Open Access Article
Open Access ArticleMachine learning-based QSAR modeling assay for the nitrification inhibition of 2,4,5-trichloroaniline-derived eco-friendly Schiff bases
Tilak Mondal*a,
Rajesh Kumarb,
Santanu Mukherjee *cd,
Nilimesh Mridhaa,
Namrata Laskara,
Avijit Dasa,
Pradip Basake,
Atul Singhaa,
Kanchan Sinhaf and
Dinesh Babu Shakyawara
*cd,
Nilimesh Mridhaa,
Namrata Laskara,
Avijit Dasa,
Pradip Basake,
Atul Singhaa,
Kanchan Sinhaf and
Dinesh Babu Shakyawara
aICAR-National Institute of Natural Fibre Engineering and Technology, Kolkata, India. E-mail: tilakmondal1987@gmail.com
bIndian Agricultural Research Institute, New Delhi, India
cSchool of Agriculture Sciences, Shoolini University of Biotechnology and Management Sciences, Bajhol, PO Sultanpur, Solan 173229, Himachal Pradesh, India. E-mail: santanu@shooliniuniversity.com
dDepartment of Environmental and Civil Engineering, Faculty of Engineering, Toyama Prefectural University, 5180 Kurokawa, Imizu, Toyama 939-0398, Japan
eUttar Bangha Krishi Viswavidyalaya, Cooch Behar, West Bengal, India
fICAR-Indian Agricultural Statistics Research Institute, New Delhi, India
First published on 23rd February 2026
Abstract
Twenty-one Schiff bases were synthesized from 2,4,5-trichloroaniline and characterized by instrumental techniques such as 1H NMR, 13C NMR and IR spectroscopy. The compounds were tested for their ability to inhibit nitrification in soil nitrifying bacteria under controlled laboratory conditions over 28 days at three different concentrations (1%, 5%, and 10%), with N-serve and Dicyandiamide (DCD) used as nitrification inhibitors. All the tested compounds showed markedly higher ammonium-N levels (33–185 mg kg−1) and lower nitrate-N levels (23–137 mg kg−1) compared to the control treatment with urea alone. The synthesized compounds were found to be extremely efficient nitrification inhibitors (15–77%) and enhanced soil microbial activities including soil respiration (32.2–34.0 µg CO2-C per g), dehydrogenase activity (43–46.4 µg TPF per g soil per 24 h) and microbial biomass carbon (537.2–556.3 µg gm−1), compared to other treatments, demonstrating superior microbial activity and soil health. The most important compounds identified from the series were C21, C20, and C3. Pair t-test was performed within the treatment concentration and found that all the treatments differed significantly in terms of mean responses from each other. Statistical and machine leaning models such as stepwise multiple linear regression, artificial neural network (ANN), support vector regression and random forest were used to validate the quantitative structure activity relationship (QSAR) of the synthesized compounds with their nitrification inhibition property. ANN models developed based on feature variables selected from random forest regression were found to be the best fitted models for predicting nitrification inhibition activity at 1%, 5% and 10% of treatment doses.
1. Introduction
Nitrogen (N) is the most important primary nutrient and considered an essential element for plant growth and development. Indian soils generally have low levels of plant-available nitrogen, with a nitrogen content of approximately 0.05%.1 Therefore, application of chemical fertilizer in a judicious amount is a pre-requisite to fulfill the crop N requirement to feed nearly 1.4 billion people.2 Nitrogen fertilizers are usually supplied in three forms, urea, ammonia and nitrate. Urea undergoes enzymatic hydrolysis and is converted into ammonia and subsequently to ammonium. The plants uptake N either in ammonium (NH4+) or nitrate (NO3−) form and leftover NH4+ is converted into NO3− by the nitrification process which is facilitated by the chemolithoautotrophic groups of soil bacteria, namely Nitrosomonas and Nitrobacter. Nitrification is the biological process that converts NH4+ or nitrogenous fertilizers into nitrite (NO2−) and then into NO3−. However, excessive nitrification and the rapid formation of nitrate can lead to fertilizer waste and groundwater pollution.3,4 Nitrogenous fertilizers, especially urea, have low nitrogen use efficiency (NUE), with only 20–30% being utilized, due to losses from ammonia volatilization, nitrate leaching, and denitrification.5 Cardoen et al. (2015) discussed that up to 60–80% of nitrogen from urea fertilizers can be lost, highlighting the need for better nitrogen management.6 In wetland paddy fields, only 30–40% of applied nitrogen is used effectively, with the rest lost through emissions, runoff, or leaching.7,8 Therefore, improvement of nutrient management is essential to increase the crop productivity which can be achieved by increasing NUE of applied fertilizer. The low NUE due to nitrification and other losses worth Rs. 85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 crores and impose significant impact on nation's economy and environment.9,10
000 crores and impose significant impact on nation's economy and environment.9,10
For increasing the NUE, use of nitrification inhibitors along with nitrogenous fertilizers is the most suitable approach among the different cultural and chemical management practices followed in the field.11–13 Nitrification inhibitors are substances designed to slow down the conversion of ammonium to nitrite by specifically targeting Nitrosomonas bacteria. These inhibitors do not affect the subsequent transformation of nitrite to nitrate. By delaying the oxidation of ammonium-N to nitrite-N, they enhance NUE in soil. Several commercial nitrification inhibitors are available to reduce nitrogen losses from fertilizers, targeting the enzyme ammonia monooxygenase (AMO), which catalyzes the initial oxidation of ammonia to hydroxylamine.14 Common inhibitors like nitrapyrin, dicyandiamide, and etridiazole15 are incorporated into fertilizers to improve NUE in crops. However, these inhibitors are not widely adopted at the farmer level due to their environmental persistence, potential impact on non-target soil microorganisms,16 and issues like complex synthesis, high toxicity, low stability, high application rates, and costly formulations.
Schiff bases are widely utilized in enhancing human welfare and agricultural systems due to their broad range of biological activities. These compounds are known for their antibacterial,17 antiviral, and antimicrobial properties.18,19 Many compounds containing the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N group have been identified as effective nitrification inhibitors20,21 and have demonstrated antifungal, pesticidal activities.22,23 The synthesis and study of imines, formed by the reaction of carbonyl compounds with primary amines have garnered significant interest in recent years. The carbon–nitrogen double bond in imines provides a key site for both chemical and biological reactivity.19,25 The conventional synthesis of Schiff bases is straightforward and environmentally friendly.24,25 To explore the wide spectrum bioactive potential of Schiff bases, our present study was formulated. In this novel study, the nitrification inhibitory activity of synthesized 2,4,5-trichloro aniline based Schiff bases against the nitrifying soil bacteria have been reported. We have also validated their nitrification inhibition activity with the developed QSAR models by multiple linear regression analysis using their various distance based topological indices, steric and hydrophobic parameters. The models were justified with the application of principal component regression analysis.
N group have been identified as effective nitrification inhibitors20,21 and have demonstrated antifungal, pesticidal activities.22,23 The synthesis and study of imines, formed by the reaction of carbonyl compounds with primary amines have garnered significant interest in recent years. The carbon–nitrogen double bond in imines provides a key site for both chemical and biological reactivity.19,25 The conventional synthesis of Schiff bases is straightforward and environmentally friendly.24,25 To explore the wide spectrum bioactive potential of Schiff bases, our present study was formulated. In this novel study, the nitrification inhibitory activity of synthesized 2,4,5-trichloro aniline based Schiff bases against the nitrifying soil bacteria have been reported. We have also validated their nitrification inhibition activity with the developed QSAR models by multiple linear regression analysis using their various distance based topological indices, steric and hydrophobic parameters. The models were justified with the application of principal component regression analysis.
2. Materials and methods
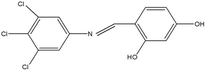
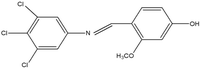
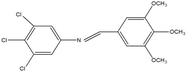
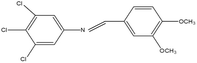
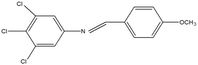
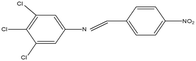
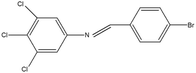
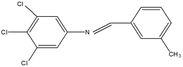
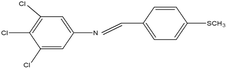
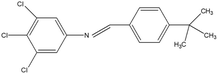
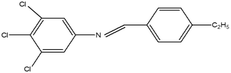
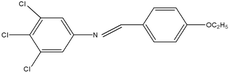
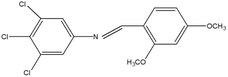
2,4,5-Trichloroaniline, twenty-one different aldehydes, methanol, ethanol, hexane, ethyl acetate, nitrapyrin, Dicyandiamide (DCD), sodium sulphate, phenol crystal, sodium nitroprusside, hypochlorite (alkaline), ammonium chloride, sulfanilic acid, N-(1-Naphthyl) ethylenediamine dihydrochloride (NED), sodium nitrite, sodium hydroxide and potassium nitrate were procured from Sigma-Aldrich Chemicals Pvt. Ltd, SD Fine Chemicals Ltd, Qualigens Fine Chemicals, and Merck India Ltd respectively. All procured chemicals were of analytical grade (>99% purity).The reactions were conducted using conventional methods (Aggarwal et al. 2009)24 with slight modification as per the method previously published by the authors (Mondal et al. 2024).25 A mixture of 2,4,5-trichloroaniline, and various substituted benzaldehydes in methanol was stirred for 4–6 h at ambient temperature (Fig. 1 and Table 1). The progress of various reactions was examined by thin-layer chromatography (TLC) method. The resulting solid precipitate was filtered, washed with methanol, and further purified by recrystallization in ethanol.25 The reactions were carefully monitored using TLC chamber, employing ethyl acetate and hexane (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50) as the developing solvent system and visualized the reaction plate in iodine chamber. The synthesized Schiff base compounds were confirmed by measuring their melting points by an electro-thermal melting point apparatus (Lab india MPA 120). NMR spectra were recorded on a Bruker Avance 400 MHz instrument, with the compounds dissolved in CDCl3 or DMSO-d6. IR spectra were obtained using a Bruker Alpha FT-IR spectrophotometer in automatic mode.
50) as the developing solvent system and visualized the reaction plate in iodine chamber. The synthesized Schiff base compounds were confirmed by measuring their melting points by an electro-thermal melting point apparatus (Lab india MPA 120). NMR spectra were recorded on a Bruker Avance 400 MHz instrument, with the compounds dissolved in CDCl3 or DMSO-d6. IR spectra were obtained using a Bruker Alpha FT-IR spectrophotometer in automatic mode.
Soil for the in-vitro incubation experiments was collected from the Indian Agricultural Research Institute (IARI) farm, New Delhi (28.64° N and 77.12° E). All the collected soil samples were subjected to the physico-chemical parameters analysis in the laboratory and had the following characteristics such as 60.8% sand, 18.7% silt, 20.5% clay, 35.5% water holding capacity, 1.51 mg kg−1, bulk density, 0.5% organic carbon, 553.72 mg kg−1, available nitrogen, pH 7.9 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2.5 soil-to-water ratio), and an electrical conductivity (EC) of 0.35 dS m−1 at 25 °C. The experiments were carried out using a completely randomized design (CRD) with three replicates (n = 3). Fertilizer-N was added at a rate of 1000 mg kg−1 along with urea-N in each sample. Both synthesized Schiff base compounds (C1 to C21) and control samples were also included for the experiment. Nitrapyrin and DCD commercially available nitrification inhibitors were used as a reference inhibitor.26 Samples were placed in plastic beakers with air-dried and sieved (2 mm mesh sieve) soil. The desired amounts of test compounds (C1–C21) were added to the soil and mixed uniformly. Distilled water and urea (source of N) were then incorporated into the soil and mixed well, adjusting the moisture content to approximately 70%. The sample containers were incubated at a temperature of 28 ± 1 °C with a relative humidity of 98% in a controlled environment. Samples were collected at intervals on the 7th, 14th, 21st, and 28th days of incubation to measure ammonium, nitrate, and nitrite-N levels.
2.5 soil-to-water ratio), and an electrical conductivity (EC) of 0.35 dS m−1 at 25 °C. The experiments were carried out using a completely randomized design (CRD) with three replicates (n = 3). Fertilizer-N was added at a rate of 1000 mg kg−1 along with urea-N in each sample. Both synthesized Schiff base compounds (C1 to C21) and control samples were also included for the experiment. Nitrapyrin and DCD commercially available nitrification inhibitors were used as a reference inhibitor.26 Samples were placed in plastic beakers with air-dried and sieved (2 mm mesh sieve) soil. The desired amounts of test compounds (C1–C21) were added to the soil and mixed uniformly. Distilled water and urea (source of N) were then incorporated into the soil and mixed well, adjusting the moisture content to approximately 70%. The sample containers were incubated at a temperature of 28 ± 1 °C with a relative humidity of 98% in a controlled environment. Samples were collected at intervals on the 7th, 14th, 21st, and 28th days of incubation to measure ammonium, nitrate, and nitrite-N levels.
To extract nitrate, nitrite, and ammonium-N from soil samples, 50 mL 2 M aq. sodium sulphate (Na2So4) solution was added into the soil. The mixture was shaken for one hr and the extract was filtered using Whatman no. 1 filter paper (125 mm) and the filtrate was stored in capped PET bottles in a refrigerator at 5 °C until further analysis. The indophenol blue method, the sulfanilic acid method and the phenol disulphonic acid methods were applied for ammonium-N, nitrite-N and nitrate-N respectively. The absorbance was measured at 630 nm, 550 nm, and 410 nm, respectively, using a spectrophotometer (Agilent Technologies, Cary 60 UV-vis). To calculate the percentage of nitrification inhibition, determine the levels of NH4+-N, NO3−-N, and NO2−-N from standard curves and express them in mg kg−1. The nitrification rate computed using the following formula,27 NR = (NO3−-N + NO2−-N) × 100/(total-N) where, total-N = NH4+-N + NO3−-N + NO2−-N. The percentage of nitrification inhibition (NI) was estimated as, NI % = ((NR in control − NR in treated)/NR in control) × 100. In this formula where, nitrification rate (NR) in control refers to the nitrification rate observed in the control samples and NR in treatment refers to the nitrification rate observed in the treated samples.
2.1. Assessment of soil microbial activities
In this study, soil dehydrogenase activity (DHA), soil respiration, and microbial biomass carbon (MBC) were analyzed using standard methods.28 Soil samples were collected from highest treatment dose i.e. 10% after the nitrification study at 28th days. These samples were air-dried, sieved, and stored for microbial analysis. For DHA, 3 g of soil was incubated with 3% of triphenyl tetrazolium chloride (1 mL) at 37 °C for 24 h. The produced formazan was then extracted with methanol, and its absorbance was measure at 485 nm using a spectrophotometer.28 Soil respiration was measured by placing 50 g of soil in an incubation jar with 10 mL of 0.5 M sodium hydroxide (NaOH) to trap CO2 at 25 °C for 7 days. Following incubation, barium chloride (BaCl2) was added to the NaOH solution to precipitate carbonate, and the remaining NaOH was titrated with 0.1 M hydrochloric acid (HCl) using phenolphthalein as an indicator.29 The MBC was determined using the chloroform fumigation-extraction method. In this process, 10 g of soil was fumigated with chloroform for 24 hours, and then extracted with 0.5 M potassium sulfate. The organic carbon in the extracts was measured using a dichromate digestion method, and MBC was calculated as the variation between the treated and non-treated samples, using a kEC (extractable C to biomass C conversion factor) factor of 0.45.302.2. Quantitative structural activity relationship (QSAR)
Various physicochemical descriptors and topological parameters were computed using Marvin Sketch software. The calculated values are presented in (SI Table 2). These values are further used to develop suitable statistical models31 for predicting the potential activity of prepared Schiff base compounds for their nitrification inhibition.2.3. Statistical analysis
The experimental data were analyzed using statistical procedures outlined by Gomez and Gomez (1984).32 The analysis of variance and Duncan's Multiple Range Test (DMRT) were carried out using SPSS version 16.0 to compare treatment means at a 5% significance level. Paired t-test was performed to identify the treatment differences in nitrification inhibition activity. Nitrification inhibition activity was predicted using stepwise regression model as well as machine learning models such as random forest, support vector regression (SVR) and artificial neural network (ANN). Stepwise regression and machine learning models have been applied to check the significance of synthesized Schiff considering the bases at 1%, 5% and 10% doses as dependent variable and physiochemical factor as explanatory variable. Out of total 21 observations, 17 were randomly chosen as training dataset and remaining were used as testing dataset. The data was partitioned into 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 as training: testing ratio. Models were fitted based on the training dataset and further, these were validated using test dataset. Before performing ANN, all parameters were scaled as a part of the data analysis process.
20 as training: testing ratio. Models were fitted based on the training dataset and further, these were validated using test dataset. Before performing ANN, all parameters were scaled as a part of the data analysis process.
3. Results and discussion
3.1. Synthesis of 2,4,5-trichloro aniline based Schiff bases
Schiff bases derived from 2,4,5-trichloroaniline were synthesized by condensing aniline with various aldehydes, resulting in twenty-one novel azomethine compounds presented in Fig. 1. Reactions were completed within 4–6 hours at room temperature, yielding 45% to 80% with melting points ranging from 60 °C to 186 °C (Mondal et al. 2024).25 Purity was assessed with retardation factors (Rf) between 0.4 and 0.7, as shown in Table 3. Dhiraj et al. (2020) have also reported a similar type of results for m-nitroaniline-based Schiff base compounds.33 All synthesized compounds were characterized using physicochemical and spectral studies (Tables 2, 3 and SI Table 1). Spectral analysis confirmed the identity of the compounds, showing a characteristic azomethine group (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) band at 1550–1610 cm−1 and broad band at around 3300–3400 cm−1 due to O–H stretching vibration of phenolic group in IR spectra presented in Fig. 2, matched with the findings of Aggarwal et al. (2009) and Jyoti Kumari (2024).24,34,35 The 1H NMR and 13C NMR spectra as shown in Fig. 3 and 4 provided singlet peaks at δ 8.34–8.45 and 163–165, respectively, consistent with findings Ommenya et al. (2020) and Rani et al. (2024).36,37
N) band at 1550–1610 cm−1 and broad band at around 3300–3400 cm−1 due to O–H stretching vibration of phenolic group in IR spectra presented in Fig. 2, matched with the findings of Aggarwal et al. (2009) and Jyoti Kumari (2024).24,34,35 The 1H NMR and 13C NMR spectra as shown in Fig. 3 and 4 provided singlet peaks at δ 8.34–8.45 and 163–165, respectively, consistent with findings Ommenya et al. (2020) and Rani et al. (2024).36,37
| S. no. | Substituted aldehyde (R) | mpa (°C) | Rfb | % Yield |
|---|---|---|---|---|
a mp = observed melting point.b Solvent system = [ethyl acetate![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) hexane; 50 hexane; 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50]. 50]. |
||||
| 1 | 2,4-(OH)2C6H3 | 185–187 | 0.70 | 81 |
| 2 | 2-OCH3-4-OH C6H3 | 180–182 | 0.61 | 48 |
| 3 | 3,4,5-(OCH3)3C6H2 | 116–118 | 0.47 | 72 |
| 4 | 3,4-(OCH3)2 C6H3 | 108–110 | 0.43 | 75 |
| 5 | 4-OCH3C6H4 | 97–98 | 0.52 | 67 |
| 6 | 4-NO2C6H4 | 62–65 | 0.72 | 82 |
| 7 | 4-BrC6H4 | 68–70 | 0.6 | 75 |
| 8 | 3-CH3C6H4 | 99–101 | 0.53 | 73 |
| 9 | 4-ClC6H4 | 102–105 | 0.58 | 80 |
| 10 | 2-Pyridyl | 108 | 0.39 | 67 |
| 11 | 1-Napthyl | 135–137 | 0.45 | 77 |
| 12 | 4-CF3C6H4 | 77–79 | 0.48 | 87 |
| 13 | 4-SCH3C6H4 | 140–143 | 0.67 | 61 |
| 14 | 4-C(CH3)3C6H4 | 70–71 | 0.41 | 58 |
| 15 | 4-C2H5 | 65–66 | 0.52 | 47 |
| 16 | 4-OC2H5C6H4 | 81–83 | 0.69 | 72 |
| 17 | 4-OHC6H4 | 75–78 | 0.54 | 82 |
| 18 | 4-CH3C6H4 | 87–89 | 0.42 | 78 |
| 19 | 2-Naphthyl | 165–169 | 0.65 | 84 |
| 20 | 4-(CH3)2CH C6H4 | 59–61 | 0.71 | 46 |
| 21 | 2,4-(OCH3)2C6H3 | 140–142 | 0.52 | 79 |
| S. no. | R | IR (cm−1) (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) N) |
1H NMR (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, δ, ppm) N, δ, ppm) |
13C NMR (CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, δ, ppm) N, δ, ppm) |
|---|---|---|---|---|
| 1 | 2,4-(OH)2C6H3 | 1574 | 8.9 (singlet) | 165.45 |
| 2 | 2-OCH3-4-OH C6H3 | 1595 | 8.42 (singlet) | 164.2 |
| 3 | 3,4,5-(OCH3)3C6H2 | 1582 | 8.49 (singlet) | 164.48 |
| 4 | 3,4-(OCH3)2 C6H3 | 1597 | 8.48 (singlet) | 164.17 |
| 5 | 4-OCH3C6H4 | 1558 | 8.51 (singlet) | 163.96 |
| 6 | 4-NO2C6H4 | 1524 | 8.4 (singlet) | 162.7 |
| 7 | 4-BrC6H4 | 1558 | 8.60 (singlet) | 163.84 |
| 8 | 3-CH3C6H4 | 1541 | 8.32 (singlet) | 163.39 |
| 9 | 4-ClC6H4 | 1541 | 8.62 (singlet) | 163.68 |
| 10 | 2-Pyridyl | 1556 | 8.64 (singlet) | 155.25 |
| 11 | 1-Napthyl | 1541 | 8.41 (singlet) | 164.93 |
| 12 | 4-CF3C6H4 | 1542 | 8.716 (singlet) | 163.68 |
| 13 | 4-SCH3C6H4 | 1553 | 8.31 (singlet) | 162.4 |
| 14 | 4-C(CH3)3C6H4 | 1606 | 8.56 (singlet) | 164.58 |
| 15 | 4-C2H5 | 1558 | 8.42 (singlet) | 163.69 |
| 16 | 4-OC2H5C6H4 | 1601 | 8.46 (singlet) | 163.69 |
| 17 | 4-OHC6H4 | 1541 | 8.3 (singlet) | 163.43 |
| 18 | 4-CH3C6H4 | 1557 | 8.32 (singlet) | 163.04 |
| 19 | 2-Naphthyl | 1554 | 8.78 (singlet) | 163.61 |
| 20 | 4-(CH3)2CH C6H4 | 1606 | 8.31 (singlet) | 163.48 |
| 21 | 2,4-(OCH3)2C6H3 | 1607 | 8.47 (singlet) | 188.38 |
3.2. Ammonium-N content
All the synthesized Schiff bases (SI Table 3), across the time and concentration showed considerably better NH4+-N content (33–185 mg kg−1) in comparison to untreated samples (8–91 mg kg−1) throughout the incubation period. The respective NH4+-N content for the reference inhibitors i.e. nitrapyrin and DCD were 110–185 and 71–174 mg kg−1 respectively during the study period. In the experiment, compounds C3, C5, C15, C18, and C21 effectively maintained NH4+-N levels throughout the incubation period. Their performance at respective doses was comparable to the reference inhibitors, nitrapyrin and DCD (SI Table 3). At 10% concentration level, the reference inhibitor nitrapyrin maintained the highest NH4+-N (141–182 mg kg−1) during the incubation period.Among the synthesized potential Schiff bases, C6 was best at on 7th day with maximum NH4+-N (185 mg kg−1) but its effect was not persistent in conserving the NH4+-N. The NH4+-N content for C6 was 104 mg kg−1 as compared 120 mg kg−1 for the best performer on 28th day. Compound C18 was better which retained 167, 148 and 120 mg kg−1 NH4+-N on 14th, 21st and 28th day respectively. Its performance was superior to DCD on 7th, 14th and 21st day and at par with DCD on the 28th day as evident from CD values (SI Table 2). The next best in performance was C15, with values 120–176 mg kg−1 NH4+-N being comparable with DCD on 7th, 14th and 28th day with little moderate performance on 21st day. The synthesized Schiff base compound C3, with NH4+-N levels of 114–180 mg kg−1, remained in the top five through the 21st day. Similarly, compound C5, maintaining NH4+-N levels of 114–185 mg kg−1 during incubation, up to the 14th day. The remaining compounds showed moderate effectiveness, performing below the reference standard inhibitors (nitrapyrin and DCD) in maintaining NH4+-N. However, all tested compounds were effective in maintainingNH4+-N levels, ranging from 83–185 mg kg−1 at various stages of incubation, compared to urea alone, which had levels of 8–91 mg kg−1.
The conservation of NH4+-N with synthesized compounds was observed in a dose dependent nature (SI Table 3).20,21 At 5% level of concentration the NH4+-N content for nitrapyrin and DCD were 126–180 and 104–148 mg kg−1 respectively. Among the series, compound C3, was most active on all the days except on 14th day. The other potent molecules were C15, C21, C18 and C5 with respective NH4+-N, 99–163, 94–163, 95–158 and 93–160 mg kg−1. The remaining Schiff bases exhibited moderate activity in retaining NH4+-N. The tested compounds demonstrated consistent performance at the lower concentration level of 1%, comparable to their effectiveness at higher doses. Significantly higher NH4+-N was detected with the compounds C3, C15, C20 and C21 on all the days. The respective NH4+-N content was 67–138, 74–138, 74–134 and 79–145 mg kg−1 during the entire incubation period. Similar findings were reported by Li et al. (2024), the application of 3,4-dimethylpyrazole (DMP)-based nitrification inhibitors during a 35 day incubation study significantly increased soil NH4+-N concentrations in the first 25 days and decreased concentrations thereafter.38 The reference standards, nitrapyrin and DCD showed 110–161 and 71–131 mg kg−1 NH4+-N on all the sampling days.
3.3. Nitrite and nitrate-N content
The nitrite-N content was insignificant (<0.5 mg kg−1) across all the treatments on each sampling day.At a higher concentration level (10%), compounds C1, C3, C5, C6, C15, C18, and C20 demonstrated significant effectiveness in droppingNO3−-N content all over the incubation period. Among these, compound C1 was the most potent, lowering the NO3−-N levels to 6 mg kg−1, 11 mg kg−1, 21 mg kg−1, and 33 mg kg−1 on the 7th, 14th, 21st, and 28th days, respectively. Other effective compounds C3, C20, C15, C16, C6, and C21 showed nitrate-N levels ranging from 9–37 mg kg−1, 11–42 mg kg−1, 12–42 mg kg−1, 6–47 mg kg−1, 6–48 mg kg−1, and 10–50 mg kg−1 during the same period. The remaining compounds were less effective in reducing NO3−-N compared to the reference inhibitors nitrapyrin (2–29 mg kg−1) and DCD (7–50 mg kg−1). Compound C16 was found the least effective, with NO3−-N levels of 24 mg kg−1, 43 mg kg−1, 67 mg kg−1, and 91 mg kg−1 on the 7th, 14th, 21st, and 28th days, respectively.
At medium concentration (5%) tested Schiff bases showed similar effect as in higher dose. The standard inhibitor, nitrapyrin provided best results at the 5% level, reducing NO3−-N content to 8–52 mg kg−1. The significantly active compounds were C21, C20, C6, C18 and C3. Among these, most active compounds were C20 and C21 with NO3−-N content of 11, 23, 46, 66 mg kg−1 and 10, 24, 46, 67 mg kg−1 on the respective sampling days. They showed better activity than reference compound i.e. DCD (22–72 mg kg−1). Remaining test materials were low to quite active in lowering the NO3−-N content (SI Table 4).
Compounds C17 and C10 were observed the lowest effective in minimizing NO3−-N level, with values ranging from 56–116 mg kg−1 and mg kg−1, respectively, throughout the incubation period. At the lower treatment doses (1%), the tested compounds performed similarly to higher doses, showing nitrate-N levels between 23–137 mg kg−1 during the study period. Among them, compounds C21, C20, and C3 were particularly effective in reducing NO3−-N.
3.4. Nitrification inhibition
All the synthesized Schiff bases were efficient NI, presented in Table 4 and showed 4–95, 8–90, 18–85 and 15–77% reduction in nitrification process at different incubation days (7, 14, 21 and 28th days). Nitrapyrin and DCD as a reference inhibitor showed 76–97, 70–91, 65–88, 60–82% and 51–91, 43–84, 45–77, 36–69% nitrification inhibition on days 7, 14, 21, and 28, respectively. Among the series, C3, C20 and C21 were emerged as promising nitrification inhibitors and proved highly effective at low doses. The incubation study by Cui et al. (2021) found similar results: they observed that the use of NIs (nitrapyrin, 3,4-dimethylpyrazole phosphate, and dicyandiamide) with ammonium sulfate significantly increased the releases of NH4+-N contents and reduced NO3-N content by reducing soil nitrification process.40| Sl. no. | R | Dose (%) | Nitrification inhibition % | |||
|---|---|---|---|---|---|---|
| 7th day | 14th day | 21st day | 28th day | |||
| a Data are presented as mean ± standard error (n = 3), means with the same letter are not significantly different (P < 0.05) level of significance according to DMRT. | ||||||
| 1 | 2,4-(OH)2C6H3 | 1 | 69.40 ± 0.73ghi | 59.10 ± 1.04d | 51.10 ± 1.55c | 42.25 ± 0.55c |
| 5 | 86.31 ± 0.58cd | 77.23 ± 0.82cd | 68.56 ± 0.95b | 58.64 ± 0.77b | ||
| 10 | 96.10 ± 0.36b | 90.97 ± 0.75b | 84.34 ± 0.92b | 72.04 ± 1.33b | ||
| 2 | 2-OCH3-4-OH C6H3 | 1 | 65.17 ± 0.60j | 49.75 ± 0.92g | 33.76 ± 0.51hi | 20.11 ± 0.41ij |
| 5 | 85.34 ± 1.38cd | 65.36 ± 0.99j | 48.90 ± 0.78ij | 25.43 ± 0.60l | ||
| 10 | 91.40 ± 1.22cde | 86.20 ± 0.47c | 73.84 ± 0.52ef | 57.12 ± 0.71ef | ||
| 3 | 3,4,5-(OCH3)3C6H2 | 1 | 74.01 ± 0.85d | 62.81 ± 1.24c | 55.70 ± 0.96b | 45.75 ± 1.01b |
| 5 | 88.30 ± 0.81bc | 76.37 ± 0.73cd | 65.62 ± 1.25c | 58.97 ± 0.73b | ||
| 10 | 98.58 ± 0.15a | 93.74 ± 0.91a | 86.53 ± 0.90a | 74.21 ± 1.11b | ||
| 4 | 3,4-(OCH3)2 C6H3 | 1 | 59.10 ± 0.92kl | 41.92 ± 0.72h | 28.10 ± 0.67jk | 15.65 ± 0.23k |
| 5 | 74.12 ± 1.47ij | 58.73 ± 0.81kl | 43.00 ± 0.34k | 31.87 ± 0.61j | ||
| 10 | 88.63 ± 0.46fgh | 79.05 ± 0.83fg | 64.47 ± 0.41k | 48.49 ± 0.27h | ||
| 5 | 4-OCH3C6H4 | 1 | 68.81 ± 0.27hi | 54.43 ± 0.37e | 38.03 ± 0.57g | 31.30 ± 1.14g |
| 5 | 77.00 ± 1.01fhh | 66.39 ± 0.76hi | 55.80 ± 0.77fg | 34.59 ± 0.50efg | ||
| 10 | 96.58 ± 0.44ab | 91.81 ± 1.37b | 78.37 ± 0.70d | 62.03 ± 0.46d | ||
| 6 | 4-NO2C6H4 | 1 | 73.14 ± 0.89de | 50.07 ± 0.72g | 28.23 ± 0.57jk | 23.94 ± 0.81h |
| 5 | 87.67 ± 0.62bcd | 71.07 ± 1.85f | 55.93 ± 1.11fg | 34.59 ± 1.08i | ||
| 10 | 92.78 ± 0.97cd | 83.05 ± 0.83e | 73.39 ± 0.53ef | 57.47 ± 0.77ef | ||
| 7 | 4-BrC6H4 | 1 | 71.37 ± 0.61efg | 43.68 ± 0.19h | 23.93 ± 0.54l | 14.69 ± 0.63k |
| 5 | 87.45 ± 1.01bcd | 63.54 ± 0.40j | 47.85 ± 0.50j | 28.31 ± 1.23k | ||
| 10 | 89.99 ± 0.87efg | 72.17 ± 1.04i | 58.25 ± 0.83l | 43.95 ± 1.06i | ||
| 8 | 3-CH3C6H4 | 1 | 60.72 ± 1.21k | 48.10 ± 0.31g | 29.58 ± 0.77j | 20.73 ± 0.42i |
| 5 | 78.65 ± 1.03efg | 66.20 ± 0.86i | 47.75 ± 0.13j | 31.07 ± 0.76j | ||
| 10 | 76.54 ± 0.40l | 69.17 ± 1.15jk | 60.47 ± 0.96l | 50.82 ± 0.71h | ||
| 9 | 4-ClC6H4 | 1 | 53.60 ± 0.44n | 33.24 ± 0.20k | 14.71 ± 0.44n | 4.78 ± 0.66n |
| 5 | 73.55 ± 0.05ij | 53.71 ± 0.31n | 36.38 ± 0.23m | 10.91 ± 0.50° | ||
| 10 | 91.02 ± 0.97de | 79.79 ± 0.44f | 60.06 ± 0.28l | 37.95 ± 0.51kj | ||
| 10 | 2-Pyridyl | 1 | 46.11 ± 0.65° | 30.44 ± 0.64l | 18.51 ± 0.33m | 11.28 ± 0.23l |
| 5 | 67.53 ± 0.65k | 48.70 ± 0.14° | 31.94 ± 0.26n | 16.47 ± 0.47n | ||
| 10 | 87.86 ± 0.28ghi | 73.09 ± 0.74i | 55.01 ± 0.33m | 38.54 ± 0.41kj | ||
| 11 | 1-Naphthyl | 1 | 56.30 ± 0.37n | 39.05 ± 0.42i | 23.46 ± 0.31l | 18.23 ± 0.36j |
| 5 | 72.75 ± 0.13j | 56.02 ± 0.36mn | 38.53 ± 0.36l | 27.17 ± 1.26kl | ||
| 10 | 77.61 ± 0.45l | 71.02 ± 0.33ij | 60.45 ± 0.59l | 44.88 ± 0.32i | ||
| 12 | 4-CF3C6H4 | 1 | 68.15 ± 0.25i | 50.52 ± 0.46fg | 35.29 ± 0.22h | 28.81 ± 0.59fg |
| 5 | 79.95 ± 0.93e | 69.48 ± 1.05fg | 50.48 ± 0.53i | 37.79 ± 0.85g | ||
| 10 | 96.05 ± 0.65b | 77.13 ± 1.02gh | 69.70 ± 1.20hi | 58.23 ± 0.67ef | ||
| 13 | 4-SCH3C6H4 | 1 | 58.49 ± 0.73l | 37.12 ± 0.29ij | 27.04 ± 1.72k | 19.07 ± 1.04ij |
| 5 | 75.72 ± 0.16hi | 60.79 ± 0.37k | 41.91 ± 0.24k | 31.29 ± 0.88j | ||
| 10 | 91.62 ± 0.47cde | 79.92 ± 0.60f | 65.68 ± 0.12kj | 48.83 ± 0.17h | ||
| 14 | 4-C(CH3)3C6H4 | 1 | 70.57 ± 0.69fgh | 54.03 ± 0.74e | 39.87 ± 0.67fg | 30.54 ± 0.62efg |
| 5 | 87.86 ± 0.16bcd | 73.69 ± 1.01e | 59.42 ± 1.41e | 40.21 ± 0.56ef | ||
| 10 | 88.15 ± 0.38ghi | 82.52 ± 0.41e | 74.75 ± 0.93ef | 63.60 ± 1.05d | ||
| 15 | 4-C2H5C6H4 | 1 | 77.54 ± 0.91b | 62.16 ± 1.86c | 41.00 ± 0.22f | 33.91 ± 0.58d |
| 5 | 89.55 ± 0.38b | 78.64 ± 1.60c | 57.84 ± 0.91ef | 40.97 ± 0.80e | ||
| 10 | 86.05 ± 1.50ij | 76.56 ± 1.28h | 67.77 ± 0.48ij | 59.08 ± 0.43e | ||
| 16 | 4-OC2H5C6H4 | 1 | 41.81 ± 0.91p | 26.33 ± 0.44m | 15.81 ± 0.41n | 12.18 ± 0.48l |
| 5 | 60.25 ± 0.46l | 44.66 ± 0.07p | 31.51 ± 0.11n | 20.00 ± 0.18m | ||
| 10 | 79.88 ± 0.30k | 67.04 ± 0.10k | 52.27 ± 0.41n | 37.00 ± 0.12k | ||
| 17 | 4-OHC6H4 | 1 | 70.54 ± 0.35fgh | 52.86 ± 0.76ef | 37.99 ± 0.48g | 33.21 ± 1.24efg |
| 5 | 85.16 ± 0.96d | 70.54 ± 0.72fg | 55.03 ± 0.23gh | 43.51 ± 0.89d | ||
| 10 | 86.96 ± 1.01hij | 79.49 ± 0.58fg | 72.81 ± 0.36fg | 59.46 ± 0.35e | ||
| 18 | 4-CH3C6H4 | 1 | 74.36 ± 1.11d | 58.97 ± 0.42d | 40.18 ± 0.31f | 25.06 ± 0.89g |
| 5 | 87.11 ± 2.16bcd | 75.54 ± 0.47de | 58.68 ± 0.33e | 35.09 ± 0.04hi | ||
| 10 | 90.59 ± 0.63def | 83.33 ± 0.66de | 74.62 ± 1.32ef | 56.49 ± 1.12f | ||
| 19 | 2-Naphthyl | 1 | 68.07 ± 0.59i | 48.23 ± 0.43g | 32.45 ± 0.79i | 28.32 ± 0.52g |
| 5 | 76.01 ± 1.06ghi | 68.59 ± 0.40fghi | 53.36 ± 0.58h | 39.10 ± 1.01efg | ||
| 10 | 85.09 ± 0.37j | 78.62 ± 0.77fgh | 71.04 ± 0.37gh | 53.06 ± 0.38g | ||
| 20 | 4-(CH3)2CH C6H4 | 1 | 77.05 ± 0.70b | 64.21 ± 0.60c | 48.65 ± 0.17d | 31.67 ± 0.92de |
| 5 | 79.59 ± 0.80ef | 68.06 ± 0.92ghi | 58.46 ± 1.26e | 44.76 ± 0.92d | ||
| 10 | 87.54 ± 0.55hi | 82.44 ± 0.28e | 74.14 ± 1.24ef | 67.23 ± 0.82c | ||
| 21 | 2,4-(OCH3)2C6H3 | 1 | 67.72 ± 0.40i | 54.06 ± 1.07e | 43.78 ± 0.85e | 32.62 ± 1.02de |
| 5 | 78.97 ± 0.60ef | 68.85 ± 0.71fgh | 55.86 ± 0.74fg | 48.55 ± 0.62c | ||
| 10 | 90.33 ± 0.57ef | 83.37 ± 0.65de | 68.40 ± 0.63i | 49.66 ± 0.59h | ||
| Std.1 | Nitrapyrin | 1 | 89.22 ± 1.41a | 74.71 ± 1.41a | 61.05 ± 1.02a | 48.01 ± 1.42a |
| 5 | 94.27 ± 0.83a | 84.86 ± 0.65a | 72.27 ± 0.90a | 62.55 ± 0.56a | ||
| 10 | 97.78 ± 0.21ab | 93.12 ± 0.60ab | 87.27 ± 0.88a | 77.28 ± 1.19a | ||
| Std.2 | DCD | 1 | 71.78 ± 0.45ef | 58.64 ± 1.30d | 43.12 ± 0.73e | 31.08 ± 0.62ef |
| 5 | 86.82 ± 1.19bcd | 75.52 ± 0.97de | 61.87 ± 0.77d | 37.09 ± 0.29gh | ||
| 10 | 92.82 ± 1.06cd | 86.13 ± 0.67c | 75.50 ± 0.80e | 61.89 ± 1.08d | ||
| LSD (5%) | — | 3.1 | 4.3 | 2.7 | 1.4 | |
At 10% concentration level, all the compounds were highly effective and showed 47–77% NI as compared to nitrapyrin (82%) and DCD (69%). Among the series, compound C1 was the most effective, exhibiting NI percentages of 94%, 90%, 85%, and 77% on days 7, 14, 21, and 28, in that order. This was followed by compound C3, which showed NI percentages ranging from 91% to 74% over the same period. Though, this compound was not able to maintain the same potency at lower doses. Other active novel synthesized compounds were C18 with 70–93% NI; C15 with 71–94% NI and C20 with 72–87% NI during incubation period.
At medium concentration level, C21 compounds showed highest NI i.e. 89, 78, 67 and 56% at different incubation days respectively (Table 4). The performance was quite comparable with nitrapyrin but more effective than DCD up to 21 days. The next active compounds were C20 and C3 with 56–86% and 55–81% NI respectively during the study period. Rest of the compounds exhibited low to moderate nitrification inhibition potential in comparison to nitrapyrin and DCD.
Based on overall performance at 1% dose, C21 performed the best with NI value 72, 60, 53 and 43% on the same days respectively, followed by C20, C3, C15 and C18 showed moderate to high NI activity (Table 4). All the synthesized Schiff base compounds showed an increase in NI with the increase in dose, though the rate of increase varied with the different compounds, which also matched with the findings of Sidhu and Wille's research on 1,4-disubstituted 1,2,3-triazoles as a novel class of nitrification inhibitors (NIs).41
4. Assessment of soil health indicator parameters
The impacts of different treatments on key soil health indicators, including soil respiration, dehydrogenase activity, and microbial biomass carbon (MBC) presented in Table 5, revealed significant variation between the treatments according to DMRT results. The C3 treatment demonstrated the highest values across all indicators, with soil respiration at 34.0 (µg CO2-C per g), dehydrogenase activity at 46.4 (µg TPF per g soil per 24 h) and MBC at 556.3 (µg gm−1), indicating superior microbial activity and soil health, significantly differing from all other treatments. Li et al. (2024) also observed the similar kind of findings, they reported 3,4-dimethylpyrazole phosphate, a novel nitrification inhibitor has potential to enhance the soil microbial activities which helps to promote the plant root and shoot growth.38 The C20 and C21 treatments showed moderate values, with soil respiration at 32.8 and 32.6 (µg CO2-C per g), dehydrogenase activity at 44.2 and 43.0 (µg TPF per g soil per 24 h), and MBC at 542.5 and 537.2 (µg gm−1) respectively. These values were significantly higher than those of the standard inhibitors DCD and nitrapyrin. The DCD treatment exhibited soil respiration at 27.6 (µg CO2-C per g), dehydrogenase activity at 40.9 (µg TPF per g soil per 24 h), and MBC at 498.2 (µg gm−1), while the nitrapyrin treatment had soil respiration at 27.3 (µg CO2-C per g), dehydrogenase activity at 39.0 (µg TPF per g soil per 24 h), and MBC at 470.6 (µg gm−1), indicating suppressed microbial functions and significant differences from other treatments.42,43 The urea alone control showed intermediate values, with soil respiration at 30.9 (µg CO2-C per g), dehydrogenase activity at 43.5 (µg TPF per g soil per 24 h), and MBC at 530.6 (µg gm−1), which were higher than the standard inhibitors but significantly lower than the C3 treatment. The initial status had soil respiration at 29.1 (µg CO2-C per g), dehydrogenase activity at 44.8 (µg TPF per g soil per 24 h), and MBC at 535.9 (µg gm−1), showing no significant differences from C20 and C21 but significantly differing from DCD, nitrapyrin, and urea alone. Overall, the 2,4,5-trichloro aniline-based nitrification inhibitors, particularly C3, significantly improved soil health indicators compared to the standard inhibitors and control, with clear statistical differences observed among the treatments.| Test compound | Soil respiration (µg CO2-C per g) | Dehydrogenases (µg TPF per g soil per 24 h) | MBC (µg gm−1) |
|---|---|---|---|
| a Data are means ± S.Em with three replications and the letters a, b, c, d, and e indicate Duncan grouping of treatment differences. Means with the same letter are not significantly different. | |||
| C3 | 34.0 ± 0.83a | 46.4 ± 0.79a | 556.3 ± 8.57a |
| C20 | 32.8 ± 1.61b | 44.2 ± 0.81b | 542.5 ± 6.48b |
| C21 | 32.2 ± 1.10b | 43.0 ± 0.31b | 537.2 ± 8.84b |
| DCD (Std) | 27.6 ± 0.67c | 40.9 ± 1.17c | 498.2 ± 7.92d |
| Nitrapyrin (Std) | 27.3 ± 0.79c | 39.0 ± 0.90d | 470.6 ± 6.10e |
| Urea alone (control) | 30.9 ± 0.91b | 43.5 ± 1.75b | 530.6 ± 7.21c |
| Initial status | 29.1 ± 1.79b | 44.8 ± 1.28b | 535.9 ± 9.08b |
5. Structural activity relationships
An increase in the number of methoxy and hydroxy group in the phenyl ring of 2,4,5 trichloro aniline, increases the microbial inhibitory activity significantly (Fig. 5).25 The compound containing three methoxy groups (C3; 46, 59 & 74%) in the ring showed the highest nitrification inhibition activity as compare to the compounds having one (C5; 29, 39 & 62%) and two methoxy groups (C4; 16, 32 & 48%) in the phenyl ring at 1, 5 and 10% treatment doses on 28 days.20,24 Introduction of ethoxy group (C17, 31, 44 & 59%) and hydroxy group (C18; 29, 39 & 62%) in place of methoxy group (C5) showed similar affect at all treatments level. However substitution of two hydroxy group (C1; 42, 59 & 72%) increase the inhibition activity tremendously reported by Sonnekar et al. 2013.44,45 The nitrification inhibition activity also influence on the number of alkyl chain in the aromatic ring, study of the effect of carbon chain was presented in Fig. 6. The result revealed that introduction of more number of carbon in the alkyl group enhances the activity, similar findings was observed by Datta et al. (2001).21 The compound having double methyl group (C20; 32, 45 & 67%) and three methyl group (C14; 31, 40 & 64%)providing good result as compare to single (C18; 25, 35 & 56%) containing Schiff bases. But substitution of hetero cyclic group retarded the inhibitory activity extensively (Fig. 6). Other side electron withdrawing group reduces the nitrification inhibition percentage except compound containing nitro group (C6; 24, 35 & 57%) in their phenyl ring (Fig. 7). | ||
| Fig. 5 Effect of number and position of hydroxy and alkoxy groups on nitrification inhibition (bar indicates standard error of mean). | ||
 | ||
| Fig. 6 Effect of carbon chain of alkyl groups and heterocyclic substitution on nitrification inhibition (bar indicates standard error of mean). | ||
 | ||
| Fig. 7 Influence of substituted electron withdrawing groups of 2,4,5-trichloro aniline based Schiff base on nitrification inhibition % (bar indicates standard error of mean). | ||
5.1. Statistical analysis within the treatment concentration
At different treatments concentrations combinations paired t-test has been performed to check whether there is any significance difference in mean for any treatment combination mentioned in the SI Table 5. From the table, it can be concluded that all the treatments differ significantly in terms of mean response from each other. The treatment mean difference of 1% is less than at 5% and 10% respectively and the treatment mean difference at 5% is also less than 10%.5.2. QSAR model and validation
Stepwise regression model at 1% treatment dose
| Nitrification inhibition (NI) % = −29.87 + 1.82VV − 10.89Pol − 0.04Sz |
Stepwise regression model at 5% treatment dose
Nitrification inhibition (NI) % = 48.37 + 0.22 MW + 4.43VV − 30.32Pol + 0.60Polar.2D.SA + 23.03![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P − 57.39BI − 0.09WI + 8.29Wp − 7.71PI P − 57.39BI − 0.09WI + 8.29Wp − 7.71PI |
Stepwise regression model at 10% treatment dose
Nitrification inhibition (NI) % = −401.70 − 56.96 MR + 13.07VV + 33.86Pol + 56.15HbA + 94.31![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P + 0.89Sz − 2.13WI − 11.94Wp + 88.94RI + 7.73PI P + 0.89Sz − 2.13WI − 11.94Wp + 88.94RI + 7.73PI |
At 1% treatment dose, multiple R-squared and adjusted R-squared values of the fitted stepwise regression model are found to be 0.438 and 0.310 respectively. At 5% treatment dose, multiple R-squared and adjusted R-squared values of the fitted stepwise regression model are found to be 0.720 and 0.360 respectively. At 10% treatment dose, multiple R-squared and adjusted R-squared values of the fitted stepwise regression model are found to be 0.898 and 0.728 respectively. Hence, it may be concluded that the model with 10% treatment dose is the best fitted stepwise regression model.
The variable importance plots generated from the random forest models are presented in the Fig. 8. For nitrification inhibition activity at 1% treatment dose, important feature variables identified from variable importance plot based on % increase in node impurity are VV, MR, BI, and Pol. Similarly, for nitrification inhibition activity at 5% treatment dose important feature variables identified from variable importance plot are MR, VV, Pol and MW, whereas for 10% treatment dose important feature variables Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, VV, Sz and MW respectively.
P, VV, Sz and MW respectively.
6. Conclusions
A novel series of 2,4,5-trichloro aniline-based Schiff bases were successfully synthesized through a single-step green synthesis method and assessed for their nitrification inhibitory activity against soil nitrifying bacteria. The study revealed that these compounds not only demonstrated moderate to strong nitrification inhibition but also positively influenced soil microbial activities. Among the synthesized compounds, C3 showed the strongest inhibition of nitrification, being only 3–5% less effective than nitrapyrin and surpassing DCD by 19–60%. C20 was 27–33% less effective than nitrapyrin but 3–22% more effective than DCD. C21 was 23–36% less effective than nitrapyrin and 6–30% more effective than DCD at lower doses, but at a 10% concentration, it was 21% less effective than DCD. In terms of soil biological activity, C-3 improved soil respiration, dehydrogenase activity, and microbial biomass carbon by 13.3%, 7.0%, and 4.9% over urea, whereas C20 (2–10%) and C21 (1–6.7%) showed moderate improvements; conversely, nitrapyrin and DCD decreased microbial biomass by 6–11%, DHA by 7–9%, and soil respiration by up to 10%, indicating negative effects on soil microbial functioning. The eco-friendly nature of these compounds, along with their efficiency in conserving ammonium-N levels and boosting microbial activity, highlights their potential in promoting sustainable agricultural practices. By reducing nitrogen losses through nitrification, these inhibitors can help to improve nitrogen use efficiency in soils, leading to enhanced crop productivity and reduced environmental pollution. Future research should focus on molecular modifications of these compounds that can lead to the development of even more potent nitrification inhibitors, offering a valuable tool for sustainable agriculture and mitigating the environmental impacts of excessive fertilizer use. This study provides a strong foundation for the future design of eco-friendly nitrification inhibitors that can support long-term soil health and agricultural productivity.Author contributions
Tilak Mondal, Rajesh Kumar – conceptualization, guidance, methodology, graphical design, writing – original draft, review and editing, and visualization. Santanu Mukherjee, Nilimesh Mridha, Namrata Laskar, Avijit Das, Pradip Basak, Atul Singha, Kanchan Sinha and Dinesh Babu Shakyawar – facilitation, data interpretation, and writing – review and editing.Conflicts of interest
The authors have no conflicts of interest in connection to this article.Data availability
All data that sustenance the observations of the current study are incorporated in the article.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra09828a.
Acknowledgements
The authors are thankful to the Indian Council of Agricultural Research (ICAR), India for financial assistance. The author expresses gratitude to the Indian Council of Agricultural Research, New Delhi, for its financial assistance. Special thanks are also extended to ICAR-National Institute of Natural Fibre Engineering and Technology, Kolkata and the Division of Agricultural Chemicals, Indian Agricultural Research Institute, New Delhi, for providing instrumental facilities and supporting research expenses.References
- NAAS, Policy Options for Efficient Nitrogen Use, National Academy of Agricultural Sciences, New Delhi, 2005, p. 12, https://naas.org.in/PolicyPapers/policy33.pdf Search PubMed.
- P. Newbould, The use of nitrogen fertilizer in agriculture. Where do we go practically and ecologically?, Plant Soil, 1989, 115, 297–311 Search PubMed.
- D. Chen, H. Suter, A. Islam, R. Edis and J. R. Freney, Prospects of improving efficiency of fertilizer nitrogen in Australian agriculture; a review of enhanced efficiency fertilizers, Aust. J. Soil Res., 2008, 46, 289–301 Search PubMed.
- G. Chen, X. Cao and C. Song, Adverse effect of ammonia on nitrification process: the case of Chines shallow freshwater lake, Water, Air, Soil Pollut., 2010, 210, 297–306 CrossRef CAS.
- R. Prasad, Fertilizer urea, food security, health, and the environment, Curr. Sci., 1998, 75, 677–683 Search PubMed.
- D. Cardoen, P. Joshi, L. Diels, P. M. Sarma and D. Pant, Agriculture biomass in India: part 2. post-harvest losses, cost and environmental impacts, Resour., Conserv. Recycl., 2015, 101, 143–153, DOI:10.1016/j.resconrec.2015.06.002.
- M. M. Deo, D. De, I. Mani and M. A. Iqbal, Development of mechanical urea briquette applicator for SRI, Indian J. Agric. Sci., 2021, 91(2), 208–212, DOI:10.56093/ijas.v91i2.111578.
- E. Cataldo, L. Salvi, F. Paoli and M. Fucile, et al., Application of zeolites in agriculture and other potential uses: a review, Agronomy, 2021, 11(8), 1547, DOI:10.3390/agronomy11081547.
- W. R. Raun and G. V. Johnson, Improving nitrogen use efficiency for cereal production, Agron. J., 1999, 91, 357–363 Search PubMed.
- G. V. Subbarao, O. Ito, K. L. Sahrawat, W. L. Berry, K. Nakahara, T. Ishikawa, T. Watanabe, K. Suenaga, M. Rondon and I. M. Rao, Crit. Rev. Plant Sci., 2006, 25, 305 CrossRef.
- R. Kumar, C. Devakumar, V. Sharma, G. Kakkar, D. Kumar and P. Panneerselvam, Influence of Physicochemical Parameters of Neem (Azadirachta indica A Juss) Oils on Nitrification Inhibition in soil, J. Agric. Food Chem., 2007, 55, 1389–1393 Search PubMed.
- R. Kumar, C. Devakumar, D. Kumar, P. Panneerselvam, G. Kakkar and T. Arivalagan, Influence of edaphic factors on the mineralization of neem oil coated urea in four Indian soils, J. Agric. Food Chem., 2008, 56(21), 10183–10191 Search PubMed.
- R. Prasad and J. F. Power, Nitrification inhibitors for agriculture, health, and the environment, Adv. Agron., 1995, 54, 233–281 Search PubMed.
- S. C. Yildirim, J. G. Nathanael and K. Frindte, et al., 4-methyl-1-(prop-2-yn-1-yl)-1h-1,2,3-triazole (MPT): a novel, highly efficient nitrification inhibitor for agricultural applications, ACS Agric. Sci. Technol., 2024, 4(2), 255–265, DOI:10.1021/acsagscitech.3c00506.
- N. Aggarwal, R. Kumar, C. Shrivastava, P. Dureja and J. M. Khurana, Synthesis of nalidixic acid-based hydrazones as novel pesticides, J. Agric. Food Chem., 2010, 58, 3056–3061 Search PubMed.
- E. D. Souza, R. P. Soratto and A. M. Fernandes, Performance of conventional and enhanced-efficiency nitrogen fertilizers on potato tuber mineral composition and marketability, J. Sci. Food Agric., 2021, 102(8), 3078–3087, DOI:10.1002/jsfa.11648.
- D. N. Dhar and C. L. Taploo, Schiff bases and their application, J. Sci. Ind. Res., 1982, 41, 501–506 Search PubMed.
- A. M. Suzan, H. T. Wamidh, M. S. Mustafa, M. S. Mubarak, A. Murad and A. Damen, Synthesis, characterization, and antimicrobial activity of Schiff bases derived from benzaldehydes and 3,3′-diaminodipropylamine, Arabian J. Chem., 2015, 8(6), 850–857 CrossRef.
- R. Fontana, P. C. R. Marconi, A. Caputo and V. B. Gavalyan, Novel Chitosan-Based Schiff Base Compounds: Chemical Characterization and Antimicrobial Activity, Molecules, 2022, 27, 2740, DOI:10.3390/molecules27092740.
- N. Thombare, N. Aggarwal, R. Kumar and M. Gopal, Synthesis of 2-furyl-4-arylidene-5(4H)-oxazolones as new potent antibacterial agents against phyto-pathogenic and nitrifying bacteria, J. Environ. Sci. Health, Part B, 2012, 47(4), 326–335 Search PubMed.
- A. Datta, S. Walia and B. S. Parmar, Some furfural derivatives as nitrification, J. Agric. Food Chem., 2001, 49, 4726–4731 Search PubMed.
- C. Wang, L. Fan, Z. Pan, S. Fan, L. Shi, X. Li, J. Zhao, L. Wu, G. Yang and C. Xu, Synthesis of Novel Indole Schiff Base Compounds and Their Antifungal Activities, Molecules, 2022, 27, 6858, DOI:10.3390/molecules27206858.
- A. A. Jarrahpour, M. Motamedifar, K. Pakshir, N. Hadi and M. Zarei, Synthesis of Novel Azo Schiff Bases and Their Antibacterial and Antifungal Activities, Molecules, 2004, 9, 815–824 CrossRef CAS PubMed.
- N. Aggarwal, R. Kumar, P. Dureja and D. S. Rawat, Schiff bases as potential fungicides and nitrification inhibitors, J. Agric. Food Chem., 2009, 57, 8520–8525 CrossRef CAS.
- T. Mondal, R. Kumar, J. Bettanayaka, R. Gogoi, P. Koti, M. Ray, R. K. Kole and S. Mukherjee, Biodegradable Schiff bases: a novel approach for the management of pathogenic fungi (Sclerotium rolfsii and Rhizoctonia bataticola)and stored grain insect (Callosobruchus maculatus) in green gram (Vigna radiata), Environ. Sci. Pollut. Res., 2024, 31, 52540–52561, DOI:10.1007/s11356-024-34713-9.
- K. L. Sahrawat, D. R. Keeney and S. S. Adams, Ability of nitrapyrin, dicyandiamide and acetylene to retard nitrification in a mineral and an organic soil, Plant Soil, 1987, 101, 179–182 CrossRef CAS.
- K. L. Sahrawat, On the criteria for comparing the ability of compounds for retardation of nitrification in soil, Plant Soil, 1980, 55, 487–490 CrossRef CAS.
- L. E. Casida Jr., D. A. Klein and T. Santoro, Soil dehydrogenase activity, Soil Sci., 1964, 98(6), 371–376, DOI:10.1097/00010694-196412000-00004.
- J. P. E. Anderson, Soil respiration, in Methods of Soil Analysis. Part 2: Chemical and Microbiological Properties, ed. A. L. Page, R. H. Miller and D. R. Keeney, 1982, vol. 2, pp. 831–871 Search PubMed.
- E. D. Vance, P. C. Brookes and D. S. Jenkinson, An extraction method for measuring soil microbial biomass carbon, Soil Biol. Biochem., 1987, 19(6), 703–707 Search PubMed.
- C. Nantasenamat, C. Isarankura-Na-Ayudhya, T. Naenna and V. Prachayasittikul, A practical overview of quantitative structure-activity relationship, Excli J., 2009, 8, 74–88 Search PubMed.
- K. A. Gomez and A. A. Gomez, Statistical Procedures for Agricultural Research, John Wiley and Sons Inc., New York,2nd edn, 1984 Search PubMed.
- N. Dhiraj, N. Rutuja, S. Kalpesh and K. Sunil, Synthesis of Schiff’s Bases with simple synthetic approach, Asian J. Pharm. Res. Dev., 2020, 8(5), 72–74 CAS , https://ajprd.com/index.php/journal/article/view/837/683.
- J. Kumari, Synthesis and characterization of novel Schiff base ligands, Int. J. Chem. Stud., 2024, 12(6), 56–62 CAS.
- A. A. Abbas, Z. N. Mageed, O. A. Nief and H. H. Mohammed, Synthesis and characterization of Schiff bases derived from 3-(4-methoxyphenyl) acrylic acid and 3-(benzo[d][1,3]dioxol-5-yl) acrylic acid, J. Med. Chem. Sci., 2023, 6(2), 205–219, DOI:10.26655/JMCHEMSCI.2023.2.4.
- F. K. Ommenya, E. A. Nyawade, D. M. Andala and J. K. Synthesis, Characterization and Antibacterial Activity of Schiff Base, 4-Chloro-2-{(E)-[(4-Fluorophenyl) imino]methyl}phenol Metal (II) Complexes, J. Chem., 2020, 1–8, DOI:10.1155/2020/1745236.
- R. Mamatha Rani and P. Kavitha, Synthesis, characterization and antimicrobial activity of Schiff base ligand and metal complexes, Indian J. Chem., 2024, 63(3), 281–285, DOI:10.56042/ijc.v63i3.6539.
- D. Li, Q. Wang, T. Yang, R. Liu and D. Chen, Novel nitrification inhibitor produced by immobilizing 3,4-dimethylpyrazole using biogenic substances instead of chemicals have potential for sustainable agriculture, J. Environ. Chem. Eng., 2024, 12, 114011 Search PubMed.
- H. Gao, J. Li and F. Xu, Synthesis of a Novel Polymer Nitrification Inhibitor with Acrylic Acid and 3,4-Dimethylpyrazole, J. Agric. Food Chem., 2021, 69(11), 3307–3311 Search PubMed.
- L. Cui, D. Li, Z. Wu, Y. Xue, F. Xiao, L. Zhang, Y. Song, Y. Li, Y. Zheng and J. Zhang, Effects of Nitrification Inhibitors on Soil Nitrification and Ammonia Volatilization in Three Soils with Different pH, Agronomy, 2021, 11(8), 1674, DOI:10.3390/agronomy11081674.
- P. K. Sidhu and U. Wille, Exploring the Fate of Substituted 1,2,3-Triazoles: Unraveling the Soil Degradation Pathways of a Novel Class of Nitrification Inhibitors, ACS Agric. Sci. Technol., 2025, 5, 2429–2438 Search PubMed.
- R. Schmidt, X. B. Wang, P. Garbeva and É. Yergeau, The nitrification inhibitor nitrapyrin has non-target effects on the soil microbial community structure, composition, and functions, Appl. Soil Ecol., 2022, 171, 104350, DOI:10.1016/j.apsoil.2021.104350.
- G. Giannopoulos, L. Elsgaard, V. A. Tzanakakis, R. B. Franklin, B. L. Brown, G. Zanakis, N. Monokrousos, I. Anastopoulos, M. Awad, I. Ipsilantis, N. Barbayiannis and A. N. Polidoros, Impact of nitrapyrin on urea-based fertilizers in a Mediterranean calcareous soil: Nitrogen and microbial dynamics, Eur. J. Soil Sci., 2024, 75(4), e13553, DOI:10.1111/ejss.13553.
- V. N. Mkpenie, I. V. Mkpenie and E. Essien, Biological activities of (E)-N-(CH3-substituted-phenyl)-1-phenylmethanimine: Evaluation of ortho-, meta- and para- substitution effects, Der Pharma Chem., 2015, 7(6), 330–334 CAS , https://www.researchgate.net/publication/279449795.
- V. S. Sonnekar, W. N. Jadhav, S. A. Dake and R. P. Pawar, Synthesis and Antimicrobial and antifungal activities of novel Bis-imine Derivatives, Res. J. Pharm., Biol. Chem. Sci., 2013, 4(2), 1411–1418 CAS , https://www.researchgate.net/publication/286349318.
| This journal is © The Royal Society of Chemistry 2026 |