Open Access Article
Open Access ArticleThienopyrimidine scaffolds as promising antimicrobial agents: synthesis, biological evaluation, DFT analysis and molecular docking
Mostafa Sayed†
a,
Menna A. Naguib†a,
Ahmed A. K. Mohammedb,
Reda Hassaniena,
Mahmoud S. Tolba*a and
Mostafa Ahmed *a
*a
aChemistry Department, Faculty of Science, New Valley University, El-Kharja 72511, Egypt. E-mail: Mahmoud.Tolba@sci.nvu.edu.eg; drmostafa@scinv.au.edu.eg
bChemistry Department, Faculty of Science, Assiut University, Assiut 71516, Egypt
First published on 26th January 2026
Abstract
Thienopyrimidines represent an important class of nitrogen- and sulfur-containing heterocycles with broad pharmacological relevance. In this work, a new series of thienopyrimidine derivatives was efficiently synthesized through a multistep route involving cyclization, Mannich annulation, Schiff-base condensation, dipyrimidinone ring formation, and acylation, affording structurally diverse fused frameworks. The synthesized compounds were fully characterized using IR, NMR, and mass spectrometry. Their antimicrobial activity was evaluated against representative Gram-positive, Gram-negative strains and fungal strains revealing clear structure–activity relationships. Notably, the Mannich derivative 11 exhibited the highest potency toward Bacillus subtilis, while the fused dipyrimidinone compound 13 demonstrated broadened activity, including inhibition of Escherichia coli. Density functional theory (DFT) calculations provided insight into the electronic structure, intramolecular interactions, HOMO–LUMO distributions, electrostatic potential surfaces, and noncovalent interaction patterns responsible for the observed reactivity and binding behavior. Molecular docking studies against B. subtilis and E. coli DNA gyrase B revealed strong and diverse protein–ligand interactions, consistent with the biological findings, and highlighted compound 13 as the most promising candidate due to its favorable Vina score and extensive hydrogen-bonding and hydrophobic contacts. Collectively, the combined synthetic, biological, and computational analyses underscore the potential of thienopyrimidine scaffolds as valuable agents for developing new antimicrobial agents.
1 Introduction
Heterocyclic compounds are among the most versatile and biologically significant classes of organic molecules, forming the structural core of numerous therapeutic agents and natural products.1–4 Among heterocyclic scaffolds, pyrimidine derivatives hold a particularly prominent position owing to their presence in key biomolecules5—including nucleic acids,6 coenzymes, and vitamins—and their wide range of pharmacological activities,7 such as antimicrobial,8,9 anticancer,10 antiviral,11 and anti-inflammatory effects.12 This remarkable biological relevance has driven extensive synthetic research aimed at developing new pyrimidine-based derivatives with enhanced potency, improved selectivity, and more favorable pharmacokinetic properties.13,14 Notably, the fusion of the pyrimidine ring with other heterocyclic systems frequently results in compounds with superior biological activity and improved physicochemical stability.15,16 Among these fused heterocycles, thienopyrimidines,17 comprising a thiophene ring fused to a pyrimidine nucleus—have gained significant attention as privileged scaffolds in medicinal chemistry.11,18,19 The incorporation of sulfur within the heteroaromatic core modulates lipophilicity, electronic distribution, and molecular recognition, ultimately enhancing binding affinity toward diverse biological targets.20,21 Consequently, thienopyrimidine derivatives exhibit a broad spectrum of biological activities, including kinase inhibition,22 antitumor,23 antiviral, anti-inflammatory, and antimicrobial effects.24–30 Several members of this class have emerged as potent inhibitors of tyrosine kinases, phosphoinositide-3-kinase (PI3K), and vascular endothelial growth factor receptor (VEGFR), underscoring their growing therapeutic significance.31,32The rise of antimicrobial resistance (AMR) represents one of the most urgent challenges in global health, contributing to increased morbidity, mortality, and healthcare expenditures.33 The rapid emergence of multidrug-resistant bacterial and fungal pathogens underscores the critical need for new antimicrobial agents operating through novel mechanisms of action.34,35 Heterocyclic frameworks bearing nitrogen and sulfur atoms are particularly valuable in this pursuit, as their ability to engage in hydrogen bonding, π–π stacking, and hydrophobic interactions often enhances binding within enzyme active sites.36,37 Within this context, thienopyrimidines have emerged as a promising platform for the development of next-generation antimicrobial agents.38 Their modular structure allows for strategic substitutions at various ring positions, enabling fine-tuning of lipophilicity, solubility, electronic properties, and receptor affinity, thereby facilitating the optimization of antimicrobial activity.39,40
In recent years, numerous thienopyrimidine derivatives have demonstrated notable antimicrobial activity against Staphylococcus aureus, Escherichia coli, Pseudomonas aeruginosa, and Candida albicans, with some exhibiting minimum inhibitory concentrations (MICs) comparable to those of standard antibiotics.41,42 Despite these promising results, the detailed structure–activity relationship (SAR) and the molecular basis underlying their antimicrobial effects remain only partially understood. In this regard, computational molecular docking has become an indispensable tool for elucidating ligand–target interactions, offering insights into preferred binding conformations, key amino acid residues involved, and binding energy profiles that often correlate well with experimental findings.43–45
In this study, we present the synthesis, antimicrobial evaluation, DFT calculations, and molecular docking analysis of a new series of thienopyrimidine derivatives. The target compounds were prepared through a multistep synthetic protocol using readily accessible precursors, and their structures were fully characterized by IR, NMR, and mass spectrometry.46 Their antimicrobial activities were assessed against representative Gram-positive, Gram-negative and fungal strains using standard microbiological assays. In addition, molecular docking studies were performed to rationalize the observed biological activity and to elucidate potential binding interactions within the active sites of selected microbial enzymes. By integrating synthetic, biological, and computational methodologies, this work provides a comprehensive evaluation of the antimicrobial potential of thienopyrimidine scaffolds and highlights their promise as lead candidates for the development of new antimicrobial agents.47–51
2 Results and discussion
2.1 Chemistry
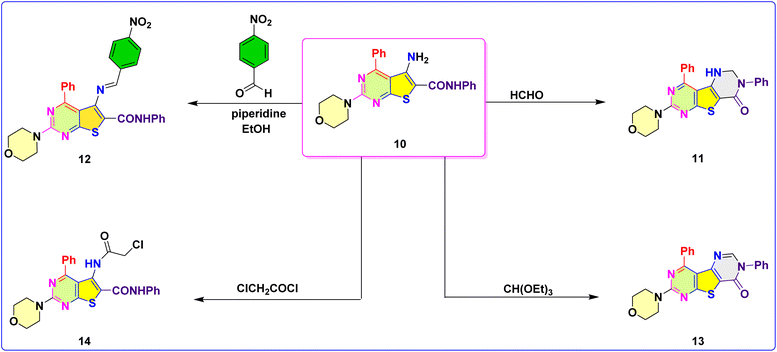
Compounds 4–8 were synthesized according to well-established literature procedures.52,53 As depicted in Scheme 1, these intermediates were subsequently employed in a mechanistically rational synthetic sequence to construct 2-((5-cyano-2-morpholino-6-phenylpyrimidin-4-yl)thio)-N-phenylacetamide (9) via alkylation of the precursor 2-morpholino-4-phenyl-6-thioxo-1,6-dihydropyrimidine-5-carbonitrile (8) with chloroacetanilide. Compound 9 then underwent a Thorpe–Ziegler-type intramolecular cyclization to afford the corresponding 5-amino-2-morpholino-N,4-diphenylthieno[2,3-d]pyrimidine-6-carboxamide (10). The structure of compound 9 was confirmed by its NMR spectra, which showed the characteristic NH proton signal at δ 8.24 ppm, along with signals in the range δ 3.97–3.66 ppm corresponding to the methylene (CH2) groups. As mentioned, the treatment of compound 9 with sodium ethoxide in refluxing ethanol led to deprotonation at the activated methylene/amide position, generating a nucleophilic anion that subsequently attacked the neighboring electrophilic carbon center, resulting in a Thorpe–Ziegler-type intramolecular cyclization. This transformation afforded 10 in good yield. The structure of 10 was confirmed by comprehensive spectroscopic characterization. The IR spectrum showed two characteristic NH2 stretching bands at 3474 and 3370 cm−1, an aliphatic C–H band at 2851 cm−1, and a strong amide C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O absorption at 1626 cm−1. The 1H NMR spectrum displayed the expected morpholine methylene resonances at δ 3.7–3.9 ppm, in addition to a broad signal at 6.10 ppm which attributed to the existence of amino group, while the mass spectrum exhibited a molecular ion peak at m/z 431, in agreement with the molecular formula C23H21N5O2S.
O absorption at 1626 cm−1. The 1H NMR spectrum displayed the expected morpholine methylene resonances at δ 3.7–3.9 ppm, in addition to a broad signal at 6.10 ppm which attributed to the existence of amino group, while the mass spectrum exhibited a molecular ion peak at m/z 431, in agreement with the molecular formula C23H21N5O2S.
As shown in Scheme 2, the Mannich reaction of compound 10 with formaldehyde in ethanol under mild reflux furnished the 7-morpholino-3,9-diphenyl-2,3-dihydrothieno[2,3-d:4,5-d′]dipyrimidin-4(1H)-one (11). The reaction proceeds through initial formation of an iminium species from formaldehyde, followed by nucleophilic attack of the amino group of 10, and intramolecular electrophilic substitution leading to annulation of the third ring. Formation of the tricyclic fused framework was supported by disappearance of the primary NH2 bands in the IR spectrum and the appearance of a new C–N stretch. The 1H NMR spectrum exhibited a new methylene singlet signal at δ 4.95 ppm, confirming Mannich incorporation, in addition to signal at 4.50 ppm characteristic for NH group. Condensation of compound 10 with p-nitrobenzaldehyde in ethanol containing catalytic glacial acetic acid led to formation of the Schiff base (E)-2-morpholino-5-((4-nitrobenzylidene)amino)-N,4-diphenylthieno[2,3-d]pyrimidine-6-carboxamide (12). The transformation followed the typical nucleophilic addition–elimination route, involving initial formation of a carbinolamine intermediate followed by dehydration to yield the azomethine functionality. The IR spectrum of 12 exhibited a strong C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching band at 1610 cm−1 alongside an amide C
N stretching band at 1610 cm−1 alongside an amide C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O band at 1654 cm−1. The 1H NMR spectrum displayed a characteristic singlet for the azomethine proton at δ 8.30 ppm, confirming imine formation. Cyclocondensation of compound 10 with triethyl orthoformate in glacial acetic acid under reflux afforded 7-morpholino-3,9-diphenylthieno[2,3-d:4,5-d′]dipyrimidin-4(3H)-one (13). The reaction likely proceeds via generation of an N-formimidate intermediate, followed by intramolecular nucleophilic attack from the amide nitrogen, cyclodehydration, and final oxidative aromatization, yielding the fused dipyrimidinone system. The IR spectrum of compound 13 revealed absorption bands at 3446 cm−1 (NH), 2849 cm−1 (aliphatic C–H), and 1677 cm−1 (C
O band at 1654 cm−1. The 1H NMR spectrum displayed a characteristic singlet for the azomethine proton at δ 8.30 ppm, confirming imine formation. Cyclocondensation of compound 10 with triethyl orthoformate in glacial acetic acid under reflux afforded 7-morpholino-3,9-diphenylthieno[2,3-d:4,5-d′]dipyrimidin-4(3H)-one (13). The reaction likely proceeds via generation of an N-formimidate intermediate, followed by intramolecular nucleophilic attack from the amide nitrogen, cyclodehydration, and final oxidative aromatization, yielding the fused dipyrimidinone system. The IR spectrum of compound 13 revealed absorption bands at 3446 cm−1 (NH), 2849 cm−1 (aliphatic C–H), and 1677 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). The 1H NMR spectrum showed morpholine methylenes at δ 4.05 and 3.81 ppm, and a C–H pyrimidine proton at δ 8.05 ppm, whereas its 13C NMR signals at δ 172.74 ppm corresponded to C
O). The 1H NMR spectrum showed morpholine methylenes at δ 4.05 and 3.81 ppm, and a C–H pyrimidine proton at δ 8.05 ppm, whereas its 13C NMR signals at δ 172.74 ppm corresponded to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O carbon, supporting formation of the dipyrimidinone system. Finally, acylation of compound 10 with 2-chloroacetyl chloride yielded 5-(2-chloroacetamido)-2-morpholino-N,4-diphenylthieno[2,3-d]pyrimidine-6-carboxamide (14) via nucleophilic acyl substitution. The amine nitrogen attacked the electrophilic carbonyl of chloroacetyl chloride, forming a tetrahedral intermediate that collapsed with chloride expulsion, consistent with a Schotten–Baumann-type acylation. IR analysis of compound 14 displayed a strong NH band at 3331 cm−1, while the 1H NMR spectrum showed two downfield NH signals at δ 9.68 and 10.07 ppm, characteristic of newly formed amide functionalities. Collectively, the sequence from compounds 9–14 demonstrates a versatile synthetic strategy enabling construction of diverse fused thienopyrimidine frameworks through sequential cyclization, Mannich annulation, Schiff base condensation, dipyrimidinone ring formation, and acylation. These chemical transformations not only expand the structural diversity of the scaffold but also generate functionalized derivatives suitable for further derivatization and biological evaluation.
O carbon, supporting formation of the dipyrimidinone system. Finally, acylation of compound 10 with 2-chloroacetyl chloride yielded 5-(2-chloroacetamido)-2-morpholino-N,4-diphenylthieno[2,3-d]pyrimidine-6-carboxamide (14) via nucleophilic acyl substitution. The amine nitrogen attacked the electrophilic carbonyl of chloroacetyl chloride, forming a tetrahedral intermediate that collapsed with chloride expulsion, consistent with a Schotten–Baumann-type acylation. IR analysis of compound 14 displayed a strong NH band at 3331 cm−1, while the 1H NMR spectrum showed two downfield NH signals at δ 9.68 and 10.07 ppm, characteristic of newly formed amide functionalities. Collectively, the sequence from compounds 9–14 demonstrates a versatile synthetic strategy enabling construction of diverse fused thienopyrimidine frameworks through sequential cyclization, Mannich annulation, Schiff base condensation, dipyrimidinone ring formation, and acylation. These chemical transformations not only expand the structural diversity of the scaffold but also generate functionalized derivatives suitable for further derivatization and biological evaluation.
2.2 Biological activity
The antimicrobial screening results of compounds 10, 11, 13, and 14 demonstrate distinct structure–activity relationships (SAR) across Gram-positive bacteria, Gram-negative bacteria, and fungal strains, with the inclusion of appropriate positive and negative controls providing a reliable framework for data interpretation (Table 1). The use of cefotaxime and fluconazole as standard antibacterial and antifungal agents, respectively, confirms the validity of the assay conditions, while the complete absence of inhibition zones for DMSO verifies that the observed activities are solely attributable to the tested compounds. The parent bifunctional amino-acetanilide precursor 10 exhibits modest antibacterial activity, showing moderate inhibition against Bacillus subtilis and weak activity toward Escherichia coli. This baseline activity can be attributed to the presence of free amino and acetanilide functionalities, which may enable limited hydrogen bonding and electrostatic interactions with bacterial cell components. Notably, compound 10 also displays measurable antifungal activity against Candida albicans, indicating that the flexible, polar scaffold may interact with fungal cell membranes or intracellular targets, albeit with moderate potency compared to fluconazole. Mannich base formation (compound 11) leads to a marked enhancement of activity against Gram-positive bacteria, particularly B. subtilis, highlighting the beneficial role of aminomethyl substitution in increasing lipophilicity and strengthening interactions with the thick peptidoglycan layer characteristic of Gram-positive cell walls. However, this modification simultaneously results in complete loss of activity against Gram-negative strains, likely due to insufficient permeability across the outer membrane barrier. Interestingly, compound 11 retains weak but detectable antifungal activity against C. albicans, suggesting that the Mannich functionality may still support limited fungal membrane interaction, though significantly less effectively than the reference antifungal agent. Cyclization of the precursor into the rigid fused pyrimidothienopyrimidine framework (compound 13) induces a pronounced shift in biological behavior. This derivative maintains good activity against B. subtilis while uniquely improving efficacy against E. coli, distinguishing it as the only compound in the series with dual activity against both Gram-positive and Gram-negative bacteria. The enhanced performance of compound 13 may arise from increased planarity, aromaticity, and electronic delocalization, which can facilitate penetration through Gram-negative membranes or interaction with intracellular bacterial targets such as nucleic acids or essential enzymes. In contrast, this rigid heterocyclic system exhibits no antifungal activity, suggesting that structural rigidity and reduced polarity may limit effective interaction with fungal cell envelopes. Acetylation of the heterocyclic system (compound 14) restores and even enhances activity against B. subtilis, likely due to increased lipophilicity and reduced hydrogen-bonding capacity, which favor penetration of Gram-positive bacterial cell walls. Nevertheless, compound 14 remains inactive against Gram-negative bacteria, reinforcing the notion that acetylation alone is insufficient to overcome the permeability barrier imposed by the outer membrane. The moderate antifungal activity observed for compound 14 against C. albicans may reflect a balance between lipophilicity and molecular flexibility, allowing partial disruption of fungal membranes. Overall, comparison with the positive controls underscores that while the synthesized compounds are less potent than cefotaxime and fluconazole, several derivatives exhibit selective and reproducible antimicrobial effects. The SAR trends clearly indicate that increased polarity and flexibility favor Gram-positive and fungal activity, whereas rigid heterocyclic fusion, as exemplified by compound 13, is crucial for extending activity toward Gram-negative organisms. These findings suggest that further structural optimization, particularly fine-tuning polarity and molecular rigidity, may lead to derivatives with broader-spectrum antimicrobial and antifungal profiles.| Standard strains | Gram positive strains | Gram negative strains | Yeast | |||
|---|---|---|---|---|---|---|
| Staphylococcus aureus | Bacillus subtilis | Pseudomonas aeruginosa | Escherichia coli | Candida albicans | ||
| Derivatives | MIC | ATCC 6538 | ATCC 6633 | ATCC 9027 | ATCC 25416 | ATCC 10231 |
| Cefotaxime | 100 µg mL−1 | 35.1 ± 0.4 | 40.7 ± 0.3 | 22.5 ± 0.5 | 34.9 ± 0.1 | — |
| Fluconazole | — | — | — | — | 38.1 ± 0.2 | |
| DMOS | 0 | 0 | 0 | 0 | 0 | |
| 10 | 0 | 12.8 ± 0.1 | 0 | 8.6 ± 0.3 | 08.7 ± 0.1 | |
| 11 | 0 | 15.3 ± 0.3 | 0 | 0 | 08.3 ± 0.2 | |
| 13 | 0 | 14.4 ± 0.3 | 0 | 11.3 ± 0.6 | 0 | |
| 14 | 0 | 15.1 ± 0.3 | 0 | 0 | 09.4 ± 0.1 | |
2.3 DFT calculations
We performed DFT calculations on the four compounds to determine their geometrical structure and reveal their electronic properties. The optimized structure, selected geometrical parameters, and the reduced density gradient (RDG) plots of compounds 11 and 14 are shown in Fig. 1. RDG plots are important tools for visualizing and interpreting noncovalent interactions within molecules and molecular complexes. By analyzing regions of low electron-density gradient, RDG plots reveal subtle interactions such as hydrogen bonds, van der Waals forces, π–π stacking, and steric repulsion, which are often difficult to observe using traditional structural analysis. The color-coded isosurfaces allow researchers to clearly distinguish between attractive and repulsive interactions, helping to explain molecular stability, binding affinity, and conformational preferences. The phenyl group has a torsional angle of 43.2° to the thienopyrimidine moiety in 11 and 45.6° in 14. The intramolecular forces within the two compounds are mostly van der Waals interactions, which are shown by green color. Compound 14 shows a hydrogen bond, represented by a blue color, between the carbonyl and imine group, with an O–HN bond distance of 1.974Å and a strength of 5.4 kcal mol−1. This intramolecular hydrogen bond could limit the ability of the compound to form hydrogen bonds with the active site of the enzyme. | ||
| Fig. 1 The optimized geometry with selected geometrical parameters and the reduced density gradient (RDG) plots of compounds 11 and 14 calculated at the B3LYP-D3(BJ)/6-31G(d) level. | ||
The distributions of the HOMO and LUMO along with their energies of the four compounds were calculated at the B3LYP-D3(BJ)/6-31G(d) level. Fig. 2 shows the HOMO–LUMO isosurface maps of compounds 10, 11, 13, and 14. The distributions of the HOMO and LUMO orbitals, along with their energy gap, play a crucial role in determining a molecule's biological activity because they influence how the molecule interacts electronically with biological targets. The HOMO distribution indicates the electron-donating regions, affecting interactions such as hydrogen bonding, π–π stacking, or nucleophilic attack, while the LUMO highlights sites that can accept electrons, which is essential for binding to electrophilic or metal-containing active sites. The HOMO–LUMO energy gap reflects the molecule's chemical reactivity and stability; a smaller gap generally corresponds to higher reactivity and easier charge transfer, which may enhance biological interactions or enzyme inhibition. Thus, analyzing HOMO and LUMO properties provides valuable insight into a compound's reactivity, binding potential, and overall pharmacological behavior.
 | ||
| Fig. 2 The HOMO and LUMO isosurface maps of compounds 10, 11, 13, and 14, their energies, and the HOMO–LUMO gap calculated at the B3LYP-D3(BJ)/6-31G(d) level. | ||
The HOMO of the four compounds is delocalized mainly on the thienopyrimidine moiety, with significant contributions from the lone pairs of the adjacent heteroatoms. The contribution of the phenyl group to the HOMO is greatly hindered by its large torsional angle to the thienopyrimidine (ranging from 41.5° in 13 to 50.3° in 10). The LUMO is delocalized on thienopyrimidine as well, but it is more spread over the conjugated system, and, unlike the HOMO, includes the phenyl group. The HOMO–LUMO energy gaps for 10, 11, 13, and 14 are 3.53, 3.56, 3.82, and 3.63 eV.
Fig. 3 depicts the molecular electrostatic potential (MESP) analysis of compounds 10, 11, 13, and 14. MESP maps are powerful tools for understanding a compound's biological activity and chemical reactivity because they visually represent the distribution of electron-rich (red color) and electron-deficient (blue color) regions across a molecule. These variations in electrostatic potential help identify sites that are prone to nucleophilic or electrophilic attack, predict hydrogen-bond donors and acceptors, and clarify how a molecule may interact with biological macromolecules such as enzymes or receptors. By highlighting potential binding hotspots and reactive centers, MESP analysis provides insight into a molecule's pharmacological behavior, guiding the design of more selective and potent bioactive compounds. Fig. 3 gives a numerical expression of the electrophilic and nucleophilic sites, and thus shows the most reactive sites, within each compound.
 | ||
| Fig. 3 The molecular electrostatic potential (MESP) maps of compounds 10, 11, 13, and 14 with ESP surface maxima and minima (in kcal mol−1) calculated at the B3LYP-D3(BJ)/6-31G(d) level. | ||
The most nucleophilic site within each compound is produced by the lone pairs of the carbonyl group. The highest negative electrostatic potential (ESP) is present in 11, with ESP = −45 kcal mol−1. The oxygen of the morpholine ring has an ESP of −29 kcal mol−1, and the two nitrogens of the pyrimidine ring produces an ESP of −22 and −23 kcal mol−1. The highest negative ESP in 10, 13, and 14 are −37, −40, and −36 kcal mol−1. The highest electrophilic sites are around hydrogen of the imine and pyrimidine-4H-one, with a positive ESP of 37 kcal mol−1. These results suggest the importance of ionic and dipole–dipole interactions between the ligand and the active site of the enzyme.
2.4 Molecular docking
Molecular docking is a computational approach commonly used to predict the spatial orientation and binding affinity between interacting molecules (Table 2). It is widely applied to identify protein binding sites and to model ligand–protein interactions. In this study, docking analyses were carried out for the four compounds to determine their binding conformations and interactions with the target protein. To assess their antimicrobial potential, simulations were performed against two DNA gyrase B structures: PDB ID 4URO for B. subtilis and PDB ID 4DUH for E. coli. DNA gyrase B is an essential enzyme in Gram-positive and Gram-negative bacteria, required for DNA replication and chromosomal compaction. Its inhibition prevents the relaxation of supercoiled DNA, thereby disrupting replication, blocking cell division, and ultimately causing bacterial cell death.54 Ten binding poses were generated for each protein–ligand pair using the CB-Dock2 server. The pose with the most favorable binding energy and lowest Vina score, indicating the strongest interaction within the active site, was selected. The Vina scores for all compounds are provided in Table 1. This docking procedure was validated by us in several previous studies.55–57 Moreover, we performed docking of the crystallized cofactors with the same parameters: novobiocin with 4URO and 4-[4′-methyl-2'-(propanoylamino)-4,5′-bi-1,3-thiazol-2-yl]aminobenzoic acid with 4DUH. Overall, all compounds demonstrated strong binding affinities to the enzyme's active site, confirming their suitability as inhibitors for this protein.| Compound | Dipole moment (debye) | Vina score (kcal mol−1) | |
|---|---|---|---|
| B. subtilis | E. coli | ||
| 10 | 1.9 | −7.5 | −8.1 |
| 11 | 6.6 | −8.6 | −8.2 |
| 13 | 3.8 | −8.3 | −10.4 |
| 14 | 2.0 | −7.6 | −7.5 |
| Novobiocin | — | −9.9 | — |
| Cofactor | — | — | −8.5 |
All compounds exhibited strong and diverse interactions with the staph gyrase B. The details of the interactions of the most potent compounds, 10, 11, 13, and 14 with the active site of the enzyme are depicted in Fig. 4. Compound 10 forms two hydrogen bonds with the enzyme: the amino group with ASP 89 and the O of the morpholine ring with ARG 84. The phenyl groups of the compound exhibited hydrophobic interactions with ASP 89, GLU 193, ARG 200, and GLU 201. The thienopyrimidine moiety showed hydrophobic interactions with PRO 87 and cation–π interactions with ARG 144 and ARG 200. The later forms cation–π interactions with the phenyl group as well. In compound 11, which has the highest biological activity and lowest Vina score, the carbonyl group is hydrogen bonded to the hydroxyl group of THR 173 and GLU 58. The thienopyrimidine moiety exhibits cation–π interactions with ARG 84. The phenyl group shows hydrophobic interactions with ILE 86, ILE 175, and ASN 54.
 | ||
| Fig. 4 The geometry of the most stable binding pose of compounds 10, 11, 13, and 14 against staph gyrase B (PDB code: 4URO) along with types of interaction using AutoDock Vina. | ||
In 13, the carbonyl group forms hydrogen bonds with ILE 86 and GLN 3. The nitrogen of the pyrimidinone ring exhibits ionic interactions with the carboxyl group of GLU 58. The thienopyrimidine moiety exhibits cation–π interactions with ARG 84. The two phenyl groups show several hydrophobic interactions: ASP 57, ALA 61, ASN 54, ILE 86, THR 173, ILE 175. Compound 14 forms only one hydrogen bond: the carbonyl group with the amino group of ARG 84. The thienopyrimidine moiety shows cation–π interactions with the same amino acid. The phenyl group exhibits cation–π interactions with ARG 200 and hydrophobic interactions with PRO 87.
To investigate the binding affinity and molecular interactions of the four compounds with E. coli, we performed molecular docking with gyrase B (PDB code: 4DUH). The binding affinities of the four compounds are listed in Table 1 and the geometry of the most stable pose and the type of interactions with the active site of the enzyme are depicted in Fig. 5. Compound 10 forms hydrogen bonds with GLU 50 and ALA 100. The thienopyrimidine exhibits hydrophobic interactions with LYS 103; the phenyl groups with PRO 79, ILE 78, GLU 50, and ASN 46. The interaction of 11 with the active site of the enzyme is dominated by hydrophobic interactions: the phenyl groups with ILE 94, LYS 103, ILE 78, THR 165, VAL 167, VAL 43, VAL 71, ALA 47, and ASN 46. The compound forms hydrogen bonds with THR 165 and GLY 101 and cation–π interactions with LYS 103. Compound 13 is hydrogen bonded to THR 165, LYS 103 and GLY 101. The thienopyrimidine has cation–π interactions with LYS 103. The two phenyl groups form several hydrophobic interactions: LYS 103, ILE 94, THR 165, VAL 167, VAL 43, VAL 71, ALA 47, and ASN 46. In 14, a hydrogen bond is formed between the imine group and GLU 58. The two phenyl groups show hydrophobic interactions with GLU 58, ILE 60, GLN 72, and ASP 74.
 | ||
| Fig. 5 The geometry of the most stable binding pose of compounds 10, 11, 13, and 14 against the E. coli DNA gyrase B (PDB code: 4DUH) along with types of interaction using AutoDock Vina. | ||
Our investigation highlights the importance of electronic distributions of the compound for the type of interaction with the active site of the receptor and its biological efficacy. The most potent compound for B. subtilis was 11. This compound possesses the highest dipole moment, 6.6 debye, and the highest negative electrostatic potential, −45 kcal mol−1 among the four molecules. These results demonstrate the importance of ionic interactions for the inhibitors of B. subtilis.
3 Conclusions
In summary, a series of novel thienopyrimidine derivatives was successfully synthesized and systematically evaluated to explore their antimicrobial potential and molecular interaction profiles. Structural modifications across the scaffold produced clear SAR trends, demonstrating that increased polarity through Mannich substitution selectively enhanced Gram-positive activity, whereas rigid fused heterocycles improved the ability to inhibit Gram-negative organisms. Compound 11 exhibited the highest activity against B. subtilis, while compound 13 emerged as the most promising broad-spectrum candidate due to its enhanced inhibition of E. coli and superior docking affinity toward DNA gyrase B. DFT analyses corroborated these findings by revealing key electronic features, reactive sites, and stabilizing noncovalent interactions that influence binding affinity and biological response. Molecular docking further confirmed strong engagement of the synthesized molecules with essential bacterial targets, providing mechanistic insight into their inhibitory potential. Overall, this integrated experimental–computational study highlights thienopyrimidine scaffolds as compelling platforms for future antimicrobial drug development and offers valuable guidance for rational structural optimization.4 Experimental
4.1 Synthesis of the target compounds
Compounds 4–8 were prepared following previously reported procedure.52,53 These intermediates are known in the literature, and their physical and spectral properties were consistent with the reported values. These intermediates were used directly in subsequent reactions without further modification, ensuring a smooth progression toward the target compounds.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) mixture to afford the pure product as white crystals (1.0 g, 71% yield) with a melting point of 218–220 °C. 1H NMR (400 MHz, chloroform-d) δ 8.24 (s, 1H), 7.94 (dt, J = 3.3, 1.5 Hz, 2H), 7.52–7.43 (m, 5H), 7.38–7.28 (m, 2H), 7.17–7.10 (m, 1H), 3.99–3.88 (m, 4H), 3.69 (m, 6H). 13C NMR (126 MHz, chloroform-d) δ 164.10, 163.98, 147.30, 137.28, 135.23, 133.96, 130.38, 129.55, 128.93, 128.71, 128.60, 128.53, 120.86, 112.99, 91.82, 67.12, 66.88, 44.58, 20.96.
1) mixture to afford the pure product as white crystals (1.0 g, 71% yield) with a melting point of 218–220 °C. 1H NMR (400 MHz, chloroform-d) δ 8.24 (s, 1H), 7.94 (dt, J = 3.3, 1.5 Hz, 2H), 7.52–7.43 (m, 5H), 7.38–7.28 (m, 2H), 7.17–7.10 (m, 1H), 3.99–3.88 (m, 4H), 3.69 (m, 6H). 13C NMR (126 MHz, chloroform-d) δ 164.10, 163.98, 147.30, 137.28, 135.23, 133.96, 130.38, 129.55, 128.93, 128.71, 128.60, 128.53, 120.86, 112.99, 91.82, 67.12, 66.88, 44.58, 20.96.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide). 1H NMR (400 MHz, DMSO-d6) δ 8.29 (d, J = 8.8 Hz, 2H), 7.68–7.63 (m, 3H), 7.63–7.58 (m, 4H), 7.37 (d, J = 4.8 Hz, 4H), 7.25–7.14 (m, 1H), 6.80 (d, J = 5.5 Hz, 1H), 6.50 (d, J = 5.2 Hz, 1H), 3.82 (t, J = 4.6 Hz, 4H), 3.66 (dd, J = 5.7, 3.9 Hz, 4H). 13C NMR (126 MHz, CDCl3) δ 173.79, 163.51, 161.01, 159.43, 144.98, 140.50, 137.71, 130.48, 128.96, 128.93, 128.59, 125.91, 124.60, 111.30, 103.94, 66.88, 62.26, 44.61. MS (ESI): calculated for C30H24N6O4S, M+ = 564.16; found m/z = 564.16.
O amide). 1H NMR (400 MHz, DMSO-d6) δ 8.29 (d, J = 8.8 Hz, 2H), 7.68–7.63 (m, 3H), 7.63–7.58 (m, 4H), 7.37 (d, J = 4.8 Hz, 4H), 7.25–7.14 (m, 1H), 6.80 (d, J = 5.5 Hz, 1H), 6.50 (d, J = 5.2 Hz, 1H), 3.82 (t, J = 4.6 Hz, 4H), 3.66 (dd, J = 5.7, 3.9 Hz, 4H). 13C NMR (126 MHz, CDCl3) δ 173.79, 163.51, 161.01, 159.43, 144.98, 140.50, 137.71, 130.48, 128.96, 128.93, 128.59, 125.91, 124.60, 111.30, 103.94, 66.88, 62.26, 44.61. MS (ESI): calculated for C30H24N6O4S, M+ = 564.16; found m/z = 564.16.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O amide). 1H NMR (400 MHz, chloroform-d) δ 8.05 (s, 1H), 7.96–7.83 (m, 2H), 7.59–7.45 (m, 6H), 7.44–7.37 (m, 2H), 4.04 (t, J = 4.8 Hz, 4H), 3.82 (t, J = 4.8 Hz, 4H). 13C NMR (101 MHz, DMSO) δ 172.74, 158.51, 148.29, 136.20, 129.65, 128.62, 126.91, 126.78, 115.61, 71.88, 65.34, 62.45. MS (ESI): calculated for C24H19N5O2S, M+ = 441.13; found m/z = 441.17.
O amide). 1H NMR (400 MHz, chloroform-d) δ 8.05 (s, 1H), 7.96–7.83 (m, 2H), 7.59–7.45 (m, 6H), 7.44–7.37 (m, 2H), 4.04 (t, J = 4.8 Hz, 4H), 3.82 (t, J = 4.8 Hz, 4H). 13C NMR (101 MHz, DMSO) δ 172.74, 158.51, 148.29, 136.20, 129.65, 128.62, 126.91, 126.78, 115.61, 71.88, 65.34, 62.45. MS (ESI): calculated for C24H19N5O2S, M+ = 441.13; found m/z = 441.17.4.2 Antimicrobial activity
Four new synthesized compounds were assessed for their antimicrobial activity against some standard reference strains, comprising two Gram-positive bacteria, Staphylococcus aureus ATCC 6538 (S. aureus) and Bacillus subtilis ATCC 6633 (B. subtilis), as well as two Gram-negative bacteria, Escherichia coli ATCC 8739 (E. coli) and Pseudomonas aeruginosa ATCC 9027 (P. aeruginosa). Additionally, Candida albicans ATCC 10231 (C. albicans) was included as a representative yeast strain. The antimicrobial efficacy against these standard strains was determined by comparison with control activity alone. Cultures of the microorganisms were incubated overnight, then suspended and adjusted to a turbidity equivalent to 0.5 McFarland standards (1.5 × 108 CFU mL−1).4.3 DFT calculations
We performed density functional theory (DFT) calculations on all compounds to reveal their structure and gain insights into their properties. The ground state geometry of all structures was optimized using DFT with the B3LYP functional, the D3BJ correction, and the basis set 6-31G(d). This dispersion correction is necessary to account for long-range and noncovalent interactions. Several conformers were considered for the ground state geometry, and the one with lowest energy was selected. The lack of imaginary frequencies proved that these geometries are true minima. All geometry optimizations were performed using the Gaussian 16 software. Molecular electrostatic potential surface maps, RDG plots, and the strength of the intramolecular hydrogen bond in 14 were obtained by the Multiwfn program based on the DFT calculations.584.4 Molecular docking
We performed molecular docking for all four derivatives to reveal binding geometries and protein–ligand interactions. Docking was done using AutoDock Vina59,60 through the CB-Dock2 server.61 We used the previously optimized geometries of the derivatives. The crystal structure of the staph gyrase B (PDB code: 4URO) and the E. coli DNA gyrase B (PDB code: 4DUH) were obtained from the protein data bank. The proteins and all ligands were prepared by the default settings of the CB-Dock2 server. We performed an auto blind docking, in which no pocket is provided for the program. For each compound, ten binding models were predicted and the binding energy of each model was calculated.Conflicts of interest
There are no conflicts of interest to declare.Data availability
All the data associated with this article is available in the main article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra09678b.References
- M. Abdel-Megid, K. M. Elmahdy, A. M. Elkazak, H. M. Seada and F. O. Mohamed, Chemistry of Thienopyrimidines and Their Biological Applications, J. pharm. appl. chem., 2016, 2, 78–102, DOI:10.18576/jpac/020301.
- E. Vitaku, D. T. Smith and J. T. Njardarson, Analysis of the Structural Diversity, Substitution Patterns, and Frequency of Nitrogen Heterocycles among U.S. FDA Approved Pharmaceuticals, J. Med. Chem., 2014, 57, 10257–10274, DOI:10.1021/jm501100b.
- M. M. Heravi and V. Zadsirjan, Prescribed drugs containing nitrogen heterocycles: an overview, RSC Adv., 2020, 10, 44247–44311, 10.1039/D0RA09198G.
- E. Kabir and M. Uzzaman, A review on biological and medicinal impact of heterocyclic compounds, Results Chem., 2022, 4, 100606, DOI:10.1016/j.rechem.2022.100606.
- S. Nadar and T. Khan, Pyrimidine: An elite heterocyclic leitmotif in drug discovery-synthesis and biological activity, Chem. Biol. Drug Des., 2022, 100, 818–842, DOI:10.1111/cbdd.14001.
- S. Iacob Ciobotaru, et al., Hybrid Molecules with Purine and Pyrimidine Derivatives for Antitumor Therapy: News, Perspectives, and Future Directions, Molecules, 2025, 30, 1–25, DOI:10.3390/molecules30132707.
- G. J. Martis, P. S. Mugali and S. L. Gaonkar, Recent Progress in the Synthesis, Functionalization, and Biological Outlook of Pyrimidines, ACS Omega, 2025, 10, 46248–46271, DOI:10.1021/acsomega.5c04880.
- E. M. Mohi El-Deen, E. A. Abd El-Meguid, S. Hasabelnaby, E. A. Karam and E. S. Nossier, Synthesis, Docking Studies, and In Vitro Evaluation of Some Novel Thienopyridines and Fused Thienopyridine-Quinolines as Antibacterial Agents and DNA Gyrase Inhibitors, Molecules, 2019, 24(20), 1–20, DOI:10.3390/molecules24203650.
- A. T. Alsaggaf, et al., New class of Thienopyridines: Design, synthesis, antimicrobial activity and molecular docking study, J. Mol. Struct., 2025, 1323, 140741, DOI:10.1016/j.molstruc.2024.140741.
- H. Amawi, et al., Thienopyrimidine derivatives exert their anticancer efficacy via apoptosis induction, oxidative stress and mitotic catastrophe, Eur. J. Med. Chem., 2017, 138, 1053–1065, DOI:10.1016/j.ejmech.2017.07.028.
- S. Nadar, M. Borkar and T. Khan, Thienopyrimidine: Unveiling the Versatile Potential of a Promising Heterocyclic Scaffold in Drug Discovery, Chem. Biol. Drug Des., 2025, 105, e70146, DOI:10.1111/cbdd.70146.
- H. u. Rashid, et al., Research developments in the syntheses, anti-inflammatory activities and structure–activity relationships of pyrimidines, RSC Adv., 2021, 11, 6060–6098, 10.1039/D0RA10657G.
- G. Rathika, et al., Exploring the anticancer potential of pyrimidine derivatives via molecular docking and dynamics, Discov. Appl. Sci., 2025, 7, 1319, DOI:10.1007/s42452-025-07840-2.
- A. M. Abdelhamed, A. A. Helwa, H. H. Kadry and R. A. Hassan, Pyrazolopyrimidines: A Promising Frontier in Cancer Treatment-Reviewing Their Potential as Inhibitors of Serine/Threonine Kinases, Chem. Biodiversity, 2025, 22, e202403071, DOI:10.1002/cbdv.202403071.
- T. H. Iorkula, et al., Advances in pyrazolo[1,5-a]pyrimidines: synthesis and their role as protein kinase inhibitors in cancer treatment, RSC Adv., 2025, 15, 3756–3828, 10.1039/D4RA07556K.
- V. Sorhie, et al., A critical review on the sustainable synthesis of pyrimidine-based heterocycles and their biological activities, Tetrahedron, 2025, 179, 134626, DOI:10.1016/j.tet.2025.134626.
- F. S. Tokalı, et al., Design and synthesis of new thienopyrimidine derivatives as potential anticancer agents: From cytotoxicity screening to VEGFR inhibition modeling, J. Mol. Struct., 2026, 1352, 144425, DOI:10.1016/j.molstruc.2025.144425.
- O. H. Rizk, O. G. Shaaban and I. M. El-Ashmawy, Design, synthesis and biological evaluation of some novel thienopyrimidines and fused thienopyrimidines as anti-inflammatory agents, Eur. J. Med. Chem., 2012, 55, 85–93, DOI:10.1016/j.ejmech.2012.07.007.
- S. Thakur, et al., Medicinal chemistry-based perspectives on thiophene and its derivatives: exploring structural insights to discover plausible druggable leads, RSC Med. Chem., 2025, 16, 481–510, 10.1039/d4md00450g.
- Jatin, et al., Thiochromenes and thiochromanes: a comprehensive review of their diverse biological activities and structure-activity relationship (SAR) insights, RSC Med. Chem., 2025, 16, 1941–1968, 10.1039/d4md00995a.
- H. T. Hamad, The anti-cancer effectiveness of some heterocyclic compounds containing sulfur atom, Results Chem., 2025, 15, 102182, DOI:10.1016/j.rechem.2025.102182.
- Y.-H. Liu, et al., Thienopyrimidine: A promising scaffold in the development of kinase inhibitors with anticancer activities, Bioorg. Med. Chem., 2025, 121, 118109, DOI:10.1016/j.bmc.2025.118109.
- M. M. Fouad, H. A. Ghabbour, I. A. Shehata and M. B. El-Ashmawy, Synthesis and in vitro antitumor evaluation of new thieno[2,3-d]pyrimidine derivatives as EGFR and DHFR inhibitors, Bioorg. Chem., 2024, 148, 107401, DOI:10.1016/j.bioorg.2024.107401.
- H. Amawi, et al., Thienopyrimidine derivatives exert their anticancer efficacy via apoptosis induction, oxidative stress and mitotic catastrophe, Eur. J. Med. Chem., 2017, 138, 1053–1065, DOI:10.1016/j.ejmech.2017.07.028.
- W. M. Alamoudi, Synthesis of Thienopyrimidine Derivatives and Their Antibacterial Activity: A Focus on Mannich Bases and Molecular Docking, Arabian J. Sci. Eng., 2025 DOI:10.1007/s13369-025-10574-3.
- W. M. Alamoudi, Synthesis of Thienopyrimidine Derivatives and Their Antibacterial Activity: A Focus on Mannich Bases and Molecular Docking, Arabian J. Sci. Eng., 2025, 1–16 Search PubMed.
- R. A. Hassan, et al., Synthesis and evaluation of antibacterial activity of new thieno [2, 3-d] pyrimidine hybrid compounds targeting dihydropteroate synthase to combat antibiotic resistance, Bioorg. Chem., 2025, 108549 Search PubMed.
- J. Qurban, et al., Antimicrobial activity, synthesis, and docking study of some novel arylazo-1, 3-thiazolopyrimidine and arylazo-1, 3-thiazolopyridopyrimidine derivatives, BMC Chem., 2025, 19, 148 Search PubMed.
- H. M. Abosalim, et al., Exploring Pyrazoline-Thiophene Hybrids as CDK2 Inhibitors: Synthesis, Mechanistic, Biological Studies, and Computational Insights, RSC Med. Chem., 2026 10.1039/D5MD00910C.
- N. T. Hatvate, T. S. Shevkar & H. Akolkar, in Examining Biological Relevance of Fused S-Heterocycles 219-270, IGI Global Scientific Publishing, 2025 Search PubMed.
- A. Glaviano, et al., PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer, Mol. Cancer, 2023, 22, 138, DOI:10.1186/s12943-023-01827-6.
- C. Tomuleasa, et al., Therapeutic advances of targeting receptor tyrosine kinases in cancer, Signal Transduct. Targeted Ther., 2024, 9, 201, DOI:10.1038/s41392-024-01899-w.
- K. W. K. Tang, B. C. Millar and J. E. Moore, Antimicrobial Resistance (AMR), Br. J. Biomed. Sci., 2023, 80, 11387, DOI:10.3389/bjbs.2023.11387.
- K. Ralhan, et al., Navigating Antibacterial Frontiers: A Panoramic Exploration of Antibacterial Landscapes, Resistance Mechanisms, and Emerging Therapeutic Strategies, ACS Infect. Dis., 2024, 10, 1483–1519, DOI:10.1021/acsinfecdis.4c00115.
- M. E. Elshobary, et al., Combating Antibiotic Resistance: Mechanisms, Multidrug-Resistant Pathogens, and Novel Therapeutic Approaches: An Updated Review, Pharmaceuticals, 2025, 18(402), 1–39, DOI:10.3390/ph18030402.
- R. J. Obaid, et al., Inhibitory potential of nitrogen, oxygen and sulfur containing heterocyclic scaffolds against acetylcholinesterase and butyrylcholinesterase, RSC Adv., 2022, 12, 19764–19855, 10.1039/D2RA03081K.
- J. K. Nkoana, G. K. More, A. A. Elhenawy and M. J. Mphahlele, Development of the 5-Nitrosalicaldehyde Arylhydrazones as Anticancer (MCF-7 and HCT116) Agents Targeting Inhibition of VEGFR-2 and Cytokines (IL-6 and TNF-α), ACS Omega, 2025, 10, 58943–58962, DOI:10.1021/acsomega.5c07719.
- P. Lagardère, C. Fersing, N. Masurier and V. Lisowski, Thienopyrimidine: A Promising Scaffold to Access Anti-Infective Agents, Pharmaceuticals, 2022, 15, 35 Search PubMed.
- N. Alotaiq and D. Dermawan, Computational Investigation of Montelukast and Its Structural Derivatives for Binding Affinity to Dopaminergic and Serotonergic Receptors: Insights from a Comprehensive Molecular Simulation, Pharmaceuticals, 2025, 18(559), 1–36, DOI:10.3390/ph18040559.
- D. J. Huggins, W. Sherman and B. Tidor, Rational Approaches to Improving Selectivity in Drug Design, J. Med. Chem., 2012, 55, 1424–1444, DOI:10.1021/jm2010332.
- S. Kumar and B. Narasimhan, Therapeutic potential of heterocyclic pyrimidine scaffolds, Chem. Cent. J., 2018, 12, 38, DOI:10.1186/s13065-018-0406-5.
- R. Shaw, et al., Recent advancements in the synthesis of fused thienopyridines and their therapeutic applications, Eur. J. Med. Chem. Rep., 2024, 12, 100185, DOI:10.1016/j.ejmcr.2024.100185.
- C. M. F. Ancajas, A. S. Oyedele, C. M. Butt and A. S. Walker, Advances, opportunities, and challenges in methods for interrogating the structure activity relationships of natural products, Nat. Prod. Rep., 2024, 41, 1543–1578, 10.1039/D4NP00009A.
- E. S. Mozafari, M. T. Baei and E. T. Lemeski, Computational study of the therapeutic properties of allicin and its nanocomplexes using DFT and molecular docking techniques, Sci. Rep., 2025, 15, 23034, DOI:10.1038/s41598-025-03293-0.
- G. Kumar, P. Kumar, A. Soni, V. Sharma and M. Nemiwal, Efficient synthesis and molecular docking analysis of quinazoline and azole hybrid derivatives as promising agents for anti-cancer and anti-tuberculosis activities, J. Mol. Struct., 2024, 1310, 138289, DOI:10.1016/j.molstruc.2024.138289.
- O. Younis, et al., Solid-State Luminescent Materials Containing Both Indole and Pyrimidine Moieties: Design, Synthesis, and Density Functional Theory Calculations, ACS Omega, 2022, 7, 15016–15026, DOI:10.1021/acsomega.2c00775.
- M. Ahmed, et al., Synthesis, antimicrobial activity, molecular docking and molecular dynamics studies of novel bioactive compounds derived from propylthiouracil, J. Mol. Struct., 2025, 1333, 141779, DOI:10.1016/j.molstruc.2025.141779.
- M. Ahmed, et al., Synthesis, Characterization, and Antimicrobial Activity of New Thienopyrimidine Derivatives, Polycyclic Aromat. Compd., 2022, 42, 3079–3088, DOI:10.1080/10406638.2020.1852587.
- S. Mostafa, et al., Anti-inflammatory Activity and Computational Biology Study of Indole/Pyrimidine Hybrids, Curr. Org. Chem., 2024, 28, 56–64, DOI:10.2174/0113852728289430231227042754.
- A. F. Saber, et al., A facile method for preparation and evaluation of the antimicrobial efficiency of various heterocycles containing thieno[2,3-d]pyrimidine, Synth. Commun., 2021, 51, 398–409, DOI:10.1080/00397911.2020.1829645.
- M. Sayed, A. M. Kamal El-Dean, M. Ahmed and R. Hassanien, Design, synthesis, and characterization of novel pyrimidines bearing indole as antimicrobial agents, J. Chin. Chem. Soc., 2019, 66, 218–225, DOI:10.1002/jccs.201800115.
- A. A. Saddik, A. M. Kamal El-Dean, W. A. El-Said, K. M. Hassan and M. S. Abbady, Synthesis, Antimicrobial, and Anticancer Activities of a New Series of Thieno[2,3-d] Pyrimidine Derivatives, J. Heterocycl. Chem., 2018, 55, 2111–2122, DOI:10.1002/jhet.3256.
- A. A. Saddik, et al., Synthesis and Cytotoxicity of Some Thieno[2,3-d]pyrimidine Derivatives, J. Chin. Chem. Soc., 2017, 64, 87–93, DOI:10.1002/jccs.201600279.
- F. Collin, S. Karkare and A. Maxwell, Exploiting bacterial DNA gyrase as a drug target: current state and perspectives, Appl. Microbiol. Biotechnol., 2011, 92, 479–497, DOI:10.1007/s00253-011-3557-z.
- M. K. Mohamed, A. A. K. Mohammed, M. M. M. Aly, A.-A. M. Gaber and A. A. Saddik, Synthesis, DFT Calculations, and Molecular Docking Study of New Aggregation-Induced Emission Lumiogens Based on Thieno[2,3-d]pyrimidine Moiety, Asian J. Org. Chem., 2025, 14, e202400670, DOI:10.1002/ajoc.202400670.
- A. Z. Ebrahim, A. A. K. Mohammed, M. Ahmed, A. M. Kamal El-Dean and M. S. Tolba, New AIEgens based on thieno[2,3-d]pyrimidine derivatives: Synthesis, DFT calculations, and molecular docking, J. Photochem. Photobiol., A, 2025, 459, 116065, DOI:10.1016/j.jphotochem.2024.116065.
- A. Mahdy, J. A. Zahra, R. N. Haddadin, A. A. K. Mohammed and M. G. Mohamed, Molecular design and structural characterization of thiocarbohydrazide-derived azomethine-linked polybenzoxazine with antibacterial properties for cotton fabric applications, J. Mol. Struct., 2026, 1352, 144496, DOI:10.1016/j.molstruc.2025.144496.
- T. Lu and F. Chen, Multiwfn: A multifunctional wavefunction analyzer, J. Comput. Chem., 2012, 33, 580–592, DOI:10.1002/jcc.22885.
- O. Trott and A. J. Olson, AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 2010, 31, 455–461, DOI:10.1002/jcc.21334.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings, J. Chem. Inf. Model., 2021, 61, 3891–3898, DOI:10.1021/acs.jcim.1c00203.
- Y. Liu, et al., CB-Dock2: improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting, Nucleic Acids Res., 2022, 50, W159–W164, DOI:10.1093/nar/gkac394.
Footnote |
| † Theses authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |