Open Access Article
Open Access ArticleA Brønsted-based deep eutectic solvent as a green catalyst for the sustainable one-pot synthesis of 1,2,4,5-tetrasubstituted imidazole derivatives: in vitro cytotoxicity and in silico binding studies on HepG2 cells
Tuan-Anh Ngoc Tran ab,
Linh Dieu Nguyen
ab,
Linh Dieu Nguyen ab,
Khoi Nam Nguyen
ab,
Khoi Nam Nguyen ab,
Phu Hoang Dang
ab,
Phu Hoang Dang ab and
Hai Truong Nguyen
ab and
Hai Truong Nguyen *ab
*ab
aDepartment of Organic Chemistry, Faculty of Chemistry, University of Science, Ho Chi Minh City 700000, Vietnam. E-mail: ngthai@hcmus.edu.vn
bVietnam National University, Ho Chi Minh City 700000, Vietnam
First published on 18th March 2026
Abstract
Liver cancer remains a major global health challenge, necessitating the development of new therapeutic agents. In this study, a series of 1,2,4,5-tetrasubstituted imidazole derivatives were synthesized via a green, one-pot four-component reaction using the recyclable deep eutectic solvent [Bet]2[PTSA] under solvent-free conditions, affording high yields. Among them, compounds 16d, 16j, and 16n exhibited notable cytotoxicity against HepG2 cells, with 16n showing the strongest activity (IC50 = 32.54 µM), outperforming camptothecin. Molecular docking and dynamics simulations revealed strong and stable binding of 16n to GLUT1, involving key residues such as Trp388, Asn288, and Trp412. Structure-activity relationship (SAR) analysis indicated that phenolic hydroxyl and para-methoxy substituents enhance anticancer activity. These results identify compound 16n as a promising lead for targeted liver cancer therapy.
1 Introduction
Hepatocellular carcinoma (HCC) represents a significant global health challenge, ranking as the most common primary liver malignancy and a leading cause of cancer-related mortality.1 Despite advancements in diagnostic imaging and treatments such as surgical resection and liver transplantation, the prognosis for advanced HCC remains poor. Systemic therapies, including chemotherapy and targeted agents like Sorafenib, are frequently employed; however, their efficacy is often compromised by high off-target toxicity, low tumor permeability, and the rapid development of multidrug resistance.2–5 Consequently, there is an imperative necessity to persist in the pursuit of novel pharmacological therapies that are economically viable, more efficient, and less detrimental. HepG2 is a widely used human liver cancer cell line in biomedical research, derived from a 15 year-old Caucasian male with liver carcinoma. Known for their epithelial morphology and capacity to secrete liver-specific proteins like albumin and alpha-fetoprotein, HepG2 cells are a valuable model for studying liver-specific functions, drug metabolism, and liver-related diseases such as hepatocellular carcinoma. Targeting the glucose transporter GLUT1 has emerged as a promising therapeutic strategy for hepatocellular carcinoma, as cancer cells exhibit elevated glucose uptake to sustain rapid proliferation. Inhibition of GLUT1 disrupts tumor energy metabolism, leading to reduced cell growth and enhanced sensitivity to anticancer treatments. Therefore, the GLUT1 pathway represents an alternative and complementary approach for liver cancer therapy.6 The increased activity in cancer cells leads to enhanced glucose transport and consumption following the Warburg effect.7 GLUT1 is a member of the SLC2 family of glucose transporters (GLUTs) encoded by the SLC2A1 gene. The overall structure of GLUT1 consists of 12 transmembrane helices (TM1-TM12) divided into two domains (C and N). TM7 and TM10 are segments that exhibit flexibility, referred to as TM7a/7b and TM10a/10b, respectively.7–9 The substrate transport mechanism follows an alternating access model, where substrate binding on one side of the membrane induces structural changes from the outside to the inside, allowing glucose to be transported into the cell. This is similarly demonstrated in the homolog XylE from E. coli.9 The inward-open IOP conformation of GLUT1 has been shown to have the best structural interaction with ligands compared to the outward-occluded OCC, outward-open OOP, and partially inward-occluded PIO conformation previously studied. This study utilized protein data from the Protein Data Bank with the protein code (PDB: 5EQG), classified as a transport protein from Homo sapiens, co-crystallized with the ligand 5RE.10 Mutagenesis studies at Trp388 suggest that this residue plays a crucial role in the interconversion between IOP and OOP during glucose absorption, demonstrating its influence on glucose transport activity. Additionally, other amino acids in the C domain, such as Trp412, Thr137, Gln282, Phe291, Phe379, and Glu380, serve as significant anchor points for inhibitors that prevent the transition from IOP to OOP. Besides Trp388 in TM10, which specifically binds in the IOP conformation, Trp412 and Phe379 in TM11 and TM10, respectively, are studied for mutations at these positions, which can severely impair glucose transport and activity. Based on data collected from various reports, typical interactions such as π–π stacking and hydrogen bonding with Trp388, Trp412, and Phe379 are prioritized in silico screening.11,12Heterocyclic blocks present a promising avenue for discovering novel molecules for cancer therapy. In recent years, numerous small molecules featuring heterocyclic rings have been suggested for the advancement of new candidates exhibiting anti-cancer activity in drug discovery initiatives.13 Heterocyclic compounds, commonly referred to as heterocycles, are organic chemical entities characterized by a cyclic structure containing one or more heteroatoms.14 Heterocycles encompass various agricultural, pharmaceutical, and veterinary applications.15 Currently, heterocycles are utilized in the manufacturing process of a wide variety of organic chemical compounds.16 Heterocycles are common in natural products and have found widespread application in drug discovery and medicinal chemistry. This category includes physiologically natural products, active complexes, and chemicals that are utilized extensively in medical chemistry.17,18 Among these heterocyclic compounds that include nitrogen: imidazoles, indoles, pyrroles, pyrazoles, and pyrrolidines have become increasingly important in various study fields, including chemical synthesis and medicine.17 For instance, benzimidazole compounds exhibit significant anticancer activity and have been studied for potential therapeutic applications, including antiviral, anticancer, or antidiabetic.19,20 Chrysin benzimidazole derivatives were synthesized by Zhe Wanga, which showed that some novel compounds can impede tumor proliferation.21 A potential heterocyclic framework is imidazole, which is a widely used structural motif in pharmaceuticals, natural products, endogenous chemicals, and polymers.22 Among them, the imidazole compound is of particular interest due to its ability to interact with many different biological targets through hydrogen bonding and π–π stacking interactions. Imidazole is a five-membered heterocyclic ring containing three carbon and two nitrogen atoms at non-adjacent positions. It is a versatile framework with diverse applications in both chemistry and bioactivity. Imidazole and its derivatives are also present in pharmaceutical medicines, including clonidine (antihypertensive),23 metronidazole (antibacterial, antifungal),24,25 dacarbazine (anticancer),26 celecoxib (anti-inflammatory),27 clotrimazole (antifungal),28 and dipyrone (antipyretic)29 (Fig. 1). The imidazole ring is a common structural motif in various drugs, such as antifungal medications, antihistamines, and antiviral agents, highlighting its significance in pharmaceutical research and development.30 Especially, biological activities of imidazole scaffolds have been reported to exhibit a broad range of anticancer, including antitumor activity and anticancer agents.
 | ||
| Fig. 1 Drugs featuring an imidazole scaffold.31 | ||
Substituents were introduced onto the imidazole scaffold to modulate biological activity and to investigate structure-activity relationships (SAR). Variations in the electronic and steric properties of these substituents contribute to enhanced interactions with the molecular target, improved physicochemical properties, and increased biological applicability of the imidazole derivatives. For the synthesis of imidazole, especially 1,2,4,5-tetrasubstituted imidazoles, deep eutectic solvents can be applied as a catalyst in the process of synthesis. The incorporation of deep eutectic solvents (DESs) as eco-friendly catalytic media has recently revolutionized multicomponent reactions (MCRs), enhancing both their efficiency and environmental sustainability. MCRs constitute a highly versatile synthetic strategy that enables the one-pot assembly of complex and structurally diverse molecules from multiple reactants. Owing to their inherent advantages-such as high atom economy, mild reaction conditions, and minimal waste formation-MCRs have become a cornerstone of green organic synthesis.32 DESs are defined as a novel and sustainable class of media, representing eutectic mixtures of two or more components, which are typically a hydrogen bond acceptor (HBA), such as choline chloride, and a hydrogen bond donor (HBD), like urea or glycerol. The essence of their formation lies in the vigorous, directional hydrogen-bonding network established between the components, which dramatically decreases the freezing point of the mixture, resulting in a liquid at or near room temperature, often with similar physicochemical properties to ionic liquids but at a fraction of the cost and complexity. The fundamental role of DESs is to act as green, non-volatile, non-flammable, and often biodegradable alternatives to conventional hazardous organic solvents. The applications of DESs are extensive, spanning extraction, electrochemistry, and material science, yet their utility is most profound in organic synthesis catalysis. Here, DESs serve as versatile, recyclable reaction media that significantly enhance sustainability. They are not merely inert solvents; their unique, highly polar, and structured environment can stabilize catalysts and activate reactants through hydrogen bonding, often leading to improved yields and enhanced product selectivity.33 Furthermore, DESs containing acidic HBDs can act as Brønsted acid catalysts themselves, eliminating the need for external catalysts in reactions like multicomponent condensations.34 This combination of simple formation, an environmentally superior role, and exceptional chemical tunability makes DESs a critical, cutting-edge platform for developing highly efficient, economic, and environmentally responsible catalytic processes in modern synthetic chemistry.
The combination of betaine (as the hydrogen bond acceptor) and p-toluenesulfonic acid (PTSA, as the hydrogen bond donor) is an excellent choice for creating a DES for organic synthesis due to its strong potential for Brønsted acid catalysis and its favorable physical properties. Betaine, a zwitterion, provides a high concentration of the quaternary ammonium cation, offering a highly polar, charged environment that can effectively stabilize transition states and polar intermediates.35 Crucially, PTSA is a strong organic acid, ensuring that the resulting DES is highly acidic. This high acidity is essential for catalyzing a wide range of organic reactions, such as esterification, Biginelli reactions, or other processes that require a powerful Brønsted acid catalyst.36,37 The strong, favorable hydrogen-bonding interaction between the carboxylate group of betaine and the sulfonic acid proton of PTSA significantly depresses the melting point of the mixture, forming a liquid DES at or near room temperature. This low melting point, combined with the DES's non-volatility and potential for reuse, makes it a greener, more efficient alternative to conventional volatile organic solvents and corrosive liquid acids.
To address the limitations of current HCC therapies, this study focuses on the synthesis of a novel series of 1,2,4,5-tetrasubstituted imidazoles synthesized from benzil, benzaldehyde, aniline and ammonium acetate with the support of a deep eutectic solvent [Bet]2[PTSA] via a one-pot multicomponent reaction. The use of DES provided a mild, efficient, and environmentally friendly route for imidazole formation, offering advantages such as high yield, simple work-up, and catalyst recyclability. The primary objective of this work is to synthesize these novel imidazole derivatives and evaluate their cytotoxic potential against the HepG2 human liver carcinoma cell line. The anticancer potential of the synthesized imidazole derivatives was evaluated through in vitro cytotoxic assays, identifying several compounds with remarkable inhibitory activity against HepG2 cells. Furthermore, molecular docking studies were conducted to predict their interactions with relevant target proteins, and selected candidates were further validated in vitro using three different liver cancer cell lines.
2 Result and discussion
2.1. Characterization of [Bet]2[PTSA]-deep eutectic solvent
Fig. 2A illustrates the structure of the DES catalyst. Fig. 2B serves as spectroscopic proof, presenting the Fourier Transform Infrared (FT-IR) profiles of the synthesized deep eutectic solvent (DES), [Bet]2[PTSA], alongside its precursor components: betaine (Bet) and p-toluenesulfonic acid (PTSA). The vibrational peak, which is clearly detected at 1132 cm−1 and 1723 cm−1 indicates the existence of C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O in the carboxylic group and C–N–C bonds belonging to betaine's structure.38 The typical vibration observed at 662 cm−1 belongs to the C–S bond existing in the structure of PTSA.39 In the FT-IR spectrum of pure PTSA, the peak at 835 cm−1 is attributed to the S–OH vibration, while the antisymmetric and symmetric stretching vibrations of the S
O in the carboxylic group and C–N–C bonds belonging to betaine's structure.38 The typical vibration observed at 662 cm−1 belongs to the C–S bond existing in the structure of PTSA.39 In the FT-IR spectrum of pure PTSA, the peak at 835 cm−1 is attributed to the S–OH vibration, while the antisymmetric and symmetric stretching vibrations of the S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group appear at 1173 cm−1 and 1118 cm−1, respectively. In contrast, the S–OH vibration disappears in the DES spectrum, signaling the involvement of this group in hydrogen bonding between the two components. Furthermore, the S
O group appear at 1173 cm−1 and 1118 cm−1, respectively. In contrast, the S–OH vibration disappears in the DES spectrum, signaling the involvement of this group in hydrogen bonding between the two components. Furthermore, the S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibrations shift to 1148 cm−1 and 1130 cm−1, providing further evidence of the intermolecular interactions within the eutectic system.40 The successful formation of the DES is emphatically demonstrated by the final product's spectrum. It meticulously retains the principal functional group characteristic peaks from both betaine and PTSA, providing unambiguous evidence of the successful stoichiometric combination required to form the [Bet]2[PTSA] compounds.
O stretching vibrations shift to 1148 cm−1 and 1130 cm−1, providing further evidence of the intermolecular interactions within the eutectic system.40 The successful formation of the DES is emphatically demonstrated by the final product's spectrum. It meticulously retains the principal functional group characteristic peaks from both betaine and PTSA, providing unambiguous evidence of the successful stoichiometric combination required to form the [Bet]2[PTSA] compounds.
 | ||
| Fig. 2 (A) Structure of [Bet]2[PTSA]; (B) The FT-IR spectra and (C) TGA curves of betaine, PTSA, and [Bet]2[PTSA]. | ||
Thermal gravimetric analysis (TGA) showed the thermal stability of the [Bet]2[PTSA] catalyst (Fig. 2C). The obtained curve shows the first weight loss below 100 °C, which may be ascribed to the removal of residual solvents, gases, and water. Upon comparison with the individual components, the thermal decomposition started at 250 °C for PTSA and 300 °C for betaine. This finding provides clear evidence that the combination of betaine and PTSA at 100 °C leads to the formation of a eutectic mixture facilitated by hydrogen-bond interactions, which in turn significantly lowers the thermal decomposition temperature of the resulting product. These results indicate that the TGA curve of the catalyst confirms its thermal stability up to 150 °C, suggesting that [Bet]2[PTSA] can be efficiently applied for a range of organic transformations across different temperature conditions.41
As can be seen in Fig. 3, the formation of [Bet]2[PTSA] from betaine and p-toluenesulfonic acid (PTSA) is clearly evidenced by the 1H-NMR spectral changes. In the spectrum of pure betaine, the characteristic singlet of the –N(CH3)3 and methylene (–CH2–) groups appear at 3.32 and 4.23 ppm, respectively, while PTSA exhibits aromatic proton signals at 7.64–7.29 ppm and a methyl (-CH3) signal near 2.32 ppm. Upon formation of [Bet]2[PTSA], noticeable upfield shifts of the –N(CH3)3 and methylene protons of betaine are observed, together with slight changes in the aromatic proton region of PTSA. These shifts indicate strong ionic interactions and proton transfer between the sulfonic acid group (–SO3H) of PTSA and the carboxylate functionality of betaine. The disappearance (or strong attenuation) of the acidic proton signal of PTSA further confirms proton transfer, leading to the formation of the protic ionic salt [Bet]2[PTSA].42,43
2.2. Synthesis of 1,2,4,5-tetrasubstituted imidazoles
The compounds used in the experimental section for the synthesis of imidazole include benzaldehyde, aniline, benzil, and ammonium acetate. The study began by testing three experimental techniques, namely microwave irradiation, sonication, and vertical heating assisted by magnetic stirring, to identify an optimal procedure. The conventional heating method proved to be the most effective, yielding the highest product conversion, approximately 75% (Entries 1–2 and 5, Table 1). Conversely, both microwave and sonication approaches led to lower reaction performance, which is likely attributed to poor temperature uniformity and inadequate surface contact between reactants, respectively. Consequently, the standard heating magnetic stirrer was selected for all subsequent optimization and experimental procedures.| Entry | Time (h) | Temperature (°C) | %mol of [Bet]2[PTSA] | Yieldb (%) |
|---|---|---|---|---|
| a Reaction conditions: Benzil (1 mmol), benzaldehyde (1 mmol), aniline (1 mmol), ammonium acetate (1 mmol), solvent-free.b Isolated yield by crystallization in acetone (10–15 mL).c Microwave method.d Sonication method.e The heating and stirring method. | ||||
| 1 | 3 | 100 | 3.5 | Tracec |
| 2 | 3 | 100 | 3.5 | 16d |
| 3 | 1 | 100 | 3.5 | 32 |
| 4 | 2 | 100 | 3.5 | 55 |
| 5 | 3 | 100 | 3.5 | 75e |
| 6 | 4 | 100 | 3.5 | 66 |
| 7 | 5 | 100 | 3.5 | 61 |
| 8 | 6 | 100 | 3.5 | 56 |
| 9 | 3 | RT | 3.5 | 11 |
| 10 | 3 | 60 | 3.5 | 30 |
| 11 | 3 | 80 | 3.5 | 64 |
| 12 | 3 | 120 | 3.5 | 64 |
| 13 | 3 | 100 | 0 | 41 |
| 14 | 3 | 100 | 0.7 | 55 |
| 15 | 3 | 100 | 16 | 55 |
| 16 | 3 | 100 | 21 | 44 |
| 17 | 3 | 100 | 35 | 37 |
The model reaction was systematically examined under different experimental parameters to determine the optimal pathway. Through this optimization, substituted 1,2,4,5-tetrasubstituted imidazole derivatives were efficiently synthesized within a shorter reaction time, affording moderate to significant yields as summarized in Table 1. A series of reactions using representative substrates was then performed under varied conditions, including reaction time (Entries 3–8, Table 1), temperature (Entries 5 and 9–12, Table 1), and the amount of deep eutectic solvent (DES) serving as the catalyst (Entries 5 and 13–17, Table 1).
Furthermore, the reactions were tested in both protic and aprotic solvents, as well as in the presence of different catalytic systems. Among the examined catalysts, [Bet]2[PTSA] at 3.5 mol% displayed superior catalytic activity and selectivity toward the target imidazole derivatives. The highest yield (75%) was obtained under solvent-free conditions. In contrast, a marked decline in catalytic performance was observed when the reactions were carried out in solvents (Entries 1–8, Table 2). At elevated temperatures (100 °C), the deep eutectic structure was destabilized, leading to partial decomposition of its organic constituents in the solvent medium. This behavior can be explained by the reduction in the mass stability of [Bet]2[PTSA] at both ambient and reflux temperatures. Furthermore, various components involved in the synthesis of [Bet]2[PTSA] were examined to validate the efficiency of the catalyst produced (Entries 8–14, Table 2). Additionally, with the contribution of Brønsted acid in the efficiency of 1,2,4,5-tetrasubstituted imidazole reaction, various types of Brønsted DES were applied as catalysts for comparison purposes. As demonstrated by the data in Table 2, utilizing a DES catalyst, composed of both betaine and PTSA, resulted in a superior yield of the main product compared to employing either betaine or PTSA in isolation for the catalytic reaction. Moreover, substituting the [Bet]2[PTSA] catalyst with structurally analogous catalysts, but possessing a modified number of Brønsted acid sites, led to a discernible and substantial reduction in the main product yield. The investigations prove the crucial role of Brønsted acid sites in the synthesis reaction.
| Entry | Catalysts | %mol of [Bet]2[PTSA] | Solvents | Yieldb (%) |
|---|---|---|---|---|
| a Reaction conditions: Benzil (1 mmol), benzaldehyde (1 mmol), aniline (1 mmol), ammonium acetate (1 mmol), catalysts (3.5% mol), solvents (3 mL) at 100 °C, 3 h.b Isolated yield by crystallization in acetone (10–15 mL).c Reactions involving volatile solvents were carried out using a reflux system. | ||||
| 1 | None | 0 | Ethanolc | 12 |
| 2 | None | 0 | Methanolc | 6 |
| 3 | None | 0 | Tetrahydrofuranc | Trace |
| 4 | None | 0 | n-Hexanec | 27 |
| 5 | None | 0 | 1,4-Dioxanec | 19 |
| 6 | None | 0 | Acetonec | 3 |
| 7 | None | 0 | Ethyl acetatec | 8 |
| 8 | [Bet]2[PTSA] | 3.5 | None | 75 |
| 9 | Betaine | 3.5 | None | 45 |
| 10 | PTSA | 3.5 | None | 40 |
| 11 | [HACl][Oxalic] | 3.5 | None | 52 |
| 12 | [HACl][Glycine] | 3.5 | None | 30 |
| 13 | [Bet][Maleic acid] | 3.5 | None | 34 |
| 14 | [ChCl][PTSA] | 3.5 | None | 38 |
| 15 | None | 0 | None | <10 |
The catalyst was recovered after finishing the experiment, and its structure was characterized by FT-IR analysis (Fig. 4). It shows that the typical signals for peaks in the [Bet]2[PTSA] structure have changed significantly. Herein, the absorption vibration of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bond at 1723 cm−1 considerably fell, proving a distinct change in the structure of DES compared to the original. The changes of intensities and wavenumbers of S
O bond at 1723 cm−1 considerably fell, proving a distinct change in the structure of DES compared to the original. The changes of intensities and wavenumbers of S![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O bonds observed in the FT-IR spectrum of the recovered catalyst were attributed to partial disruption or rearrangement of the hydrogen-bonding network within the DES structure under prolonged heating conditions. Besides, the disappearance of some signals in recover spectrum in comparison of raw components can be explained by the change in the chemical structure of DES due to the high temperature. As a result, the efficiency of the main product was reduced due to changes in the catalyst structure.
O bonds observed in the FT-IR spectrum of the recovered catalyst were attributed to partial disruption or rearrangement of the hydrogen-bonding network within the DES structure under prolonged heating conditions. Besides, the disappearance of some signals in recover spectrum in comparison of raw components can be explained by the change in the chemical structure of DES due to the high temperature. As a result, the efficiency of the main product was reduced due to changes in the catalyst structure.
Under the optimized conditions described above, a scale-up experiment was performed through the condensation of benzil (1 mmol), benzaldehyde (1 mmol), aniline (1 mmol), and ammonium acetate (1 mmol) using [Bet]2[PTSA] (3.5 mol%) as a catalyst at 100 °C for 3 h, affording compound 16a (Fig. 5). The reaction was subsequently conducted at different scales (1, 5, 10, and 20 mmol), and the isolated yields of product 16a ranged between 70% and 75%.
Based on the optimal conditions which were obtained through the above investigation, 1,2,4,5-tetra substituted imidazole derivatives were synthesized under the support of [Bet]2[PTSA] 3.5% mol without solvent (Scheme 1). At first, with aniline as a co-reactant, based on the main product yield formed from the synthesis reactions of 1,2,4,5-tetrasubstituted imidazole derivatives, it can be concluded that aromatic aldehyde derivatives bearing electron-donating groups generally provide a higher main product yield compared to the yield from reactions involving electron-withdrawing groups (16a–h). Specifically, for reactions using aniline as a co-reactant and varying the electron-withdrawing substituents on benzaldehyde, the main product yield obtained is in the range of 45–57%. Conversely, an electron-donating group such as 4-Me gives a relatively high yield of 65%. Although the nitro group is a strong electron-withdrawing substituent, the nitro group at the para-position derivative exhibited a higher yield than expected, at about 60%. This can be explained by the fact that the para-position of NO2 minimizes steric hindrance, allowing efficient molecular interactions during condensation and cyclization. Therefore, despite its electron-withdrawing nature, the enhanced carbonyl activation and steric advantages at the p-position outweigh the deactivating effect on the reaction, leading to a comparatively higher yield for the 4-NO2-substituted product. The effect of the hydroxyl group in the ortho and meta-positions was also investigated. The results showed that the hydroxyl group at the ortho-position yielded a lower product compared to the meta-position. This can be explained by the fact that 2-hydroxybenzaldehyde forms an intramolecular hydrogen bond, which subsequently reduces the activity (reactivity) of the aldehyde group, thus leading to a lower yield of the main product.
When aniline was altered by 4-aminophenol (16i–o), the yield of derivatives significantly enhanced. Typically, the reaction of four components, including benzil, 4-methoxylbenzaldehyde, 4-aminophenol and ammonium acetate with 3.5% mol of [Bet]2[PTSA] at the same temperature as the above investigations illustrated the highest yield, approximately 86%. This can be attributed to the strong electron-donating resonance effect of the phenolic –OH group, which enhances the nucleophilicity of the amino functionality and thereby promotes the initial condensation with the aldehyde to form the imine intermediate more efficiently. However, the notably low yield (10%) obtained when employing 4-fluorobenzaldehyde as a substrate can be attributed to the strong electron-withdrawing inductive effect (−I) of fluorine, which significantly decreases the electron density of the carbonyl group and thereby diminishes its electrophilicity toward nucleophilic attack by aniline. Although fluorine possesses a resonance-donating (+R) ability, this effect is extremely weak and insufficient to counterbalance its dominant −I effect, resulting in a markedly reduced reactivity of the aldehyde compared to other para-substituents. Additionally, the high electronegativity of fluorine may lead to strong dipole–dipole and hydrogen-bonding interactions with the catalytic DES components, restricting effective molecular collisions between the reactants and further suppressing product formation. Consequently, 4-fluoro substitution exerts a more detrimental influence on the reaction efficiency than other halogens or electron-donating groups, leading to a significantly lower product yield.
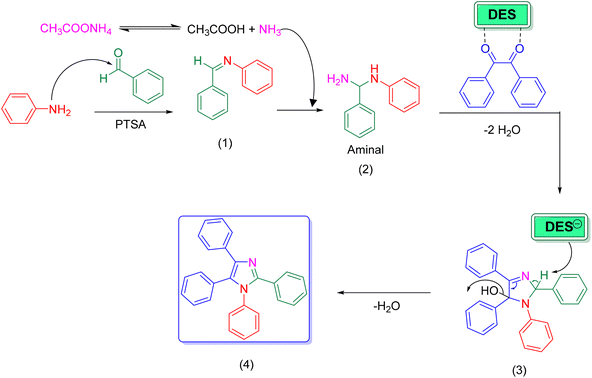
The proposed mechanism for the synthesis of 1,2,4,5-tetrasubstituted imidazoles using [Bet]2[PTSA] as a catalyst is presented in Scheme 2. The reaction pathway is initiated by the condensation between aniline and benzaldehyde. In this step, the PTSA component of the DES acts as a Brønsted acid catalyst, activating the carbonyl group of benzaldehyde through protonation, which facilitates nucleophilic attack by aniline to form the imine intermediate (1).44,45 At the same time, ammonium acetate undergoes thermal decomposition, establishing an equilibrium that releases ammonia. This ammonia then performs a nucleophilic attack on the imine (1) to yield the aminal intermediate (2). Subsequently, intermediate (2) undergoes a cascade transformation involving condensation with benzil, intramolecular cyclization, and the elimination of two water molecules. Throughout this process, the DES continues to play a crucial role by activating carbonyl groups through hydrogen bonding interactions. The final product (4) is 1,2,4,5-tetraphenyl-1H-imidazole (4).
2.3. Assessment of green metrics
To evaluate the environmental sustainability and efficiency of the developed synthetic protocol, several green chemistry metrics were calculated, including the E-factor, atom economy, atom efficiency, process mass intensity, carbon efficiency, reaction mass efficiency, and eco-score. The green chemistry assessment of the 1,2,4,5-tetrasubstituted imidazole synthesis using DES [Bet]2[PTSA] as a catalyst reveals that this reaction proceeds with notable environmental efficiency. The process achieved a very low E-factor (0.27), indicating that only a small amount of by-products or waste was formed (water and acetic acid). The atom economy (78.58%) and atom efficiency (67.57%) values suggest that the majority of the reactant atoms were successfully incorporated into the desired product. Meanwhile, the process mass intensity (1.48) shows that the overall material consumption was minimal, and the carbon efficiency (80.3%) highlights effective carbon utilization throughout the synthesis. The reaction mass efficiency (67.58%) also reflects a favorable balance between yield and resource use. Taken together, these parameters correspond to an eco-score of 69%, confirming that the synthetic route catalyzed by DES [Bet]2[PTSA] is both resource-efficient and environmentally sustainable, offering a promising green alternative for multicomponent organic transformations.462.4. Design strategy
Numerous studies in Fig. 6 have reported findings regarding the inhibition of cancer using various specific cell lines, especially human hepatocarcinoma cell lines (HepG2), employing various mechanisms and pathways with potential imidazole derivative compounds. Yi-Chen Liu et al. published compound 7, a benzobis (imidazole) derivative, which exhibited inhibition of the STAT3 signaling pathway, had functions in the immune system and influence the differentiation of T cells and the response of immune cells, making it important in inflammation and autoimmunity (IC50 = 4.30 ± 1.72 µM).47 Wen Gu et al. synthesized benzimidazole compounds combined with various heterocyclic moieties to test their activity on SMMC-7721 and HepG2 cell lines. Compounds 8 (IC50 = 0.36 ± 0.13 µM) and 9 (IC50 = 0.12 ± 0.03 µM) arrested cells in the G2/M phase, leading to accumulation and a decrease in concentrations in subsequent phases, thereby affecting cell division and hindering tumor growth.48 Furthermore, other studies have shown that compound 10 displayed superior anti-HepG2 activity compared to 5-FU and cisplatin, whereas compound 11 also exhibited strong cytotoxic effects against HepG2 cells.49,50 New strategies have emerged from the synthesis methods and hybridization approaches surrounding the imidazole framework, capturing the attention of researchers. By leveraging the unique structural characteristics and biological interactions of imidazole with various inhibitory and regulatory pathways, these studies have the potential to positively impact the disease, paving the way for a diverse array of lead compounds in the treatment of hepatocellular carcinoma.2.5. Docking study
The interaction of compounds 16a–o, serving as ligands, with the GLUT1 protein was investigated using data from the Protein Data Bank (PDB ID 5EQG). The compound 1,2,4,5-tetrasubstituted imidazole was substituted into the active site co-crystallized 5RE in complex with the protein. The docking results presented in Table 3 indicated that the synthesized compounds adhered to the expected protocol, interacting with characteristic amino acids as reported in studies related to GLUT1. Table 4 presents detailed information on the potential compounds with favorable binding energies, such as 16d, 16e, 16f, 16j, and 16n.| Compound | Binding energy | Interacting residues |
|---|---|---|
| 16a | −7.66 | — |
| 16b | −7.62 | — |
| 16c | −7.57 | Asn411 |
| 16d | −8.23 | Asn288 |
| 16e | −8.07 | — |
| 16f | −8.16 | — |
| 16g | −7.89 | — |
| 16h | −7.90 | — |
| 16i | −7.84 | — |
| 16j | −8.33 | — |
| 16k | −7.81 | — |
| 16l | −7.69 | Glu380 |
| 16m | −7.50 | Glu380 |
| 16n | −8.23 | Asn288 |
| 16o | −7.70 | Trp388 |
| Comp | Ligand sites | Receptor sites | Type of interaction | Distance (Å) | Binding energy (kcal mol−1) | Total binding energy (kcal mol−1) |
|---|---|---|---|---|---|---|
| 16d | 6-Ring | Asn288 | Π–H | 4.35 | −0.6 | −8.23 |
| 16e | — | — | — | — | — | −8.07 |
| 16f | — | — | — | — | — | −7.77 |
| 16j | — | — | — | — | — | −8.16 |
| 16n | 6-Ring | Asn288 | Π–H | 4.52 | −0.5 | −8.23 |
The binding affinities of all the compounds were better than those of α-glucose and β-glucose reported in the previous study. The competitive inhibition between sugar molecules and these compounds is crucial for cancer treatment by reducing sugar absorption.9 Numerous studies have identified Trp388 as a critical site that appears frequently; however, we also highlight other amino acids such as Trp412, Thr137, Asn288, Gln282, Phe291, and Phe379. Interactions with these specific anchor points are expected to constrain the structural transition from the inward-open (IOP) to the outward-open (OOP) conformation, enhancing the success of the method. Residues at these positions also contribute to the interaction energy through physical bonds, ionic interactions, and hydrophobic contacts.51
The 2D and 3D views in Fig. 7 for compounds 16d, 16j, and 16n, which demonstrated actual biological activity as confirmed in the following section, highlight the key amino acid residues involved in the interactions, including Trp388, Trp412, Asn288, Glu380, Thr30, and Phe379. However, the affinity interaction investigations with this active site were limited to a static state and did not account for the dynamics of the ligand and protein. In a suitable state, interactions with important residues can be recorded, although this was not associated with the best energy state among the five poses, which would be detailed in Table S3. Additionally, the RMSD (Root Mean Square Deviation) is a standard method that involves comparing the docked pose of a ligand with its cocrystal structure. A pose is considered good if its RMSD is less than 2.0 Å.9 All docking cases of the 15 compounds exhibited RMSD values less than 2 Å, indicating that all poses were well-aligned and reliable. From the docking studies, the scope of in vitro testing was narrowed down. Among the 15 synthesized compounds, biological activity assays were conducted on compounds 16a, 16d, 16e, 16f, 16j, and 16n.
 | ||
| Fig. 7 The 2D and 3D views of the docking model of the interaction of the target compounds with HepG2 (PDB code 5EQG) compounds 16d, 16j, and 16n. | ||
2.6. Anticancer activity
Based on the results from the docking studies, the five compounds mentioned above, along with the compound without the substituent (16a), will be subjected to activity testing. The anticancer potency of 6 compounds synthesized from the four-component condensation reaction on the HepG2 liver cell line was measured by the SRB method. For comparison purposes, Camptothecin, the positive control, manifested cytotoxicity with IC50 values of 50.31 µM on HepG2 cell lines. As shown in Table 5, there are some novel compounds that have been identified to possess considerably stronger cytotoxicity than the positive control in vitro. Herein, 16d, 16j, and 16n exhibited superior cytotoxicity percentages of 59.73%, 57.84%, and 78.19%, respectively, at a concentration of 100 µM. We proceeded further to investigate the IC50 values of these three compounds (Table 6).| Entry | Compounds | R | R’ | I (%)a |
|---|---|---|---|---|
| a I (%) stands for Inhibition percentage.b DMSO is the negative control.c Camptothecin is the positive control. | ||||
| 1 | 16a | 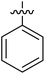 |
H | 26.65 ± 4.85 |
| 2 | 16d |  |
H | 59.73 ± 2.42 |
| 3 | 16e | 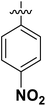 |
H | 8.79 ± 4.80 |
| 4 | 16f | 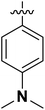 |
H | 10.55 ± 5.72 |
| 5 | 16j | 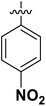 |
OH | 57.84 ± 1.47 |
| 6 | 16n | 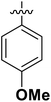 |
OH | 78.19 ± 2.20 |
| 7 | DMSOb | −0.21 ± 1.48 | ||
| 8 | Camptothecinc | 50.31 ± 1.00 | ||
| Compounds | Concentration (µM) | IC50 (µM) | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 10 | 20 | 30 | 40 | 50 | 60 | 75 | 80 | 100 | ||
| 16d | — | 0.66 ± 3.48 | — | 18.05 ± 0.87 | — | 30.26 ± 1.24 | — | 44.72 ± 2.97 | 59.73 ± 2.42 | 85.63 |
| 16j | — | 1.86 ± 4.99 | — | 18.28 ± 4.12 | — | 33.86 ± 0.95 | — | 41.69 ± 2.79 | 57.84 ± 1.47 | 87.96 |
| 16n | 3.77 ± 2.83 | 34.37 ± 1.22 | 51.16 ± 1.04 | 62.43 ± 3.35 | 68.95 ± 3.88 | — | 70.66 ± 2.23 | — | 78.19 ± 2.20 | 32.54 |
The IC50 values for 16d, 16j, and 16n are presented in Table 6 and Fig. 8, highlighting the superior activity of compound 16n compared to 16d and 16j. The docking study demonstrates the binding energy of the ligand components, with three compounds showing a high percentage of cytotoxicity. However, the incomplete consistency observed in the docking study (16j (−8.33) < 16d (−8.23) and 16d (−8.23)) indicates that the simulation results are primarily useful for screening and guiding purposes, as they mainly reflect the predicted binding energy and interaction patterns of the compounds at the active site. Predicting interaction potential using screening software is a crucial step in selecting this promising compound for activity testing. To explain why compound 16n exhibits the lowest IC50 compared to the other two compounds, it is necessary to consider the factors of molecular dynamics and the relationships that influence the parts affecting activity.
 | ||
| Fig. 8 The dose response curves of the cytotoxicity of compounds 16d, 16j, and 16n towards the HepG2 tumor cell line. | ||
2.7. Molecular dynamic
Fig. 9 illustrated the molecular dynamics of the protein-ligand complexes of three compounds (16d, 16j, 16n) over 100 ns. The results indicated varying stability among the components, allowing for conclusions about the active framework of the compounds. Compound 16d showed significant fluctuations, reaching up to 2 Å during the 100 ns, suggesting excessive movement that led to the instability of the protein-ligand complex. This was consistent with its status as the compound with the lowest activity. Hydrogen bond analysis also reveals a very low density of hydrogen bonding with the protein throughout the simulation. | ||
| Fig. 9 The 2D and 3D views of the docking model of the interaction of the target compounds with HepG2 (PDB code 5EQG) compounds 16d, 16j, and 16n. | ||
For compound 16j, which had comparable activity, stability was observed during the first 60 ns in the simulated environment. However, abnormal fluctuations occurred thereafter, indicating that extending the duration of kinetic study may reveal complete turbulence within this complex. The hydrogen bond density of 16j was more consistent than that of 16d, with the ability to form up to two hydrogen bonds. However, a sparse distribution was identified around 80 ns, corresponding to an instability, respectively RMSD axis.
The compound with the best IC50, 16n, exhibited fluctuations around the backbone for the first 10 ns but stabilizes for the remainder of the simulation. Compound 16n also consistently formed two hydrogen bonds, explaining its superior activity compared to 16d and 16j. Additionally, hydrogen bonds with residues for all three compounds were presented in Fig S1–S3, where the two compounds containing hydroxyl groups on the aromatic ring (R′) demonstrated characteristic interactions with residues Trp388, Thr30, Tyr292, Asn415, and Thr137. These structural modifications can significantly influence the activity of the compounds.
2.8. Structure-activity relationship
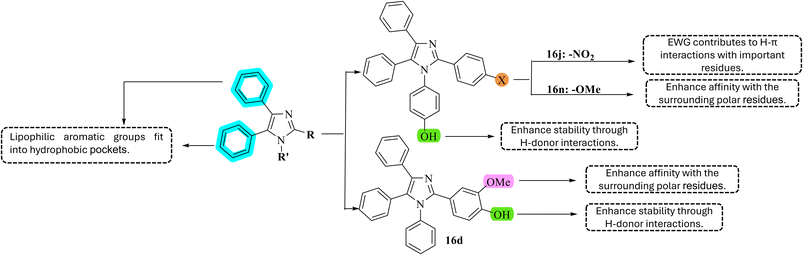
Scheme 3 illustrates the relationship between activity and structure of the three potential compounds. The hydroxyl group enhanced activity, as observed in the docking studies and molecular dynamics. Comparing 16d and 16n, which are isomers of each other, it is evident that having two electron-donating groups on one aromatic ring is not able to exhibit effective activity. In Fig. 6 and S1–S3, it can be seen that the hydroxyl group on the aromatic ring R′ interacted with the residues Asn415, Thr30, and Tyr292, with Thr30 showing the most significant interaction at 47.96% and 58.80% for 16j and 16n, respectively. Additionally, previous studies have indicated that the loss or replacement of the phenolic hydroxyl group would reduce the inhibitory activity against GLUT1, suggesting that introducing additional hydroxyl groups on the aromatic ring (R) is a promising approach.23 The interactions of the methoxy and nitro groups with the residues Thr137 and Trp412, which are both critical amino acids previously studied, reveal significant differences. However, the contrasting electronic effects of the nitro group, which withdraws electrons, and the methoxy group, which donates electrons, lead to a notable discrepancy in the IC50 values between the two compounds: 16n (IC50 = 32.54 µM) compared to 16j (IC50 = 87.96 µM). This difference may be attributed to the electron-withdrawing nature of the nitro group affecting either the aromatic ring or the structure of the imidazole scaffold. The resonance, electron withdrawal, and charge distribution of the nitrogen atom in this group may destabilize it in an electron-rich environment.52,53 Compounds with substituents containing the heteroatom nitrogen, such as 16e (I% = 8.79 ± 4.80) and 16f (I% = 10.55 ± 5.72), exhibited very low anticancer activity. Three compounds, 16e, 16f, and 16j, contain nitrogen-containing substituents, but only 16j exhibited significant activity when a hydroxyl group was introduced. Compared to the unsubstituted compound 16a (I% = 26.65 ± 4.85), it is evident that adding nitrogen-containing substituents (16e, 16f) reduces activity. However, comparing 16e, which contains a nitro group, with 16j, which also has a nitro group but additionally a OH group at R′, the activity increases again (I% = 57.84 ± 1.47, IC50 = 87.96 µM). The compounds were also found to interact with the indole group of Trp388, which could enhance the affinity and selective binding between the ligand and the transmembrane helix 10 (TM10) region.7 As observed in previous studies, Trp388 consistently serves as a key residue and anchoring point. Furthermore, the imidazole core has been shown to be suitable for the discovery and development of potential compounds, as the influence of the nitrogen-containing heterocycle on Trp388 has been demonstrated in other studies.7,54 Free electrons on the high electronegativity heteroatom form a direct bond with hydrogen on Trp388.3 Experimental
3.1. Preparation of [Bet]2[PTSA] catalyst
The mixture of betaine (2 mmol, 0.2322 g), and PTSA (1 mmol, 0.1620 g) were heated at 100 °C for 30 minutes using magnetic stirring until the mixture is homogeneous. Then, the mixture was cooled at room temperature. After that, the viscous liquid was washed several times with diethyl ether (3 × 5 mL) and ethyl acetate (3 × 5 mL) to remove the excess substrates for obtaining [Bet]2[PTSA]. The structure was determined using TGA, FT-IR, 1H and 13C-NMR spectra.3.2. Preparation of 1,2,4,5-tetrasubstituted imidazoles
In a 15 mL-COD tube, the mixture of benzil (1 mmol), aldehydes (1 mmol), anilines (1 mmol), ammonium acetate (1 mmol), and [Bet]2[PTSA] (3.5% mol) was reacted at 100 °C for 3 h and stirring about 400 rpm. The reaction mixture was monitored by thin-layer chromatography (TLC). For reactions using volatile solvents (Table 2), a reflux condenser was attached. Then, 15 mL of acetone was added, and the mixture was stirred for a few minutes at room temperature. Then, the [Bet]2[PTSA] catalyst can be obtained through filter paper. Following filtration to remove the DES, the filtrate was evaporated to dryness to yield the imidazole product as a crystalline solid, which was then recrystallized from acetone (10–15 mL) to afford the pure compound. The structure of products was determined using 1H and 13C-NMR spectra, melting point.3.3. Anti-cancer activity assay
Liver cancer (HepG2) provided by ATCC (USA), which were cultured in E'MEM (HepG2) medium supplemented with L-glutamine (2 mM), HEPES (20 mM), amphotericin B (0.025 µg mL−1), penicillin G (100 UI/mL) streptomycin (100 µg mL−1) 10% (v/v) fetal bovine serum FBS and incubated at 37 °C, 5% CO2.Single cells were seeded in 96-well culture plates at a density of 104 cells per well (for HepG2 cell lines). After 24 hours of culture, the cell population was incubated with the test substance at various concentrations for 48 hours. Then, the total protein from the test cells was fixed with 50% cold trichloroacetic acid (Sigma) solution and stained with 0.2% Sulforhodamine B (Sigma) solution. The results were read by ELISA reader at two wavelengths of 492 nm and 620 nm. The experiments were repeated three times, and the results are presented as mean ± standard deviation.
After obtaining the optical density values at wavelengths 492 nm and 620 nm (denoted as OD492 and OD620):
| Calculate the value OD = OD492 − OD620 | (1) |
| Calculate OD492 (or OD620) = ODtb − ODblank | (2) |
Calculate the percentage of cytotoxicity according to the formula:
• ODtb: OD value of the well containing cells.
• ODblank: OD value of the blank well (no cells).
• ODTN: OD value of the test sample calculated from formulas (1) and (2).
• ODC: OD value of the control sample calculated from formulas (1) and (2).
IC50 was determined using Prism software with a non-linear multi-parameter regression method and R2 > 0.9.
3.4. Docking study
Docking studies were conducted on an 11th Gen Intel Core™ i5-1135G7@2.40 GHz with Windows 11 (version 24H2). The software used was MOE – Molecular Operating Environment (version 2019.0102) by Chemical Computing Group. The target of the study was GLUT1 – Glucose Transporter 1, with the protein code 5EQG from the Protein Data Bank. The 2D structures of the compounds were drawn and constructed using PerkinElmer ChemDraw Professional (version 20.0.0.41), then converted to SMILES format and accessed via the Builder interface for conversion to 3D structures.The process for defining the active site involved using Site Finder to select the position where the ligand was co-crystallized. Once the active site was identified, it was marked as a “Dummy atom”. The docking function in MOE performed virtual docking with the Triangle Matcher placement method. The scoring methods employed included London dG, Induced Fit refinement, and GBVI/WSA dG scoring against dummy atoms surrounding the substrate binding site. Potential docking positions were identified based on the lowest energy-scoring algorithm. Additionally, it was ensured that the docking pocket provided adequate space for compound binding and facilitated reasonable interactions with the protein residues.
3.5. Molecular dynamics simulation
MD simulations were performed to evaluate the stability of the ERα LBDs complexed with chemicals 16d, 16j, and 16n. The simulations employed the CUDA-enabled NAMD 2.14 functionality in MOE 2022.02.55 The chosen compounds were solubilized with water molecules within an 8 Å radius and exposed to periodic boundary conditions. To preserve system neutrality, NaCl was incorporated at a concentration of 0.1 mol L−1 to equilibrate protein charges, employing the particle mesh Ewald method for the rectification of electrostatic interactions. Light bonds were restricted during all simulations. The simulation time interval was established at 2 femtoseconds. The simulation technique comprised an initial 10 ps minimization at 0 K, followed by a progressive heating of the entire system to 300 K over 100 ps. Subsequently, a 100 ps equilibration under NVT circumstances and an additional 200 ps equilibration under NPT conditions were performed to achieve system equilibrium. A 100 ns molecular dynamics simulation under NPT conditions was conducted. The temperature and pressure were maintained at 300 K and 100 kPa, respectively. The root means square deviation (RMSD) and root means square fluctuation (RMSF) values acquired during the manufacturing phase were evaluated to determine the stability of the ligand–receptor complex during the MD simulation.4 Conclusion
This study successfully addresses the dual challenge of developing potent anticancer agents while adhering to sustainable synthesis principles. Unlike conventional methods that rely on hazardous volatile organic solvents, our protocol utilizes the deep eutectic solvent [Bet]2[PTSA] as a catalyst with 69% yield of main synthesized product. In terms of therapeutic potential, the synthesized 1,2,4,5-tetrasubstituted imidazole derivatives demonstrated remarkable efficacy against hepatocellular carcinoma. Notably, compound 16n exhibited superior cytotoxicity (IC50 = 32.54 µM) compared to the standard drug Camptothecin (IC50 = 32.54 µM). This finding fills a critical gap in the search for novel small molecules that can overcome the limitations of current HCC therapies, such as drug resistance. Furthermore, the integration of in silico models provided a plausible molecular mechanism, identifying GLUT1 as a key target and highlighting the role of specific residues in ligand binding. The research utilized the SRB assay method to quantify changes in cell viability, thereby assessing the cytotoxicity of test substances. The compounds that showed promising results in terms of cytotoxicity against the HepG2 liver cancer cell line were 16d, 16j, and 16n. These compounds exhibited cytotoxicity percentages of 59.73%, 57.84%, and 78.19%, respectively which were higher than the positive control. Additionally, the exploration of molecular interactions through docking studies underlines the relevance of specific binding energies in understanding the pharmacological potential of compounds. The findings not only contribute to the body of knowledge regarding potential anticancer agents but also emphasize the significance of targeted therapies in combating various cancers. Additionally, while molecular dynamics simulations suggest stable binding to GLUT1, direct experimental validation of this enzymatic inhibition is required. Future research should focus on in vivo assays to validate the therapeutic window of these compounds and further optimization of the imidazole scaffold to enhance selectivity. Ultimately, this work contributes a foundational framework for the green synthesis of targeted anticancer therapies, paving the way for more sustainable drug discovery programs.Author contributions
Tuan-Anh Ngoc Tran, Linh Dieu Nguyen, Khoi Nam Nguyen: investigation, methodology, resources, formal analysis, validation, data curation, and writing – original draft. Phu Hoang Dang: methodology, resources, formal analysis, validation, data curation, writing – review & editing. Hai Truong Nguyen: methodology, resources, formal analysis, validation, data curation, writing – review & editing, and supervision.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Abbreviation
| 5-FU | Fluorouracil |
| A-549 | Lung cancer |
| AFP | Alpha-fetoprotein |
| Asn411 | Asparagine 411 |
| ATCC | American Type Culture Collection |
| Bcl-2 | Regulation of apoptosis |
| Bet | Betain |
| DESs | Deep eutectic solvents |
| DMSO | Dimethyl sulfoxide |
| ELISA | Enzyme-linked Immunosorbent assay |
| EtOH | Ethanol |
| FBS | Fetal Bovine serum |
| FT-IR | Fourier Transport Infrared |
| GBVI/WSA | Force field-based scoring function |
| Gln161 | Glutamine 161 |
| Gln282 | Glutamine 282 |
| Glu380 | Glutamate 380 |
| GLUT1 | Glucose Transporter 1 |
| GLUTs | GLUT family |
| HBA | Hydrogen bond acceptor |
| HBD | Hydrogen bond donor |
| HCC | Hepatocellular carcinoma |
| HepG2 | Human liver cancer cell line |
| His160 | Histidine 160 |
| IC50 | half maximal inhibitory concentration |
| IOP | The inward-open conformation |
| lle164 | L-Leucine 164 |
| MCRs | multicomponent reactions |
| MOE | Molecular Operating Environment |
| OCC | The outward-occluded conformation |
| OD | Optical density |
| OOP | The outward-open conformation |
| PDB | Protein Data Bank |
| Phe26 | Phenylalanine 26 |
| Phe379 | Phenylalanine 379 |
| PIO | The partially inward-occluded conformation |
| Pro385 | Proline 385 |
| PTSA | p-Toluenesulfonic acid |
| RMSD | Root Mean Square Deviation |
| ROS | Reactive oxygen species |
| RPMI | Roswell Park Memorial Institute |
| SLC2 | Solute carrier family 2 |
| SLC2A1 | Solute carrier family 2-member 1 |
| SRB | Sulforhodamine B |
| TGA | Thermal gravimetric analysis |
| Thr137 | Threonine 137 |
| Thr30 | Threonine 30 |
| TM | Transmembrane |
| TM10a/10b | Transmembrane 10a/10b |
| TM1-TM12 | Transmembrane 1–12 |
| TM7a/7b | Transmembrane 7a/7b |
| Trp388 | Tryptophan 388 |
| Trp412 | Tryptophan 412 |
| XyLE | An important enzyme in the degradation of toluenes, benzoates, and their methyl derivatives |
Data availability
The authors confirm that the data supporting the findings of this study are available within the article. Raw data that support the findings of this study are available from the corresponding author, upon reasonable request.Supplementary information (SI): general procedures, E-factor calculations, full characterization of both the catalyst and the synthesized compounds, copies of 1H, and 13C NMR spectra. See DOI: https://doi.org/10.1039/d6ra00422a.
References
- S. Pilleron, E. Soto-Perez-de-Celis, J. Vignat, J. Ferlay, I. Soerjomataram, F. Bray and D. Sarfati, Int. J. Cancer, 2021, 148, 601–608 CrossRef CAS PubMed.
- U. Anand, A. Dey, A. K. S. Chandel, R. Sanyal, A. Mishra, D. K. Pandey, V. De Falco, A. Upadhyay, R. Kandimalla and A. Chaudhary, Genes Dis., 2023, 10, 1367–1401 CrossRef CAS PubMed.
- K. Bukowski, M. Kciuk and R. Kontek, Int. J. Mol. Sci., 2020, 21, 3233 CrossRef CAS PubMed.
- F. D. Makurvet, Med. Drug Discovery, 2021, 9, 100075 CrossRef CAS.
- J. M. Llovet, R. Montal, D. Sia and R. S. Finn, Nat. Rev. Clin. Oncol., 2018, 15, 599–616 CrossRef PubMed.
- M. H. A. Soliman, I. A. I. Ali, S. S. A. El-Sakka and O. E.-S. Mohamed, J. Mol. Struct., 2022, 1254, 132325 CrossRef CAS.
- Y. Meng, X. Xu, H. Luan, L. Li, W. Dai, Z. Li and J. Bian, Future Med. Chem., 2019, 11, 2333–2352 CrossRef CAS PubMed.
- D. Deng, P. Sun, C. Yan, M. Ke, X. Jiang, L. Xiong, W. Ren, K. Hirata, M. Yamamoto and S. Fan, Nature, 2015, 526, 391–396 CrossRef CAS PubMed.
- S. Almahmoud, W. Jin, L. Geng, J. Wang, X. Wang, J. L. Vennerstrom and H. A. Zhong, Bioorg. Med. Chem., 2020, 28, 115395 CrossRef CAS PubMed.
- K. Kapoor, J. S. Finer-Moore, B. P. Pedersen, L. Caboni, A. Waight, R. C. Hillig, P. Bringmann, I. Heisler, T. Müller and H. Siebeneicher, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 4711–4716 CrossRef CAS PubMed.
- S. Almahmoud, X. Wang, J. L. Vennerstrom and H. A. Zhong, Molecules, 2019, 24, 2159 CrossRef CAS PubMed.
- H. Katagiri, T. Asano, H. Ishihara, J. L. Lin, K. Inukai, M. F. Shanahan, K. Tsukuda, M. Kikuchi, Y. Yazaki and Y. Oka, Biochem. J., 1993, 291, 861–867 CrossRef CAS PubMed.
- S. L. Schreiber, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 6699–6702 CrossRef CAS PubMed.
- P. Beigiazaraghbelagh and A. Poursattar Marjani, Res. Chem. Intermed., 2024, 50, 485–531 CrossRef CAS.
- J. Baranwal, S. Kushwaha, S. Singh and A. Jyoti, Curr. Phys. Chem., 2023, 13, 2–19 CrossRef CAS.
- P. V. Ledade, T. L. Lambat, J. K. Gunjate, P. K. Chopra, A. V. Bhute, M. R. Lanjewar, P. M. Kadu, U. J. Dongre and S. H. Mahmood, Curr. Org. Chem., 2023, 27, 206–222 CrossRef CAS.
- A. Mermer, T. Keles and Y. Sirin, Bioorg. Chem., 2021, 114, 105076 Search PubMed.
- M.-M. Li, X. Chen, Y. Deng and J. Lu, RSC Adv., 2021, 11, 38060–38078 RSC.
- S. R. Brishty, M. J. Hossain, M. U. Khandaker, M. R. I. Faruque, H. Osman and S. A. Rahman, Front. Pharmacol, 2021, 12, 762807 Search PubMed.
- A.-S. M. Abas, F. Z. Mohamed, A. R. Bayomi and E. Gad, Biochem. Lett., 2022, 18, 33–55 Search PubMed.
- Z. Wang, X. Deng, S. Xiong, R. Xiong, J. Liu, L. Zou, X. Lei, X. Cao, Z. Xie and Y. Chen, Nat. Prod. Res., 2018, 32, 2900–2909 Search PubMed.
- N. Saha, P. J. Wanjari, G. Dubey, N. Mahawar and P. V. Bharatam, J. Mol. Struct., 2023, 1272, 134092 CrossRef CAS.
- F. Petitjeans, J.-Y. Martinez, M. D. des Déserts, S. Leroy, L. Quintin and J. Escarment, Crit. Care Med., 2020, 48, e991–e993 CrossRef CAS PubMed.
- U. Kalinowska-Lis, A. Felczak, L. Chęcińska, K. Zawadzka, E. Patyna, K. Lisowska and J. Ochocki, Dalton Trans., 2015, 44, 8178–8189 Search PubMed.
- D. Y. Alawadi, H. A. Saadeh, H. Kaur, K. Goyal, R. Sehgal, T. Ben Hadda, N. A. ElSawy and M. S. Mubarak, Med. Chem. Res., 2015, 24, 1196–1209 CrossRef CAS.
- A. Mehrabifar and M. Rahmati, MedBioTech J., 2017, 1, 15–21 Search PubMed.
- S. S. Alsayed, H. A. Elshemy, M. A. Abdelgawad, M. S. Abdel-Latif and K. R. Abdellatif, Bioorg. Chem., 2017, 70, 173–183 CrossRef CAS PubMed.
- L. Kasper, P. Miramón, N. Jablonowski, S. Wisgott, D. Wilson, S. Brunke and B. Hube, J. Med. Microbiol., 2015, 64, 714–723 Search PubMed.
- H.-J. Gillmann, J. Reichart, A. Leffler and T. Stueber, PLoS One, 2022, 17, e0264440 CrossRef CAS PubMed.
- A. Sharma, V. Kumar, R. Kharb, S. Kumar, P. Chander Sharma and D. Pal Pathak, Curr. Pharm. Des., 2016, 22, 3265–3301 CrossRef CAS PubMed.
- A. Siwach and P. K. Verma, BMC Chem., 2021, 15, 12 CrossRef CAS PubMed.
- F. G. Calvo-Flores and C. Mingorance-Sánchez, ChemistryOpen, 2021, 10, 815–829 CrossRef CAS PubMed.
- T. El Achkar, H. Greige-Gerges and S. Fourmentin, Environ. Chem. Lett., 2021, 19, 3397–3408 CrossRef CAS.
- R. Javahershenas, J. Mol. Liq., 2023, 385, 122398 Search PubMed.
- M. Alagawany, S. S. Elnesr, M. R. Farag, K. El-Naggar, A. E. Taha, A. F. Khafaga, M. Madkour, H. M. Salem, A. M. El-Tahan and M. T. El-Saadony, J. Therm. Biol., 2022, 104, 103168 CrossRef CAS PubMed.
- S. Pal, D. Das and S. Bhunia, Org. Biomol. Chem., 2024, 22, 1527–1579 Search PubMed.
- Deepa, M. Aalam and S. Singh, ChemistrySelect, 2022, 7, e202103918 CrossRef CAS.
- M. Li, S. He, L. Li, J. Zhuang and H. Wang, J. Food Meas. Charact., 2024, 18, 9752–9765 Search PubMed.
- S. Kang, G. Zhang, X. Yang, H. Yin, X. Fu, J. Liao, J. Tu, X. Huang, F. G. F. Qin and Y. Xu, Energy Fuels, 2017, 31, 2847–2854 CrossRef CAS.
- H. Qin, Z. Song, Q. Zeng, H. Cheng, L. Chen and Z. Qi, AIChE J., 2019, 65, 675–683 CrossRef CAS.
- B. Joos, T. Vranken, W. Marchal, M. Safari, M. K. Van Bael and A. T. Hardy, Chem. Mater., 2018, 30, 655–662 CrossRef CAS.
- H. Meng, S. Xue, S. Ding, Q. Wu, Z. Huang and Q. Zhang, J. Chromatogr. A., 2025, 466130 CrossRef CAS PubMed.
- Q. Wu, D. Zhang, S. Sun, C. Liu and C. Wang, Catalysts, 2023, 13, 1002 CrossRef CAS.
- B. Das, J. Kashanna, R. A. Kumar and P. Jangili, Monatsh. fur Chem, 2013, 144, 223–226 CrossRef CAS.
- S. Saeedi and A. Rahmati, RSC Adv., 2022, 12, 11740–11749 RSC.
- K. Van Aken, L. Strekowski and L. Patiny, Beilstein J. Org. Chem., 2006, 2, 3 CrossRef PubMed.
- Y.-C. Liu, Y.-D. Yang, W.-Q. Liu, T.-T. Du, R. Wang, M. Ji, B.-B. Yang, L. Li and X.-G. Chen, Bioorg. Med. Chem., 2022, 65, 116757 CrossRef CAS PubMed.
- W. Gu, T.-T. Miao, D.-W. Hua, X.-Y. Jin, X.-B. Tao, C.-B. Huang and S.-F. Wang, Bioorg. Med. Chem. Lett., 2017, 27, 1296–1300 CrossRef CAS PubMed.
- W.-B. Kuang, R.-Z. Huang, Y.-L. Fang, G.-B. Liang, C.-H. Yang, X.-L. Ma and Y. Zhang, RSC Adv., 2018, 8, 24376–24385 RSC.
- A. Negi, J. M. Alex, S. M. Amrutkar, A. T. Baviskar, G. Joshi, S. Singh, U. C. Banerjee and R. Kumar, Bioorg. Med. Chem., 2015, 23, 5654–5661 CrossRef CAS PubMed.
- M. K. Temre, B. Devi, V. K. Singh, Y. Goel, S. Yadav, S. K. Pandey, R. Kumar, A. Kumar and S. M. Singh, J. Biomol. Struct. Dyn., 2023, 41, 11262–11273 CrossRef CAS PubMed.
- C. Zhang, J. Phys. Chem. A, 2006, 110, 14029–14035 CrossRef CAS PubMed.
- S. Noriega, J. Cardoso-Ortiz, A. López-Luna, M. D. Cuevas-Flores and J. A. Flores De La Torre, Pharmaceuticals, 2022, 15, 717 CrossRef CAS PubMed.
- G. Muheyuddeen, M. Y. Khan, T. Ahmad, S. Srivastava, S. Verma, M. S. Ansari and N. Sahu, Sci. Rep., 2024, 14, 23121 CrossRef CAS PubMed.
- J. C. Phillips, R. Braun, W. Wang, J. Gumbart, E. Tajkhorshid, E. Villa, C. Chipot, R. D. Skeel, L. Kalé and K. Schulten, J. Comput. Chem., 2005, 26, 1781–1802 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |