 Open Access Article
Open Access ArticleSustainable and innovative electrochemical approach for quantifying glycine betaine in bacterial cultures
El-Sayed Khafagya,
Amr Selim Abu Lilab,
Ashraf M. Ashmawyc,
Ragab A. M. Saidde and
Mahmoud Rabee *f
*f
aDepartment of Pharmaceutics, College of Pharmacy, Prince Sattam Bin Abdulaziz University, Al-kharj 11942, Saudi Arabia
bDepartment of Pharmaceutics, College of Pharmacy, University of Ha'il, Ha'il 81442, Saudi Arabia
cChemistry Department, Faculty of Science, Al-Azhar University, Nasr City 11884, Cairo, Egypt
dPharmaceutical Analytical Chemistry Dept., Faculty of Pharmacy, Al-Azhar University, 11751, Cairo, Egypt
eChemistry Department, Faculty of Pharmacy, Heliopolis University, 11785, Cairo, Egypt
fResearch and Development Dept., Heliopolis University, 11785, Cairo, Egypt. E-mail: Mahmoud.rabee@hu.edu.eg; Tel: +00201125490478
First published on 19th March 2026
Abstract
Monitoring bacterial biosynthesis in real-world developments using electrochemical sensing has been a challenging task. Our research focused on fabricating and optimizing a sustainable and economical electrochemical sensor (GB-CG) to quantify glycine betaine (GB) amount in various bacterial cultures. The sensor's design was based on the ion-association complex between GB and phosphotungstic acid (PTA) anion as an ion exchange site, using polyvinyl chloride (PVC) as the main polymeric matrix and dioctyl phthalate (DOP) as a solvent mediator/plasticizer. It exhibited a rapid, linear, and stable linear Nernstian response (57.52 mV per decade) over a wide concentration range (1 × 10−7 to 1 × 10−1 M), with a detection limit of 1 × 10−8 M. The performance of the proposed sensor was evaluated in terms of selectivity, response time, operational lifespan, and pH/temperature working range, together with the key validation parameters. In comparison to a reported HPLC method, the sensor was efficiently utilized to determine the GB amount in three different bacterial cultures. The selected bacterial species were Escherichia coli, Bacillus subtilis, and Corynebacterium glutamicum. The sensor was also evaluated by using a Trichromatic Sustainability Assessment (TSA) protocol to ensure its environmental compatibility, sustainability, and practicality. Also, the proposed sensor was statistically compared to recently reported GB sensors, proving its reliability and optimal performance.
1. Introduction
Glycine betaine (GB, betaine, trimethylglycine, N,N,N-trimethylammonioacetate) is a small, highly polar quaternary ammonium osmoprotectant that accumulates intracellularly in different bacteria, plants, and animals.1 It plays an important role in the stabilization processes of the membranes and protein structures inside the cell.2 It also helps the cells maintain their osmotic balance without affecting the physiological processes and protects them from different abiotic stresses.3GB has become a high-demand product recently due to its numerous beneficial applications in different fields.4 In agriculture, it plays an important role in improving the plant tolerance against different abiotic stresses (high salinity, severe heat/dryness).5 It also protects the plants from the heavy metals' toxicity by stabilizing the intracellular structures and boosting the antioxidant enzymatic activities.6 In the industry, GB is widely used as a protein stabilizer for several products. It is also used for the production of eco-friendly solvents called deep eutectic solvents (DES) and natural deep eutectic solvents (NADES).7 These solvents are used as effective green alternatives to the toxic organic solvents in extracting natural compounds and raw materials.8 GB has gained considerable interest in the pharmaceutical field as a potential therapeutic agent for treating homocystinuria.9 It is a rare genetic syndrome characterized by elevated homocysteine levels, which can lead to many cardiovascular complications.10 GB can reduce the homocysteine levels in the blood and also improve the blood flow by reducing platelet aggregation and preventing the formation of thrombosis without undesirable side effects.11 This information boosts many clinical trials to study the option of using GB as a potential anticoagulant and anti-hemorrhagic agent for long-term treatments.12
The main traditional source for GB is extracting it from natural plant-based materials, such as sugar beet molasses and brown marine algae.13 Chemical synthesis can be carried out to meet the industrial demand, but it suffers from several drawbacks, like complex purification steps, generation of toxic by-products, and high production costs.14 The GB biosynthesis through biotechnological routes becomes a more favorable and cost-effective alternative.15 One of the recent biotechnological advancements is “bacterial milking.” It is a procedure that enables the repeated extraction of the intracellular GB from the living bacterial cells that grow under osmotic stress. This increases the yield sustainably without harming the living bacterial cells. Commonly used bacteria for this purpose include Escherichia coli, Bacillus subtilis, and Corynebacterium glutamicum.16,17
Introducing rapid and selective analytical approaches to quantify the GB amount during large-scale biosynthesis production is a critical step. Different analytical methods have been reported for quantifying GB in bacterial cultures. These include high-performance liquid chromatography (HPLC),18–20 liquid chromatography-mass spectrometry(LC-MS),21–24 and spectrophotometric assays.25 However, these methods have many limitations, such as tedious sample preparation, expensive equipment, and the need for trained personnel.26 Electrochemical sensors provide many benefits over those conventional methods. They are characterized by their simple instrumentation and operation, reasonable selectivity, and short time of analysis. These benefits make them suitable for reliable large-scale bacterial biosynthesis monitoring.27 A few electrochemical sensors have been recently developed for determining GB in different matrices like plant tissue, blood, and urine samples.28–30 Despite their high sensitivity, these sensors face challenges in fabrication and practical utility, limiting their suitability for quick on-site analysis. Also, none of these sensors has been used to determine GB in bacterial cultures. Potentiometric sensors provide many benefits over those existing sensors. These benefits include easy fabrication and operation, cost-effectiveness, quick analysis time, improved selectivity, and a low detection limit.31
Our research focused on developing a sustainable and economical electrochemical sensor for the rapid, in situ quantification of GB in bacterial cultures without the need for complex sample preparation or derivatization procedures. The developed sensor may be very useful in selecting and screening the high-yield species or strains of GB. It could also be valuable for microbiologists conducting research on halophilic or osmoprotectant-producing bacteria. The GB-CG sensor could also help enhance bacterial biosynthesis and facilitate advancements in the large-scale GB production. The sensor's design was based on an ion-association complex between GB and the phosphotungstic acid (PTA) anion, which acts as an ion–exchange site. This complex was then incorporated into a polymeric matrix using polyvinyl chloride (PVC) as the main component and dioctyl phthalate (DOP) as a solvent mediator and plasticizer. The resulting membrane was then glued to a graphite rod. A comprehensive assessment of the proposed sensor was conducted to ensure its optimal performance. In comparison to a reported HPLC method,32 the sensor effectively quantified the GB amount in three different bacterial cultures: Escherichia coli, Bacillus subtilis, and Corynebacterium glutamicum. Table 1 demonstrates a comparison between the GB-CG sensor and chromatographic methods in terms of analysis time, energy consumption, solvent usage, total costs, and waste production. The sensor was also statistically compared to other recently reported GB sensors to confirm its optimum performance.28–30
| GB-CG sensor | Chromatographic methods | |
|---|---|---|
| a Energy is measured in kilowatt-hours (kWh), a unit equivalent to 3600 kilojoules. | ||
| Analysis time | Seconds | Minutes to hours |
| Energy consumption a | < 0.1 kWh per sample | > 1.5 kWh per sample |
| Solvents usage | Minimal | Intermediate to high |
| Total costs | Minimal | Intermediate to high |
| Waste production | Minimal | Intermediate to high |
2. Experimental
2.1. Instruments
The following instruments were used for the respective procedures. Potentiometric measurements were performed using a JENWAY 3540 mV pH−1 bench meter with an Ag/AgCl reference electrode (UK). Stoichiometric/conductometric titration was conducted with a JENWAY 4520 bench conductivity meter (UK). Elemental analysis of the ion associate was carried out using a Vario EL cube elemental analyzer (Germany). Bacterial sample preparation was performed using a Thermo Scientific Sorvall ST 8 centrifuge (USA).2.2. Materials
2.3. Standard solutions
A 1 × 10−1 M stock GB solution was prepared by dissolving 1171.5 mg of the standard powder in 100 mL of distilled water. From this stock, working solutions with concentrations varying from 1 × 10−10 to 1 × 10−2 M were freshly prepared by serial dilution of the stock standard solution using the same solvent.2.4. Samples solutions
Escherichia coli, Bacillus subtilis, and Corynebacterium glutamicum were obtained and stored on Luria-Bertani (LB) medium agar plates. They were cultured aseptically in three separate 100 mL conical flasks using LB medium. These flasks were then incubated aerobically at 37 °C and 200 rpm for 24 hours to promote growth. The seed cultures of each bacterium were aseptically transferred into 250 mL flasks containing 100 mL of LB medium. The medium was supplemented with 0.4% glucose as a carbon source, 0.2% choline chloride as a precursor for GB production, and 10% (w/v) NaCl to induce osmotic stress and choline oxidation. The cultures of each bacterium type (n = 5) were incubated aerobically at 37 °C with shaking at 200 rpm for 24 hours to ensure complete intracellular GB accumulation. After incubation, bacterial cells of each bacterium were harvested by centrifugation at 4000 rpm for 20 minutes. The supernatant was discarded, and the resulting bacterial pellet was collected. The harvested bacterial pellet for each bacterium was weighed, and its wet weight (in g) was recorded. The bacterial pellet of each bacterium was then separately re-suspended in 10 mL of distilled water. Bacterial cell lysis was performed by ultrasonic bath for 10 minutes to release the intracellular GB. The obtained lysate was centrifuged again at 4000 rpm for 20 minutes to remove cell debris. The clear supernatants, containing the extracted GB, were transferred to clean 25 mL Falcon tubes. The pH of each sample was brought to 5.5 by dropwise addition of 0.1 N NaOH or 0.1 N HCl to be suitable for the working pH range (3–8) of the proposed sensor. The procedure was summarized in Fig. 1. The preconditioned GB-CG sensor was applied to determine the GB amount (mg g−1) in each sample at room temperature (25 °C). The samples (n = 5) were additionally analyzed using a reported HPLC method,32 and the acquired results were compared with those obtained from the proposed GB-CG sensor.2.5. Procedures
3. Results and discussion
The quaternary amino group (R-N+(CH3)3) in the GB structure indicates a potential to form a water-insoluble ion-associate with anionic substances. PTA, a common reagent, reacts with nitrogenous basic compounds to yield such water-insoluble complexes.33 This principle suggests that the ion association can be utilized to develop an economical, sustainable potentiometric sensor (GB-CG) for the on-site measurement of GB levels in bacterial cultures during production.The stoichiometry of the formed GB-PTA ion-associate was determined using conductometric titration according to the described procedure in ref. 34–36. Fig. 2(A) presents the conductometric titration curve, where conductance values (mS) were plotted versus the GB![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PTA molar ratio. A distinct break in the conductance curve appeared at a 1
PTA molar ratio. A distinct break in the conductance curve appeared at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 GB
1 GB![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PTA ratio, indicating 1
PTA ratio, indicating 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry. This finding was further verified by elemental analysis, with results provided in Table S1. Elemental analysis is a crucial and definitive characterization tool for ion-associate-based sensors. It provides quantitative evidence of the chemical composition and purity of the electro-active material, directly affecting the density and availability of fixed anionic exchange sites in the polymeric membrane, which in turn influences sensor performance.37 While Fourier Transform Infrared (FTIR) spectroscopy can identify functional groups, it may not clearly show shifts upon ionic association and does not measure bulk purity.38 Scanning Electron Microscopy (SEM) with Energy Dispersive X-ray (EDX) Analysis can reveal surface morphology but may not directly correlate with electrochemical performance once the material is plasticized within a PVC matrix. It confirms surface elemental presence but lacks the detailed stoichiometric and purity data provided by bulk elemental analysis.39
1 stoichiometry. This finding was further verified by elemental analysis, with results provided in Table S1. Elemental analysis is a crucial and definitive characterization tool for ion-associate-based sensors. It provides quantitative evidence of the chemical composition and purity of the electro-active material, directly affecting the density and availability of fixed anionic exchange sites in the polymeric membrane, which in turn influences sensor performance.37 While Fourier Transform Infrared (FTIR) spectroscopy can identify functional groups, it may not clearly show shifts upon ionic association and does not measure bulk purity.38 Scanning Electron Microscopy (SEM) with Energy Dispersive X-ray (EDX) Analysis can reveal surface morphology but may not directly correlate with electrochemical performance once the material is plasticized within a PVC matrix. It confirms surface elemental presence but lacks the detailed stoichiometric and purity data provided by bulk elemental analysis.39
 | ||
| Fig. 2 (A) Conductometric curve for titration of 10 mL of 10−1 M GB with 10−1 M PTA, (B) typical calibration curve for the GB-CG sensor. | ||
The GB-PTA complex results from van der Waals interactions between the quaternary amino group (R-N+(CH3)3) of GB and the anionic PTA−, as illustrated in Fig. 3(A). The sensing mechanism and response of the GB-CG sensor depended on an ionic charge transfer that occurred when GB molecules in the sample solution bound to receptor sites within the GB-PTA polymeric membrane. This charge transfer established an ion-exchange equilibrium, causing a buildup of positive charge at the membrane interfaces. Consequently, conducting electrons from the graphite rod were attracted to these positively charged sites, generating an electric field and altering the potential at the sensor surface, as described in Fig. 3(B).
 | ||
| Fig. 3 (A) Reaction pathway for the GB-PTA association, (B) sensing mechanism and response of the GB-CG sensor. | ||
Based on this data, an electrochemical/potentiometric cell was conceptualized, comprising two primary components. The first is the GB-CG sensor, which generates a sample-dependent potential, and the second is an Ag/AgCl reference electrode, which provides a constant, sample-independent potential (k). The potential difference between these electrodes (ECell) was measured using a potentiometer. The relationship between (ECell) and glycine betaine concentration [GB] follows the Nernst equation for potentiometry, demonstrating a linear correlation as shown in eqn (1):
Ecell = k + 0.05916![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log[GB]Sample log[GB]Sample
| (1) |
3.1. Sensor performance evaluation
The analytical performance of the GB-CG sensor was assessed in accordance with the IUPAC guidelines.40 Key parameters assessed included selectivity, response time/range, detection limit, operational pH/temperature range, stability, operational lifetime, accuracy, and precision. Fig. 2(B) presents the calibration curve for the sensor, displaying potential (mV) versus the negative logarithm of GB concentration. The calibration slope was measured at 57.52 mV per decade, closely approaching the theoretical Nernstian value of 59.1 mV per decade. The sensor demonstrated a linear Nernstian response across a broad concentration range from 1 × 10−7 M to 1 × 10−1 M, with a detection limit of 1 × 10−8 M. Excellent linearity was confirmed by a coefficient of determination (r2) of 0.9996. A complete summary of the sensor's performance characteristics is provided in Table 2.| Performance features | The GB-CG sensor |
|---|---|
| Linearity (M) | 10−7–10−1 |
| Slope (mV per decade) | −57.52 |
| Intercept | 258.7 |
| Correlation coefficient (r2) | 0.9996 |
| LOD | 10−8 M |
| LOQ | 10−7 M |
| Accuracy (mean ± SD) | 98.56 ± 0.96 |
| Repeatability (% RSD) | 0.322 |
| Intermediate precision (% RSD) | 1.122 |
| Operational pH range | 3–8 |
| Response time (second) | 10 |
| Operational temp. Range (°C) | 10–40 |
| Stability and life-time (days) | 75 |
3.2. Optimization of sensor membrane composition
The GB-CG sensor's polymeric membrane consists of two key elements: the GB-PTA ion-associate and solvent mediators/plasticizers. For an effective active membrane, it is essential to employ an ion-associate that exhibits stability, rapid exchange kinetics, suitable hydrophobicity, solubility, and overall durability.41 In this study, phosphotungstic acid (PTA) was selected as the optimal electroactive compound for forming the ion-associate with GB, owing to its high stability and rapid exchange kinetics. Other compounds, like sodium tetraphenyl borate, molybdic acid, silicotungstic acid, and tributyl phosphate produced unstable ion-associates with significantly non-Nernstian responses.The selection of an optimal ion-pairing agent for GB was guided by four fundamental chemical requirements: (i) sufficient charge density to form a stable electrostatic complex with the quaternary amino group (R-N+(CH3)3) of GB, (ii) adequate lipophilicity to ensure retention within the hydrophobic PVC membrane, (iii) structural complementarity with GB for ordered packing, and (iv) rapid, reversible ion-exchange kinetics. Systematic evaluation of alternative reagents revealed their failure to satisfy one or more of these criteria. Sodium tetraphenyl borate, despite its established utility in ion-selective electrodes, formed a GB-TPB ion-pair with inadequate lipophilicity and poor steric complementarity between the compact GB and the bulky tetraphenylborate anion, resulting in rapid leaching and unstable potentials. Molybdic acid, existing as small, highly charged molybdate anions, produced extremely hydrophilic ion-pairs with high hydration energies that strongly favored aqueous partitioning over membrane retention. Silicotungstic acid, while structurally analogous to phosphotungstic acid, carries a higher negative charge, which increases hydration and reduces lipophilicity, compromising membrane compatibility. Tributyl phosphate, as a neutral organophosphate ester, lacks anionic exchange sites entirely and therefore cannot function as an ion-exchanger, yielding only non-specific responses. In contrast, phosphotungstic acid uniquely satisfies all four requirements: it provides optimal charge density for stable electrostatic complexation; its large, delocalized polyoxometalate structure confers sufficient lipophilicity for membrane retention; its approximately spherical geometry offers complementary surface interactions with the GB; and its oxygen-bridged tungsten-oxo framework facilitates rapid, reversible ion-exchange kinetics, collectively yielding the observed near-Nernstian response and excellent membrane stability.
Polyvinyl chloride (PVC) with high-molecular-weight was selected as the main polymeric matrix, as it offers sufficient structural support and effectively entraps the other membrane components. An effective membrane plasticizer must impart flexibility and increase the distribution constant of the ion exchanger.42 In this research, DOP proved to be the optimal plasticizer, as it produced a favorable Nernstian response. This performance is due to its suitable lipophilicity, which facilitates the solvation and mobility of the ion-associate inside the polymeric membrane.
The GB-CG sensor's polymeric membrane was prepared by combining GB-PTA ion-associate, DOP, and PVC in specific weight proportions. In this study, we systematically varied the weight percentage of GB-PTA ion-associate while maintaining a constant 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio between DOP and PVC (Table 3). The membrane formulation containing 20% GB-PTA exhibited optimal performance, demonstrating a Nernstian slope of −57.52 mV per decade.
1 ratio between DOP and PVC (Table 3). The membrane formulation containing 20% GB-PTA exhibited optimal performance, demonstrating a Nernstian slope of −57.52 mV per decade.
| Membrane composition (%w/w) | Linearity (M) | Slope (mV per decade) | Intercept | Correlation coefficient (r2) | ||
|---|---|---|---|---|---|---|
| GB- PTA | PVC | DOP | ||||
| 5 | 47.5 | 47.5 | 10−7–10−1 | −50.2 | 246.45 | 0.9982 |
| 10 | 45 | 45 | 10−7–10−1 | −53.3 | 247.3 | 0.9921 |
| 15 | 42.5 | 42.5 | 10−7–10−1 | −54.12 | 246.5 | 0.9987 |
| 20 | 40 | 40 | 10−7–10−1 | −57.52 | 258.7 | 0.9996 |
| 25 | 37.5 | 37.5 | 10−7–10−1 | −52.25 | 244.5 | 0.9992 |
| 30 | 35 | 35 | 10−7–10−1 | −51.14 | 238.5 | 0.9971 |
An optimal polymeric membrane thickness of approximately 1 mm was determined to balance response time and sensitivity while preserving the coating's structural integrity. Excessive thickness can prolong response time by increasing the analyte's diffusion path through the membrane and may reduce sensitivity by attenuating the analyte concentration that reaches the electrode surface. On the other hand, thinner coatings, while potentially faster and more sensitive, often lack the robustness required for sustained operational durability.
3.3. Effect of conditioning time on sensor performance
Prior to initial use, the newly fabricated GB-CG sensor must be soaked in a GB solution to activate its membrane surface, forming a thin gel layer essential for ion exchange. The sensor was conditioned in a 1 × 10−1 M GB solution and calibrated periodically until the response slope deviated significantly from Nernstian behavior. As shown in Table S2, a conditioning period of two hours was found to be optimal. As it yields a near-ideal Nernstian behavior with a slope of −57.52 mV per decade and a high coefficient of determination (r2 = 0.9996).3.4. Sensor selectivity assessment
The selectivity of the GB-CG sensor was evaluated against various potential interfering substances following the IUPAC-recommended Separate Solution Method (SSM). This procedure involved measuring the potentiometric response of both a 1 × 10−3 M GB solution and individual 1 × 10−3 M solutions of each interfering species. The corresponding selectivity coefficients (KC+pot.GB,X) were subsequently calculated using eqn (2) 40:
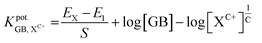 | (2) |
| Interferents a | Selectivity coefficientsb (KC+pot.GB,X) |
|---|---|
| GB-CG sensor | |
| a All solutions were prepared at a concentration of 10−3 M.b Each value is the mean of three replicate measurements. | |
| Na+ | 2.4 × 10−2 |
| Mg2+ | 1.1 × 10−3 |
| Ni2+ | 4.2 × 10−3 |
| K+ | 5.4 × 10−3 |
| Cu2+ | 1.9 × 10−2 |
| Ca2+ | 3.4 × 10−2 |
| Choline | 2.9 × 10−1 |
| Glycine | 3.6 × 10−1 |
| Dimethyl glycine | 1.2 × 10−1 |
| Carnitine | 1.6 × 10−1 |
| Glucose | 7.6 × 10−3 |
| Fructose | 5.3 × 10−3 |
| Sucrose | 2.5 × 10−4 |
| L-alanine | 1.3 × 10−3 |
| Asparagine | 3.4 × 10−3 |
| L-cysteine | 7.2 × 10−3 |
The exceptional selectivity of the GB-CG sensor can be explained by three key principles: ion-ionophore complementarity, ion hydration energetics, and lipophilicity-driven partitioning.43 The experimental selectivity coefficients (KC+pot.GB,X) obtained for the tested potential interferents align with these principles. Metal cations exhibited low selectivity coefficients due to their high hydration energies, such as Mg2+ with a ΔGhyd of −1900 kJ mol−1 and Ca2+ with −1500 kJ mol−1, making it energetically unfavorable for them to lose their hydration shells and enter the membrane, leading to their preferential retention in the aqueous phase.34
For structural analogs, the sensor's selectivity for GB molecules is attributed to ion-ionophore complementarity “lock-and-key” mechanism.44 The sensor's membrane contains a GB-PTA ion associate, which acts as a specific receptor site with defined steric and electronic complementarity to GB. Choline, on the other hand, despite having a quaternary ammonium group, lacks a carbonyl group and cannot engage in the necessary hydrogen bonding or dipole interactions with the PTA framework that stabilize the ion-associate complex. This lack of ion-ionophore complementarity leads to a lower binding affinity and less partitioning into the membrane phase. Glycine lacks methylation on its nitrogen atom, leading to a primary ammonium group with reduced cationic character and limited steric bulk. This hinders the formation of an effective ion-pair. Dimethylglycine, with two methyl groups on its nitrogen, has an incomplete methylation pattern that does not provide the optimal steric fit needed for stable complexation in the receptor site. Carnitine, with its quaternary ammonium group and carboxylate functionality, has a long carbon chain that ends with a distal carboxylate. This structure creates steric hindrance and unfavorable electrostatic repulsion, making it challenging to fit into the binding site due to geometric constraints.
Other common bacterial matrix components such as sugars and amino acids, have very low selectivity coefficient values ranging between 10−4 and 10−3, which means they do not significantly interfere. This can be attributed to their high hydrophilicity (low lipophilicity), which provides no thermodynamic driving force for partitioning into the hydrophobic membrane.45
3.5. Sensor response time
The sensor response time was determined by recording the time required to achieve a stable potential reading within a ±3 mV variation. The sensor was tested in successive GB solutions across a concentration range from 1 × 10−7 M to 1 × 10−1 M while monitoring the potential response. For all concentrations tested, the sensor reached equilibrium within 10 seconds, as demonstrated in Fig. 4(A). The measured potentials remained stable within the ±3 mV threshold for a minimum period of 300 seconds (5 minutes). | ||
| Fig. 4 (A) The GB-CG sensor response time, (B and C) the pH and temperature profiles of the GB-CG sensor, (D) stability and life-time profile of the GB-CG sensor. | ||
3.6. Sensor measurement reproducibility
The sensor measurement reproducibility was evaluated by measuring a 1 × 10−2 M GB solution immediately after measuring a 1 × 10−3 M GB solution. For six replicate measurements, the percent relative standard deviation (%RSD) was 0.729% at 1 × 10−3 M and 0.515% at 1 × 10−2 M. Both %RSD values being below 2% confirms the sensor's excellent measurement reproducibility.3.7. Influence of pH on the GB-CG sensor response
The influence of pH on the GB-CG sensor's performance was studied to identify its optimal operational range. Sensor potential was recorded for a 1 × 10−2 M GB solution across pH levels from 0.5 to 12. As shown in Fig. 4(B), the sensor showed a consistent and reliable response from approximately pH 2.7 to pH 8. To ensure reliability and buffer capacity under all conditions, the operational range was reported as pH 3–8. The sensor maintains a stable response even at pH 8, where GB exists predominantly as a zwitterion. This behavior is explained by the sensor's recognition mechanism: the quaternary ammonium group of GB carries a permanent positive charge independent of pH, while the distal carboxylic acid group undergoes deprotonation above pH 2 (pKa ∼1.8), resulting in a net neutral zwitterion. The sensing mechanism, as depicted in Fig. 3(B), relies on the ion-exchange interaction between the GB quaternary ammonium group and the anionic PTA sites in the polymeric membrane. The protonation state of the distal carboxyl group does not affect this recognition event, allowing the sensor to selectively respond to the quaternary ammonium moiety over a broad pH range.Outside this range, at pH values below 3 and above 8, the measured potential gradually decreased and became unstable. This deviation is attributed to protonation and deprotonation of the PTA itself in the ion-associate, leading to charge loss and eventual dissociation of the GB-PTA complex. Therefore, all subsequent measurements were conducted within the pH 3–8 range to ensure reliable sensor performance.
3.8. Influence of temperature on the GB-CG sensor response
The influence of temperature on the GB-CG sensor's performance was studied to identify its optimal operational range. Sensor potential was recorded for a 1 × 10−2 M GB solution across temperature levels from 5 °C to 55 °C. As shown in Fig. 4(C), the sensor exhibited a stable response between 10 °C and 40 °C. Below 10 °C, the potential gradually decreased and became unstable, potentially due to partial surface freezing of the solution restricting GB mobility. Above 40 °C, a marked increase in potential was observed, likely resulting from accelerated GB diffusion and possible thermal effects on the sensor membrane that could reduce its operational lifetime.3.9. Sensor stability and lifetime
The sensor stability and lifetime were assessed by tracking its calibration graph slopes over a 75-day period. Throughout this duration, the sensor maintained stable Nernstian slopes near optimal values, confirming its reliable functionality for up to 75 days. Beyond this period, deviations from Nernstian behavior were observed, as illustrated in Fig. 4(D). This long-term stability was due to the strong binding between the membrane components and the hydrophobic PTA anion, which certainly anchor the GB-PTA complex at specific sites rather than allowing free diffusion. However, PTA transport involves jumps between binding sites, indicating a degree of mobility that may lead to gradual leaching over extended periods (>75 days), particularly under dynamic operational stress. Prolonged use beyond 75 days may also reduce or damage the membrane's mechanical properties, indirectly affecting the GB-PTA complex stability. Such damage could modify the membrane matrix, potentially inducing the complex leaching or dissociation.3.10. Method validation
3.11. Application
The fabricated sensor (GB-CG) was effectively used to determine the GB amount in three distinct bacterial cultures: Escherichia coli, Bacillus subtilis, and Corynebacterium glutamicum. As summarized in Table 5, the GB amounts are expressed in mg g−1 for each bacterium. Corynebacterium glutamicum exhibited the highest GB level among the three. To confirm the reliability of these findings, all bacterial samples were additionally prepared and analyzed via a previously established HPLC method.32 The chromatographic results aligned closely with those from the GB-CG sensor, as presented in Table 5. The chromatographic analysis of the bacterial samples was also illustrated in Fig. S1. Furthermore, the accuracy of the sensor was evaluated using the standard addition technique. Each bacterial sample solution was spiked with a GB standard solution at three different concentrations. Percent recoveries for GB were calculated using the corresponding regression equation after subtracting the amounts of the added standard. The standard addition method confirmed that the sensor's potential values accurately corresponded to the GB quantity, with no observable interference from other components in the bacterial matrix.| Samples | Amount of GB, mg g−1 (mean ± SD) | |
|---|---|---|
| GB-CG sensor | HPLC32 | |
| Escherichia coli | 15.266 ± 0.079 | 15.349 ± 0.127 |
| Bacillus subtilis | 19.015 ± 0.030 | 18.983 ± 0.275 |
| Corynebacterium glutamicum | 23.278 ± 0.123 | 23.429 ± 0.233 |
3.12. Trichromatic sustainabilityassessment (TSA) for the GB-CG sensor
The TSA is a recent, comprehensive method used primarily in analytical chemistry to evaluate a procedure's overall sustainability based on three distinct “color” metrics: green, blue, and white.463.13. A statistical comparison
To comprehensively evaluate the analytical performance and practical utility of the proposed GB-CG sensor, a detailed comparison was conducted with recently reported GB sensors from the literature (ref. 28–30), as summarized in Table 6. These comparisons are contextualized by briefly describing the sensing principles and target matrices of each reported method, as they differ substantially from the potentiometric approach presented herein.| Ref. | Linearity range (M) | LOD (M) | pH range | Life-time (days) | AGREE score | BAGI score | RGB12 score |
|---|---|---|---|---|---|---|---|
| GB-CG | 1.0 × 10−7–1.0 × 10−1 | 1.0 × 10−8 | 3.0–8.0 | 75 | 0.78 | 77.5 | 90.0 |
| Ref. 28 | 1.0 × 10−5–6.0 × 10−5 | 7.6 × 10−8 | — | — | 0.65 | 72.5 | 88.5 |
| Ref. 29 | 1.0 × 10−15–1.0 × 10−2 | 7.1 × 10−16 | 7.1–7.5 | 14 | 0.72 | 75.0 | 84.7 |
| Ref. 30 | 2.8 × 10−4–2.8 × 10−3 | 1.5 × 10−4 | 7.1–7.3 | 1 | 0.68 | 72.5 | 85.5 |
Raja et al. (ref. 28) developed an electrochemical sensor based on a molecularly imprinted polymer (MIP) fabricated on a glassy carbon electrode, which was successfully applied to GB determination in Adhatoda vasica leaf extracts. This sensor demonstrated good selectivity owing to the MIP recognition sites but exhibited a relatively narrow linear range (1.0 × 10−5 to 6.0 × 10−5 M) and a limited operational lifetime of only 5 days. Ai et al. (ref. 29) reported an exceptionally sensitive photoelectrochemical sensor utilizing an advanced nanocomposite architecture, achieving detection limits in the femtomolar range (7.1 × 10−16 M) with a remarkably broad linear response from 1.0 × 10−15 to 1.0 × 10−2 M. This sensor was validated in human urine and serum samples, demonstrating the power of photoelectrochemical transduction for ultra-trace analysis, though its fabrication complexity and cost may limit routine adoption in resource-limited settings. Ahmad et al. (ref. 30) described a potentiometric sensor incorporating a multi-walled carbon nanotube (MWCNT) composite within a polymeric membrane, which was applied to GB determination in pharmaceutical formulations and plant tissue samples. While this sensor offered the simplicity inherent to potentiometry, its linear range (2.8 × 10−4 to 2.8 × 10−3 M) and detection limit (1.5 × 10−4 M) were modest compared to other reported methods.
In contrast to these existing sensors, the GB-CG sensor presented in this work offers several distinct advantages: (i) it represents the first potentiometric sensor designed explicitly for, and experimentally validated in, complex bacterial culture matrices without requiring extensive sample pretreatment; (ii) it achieves a wide linear range (1.0 × 10−7 to 1.0 × 10−1 M) with a detection limit of 1.0 × 10−8 M, well-suited for monitoring GB production in bacterial fermentation processes; (iii) its fabrication is remarkably simple and cost-effective, using readily available graphite rods and common chemicals; and (iv) it demonstrates superior sustainability metrics, including an AGREE greenness score of 0.78, a BAGI practicality score of 77.5, and an RGB12 whiteness score of 90.0, reflecting its environmental compatibility, operational simplicity, and economic viability. While the photoelectrochemical sensor reported by Ai et al. (ref. 29) offers superior sensitivity for trace analysis in clinical matrices, the GB-CG sensor provides an optimal balance of analytical performance, practical simplicity, and sustainability for routine monitoring of bacterial cultures, an application space previously unexplored with potentiometric methods. Additionally, with an extended operational lifetime of 75 days and a rapid 10-second response time, the GB-CG sensor is well-positioned for real-time, on-site monitoring in bioprocessing environments where robustness, simplicity, and cost-effectiveness are paramount considerations.
4. Conclusion
This research aimed to develop and optimize a polymer-coated graphite sensor (GB-CG) to be utilized for the direct in situ quantification of the GB amount in various bacterial cultures without the need for complex sample preparation steps. The GB-CG sensor was simply fabricated by using a basic graphite rod and commonly available chemicals. Its performance was evaluated using a standard laboratory pH/potentiometer, and the overall operation was cost-effective. The GB-CG sensor showed a fast, linear, and selective Nernstian response over a wide concentration range (1 × 10−7 to 1 × 10−1 M) with a low detection limit of 1 × 10−8 M. The sensor was effectively employed to determine the GB amounts in various bacterial cultures, producing results consistent with a reported HPLC method. Comparative studies under different experimental conditions demonstrated the advantages of the GB-CG sensor over previously reported GB sensors. However, validation in real-life conditions, such as production facilities, is essential. Future research has to explore its use to determine the GB amount in larger sample sizes and various types of bacterial cultures and also has to improve the sensor durability for precise monitoring in complex environments.Author contributions
El-Sayed Khafagy: methodology, validation, project administration, writing – review & editing. Amr Selim Abu Lila: conceptualization, writing – original draft. Ashraf M. Ashmawy: methodology, validation, project administration, writing – review & editing. Ragab A. M. Said: investigation, formal analysis, methodology, writing – review & editing. Mahmoud Rabee: methodology, validation, resources, writing – original draft. All authors contributed to the final version of the manuscript.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships.Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its Supplementary information (SI) file. Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00352d.Acknowledgements
This study is supported via funding from Prince Sattam bin Abdulaziz University project number (PSAU/2026/R/1447)References
- P. K. Chauhan, S. K. Upadhyay, M. Tripathi, R. Singh, D. Krishna, S. K. Singh and P. Dwivedi, Biotechnol. Genet. Eng. Rev., 2023, 39, 311–347 CrossRef CAS PubMed.
- X. Liang, S. Yu, Y. Ju, Y. Wang and D. Yin, Plants, 2025, 14, 2829 CrossRef PubMed.
- M. G. Annunziata, L. F. Ciarmiello, P. Woodrow, E. Dell'Aversana and P. Carillo, Front. Plant Sci., 2019, 10, 230 CrossRef PubMed.
- T. H. H. Chen and N. Murata, Plant Cell Environ., 2011, 34, 1–20 CrossRef CAS PubMed.
- M. Ashraf and M. R. Foolad, Environ. Exp. Bot., 2007, 59, 206–216 CrossRef CAS.
- M. Hasanuzzaman, K. Nahar, M. M. Alam, R. Roychowdhury and M. Fujita, Int. J. Mol. Sci., 2013, 14, 9643–9684 CrossRef PubMed.
- Y. Dai, J. Van Spronsen, G.-J. Witkamp, R. Verpoorte and Y. H. Choi, Anal. Chim. Acta, 2013, 766, 61–68 CrossRef CAS PubMed.
- Y. H. Choi, J. van Spronsen, Y. Dai, M. Verberne, F. Hollmann, I. W. C. E. Arends, G.-J. Witkamp and R. Verpoorte, Plant Physiol., 2011, 156, 1701–1705 CrossRef CAS PubMed.
- U. Schwab, A. Törrönen, L. Toppinen, G. Alfthan, M. Saarinen, A. Aro and M. Uusitupa, Am. J. Clin. Nutr., 2002, 76, 961–967 CrossRef CAS PubMed.
- S. van Calcar, Nutrition Management of Patients with Inherited Metabolic Disorders, 2009, 237 Search PubMed.
- M. R. Olthof and P. Verhoef, Curr. Drug Metabol., 2005, 6, 15–22 CrossRef CAS PubMed.
- P. Detopoulou, D. B. Panagiotakos, S. Antonopoulou, C. Pitsavos and C. Stefanadis, Am. J. Clin. Nutr., 2008, 87, 424–430 CrossRef CAS PubMed.
- F. Zhao, J. Luo, E. Ibrahim, L. Chen, Y. Shen, M. Ibrahim, W. B. Alonazi, J. Lu, Y. Luo and H. Wu, Crop Health, 2025, 3, 1–14 CrossRef PubMed.
- J. Boch, B. Kempf, R. Schmid and E. Bremer, J. Bacteriol., 1996, 178, 5121–5129 Search PubMed.
- S. A. Adu, M. S. Twigg, P. J. Naughton, R. Marchant and I. M. Banat, Molecules, 2023, 28, 4463 CrossRef CAS PubMed.
- T. Han and S. Y. Lee, Metab. Eng., 2023, 79, 78–85 CrossRef CAS PubMed.
- A. Rasheed, H. Li, M. M. Tahir, A. Mahmood, M. Nawaz, A. N. Shah, M. T. Aslam, S. Negm, M. Moustafa and M. U. Hassan, Front. Plant Sci., 2022, 13, 976179 CrossRef CAS PubMed.
- M.-A. Bessieres, Y. Gibon, J. C. Lefeuvre and F. Larher, J. Agric. Food Chem., 1999, 47, 3718–3722 CrossRef CAS PubMed.
- F. A. Mehta, B. G. Patel, S. S. Pandya and K. B. Ahir, J. Planar Chromatogr. - Mod. TLC, 2011, 24, 136–139 CrossRef CAS.
- F. J. De Zwart, S. Slow, R. J. Payne, M. Lever, P. M. George, J. A. Gerrard and S. T. Chambers, Food Chem., 2003, 83, 197–204 CrossRef CAS.
- S. J. Bruce, P. A. Guy, S. Rezzi and A. B. Ross, J. Agric. Food Chem., 2010, 58, 2055–2061 Search PubMed.
- S. L. MacKinnon and C. Craft, in Natural Products from Marine Algae: Methods and Protocols, Springer, 2015, pp. 267–275 Search PubMed.
- R. L. Airs and S. D. Archer, Limnol Oceanogr. Methods, 2010, 8, 499–506 CrossRef CAS.
- J. Liu, M. Zhao, J. Zhou, C. Liu, L. Zheng and Y. Yin, J. Chromatogr. B, 2016, 1035, 42–48 CrossRef CAS PubMed.
- M. G. Valadez-Bustos, G. A. Aguado-Santacruz, A. Tiessen-Favier, A. Robledo-Paz, A. Munoz-Orozco, Q. Rascón-Cruz and A. Santacruz-Varela, Anal. Biochem., 2016, 498, 47–52 CrossRef CAS PubMed.
- B. C. Pattnayak and S. Mohapatra, Spectrochim. Acta, Part A, 2023, 286, 122009 CrossRef CAS PubMed.
- I. Kathuria, S. Rani, R. K. Srivastava and S. Kumar, Microchem. J., 2024, 199, 110217 CrossRef CAS.
- L. Raja, S. Balamuthu, S. Venkatesan and L. Ming, J. Iran. Chem. Soc., 2024, 21, 2325–2333 CrossRef CAS.
- G. Ai, Y. Zhou, H. Zhang, Q. Wei, B. Luo, Y. Xie, C. Wang, X. Xue and A. Li, Food Chem., 2024, 435, 137554 CrossRef CAS PubMed.
- M. Ahmad, S. Ameen, T. Omar, P. Khan and A. Ahmad, Biosens. Bioelectron., 2016, 86, 169–175 CrossRef CAS PubMed.
- B. Mohan, V. Virender, R. K. Gupta, V. Pandey, A. J. L. Pombeiro and P. Ren, Trends Food Sci. Technol., 2025, 159, 104936 CrossRef CAS.
- T. P. Chendrimada, M. G. Neto, G. M. Pesti, A. J. Davis and R. I. Bakalli, J. Sci. Food Agric., 2002, 82, 1556–1563 CrossRef CAS.
- M. S. García, J. A. Ortuño, M. I. Albero and M. S. Abuherba, Sensors, 2009, 9, 4309–4322 CrossRef PubMed.
- S. M. Riad, F. I. Khattab, H. Salem and H. T. Elbalkiny, Anal. Bioanal. Electrochem., 2014, 6, 559–572 Search PubMed.
- R. O. El-Attar and E. A. Khaled, Egypt. J. Chem., 2021, 64, 6505–6513 CrossRef.
- O. I. A. Sattar, H. H. M. Abuseada, M. S. Emara and M. Rabee, J. Pharm. Biomed. Anal., 2022, 213, 114680 CrossRef CAS PubMed.
- W. Kandioller, J. Theiner, B. K. Keppler and C. R. Kowol, Inorg. Chem. Front., 2022, 9, 412–416 RSC.
- M. A. Peshkova, A. I. Korobeynikov and K. N. Mikhelson, Electrochim. Acta, 2008, 53, 5819–5826 CrossRef CAS.
- L. Perez-Marin, H. Lopez-Valdivia, P. Avila-Perez, E. Otazo-Sánchez, G. Macedo-Miranda, O. Gutierrez-Lozano, J. A. Chamaro, J. De la Torres-Orozco and L. Carapia-Morales, Analyst, 2001, 126, 501–504 RSC.
- E. Lindner and Y. Umezawa, Pure Appl. Chem., 2008, 80, 85–104 CrossRef CAS.
- H. K. Ibrahim, M. M. Abdel-Moety, S. A. Abdel-Gawad, M. A. Al-Ghobashy and M. A. Kawy, Environ. Sci. Pollut. Res., 2017, 24, 7023–7034 CrossRef CAS PubMed.
- D. Nashed, I. Noureldin and A. A. Sakur, BMC Chem., 2020, 14, 1–9 Search PubMed.
- E. Bakker, E. Pretsch and P. Bühlmann, Anal. Chem., 2000, 72, 1127–1133 CrossRef CAS PubMed.
- A. V Bondar, V. M. Keresten and K. N. Mikhelson, J. Anal. Chem., 2022, 77, 145–154 CrossRef.
- L. Mendecki, N. Callan, M. Ahern, B. Schazmann and A. Radu, Sensors, 2016, 16, 1106 CrossRef PubMed.
- D. S. El-Kafrawy and A. H. Abo-Gharam, BMC Chem., 2025, 19, 1–16 Search PubMed.
- F. Pena-Pereira, W. Wojnowski and M. Tobiszewski, Anal. Chem., 2020, 92, 10076–10082 CrossRef CAS PubMed.
- N. Manousi, W. Wojnowski, J. Płotka-Wasylka and V. Samanidou, Green Chem., 2023, 25, 7598–7604 RSC.
- L. Yin, L. Yu, Y. Guo, C. Wang, Y. Ge, X. Zheng, N. Zhang, J. You, Y. Zhang and M. Shi, J. Pharm. Anal., 2024, 101013 CrossRef PubMed.
- P. M. Nowak and F. Arduini, Green Anal. Chem., 2024, 10, 100120 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |


