Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Targeting cell cycle and apoptotic pathways with newly synthesized diselenide-linked imidazolone analogues with strong CDK6-targeting potential
Marwa Abdel-Motaal†
ab,
Saad Shaaban† *c,
Samia S. Hawas†d,
Asma M. Elsharife,
Marwa Sharakyf,
Fatema S. Alatawig,
Mohamed E. Eissah,
Arwa Omar Al Khatibi,
Hany M. Abd El-Lateefc,
Medhat Asemj and
Ahmed A. Al-Karmalawy
*c,
Samia S. Hawas†d,
Asma M. Elsharife,
Marwa Sharakyf,
Fatema S. Alatawig,
Mohamed E. Eissah,
Arwa Omar Al Khatibi,
Hany M. Abd El-Lateefc,
Medhat Asemj and
Ahmed A. Al-Karmalawy *dk
*dk
aDepartment of Chemistry, College of Science, Qassim University, Buraidah 51452, Qassim, Saudi Arabia
bOrganic Chemistry Division, Department of Chemistry, College of Science, Mansoura University, Mansoura, Egypt
cDepartment of Chemistry, College of Science, King Faisal University, Al-Ahsa 31982, Saudi Arabia. E-mail: sibrahim@kfu.edu.sa
dDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy, Horus University-Egypt, New Damietta 34518, Egypt. E-mail: akarmalawy@horus.edu.eg
eDepartment of Chemistry, College of Science, Imam Abdulrahman Bin Faisal University, Dammam 31441, Saudi Arabia
fCancer Biology Department, Pharmacology Unit, National Cancer Institute (NCI), Cairo University, Cairo, Egypt
gDepartment of Biochemistry, Faculty of Science, University of Tabuk, Tabuk, Saudi Arabia
hDepartment of Chemistry, College of Science, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh 11623, Saudi Arabia
iFaculty of Pharmacy, Al-Ahliyya Amman University, Amman, Jordan
jDepartment of Civil Engineering, College of Engineering and Information Technology, Onaizah Colleges, Qassim 56447, Saudi Arabia
kDepartment of Pharmaceutical Chemistry, College of Pharmacy, The University of Mashreq, Baghdad 10023, Iraq
First published on 4th February 2026
Abstract
A novel panel of diselenide-linked imidazolone derivatives was synthesized and biologically profiled, revealing a promising new chemotype with broad-spectrum anticancer activity. Among the series, compounds 6b, 6d, and 6g demonstrated exceptional growth-inhibitory (GI) potency, achieving GI% values of 80.32%, 79.24%, and 86.40%, respectively—substantially outperforming doxorubicin (61.49%). Notably, 6g emerged as the lead candidate, exhibiting robust cytotoxicity across diverse cancer models with IC50 values of 6.49 µM (PC3), 6.58 µM (MCF7), 5.38 µM (A549), and 7.25 µM (HCT116). Mechanistic studies in A549 cells indicated that 6g simultaneously modulates multiple oncogenic pathways: it markedly downregulated CDK2, CDK4, and CDK6 (1.57–4.12 fold), while upregulating caspase-3, caspase-8, and caspase-9 (1.60–1.64 fold), collectively supporting its dual action on cell-cycle blockade and apoptotic activation. Furthermore, a 1.68-fold reduction in VEGFR-2 expression underscores its additional anti-angiogenic potential. Flow cytometry corroborated these findings, revealing a dramatic S-phase arrest, with the S-phase population rising from 4.61% to 42.09% upon treatment. Several other analogues, including 6d, 6e, 6i, and 6j, also displayed potent cytotoxicity (IC50 < 10 µM), highlighting the broader therapeutic relevance of this scaffold. Collectively, these data position 6g as a compelling multi-target anticancer lead that integrates apoptosis induction, cell-cycle regulation, and angiogenesis suppression—supporting its potential for development as a next-generation broad-spectrum anticancer agent.
1. Introduction
Cancer remains one of the leading causes of morbidity and mortality worldwide, characterized by uncontrolled cellular proliferation, evasion of apoptosis, and the ability to invade surrounding tissues.1–4 Among the various types, lung cancer continues to pose a major global health challenge, with non-small cell lung cancer (NSCLC) accounting for nearly 85% of all cases.5,6 The A549 cell line, derived from human alveolar adenocarcinoma, is widely used as a representative in vitro model for studying NSCLC biology and evaluating potential anticancer agents.5,7A hallmark of malignant transformation is the loss of normal cell-cycle regulation, which allows cancer cells to proliferate indefinitely.8,9 The progression of the cell cycle is tightly controlled by a network of cyclins and cyclin-dependent kinases (CDKs).10,11 Among these, cyclin-dependent kinase 6 (CDK6) plays a central role in driving the transition from the G1 to S phase by phosphorylating the retinoblastoma (Rb) protein, leading to the release of E2F transcription factors and activation of genes required for DNA synthesis.12,13 Overexpression or dysregulation of CDK6 has been documented in several human malignancies, including lung, breast, and hematologic cancers, making it an attractive therapeutic target.14,15
Moreover, cell-cycle arrest often triggers apoptosis, a programmed cell-death mechanism essential for eliminating damaged or abnormal cells.16–18 Apoptosis involves a cascade of molecular events regulated by pro- and anti-apoptotic proteins such as Bax, Bcl-2, and caspase-3, which serve as key biomarkers for evaluating the apoptotic potential of anticancer agents.19,20 In particular, a decrease in Bcl-2 levels coupled with increased Bax expression and activation of caspase-3 is indicative of apoptosis induction.21,22
Recent advances in targeted therapy have demonstrated that selective CDK4/6 inhibitors, such as palbociclib, can induce cell-cycle arrest and apoptosis in a variety of tumor models, including A549 cells.23,24 This underscores the therapeutic significance of targeting CDK6 to restore normal cell-cycle control and promote apoptotic cell death in cancerous cells.25,26 Therefore, designing novel compounds that interfere with CDK6 function represents a promising strategy for the development of effective anticancer agents against NSCLC and other CDK6-driven malignancies.27
1.1. Rationale design
We designed and synthesized a novel series of diselenide-linked imidazolone derivatives that modulate CDK6-associated cell-cycle signaling. The design concept was derived from the co-crystallized ligand palbociclib, which stabilizes the CDK6 receptor (PDB ID: 5L2I) through a network of crucial interactions. Specifically, palbociclib forms two key hinge-region hydrogen bonds with Val101 and Asp163, in addition to a polar contact with Glu99, which collectively anchor the inhibitor within the ATP-binding cleft. Furthermore, multiple hydrophobic residues, including Val27, Val77, Leu152, Phe98, Ala41, and Ala162, surround the aromatic scaffold, forming a nonpolar pocket that enhances ligand accommodation and stability (Fig. 1). | ||
| Fig. 1 Representation of newly synthesized diselenide-linked imidazolone candidates as potential CDK6 suppression and cell-cycle disruption anticancer agents. | ||
Based on these binding insights, the newly designed scaffold was tailored to preserve the essential pharmacophoric features of palbociclib. The amide carbonyl group and the imidazolone nitrogen within the newly designed compounds can serve as hydrogen bond acceptor and donor sites, respectively, allowing the formation of interactions analogous to those observed between palbociclib and the hinge residues.
The incorporation of a Se–Se (diselenide) linker introduces conformational rigidity and potential redox responsiveness, which may contribute to enhanced interaction stability and improved cellular uptake. Meanwhile, the terminal aromatic or heteroaromatic substituents (Ar) were designed to fit into the hydrophobic pocket, promoting favorable π–alkyl and van der Waals interactions that complement the polar hinge contacts.
Collectively, these structural features were strategically integrated to mirror the ATP-competitive binding mode of palbociclib, while providing new opportunities to optimize structure–activity relationships (SAR) through variation in the aromatic substituents. Hence, this rational design approach combines palbociclib's hinge-binding motif with a diselenide-linked bis-imidazolone framework, aiming to yield novel analogues with improved stability and potent anticancer efficacy against CDK6-driven malignancies.
2. Results and discussion
2.1. Chemistry
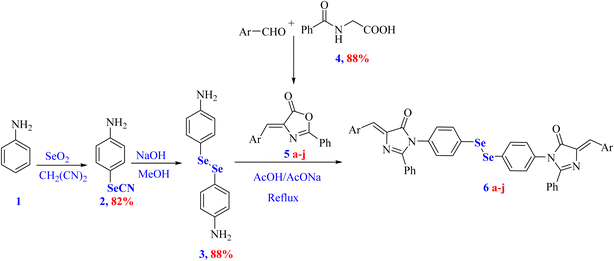
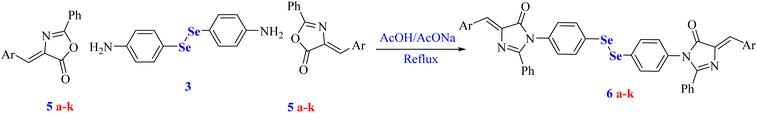
Heterocyclic scaffolds remain indispensable in medicinal chemistry due to their broad synthetic versatility and significant therapeutic relevance.28,29 Accordingly, our study reports the synthesis of a new series of organoselenium-based imidazolones 6a–j starting from hippuric acid (4), various 4-arylidene oxazolones 6a–j with 4,4′-diselenediyldianiline (3) (Scheme 1). In modern medicinal chemistry and advanced drug discovery, the integration of multiple bioactive pharmacophores into a single molecular framework represents a promising approach.30,31 Such a strategy is frequently employed to develop more potent candidates with improved biological efficacy, enhanced selectivity, thereby overcoming limitations related to adverse side effects and drug resistance.In this context, we envisage designing and synthesizing a series of novel diselenide-based bis-arylidene-4-imidazolone hybrids that incorporate both a 4-imidazolone moiety and a diselenide linkage.32 Organic diselenides have garnered considerable attention in recent years due to their promising pharmacological properties, including antioxidant, antibacterial, and anti-inflammatory properties.32–34 Despite this promise, further development is often limited by the use of costly and hazardous OSe reagents, demanding reaction conditions, and limited functional-group tolerance.35,36 Therefore, there is continuing interest in simple synthetic strategies that employ stable, non-toxic OSe sources compatible with diverse substrates.33,34,37,38 In this regard, diaryl diselenides represent valuable precursors for numerous OSe frameworks.39 Their high stability facilitates storage and enables efficient optimization of new transformations, and they are commonly used to access a variety of selenium-containing compounds, including organoselenides, selenocyanates, and selenonic acids.40
Meanwhile, imidazolones constitute a privileged class of organic heterocycles, found in plants and other living organisms, and they are widely known for their broad spectrum of pharmacological activities.41 These scaffolds are also an integral part of many nucleotides (e.g., purine), and several imidazole-based drugs (e.g., fadrozole, dacarbazine, quizartinib, indimitecan, tipifarnib, nilotinib, and ponatinib) are already used for the treatment of a wide range of cancers, including breast cancer, melanoma, lymphomas, and leukemias.42,43
Accordingly, the synthesis of diselenide-based bis-arylidene-4-imidazolone conjugates 6a–j is described herein, as illustrated in Scheme 1.
In the initial step, hippuric acid (4) was allowed to react with various aryl aldehydes in the presence of sodium acetate and acetic anhydride to furnish the key intermediate 4-arylidene oxazolones 5a–j. Subsequently, an acetic acid suspension of appropriate 4-arylidene oxazolones 5a–j was refluxed with 4,4′-diselenediyldianiline (3) in the presence of freshly added sodium acetate, afforded the target compounds 6a–j. The yields of the final compounds were high, ranging from 63–95%, as shown in Scheme 1. The most evidence for the formation of final imidazolone targets 6a–j was their IR spectra, which affirms the disappearance of cyclic ester absorption peaks present in oxazolone structures.
The IR spectrum of 6a showed a carbonyl stretching band at 1715 cm−1 for the lactam, along with the disappearance of the lactone carbonyl band of the corresponding oxazolone previously observed at 1794 cm−1. In addition, the absorption bands of the amino groups of 4,4′-diselenediyldianiline (3) at 3413 and 3404 cm−1 were no longer present.
Furthermore, we observed a characteristic band at 1639, 1596 cm−1 corresponding to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N and C
N and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C of arylidene groups, respectively. Similarly, IR spectra of imidazolones 6b–6j exhibit sharp stretching frequencies of the lactam C
C of arylidene groups, respectively. Similarly, IR spectra of imidazolones 6b–6j exhibit sharp stretching frequencies of the lactam C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group in the range 1690–1730 cm−1, and this differs from the corresponding oxazolones, which show their lactone carbonyl stretching bands in the range (1780–1796 cm−1), which confirms the formation of imidazolone moieties.
O group in the range 1690–1730 cm−1, and this differs from the corresponding oxazolones, which show their lactone carbonyl stretching bands in the range (1780–1796 cm−1), which confirms the formation of imidazolone moieties.
Moreover, the other bands attributed to some function groups exist, such as characteristic stretching bands at (1651–1632 cm−1) for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N groups, and the exocyclic C
N groups, and the exocyclic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C groups appeared at (1596–1515 cm−1). Additionally, the IR spectra show another stretching frequency characterized for each target, such as (827.9 cm−1) attributable to C–Cl for 6b, C–NO2 appeared at (1484, 1396.9 cm−1) for 6c, and (1489.9–1371.3) for 6d. Also, the NH group of the phenothiazine moiety of the compound 6i displayed an absorption band at 3330 cm−1. 1H-NMR and 13C-NMR spectra of the synthesized imidazolones give further support for structural elucidation. Their 1H-NMR spectra confirmed that most peaks are located in the aromatic region with complex multiplet signals at δ 6.9–8.9 ppm, and the difference between oxazolones 5a–j and imidazolones 6a–j is the integration of the signals in the aromatic region beside the presence of singlet signals in the range of δ 8.2–9.1 ppm due to the exocyclic –CH
C groups appeared at (1596–1515 cm−1). Additionally, the IR spectra show another stretching frequency characterized for each target, such as (827.9 cm−1) attributable to C–Cl for 6b, C–NO2 appeared at (1484, 1396.9 cm−1) for 6c, and (1489.9–1371.3) for 6d. Also, the NH group of the phenothiazine moiety of the compound 6i displayed an absorption band at 3330 cm−1. 1H-NMR and 13C-NMR spectra of the synthesized imidazolones give further support for structural elucidation. Their 1H-NMR spectra confirmed that most peaks are located in the aromatic region with complex multiplet signals at δ 6.9–8.9 ppm, and the difference between oxazolones 5a–j and imidazolones 6a–j is the integration of the signals in the aromatic region beside the presence of singlet signals in the range of δ 8.2–9.1 ppm due to the exocyclic –CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C protons, which confirms the formation of the target compounds by condensation of the oxazolones with the 4,4′-diselenediyldianiline supported with the disappearance of the –NH2 proton signals of the diselenide compound 3. On the other hand, the 1H-NMR spectrum of the formed imidazolones showed characteristic signals according to the condensed aromatic aldehyde fragment. Trimethoxy protons in the case of compound 6e were observed as two singlet signals at δ 3.7 ppm (2-OCH3) and 3.8 ppm (4-OCH3), which revealed the formation of the bis-arylidene–imidazolone derivatives. Further, we detected a singlet signal at δ 7.2 ppm for compound 6f belongs to (2O–CH2–O) protons. The two NH protons of the phenothiazine fragment for compound 6i were observed at δ 9.6 ppm. Besides, a singlet signal due to two acetyl protons for compound 6j was displayed at δ 2.09 ppm, which indicates the occurrence of acetylation on the –NH group of the indole fragment after imidazolone cyclization. 13C-NMR spectra of the formed structures proved, besides the aromatic signals' presence of carbonyl amide for all targets, approaching δ 169 ppm. Additionally, aliphatic carbons of some compounds were observed, including two characteristic signals at δ 65.3 ppm due to the four methoxy carbons and 60.7 ppm due to the two methoxy carbons for compound 6e. Also, it is observed that there is a specific signal at δ 99.06 ppm owing to the methylene carbon of the dioxolo group for compound 6f. Finally, the methyl carbon of the acetyl group for compound 6i was observed at δ 24.5 ppm, which gave good evidence for the acetylation of the –NH group (Table 1).
C protons, which confirms the formation of the target compounds by condensation of the oxazolones with the 4,4′-diselenediyldianiline supported with the disappearance of the –NH2 proton signals of the diselenide compound 3. On the other hand, the 1H-NMR spectrum of the formed imidazolones showed characteristic signals according to the condensed aromatic aldehyde fragment. Trimethoxy protons in the case of compound 6e were observed as two singlet signals at δ 3.7 ppm (2-OCH3) and 3.8 ppm (4-OCH3), which revealed the formation of the bis-arylidene–imidazolone derivatives. Further, we detected a singlet signal at δ 7.2 ppm for compound 6f belongs to (2O–CH2–O) protons. The two NH protons of the phenothiazine fragment for compound 6i were observed at δ 9.6 ppm. Besides, a singlet signal due to two acetyl protons for compound 6j was displayed at δ 2.09 ppm, which indicates the occurrence of acetylation on the –NH group of the indole fragment after imidazolone cyclization. 13C-NMR spectra of the formed structures proved, besides the aromatic signals' presence of carbonyl amide for all targets, approaching δ 169 ppm. Additionally, aliphatic carbons of some compounds were observed, including two characteristic signals at δ 65.3 ppm due to the four methoxy carbons and 60.7 ppm due to the two methoxy carbons for compound 6e. Also, it is observed that there is a specific signal at δ 99.06 ppm owing to the methylene carbon of the dioxolo group for compound 6f. Finally, the methyl carbon of the acetyl group for compound 6i was observed at δ 24.5 ppm, which gave good evidence for the acetylation of the –NH group (Table 1).
2.2. Biological assays
| Cell line/comp. | 6a | 6b | 6c | 6d | 6e | 6f | 6g | 6h | 6i | 6j | DOX | Average |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| HCT116 | 73.06 | 86.12 | 71.14 | 74.27 | 73.09 | 48.00 | 81.57 | 74.41 | 81.13 | 78.15 | 64.36 | 73.72 |
| MCF7 | 73.69 | 81.26 | 68.21 | 80.33 | 76.69 | 48.23 | 91.10 | 72.07 | 71.4 | 76.73 | 58.65 | 73.31 |
| HuH7 | 64.97 | 78.91 | 74.86 | 64.81 | 59.31 | 9.91 | 83.17 | 72.62 | 76.04 | 60.22 | 68.00 | 65.16 |
| PC3 | 90.45 | 80.39 | 83.55 | 87.56 | 87.95 | 56.03 | 89.58 | 83.41 | 90.51 | 85.11 | 68.92 | 82.13 |
| A549 | 89.93 | 89.02 | 85.17 | 90.10 | 83.46 | 52.02 | 92.11 | 88.31 | 90.77 | 87.56 | 39.14 | 81.18 |
| MG63 | 68.83 | 66.19 | 73.47 | 78.34 | 47.34 | 9.84 | 80.84 | 60.16 | 61.04 | 79.59 | 69.86 | 63.23 |
| Average | 76.82 | 80.32 | 76.07 | 79.24 | 71.31 | 37.34 | 86.40 | 75.16 | 78.48 | 77.89 | 61.49 | |
| HSF | 69.98 | 74.54 | 61.71 | 76.41 | 61.65 | 5.93 | 74.42 | 68.00 | 68.22 | 67.49 | 29.57 | 59.81 |
A detailed analysis of the structural features of the synthesized diselenide–imidazolone derivatives revealed that their cytotoxic activity is strongly governed by both the electronic nature and steric environment of the substituents on the two aromatic rings attached to the imidazolone core. Interestingly, both electron-donating and electron-withdrawing groups contributed to pronounced cytotoxicity, albeit likely through distinct mechanisms. Electron-donating substituents tend to enhance lipophilicity and facilitate cell membrane permeation, whereas electron-withdrawing substituents increase the electrophilicity of the Se–C bond, thereby intensifying oxidative stress or enabling enzyme inhibition in cancer cells.
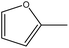
Among all the tested analogues, compound 6g, featuring two furan rings symmetrically positioned on both sides of the molecule, emerged as the most potent member of the series. This unique configuration may favor π–π stacking interactions and optimize binding affinity toward intracellular targets, leading to strong inhibition across multiple cancer cell lines.
Compounds containing electron-withdrawing substituents, such as –Cl or –NO2 (notably in 6b, 6c, and 6d), also demonstrated remarkable inhibitory effects, particularly against HCT116, MCF7, and A549 cell lines. The increased electrophilicity imparted by these groups may facilitate redox modulation and trigger oxidative damage in malignant cells, accounting for their strong cytotoxic responses.
In contrast, unsubstituted derivatives such as 6a and 6h exhibited moderate activity. This reduction in activity likely arises from bulky structural features that restrict optimal target interaction and impede cell penetration.
Notably, 6g displayed moderate cytotoxicity against normal HSF cells (GI% = 74.42%), indicating some selectivity toward cancer cells. Similarly, 6b and 6d exhibited GI% values of 74.54% and 76.41% in HSF cells, respectively, suggesting lower selectivity despite strong cancer cell inhibition. In contrast, compounds 6a, 6c, 6e, and 6h–6j showed moderate cytotoxicity in both cancer and normal cells, while the sterically hindered 6f displayed the lowest activity (average GI% = 37.34% in cancer cells and 5.93% in HSF cells), reflecting reduced cell penetration and target interaction. The structure–activity relationship is illustrated in Fig. 2.
 | ||
| Fig. 2 Structure–activity relationship of the newly synthesized diselenide-linked imidazolone derivatives. | ||
Selenium-containing compounds are known to exert cytotoxicity through redox-related mechanisms, including induction of reactive oxygen species (ROS). While direct ROS measurements were not performed in this study, the observed cytotoxicity patterns suggest that redox modulation may contribute to the antiproliferative effects of the synthesized diselenide–imidazolone derivatives. To evaluate selectivity, we calculated a selectivity index (SI = GI% in normal HSF/average GI% in cancer cells) for each compound (Table 3). Compound 6g demonstrated high potency across cancer cell lines (average GI% = 86.40%) with moderate normal cell toxicity (GI% HSF = 74.42%, SI = 0.86), indicating a favorable therapeutic window. In contrast, compound 6f exhibited very low activity in both cancer and normal cells (SI = 0.16), while DOX showed lower potency with a higher relative selectivity (SI = 0.48).
| Compound | Average GI% cancer | GI% HSF | aSI (HSF/cancer) |
|---|---|---|---|
| a SI values closer to 1 indicate lower selectivity, whereas lower SI values indicate higher selectivity toward cancer cells. | |||
| 6a | 76.82 | 69.98 | 0.91 |
| 6b | 80.32 | 74.54 | 0.93 |
| 6c | 76.07 | 61.71 | 0.81 |
| 6d | 79.24 | 76.41 | 0.96 |
| 6e | 71.31 | 61.65 | 0.86 |
| 6f | 37.34 | 5.93 | 0.16 |
| 6g | 86.40 | 74.42 | 0.86 |
| 6h | 75.16 | 68.00 | 0.90 |
| 6i | 78.48 | 68.22 | 0.87 |
| 6j | 77.89 | 67.49 | 0.87 |
| DOX | 61.49 | 29.57 | 0.48 |
Overall, these results suggest that structural modifications can modulate both potency and selectivity. Importantly, despite the known redox-related toxicity of selenium compounds, the moderate cytotoxicity toward normal HSF cells observed for the most active analogues, particularly 6g, supports their potential for further development as anticancer agents, while highlighting the need for careful dose optimization and safety evaluation in future studies.
 | ||
| Fig. 3 Assessment of the inspected compounds 6a–j cytotoxic inhibitory concentration 50 (IC50) against the cancer cell lines: PC3, MCF7, A549, and HCT116 using DOX as a reference. | ||
Among the tested compounds, 6g emerged as one of the most potent derivatives, displaying the highest average GI% (86.40%) and strong cytotoxic activity with IC50 values of 6.49 µM (PC3), 6.58 µM (MCF7), 5.38 µM (A549), and 7.25 µM (HCT116). Likewise, 6e exhibited the lowest IC50 values overall (4.57, 5.83, 5.80, and 11.63 µM against PC3, MCF7, A549, and HCT116, respectively), indicating high potency comparable to doxorubicin (4.88, 17.59, 28.38, and 13.40 µM).
Other derivatives, including 6d (5.56, 6.69, 5.52, and 18.86 µM), 6i (6.50, 5.42, 5.14, and 7.93 µM), and 6j (6.68, 7.24, 6.49, and 6.08 µM), also demonstrated promising cytotoxic profiles with IC50 values mostly in the low micromolar range.
In contrast, 6c and 6h showed noticeably weaker activity, particularly against HCT116 cells (30.72 and 42.64 µM, respectively).
Overall, compounds 6d, 6e, 6g, 6i, and 6j can be considered the most potent members of this series, exhibiting IC50 values below 10 µM in several cancer cell lines, suggesting their strong antiproliferative potential.
Specifically, cell cycle-related proteins CDK2, CDK4, and CDK6 were downregulated with fold changes of 1.57, 1.76, and 4.12, respectively, indicating partial suppression of cell cycle progression. Pro-apoptotic proteins (caspase-3, caspase-8, and caspase-9) were modestly upregulated (fold changes 1.60, 1.62, and 1.64), respectively, suggesting activation of both intrinsic and extrinsic apoptotic pathways. VEGFR-2 expression decreased by 1.68-fold, which may indicate a potential impact on angiogenic signaling; however, functional assays such as tube formation or migration would be required to confirm anti-angiogenic activity.45,46
Overall, these findings indicate that 6g may modulate multiple regulatory pathways, including apoptosis, cell cycle progression, and angiogenesis, although the observed effects are moderate and warrant further functional validation to establish causal relationships (Fig. 4).
 | ||
| Fig. 5 (A) Cell cycle analysis of treated & untreated A549 cells with compound 6g. (B) Histograms of A549 treated with compound 6g (left side) vs. control (right side). | ||
While these results indicate S-phase accumulation, it likely reflects replication stress or downstream cell cycle regulation rather than direct CDK6 inhibition, as CDK4/6 classically regulate the G1–S transition. Selenium-containing compounds, including diselenides, are known to induce oxidative stress and DNA damage, which can converge to produce S-phase arrest.50–52 Therefore, the observed effect supports the conclusion that 6g exerts multifactorial cytotoxic effects, rather than exclusive CDK4/6 inhibition. The experiment was performed three times to ensure reproducibility, and the corresponding histograms are provided in the SI.
2.3. Molecular docking
A molecular docking study was conducted to investigate the apoptotic potential of analogue 6g against the CDK6 target receptor (PDB ID: 5L2I). Compound 6g was selected for molecular docking studies because it exhibited the highest GI% across cancer cell lines and the most favorable selectivity profile toward normal HSF cells among all tested analogues. Accordingly, 6g served as a representative lead compound to explore potential molecular interactions with the CDK6 target receptor.Compound 6g achieved a binding score of −9.55 kcal mol−1 (RMSD = 1.69 Å), which was very promising and comparable to that of the CDK6 co-crystal palbociclib inhibitor (binding score = −10.53 kcal mol−1 and RMSD = 1.71 Å). Analogue 6g got stabilized inside the binding pocket of the CDK6 receptor through the formation of three hydrogen bonds with Thr107, Asp104, and Glu18. Also, it formed four pi–hydrogen interactions with Asp163, Ala162, Leu152, and Gly20. On the other side, the CDK6 co-crystal palbociclib inhibitor showed five hydrogen bonds with Asp163, Val101 (2), His100, and Lys29. Besides, it formed one pi–hydrogen interaction with Gln103 (Fig. 6). Accordingly, the docking study results were in accordance with the previously discussed in vitro ones, indicating a potential suppression activity for compound 6g towards the CDK6 target receptor. While these docking results suggest that 6g could interact with CDK6, it is important to note that docking alone cannot confirm target engagement or selectivity. Therefore, the observed in vitro cytotoxicity may arise from CDK6 inhibition or from other nonspecific effects. These results serve as a computational prediction supporting further experimental validation of 6g as a potential CDK6 inhibitor.
 | ||
| Fig. 6 3D binding interactions of analogue 6g and the co-crystallized palbociclib within the active site of CDK6 receptor (PDB ID: 5L2I). | ||
3. Conclusion
The present study highlights the strong therapeutic promise of the newly synthesized diselenide-linked imidazolone derivatives, several of which demonstrated superior cytotoxic performance compared to DOX. Among the evaluated analogues, 6b, 6d, and 6g were the most active, exhibiting average GI% values of 80.32%, 79.24%, and 86.40%, respectively, underscoring their broad anticancer potential. Compound 6g, in particular, emerged as a standout candidate, displaying potent activity across diverse cancer cell lines with IC50 values of 6.49 µM (PC3), 6.58 µM (MCF7), 5.38 µM (A549), and 7.25 µM (HCT116). Mechanistic investigations revealed that 6g profoundly disrupts multiple oncogenic signaling axes. It significantly downregulated CDK2, CDK4, and CDK6 (1.57-, 1.76-, and 4.12-fold decreases) as well as VEGFR-2 (1.68-fold decrease), consistent with both cell cycle suppression and anti-angiogenic action. Simultaneously, caspase-3, caspase-8, and caspase-9 were upregulated (1.60–1.64-fold), verifying activation of apoptotic pathways. Flow cytometric analysis further confirmed these molecular events, revealing a striking S-phase arrest, where the S-phase fraction increased from 4.61% to 42.09%, accompanied by marked reductions in G0 and G1 populations (13.78% and 42.69%, respectively). Collectively, these findings establish that the diselenide-linked imidazolone scaffold—particularly compound 6g—acts through a coordinated, multi-target mechanism involving apoptosis induction, cell cycle blockade, and inhibition of angiogenesis. Such a profile positions this chemotype as a promising platform for further optimization and development of next-generation anticancer agents.4. Materials and methods
4.1. Chemistry
4.1.3.1. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-benzylidene-2-phenyl-3,5-dihydro-4H-imidazol-4-one)one (6a). Bage powder; yield: 88%; m.p. 98–99 °C (EtOH). FT-IR (KBr, cm−1): 1715 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1639 (C
O stretching, cyclic amide), 1639 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.16 (S, S, 4H, 2H(CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.16 (S, S, 4H, 2H(CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 2H(Ar–H)), 8.16 (d, 2H, Ar–H), 7.6 (t, 2H, Ar–H), 7.5 (t, 2H, Ar–H), 7.4 (m, 12H, Ar–H), 7.3 (t, 4H, Ar–H), 7.1 (m, 4H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 167.4, 163.56, 161.1, 138.8, 136.1, 134.6, 134.2, 133.6, 132.8, 132.1, 131.2, 129.8, 129.5, 129, 128.2, 125.6, 120.6.
C), 2H(Ar–H)), 8.16 (d, 2H, Ar–H), 7.6 (t, 2H, Ar–H), 7.5 (t, 2H, Ar–H), 7.4 (m, 12H, Ar–H), 7.3 (t, 4H, Ar–H), 7.1 (m, 4H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 167.4, 163.56, 161.1, 138.8, 136.1, 134.6, 134.2, 133.6, 132.8, 132.1, 131.2, 129.8, 129.5, 129, 128.2, 125.6, 120.6.
4.1.3.2. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-(4-chlorobenzylidene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6b). Yellow crystals; yield: 60%; m.p. 230–232 °C (EtOH). FT-IR (KBr, cm−1): 2980 (CH–Ar), 1711 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1641 (C
O stretching, cyclic amide), 1641 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.40 (S, 4H, 2H(CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.40 (S, 4H, 2H(CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 2H(Ar–H)), 7.6 (d, 2H, Ar–H), 7.5 (d, 4H, Ar–H), 7.4 (t, 8H, Ar–H), 7.3 (t, 4H, Ar–H), 7.2 (m, 6H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 161.5, 139.2, 136.1, 135.6, 134.3, 133.4, 132.2, 129.4, 128, 128.8, 126.6.
C), 2H(Ar–H)), 7.6 (d, 2H, Ar–H), 7.5 (d, 4H, Ar–H), 7.4 (t, 8H, Ar–H), 7.3 (t, 4H, Ar–H), 7.2 (m, 6H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 161.5, 139.2, 136.1, 135.6, 134.3, 133.4, 132.2, 129.4, 128, 128.8, 126.6.
4.1.3.3. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-(4-nitrobenzylidene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6c). Yellow crystals; yield: 93%; m.p. 250–252 °C (AcOH). FT-IR (KBr, cm−1): 1711.12 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1640 (C
O stretching, cyclic amide), 1640 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.24 (S, 4H, 2H(CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.24 (S, 4H, 2H(CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 2H(Ar–H), 8.00 (d, 4H, Ar–H), 7.8 (d, 4H, Ar–H), 7.7 (d, 4H, Ar–H), 7.6 (t, 4H, Ar–H), 7.5 (t, t, 10H, Ar–H), 7.14 (s, 2H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 166.5, 164.6, 146.9, 142, 140.01, 134.8, 133.6, 133.4, 132.5, 130.7, 128.9, 128.4, 128.09, 125.11, 124.68, 124.1, 121.5, 121.1.
C), 2H(Ar–H), 8.00 (d, 4H, Ar–H), 7.8 (d, 4H, Ar–H), 7.7 (d, 4H, Ar–H), 7.6 (t, 4H, Ar–H), 7.5 (t, t, 10H, Ar–H), 7.14 (s, 2H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 166.5, 164.6, 146.9, 142, 140.01, 134.8, 133.6, 133.4, 132.5, 130.7, 128.9, 128.4, 128.09, 125.11, 124.68, 124.1, 121.5, 121.1.
4.1.3.4. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-(2-nitrobenzylidene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6d). Deep yellow crystals; yield: 85%; m.p. 124–125 °C (EtOH). FT-IR (KBr, cm−1): 1728.6 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1634 (C
O stretching, cyclic amide), 1634 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.15 (S, 2H, (CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 8.15 (S, 2H, (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 8.13 (t, 2H, Ar–H), 7.9 (t, 2H, Ar–H), 7.5 (m, 12H, Ar–H), 7.4 (t, 4H, Ar–H), 7.2–7.35 (m, 6H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 169.8, 163.2, 149.8, 141.09, 140.4, 136.2, 134.2, 133.8, 132.4, 132.16, 131.2, 130.9, 129.5, 129.2, 128.9, 128.3, 125.3, 121.8, 120.6, 120.2.
C), 8.13 (t, 2H, Ar–H), 7.9 (t, 2H, Ar–H), 7.5 (m, 12H, Ar–H), 7.4 (t, 4H, Ar–H), 7.2–7.35 (m, 6H, Ar–H); 13C-NMR (DMSO-d6, 100 MHz) d: 169.8, 163.2, 149.8, 141.09, 140.4, 136.2, 134.2, 133.8, 132.4, 132.16, 131.2, 130.9, 129.5, 129.2, 128.9, 128.3, 125.3, 121.8, 120.6, 120.2.
4.1.3.5. 3,3′-(Diselanediylbis(4,1-phenylene))bis(2-phenyl-5-(3,4,5-trimethoxybenzylidene)-3,5-dihydro-4H-imidazol-4-one) (6e). Orange crystals; yield: 83%; m.p. 220–222 °C (EtOH). FT-IR (KBr, cm−1): 1714.15 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1634 (C
O stretching, cyclic amide), 1634 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 3.7 (s, 6H, 2-OCH3), 3.8 (s, 12H, 4-OCH3), 7.27 (m, 4H, Ar–H), 7.4 (t, 4H, Ar–H), 7.5 (m, 8H, Ar–H),7.6 (d, 2H, Ar–H), 7.66 (d, 4H, Ar–H), 7.8 (s, 2H, CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 3.7 (s, 6H, 2-OCH3), 3.8 (s, 12H, 4-OCH3), 7.27 (m, 4H, Ar–H), 7.4 (t, 4H, Ar–H), 7.5 (m, 8H, Ar–H),7.6 (d, 2H, Ar–H), 7.66 (d, 4H, Ar–H), 7.8 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C); 13C-NMR (DMSO, 100 MHz) d: 56.15, 60.7, 110.5, 128.5, 129.1, 129.5, 130.01, 130.67, 132.0, 132.14, 133.8, 134.6, 134.7, 137.9, 140.4, 153.2, 160.2, 169.9.
C); 13C-NMR (DMSO, 100 MHz) d: 56.15, 60.7, 110.5, 128.5, 129.1, 129.5, 130.01, 130.67, 132.0, 132.14, 133.8, 134.6, 134.7, 137.9, 140.4, 153.2, 160.2, 169.9.
4.1.3.6. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-(benzo[d][1,3]dioxol-5-ylmethylene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6f). Yellow crystals; yield: 95%; m.p. 170–172 °C (AcOH). FT-IR (KBr, cm−1): 1716.4 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1640 (C
O stretching, cyclic amide), 1640 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.2 (s, 4H, 2-OC
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.2 (s, 4H, 2-OC![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 2O–), 8.13 (t, 2H, Ar–H), 8.3 (m, 6H, Ar–H), 8.6 (m, 6H, Ar–H),8.7 (t, 3H, Ar–H), 9 (m, 4H, Ar–H), 9.1 (s, 2H, CH
2O–), 8.13 (t, 2H, Ar–H), 8.3 (m, 6H, Ar–H), 8.6 (m, 6H, Ar–H),8.7 (t, 3H, Ar–H), 9 (m, 4H, Ar–H), 9.1 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C); 13C-NMR (DMSO-d6, 100 MHz) d: 99.06, 105.3, 106, 116.2, 120, 123.4, 123.9, 124.3, 125.2, 125.4, 126.2, 127.8, 128.9, 132.4.
C); 13C-NMR (DMSO-d6, 100 MHz) d: 99.06, 105.3, 106, 116.2, 120, 123.4, 123.9, 124.3, 125.2, 125.4, 126.2, 127.8, 128.9, 132.4.
4.1.3.7. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-(furan-2-ylmethylene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6g). Yellow powder; yield: 75%; m.p. 118–119 °C (AcOH). FT-IR (KBr, cm−1): 1714.7(C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1637.31 (C
O stretching, cyclic amide), 1637.31 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.15 (t, 2H, Ar–H), 7.2 (m, 4H, Ar–H), 7.3 (m, 4H, Ar–H), 7.5 (m, 10H, Ar–H), 7.6 (t, 4H, Ar–H), 7.7 (d, 2H, Ar–H), 8.1 (s, 2H, CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.15 (t, 2H, Ar–H), 7.2 (m, 4H, Ar–H), 7.3 (m, 4H, Ar–H), 7.5 (m, 10H, Ar–H), 7.6 (t, 4H, Ar–H), 7.7 (d, 2H, Ar–H), 8.1 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C); 13C-NMR (DMSO-d6, 100 MHz) d: 112.9, 114.5, 120.02, 128.14, 128.9, 129.1, 129.5, 130.6, 131.1, 132.1, 133.5, 133.8, 134.5, 136, 147.7, 147.7, 150.9, 169.4.
C); 13C-NMR (DMSO-d6, 100 MHz) d: 112.9, 114.5, 120.02, 128.14, 128.9, 129.1, 129.5, 130.6, 131.1, 132.1, 133.5, 133.8, 134.5, 136, 147.7, 147.7, 150.9, 169.4.
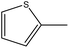
4.1.3.8. 3,3′-(Diselanediylbis(4,1-phenylene))bis(2-phenyl-5-(thiophen-2-ylmethylene)-3,5-dihydro-4H-imidazol-4-one) (6h). Deep yellow crystals; yield: 92%; m.p. 310–312 °C (AcOH). FT-IR (KBr, cm−1): 1730.8 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1632.3 (C
O stretching, cyclic amide), 1632.3 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.2 (t, 4H, Ar–H), 8.13 (t, 2H, Ar–H), 8.6 (d, 4H, Ar–H),8.6 (m, 8H, Ar–H), 8.9 (m, 4H, Ar–H), 9 (d, 2H, Ar–H), 9.4 (s, 2H, CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 7.2 (t, 4H, Ar–H), 8.13 (t, 2H, Ar–H), 8.6 (d, 4H, Ar–H),8.6 (m, 8H, Ar–H), 8.9 (m, 4H, Ar–H), 9 (d, 2H, Ar–H), 9.4 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C); 13C-NMR (DMSO-d6, 100 MHz) d: 73.9, 101.7, 102.9, 109.3, 112.3, 112.6, 114.8, 115.6, 119.6, 129.1, 129.9, 145.8, 152.7, 166.6, 168.2, 175.7.
C); 13C-NMR (DMSO-d6, 100 MHz) d: 73.9, 101.7, 102.9, 109.3, 112.3, 112.6, 114.8, 115.6, 119.6, 129.1, 129.9, 145.8, 152.7, 166.6, 168.2, 175.7.
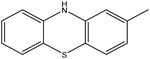
4.1.3.9. 3,3′-(Diselanediylbis(4,1-phenylene))bis(5-((10H-phenothiazin-2-yl)methylene)-2-phenyl-3,5-dihydro-4H-imidazol-4-one) (6i). Deep blue crystals; yield: 65%; m.p. 198–200 °C (DMF). FT-IR (KBr, cm−1): 3330.2 (NH), 1732.8 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1651.7 (C
O stretching, cyclic amide), 1651.7 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 6.7 (m, 4H, Ar–H), 6.9 (t, 6H, Ar–H), 6.9 (t, 4H, Ar–H), 7.00 (t, 4H, Ar–H), 7.1 (s, 2H, Ar–H), 7.3 (s, 2H, Ar–H),7.5 (d, 4H, Ar–H), 7.6 (t, 6H, Ar–H), 7.9 (d, 2H, Ar–H), 7.9 (s, 2H, CH
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 6.7 (m, 4H, Ar–H), 6.9 (t, 6H, Ar–H), 6.9 (t, 4H, Ar–H), 7.00 (t, 4H, Ar–H), 7.1 (s, 2H, Ar–H), 7.3 (s, 2H, Ar–H),7.5 (d, 4H, Ar–H), 7.6 (t, 6H, Ar–H), 7.9 (d, 2H, Ar–H), 7.9 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 8.1 (d, 4H, Ar–H), 9.6 (s, 2H, –NH); 13C-NMR (DMSO-d6, 100 MHz) d: 114.4, 114.9, 115.6, 116.1, 117.1, 123.3, 123.5, 126.7, 127.8, 128.1, 128.3, 129.8, 130.2, 140, 147.7, 190.4.
C), 8.1 (d, 4H, Ar–H), 9.6 (s, 2H, –NH); 13C-NMR (DMSO-d6, 100 MHz) d: 114.4, 114.9, 115.6, 116.1, 117.1, 123.3, 123.5, 126.7, 127.8, 128.1, 128.3, 129.8, 130.2, 140, 147.7, 190.4.
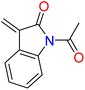
4.1.3.10. 3,3′-((Diselanediylbis(4,1-phenylene))bis(5-oxo-2-phenyl-1,5-dihydro-4H-imidazole-1-yl-4-ylidene))bis(1-acetylindolin-2-one) (6j). Brown crystals; yield: 60%; m.p. 124–125 °C (EtOH). FT-IR (KBr, cm−1): 1717.5 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching, cyclic amide), 1694
O stretching, cyclic amide), 1694 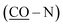 , 1637.4 (C
, 1637.4 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 2.09 (s, 6H, COC
N stretching); 1H-NMR (DMSO-d6, 400 MHz), d (ppm): 2.09 (s, 6H, COC![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 3), 6.9 (t, 2H, Ar–H), 7.2 (d, 2H, Ar–H), 7.3 (d, 4H, Ar–H), 7.00 (t, 4H, Ar–H), 7.5 (m, 8H, Ar–H), 7.6 (m, 6H, Ar–H),7.66 (d, 2H, Ar–H), 7.9 (s, 2H, CH
3), 6.9 (t, 2H, Ar–H), 7.2 (d, 2H, Ar–H), 7.3 (d, 4H, Ar–H), 7.00 (t, 4H, Ar–H), 7.5 (m, 8H, Ar–H), 7.6 (m, 6H, Ar–H),7.66 (d, 2H, Ar–H), 7.9 (s, 2H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C); 13C-NMR (DMSO-d6, 100 MHz) d: 224.5, 120.1, 120.4, 129.8, 133.6, 133.8, 134.5, 139.4, 140.1, 169.0.
C); 13C-NMR (DMSO-d6, 100 MHz) d: 224.5, 120.1, 120.4, 129.8, 133.6, 133.8, 134.5, 139.4, 140.1, 169.0.
4.2. Biological assays
4.3. Molecular docking
To further investigate the apoptotic potential of compound 6g, a molecular docking study was conducted against the CDK6 target receptor using the AutoDock Vina61 and PyMOL.62 Analogue 6g was first prepared by energy minimization and optimization of partial charges.63,64 The PDB was searched, and the CDK6 receptor with ID (5L2I) was selected and prepared through hydrogenation (3D), correction, and energy minimization.65 The co-crystal inhibitor (palbociclib) of 5L2I was inserted as a positive reference to analogue 6g. Finally, a redocking procedure for the CDK6 co-crystal palbociclib clarified the validity of the applied software (RMSD < 2 Å).66Author contributions
Conceptualization: Ahmed A. Al-Karmalawy and Saad Shaaban; supervision: Marwa Abdel-Motaal, Saad Shaaban, and Ahmed A. Al-Karmalawy; data curation, visualization, methodology, and writing – review & editing: Marwa Abdel-Motaal, Saad Shaaban, Samia S. Hawas, Asma M. Elsharif, Marwa Sharaky, Fatema S. Alatawi, Mohamed E. Eissa, Arwa Omar Al Khatib, Hany M. Abd El-Lateef, Medhat Asem, and Ahmed A. Al-Karmalawy. Finally, all authors revised and approved the final submitted version of the manuscript.Conflicts of interest
The authors declared no conflict of interest.Data availability
The data supporting this article have been included in the manuscript and the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra10063a.Funding
This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia [Grant No. KFU260238].Acknowledgements
This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia [Grant No. KFU260238].References
- F. H. Al-Ostoot, S. Salah and S. A. Khanum, Cancer Invest., 2024, 42, 559–604 Search PubMed.
- S. M. Alzahrani, H. A. Al Doghaither and A. B. Al-Ghafari, Mol. Clin. Oncol., 2021, 15, 271 Search PubMed.
- D. E.-S. Ellakwa, N. Mushtaq, S. Khan, A. Jabbar, M. A. Abdelmalek, A.-H. S. Wadan, T. E. Ellakwa and A. Raza, Naunyn-Schmiedeberg's Arch. Pharmacol., 2024, 397, 5617–5630 CrossRef CAS PubMed.
- M. M. Hammoud, A. S. Nageeb, M. A. Morsi, E. A. Gomaa, A. A. Elmaaty and A. A. Al-Karmalawy, New J. Chem., 2022, 46, 11422–11436 Search PubMed.
- A. Leiter, R. R. Veluswamy and J. P. Wisnivesky, Nat. Rev. Clin. Oncol., 2023, 20, 624–639 Search PubMed.
- H. Padinharayil, J. Varghese, M. C. John, G. K. Rajanikant, C. M. Wilson, M. Al-Yozbaki, K. Renu, S. Dewanjee, R. Sanyal, A. Dey, A. G. Mukherjee, U. R. Wanjari, A. V. Gopalakrishnan and A. George, Genes Dis., 2023, 10, 960–989 CrossRef CAS PubMed.
- P. Das, S. Najafikhoshnoo, J. A. Tavares-Negrete, Q. Yi and R. Esfandyarpour, Bioprinting, 2022, 28, e00243 Search PubMed.
- F. Abid, M. Khan and M. Shabbir, in Nanotheranostics and Precision Oncology, eds. R. Riaz, M. Shabbir, Y. Badshah, A. Ahmad and K. Khan, Academic Press, 2026, pp. 29–56, DOI:10.1016/B978-0-443-34671-2.00003-7.
- K. H. Walen, Semin. Cancer Biol., 2022, 81, 73–82 CrossRef CAS PubMed.
- P. Łukasik, M. Załuski and I. Gutowska, Int. J. Mol. Sci., 2021, 22, 2935 CrossRef PubMed.
- Z. Wang, Cells, 2021, 10, 3327 CrossRef CAS PubMed.
- A. Lashen, M. Algethami, S. Alqahtani, A. Shoqafi, A. Sheha, J. N. Jeyapalan, N. P. Mongan, E. A. Rakha and S. Madhusudan, Int. J. Mol. Sci., 2024, 25, 4060 Search PubMed.
- R. Janostiak, A. Torres-Sanchez, F. Posas and E. de Nadal, Cancers, 2022, 14, 1265 Search PubMed.
- S. Goel, J. S. Bergholz and J. J. Zhao, Nat. Rev. Cancer, 2022, 22, 356–372 CrossRef CAS PubMed.
- A. Fassl, Y. Geng and P. Sicinski, Science, 2022, 375(6577), eabc1495 CrossRef CAS PubMed.
- K. Gousias, T. Theocharous and M. Simon, Biomedicines, 2022, 10, 564 CrossRef CAS PubMed.
- L. V. Loftus, S. R. Amend and K. J. Pienta, Int. J. Mol. Sci., 2022, 23, 4723 CrossRef CAS PubMed.
- S. G. Elia, A. A. Al-Karmalawy, M. Y. Nasr and M. F. Elshal, J. Biochem. Mol. Toxicol., 2022, 36, e22938 Search PubMed.
- M. Mustafa, R. Ahmad, I. Q. Tantry, W. Ahmad, S. Siddiqui, M. Alam, K. Abbas, M. I. Moinuddin, S. Habib and S. Islam, Cells, 2024, 13(22), 1838 Search PubMed.
- M. Saddam, S. K. Paul, M. A. Habib, M. A. Fahim, A. Mimi, S. Islam, B. Paul and M. M. U. Helal, Egypt. J. Med. Hum. Genet., 2024, 25, 12 CrossRef.
- P. Hussar, Encyclopedia, 2022, 2, 1624–1636 Search PubMed.
- W. Yu, H. Zhu, R. Huang, B. Yan, B. Xu, Y. Shi, J. Mao, Z. Liu and J. Wang, Toxicon, 2024, 237, 107561 Search PubMed.
- S. Sağ, İ. Bayav, A. G. Tomatır and Y. Dodurga, Bratislava Med. J., 2025, 126(11), 2945–2953 Search PubMed.
- C. M. Roggero, A. B. Ghosh, A. Devineni, S. Ma, E. Blatt, G. V. Raj and Y. Yin, Transl. Oncol., 2025, 52, 102231 CrossRef CAS PubMed.
- B. Liu, Z. Peng, H. Zhang, N. Zhang, Z. Liu, Z. Xia, S. Huang, P. Luo and Q. Cheng, Mol. Cancer, 2025, 24, 106 Search PubMed.
- V. Nardone, M. Barbarino, A. Angrisani, P. Correale, P. Pastina, S. Cappabianca, A. Reginelli, L. Mutti, C. Miracco, R. Giannicola, A. Giordano and L. Pirtoli, Int. J. Mol. Sci., 2021, 22, 8391 Search PubMed.
- J. J. Asciolla, X. Wu, C. Adamopoulos, E. Gavathiotis and P. I. Poulikakos, Nat. Cancer, 2025, 6, 24–40 CrossRef PubMed.
- W. S. Hamama, G. G. El-Bana, S. Shaaban, O. Habib and H. H. Zoorob, RSC Adv., 2016, 6, 24010–24049 RSC.
- W. S. Hamama, G. G. El-Bana, S. Shaaban and H. H. Zoorob, J. Heterocycl. Chem., 2018, 55, 971–982 CrossRef CAS.
- S. Shaaban, A. Y. M. Alabdali, M. H. Mousa, H. Ba-Ghazal, Y. S. Al-Faiyz, I. Elghamry, H. A. Althikrallah, A. O. A. Khatib, M. Alaasar and A. A. Al-Karmalawy, Drug Dev. Res., 2025, 86, e70075 CrossRef CAS PubMed.
- S. Shaaban, H. M. A. E.-L. Ahmed and B. Alshemary, US Pat., 11964947, 2024 Search PubMed.
- D. A. Refaay, D. M. Ahmed, A. M. Mowafy and S. Shaaban, Med. Chem. Res., 2022, 1–11 Search PubMed.
- S. Shaaban, M. A. Arafat and W. S. Hamama, Arkivoc, 2014, 2014, 470–505 Search PubMed.
- S. Shaaban, H. M. A. El-Lateef, M. M. Khalaf, M. Gouda and I. Youssef, Polymers, 2022, 14, 2208 Search PubMed.
- H. M. Abd El-Lateef, M. M. Khalaf, M. Gouda, K. Shalabi, F. E. T. Heakal, A. S. Al-Janabi and S. Shaaban, Constr. Build. Mater., 2023, 366, 130135 CrossRef CAS.
- S. Shaaban, K. Shalabi, T. A. Yousef, M. Abou-Krisha, A. A. Alanazi, H. A. Althikrallah, M. Alaasar, A. M. Abu-Dief and A. S. Al-Janabi, J. Taiwan Inst. Chem. Eng., 2024, 165, 105766 CrossRef CAS.
- S. Shaaban, K. T. Abdullah, K. Shalabi, T. A. Yousef, O. K. Al Duaij, G. M. Alsulaim, H. A. Althikrallah, M. Alaasar, A. S. Al-Janabi and A. M. Abu-Dief, Appl. Organomet. Chem., 2024, e7712 CrossRef CAS.
- S. Shaaban, H. Ba-Ghazal, Y. S. Al-Faiyz, A. A. Al-Karmalawy, N. Amri and I. Youssef, Tetrahedron, 2024, 133957 Search PubMed.
- S. Shaaban, H. A. Althikrallah, A. Negm, A. Abo Elmaaty and A. A. Al-Karmalawy, RSC Adv., 2024, 14, 18576–18587 Search PubMed.
- S. Shaaban, M. A. Arafat, H. E. Gaffer and W. S. Hamama, Der Pharma Chem., 2014, 6, 186–193 Search PubMed.
- Z. Murtazaeva, A. Nasrullaev, A. Buronov, S. Gaybullaev, L. Nie, S. Numonov, Z. Khushnazarov, D. Turgunov, R. Kuryazov and J. Zhao, Molecules, 2025, 30, 2245 CrossRef CAS PubMed.
- V. Vainauskas, P. Kavaliauskas, B. Grybaitė, V. Petraitis, R. Petraitienė, R. Grigalevičiūtė, R. Prakapaitė, W. Acevedo and V. Mickevičius, ChemMedChem, 2025, 20, e202500260 Search PubMed.
- B. Liu, J. Wu, T. Zhu, X. Tian, N. Chen, W. Li and T. Li, Mini-Rev. Org. Chem., 2025, 22, 427–446 Search PubMed.
- P. Skehan, R. Storeng, D. Scudiero, A. Monks, J. McMahon, D. Vistica, J. T. Warren, H. Bokesch, S. Kenney and M. R. Boyd, J. Natl. Cancer Inst., 1990, 82, 1107–1112 CrossRef CAS PubMed.
- H. Kemel, L. Benguedouar, D. Boudjerda, S. Menadi, E. Cacan and M. Sifour, Med. Oncol., 2024, 41, 144 Search PubMed.
- G. Natale and G. Bocci, in Tumor Angiogenesis Assays: Methods and Protocols, ed. D. Ribatti, Springer US, New York, NY, 2023, pp. 1–37, DOI:10.1007/978-1-0716-2703-7_1.
- S. Nazreen, A. S. A. Almalki, S. E. I. Elbehairi, A. A. Shati, M. Y. Alfaifi, A. A. Elhenawy, N. I. Alsenani, A. Alfarsi, A. Alhadhrami, E. A. Alqurashi and M. M. Alam, Molecules, 2022, 27, 6899 Search PubMed.
- F. Alminderej, S. Ghannay, M. O. Elsamani, F. Alhawday, A. E. A. E. Albadri, S. E. I. Elbehairi, M. Y. Alfaifi, A. Kadri and K. Aouadi, Pharmaceuticals, 2023, 16, 1025 Search PubMed.
- T. Keleş, B. Barut, A. Özel and Z. Biyiklioglu, Bioorg. Chem., 2021, 107, 104637 CrossRef PubMed.
- K. Wang, X. T. Fu, Y. Li, Y. J. Hou, M. F. Yang, J. Y. Sun, S. Y. Yi, C. D. Fan, X. Y. Fu, J. Zhai and B. L. Sun, Neurochem. Res., 2016, 41, 1439–1447 Search PubMed.
- Y. Qi, N. W. Schoene, F. M. Lartey and W. H. Cheng, J. Biol. Chem., 2010, 285, 33010–33017 CrossRef CAS PubMed.
- L. Letavayová, V. Vlcková and J. Brozmanová, Toxicology, 2006, 227, 1–14 Search PubMed.
- P. Skehan, R. Storeng, D. Scudiero, A. Monks, J. McMahon, D. Vistica, J. T. Warren, H. Bokesch, S. Kenney and M. R. Boyd, J. Natl. Cancer Inst., 1990, 82, 1107–1112 Search PubMed.
- M. M. Hammoud, M. Khattab, M. Abdel-Motaal, J. Van der Eycken, R. Alnajjar, H. S. Abulkhair and A. A. Al-Karmalawy, J. Biomol. Struct. Dyn., 2023, 41, 5199–5216 CAS.
- N. El-Gohary, S. Hawas, M. Gabr, M. Shaaban and M. El-Ashmawy, Bioorg. Chem., 2019, 92, 103109 CrossRef CAS PubMed.
- S. S. Hawas, N. S. El-Gohary, M. T. Gabr, M. I. Shaaban and M. B. El-Ashmawy, Synth. Commun., 2019, 49, 2466–2487 Search PubMed.
- S. Shaaban, H. A. Althikrallah, A. Negm, A. A. Elmaaty and A. A. Al-Karmalawy, RSC Adv., 2024, 14, 18576–18587 RSC.
- P. Karousi, C. K. Kontos, S. T. Nikou, T. Carell, D. C. Sideris and A. Scorilas, Funct. Integr. Genomics, 2025, 25, 1–18 Search PubMed.
- S. S. Hawas, S. M. El-Sayed, P. A. Elzahhar and M. A. Moustafa, Bioorg. Chem., 2023, 141, 106874 Search PubMed.
- A. Ligasová, I. Frydrych and K. Koberna, Int. J. Mol. Sci., 2023, 24, 3674 CrossRef PubMed.
- R. Huey, G. M. Morris and S. Forli, The Scripps Research Institute Molecular Graphics Laboratory, 2012, vol. 10550, p. 1000 Search PubMed.
- S. Yuan, H. S. Chan and Z. Hu, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2017, 7, e1298 Search PubMed.
- D. B. Mahmoud, M. M. Bakr, A. A. Al-Karmalawy, Y. Moatasim, A. El Taweel and A. Mostafa, AAPS PharmSciTech, 2022, 23, 1–12 CrossRef PubMed.
- T. Al-Warhi, M. M. Elbadawi, A. Bonardi, A. Nocentini, A. A. Al-Karmalawy, N. Aljaeed, O. J. Alotaibi, H. A. Abdel-Aziz, C. T. Supuran and W. M. Eldehna, J. Enzyme Inhib. Med. Chem., 2022, 37, 2635–2643 CrossRef CAS PubMed.
- A. Mahmoud, E. Kotb, A. I. Alqosaibi, A. A. Al-Karmalawy, I. S. Al-Dhuayan and H. Alabkari, Heliyon, 2021, 7, e08148 CrossRef CAS PubMed.
- M. M. Hammouda, A. A. Elmaaty, M. S. Nafie, M. Abdel-Motaal, N. S. Mohamed, M. A. Tantawy, A. Belal, R. Alnajjar, W. M. Eldehna and A. A. Al-Karmalawy, Bioorg. Chem., 2022, 127, 105995 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |