 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Molecular dynamics simulations of temperature-dependent PET binding in PETase, ThermoPETase, and FAST-PETase
Athina Karaoli ad,
Dimitris G. Mintis
ad,
Dimitris G. Mintis ab,
Haralampos Tzoupisa,
Chris T. Kiranoudisd,
Iseult Lynch
ab,
Haralampos Tzoupisa,
Chris T. Kiranoudisd,
Iseult Lynch be,
Georgia Melagraki
be,
Georgia Melagraki f and
Andreas Afantitis
f and
Andreas Afantitis *abcg
*abcg
aNovaMechanics Ltd, Nicosia 1070, Cyprus. E-mail: afantitis@novamechanics.com
bEntelos Institute, Nicosia 2102, Cyprus. E-mail: i.lynch@bham.ac.uk
cNovaMechanics MIKE, Piraeus 18545, Greece
dSchool of Chemical Engineering, National Technical University, Zografou 15780, Athens, Greece
eSchool of Geography, Earth and Environmental Sciences, University of Birmingham, Birmingham B15 2TT, UK
fDivision of Physical Sciences and Applications, Hellenic Military Academy, Vari 16672, Greece
gDepartment of Pharmacy, Frederick University, Nicosia 1036, Cyprus
First published on 23rd March 2026
Abstract
Polyethylene terephthalate (PET) biodegradation has gained significant attention following the 2016 discovery of the PET-hydrolyzing enzyme IsPETase from Ideonella sakaiensis 201-F6. Although IsPETase operates under mild temperature conditions, its limited catalytic activity and poor thermal stability restrict large-scale industrial applications. To overcome these limitations, several engineered variants have been developed. In this study, the wild-type IsPETase was systematically compared with two engineered variants, ThermoPETase and FAST-PETase, under four temperature conditions (300, 313, 323, and 333 K) using fully detailed atomistic molecular dynamics (MD) simulations. A PET dimer was docked into the active site of each enzyme, followed by 150 ns of restrained and unrestrained MD simulations to evaluate structural stability and substrate interactions. Both engineered variants exhibited improved thermal stability relative to IsPETase, with FAST-PETase demonstrating the highest stability across all temperatures. Increased flexibility of the catalytic triad was observed in both engineered variants, suggesting enhanced catalytic adaptability compared to IsPETase. Key residues, Tyr87 and Met161, were identified as essential for stabilizing the PET dimer within the active-site cavity. At elevated temperature (333 K), IsPETase showed increased flexibility, leading to disrupted PET dimer binding. In contrast, ThermoPETase and FAST-PETase preserved stable PET dimer positioning at elevated temperatures, with FAST-PETase displaying the most favourable structural characteristics across all temperature conditions. Furthermore, binding energy calculations revealed a correlation between structural stability and reduced enthalpic binding energies at each enzyme's optimal temperature. Based on these analyses, the optimal temperatures for PET degradation were found to be 323 K for IsPETase, 333 K for ThermoPETase, and 300–313 K for FAST-PETase. These findings provide molecular-level insights into the structure–stability–activity relationships of PETase variants and highlight key determinants for the rational design of improved PET-degrading enzymes.
1 Introduction
It is estimated that by 2050, approximately 1.2 trillion tons of plastic waste will have accumulated in landfills and natural ecosystems,1 posing significant health risks to ecosystems and humans.2 The low cost and versatility of plastics3 make them a convenient solution for everyday human needs, with polyethylene terephthalate (PET) being the most widely produced thermoplastic polyester, due to its extensive use in the manufacture of water bottles, food packaging and textiles.4,5 Despite its widespread use, PET exhibits high resistance to degradation1 due to its physicochemical characteristics, particularly its high crystallinity and chemical stability.5 Environmental weathering processes, such as UV radiation, temperature fluctuations and mechanical abrasion,6 can lead to the fragmentation of PET into micro- and nano-plastics, contributing substantially to environmental pollution.7 As a result, these persistent fragments accumulate in diverse environmental compartments, including landfills8 and marine ecosystems,9 where they can persist for decades.10 Numerous studies11,12 have shown that micro- and nano-plastics can pose serious risks to both wildlife and humans through the contamination of drinking water sources and their entry into the food chain.11,12 The small size and large surface area of these particles also increases their potential to adsorb toxic pollutants such as perfluoroalkyl substances (PFAS) and polycyclic aromatic hydrocarbons (PAH), further intensifying their environmental impact.13–17The primary strategies for managing plastic waste are currently mechanical and chemical recycling.18 Mechanical recycling involves the grinding of plastics to convert them into other products.18 However, repeated processing diminishes the material quality.18 Chemical recycling methods such as chemolysis and pyrolysis, break down plastics into their constituent monomers, allowing the production of new plastics or other synthetic chemicals with properties similar to virgin materials.18–20 However, these approaches require high energy consumption and raise environmental concerns associated with CO2 emissions and the generation of toxic byproducts.19,21,22 An innovative biotechnological approach, enzymatic biodegradation, has been developed as a more environmentally friendly alternative for plastic management, relying on microbial enzymes capable of hydrolyzing plastic polymers.21,23
Polyester hydrolases, cutinases and lipases are some of the key enzymes involved in the degradation of PET.24,25 In 2005, the first enzyme, TfH hydrolase, isolated from Thermobifida fusca DSM43793 was recognized for its ability to degrade PET into its monomers (i.e. terephthalic acid and ethylene glycol).26 In the following years, enzymes such as the metagenomic leaf-branch compost cutinase (LCC)27 and TfCut2 isolated from Thermobifida fusca KW3 (ref. 28) also demonstrated significant PET-degrading capabilities. Among these, an enzyme called IsPETase, isolated from the bacterium Ideonella sakaiensis 201-F6 in 2016, gained attention for its ability to degrade PET at ambient temperatures.29 Nevertheless, for large-scale industrial applications, enzymes must be cost-effective, highly active, and stable under specific conditions.30 The wild-type enzymes often fail to meet these requirements due to low activity, poor stability, and a narrow substrate range.30 In the case of IsPETase, researchers have engineered several variants through targeted mutations such as ThermoPETase,31 DuraPETase32 and FAST-PETase,33 each exhibiting enhanced performance compared to the wild-type. These improvements are achieved by introducing mutations that increase the rigidity of flexible loops,34 thereby enhancing thermostability and shifting the enzyme's optimal activity toward temperatures closer to the glass transition temperature of PET (70 °C),31,35 resulting in higher catalytic efficiency.
The discovery of enzymes capable of degrading PET has stimulated significant interest in elucidating the atomistic mechanisms that govern enzyme–PET interactions. Over the past five years, an increasing number of fully atomistic Molecular Dynamics (MD) simulations has been conducted to study structural flexibility, substrate binding, and conformational dynamics of PET-degrading enzymes.36 In addition, significant progress has been achieved through the application of hybrid Quantum Mechanics/Molecular Mechanics (QM/MM) simulations aimed at investigating catalytic pathways of PET hydrolysis at the atomistic scale.37,38 Table S1 in the SI provides a comprehensive overview of MD-based studies focused on PET degradation, detailing critical parameters including the specific substrate modelled, the enzyme employed, docking protocols used, molecular force fields applied, simulation duration and temperature, as well as the software platforms utilized. Annotations are also provided for studies that incorporate QM/MM methodologies to model the catalytic mechanisms. Furthermore, Table S2 in the SI presents a list of studies that integrate MD simulations with experimental data, such as X-ray crystallography39 or enzymatic activity assays,40–42 offering valuable validation and mechanistic insight, outlining similar information as in Table S1.
As presented in Tables S1 and S2, the wild-type PETase enzyme, IsPETase,43–49 is extensively studied using high-resolution crystal structures (e.g., protein database (PDB) structures: 5XJH, 6EQE). These studies have provided detailed insights into the enzyme's relatively high activity at mesophilic temperatures (i.e., 20 to 45 °C),50 and its early success in demonstrating PET hydrolysis under mild conditions.40 In recent years, attention has shifted toward enzymes with improved thermostability and catalytic efficiency, such as LCC,39,51,52 DuraPETase,53–55 ThermoPETase,49,53 FAST-PETase,47,53,56 and PES-H1.51,57,58 Notably, enzymes such as PHL7 (ref. 59 and 60) and TurboPETase,61 which are either derived from thermophilic organisms or designed through AI-guided engineering, have gained attention for their robust activity at elevated temperatures (e.g., 65/70 °C).59 Similarly, Combi-PETase,62 a machine-learning-assisted engineered variant, has been investigated, reflecting the growing interest in predictive enzyme engineering. Additionally, enzymes such as TfCut2,63–66 TfCa,67 HiCut,68 and FoCut69 have been explored to broaden the phylogenetic and mechanistic landscape of PET biodegradation. Some studies also explore less-characterized or predicted enzymes derived from metagenomic datasets or de novo structure prediction (e.g., AlphaFold2-based models), reflecting a trend toward integrating computational enzyme design with atomistic modeling workflows.70–72 In terms of substrate complexity, there has been a clear progression from early studies using minimal PET fragments (e.g., MHET, BHET, 2PET)48,73–77 to more realistic representations such as amorphous oligomers of 4PET-9PET,62,64,78,79 and even crystalline polymer chains.4
The catalytic mechanism of IsPETase that promotes the enzymatic degradation of PET is driven by the catalytic triad Ser, His and Asp. Experimental investigations by Han et al.80 and Joo et al.81 proposed a two-step catalytic cycle involving acylation and deacylation, each proceeding through a tetrahedral transition state (TS), as shown in Fig. 1. In this model, the Ser residue attacks the ester bond of the PET dimer substrate, forming an acyl–enzyme intermediate (AEI) and releasing the first product, bis(2-hydroxyethyl) terephthalate (BHET), while His and Asp facilitate proton transfer within the catalytic triad. Subsequent deacylation by water regenerates the active site and releases the second product, which is terephthalic acid (TPA). This mechanism has been also supported by several QM/MM studies.56,76,77,82 However, other computational studies have suggested a more complex mechanism involving two distinct transition states and a short-lived tetrahedral intermediate (TI) in both catalytic steps, raising questions about whether key structures represent true intermediates or transient states.4,44,45,57,73,74,83–85
Tables S3 and S4 present a review of studies that computed key interatomic distances for IsPETase and its engineered variants, ThermoPETase and FAST-PETase, across different temperatures and PET oligomers. For IsPETase, most reported simulations involve oligomers of 2–4 PET monomers at 300–353 K,43,44,53,56,76,82,83,85 whereas studies of ThermoPETase are mainly restricted to tetramers (4-unit oligomers) of PET and those of FAST-PETase to trimer and tetramer PET.53,56 These structural parameters, which include distances within the catalytic triad (Ser160–His237–Asp206)† and substrate-stabilizing contacts involving Tyr87 (ref. 1) and Met161,1 provide valuable insight into temperature-induced conformational rearrangements relevant to the catalytic hydrolysis of PET polymer. At moderate temperatures (300–313 K), triad-associated distances remain short (1.6–3.3 Å) except in da Costa et al.,43 (4.2–8.0 Å), suggesting geometries compatible with catalysis. However, at elevated temperatures (353 K), Berselli et al.,53 reported substantially larger distances, frequently exceeding 9–15 Å, indicating the disruption of the catalytic triad interactions with PET polymer and the subsequent reduction in catalytic efficiency. Temperature appears to have a limited effect on the distance between the hydroxyl oxygen atom of the side chain of Ser160 and the PET carbonyl carbon (∼2.5–3.3 Å) and stabilizing contacts with Tyr87 or Met161 (∼1.8–2.2 Å).56,76,82,83 Notably FAST-PETase exhibits the most robust geometries under temperature stress.56 Interestingly, ThermoPETase, despite its design engineering it for improved stability, demonstrates significant triad disruption at high temperature, sometimes rivaling or exceeding that observed in IsPETase.53 Collectively, these findings underscore the lack of systematic comparisons across enzyme variants at identical PET oligomer size and temperature conditions. To address this, the present study undertakes a systematic temperature-resolved comparison of IsPETase, ThermoPETase, and FAST-PETase in complex with dimeric PET at 300, 313, 323 and 333 K, providing a comprehensive assessment of structural dynamics and enzyme–substrate interactions across physiologically and industrially relevant temperatures. In contrast, previous atomistic studies have mainly focused on either individual PET hydrolases or on multi-enzyme systems examined at a more limited set of temperatures (Tables S1 and S2) and primarily on catalytic-triad-to-PET distances, without a unified comparison of these three widely used PETase variants.
The choice of considering a PET dimer was based on the fact that it represents the minimal oligomer that still contains a single scissile ester bond and reproduces the local chemistry of PET hydrolysis in the active site, while remaining fully compatible with the catalytic mechanism and active-site interactions reported in previous MD and QM/MM studies (Tables S1 and S2). Dimeric PET (or closely related 2MHET/2BHET fragments) has been widely adopted as a model substrate in atomistic simulations of PET hydrolases, where it has been shown to capture the key catalytic triad geometry, hydrogen-bonding network, and transition-state stabilization around the cleaved ester bond.25,45,76,77,82 Comparative simulations using longer oligomers (3-4PET) and amorphous or crystalline 6-9PET chains4,52,53,85,86 indicate that increasing chain length primarily refines distal contacts along the binding pocket and captures polymer packing effects, whereas the identity of the catalytic residues, the scissile bond environment, and the core acylation/deacylation mechanism remain consistent with PET dimer models.44,56,82 The temperatures selected in the current study were 300 K, 313 K, 323 K and 333 K. Temperatures around 300–313 K (27–40 °C) correspond to mesophilic and mildly elevated conditions under which IsPETase and other PET hydrolases have been shown to retain significant activity on PET films and oligomers, whereas temperatures between 323 and 333 K (50–60 °C) several engineered variants, such as ThermoPETase and FAST-PETase, demonstrate enhanced depolymerisation of amorphous and semi-crystalline PET while remaining structurally stable, as reported in previous experimental and computational studies.53 Temperatures substantially above 333–338 K (60–65 °C) would approach or exceed the glass transition temperature of PET (∼343 K) and the onset of pronounced enzyme destabilisation, where changes in polymer crystallinity, surface softening and enzyme unfolding become dominant and thus the polymer can no longer be reliably represented by a solvated PET model.40,52,65,87–90 Though several engineered PET hydrolases, including HotPETase, Combi-PETase, TmFae-PETase and some FAST-PETase variants, are applied experimentally at temperatures approaching or exceeding the glass transition temperature of PET, the goal of the present work is not to reproduce specific industrial process conditions, but to provide a systematic, temperature-resolved comparison of IsPETase, ThermoPETase and FAST-PETase under a common temperature window. Therefore, the PET dimer–enzyme complex is considered only within the 300–333 K range to provide a realistic local description of the PET–enzyme interface, capturing temperature-dependent changes in active-site geometry, hydrogen-bonding patterns and enzyme–substrate packing, while long-range effects associated with polymer crystallinity, surface morphology and multi-chain packing are beyond the scope of the present study.
Fully detailed atomistic simulations are conducted in the present study to capture enzyme–substrate interactions across a physiologically and industrially relevant temperature range (300 K, 313 K, 323 K and 333 K). Through these simulations, the effects of temperature on enzyme–substrate interactions, active site conformational flexibility, and binding stability are systematically examined. Particular attention is given to identifying temperature-induced alterations in the hydrogen-bonding network and the global conformational stability of the enzyme–substrate complexes. The structure of this work is as follows: Section 2 provides a detailed description of the computational methodology and system set up; Section 3 presents the simulation results and their analysis and Section 4 outlines the most important findings of the work and possible future research plans and directions.
2 Materials and methods
2.1 Simulation details
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 550 water molecules modelled with the SPC/E water model. The simulation box was approximately twice the polymer chain's end-to-end distance to prevent overlap between the polymer and its periodic images. Positional restraints were imposed on all nonhydrogen atoms (1000 kJ mol−1 nm−2) of the enzyme–PET complex structures. The systems were then energy minimized with the steepest descent algorithm allowing water molecules and ions to move freely until the maximum force converged to below 20 kJ mol−1 nm−1. Equilibration of the protein–ligand-constrained systems was conducted for 2 ns under NVT ensemble. In the NVT statistical ensemble, the system was gradually heated from 0 to 300 K using the V-rescale thermostat,108 followed by production MD runs conducted under NPT statistical ensemble at 300 K and 1 bar which controlled by the V-rescale thermostat108 and Parrinello–Rahman barostat, for a total of 150 ns.109 The positional restraints were maintained during energy minimization, NVT equilibration, and the first 50 ns of the NPT production runs. During this restrained simulation, only solvent molecules and ions were free to move, allowing proper solvent relaxation. After the 50 ns restrained NPT run, all restraints were released, and the systems were simulated for an additional 100 ns to capture the dynamical behavior of the enzyme–PET interactions. Four different temperatures were investigated: 300 K, 313 K, 323 K, and 333 K. In total, 12 fully detailed atomistic MD simulations were conducted. The LINCS algorithm110 and Particle Mesh Ewald (PME) method111 were used, with an integration step of 1 fs and a cutoff distance of 10 Å. The first 50 ns were conducted with restraints, while the remaining 100 ns were performed without restraints. The same protocol was applied to simulate all enzyme PET-systems.
550 water molecules modelled with the SPC/E water model. The simulation box was approximately twice the polymer chain's end-to-end distance to prevent overlap between the polymer and its periodic images. Positional restraints were imposed on all nonhydrogen atoms (1000 kJ mol−1 nm−2) of the enzyme–PET complex structures. The systems were then energy minimized with the steepest descent algorithm allowing water molecules and ions to move freely until the maximum force converged to below 20 kJ mol−1 nm−1. Equilibration of the protein–ligand-constrained systems was conducted for 2 ns under NVT ensemble. In the NVT statistical ensemble, the system was gradually heated from 0 to 300 K using the V-rescale thermostat,108 followed by production MD runs conducted under NPT statistical ensemble at 300 K and 1 bar which controlled by the V-rescale thermostat108 and Parrinello–Rahman barostat, for a total of 150 ns.109 The positional restraints were maintained during energy minimization, NVT equilibration, and the first 50 ns of the NPT production runs. During this restrained simulation, only solvent molecules and ions were free to move, allowing proper solvent relaxation. After the 50 ns restrained NPT run, all restraints were released, and the systems were simulated for an additional 100 ns to capture the dynamical behavior of the enzyme–PET interactions. Four different temperatures were investigated: 300 K, 313 K, 323 K, and 333 K. In total, 12 fully detailed atomistic MD simulations were conducted. The LINCS algorithm110 and Particle Mesh Ewald (PME) method111 were used, with an integration step of 1 fs and a cutoff distance of 10 Å. The first 50 ns were conducted with restraints, while the remaining 100 ns were performed without restraints. The same protocol was applied to simulate all enzyme PET-systems.2.2 MD analysis
After completing the production MD runs of the 12 simulated systems, post-analysis was conducted on the final 10 ns equilibrated trajectory, during which no positional restraints were applied, allowing the systems to evolve freely under near-physiological conditions. Once equilibration was confirmed, further analysis was conducted, including measurements of catalytic distances and the calculation of binding energies. Plots were created using the matplotlip library in Python.113
 | (1) |
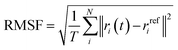 | (2) |
 | (3) |
 | (4) |
 | (5) |
 | (6) |
| ΔGbind = ΔEMM + ΔGsolv − TΔS | (7) |
| ΔEMM = ΔEcovalent + ΔEelec + ΔEvdW | (8) |
The covalent term includes changes in bonds (ΔEbond), angles (ΔEangle), and torsion (ΔEtorsion) energies.128,129
| ΔEcovalent = ΔEbond + ΔEangle + ΔEtorsion | (9) |
| ΔGsolv = ΔGpolar + ΔGnon-polar | (10) |
The polar contribution to the solvation free energy is calculated using the Poisson–Boltzmann implicit solvent model, while the non-polar contribution is calculated based on the SASA.128 The polar solvation component specifies the interaction energy of the solute's charge distribution in the continuum solvent and is found by evaluation of the Poisson–Boltzmann equation (PBE).129 The PBE is based on the fundamental Poisson equation:125,129
| ∇ε(r)∇φ(r) = −4πρ(r) | (11) |
| ∇ε(r)∇φ(r) − εsolvk2φ(r) = −4πρ(r) | (12) |
 | (13) |
 | (14) |
 | (15) |
| ΔGnon-polar = γ × SASA + b | (16) |
3 Results and discussion
3.1 Molecular docking
Molecular docking was first conducted to generate plausible binding poses of the PET dimer in the active sites of IsPETase, ThermoPETase and FAST-PETase. The lowest-energy poses obtained with AutoDock Vina consistently placed the PET dimer in the canonical binding pocket adjacent to the Ser160–His237–Asp206 catalytic triad, in agreement with crystallographic and computational studies of PET hydrolases.80,81 In all three complexes, one terephthalate unit is oriented such that its carbonyl group is directed toward the nucleophilic Ser160, while the aromatic ring lies within van der Waals contact of Tyr87 and Met161, forming a hydrophobic pocket that stabilizes PET chains in PETase enzymes.81 Visual inspection of the docked poses (Fig. 3 and S2) shows that the overall placement of the PET dimer is qualitatively similar across the three enzymes, with only modest differences in tilt and depth of insertion into the pocket. The catalytic Ser side chain is positioned close enough to at least one ester carbonyl of the PET to enable reaction, while the His and Asp residues of the enzymes remain suitably oriented to support proton transfer. This ensures (at the docking level) that all three variants are capable of adopting geometries compatible with the first catalytic step, in line with previous structural analyses.31,33,81 The predicted docking scores fall within a relatively narrow range with −4.91 kcal mol−1 for IsPETase:2PET complex, −4.57 kcal mol−1 for ThermoPETase:2PET complex and −4.74 kcal mol−1 for FAST-PETase:2PET. Docking scoring functions are known to have limited quantitative reliability, and deviations of a few tenths lie well within the intrinsic uncertainty of the method.104 At this stage, the docking fulfilled its primary role in the present work of generating enzyme–substrate complexes that are consistent with experimental structural data, feature catalytically competent arrangements of the triad and PET dimer, and are suitable as initial configurations for subsequent MD simulations. | ||
| Fig. 3 Binding modes of PET dimer complexed with the enzymes IsPETase, ThermoPETase, and FAST-PETase, as determined by molecular docking using AutoDock Vina. | ||
3.2 Analysis of the MD simulations
Across all simulations, backbone RMSD and Rg reach stable plateaus and remain within narrow ranges (∼0.7–1.2 Å for RMSD, 16.7–16.9 Å for Rg; Table 1 and Fig. S3, S4), indicating that none of the complexes undergo global unfolding at the temperatures examined. IsPETase:2PET systematically shows higher RMSD than the engineered variants, in line with its lower thermal robustness reported in previous studies.40,76,82 ThermoPETase:2PET and FAST-PETase:2PET maintain more compact conformations with marginally lower Rg and SASA values than the wild-type enzyme (Table 1 and Fig. S5), reflecting a more compact and tightly packed surface, consistent with earlier findings.31,33 This increased compactness is in line with the design of these variants as thermally stabilized scaffolds and suggests a more persistent binding interface for PET at elevated temperature, as previously reported.47,49
| System | Temp (K) | RMSD (Å) | Rg (Å) |
|---|---|---|---|
| IsPETase:2PET | 300 | 1.10 ± 0.06 | 16.84 ± 0.03 |
| 313 | 1.00 ± 0.05 | 16.93 ± 0.04 | |
| 323 | 1.15 ± 0.05 | 16.93 ± 0.03 | |
| 333 | 1.21 ± 0.07 | 16.91 ± 0.04 | |
| ThermoPETase:2PET | 300 | 0.83 ± 0.06 | 16.75 ± 0.03 |
| 313 | 0.84 ± 0.07 | 16.77 ± 0.04 | |
| 323 | 0.88 ± 0.07 | 16.81 ± 0.04 | |
| 333 | 0.88 ± 0.10 | 16.82 ± 0.04 | |
| FAST-PETase:2PET | 300 | 0.72 ± 0.05 | 16.79 ± 0.03 |
| 313 | 0.84 ± 0.07 | 16.76 ± 0.03 | |
| 323 | 0.83 ± 0.06 | 16.80 ± 0.04 | |
| 333 | 0.77 ± 0.06 | 16.83 ± 0.03 |
Although RMSD and Rg clearly distinguish the global stability of the wild-type and engineered variants, it is essential to recognize that these measurements capture only global structural drift.117 PET hydrolases often rely on subtle, localized rearrangements, particularly within the catalytic triad and substrate-binding loops, that may not significantly alter RMSD or Rg but can strongly influence catalytic competence.130 Thus, while the data in Table 1 confirm that all systems remain structurally intact and that ThermoPETase and FAST-PETase exhibit better thermal robustness, these global descriptors cannot determine whether catalytically relevant geometries are preserved. For that, residue-level flexibility and catalytic distance analyses are required, which are addressed in Sections 3.2.2 and 3.2.3, respectively.
Notably, from the analyses conducted so far neither global rigidity nor residual flexibility could fully explain the catalytic behavior of the enzyme complexes studied. The overall flexibility for all complexes studied here remains broadly stable across the four temperatures examined, with no indication of global unfolding or large-scale destabilization. It is observed that the engineered variants, particularly FAST-PETase, tend to exhibit marginally higher flexibility in the catalytic triad region (especially at high temperatures) compared with the wild-type enzyme, which may allow the active site to adopt a more adaptable catalytic geometry. However, the mutation sites themselves do not display a uniform rigidity that could explain the enhanced thermostability or improved catalytic performance observed experimentally31,33 for these variants. In contrast, the wild-type enzyme shows increased flexibility of substrate-stabilizing residues at elevated temperatures, suggesting that excessive mobility in these regions may hinder catalytic readiness. This is confirmed by the dynamical cross-correlation matrix (DCCM) analysis shown in Fig. S9. At 333 K, low to no correlation is observed between the motions of the mutated residues and the rest of the protein, indicating increased flexibility compared with 300 K, where strong positive correlation reflects the relative stability of these regions. In the engineered enzymes, however, the correlation in the mutated regions increases at 333 K compared with 300 K, indicating enhanced rigidity of these residues, consistent with the intended stabilizing role of the mutations.
Overall, the RMSF analysis demonstrates that while engineered PETases maintain more controlled local fluctuations than the wild-type enzyme, RMSF alone cannot provide a complete mechanistic explanation for their improved catalytic properties. The results underscore the necessity of complementing the RMSF analysis with additional assessments, such as quantifying enzyme–substrate distances (presented in Section 3.2.3) and characterizing hydrogen-bonding patterns (presented in Section 3.2.4) to develop a more comprehensive understanding of the catalytic behavior in PET hydrolases under varying temperature conditions.
Fig. 6 compares the catalytic and stabilising distances in IsPETase, ThermoPETase and FAST-PETase across the simulated temperatures. In IsPETase, catalytically competent Ser–PET and Tyr87–PET distances are sampled most clearly at 323 K, where the nucleophile and oxyanion-stabilising Tyr87 adopt a more compact arrangement around the PET dimer, whereas at 300 K the substrate remains further from Ser160 and the Ser160–His237 separation is larger, indicating a less pre-organised catalytic geometry. Previous QM/MM studies82,83 have reported more compact catalytic distances for IsPETase at 300 K therefore, the present MD results should only be considered as a qualitative indication of temperature-dependent pre-organization. At 333 K, PET–Tyr87 and PET–Met161 distances increase and Ser–PET separations expand, consistent with reduced stabilisation of the substrate and temperature-induced destabilisation of substrate-orienting residues as reported previously for IsPETase.43 In ThermoPETase, catalytic and stabilising distances are systematically shorter than in the wild-type across the temperature range as confirmed from the time evolution of the distance between Ser and the ester bond of PET (Fig. S11) and become most favourable at 333 K, where Tyr87–PET and Met161–PET contacts (d6 = 2.12 Å and d5 = 2.58 Å) remain within hydrogen-bonding distances and Ser160–PET (d3 = 3.70 Å, d4 = 3.15 Å) separations fall in the range typically associated with pre-reactive configurations.48 FAST-PETase consistently exhibits the shortest catalytic distances at all temperatures (Table S7), with particularly favourable geometries at 300–313 K, indicating a highly pre-organised active site even under mild conditions, in agreement with its experimentally reported high activity in this regime.33,53 Unlike the IsPETase:2PET complex, PET in both ThermoPETase:2PET and FAST-PETase:2PET complexes remains stably positioned within the binding pocket at all temperatures examined. This observation is consistent with the distance distribution (Fig. S12) between the hydroxyl oxygen of the catalytic Ser and the ester carbon, which reveals a more favorable catalytic geometry for the engineered enzymes compared to IsPETase across all temperatures, particularly at 333 K, with modal distances of 7.12 Å for the wild-type, 2.97 Å for ThermoPETase, and 3.02 Å for FAST-PETase.
To gain further insight into how closely PET interacts with the enzymes, the radial distribution function g(r) was used to examine the spatial distribution of water molecules around the PET substrate (Fig. S13). At the mild temperature of 300 K, the function shows that water molecules are located closer to PET in the wild-type enzyme systems, with a peak at short distances from PET, indicating that water penetrates between PET and the protein pocket. In contrast, for the engineered enzymes, water molecules are distributed at larger distances from PET, reflecting a more favorable positioning and configuration of the substrate within the active site, where water molecules are largely excluded. At 333 K, the function confirms that PET tends to move away from the wild-type protein, resulting in increased water density near the PET, whereas the distribution around the engineered enzymes remains largely unchanged, consistent with the distances reported in Table S7.
Several studies have employed hybrid QM/MM approaches44,82,83,85 for reporting catalytic distances in PETases which directly sample reactive conformations and often yield shorter Ser160–PET/His237–Ser160 distances than those obtained from classical MD simulations. Consequently, quantitative comparisons with previous works must be made cautiously, particularly given differences in simulated temperature and substrate length (monomer/dimer/tetramer). Nevertheless, the trends observed here, such as the diminished catalytic organization at high temperature in the wild-type and improved geometries in engineered variants, are consistent with existing computational and experimental studies of PETase activity.88,131 Overall, the data presented here suggest that ThermoPETase maintains a more favorable geometry at higher temperatures, consistent with its engineered thermostability while FAST-PETase exhibits consistently optimal catalytic distances across all temperatures, including 300 K, reflecting its enhanced structural pre-organization. However, it is important to emphasize that the geometric proximity analysis conducted here does not guarantee catalytic activity, as electronic reorganization and transition-state stabilization cannot be captured with classical MD.36 Distance-based analyses must therefore be interpreted using structural descriptors rather than mechanistic predictors.
| System | Temp (K) | % of HB occurrence | ||
|---|---|---|---|---|
| Met161–PET | Tyr87–PET | Ser207–PET | ||
| IsPETase:2PET | 300 | |||
| 313 | 36.73 ± 1.08 | |||
| 323 | 35.08 ± 1.07 | 12.39 ± 0.74 | ||
| 333 | ||||
| ThermoPETase:2PET | 300 | 29.34 ± 1.02 | 12.14 ± 0.73 | |
| 313 | 47.63 ± 1.12 | |||
| 323 | 17.29 ± 0.85 | 21.39 ± 0.92 | ||
| 333 | 35.13 ± 1.07 | 56.97 ± 1.10 | ||
| FAST-PETase:2PET | 300 | 53.12 ± 1.12 | 71.41 ± 1.01 | 17.04 ± 0.84 |
| 313 | 45.58 ± 1.11 | 71.61 ± 1.01 | 20.29 ± 0.90 | |
| 323 | 35.38 ± 1.07 | 75.36 ± 0.96 | 19.29 ± 0.88 | |
| 333 | 26.64 ± 0.99 | 57.62 ± 1.10 | ||
Overall, the findings presented here align well with recent mechanistic and structural studies of PET hydrolases. Earlier studies48,74,132 have shown that the backbone amines of Met161 and Tyr87 form persistent HBs with the PET carbonyl oxygen, thereby stabilizing the oxyanion and facilitating catalysis. In particular, umbrella-sampling QM/MM studies confirmed that these HBs are maintained throughout both acylation and deacylation steps, underscoring their mechanistic significance.56,82 Nevertheless, despite the improved HB patterns in the engineered variants, some limitations must be recognized. First, HB occupancy values derived from classical MD provide only a time-averaged, geometric proxy for binding strength and they do not capture full energetics, nor do they ensure that the bound configurations correspond to catalytically competent states. Second, our simulations employ a PET dimer, whereas realistic substrates in biological or industrial settings often involve longer oligomers or even polymer chains which will significantly alter binding dynamics, chain flexibility, and steric constraints therefore altering HB patterns and anchoring stability. In light of these limitations, the HB analysis should be considered as indicative of, but not definitive proof of, catalytic activity. While the high HB occupancy in FAST-PETase strongly supports a stable enzyme–substrate complex, only hybrid methods (e.g., QM/MM free-energy sampling) can establish whether these configurations progress to a transition state and achieve productive catalysis.
 | ||
| Fig. 7 Comparison of the enthalpic energy for the 12 simulation systems (3 enzymes at 4 temperatures) as computed using the PBSA method. | ||
For IsPETase, the most favourable enthalpic contribution is obtained at 323 K (Fig. 7 and Table S8), in qualitative agreement with the structural analysis, which identified this temperature as providing the shortest Ser160–PET and Tyr87–PET distances and the highest Tyr87–PET HB occupancy (Fig. 6 and Table 2). At 323 K, van der Waals interactions dominate the enthalpic term and the polar solvation penalty is reduced relative to 300 K, indicating that PET binding at this temperature requires less desolvation of the binding pocket. A similar predominance of van der Waals contributions in PETase–substrate interactions has been reported in other MM-PBSA studies of PET-binding enzymes.133,134 At 300 K, the less favourable enthalpic value coincides with the larger Ser160–PET and Tyr87/Met161–PET distances and low HB occupancies, indicating suboptimal conformation. At 333 K, the enthalpic term does not improve despite the higher temperature; this is consistent with the observed drift of PET away from the active site and the loss of stabilising HBs. For ThermoPETase, the most favorable enthalpic contribution is found at 333 K (−20.92 kcal mol−1), again in qualitative agreement with the distance and HB analyses. The short Ser160–PET, Tyr87–PET, and Met161–PET distances and the high HB occupancies observed at 333 K (Sections 3.2.3 and 3.2.4) are consistent with this enthalpic minimum, suggesting that substrate packing in the binding pocket is most effective at the highest temperature examined. These findings are in line with experimental and computational evidence indicating that ThermoPETase is designed to maintain or even improve catalytic performance at elevated temperatures relative to the wild-type enzyme.33 In FAST-PETase, the most favorable enthalpic values are obtained at 300 K and 313 K (−23.38 kcal mol−1 and −23.54 kcal mol−1, respectively), which are clearly more stabilizing than those of the other two enzymes at any temperature. In these conditions, van der Waals interactions again dominate the enthalpic term, reflecting tight packing of the PET dimer in the binding pocket. The shortest Ser160–PET and Tyr87/Met161–PET distances and the highest HB occupancies are also observed at these lower temperatures, indicating a strong correspondence between geometric pre-organization and enthalpic stabilization. This is consistent with computational and experimental studies which found that FAST-PETase combines enhanced thermostability with high activity under relatively mild conditions, with an active site that remains pre-organized for PET binding over a broad temperature window.33,56
Although the findings reported here are consistent with the structural and HB analyses presented in previous sections, several methodological limitations must be acknowledged. First, the present MM-PBSA calculations neglect explicit entropic contributions, so the reported values correspond to enthalpic components (ΔHtotal) rather than the total binding free energy (ΔGbind). Entropy can be a substantial factor in enzyme–substrate systems and often leads to partial enthalpy–entropy compensation.135 Second, endpoint methods such as MM-PBSA are known to be sensitive to molecular force-field choice, dielectric constants, and treatment of solvation, and are best employed for relative rather than absolute comparisons.125 Finally, the use of a PET dimer as a model substrate does not fully capture the conformational and enthalpic complexity associated with longer oligomers or semi-crystalline PET surfaces, which have been shown to exhibit different binding characteristics.134 Overall, the MM-PBSA analysis presented here is used only as a qualitative, supportive measure of relative substrate binding stability that complements the geometric and hydrogen-bonding analyses, rather than as a direct indicator of catalytic efficiency or optimal operating temperature. A more complete thermodynamic description would require explicit evaluation of entropic contributions and, ideally, free-energy methods or QM/MM approaches that can more rigorously capture the coupling between binding, conformational changes, and chemical reactivity.
4 Conclusion
In this study, fully detailed atomistic MD simulations were conducted to investigate systematically the temperature-dependent catalytic behavior of three PET-hydrolyzing enzymes: the wild-type PETase (IsPETase) and two engineered variants (ThermoPETase and FAST-PETase) in complex with a PET dimer substrate, across four temperatures (300, 313, 323, and 333 K). Plausible enzyme–substrate complexes were first generated by molecular docking, after which global structural stability of the three PET-hydrolyzing enzymes was evaluated through RMSD, Rg and secondary-structure analysis (DSSP). Local residue-level flexibility was evaluated through RMSF analysis, while catalytic behavior was examined through key geometric distance metrics and hydrogen-bond analysis. Finally, MM-PBSA calculations were employed to estimate enthalpic contributions to binding and to support a comparative thermodynamic interpretation across enzyme variants and temperature conditions.The PET-hydrolyzing enzymes studied here maintained their natural fold state with temperature, while the PET dimer remained in an extended conformation within the enzyme cavity. Both engineered variants displayed significantly improved thermal stability relative to IsPETase, while FAST-PETase presented the greatest stability across all temperatures. Both engineered variants also displayed higher catalytic triad flexibility relative to the wild-type, corresponding to enhanced catalytic behavior. This was further supported by monitoring Tyr87 and Met161, key residues for stabilizing the PET dimer within the enzyme's active-site cavity. In IsPETase, elevated temperature (333 K) increased the flexibility of these residues, leading to disrupted PET positioning and destabilization, with no HB formation observed between Tyr87 or Met161 and the substrate (PET dimer). In contrast, both engineered variants maintained shorter PET–residue distances (∼2.5 Å in ThermoPETase at higher temperatures and ∼2.0 Å in FAST-PETase at lower temperatures), accompanied by a high frequency of HB formation. Importantly, the shortest PET–residue distances and highest HB occupancies correlated with the lowest enthalpic energy at each enzyme's optimal temperature. Based on these data, the potentially optimal temperatures for PET degradation are 323 K for IsPETase, 333 K for ThermoPETase, and 300–313 K for FAST-PETase. In all cases, PET binding is primarily found to be driven by hydrophobic interactions. Overall, these findings provide a complementary perspective to previous atomistic studies of PET hydrolases (Tables S1 and S2), which have typically focused either on individual PET hydrolases at one or two temperatures, or on multi-enzyme comparisons restricted to two temperature points and primarily analyzing catalytic-triad-to-PET distances, none of which has provided a unified, temperature-resolved comparison of the three widely used PET hydrolases examined here.
In future work, a systematic assessment of the effect of PET's degree of polymerization will be conducted for the three PET-hydrolyzing enzymes investigated here. Hybrid QM/MM calculations combined with free-energy sampling will be also employed to quantify acylation and deacylation barriers across temperatures and variants, thereby moving beyond geometric descriptors toward direct evaluation of chemical reactivity and transition-state stabilization.
Author contributions
Conceptualization, D. G. M., A. K., I. L., A. A.; methodology, A. K., D. G. M., H. T.; formal analysis, A. K.; funding: G. M., I. L., A. A.; supervision, D. G. M., H. T., A. A.; writing-original draft preparation, A. K.; writing-review and editing, H. T., D. G. M., C. T. K., I. L., G. M. and A. A. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
No conflict of interest to declare.Abbreviations
| AEI | Acyl–enzyme intermediate |
| BHET | Bis(2-hydroxyethyl) terephthalate |
| DSSP | Dictionary of secondary structure of proteins |
| EG | Ethylene glycol |
| HB | Hydrogen bond |
| MD | Molecular dynamics |
| MHET | Mono(2-hydroxyethyl) terephthalate |
| PAH | Polycyclic aromatic hydrocarbons |
| PET | Polyethylene terephthalate |
| PFAS | Perfluoroalkyl substances |
| Rg | Radius of gyration |
| RMSD | Root mean square deviation |
| RMSF | Root mean square fluctuation |
| SASA | Solvent-accessible surface area |
| TI | Tetrahedral intermediate |
| TS | Transition state |
| TPA | Terephthalic acid |
| QM/MM | Quantum mechanics/molecular mechanics |
Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6ra00343e.Acknowledgements
This work received funding from the Horizon Europe research and innovation program under the Marie Sklodowska-Curie Doctoral Network PlasticUnderground (Grant Agreement No. 101072777) and additional funding from UKRI EPSRC via the Horizon Europe Guarantee Fund (project number EP/X03626X/1). The Asclepios KNIME nodes presented in this work were developed within the European Union's Horizon Europe research and innovation program as part of the SSbD4CheM project (Grant Agreement No. 101138475), advancing digital, in silico decision-support tools that translate molecular-scale simulations into actionable design knowledge for safe and sustainable chemicals and materials design. This work was supported by computing time awarded on the Cyclone Supercomputer of the High-Performance Computing Facility of The Cyprus Institute under production access project ID p197.References
- R. Geyer, J. R. Jambeck and K. L. Law, Production, use, and fate of all plastics ever made, Sci. Adv., 2017, 3(7), e1700782, DOI:10.1126/sciadv.1700782.
- S. S. Ali, T. Elsamahy, E. Koutra, M. Kornaros, M. El-Sheekh, E. A. Abdelkarim, D. Zhu and J. Sun, Degradation of conventional plastic wastes in the environment: a review on current status of knowledge and future perspectives of disposal, Sci. Total Environ., 2021, 771, 144719, DOI:10.1016/j.scitotenv.2020.144719.
- R. W. Chia, J.-Y. Lee, H. Kim and J. Jang, Microplastic pollution in soil and groundwater: a review, Environ. Chem. Lett., 2021, 19(6), 4211–4224, DOI:10.1007/s10311-021-01297-6.
- M. Zheng, Y. Li, W. Dong, Q. Zhang and W. Wang, Hydrolase-Catalyzed Depolymerization Mechanism toward Crystalline and Amorphous Polyethylene Terephthalate, ACS Sustain. Chem. Eng., 2024, 12(27), 10252–10259, DOI:10.1021/acssuschemeng.4c02986.
- J. Qiu, Y. Chen, L. Zhang, J. Wu, X. Zeng, X. Shi, L. Liu and J. Chen, A comprehensive review on enzymatic biodegradation of polyethylene terephthalate, Environ. Res., 2024, 240(Pt 2), 117427, DOI:10.1016/j.envres.2023.117427.
- Y. K. Song, S. H. Hong, M. Jang, G. M. Han, S. W. Jung and W. J. Shim, Combined Effects of UV Exposure Duration and Mechanical Abrasion on Microplastic Fragmentation by Polymer Type, Environ. Sci. Technol., 2017, 51(8), 4368–4376, DOI:10.1021/acs.est.6b06155.
- J. Duan, N. Bolan, Y. Li, S. Ding, T. Atugoda, M. Vithanage, B. Sarkar, D. C. W. Tsang and M. B. Kirkham, Weathering of microplastics and interaction with other coexisting constituents in terrestrial and aquatic environments, Water Res., 2021, 196, 117011, DOI:10.1016/j.watres.2021.117011.
- M. Blasing and W. Amelung, Plastics in soil: analytical methods and possible sources, Sci. Total Environ., 2018, 612, 422–435, DOI:10.1016/j.scitotenv.2017.08.086.
- L. Peng, D. Fu, H. Qi, C. Q. Lan, H. Yu and C. Ge, Micro-and nano-plastics in marine environment: source, distribution and threats—a review, Sci. Total Environ., 2020, 698, 134254, DOI:10.1016/j.scitotenv.2019.134254.
- C. Papavasiliou, D. G. Mintis, A. Tsoumanis, A. Karaoli, I. Lynch, S. Krause, D. D. Varsou, G. Melagraki, M. Kavousanakis and A. Afantitis, MicroPlasticFate web application: multimedia environmental fate modelling of microplastic particles via the Enalos Cloud platform, Bioresour. Technol. Rep., 2025, 102157, DOI:10.1016/j.biteb.2025.102157.
- M. S. Yee, L. W. Hii, C. K. Looi, W. M. Lim, S. F. Wong, Y. Y. Kok, B. K. Tan, C. Y. Wong and C. O. Leong, Impact of Microplastics and Nanoplastics on Human Health, Nanomaterials, 2021, 11(2), 496, DOI:10.3390/nano11020496.
- R. Kumar, C. Manna, S. Padha, A. Verma, P. Sharma, A. Dhar, A. Ghosh and P. Bhattacharya, Micro (nano) plastics pollution and human health: how plastics can induce carcinogenesis to humans?, Chemosphere, 2022, 298, 134267, DOI:10.1016/j.chemosphere.2022.134267.
- M. Ateia, T. Zheng, S. Calace, N. Tharayil, S. Pilla and T. Karanfil, Sorption behavior of real microplastics (MPs): insights for organic micropollutants adsorption on a large set of well-characterized MPs, Sci. Total Environ., 2020, 720, 137634, DOI:10.1016/j.scitotenv.2020.137634.
- L. Sørensen, E. Rogers, D. Altin, I. Salaberria and A. M. Booth, Sorption of PAHs to microplastic and their bioavailability and toxicity to marine copepods under co-exposure conditions, Environ. Pollut., 2020, 258, 113844, DOI:10.1016/j.envpol.2019.113844.
- D. G. Mintis, N. Cheimarios, A. Tsoumanis, A. G. Papadiamantis, N. W. van den Brink, H. J. van Lingen, G. Melagraki, I. Lynch and A. Afantitis, NanoBioAccumulate: modelling the uptake and bioaccumulation of nanomaterials in soil and aquatic invertebrates via the Enalos DIAGONAL Cloud platform, Comput. Struct. Biotechnol. J., 2024, 25, 243–255, DOI:10.1016/j.csbj.2024.09.028.
- D. G. Mintis, D.-D. Varsou, P. D. Kolokathis, A. Tsoumanis, G. Melagraki, J. P. Seif, A. J. del Real, I. Lynch and A. Afantitis, LungDepo: modelling the regional particle deposition in the human lung via the Enalos Cloud platform, Environ. Sci.: Nano, 2025, 12(8), 3921–3937, 10.1039/D5EN00299K.
- D. G. Mintis, C. V. Papavasiliou, D. Dimitra, A. Tsoumanis, G. Melagraki, P. J. Seif, M. Majó, J. A. del Real, T. Serchi, R. Hischier, I. Lynch and A. Afantitis, SimpleBox4Planet: Environmental Fate Modelling of PFAS and Their Alternatives via the Enalos Cloud Platform, RSC Sustain., 2025, 4, 906–927, 10.1039/D5SU00622H.
- K. Ragaert, L. Delva and K. Van Geem, Mechanical and chemical recycling of solid plastic waste, Waste Manag., 2017, 69, 24–58, DOI:10.1016/j.wasman.2017.07.044.
- J. Hopewell, R. Dvorak and E. Kosior, Plastics recycling: challenges and opportunities, Philos. Trans. R. Soc. Lond. B Biol. Sci., 2009, 364(1526), 2115–2126, DOI:10.1098/rstb.2008.0311.
- G. P. Karayannidis and D. S. Achilias, Chemical recycling of poly(ethylene terephthalate), Macromol. Mater. Eng., 2007, 292(2), 128–146, DOI:10.1002/mame.200600341.
- R. P. Magalhaes, J. M. Cunha and S. F. Sousa, Perspectives on the Role of Enzymatic Biocatalysis for the Degradation of Plastic PET, Int. J. Mol. Sci., 2021, 22(20), 11257, DOI:10.3390/ijms222011257.
- S. D. A. Sharuddin, F. Abnisa, W. M. A. W. Daud and M. K. Aroua, A review on pyrolysis of plastic wastes, Energy Convers. Manag., 2016, 115, 308–326, DOI:10.1016/j.enconman.2016.02.037.
- J. Mican, J. Da'san MM, W. Liu, G. Weber, S. Mazurenko, U. T. Bornscheuer, J. Damborsky, R. Wei and D. Bednar, Exploring new galaxies: perspectives on the discovery of novel PET-degrading enzymes, Appl. Catal. B Environ., 2024, 342, 123404, DOI:10.1016/j.apcatb.2023.123404.
- Y. Zhang, J. N. Pedersen, B. E. Eser and Z. Guo, Biodegradation of polyethylene and polystyrene: from microbial deterioration to enzyme discovery, Biotechnol. Adv., 2022, 60, 107991, DOI:10.1016/j.biotechadv.2022.107991.
- N. Wang, Y. Li, M. Zheng, W. Dong, Q. Zhang and W. Wang, BhrPETase catalyzed polyethylene terephthalate depolymerization: a quantum mechanics/molecular mechanics approach, J. Hazard. Mater., 2024, 477, 135414, DOI:10.1016/j.jhazmat.2024.135414.
- R. J. Müller, H. Schrader, J. Profe, K. Dresler and W. D. Deckwer, Enzymatic degradation of poly(ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca, Macromol. Rapid Commun., 2005, 26(17), 1400–1405, DOI:10.1002/marc.200500410.
- S. Sulaiman, S. Yamato, E. Kanaya, J. J. Kim, Y. Koga, K. Takano and S. Kanaya, Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach, Appl. Environ. Microbiol., 2012, 78(5), 1556–1562, DOI:10.1128/AEM.06725-11.
- J. Then, R. Wei, T. Oeser, M. Barth, M. R. Belisario-Ferrari, J. Schmidt and W. Zimmermann, Ca2+ and Mg2+ binding site engineering increases the degradation of polyethylene terephthalate films by polyester hydrolases from Thermobifida fusca, Biotechnol. J., 2015, 10(4), 592–598, DOI:10.1002/biot.201400620.
- S. Yoshida, K. Hiraga, T. Takehana, I. Taniguchi, H. Yamaji, Y. Maeda, K. Toyohara, K. Miyamoto, Y. Kimura and K. Oda, A bacterium that degrades and assimilates poly(ethylene terephthalate), Science, 2016, 351(6278), 1196–1199, DOI:10.1126/science.aad6359.
- K. Grigorakis, C. Ferousi and E. Topakas, Protein Engineering for Industrial Biocatalysis: Principles, Approaches, and Lessons from Engineered PETases, Catalysts, 2025, 15(2), 147, DOI:10.3390/catal15020147.
- H. F. Son, I. J. Cho, S. Joo, H. Seo, H.-Y. Sagong, S. Y. Choi, S. Y. Lee and K.-J. Kim, Rational protein engineering of thermo-stable PETase from Ideonella sakaiensis for highly efficient PET degradation, ACS Catal., 2019, 9(4), 3519–3526, DOI:10.1021/acscatal.9b00568.
- Y. Cui, Y. Chen, X. Liu, S. Dong, Y. e. Tian, Y. Qiao, R. Mitra, J. Han, C. Li and X. Han, et al., Computational Redesign of a PETase for Plastic Biodegradation under Ambient Condition by the GRAPE Strategy, ACS Catal., 2021, 11(3), 1340–1350, DOI:10.1021/acscatal.0c05126.
- H. Lu, D. J. Diaz, N. J. Czarnecki, C. Zhu, W. Kim, R. Shroff, D. J. Acosta, B. R. Alexander, H. O. Cole and Y. Zhang, et al., Machine learning-aided engineering of hydrolases for PET depolymerization, Nature, 2022, 604(7907), 662–667, DOI:10.1038/s41586-022-04599-z.
- H. Yu, Y. Yan, C. Zhang and P. A. Dalby, Two strategies to engineer flexible loops for improved enzyme thermostability, Sci. Rep., 2017, 7(1), 41212, DOI:10.1038/srep41212.
- H. T. H. Nguyen, P. Qi, M. Rostagno, A. Feteha and S. A. Miller, The quest for high glass transition temperature bioplastics, J. Mater. Chem. A, 2018, 6(20), 9298–9331, 10.1039/c8ta00377g.
- S. A. Hollingsworth and R. O. Dror, Molecular dynamics simulation for all, Neuron, 2018, 99(6), 1129–1143, DOI:10.1016/j.neuron.2018.08.011.
- G. Groenhof, Introduction to QM/MM simulations, Methods Mol. Biol., 2013, 924, 43–66, DOI:10.1007/978-1-62703-017-5_3.
- T. Laino, F. Mohamed, A. Laio and M. Parrinello, An Efficient Real Space Multigrid QM/MM Electrostatic Coupling, J. Chem. Theory Comput., 2005, 1(6), 1176–1184, DOI:10.1021/ct050123f.
- V. Tournier, C. M. Topham, A. Gilles, B. David, C. Folgoas, E. Moya-Leclair, E. Kamionka, M. L. Desrousseaux, H. Texier and S. Gavalda, et al., An engineered PET depolymerase to break down and recycle plastic bottles, Nature, 2020, 580(7802), 216–219, DOI:10.1038/s41586-020-2149-4.
- Z. Qu, L. Zhang and Y. Sun, Molecular Insights into the Enhanced Activity and/or Thermostability of PET Hydrolase by D186 Mutations, Molecules, 2024, 29(6), 1338, DOI:10.3390/molecules29061338.
- K. Ding, Z. Levitskaya, B. Sana, R. R. Pasula, S. Kannan, A. Adam, V. V. Sundaravadanam, C. Verma, S. Lim and J. F. Ghadessy, Modulation of PETase active site flexibility and activity on morphologically distinct polyethylene terephthalate substrates by surface charge engineering, Biochem. Eng. J., 2024, 209, 109420, DOI:10.1016/j.bej.2024.109420.
- M. E. Sevilla, M. D. Garcia, Y. Perez-Castillo, V. Armijos-Jaramillo, S. Casado, K. Vizuete, A. Debut and L. Cerda-Mejia, Degradation of PET Bottles by an Engineered Ideonella sakaiensis PETase, Polymers, 2023, 15(7), 1779, DOI:10.3390/polym15071779.
- C. H. S. da Costa, A. M. Dos Santos, C. N. Alves, S. Marti, V. Moliner, K. Santana and J. Lameira, Assessment of the PETase conformational changes induced by poly(ethylene terephthalate) binding, Proteins, 2021, 89(10), 1340–1352, DOI:10.1002/prot.26155.
- S. Boneta, K. Arafet and V. Moliner, QM/MM Study of the Enzymatic Biodegradation Mechanism of Polyethylene Terephthalate, J. Chem. Inf. Model., 2021, 61(6), 3041–3051, DOI:10.1021/acs.jcim.1c00394.
- S. Feng, Y. Yue, M. Zheng, Y. Li, Q. Zhang and W. Wang, Is PETase-and Is MHETase-catalyzed cascade degradation mechanism toward polyethylene terephthalate, ACS Sustain. Chem. Eng., 2021, 9(29), 9823–9832, DOI:10.1021/acssuschemeng.1c02420.
- K. Charupanit, V. Tipmanee, T. Sutthibutpong and P. Limsakul, In silico identification of potential sites for a plastic-degrading enzyme by a reverse screening through the protein sequence space and molecular dynamics simulations, Molecules, 2022, 27(10), 3353, DOI:10.3390/molecules27103353.
- C. Orlando, M. Prejano, N. Russo and T. Marino, On the Role of Temperature in the Depolymerization of PET by FAST-PETase: An Atomistic Point of View on Possible Active Site Pre-Organization and Substrate-Destabilization Effects, Chembiochem, 2023, 24(20), e202300412, DOI:10.1002/cbic.202300412.
- B. Braga, M. Silva and M. Amaral, Variations of the canonical triad of IsPETase: in silico insights with molecular dynamics simulation, Chin. J. Phys., 2023, 84, 282–291, DOI:10.1016/j.cjph.2023.05.022.
- S. Xu, C. Huo and X. Chu, Unraveling the Interplay between Stability and Flexibility in the Design of Polyethylene Terephthalate (PET) Hydrolases, J. Chem. Inf. Model., 2024, 64(19), 7576–7589, DOI:10.1021/acs.jcim.4c00877.
- X. Meng, L. Yang, H. Liu, Q. Li, G. Xu, Y. Zhang, F. Guan, Y. Zhang, W. Zhang, N. Wu and J. Tian, Protein engineering of stable IsPETase for PET plastic degradation by Premuse, Int. J. Biol. Macromol., 2021, 180, 667–676, DOI:10.1016/j.ijbiomac.2021.03.058.
- A. Jackering, F. Gottsch, M. Schaffler, M. Doerr, U. T. Bornscheuer, R. Wei and B. Strodel, From Bulk to Binding: Decoding the Entry of PET into Hydrolase Binding Pockets, JACS Au, 2024, 4(10), 4000–4012, DOI:10.1021/jacsau.4c00718.
- M. Sahihi, P. Fayon, L. Nauton, F. Goujon, J. Devemy, A. Dequidt, P. Hauret and P. Malfreyt, Probing Enzymatic PET Degradation: Molecular Dynamics Analysis of Cutinase Adsorption and Stability, J. Chem. Inf. Model., 2024, 64(10), 4112–4120, DOI:10.1021/acs.jcim.4c00079.
- A. Berselli, M. J. Ramos and M. C. Menziani, Novel Pet-Degrading Enzymes: Structure-Function from a Computational Perspective, Chembiochem, 2021, 22(12), 2032–2050, DOI:10.1002/cbic.202000841.
- Y. Liu, Z. Liu, Z. Guo, T. Yan, C. Jin and J. Wu, Enhancement of the degradation capacity of IsPETase for PET plastic degradation by protein engineering, Sci. Total Environ., 2022, 834, 154947, DOI:10.1016/j.scitotenv.2022.154947.
- S. Schreiber, D. Gercke, F. Lenz and J. Jose, Application of an alchemical free energy method for the prediction of thermostable DuraPETase variants, Appl. Microbiol. Biotechnol., 2024, 108(1), 305, DOI:10.1007/s00253-024-13144-z.
- R. García-Meseguer, E. Ortí, I. Tuñón, J. J. Ruiz-Pernía and J. Aragó, Insights into the enhancement of the poly(ethylene terephthalate) degradation by FAST-PETase from computational modeling, J. Am. Chem. Soc., 2023, 145(35), 19243–19255, DOI:10.1021/jacs.3c04427.
- A. Jackering, M. van der Kamp, B. Strodel and K. Zinovjev, Influence of Wobbling Tryptophan and Mutations on PET Degradation Explored by QM/MM Free Energy Calculations, J. Chem. Inf. Model., 2024, 64(19), 7544–7554, DOI:10.1021/acs.jcim.4c00776.
- L. Pfaff, J. Gao, Z. Li, A. Jackering, G. Weber, J. Mican, Y. Chen, W. Dong, X. Han and C. G. Feiler, et al., Multiple Substrate Binding Mode-Guided Engineering of a Thermophilic PET Hydrolase, ACS Catal., 2022, 12(15), 9790–9800, DOI:10.1021/acscatal.2c02275.
- P. K. Richter, P. Blazquez-Sanchez, Z. Zhao, F. Engelberger, C. Wiebeler, G. Kunze, R. Frank, D. Krinke, E. Frezzotti and Y. Lihanova, et al., Structure and function of the metagenomic plastic-degrading polyester hydrolase PHL7 bound to its product, Nat. Commun., 2023, 14(1), 1905, DOI:10.1038/s41467-023-37415-x.
- Z. Wang, J. Zhang, S. You, R. Su and W. Qi, Energy-guided accumulated mutation strategy achieves a highly efficient polyethylene terephthalate-degrading enzyme, Biochem. Eng. J., 2025, 219, 109708, DOI:10.1016/j.bej.2025.109708.
- Y. Cui, Y. Chen, J. Sun, T. Zhu, H. Pang, C. Li, W.-C. Geng and B. Wu, Computational redesign of a hydrolase for nearly complete PET depolymerization at industrially relevant high-solids loading, Nat. Commun., 2024, 15(1), 1417, DOI:10.1038/s41467-024-45662-9.
- Y. Joho, S. Royan, A. T. Caputo, S. Newton, T. S. Peat, J. Newman, C. Jackson and A. Ardevol, Enhancing PET Degrading Enzymes: A Combinatory Approach, Chembiochem, 2024, 25(10), e202400084, DOI:10.1002/cbic.202400084.
- A. James and S. De, Cation–π and hydrophobic interaction controlled PET recognition in double mutated cutinase–identification of a novel binding subsite for better catalytic activity, RSC Adv., 2022, 12(32), 20563–20577, 10.1039/d2ra03394a.
- X. Q. Chen, Z. Y. Guo, L. Wang, Z. F. Yan, C. X. Jin, Q. S. Huang, D. M. Kong, D. M. Rao and J. Wu, Directional-path modification strategy enhances PET hydrolase catalysis of plastic degradation, J. Hazard. Mater., 2022, 433, 128816, DOI:10.1016/j.jhazmat.2022.128816.
- S. Meng, Z. Li, P. Zhang, F. Contreras, Y. Ji and U. Schwaneberg, Deep learning guided enzyme engineering of Thermobifida fusca cutinase for increased PET depolymerization, Chin. J. Catal., 2023, 50, 229–238, DOI:10.1016/S1872-2067(23)64470-5.
- P. Falkenstein, Z. Zhao, A. Di Pede-Mattatelli, G. Künze, M. Sommer, C. Sonnendecker, W. Zimmermann, F. Colizzi, J. r. Matysik and C. Song, On the binding mode and molecular mechanism of enzymatic polyethylene terephthalate degradation, ACS Catal., 2023, 13(10), 6919–6933, DOI:10.1021/acscatal.3c00259.
- G. von Haugwitz, X. Han, L. Pfaff, Q. Li, H. Wei, J. Gao, K. Methling, Y. Ao, Y. Brack and J. Mican, Structural insights into (tere)phthalate-ester hydrolysis by a carboxylesterase and its role in promoting PET depolymerization, ACS Catal., 2022, 12, 15259–15270 CrossRef PubMed.
- L. Aristizábal-Lanza, S. V. Mankar, C. Tullberg, B. Zhang and J. A. Linares-Pastén, Comparison of the enzymatic depolymerization of polyethylene terephthalate and AkestraTM using Humicola insolens cutinase, Front. Chem. Eng., 2022, 4, 1048744, DOI:10.3389/fceng.2022.1048744.
- F. K. de Oliveira, L. O. Santos and J. G. Buffon, Mechanism of action, sources, and application of peroxidases, Food Res. Int., 2021, 143, 110266, DOI:10.1016/j.foodres.2021.110266.
- A. Li, Y. Sheng, H. Cui, M. Wang, L. Wu, Y. Song, R. Yang, X. Li and H. Huang, Discovery and mechanism-guided engineering of BHET hydrolases for improved PET recycling and upcycling, Nat. Commun., 2023, 14(1), 4169, DOI:10.1038/s41467-023-39929-w.
- T. Mamtimin, X. Ouyang, W. M. Wu, T. Zhou, X. Hou, A. Khan, P. Liu, Y. L. Zhao, H. Tang and C. S. Criddle, et al., Novel Feruloyl Esterase for the Degradation of Polyethylene Terephthalate (PET) Screened from the Gut Microbiome of Plastic-Degrading Mealworms (Tenebrio Molitor Larvae), Environ. Sci. Technol., 2024, 58(40), 17717–17731, DOI:10.1021/acs.est.4c01495.
- Y. Song, A. Li, H. Cui, L. Wu, B. Zhou and X. Li, Ancestral Sequence Reconstruction and Comprehensive Computational Simulations Unmask an Efficient PET Hydrolase with the Wobbled Catalytic Triad, ChemSusChem, 2025, 18(10), e202402614, DOI:10.1002/cssc.202402614.
- A. V. Pinto, P. Ferreira, R. P. Neves, P. A. Fernandes, M. J. Ramos and A. L. Magalhaes, Reaction mechanism of MHETase, a PET degrading enzyme, ACS Catal., 2021, 11(16), 10416–10428, DOI:10.1021/acscatal.1c02444.
- E. Shrimpton-Phoenix, J. B. Mitchell and M. Bühl, Computational Insights into the Catalytic Mechanism of Is-PETase: An Enzyme Capable of Degrading Poly(ethylene) Terephthalate, Chem.–Eur. J., 2022, 28(70), e202201728, DOI:10.1002/chem.202201728.
- Z. Zara, D. Mishra, S. K. Pandey, E. Csefalvay, F. Fadaei, B. Minofar and D. Reha, Surface Interaction of Ionic Liquids: Stabilization of Polyethylene Terephthalate-Degrading Enzymes in Solution, Molecules, 2021, 27(1), 119, DOI:10.3390/molecules27010119.
- C. Jerves, R. P. Neves, S. L. da Silva, M. J. Ramos and P. A. Fernandes, Rate-enhancing PETase mutations determined through DFT/MM molecular dynamics simulations, New J. Chem., 2024, 48(1), 45–54, 10.1039/d3nj04204a.
- T. Burgin, B. C. Pollard, B. C. Knott, H. B. Mayes, M. F. Crowley, J. E. McGeehan, G. T. Beckham and H. L. Woodcock, The reaction mechanism of the Ideonella sakaiensis PETase enzyme, Commun. Chem., 2024, 7(1), 65, DOI:10.1038/s42004-024-01154-x.
- V. Pirillo, M. Orlando, D. Tessaro, L. Pollegioni and G. Molla, An Efficient Protein Evolution Workflow for the Improvement of Bacterial PET Hydrolyzing Enzymes, Int. J. Mol. Sci., 2021, 23(1), 264, DOI:10.3390/ijms23010264.
- X. Guo, D. Xie and Y. Zhou, QM/MM-MD Studies on the Degradation Mechanism and Size Effect of PET by PETase, J. Phys. Chem. B, 2025, 129(22), 5400–5410, DOI:10.1021/acs.jpcb.5c00344.
- X. Han, W. Liu, J. W. Huang, J. Ma, Y. Zheng, T. P. Ko, L. Xu, Y. S. Cheng, C. C. Chen and R. T. Guo, Structural insight into catalytic mechanism of PET hydrolase, Nat. Commun., 2017, 8(1), 2106, DOI:10.1038/s41467-017-02255-z.
- S. Joo, I. J. Cho, H. Seo, H. F. Son, H. Y. Sagong, T. J. Shin, S. Y. Choi, S. Y. Lee and K. J. Kim, Structural insight into molecular mechanism of poly(ethylene terephthalate) degradation, Nat. Commun., 2018, 9(1), 382, DOI:10.1038/s41467-018-02881-1.
- C. Jerves, R. P. P. Neves, M. J. Ramos, S. da Silva and P. A. Fernandes, Reaction Mechanism of the PET Degrading Enzyme PETase Studied with DFT/MM Molecular Dynamics Simulations, ACS Catal., 2021, 11(18), 11626–11638, DOI:10.1021/acscatal.1c03700.
- M. M. Aboelnga and S. Kalyaanamoorthy, QM/MM investigation to identify the hallmarks of superior PET biodegradation activity of PETase over cutinase, ACS Sustain. Chem. Eng., 2022, 10(48), 15857–15868, DOI:10.1021/acssuschemeng.2c04913.
- Y. Wang and K. D. Good, Microplastics and PFAS air-water interaction and deposition, Sci. Total Environ., 2024, 954, 176247, DOI:10.1016/j.scitotenv.2024.176247.
- A. M. Dos Santos, C. H. da Costa, P. H. Silva, M. S. Skaf and J. Lameira, Exploring the Reaction Mechanism of Polyethylene Terephthalate Biodegradation through QM/MM Approach, J. Phys. Chem. B, 2024, 128(31), 7486–7499, DOI:10.1021/acs.jpcb.4c02207.
- E. S. M. Pinto, A. T. Mangini, L. C. C. Novo, F. G. Cavatao, M. J. Krause and M. Dorn, Assessment of Kaistella jeonii esterase conformational dynamics in response to poly(ethylene terephthalate) binding, Curr. Res. Struct. Biol., 2024, 7, 100130 CrossRef PubMed.
- Z. Qu, K. Chen, L. Zhang and Y. Sun, Computation-Based Design of Salt Bridges in PETase for Enhanced Thermostability and Performance for PET Degradation, Chembiochem, 2023, 24(21), e202300373, DOI:10.1002/cbic.202300373.
- L. Shi, P. Liu, Z. Tan, W. Zhao, J. Gao, Q. Gu, H. Ma, H. Liu and L. Zhu, Complete Depolymerization of PET Wastes by an Evolved PET Hydrolase from Directed Evolution, Angew Chem. Int. Ed. Engl., 2023, 62(14), e202218390, DOI:10.1002/anie.202218390.
- J. Zhang, H. Wang, Z. Luo, Z. Yang, Z. Zhang, P. Wang, M. Li, Y. Zhang, Y. Feng, D. Lu and Y. Zhu, Computational design of highly efficient thermostable MHET hydrolases and dual enzyme system for PET recycling, Commun. Biol., 2023, 6(1), 1135, DOI:10.1038/s42003-023-05523-5.
- S. Gao, L. Shi, H. Wei, P. Liu, W. Zhao, L. Gong, Z. Tan, H. Zhai, W. Liu and H. Liu, β-Sheet Engineering of IsPETase for PET Depolymerization, Engineering, 2024, 47, 180–193, DOI:10.1016/j.eng.2024.10.015.
- H. P. Austin, M. D. Allen, B. S. Donohoe, N. A. Rorrer, F. L. Kearns, R. L. Silveira, B. C. Pollard, G. Dominick, R. Duman and K. El Omari, et al., Characterization and engineering of a plastic-degrading aromatic polyesterase, Proc. Natl. Acad. Sci. U. S. A., 2018, 115(19), E4350–E4357, DOI:10.1073/pnas.1718804115.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. Ferrin, UCSF Chimera--a visualization system for exploratory research and analysis, J. Comput. Chem., 2004, 25(13), 1605–1612, DOI:10.1002/jcc.20084.
- E. C. Meng, T. D. Goddard, E. F. Pettersen, G. S. Couch, Z. J. Pearson, J. H. Morris and T. E. Ferrin, UCSF ChimeraX: tools for structure building and analysis, Protein Sci., 2023, 32(11), e4792, DOI:10.1002/pro.4792.
- D. A. Case, H. M. Aktulga, K. Belfon, D. S. Cerutti, G. A. Cisneros, V. W. D. Cruzeiro, N. Forouzesh, T. J. Giese, A. W. Gotz and H. Gohlke, et al., AmberTools, J. Chem. Inf. Model., 2023, 63(20), 6183–6191, DOI:10.1021/acs.jcim.3c01153.
- P. E. Lopes, O. Guvench and A. D. MacKerell Jr, Current status of protein force fields for molecular dynamics simulations, Methods Mol. Biol., 2015, 1215, 47–71, DOI:10.1007/978-1-4939-1465-4_3.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB, J. Chem. Theory Comput., 2015, 11(8), 3696–3713, DOI:10.1021/acs.jctc.5b00255.
- M. D. Hanwell, D. E. Curtis, D. C. Lonie, T. Vandermeersch, E. Zurek and G. R. Hutchison, Avogadro: an advanced semantic chemical editor, visualization, and analysis platform, J. Cheminf., 2012, 4(1), 17, DOI:10.1186/1758-2946-4-17.
- P. S. Nerenberg and T. Head-Gordon, New developments in force fields for biomolecular simulations, Curr. Opin. Struct. Biol., 2018, 49, 129–138, DOI:10.1016/j.sbi.2018.02.002.
- M. W. Schmidt, K. K. Baldridge, J. A. Boatz, S. T. Elbert, M. S. Gordon, J. H. Jensen, S. Koseki, N. Matsunaga, K. A. Nguyen and S. Su, General atomic and molecular electronic structure system, J. Comput. Chem., 1993, 14(11), 1347–1363, DOI:10.21105/joss.01612.
- A. W. Sousa da Silva and W. F. Vranken, ACPYPE-AnteChamber PYthon Parser interfacE, BMC Res. Notes, 2012, 5, 1–8, DOI:10.1186/1756-0500-5-367.
- D. G. Mintis and V. G. Mavrantzas, Effect of pH and Molecular Length on the Structure and Dynamics of Short Poly(acrylic acid) in Dilute Solution: Detailed Molecular Dynamics Study, J. Phys. Chem. B, 2019, 123(19), 4204–4219, DOI:10.1021/acs.jpcb.9b01696.
- D. G. Mintis and V. G. Mavrantzas, Phase boundary and salt partitioning in coacervate complexes formed between poly(acrylic acid) and poly(n,n-dimethylaminoethyl methacrylate) from detailed atomistic simulations combined with free energy perturbation and thermodynamic integration calculations, Macromolecules, 2020, 53(18), 7618–7634, DOI:10.1021/acs.macromol.0c00728.
- G. M. Morris, R. Huey, W. Lindstrom, M. F. Sanner, R. K. Belew, D. S. Goodsell and A. J. Olson, AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility, J. Comput. Chem., 2009, 30(16), 2785–2791, DOI:10.1002/jcc.21256.
- O. Trott and A. J. Olson, AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 2010, 31(2), 455–461, DOI:10.1002/jcc.21334.
- W. L. DeLano, Pymol: an open-source molecular graphics tool, CCP4 Newsletter on Protein Crystallography, 2002, 40(1), 82–92 Search PubMed.
- W. P. Feinstein and M. Brylinski, Calculating an optimal box size for ligand docking and virtual screening against experimental and predicted binding pockets, J. Cheminf., 2015, 7(1), 18, DOI:10.1186/s13321-015-0067-5.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, GROMACS: fast, flexible, and free, J. Comput. Chem., 2005, 26(16), 1701–1718, DOI:10.1002/jcc.20291.
- G. Bussi, D. Donadio and M. Parrinello, Canonical sampling through velocity rescaling, J. Chem. Phys., 2007, 126(1) DOI:10.1063/1.2408420.
- M. Parrinello and A. Rahman, Polymorphic transitions in single crystals: a new molecular dynamics method, J. Appl. Phys., 1981, 52(12), 7182–7190, DOI:10.1063/1.328693.
- B. Hess, H. Bekker, H. J. Berendsen and J. G. Fraaije, LINCS: A linear constraint solver for molecular simulations, J. Comput. Chem., 1997, 18(12), 1463–1472, DOI:10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H.
- T. Darden, D. York and L. Pedersen, Particle mesh Ewald: an N log (N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98, 10089, DOI:10.1063/1.464397.
- K. D. Papavasileiou, A. C. Tsoumanis, P. I. Lagarias, P. D. Kolokathis, N.-M. Koutroumpa, G. Melagraki and A. Afantitis, PFAS-Biomolecule Interactions: Case Study Using Asclepios Nodes and Automated Workflows in KNIME for Drug Discovery and Toxicology, in Computational Toxicology: Methods and Protocols, Springer, 2024, pp. 393–441 Search PubMed.
- J. D. Hunter, Matplotlib: a 2D graphics environment, Comput. Sci. Eng., 2007, 9(03), 90–95, DOI:10.1109/MCSE.2007.55.
- K. Sargsyan, C. Grauffel and C. Lim, How molecular size impacts RMSD applications in molecular dynamics simulations, J. Chem. Theor. Comput., 2017, 13(4), 1518–1524, DOI:10.1021/acs.jctc.7b00028.
- L. Martinez, Automatic identification of mobile and rigid substructures in molecular dynamics simulations and fractional structural fluctuation analysis, PLoS One, 2015, 10(3), e0119264, DOI:10.1371/journal.pone.0119264.
- P. Bauer, B. Hess and E. Lindahl, GROMACS 2022 Manual, 2022, DOI:10.5281/zenodo.6103568.
- M. Hernandez-Rodriguez, M. C. Rosales-Hernandez, J. E. Mendieta-Wejebe, M. Martinez-Archundia and J. C. Basurto, Current Tools and Methods in Molecular Dynamics (MD) Simulations for Drug Design, Curr. Med. Chem., 2016, 23(34), 3909–3924, DOI:10.2174/0929867323666160530144742.
- D. Lu, Y. Chen, S. Jin, Q. Wu, J. Wu, J. Liu, F. Wang, L. Deng and K. Nie, The evolution of cutinase Est1 based on the clustering strategy and its application for commercial PET bottles degradation, J. Environ. Manage., 2024, 368, 122217, DOI:10.1016/j.jenvman.2024.122217.
- D. G. Mintis, A. Chasapi, K. Poulas, G. Lagoumintzis and C. T. Chasapis, Assessing the direct binding of ark-like E3 RING ligases to ubiquitin and its implication on their protein interaction network, Molecules, 2020, 25(20), 4787, DOI:10.3390/molecules25204787.
- D. G. Mintis, M. Dompe, M. Kamperman and V. G. Mavrantzas, Effect of Polymer Concentration on the Structure and Dynamics of Short Poly(N,N-dimethylaminoethyl methacrylate) in Aqueous Solution: A Combined Experimental and Molecular Dynamics Study, J. Phys. Chem. B, 2020, 124(1), 240–252, DOI:10.1021/acs.jpcb.9b08966.
- W. Kabsch and C. Sander, Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features, Biopolymers, 1983, 22(12), 2577–2637, DOI:10.1002/bip.360221211.
- S. Ausaf Ali, M. Imtaiyaz Hassan, A. Islam and F. Ahmad, A review of methods available to estimate solvent-accessible surface areas of soluble proteins in the folded and unfolded states, Curr. Protein Pept. Sci., 2014, 15(5), 456–476, DOI:10.2174/1389203715666140327114232.
- B. Lee and F. M. Richards, The interpretation of protein structures: estimation of static accessibility, J. Mol. Biol., 1971, 55(3), 379–400, DOI:10.1016/0022-2836(71)90324-x.
- D. G. Mintis, M. Dompe, P. D. Kolokathis, J. van der Gucht, A. Afantitis and V. G. Mavrantzas, Effect of pH, Temperature, Molecular Weight and Salt Concentration on the Structure and Hydration of Short Poly(N,N-dimethylaminoethyl methacrylate) Chains in Dilute Aqueous Solutions: A Combined Experimental and Molecular Dynamics Study, Polymers, 2025, 17(16), 2189, DOI:10.3390/polym17162189.
- C. Wang, D. A. Greene, L. Xiao, R. Qi and R. Luo, Recent developments and applications of the MMPBSA method, Front. Mol. Biosci., 2018, 4, 87, DOI:10.3389/fmolb.2017.00087.
- B. R. Miller III, T. D. McGee Jr, J. M. Swails, N. Homeyer, H. Gohlke and A. E. Roitberg, MMPBSA. py: an efficient program for end-state free energy calculations, J. Chem. Theor. Comput., 2012, 8(9), 3314–3321, DOI:10.1021/ct300418h.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, gmx_MMPBSA: a new tool to perform end-state free energy calculations with GROMACS, J. Chem. Theor. Comput., 2021, 17(10), 6281–6291, DOI:10.1021/acs.jctc.1c00645.
- G. Poli, C. Granchi, F. Rizzolio and T. Tuccinardi, Application of MM-PBSA Methods in Virtual Screening, Molecules, 2020, 25(8), 1971, DOI:10.3390/molecules25081971.
- E. King, E. Aitchison, H. Li and R. Luo, Recent Developments in Free Energy Calculations for Drug Discovery, Front. Mol. Biosci., 2021, 8, 712085, DOI:10.3389/fmolb.2021.712085.
- Y. W. Dong, M. L. Liao, X. L. Meng and G. N. Somero, Structural flexibility and protein adaptation to temperature: molecular dynamics analysis of malate dehydrogenases of marine molluscs, Proc. Natl. Acad. Sci. U. S. A., 2018, 115(6), 1274–1279, DOI:10.1073/pnas.1718910115.
- T. Fecker, P. Galaz-Davison, F. Engelberger, Y. Narui, M. Sotomayor, L. P. Parra and C. A. Ramirez-Sarmiento, Active Site Flexibility as a Hallmark for Efficient PET Degradation by I. sakaiensis PETase, Biophys. J., 2018, 114(6), 1302–1312, DOI:10.1016/j.bpj.2018.02.005.
- A. James, A. Bhasi and S. De, Bridging the Gap in the Structure-Function Paradigm of Enzymatic PET Degradation-Aromatic Residue Driven Balanced Interactions with Catalytic and Anchoring Subsite, Chembiochem, 2024, 25(21), e202400555, DOI:10.1002/cbic.202400555.
- A. Karaoli, H. Tzoupis, K. D. Papavasileiou, A. G. Papadiamantis, D. G. Mintis, C. T. Kiranoudis, I. Lynch, G. Melagraki and A. Afantitis, Atomistic-Level Insights into the Role of Mutations in the Engineering of PET Hydrolases: A Systematic Review, Int. J. Mol. Sci., 2025, 26(16), 7682, DOI:10.3390/ijms26167682.
- X. Guo, Y. Jiang, D. Xie and Y. Zhou, Computational investigation on the binding modes of PET polymer to PETase, J. Biomol. Struct. Dyn., 2024, 42(13), 6842–6849, DOI:10.1080/07391102.2023.2240893.
- J. D. Chodera and D. L. Mobley, Entropy-enthalpy compensation: role and ramifications in biomolecular ligand recognition and design, Annu. Rev. Biophys., 2013, 42(1), 121–142, DOI:10.1146/annurev-biophys-083012-130318.
Footnote |
| † Residue numbering is the same for the three enzymes and follows the PDB files from the Protein Data Bank (PDB). |
| This journal is © The Royal Society of Chemistry 2026 |





