DOI:
10.1039/D5RA10028C
(Paper)
RSC Adv., 2026,
16, 17295-17304
Rapid and facile colorimetric biosensing of dengue NS1 antigen using gold nanoparticles: potential for point-of-care testing
Received
26th December 2025
, Accepted 25th March 2026
First published on 30th March 2026
Abstract
Dengue fever is one of the major global health concerns with increasing incidence and mortality. Early and accurate detection is essential to prevent severe outcomes, especially in resource-limited facilities. Here, we report a rapid, low-cost colorimetric biosensor based on gold nanoparticles (AuNPs) synthesized using ortho-hydroxybenzoic acid, yielding a red wine-coloured solution with a surface plasmon resonance peak at 524 nm. Surface modification with N-[(3-dimethylamino)propyl]-N′-ethylcarbodiimide hydrochloride/N-hydroxysuccinimide (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio) enabled efficient antibody conjugation to NS1 protein monoclonal antibodies. Colorimetric detection with the NS1 antigen provided results within 30 seconds, exhibited excellent linearity (R2 = 0.982), and demonstrated limits of detection and quantification of 49.65 ng mL−1 and 165.48 ng mL−1, respectively. This AuNP-based biosensor offers a fast, accurate, and user-friendly platform, potential for point-of-care dengue diagnosis, with nanoparticles remaining stable for up to 28 days.
1 ratio) enabled efficient antibody conjugation to NS1 protein monoclonal antibodies. Colorimetric detection with the NS1 antigen provided results within 30 seconds, exhibited excellent linearity (R2 = 0.982), and demonstrated limits of detection and quantification of 49.65 ng mL−1 and 165.48 ng mL−1, respectively. This AuNP-based biosensor offers a fast, accurate, and user-friendly platform, potential for point-of-care dengue diagnosis, with nanoparticles remaining stable for up to 28 days.
1. Introduction
Dengue fever (DF) is a severe mosquito-borne viral infection caused by the Flaviviridae family, transmitted primarily by female Aedes aegypti and Aedes albopictus mosquitoes.1 The dengue virus has four serotypes—DENV-1, DENV-2, DENV-3, and DENV-4, which are genetically and antigenically related.1 DF symptoms include high fever, headache, nausea, and diarrhea. These nonspecific symptoms may progress to critical conditions such as hypotension, plasma leakage, internal bleeding, and ascites.2 Four levels of severity according to the World Health Organisation (WHO) are: level I (minor bruising), level II (spontaneous skin bleeding), level III (shock symptoms), and level IV (severe acute shock), among which levels II and IV are considered acute and potentially fatal.3 As DF is prevalent in tropical and subtropical regions, including Indonesia, it is considered as endemic due to consistently high and rising case numbers.4 In fact, as of May 2025, the Ministry of Health recorded more than 56![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 DF cases and 250 deaths across 456 districts/cities, representing 87% of Indonesia's territory.5 Adding to concerns, many high-risk areas in Indonesia face resource limitations, making it challenging to access necessary detection and monitoring equipment.
000 DF cases and 250 deaths across 456 districts/cities, representing 87% of Indonesia's territory.5 Adding to concerns, many high-risk areas in Indonesia face resource limitations, making it challenging to access necessary detection and monitoring equipment.
The DF virus itself contains a single-stranded positive-sense RNA genome encoding three structural proteins (E, C, and prM) and seven non-structural proteins (NS1 to NS5).6 Among these, the NS1 protein is a secreted glycoprotein that circulates in the bloodstream during early infection and serves as a reliable biomarker due to its high sensitivity and specificity.7,8 Generally, two conventional approaches are used to detect the DF virus at a laboratory scale. The first method by serological testing, aiming IgM,9 IgG,10 IgM/IgG ratio,11 hemagglutination inhibition,12 plaque reduction neutralization test,13 and NS1-based tests.14 However, these tests often require 4–5 days post-symptom onset to yield accurate results and may suffer from low sensitivity.15 The second method is molecular detection, primarily reverse transcription-polymerase chain reaction, which offers high specificity and speed but is costly and requires sophisticated laboratory infrastructure.16
Meanwhile, the development of rapid, accessible diagnostic methods suitable for point-of-care testing (PoCT) for virus detection via colorimetric tests has attracted attention due to their simplicity.17–19 Under this approach, observations by the naked human eye or an unsophisticated camera device could easily identify a color change to determine whether a specific virus is present, and, with relatively simple algorithms, could further quantify test results inexpensively and instantly.20 One effective strategy for developing colorimetric sensors involves the use of metal nanoparticles, particularly gold nanoparticles (AuNPs), that exhibit strong surface plasmon resonance (SPR) properties.21 AuNPs are widely used in biosensor research due to their inertness, outstanding stability, biocompatibility, low toxicity, and environmental safety.22 These properties deliberately promote AuNPs as an ideal material for biorecognition and biological detection purpose. Moreover, surface modification using functional ligands or biomolecules can enhance its sensitivity and selectivity.23
Our long-standing interest has been to synthesize AuNPs by employing ortho-hydroxybenzoic acid (o-HBA) and to further explore their colorimetric biosensing capabilities. o-HBA has successfully served as reductant by allowing the conversion of Au3+ to Au0, and exhibiting good stabilizing effects due to hydroxy and carboxy groups.24 Furthermore, the synthesized AuNPs from o-HBA was found to have high crystallinity and long-term stability up to five months.25 According to this speciality, herein, we explore the development of colorimetric biosensor of NS1 antigen using AuNPs, in which chemically modified of AuNPs' surface was conducted with N-[(3-dimethylamino)propyl]-N′-ethylcarbodiimide hydrochloride and N-hydroxysuccinimide (EDC/NHS). Surface modification using the EDC/NHS cross coupling is a common method for activating carboxyl groups, in this case from o-HBA, and converting them into stable NHS esters.26 This modification enables covalent bonding to the upcoming terminal amine in NS1 monoclonal antibodies (mAb). Subsequently, bioconjugating the mAb to NS1 antigens is expected for determining the selectivity and sensitivity of the gold nanoparticles (AuNPs) used in the process.27 This work offers a rapid diagnostic method and a potential practical solution for point-of-care DF detection, particularly in laboratory-limited regions since visual color change to detect dengue NS1 antigen was observable within 30 seconds. To emphasize, the discussion would be framed around preliminary findings, focusing on new trends for which further exploration would be needed to enable practical clinical application.
2. Experimental works
2.1. Materials and instruments
The chemicals and reagents were purchased from commercial source: HAuCl4 (100 ppm, Sigma-Aldrich), o-HBA (0.01 M, pH 12, Central Drug House), EDC and NHS (Sigma-Aldrich), 0.1 M HCl (Smart Lab), 0.1 M NaOH (Merck), phosphate buffer saline (PBS, pH 7.4, Himedia), Milli-Q distilled water, NS1 monoclonal antibody (NS1 mAb, ab41616), recombinant NS1 antigen (NS1, ab64456), human serum (H6914-20 mL), cholesterol, and uric acid (Sigma-Aldrich), glucose (Tokyo Chemical Industry), and they were used without further purification. The equipment used in this study is as follows Pyrex glassware, magnetic stirrer, microtubes, micropipettes (Thermo Scientific), analytical balance, well plates, pH meter (Horiba), shaker (LabTech), silicon wafers (Semiconductor Technology and Applications), ELISA Reader (Thermo Scientific), ultraviolet-visible (UV-vis) spectrophotometer (Agilent Cary 60), Fourier Transform Infrared Spectroscopy (FT-IR) spectrometer (Shimadzu IR Prestige 21), high-resolution transmission electron microscope (HR-TEM H9500), and field emission scanning electron microscope (FESEM JIB-4610F). TEM images were analyzed using ImageJ 1.54g (Wayne Rasband, National Institutes of Health, USA).
2.2. Synthesis of AuNPs
AuNPs were synthesized following reported work by mixing 0.01 M o-HBA solution (5 mL) and 100 ppm HAuCl4 solution (5 mL).24 The resultant mixture was heated at 98 °C for 20 minutes on a water bath while maintaining the pH of 12. The color change of the mixture was monitored by UV-vis spectrophotometry over 200–800 nm.
2.3. Surface modification of AuNPs
To a solution of AuNPs solution (1 mL), a mixture of EDC (10 µL) and NHS (100 µL) was added, followed by shaking at 200 rpm for 30 min at room temperature. UV-vis analysis (200–800 nm) was performed with different EDC/NHS ratios listed in Table 1.
Table 1 EDC/NHS ratio for surface modification
| EDC/NHS ratio |
Amount of reagent (in 1 mL PBS) |
| EDC (g) |
NHS (g) |
0.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
0.005 |
0.005 |
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
0.01 |
0.005 |
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
0.02 |
0.005 |
4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
0.04 |
0.005 |
| Control |
0 |
0 |
2.4. NS1 antibody bioconjugation
NS1 mAb was diluted with PBS at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 ratio. The optimal AuNPs-EDC/NHS conjugate was prepared by mixing varying volumes of diluted NS1 mAb (Table 2) and stirring at 200 rpm for 1 h at 37 °C. UV-vis analysis (200–800 nm) was used to evaluate the optimal bioconjugation conditions, followed by characterization using FT-IR, TEM, and FESEM.
100 ratio. The optimal AuNPs-EDC/NHS conjugate was prepared by mixing varying volumes of diluted NS1 mAb (Table 2) and stirring at 200 rpm for 1 h at 37 °C. UV-vis analysis (200–800 nm) was used to evaluate the optimal bioconjugation conditions, followed by characterization using FT-IR, TEM, and FESEM.
Table 2 AuNPs-EDC/NHS and mAb NS1 variations
| Variation |
AuNPs-EDC/NHS (µL) |
NS1 mAb (µL) |
| 1 |
900 |
100 |
| 2 |
800 |
200 |
| 3 |
700 |
300 |
| 4 |
600 |
400 |
| Control |
1000 |
— |
2.5. NS1 antigen detection
NS1 antigen was diluted with PBS to final concentrations of 75, 100, 150, 200, and 250 ng mL−1. The antigen solution (100 µL) was mixed with an AuNPs-mAb solution (100 µL) in a well plate. PBS was used as the negative control. Samples were incubated on a shaker under designated time and analyzed using an ELISA Reader (200–800 nm).
2.6. Characterization of AuNPs
Samples of AuNPs, AuNPs-EDC/NHS, and AuNPs-mAb were drop-cast onto silicon wafers and air-dried. This process was repeated 3–4 times. The dried samples were analyzed using FT-IR (400–4000 cm−1), FESEM and Energy-Dispersive X-ray Spectroscopy (EDX). For TEM analysis, drops of each sample were placed on carbon-coated grids and allowed to dry at room temperature for 24 h before insertion into the TEM.
2.7. Validation methods
2.7.1. Variation of detection time. An AuNPs-mAb solution (100 µL) and a solution of 250 ng mL−1 NS1 antigen (100 µL) were mixed, and the resultant mixture was analyzed using an ELISA Reader (200–800 nm). Absorbance was recorded at 5-min intervals for a total period of 60 min.
2.7.2. Sensitivity test. A calibration curve of the absorbance data was generated to determine the linear equation and correlation coefficient. The limit of detection (LoD) and limit of quantitation (LoQ) were calculated using equations (eqn (1) and (2)), respectively, where SD indicates standard deviation, and s indicates slope of the calibration curve.28| |
 | (1) |
| |
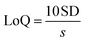 | (2) |
2.7.3. Repeatability and reproducibility test. Repeatability was assessed by conducting three replicate measurements using an ELISA Reader (200–800 nm) under identical conditions. Reproducibility was determined using three independent sets of measurements. Relative standard deviation (RSD) values were calculated using equations (eqn (3) and (4)), respectively where ![[x with combining macron]](https://www.rsc.org/images/entities/i_char_0078_0304.gif) indicates average of data set, and C indicates concentration of analyte.29
indicates average of data set, and C indicates concentration of analyte.29| |
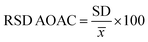 | (3) |
| |
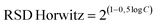 | (4) |
2.7.4. Selectivity test. Selectivity was evaluated by testing potential interferents, including glucose (250 ng mL−1), cholesterol (250 ng mL−1), and uric acid (250 ng mL−1). These matrices were compared to the NS1 antigen (250 ng mL−1) and the PBS as a control. Each test solution (100 µL) and a solution of AuNPs-mAb (100 µL) were mixed in well plates, and the mixture was shaken and analyzed using ELISA at 200–800 nm.
2.7.5. Application to human serum. NS1-negative and NS1-positive serum samples were prepared by spiking 250 ng mL−1 NS1 antigen into the diluted human serum. Each test solution (100 µL) was mixed with a solution of AuNPs-mAb (100 µL) in a well plate. Samples were shaken and analyzed using an ELISA Reader (200–800 nm).
2.7.6. Stability test. A mixture of an AuNPs-mAb solution (100 µL) and 250 ng mL−1 NS1 antigen solution (100 µL) was stored for up to 4 weeks. Samples were analyzed by ELISA at defined intervals: 0 d, 1 d, 3 d, and 1–4 weeks, with a wavelength range of 200–800 nm.
3. Results and discussion
3.1. Synthesis of AuNPs
AuNPs were successfully synthesized through the reduction of Au3+ to Au0 using o-HBA (Fig. 1a). In addition to reducing ability of o-HBA, the vicinal functionalities serve as a stabilizer of AuNPs by chelation, which helps to prevent aggregation and contributes to the long-term stability.
 |
| | Fig. 1 (a) Schematic illustration of the reaction mechanism for AuNPs synthesis, (b) reaction mechanism for modification of AuNPs surface using EDC/NHS mixture. | |
The formation of AuNPs was visually confirmed by a color change from bright yellow to wine red, indicating the presence of small, non-aggregated nanoparticles. This transformation is attributed to SPR, in which collective oscillations of conduction electrons on the nanoparticle surface interact with light, producing strong absorbance in the visible range (500–560 nm), depending on particle size.30 The optimal synthesis conditions involved 0.01 M o-HBA at pH 12 and heating at 98 °C for 20 min. UV-vis spectrophotometric analysis showed a distinct SPR absorption at 524 nm with an absorbance of 0.687 (see Fig. S2), consistent with previous reports,31 confirming the successful formation of AuNPs under the stated conditions.
3.2. Surface modification of AuNPs
In our previous work, we employed 11-mercaptoundecanoic acid (MUA) since the selected α- and β-cyclodextrins as a reductant and stabilizer was unable to achieve appropriate coupling between AuNPs and EDC/NHS.32,33 Similar to this case, MUA was once used to achieve a comparable effect on modification of AuNPs surface, in which sodium citrate acted as the reducant.34 In the present system, the carboxy group of o-HBA adds to EDC reacts to form an unstable o-acylisourea, which is subsequently attacked by NHS leading to the formation of an NHS ester on AuNPs (AuNPs-EDC/NHS), as illustrated in Fig. 1b. The surface of the AuNPs possesses amine-reactive character, enabling binding with the NS1 monoclonal antibody (mAb).35,36 Consequently, this surface modification method is advantageous because it can omit one reaction step using MUA.
Surface modification is designed to preserve or subtly alter the optical properties of AuNPs. Thus, UV-vis analysis was performed to evaluate the extent of aggregation. The optimal EDC/NHS ratio was determined based on minimal absorbance reduction, indicating minimal nanoparticle aggregation. The best performance was observed at a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 EDC/NHS ratio, yielding a peak at 522 nm with an absorbance of 0.478 (Fig. 2a). The 2 nm red shift to 519 nm and the almost identical absorbance value for the 4
1 EDC/NHS ratio, yielding a peak at 522 nm with an absorbance of 0.478 (Fig. 2a). The 2 nm red shift to 519 nm and the almost identical absorbance value for the 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 EDC/NHS ratio confirm that the SPR characteristics were unaffected. The change of color from wine red to bright pink indicates that the modification was successfully achieved.
1 EDC/NHS ratio confirm that the SPR characteristics were unaffected. The change of color from wine red to bright pink indicates that the modification was successfully achieved.
 |
| | Fig. 2 UV-vis absorption spectra to determine optimum conditions for (a) AuNPs-EDC/NHS at varying molar ratios, (b) AuNPs-mAb bioconjugates at different antibody volumes. | |
The second surface modification of AuNPs-EDC/NHS with NS1 mAb will yield AuNPs-mAb (Fig. 3a). To optimize the bioconjugation process, various concentrations by volume of NS1 mAb were tested to determine the optimal concentration. UV-vis analysis showed that adding 100 µL of mAb resulted in the highest absorbance at 523 nm (Fig. 2b), whereas increasing the volume of mAb decreased absorbance. Hence, this volume and wavelength would be selected for working conditions when binding the NS1 antigen for detection.
 |
| | Fig. 3 (a) Bioconjugation mechanism of AuNPs-EDC/NHS with anti-NS1 mAb, (b) schematic representation of AuNP-mAb aggregation upon NS1 antigen binding. | |
3.3. NS1 antigen detection
The bioconjugation of AuNPs with mAb is expected to facilitate targeted recognition of the NS1 antigen, enabling specific and accurate detection. When the NS1 antibody on the surface of the AuNPs binds to the NS1 antigen, the resulting antigen–antibody complexes form on the AuNP surface (AuNPs-mAb-NS1) (see Fig. 3b). This type of binding leads to a visible color change, as the increase in particle size and subsequent aggregation affect SPR once again.37
Absorbance changes were initially monitored at 523 nm as the working wavelength (result from Fig. 2b) nm using an ELISA reader, with readings taken over 60 minutes. However, the developed sensor demonstrated a rapid response, exhibiting a drop in absorbance upon initial contact with the target antigen and remaining stable thereafter (Fig. 4a). The data indicated that the NS1 antigen was detected within 30 seconds, with visible detection on absorbance decreasing from 0.345 to 0.070 at the highest NS1 antigen concentration added (250 ng mL−1) (see Fig. 4b). This ultrafast response surpassed our previous results, which ranged from 1 to 5 min.32,33 This output underscores the sensor's potential for point-of-care applications, offering a rapid, modest, and user-friendly equipment detection method.
 |
| | Fig. 4 (a) Detection time profile of the AuNPs-NS1 sensor in 60 min, (b) Uv-vis absorption and visual comparison before and after NS1 antigen contact with AuNPs-mAb after 30 seconds. | |
3.4. Characterization of AuNPs
3.4.1. FT-IR. FT-IR was used to analyze covalent bonding, functional groups, and chemical interactions of AuNPs before and after surface modification. Since we previously discussed the transformation of o-HBA into AuNPs, which serves as a reducing agent and stabilizer,24 here we compare the patterns of AuNPs, AuNPs-EDC/NHS, and AuNPs-mAb to identify changes in crucial functional groups (Fig. 5). Synthesized AuNPs showed a sharp C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption at 1731 cm−1, indicating a coordination to Au surface. When AuNPs are treated with EDC/NHS, the absorption at 1731 cm−1 disappeared, replaced by a new signal at 1631 cm−1 that might stand for C
O absorption at 1731 cm−1, indicating a coordination to Au surface. When AuNPs are treated with EDC/NHS, the absorption at 1731 cm−1 disappeared, replaced by a new signal at 1631 cm−1 that might stand for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O from the eliminated urea. Meanwhile, a similar study reported that 1572 cm−1 once referred to C
O from the eliminated urea. Meanwhile, a similar study reported that 1572 cm−1 once referred to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide, following a -COO-NHS ester formation.38 Eventually, these spectral changes confirmed the successful AuNPs embedding with EDC/NHS. Next, the detection of antibody binding is evidenced by the amide I band at 1637 cm−1 for C
O amide, following a -COO-NHS ester formation.38 Eventually, these spectral changes confirmed the successful AuNPs embedding with EDC/NHS. Next, the detection of antibody binding is evidenced by the amide I band at 1637 cm−1 for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O protein backbone (the typical assignment for β-sheet ranged from 1618–1642 cm−1),39 the amide II band at 1590 cm−1 for primary N–H bending,40 and the C–N stretch at 1154 cm−1, collectively demonstrating effective surface functionalization of AuNPs-mAb.
O protein backbone (the typical assignment for β-sheet ranged from 1618–1642 cm−1),39 the amide II band at 1590 cm−1 for primary N–H bending,40 and the C–N stretch at 1154 cm−1, collectively demonstrating effective surface functionalization of AuNPs-mAb.
 |
| | Fig. 5 Characteristic FT-IR absorption comparison of (a) AuNPs, (b) AuNPS-EDC/NHS, and (c) AuNPs-mAb. | |
3.4.2. TEM and FESEM EDX. TEM was used to observe particle size and morphology. As shown, AuNPs synthesized with o-HBA were spherical, uniform, and had an average diameter of 19.75 nm, with no visible aggregation (Fig. 6a). This size is smaller than previously reported values (25.04 nm),24 indicating enhanced stability. After surface modification as AuNPs-mAb, the average particle size increased slightly to 19.85 nm, with particles clustering more closely (Fig. 6b). In the AuNPs-NS1 antigen complex, larger aggregates formed with an average size of 23.24 nm, attributed to EDC/NHS and antibody-induced particle interactions (Fig. 6c).
 |
| | Fig. 6 TEM images (a) AuNPs, (b) AuNPs-mAB, and (c) AuNPs-mAb-NS1 conjugates with their corresponding particle size. | |
On the other hand, FESEM and EDX were deployed to evaluate surface morphology and elemental composition after each modification (Fig. 7a–c). Although FESEM results generally confirmed TEM findings of spherical morphology across all samples, the measured sizes of all nanoparticles appeared similar, around 17 nm, despite AuNPs-mAb and AuNPs-mAb-NS1 having their surfaces covered. This unusual result, where FESEM size measurements were smaller than TEM due to clustered nanoparticles following surface modification, led to dense surface layering, making distinct individual particles difficult to detect.
 |
| | Fig. 7 FESEM images of (a) AuNPS, (b) AuNPS-mAb, and (c) AuNPs-mAb-NS1 with their corresponding EDX analysis graphs. | |
For EDX analysis, the unmodified AuNPs showed peaks only for C (69.8%), O (21.7%), and Au (8.5%) (Fig. 7a). After bioconjugation, nitrogen (N, 0.3%) started to appear in AuNPs-mAb, indicating protein species presence, while the rest of the elemental composition gave Au (37%), O (31.8%), and C (31%) (Fig. 7b). The enhancement of Au composition is understandable, as the element distribution in the screened area is derived from a dense environment. In contrast, elemental screening for AuNPs-mAb-NS1 showed elevated N (1.7%) values but no detection of Au. The FESEM image in Fig. 7c explains the disappearance of Au. As the samples became sufficiently thick, organic coverage caused the Au peaks to fall below the detection limit, rendering them undetectable. Nevertheless, the entire interpretation could justify the conclusion that the surface modification and bioconjugation was successful.
3.5. Validation results
3.5.1. Sensitivity. The linearity test assessed the sensitivity of AuNPs-mAb in detecting varying concentrations of NS1 antigen. Absorbance at the working wavelength of 523 nm was recorded using an ELISA Reader. As the antigen concentration increased, a corresponding increase in Δ absorbance was observed due to SPR changes. The calibration curve yielded a linear regression equation: y = 0.0001x + 0.0094, with an R2 of 0.982 and a correlation coefficient (r) of 0.993, indicating excellent linearity and detection accuracy (Fig. 8). According to the standard deviation and slope of the calibration curve, the Limit of Detection (LoD) and Limit of Quantification (LoQ) were calculated. LoD refers to the minimum detectable concentration,41 while LoQ indicates the lowest quantifiable concentration.42 In this study, the LoD was 49.65 ng mL−1, and the LoQ was 165.48 ng mL−1 (Table S2). These values were compared with those reported in previous studies targeting the NS1 antigen (Table 3) to compare the relative performance of the developed sensor.
 |
| | Fig. 8 Calibration curve showing linear response of sensor when varying NS1 antigen concentrations. | |
Table 3 Comparison of several biosensor methods for NS1 antigen detection
| Method |
Material |
Detection time |
Linear range (ng mL−1) |
LoD (ng mL−1) |
| SPCE: screen-printed carbon electrodes, BSA: bovine serum albumin. TAS: thermally annealed silver, NS: nanostructures. Apt: aptamer. MIP: molecularly imprinted polymer, QDs: quantum dots, SPCEs: screen-printed carbon electrodes. ITO: indium tin oxide. AgNPs: reduced by trisodium citrate. AuNPs: reduced by α-cyclodextrins. AuNPs reduced by β-cyclodextrins. AuNPs: reduced with o-HBA. |
| Electrochemical15 |
SPCE modified with BSAa |
— |
1–200 |
0.3 |
| Localized surface plasmon resonance43 |
TAS-NSb |
30 min |
0–50000 |
1.1 |
| Aptamer44 |
AuNPs-Apt1 and AuNPs-Apt2c |
— |
1 × 10−3–1 |
0.00003 |
| Electropolymerized45 |
MIP@InZnSe@PtAg-QDs/SPCEsd |
— |
1 × 10−6–1 × 10−1 |
0.00136 |
| Immunofluorescence46 |
ITO modified with mAb and pAbe |
— |
15–500 |
15 |
| Flow chromatography47 |
AgNPs-NS1f |
— |
0–500 |
150 |
| Colorimetric32 |
AuNPs-NS1g |
5 min |
75–250 |
22.02 |
| Colorimetric33 |
AuNPs-NS1h |
1 min |
75–250 |
39.29 |
| Colorimetricthis work |
AuNPs-NS1i |
30 seconds |
75–250 |
49.65 |
Because the detection limit reflects the method's sensitivity, lower LoD values indicate higher sensitivity. Even though this study used a colorimetric method with a higher LoD than the listed techniques,15,43–46 in which resulted under a more advanced and sophisticated procedure, it still significantly outperformed other methods in terms of detection time, achieving NS1 antigen detection in just 30 seconds, even compared to the similar colorimetric techniques. Moreover, the comparison with a relatively similar facile technique such as flow chromatography,47 still gave the lower LoD value than the latter, emphasizing its advantage for rapid screening result, suitable for limited-resources setting.
3.5.2. Repeatability and reproducibility. Repeatability (intra-assay precision) and reproducibility (inter-assay precision) were evaluated to assess the sensor's consistency.48 Absorbance at 523 nm was recorded using an ELISA Reader for 100 ng mL−1 AuNPs-mAb with NS1 antigen at 75–250 ng mL−1. Repeatability was measured in triplicate on the same day, while reproducibility was assessed across three separate days (see Tables S3 and S4). According to AOAC and Horwitz guidelines,28 acceptable %RSD limits are 30% and 45%, respectively. As the result, all values fell within these limits (Tables 4 and 5), confirming good precision.
Table 4 Results repeatability tests of AuNPs-mAb-NS1 at 523 nm
| Concentration (ng mL−1) |
Average absorbance |
Standard deviation |
RSD (%) |
| AOAC |
Horwitz |
| 75 |
0.081 |
0.0004 |
0.473 |
1.044 |
| 100 |
0.078 |
0.0001 |
0.133 |
1 |
| 150 |
0.071 |
0.0005 |
0.751 |
0.941 |
| 200 |
0.066 |
0.0006 |
0.939 |
0.901 |
| 250 |
0.063 |
0.0005 |
0.814 |
0.871 |
Table 5 Results of reproducibility test of AuNPs-mAb-NS1 at 523 nm
| Concentration (ng mL−1) |
Average absorbance |
Standard deviation |
RSD (%) |
| AOAC |
Horwitz |
| 75 |
0.081 |
0.0005 |
0.630 |
1.044 |
| 100 |
0.079 |
0.0008 |
0.999 |
1 |
| 150 |
0.071 |
0.0002 |
0.299 |
0.941 |
| 200 |
0.066 |
0.0003 |
0.472 |
0.901 |
| 250 |
0.062 |
0.0005 |
0.773 |
0.871 |
3.5.3. Selectivity. The selectivity as a biosensor was evaluated using potential interfering substances, including aquabidest, glucose, cholesterol, and uric acid, which are commonly present in blood. Only the NS1 antigen showed a significant absorbance change (Δ = 0.013) (Fig. 9). This demonstrates high specificity of the sensor for the NS1 antigen with minimal cross-reactivity, making it a strong candidate for point-of-care diagnostic applications in clinical settings.
 |
| | Fig. 9 Selectivity comparison of the sensor across different matrix samples. | |
3.5.4. Application in human serum. Human serum was used to validate the biosensor's practical performance in real biological matrices because serum proteins influence fluid regulation and may affect sensor behaviour. Consequently, evaluating sensor response in human serum provides insights into its potential clinical applications.49As shown in Fig. 10, NS1-negative serum produced a negligible Δ absorbance, while both NS1-positive serum and pure NS1 solution yielded positive Δ absorbance responses (see Table S7). These results suggest that the sensor does not produce false positives and can reliably detect the NS1 antigen even in complex biological matrices. Minor response variation may be attributed to matrix effects in human serum that influence antigen–antibody interactions.
 |
| | Fig. 10 Performance comparison of the biosensor in human serum samples. | |
3.5.5. Stability. Periodic monitoring of the AuNPs-mAb sensor's signal output in response to a standard amount of NS1 as the analyte over an extended period was used as a stability test. This observation is crucial for point-of-care implementation,50 since it assesses long-term stability to determine the product's storage lifetime.51 Upon monitoring absorbance and SPR peak shifts at 523 nm under storage period in cold temperature (4 °C), the major decline around 0.023 a.u. occurs from the initial contact into the 1-day after storage (Fig. 11). After this, the consecutive 7-day period showed only slight ups and downs in absorbance, suggesting that slight nanoparticle aggregation, driven by surface charge density and enhanced van der Waals interactions, might have caused the alteration. Overall, the results indicate high storage stability from the developed biosensor up to 28 days.
 |
| | Fig. 11 Stability profile of the sensor during storage over time. | |
4. Conclusion
An alternative approach for detecting the dengue virus was successfully developed using a colorimetric method based on AuNPs. These AuNPs were synthesized using o-HBA as both a reducing and stabilizing agent, producing a wine red-colored solution with a characteristic SPR peak at 524 nm. The resulting AuNPs were uniformly spherical with an average diameter of 19.75 nm. Surface sensitivity was enhanced by EDC/NHS modification at an optimal 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, yielding a bright pink solution with an SPR peak at 522 nm. Bioconjugation was carried out with optimum volume used as 100 µL. The resulting solution retained its bright pink color and SPR peak shifted to 523 nm, indicating a stable conjugation. A slight increase in particle size from 19.75 nm (AuNPs) to 19.85 nm (AuNPs-mAb) was also observed, attributed to closer interparticle spacing caused by antibody binding. When the developed colorimetric sensor successfully detected the NS1 antigen within 30 seconds, a visible color change from bright pink to colorless was easily spotted. The method demonstrated a LoD of 49.65 ng mL−1 and a LoQ of 165.48 ng mL−1. These promising results indicate that the colorimetric sensor has the potential to be developed for point-of-care applications, given the simplicity of the detection procedure. Nonetheless, the study's limitations, such as validation only in the laboratory with human serum and potential cross-reactivity with other arboviruses like ZIKV and CHIKV, should be acknowledged. Future work should focus on validating the sensor's performance using real blood samples from suspected dengue patients to ensure reliability in clinical settings.
1 ratio, yielding a bright pink solution with an SPR peak at 522 nm. Bioconjugation was carried out with optimum volume used as 100 µL. The resulting solution retained its bright pink color and SPR peak shifted to 523 nm, indicating a stable conjugation. A slight increase in particle size from 19.75 nm (AuNPs) to 19.85 nm (AuNPs-mAb) was also observed, attributed to closer interparticle spacing caused by antibody binding. When the developed colorimetric sensor successfully detected the NS1 antigen within 30 seconds, a visible color change from bright pink to colorless was easily spotted. The method demonstrated a LoD of 49.65 ng mL−1 and a LoQ of 165.48 ng mL−1. These promising results indicate that the colorimetric sensor has the potential to be developed for point-of-care applications, given the simplicity of the detection procedure. Nonetheless, the study's limitations, such as validation only in the laboratory with human serum and potential cross-reactivity with other arboviruses like ZIKV and CHIKV, should be acknowledged. Future work should focus on validating the sensor's performance using real blood samples from suspected dengue patients to ensure reliability in clinical settings.
Author contributions
Annisa Indah Reza rewriting the manuscript and editing figures. Fitri Fadila conducted data curation and writing – original draft. Dita Cinta Toharani and Agustina Vidiawati performed experiments and formal analysis. Dita Ariyanti helped by supervised and validated the data. Nagatoshi Nishiwaki provided valuable corrections, suggestions, and supervision. Agustina Sus Andreani provided funding acquisition, supervised the research, validation of data, reviewed and editing the manuscript. All authors discussed the results and commented on the manuscript accordingly.
Conflicts of interest
All authors declare no conflicts of interest.
Data availability
All data needed to evaluate the conclusions in the paper are present in the paper and/or the supplementary information (SI). Additional data related to this paper can be requested from the corresponding author. Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra10028c.
Acknowledgements
This research was funded by Research Organization for Health from National Research and Innovation Agency (BRIN) (5/III.9/HK/2025), along with provider of facilities, and technical support through E-Layanan Sains (ELSA) system.
References
- M. A. Kabir, H. Zilouchian, M. A. Younas and W. Asghar, Biosensors, 2021, 11, 206 CrossRef CAS PubMed.
- M. R. Hasan, P. Sharma, S. Khan, U. Mushtaq Naikoo, K. Bhalla, M. Z. Abdin, N. Malhotra, T. M. Aminabhavi, N. P. Shetti and J. Narang, Sens. Diagn., 2024, 4(1), 7–23 Search PubMed.
- S. Halstead, F1000Research, 2019, 8, 1–12 Search PubMed.
- S. Vita, E. Lalle, P. Caputi, F. Faraglia, A. D'Abramo, L. Bordi, G. De Carli, G. Sberna, M. L. Giancola, G. Maffongelli, C. Mija, A. Antinori, S. Cicalini, F. Maggi, E. Girardi, F. Vairo and E. Nicastri, Travel Med. Infect. Dis., 2024, 62, 102762 CrossRef CAS.
- T. R. P. Lestari, Strategic Analysis on Current Issue, 2025, 27(11), 1–5.
- C. R. Basso, C. C. Tozato, B. P. Crulhas, G. R. Castro, J. P. A. Junior and V. A. Pedrosa, Virology, 2018, 513, 85–90 Search PubMed.
- H. B. Y. Chan, C. H. How and C. W. M. Ng, Singap. Med. J., 2017, 58(11), 632–635 CrossRef PubMed.
- H. Xu, B. Di, Y. Pan, L. Qiu, Y. Wang and W. Hao, J. Clin. Microbiol., 2006, 44(8), 2872–2878 CrossRef CAS PubMed.
- P. S. Tsai, P. X. Du, B. B. Keskin, N. Y. Lee, S. W. Wan, Y. L. Lin, H. C. Shih, G. Syu and T. S. Ho, Microchem. J., 2024, 200, 110390 CrossRef CAS.
- V. Liberal, R. Forrat, C. Zhang, C. Pan, M. Bonaparte, W. Yin, L. Zheng, V. Viscardi, Y. Wu, Y. Ataman-Önal, S. Savarino and C. Chen, Microbiol. Spectr., 2022, 10(3), 1–8 CAS.
- A. Agarwal, R. K. Jain, D. Chaurasia and D. Biswas, Indian J. Med. Microbiol., 2022, 40(4), 492–495 CrossRef CAS PubMed.
- N. Lukman, G. Salim, H. Kosasih, N. H. Susanto, I. Parwati, S. Fitri, B. Alisjahbana, S. Widjaja and M. Williams, BioMed Res. Int., 2016, 1, 5253842 Search PubMed.
- P. Keelapang, R. Kraivong, R. Pulmanausahakul, R. Sriburi, E. Prompetchara, J. Kaewmaneephong, N. Charoensri, P. Pakchotanon and T. Duangchinda, Microbiol. Spectr., 2023, 11(4), e0091823 Search PubMed.
- P. Kulkarni, S. Taklikar and D. Turbadkar, Indian J. Med. Microbiol., 2023, 45, 100376 Search PubMed.
- C. C. Santos, P. C. M. Santos, K. L. S. Rocha, R. L. D. Thomasini and L. F. Ferreira, Microchem. J., 2020, 154, 104544 CrossRef CAS.
- C. Coronel-Ruiz, M. L. Velandia-Romero, E. Calvo, S. Camacho-Ortega, S. Parra-Alvarez, E. O. Beltrán, M. A. Calderón-Pelaez, A. Porras-Ramírez, F. Cortés-Muñoz, J. P. Rojas-Hernandez, S. Velasco-Alvarez, A. Pinzón-Junca and J. E. Castellanos, Front. Trop. Dis., 2023, 4, 1–15 Search PubMed.
- Y. Choi, J. H. Hwang and S. Y. Lee, Small Methods, 2018, 2, 1700531 CrossRef.
- A. Roda, E. Michelini, M. Zangheri, M. Di Fusco, D. Calabria and P. Simoni, Trends Anal. Chem., 2016, 79, 317–325 CrossRef CAS.
- C. Wang, M. Liu, Z. Wang, S. Li, Y. Deng and N. He, Nano Today, 2021, 37, 101092 CrossRef CAS PubMed.
- V. X. T. Zhao, et al., Mater. Sci. Energy Technol., 2020, 3, 237–249 CAS.
- X. Huang, P. K. Jain, E. H. El-Sayed and M. A. El-Sayed, Nanomedicine, 2007, 2(5), 681–693 CrossRef CAS.
- H. Chen, S. Cai, J. Luo, X. Liu, L. Ou, Q. Zhang, B. Liedberg and Y. Wang, Trends Anal. Chem., 2024, 173, 1–18 CrossRef.
- R. Eivazzadeh-Keihan, Z. Saadatidizaji, M. Mahdavi, A. Maleki, M. Irani and I. Zare, Talanta, 2024, 275, 126099 CrossRef CAS PubMed.
- A. S. Andreani, E. S. Kunarti, T. Hashimoto, T. Hayashita and S. J. Santosa, J. Environ. Chem. Eng., 2021, 9, 105962 CrossRef CAS.
- A. S. Andreani, M. Wulandari, I. Istianah and I. Yati, AIP Conf. Proc., 2023, 2902, 080002 CrossRef CAS.
- D. Bartczak and A. G. Kanaras, Langmuir, 2011, 27, 10119–10123 CrossRef CAS PubMed.
- S. A. F. Kusuma, J. A. Harmonis, R. Pratiwi and A. N. Hasanah, Sensors, 2023, 23, 8172 CrossRef CAS PubMed.
- H. Moulahoum and F. Ghorbanizamani, Biosens. Bioelectron., 2024, 264, 116670 CrossRef CAS PubMed.
- Association of Official Analytical Chemists (AOAC) International, Appendix F: Guidelines for Standard Method Performance Requirements, Off. Methods Anal. AOAC Int., 2016 Search PubMed.
- P. K. Ngumbi, S. W. Mugo and J. M. Ngaruiya, IOSR J. Appl. Chem., 2018, 11(7), 25–29 CAS.
- D. K. Nguyen and C. H. Jang, Micromachines, 2021, 12(12), 1–11 Search PubMed.
- A. S. Andreani, S. U. Zahrob, A. Vidiawati and A. Fathoni, Anal. Bioanal. Chem., 2025, 12(4), 457–472 CAS.
- A. I. Alfiyan, A. S. Andreani, T. Setyaningtyas, B. Yuliarto and M. Angelina, Anal. Methods, 2025, 17, 5238–5249 RSC.
- R. Zhu, J. Song, Y. Zhou, P. Lei, Z. Li, H. W. Li, S. Shuang and C. Dong, Talanta, 2019, 204, 294–303 CrossRef CAS PubMed.
- D. Di Iorio and J. Huskens, ChemistryOpen, 2020, 9(1), 53–66 CrossRef CAS PubMed.
- A. Hushegyi, T. Bertok, P. Damborsky, J. Katrlik and J. Tkac, Chem. Commun., 2015, 51, 7474–7477 RSC.
- H. Mateos, A. Mallardi, E. Serrano-Pertierra, M. C. Blanco-López, M. Izzi, N. Cioffi and G. Palazzo, JCIS Open, 2023, 11, 100089 CrossRef.
- D. Rath, S. Kumar and S. Panda, Appl. Biochem. Biotechnol., 2019, 187, 1272–1284 CrossRef CAS PubMed.
- J. Dai, C. Chen, M. Yin, H. Li, W. Li and Y. Wang, Front. Chem., 2023, 11, 1273388 CrossRef CAS PubMed.
- J. Zhao, Y. Ruan, Z. Zheng, Y. Li, M. Sohail, F. Hu and J. Z. L. Ling, iScience, 2023, 26, 106823 CrossRef CAS PubMed.
- G. L. Long and J. D. Winefordner, Limit of Detection a Closer Look at the IUPAC Definition, Anal. Chem., 1983, 55, 712A–724A Search PubMed.
- B. Magnusson and U. Omemark, Eurachem Guide, The Fitness for Purpose of Analytical Methods–A Laboratory Guide to Method Validation & Related Topic, 2014, p. 21 Search PubMed.
- P. P. A. Suthanthiraraj and A. K. Sen, Biosens. Bioelectron., 2019, 132, 38–46 CrossRef.
- I. M. Khoris, F. Nasrin, A. D. Chowdhury and E. Y. Park, Anal. Chim. Acta, 2022, 1207, 339817 CrossRef CAS PubMed.
- K. Isla Gray and O. Adegoke, Microchem. J., 2024, 207, 111704 CrossRef CAS.
- N. T. Darwish, S. D. Sekaran, Y. Alias and S. M. Khor, J. Pharm. Biomed. Anal., 2018, 149, 591–602 CrossRef CAS.
- C. W. Yen, H. Puig, J. O. Tam, J. Gómez-Márquez, I. Bosch, K. Hamad-Schifferli and L. Gehrke, Lab Chip, 2015, 15, 1638–1641 RSC.
- R. Voet, J. A. Rhijn and H. J. Wiel, Anal. Chim. Acta, 1999, 391(2), 159–171 CrossRef.
- M. E. Rosa, M. S. M. Mendes, D. C. V. Belchior, J. A. P. Coutinho, F. A. Silva and M. G. Freire, Adv. Sample Prep., 2024, 10, 100116 Search PubMed.
- F. Saira, S. Saleemi, H. Razzaq and R. Qureshi, Turk. J. Chem., 2021, 45(1), 82–91 Search PubMed.
- A. Klos-Witkowska and V. Martsenyuk, Acta Biochim. Pol., 2024, 71, 12196 CrossRef CAS PubMed.
|
| This journal is © The Royal Society of Chemistry 2026 |
Click here to see how this site uses Cookies. View our privacy policy here.  Open Access Article
Open Access Article a,
Fitri Fadilab,
Dita Cinta Toharanic,
Dita Ariyanti
a,
Fitri Fadilab,
Dita Cinta Toharanic,
Dita Ariyanti d,
Agustina Vidiawatie,
Nagatoshi Nishiwaki
d,
Agustina Vidiawatie,
Nagatoshi Nishiwaki *fg and
Agustina Sus Andreani
*fg and
Agustina Sus Andreani *a
*a
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio) enabled efficient antibody conjugation to NS1 protein monoclonal antibodies. Colorimetric detection with the NS1 antigen provided results within 30 seconds, exhibited excellent linearity (R2 = 0.982), and demonstrated limits of detection and quantification of 49.65 ng mL−1 and 165.48 ng mL−1, respectively. This AuNP-based biosensor offers a fast, accurate, and user-friendly platform, potential for point-of-care dengue diagnosis, with nanoparticles remaining stable for up to 28 days.
1 ratio) enabled efficient antibody conjugation to NS1 protein monoclonal antibodies. Colorimetric detection with the NS1 antigen provided results within 30 seconds, exhibited excellent linearity (R2 = 0.982), and demonstrated limits of detection and quantification of 49.65 ng mL−1 and 165.48 ng mL−1, respectively. This AuNP-based biosensor offers a fast, accurate, and user-friendly platform, potential for point-of-care dengue diagnosis, with nanoparticles remaining stable for up to 28 days.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 DF cases and 250 deaths across 456 districts/cities, representing 87% of Indonesia's territory.5 Adding to concerns, many high-risk areas in Indonesia face resource limitations, making it challenging to access necessary detection and monitoring equipment.
000 DF cases and 250 deaths across 456 districts/cities, representing 87% of Indonesia's territory.5 Adding to concerns, many high-risk areas in Indonesia face resource limitations, making it challenging to access necessary detection and monitoring equipment.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 ratio. The optimal AuNPs-EDC/NHS conjugate was prepared by mixing varying volumes of diluted NS1 mAb (Table 2) and stirring at 200 rpm for 1 h at 37 °C. UV-vis analysis (200–800 nm) was used to evaluate the optimal bioconjugation conditions, followed by characterization using FT-IR, TEM, and FESEM.
100 ratio. The optimal AuNPs-EDC/NHS conjugate was prepared by mixing varying volumes of diluted NS1 mAb (Table 2) and stirring at 200 rpm for 1 h at 37 °C. UV-vis analysis (200–800 nm) was used to evaluate the optimal bioconjugation conditions, followed by characterization using FT-IR, TEM, and FESEM.

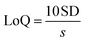
![[x with combining macron]](https://www.rsc.org/images/entities/i_char_0078_0304.gif) indicates average of data set, and C indicates concentration of analyte.29
indicates average of data set, and C indicates concentration of analyte.29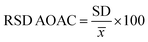
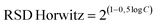

![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 EDC/NHS ratio, yielding a peak at 522 nm with an absorbance of 0.478 (Fig. 2a). The 2 nm red shift to 519 nm and the almost identical absorbance value for the 4
1 EDC/NHS ratio, yielding a peak at 522 nm with an absorbance of 0.478 (Fig. 2a). The 2 nm red shift to 519 nm and the almost identical absorbance value for the 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 EDC/NHS ratio confirm that the SPR characteristics were unaffected. The change of color from wine red to bright pink indicates that the modification was successfully achieved.
1 EDC/NHS ratio confirm that the SPR characteristics were unaffected. The change of color from wine red to bright pink indicates that the modification was successfully achieved.


![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption at 1731 cm−1, indicating a coordination to Au surface. When AuNPs are treated with EDC/NHS, the absorption at 1731 cm−1 disappeared, replaced by a new signal at 1631 cm−1 that might stand for C
O absorption at 1731 cm−1, indicating a coordination to Au surface. When AuNPs are treated with EDC/NHS, the absorption at 1731 cm−1 disappeared, replaced by a new signal at 1631 cm−1 that might stand for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O from the eliminated urea. Meanwhile, a similar study reported that 1572 cm−1 once referred to C
O from the eliminated urea. Meanwhile, a similar study reported that 1572 cm−1 once referred to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide, following a -COO-NHS ester formation.38 Eventually, these spectral changes confirmed the successful AuNPs embedding with EDC/NHS. Next, the detection of antibody binding is evidenced by the amide I band at 1637 cm−1 for C
O amide, following a -COO-NHS ester formation.38 Eventually, these spectral changes confirmed the successful AuNPs embedding with EDC/NHS. Next, the detection of antibody binding is evidenced by the amide I band at 1637 cm−1 for C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O protein backbone (the typical assignment for β-sheet ranged from 1618–1642 cm−1),39 the amide II band at 1590 cm−1 for primary N–H bending,40 and the C–N stretch at 1154 cm−1, collectively demonstrating effective surface functionalization of AuNPs-mAb.
O protein backbone (the typical assignment for β-sheet ranged from 1618–1642 cm−1),39 the amide II band at 1590 cm−1 for primary N–H bending,40 and the C–N stretch at 1154 cm−1, collectively demonstrating effective surface functionalization of AuNPs-mAb.


![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, yielding a bright pink solution with an SPR peak at 522 nm. Bioconjugation was carried out with optimum volume used as 100 µL. The resulting solution retained its bright pink color and SPR peak shifted to 523 nm, indicating a stable conjugation. A slight increase in particle size from 19.75 nm (AuNPs) to 19.85 nm (AuNPs-mAb) was also observed, attributed to closer interparticle spacing caused by antibody binding. When the developed colorimetric sensor successfully detected the NS1 antigen within 30 seconds, a visible color change from bright pink to colorless was easily spotted. The method demonstrated a LoD of 49.65 ng mL−1 and a LoQ of 165.48 ng mL−1. These promising results indicate that the colorimetric sensor has the potential to be developed for point-of-care applications, given the simplicity of the detection procedure. Nonetheless, the study's limitations, such as validation only in the laboratory with human serum and potential cross-reactivity with other arboviruses like ZIKV and CHIKV, should be acknowledged. Future work should focus on validating the sensor's performance using real blood samples from suspected dengue patients to ensure reliability in clinical settings.
1 ratio, yielding a bright pink solution with an SPR peak at 522 nm. Bioconjugation was carried out with optimum volume used as 100 µL. The resulting solution retained its bright pink color and SPR peak shifted to 523 nm, indicating a stable conjugation. A slight increase in particle size from 19.75 nm (AuNPs) to 19.85 nm (AuNPs-mAb) was also observed, attributed to closer interparticle spacing caused by antibody binding. When the developed colorimetric sensor successfully detected the NS1 antigen within 30 seconds, a visible color change from bright pink to colorless was easily spotted. The method demonstrated a LoD of 49.65 ng mL−1 and a LoQ of 165.48 ng mL−1. These promising results indicate that the colorimetric sensor has the potential to be developed for point-of-care applications, given the simplicity of the detection procedure. Nonetheless, the study's limitations, such as validation only in the laboratory with human serum and potential cross-reactivity with other arboviruses like ZIKV and CHIKV, should be acknowledged. Future work should focus on validating the sensor's performance using real blood samples from suspected dengue patients to ensure reliability in clinical settings.





