low temperature worked well for the conversion of ester group to the desired hydroxamates (2a–2r) in good to satisfactory yields (Scheme 1).20 Since the modeling studies implied an important role of hydroxamate function in forming H-bonds (Fig. 2D), we also synthesized analogues with alternative moieties at region P4. Thus, analogues 2s–2v were obtained from 1a using the appropriate nucleophiles, while acid derivative 2v was obtained by base-mediated hydrolysis of the ester 1a.
The purity (≥95%) of all compounds was confirmed using reverse-phase HPLC. The 1D NMR spectra of most of the compounds revealed the presence of two sets of NMR signals due to the cis–trans amide bond rotamers (Fig. S3).19,21 It was further confirmed by performing NMR spectroscopy of 2a at a higher temperature (60 °C) which led to the merging peaks due to the rapid interconversion of two isomers. (SI, Fig. S2)
2.4. Modeling, binding studies and SAR analysis
The binding affinity of all ligand derivatives for M. tuberculosis σA was evaluated using MST, as shown in Tables 1–4. Overall, we did not find significant correlation between experimental binding and the in silico docking score. There could be several reasons for this observation. First, pose-scoring algorithms often fail to fully account for the entropic penalties and desolvation effects associated with binding events occurring in cellular environment. More importantly, these compounds feature a tertiary amide moiety, which exists as a mixture of cis and trans rotamers. Because the ground-state energy difference (ΔG) between these rotamers is negligible in tertiary amides, both forms are sampled during ligand preparation in the Schrödinger suite and subsequently evaluated during the docking process.19 In a biological assay, the observed affinity is a macroscopic result of the equilibrium between these two species, which may interconvert at different rates depending on the specific analogue and assay conditions. However, docking scores reflect most favourable binding for a single rotamer (whether cis or trans). Furthermore, the hydrophobic nature of the substituents resulted in poor aqueous solubility for several compounds. Nevertheless, we noted a few SAR patterns that we discuss below.Initially, we screened analogues 2b–2d by varying aldehyde components at the P1 region. While analogue 2b with a 4-hydroxyphenyl ring (KD 237.3 µM), did not show good binding, compound 2c (KD 0.124 µM) and 2d (KD 0.699 µM, Table 1) displayed significant affinity as compared to the 2a. All three analogues displayed a different orientation than 2a (Fig. 5A–C). The analogue 2c featuring a pyridine ring at the P1 position displayed a distinct binding pose from 2a and suggested strong binding with σA. However, the key interactions with Glu323 and Gln322 were preserved (Fig. 5B). However, we did not synthesize more analogues in this series, given the liability associated with the inflated molecular weight and higher lipophilicity due to the replacement of –H (in 2a) with other aromatic substituents.
We subsequently probed the P2 region by incorporating different acid component in the Ugi reaction. In this series, compounds 2e, 2g, 2m, and 2n displayed better binding and fostered all key interactions like 2a (Fig. 6A–D). Compound 2g with a para-nitrophenyl substitution has KD (7.70 ± 4.48 µM) comparable to 2a but showed a very distinct binding pose than the latter. According to the modelling study, 2g missed a few crucial interactions with Glu323 and Gln362 (Fig. 6B). The analogue 2m with a smaller pyrazole at P2 showed a lower KD value (12.5 ± 66.2 µM) and displayed H-bond interactions comparable to 2a. In addition, the pyrazole ring of 2m fostered an additional H-bond with Gln322 (Fig. 6C). The compound 2n displayed binding (KD 3.5 ± 5.4 µM) comparable to 2a but a different pose resulting in fewer interactions with the conserved residues (Fig. 6D). Other molecules in the P2 series (2h, 2i, and 2l) showed very poor binding (Table 2) with σA. The investigation of the binding poses revealed that compounds 2h and 2i with 2-chloropyridine, and 2-chlorophenyl respectively, exhibited very different binding poses lacking crucial interactions with Gln362 (Fig. 6G and E). Compound 2l with thiazolidine showed comparable binding poses and retained interactions like 2a (Fig. 6F). In general, except for 2a and 2g, the six-membered aromatic rings at P2 displayed poor binding affinity. In contrast, compounds with smaller substituents, such as a methyl group (2e, KD 6.03 ± 3.81 µM) or furyl (2n, KD 3.5 ± 5.4 µM), seem to retain strong binding.
 | ||
| Fig. 6 Comparison of the binding poses 2a with (A) 2e, (B) 2g, (C) 2m, (D) 2n, (E) 2i, (F) 2l, and (G) 2h. | ||
Next, we investigated the P3 region by varying the isonitrile component of the reaction. Replacing the tert-butyl group of 2a with the benzyl 2p, cyclohexyl 2q, and naphthyl 2r rings was detrimental to the binding affinity (Table 3). Interestingly, the binding poses of all three analogues (2p, 2q and 2r) was similar to 2a (Fig. 7). The compounds 2p and 2q retained the key interactions with all three conserved amino acids while 2r displayed interactions with only two amino acids (Glu323 and Gln322, Fig. 7). Due to the limited availability of the commercially available isonitriles, we could not evaluate further diversity at this position.
Finally, we investigated the role of hydroxamate moiety (P4) in target engagement as suggested by the docking pose of 2a (Fig. 2D). Indeed, the replacement of hydroxamate with hydrazide (2s) or methoxyacetamide (2t) significantly weakened binding (Table 4) underscoring the role of free –OH of hydroxamate. Similarly, replacement of hydroxamate with a carboxylic acid (2v) or an ester groups (1a) also markedly reduced binding. Interestingly, converting hydroxamate into an amide in 2u resulted in a 3-fold improvement in binding (KD 0.9 µM) compared to 2a. Notably, 2a and 2u adopt highly similar biding poses, with hydroxamate and amide groups nearly superimposable and both fostering equivalent interactions with Glu323 (Fig. 8A). In contrast, derivatives 2s, 2t, 2v, and 1a bind in a distinct orientation that lack key H-bond interactions with conserved residues. Collectively, these data indicate that the presence of an appropriately oriented H-bond donor capable of engaging Glu322 and Gln362 is a critical requirement for strong binding to σA, a feature retained only in 2a and 2u within this subset of compounds.
2.5 In vitro transcription and RNAP-σA holoenzyme complex assembly
Next, we evaluated all compounds for their ability to inhibit transcription. The in vitro transcription assay was performed using freshly purified RNAP (comprising the α, β, β′ and ω subunits) along with the associated transcription factors σA, RbpA, and CarD to drive transcription from a DNA template with the M. tuberculosis rRNA AP3 promoter element. Multiple protein preparations were evaluated to obtain an enzyme sample that was free from the E. coli σ70 contaminant. Only batches of freshly purified enzyme that that did not show any transcript in the absence of σA in the in vitro transcription assays were subsequently used for the assays to evaluate effect of ligand binding. Synthesized RNA transcripts were quantified using the fluorescent RiboGreen® reagent, which selectively binds RNA. A decrease in fluorescence intensity relative to the control indicated inhibition of transcription by the test compounds. A critical step in this context involves DNAse treatment to the reaction mixture to ensure that there is no contamination due to traces of the promoter DNA template. As expected, no transcript was detected in the absence of rNTPs, confirming that the assay only reported on the mRNA formed due to RNAP activity (Fig. 9A and B). The well characterized anti-mycobacterial therapeutic agent, rifampicin (maintained at 1 mM), served as a positive control and was seen to completely abolish transcript formation. It is important to reiterate in this context that rifampicin is a competitive inhibitor of RNAP as opposed to the compounds evaluated in this study that are protein–protein interaction inhibitors.As expected, compound 2a displayed dose-dependent inhibition of transcription indicating with almost 50% inhibition at 500 µM and complete inhibition at 1 mM concentration (Fig. 9A). Evaluation of other analogues in the same assay highlighted 2u as another potent inhibitor of transcript formation (1 mM concentration, Fig. 2B) correlating well with its strong binding affinitiy (Table 4). In contrast, compounds such as 2b, which showed weak binding, and others like 2c, 2d, 2l, 2p and 2q, despite having relatively better binding profiles, did not produce a notable reduction in transcription. Interestingly, derivatives with substitutions at positions P2 and P3, notably 2m and 2r, exhibited moderate transcription inhibition of up to 50% (Fig. 9B).
To determine if the observed transcriptional inhibition is due to the direct disruption of the RNAP-σA interaction, we employed MST to quantify the binding affinity (KD) of the holoenzyme complex. Under baseline conditions (5% DMSO), σA bound to RNAP with an apparent KD of 109 ± 50 nM (Fig. S3A and Table S1). Pre-incubation with the negative control 2p (Table 3) resulted in a comparable binding affinity (KD of 143 ± 22 nM, Fig. S3B and Table S1). In contrast, the introduction of compound 2u led to a marked decrease in binding strength, with the apparent KD increasing to 1.78 ± 2.7 µM. This represents an approximately 16-fold reduction in binding affinity relative to the untreated control. These results together with inhibition of transcription in the in vitro assay (Fig. 9) demonstrate that 2a and 2u specifically and substantially weakens the RNAP-σA interaction.
2.6 Antibacterial studies
Finally, we evaluated the most promising analogues 2a and 2u together with other analogues in whole-cell growth inhibition assays against M. tuberculosis (H37Rv). Initial screening showed no inhibition of mycobacterial growth at concentrations up to 50 µM (data not shown). Given that 2a inhibits in vitro transcription significantly only at 700 µM (Fig. 9A), higher concentrations were tested. A 5% DMSO control (equivalent to that in 500 µM compound) inhibited M. tuberculosis growth, so results obtained with 250 µM compound concentrations were considered. While most of the compounds remain ineffective, a notable complete inhibition of growth was observed for analogue 2u at 250 µM, as assessed by visualizing the pellet formation after 10 days of incubation at 37 °C in a U-bottom 96-well plate (Fig. 10A). This data likely reflects stronger transcription inhibition by 2u at this concentration or permeation liabilities of the polar hydroxamate moiety, which impedes passive diffusion of 2a across membranes.22–24 Hydroxamates are also known to suffer hydrolytic cleavage in cellular environments, unlike the more robust amide functionality in 2u.25To investigate whether the observed lack of antibacterial potency was a result of chemical instability, particularly regarding the hydroxamate moiety in 2a, stability studies were conducted in phosphate-buffered saline (PBS). Both 2a and its amide control 2u, were monitored via HPLC over 48 hours. Neither compound exhibited significant degradation or changes in chromatographic profiles over 48 hours (Fig. S4), indicating sufficient chemical stability, at least under the assay conditions.
Given the pronounced structural similarity between RNAP homologues (despite sequence differences) and the conserved nature of the RNAP-σA binding interface across bacterial species, we also evaluated these analogues against the Gram-positive pathogen Staphylococcus aureus. Freshly grown S. aureus cultures were incubated with individual compounds at 100 µM concentration in a U-bottom 96-well plate, and growth was assessed by visualizing pellet formation after 48 h incubation at 37 °C. Among the 18 analogues tested, only 2u completely inhibited growth (Fig. 10B), consistent with its strong σA binding affinity, and potent inhibition of in vitro transcription as well as M. tuberculosis H37Rv growth. Subsequently, a minimum inhibitory concentration (MIC) was determined by incubating S. aureus with serial dilutions of 2u, which revealed that 2u inhibits growth of S. aureus at an MIC of 50 µM (Fig. 10C). Most importantly, plating an aliquot of S. aureus treated with 50 or 100 µM 2u and untreated culture on LB agar plate clearly established the bactericidal effect of the inhibitor (Fig. 10D).
Notably, compound 2a, the hydroxamate derivative of 2u, once again proved inactive in the whole cell assay, despite comparable transcription inhibition. This could be due to ineffective binding of 2a with S. aureus σA indicated by the docking studies discussed below (Section 2.7). Nevertheless, 2u's modest antibacterial potency against both pathogens validates the rational design strategy targeting the conserved RNAP-σA interface for the discovery of novel transcription inhibitors.
2.7 Molecular docking of 2u with σA of S. aureus
To predict binding modes, we performed molecular docking of both, 2a and 2u, with the σA of S. aureus (PDB 8X6F) obtained using cryo-electron microscopy.26 Superimposition of S. aureus and M. tuberculosis σA structures enabled the identification of binding site for docking into S. aureus σA (Fig. S5A and S5B).The docking pose of 2u in S. aureus σA closely resembled its pose in M. tuberculosis σA (Fig. S5C vs. Fig. 8A). In both cases, the amide –NH2 fostered a H-bond with Gln202 (S. aureus numbering; Gln362 in M. tuberculosis). Compound 2u also formed an additional H-bond with the conserved residue Asp159 (Asp319 in M. tuberculosis). In contrast, compound 2a adopted different orientations in the σA factors of both organisms (Fig. S5D vs. Fig. 8A). It lacked the H-bond with Gln202 in S. aureus σA unlike its interaction with the corresponding Gln362 in M. tuberculosis σA. However, 2a retained a H-bond with Asp159 in S. aureus σA.
Compound 2a also exhibited a slightly lower docking score with S. aureus σA compared to 2u (3.03 vs. 3.30 kcal mol−1). Together, this molecular modeling study supports the stronger interaction of 2u with S. aureus σA than 2a, consistent with its superior antibacterial activity against S. aureus.
Overall, the data suggest that both 2a and 2u remain stable under assay conditions and their poor cellular activity may be due to the reduced passive permeation. This is especially true for M. tuberculosis where the notoriously impermeable, lipid-rich mycomembrane pose a well-documented challenge in drug discovery.27 In contrast to 2a, 2u seems to maintain similar binding pose in σA factors of both S. aureus and M. tuberculosis.
3. Conclusions
In summary, this is the first report describing a rational approach for the design of non-competitive inhibitors of M. tuberculosis transcription by targeting RNAP-σA interface. Using sequence alignment and mutational analysis we established the σA residues important for interacting with RNAP. We systematically explored a focussed MCR-based library (URDL), occupying protein–protein inhibitor chemical space, for the identification of σA binders. SBVS approach combined with an efficient synthetic protocol expediated hit identification yielding a hydroxamate compound 2a (KD 2.8 µM). The MCR modularity allowed for rapid SAR exploration across P1–P4 regions, generating 22 analogues in 1–2 steps. Ligand binding and in vitro transcription assays revealed hydroxamate (2a) and amide (2u) as potent RNAP-σA disruptors, with nearly identical docking poses with significant interactions with the conserved σA residues. Unfortunately, the whole-cell activity was not significant against M. tuberculosis, likely due to restricted penetration across the mycomembrane. Nonetheless, amide 2u exhibited bactericidal activity against S. aureus (MIC 50 µM), displaying broader applicability of this scaffold for antimicrobial development.While inhibitors of RNAP-σA interface of other bacterial species are reported, a direct comparison of these inhibitors with our series is not possible given the inherent differences in the transcription machineries of M. tuberculosis and other bacterial species. Structurally, our lead inhibitors 2a and 2u are distinct than the other reported inhibitors. Nonetheless, these compounds show similarity to reported inhibitors in terms of higher micromolar concentrations required to inhibit the growth of S. aureus in culture.11,12
Overall, this series is a viable starting point for the development of broad-spectrum bacterial RNAP inhibitors. The facile, two-step synthesis demonstrated here highlights the utility of MCRs in accelerating the drug discovery cycle. Future efforts will prioritize amide-based analogues to improve cellular permeability and antibacterial potency. Additionally, combination studies evaluating potential synergy with competitive RNAP inhibitors, such as rifampicin, remains to be examined.
Conflicts of interest
Authors declare no financial or non-financial interests that are directly or indirectly related to the work submitted for publication.Abbreviations
| BPB | Bromophenol blue |
| CarD | Caspase recruitment domains |
| DMSO | Dimethyl sulfoxide |
| DTT | Dithiothreitol |
| EDTA | Ethylenediaminetetraacetic acid |
| KD | Dissociation constant |
| HEPES | 2-[4-(2-Hydroxyethyl)piperazin-1-yl]ethanesulfonic acid |
| HPLC | High-performance liquid chromatography |
| HRMS | High-resolution mass spectrometry |
| HTVS | High throughput virtual screening |
| MIC | minimum inhibitory concentration |
| MMGBSA | Molecular mechanics with generalised born and surface area |
| MST | Microscale thermophoresis |
| NMR | Nuclear magnetic resonance |
| PAGE | Polyacrylamide gel electrophoresis |
| RbpA | RNA polymerase-binding protein A |
| RNAP | RNA polymerase |
| rNTPs | A ribonucleotide triphosphate |
| SAR | Structure–activity relationship |
| SLS | Sodium lauryl sulphate |
| TLC | Thin layer chromatography |
| URDL | Ugi reaction-derived library |
| UT | Untreated |
Data availability
The data supporting this article has been included in the main article and as part of supplementary information (SI). Supplementary information: Fig. S1–S5, Tables S1 and S2, detailed computational and experimental methodologies, and all spectral data (NMR, HRMS, etc.) of final compounds. See DOI: https://doi.org/10.1039/d6ra00442c.Acknowledgements
This work is supported by the funding from the Indian Council of Medical Research (ICMR), New Delhi, India (Grant No. 67/3/2020-DDI/BMS). Funding support from the THSTI core to Ms. Eeba and NA is acknowledged. MT and MK acknowledge the fellowship provided by BITS Pilani. We acknowledge DST, New Delhi, India, for sponsoring the HRMS and LC-MS facility at BITS Pilani, Pilani, under the FIST program. We thank Dr. Bhabatosh Das at the Translational Health Science and Technology Institute, Faridabad for providing pathogenic S. aureus strain.References
- W. Lin, S. Mandal, D. Degen, Y. Liu, Y. W. Ebright, S. Li, Y. Feng, Y. Zhang, S. Mandal, Y. Jiang, S. Liu, M. Gigliotti, M. Talaue, N. Connell, K. Das, E. Arnold and R. H. Ebright, Mol. Cell, 2017, 66, 169–179 Search PubMed.
- D. X. Zhu, A. L. Garner, E. A. Galburt and C. L. Stallings, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 13573–13581 CrossRef CAS.
- J. C. Perez and E. A. Groisman, Cell, 2009, 138, 233–244 CrossRef CAS PubMed.
- F. Stephanie, U. Sumo and F. Tambunan, Life, 2022, 12, 1–21 Search PubMed.
- N. Woodford and M. J. Ellington, Clin. Microbiol. Infect., 2006, 13, 5–18 Search PubMed.
- B. P. Goldstein, J. Antibiot., 2014, 67, 625–630 CrossRef CAS PubMed.
- E. André, L. Bastide, P. Villain-guillot, J. Latouche, J. Rouby and J. Leonetti, Assay Drug Dev. Technol., 2004, 2, 629–635 Search PubMed.
- D. E. Scott, A. R. Bayly, C. Abell and J. Skidmore, Nat. Rev. Drug Discovery, 2016, 15, 533–550 CrossRef CAS PubMed.
- C. M. Labbé, M. A. Kuenemann, B. Zarzycka, G. Vriend, G. A. F. Nicolaes, D. Lagorce, M. A. Miteva, B. O. Villoutreix and O. Sperandio, Nucleic Acids Res., 2016, 44, D542–D547 Search PubMed.
- C. Ma, X. Yang and P. J. Lewis, Microbiol. Mol. Biol. Rev., 2016, 80, 139–160 Search PubMed.
- C. Ma, X. Yang, H. Kandemir, M. Mielczarek, E. B. Johnston, R. Griffith, N. Kumar and P. J. Lewis, ACS Chem. Biol., 2013, 8, 1972–1980 CrossRef CAS PubMed.
- C. Ma, X. Yang and P. J. Lewis, ACS Infect. Dis., 2016, 2, 39–46 CrossRef CAS PubMed.
- J. Ye, A. J. Chu, R. Harper, S. T. Chan, T. L. Shek, Y. Zhang, M. Ip, M. Sambir, I. Artsimovitch, Z. Zuo, X. Yang and C. Ma, J. Med. Chem., 2020, 63, 7695–7720 CrossRef CAS PubMed.
- A. Feklístov, B. D. Sharon, S. A. Darst and C. A. Gross, Annu. Rev. Microbiol., 2014, 68, 357–376 Search PubMed.
- M. Tandi, N. Tripathi, A. Gaur, B. Gopal and S. Sundriyal, Mol. Divers., 2024, 28, 37–50 CrossRef CAS PubMed.
- I. Ugi and C. Steinbrückner, Angew. Chem., 1960, 72, 267–268 Search PubMed.
- M. Tandi, V. Sharma, B. Gopal and S. Sundriyal, RSC Adv., 2025, 15, 1447–1489 Search PubMed.
- M. Tandi and S. Sundriyal, J. Indian Chem. Soc., 2021, 98, 100106 Search PubMed.
- D. Diedrich, A. Hamacher, C. G. W. Gertzen, L. A. Alves Avelar, G. J. Reiss, T. Kurz, H. Gohlke, M. U. Kassack and F. K. Hansen, Chem. Commun., 2016, 52, 3219–3222 Search PubMed.
- G. Joshi, S. Kalra, U. Prasad, P. Sharma, P. Kumar, S. Amrutkar, A. J. Ansari, S. Kumar, A. Sharon and S. Sharma, Bioorg. Chem., 2020, 94, 103409 CrossRef CAS PubMed.
- B. Yoo and K. Kirshenbaum, Curr. Opin. Chem. Biol., 2008, 12, 714–721 CrossRef CAS PubMed.
- S. Shen and A. P. Kozikowski, ChemMedChem, 2016, 11, 15–21 CrossRef CAS PubMed.
- S. Kesharwani, N. Eeba, M. Tandi, N. Agarwal and S. Sundriyal, RSC Adv., 2024, 14, 27530–27554 RSC.
- S. Kesharwani and S. Sundriyal, Eur. J. Med. Chem., 2021, 213, 113055 CrossRef CAS PubMed.
- P. Hermant, D. Bosc, C. Piveteau, R. Gealageas, B. Lam, C. Ronco, M. Roignant, H. Tolojanahary, L. Jean, P. Y. Renard, M. Lemdani, M. Bourotte, A. Herledan, C. Bedart, A. Biela, F. Leroux, B. Deprez and R. Deprez-Poulain, J. Med. Chem., 2017, 60, 9067–9089 CrossRef CAS PubMed.
- L. Yuan, Q. Liu, L. Xu, B. Wu and Y. Feng, Nat. Commun., 2024, 15, 4850 CrossRef CAS PubMed.
- S. M. Batt, D. E. Minnikin and G. S. Besra, Biochem. J., 2020, 447, 1983–2006 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |