Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Regioselective synthesis, structural investigation and binding behavior of 6-aroyl-7-aryl-5-methyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidines with BSA using various spectroscopic and in silico methods
Ranjana Aggarwal *ab,
Manisha Sharma
*ab,
Manisha Sharma a,
Garima Sumran
a,
Garima Sumran c,
Suresh Kumar
c,
Suresh Kumar d and
Parvin Kumar
d and
Parvin Kumar a
a
aDepartment of Chemistry, Kurukshetra University, Kurukshetra-136119, Haryana, India. E-mail: ranjana67in@yahoo.com; ranjanaaggarwal67@gmail.com; Tel: +91-9896740740
bCouncil of Scientific and Industrial Research (CSIR) HQ, Vigyan Suchna Bhawan, 14, Satsang Vihar Marg, New Delhi-110067, India
cDepartment of Chemistry, D. A. V. College (Lahore), Ambala City, Haryana 134 003, India
dDepartment of Chemistry, SUS Govt. College, Matak Majri, Karnal, Haryana, 132041, India
First published on 10th March 2026
Abstract
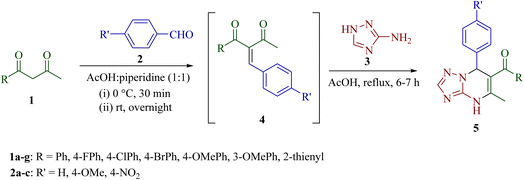
A series of 6-aroyl-7-aryl-5-methyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidines 5 was synthesized regioselectively via a one-pot multicomponent reaction between β-diketone 1, aromatic aldehyde 2, and 3-amino-1H-1,2,4-triazole 3. The regioisomeric structure of the newly synthesized compounds 5 was unambiguously determined using 1H NMR, 13C NMR, and rigorous multinuclear 2D-NMR spectroscopy [1H–13C] HMBC, [1H–13C] HSQC and [1H–15N] HMBC. The remarkable features of this protocol are high yields, operational simplicity, use of commercially available reagents and broad substrate scope. The interactions of selected compounds (5f, 5m and 5t) with bovine serum albumin (BSA) were studied by UV-vis spectroscopy, steady-state fluorescence, and molecular docking. The results indicated that compound 5t could effectively quench the intrinsic fluorescence of BSA via a static quenching process. Competitive binding studies using site markers demonstrated that compound 5t binds to site I of BSA. Binding constants for [1,2,4]triazolo[1,5-a]pyrimidines show that the affinity of 5t binding to BSA is stronger than that of 5f and 5m.
1. Introduction
[1,2,4]Triazolo[1,5-a]pyrimidine and its derivatives have profound applications in both medical and agricultural fields because of their varied biological activities and synthetic versatility.1,2 This system exhibits favourable physicochemical characteristics such as planarity, hydrogen-bonding capability, and multiple nitrogen donor sites along with multiple substitution sites (5,6,7) at the pyrimidine ring and the capacity to interact with diverse biological targets. Due to its structural similarity with purines, this scaffold has been extensively investigated as a possible isosteric replacement for the purine ring.3 [1,2,4]Triazolo[1,5-a]pyrimidines have a broad spectrum of biological activities including antiviral,4 antimicrobial,5,6 antiparasitic,7 anticancer,8 anti-inflammatory,9 and anticonvulsant properties.10 These compounds have emerged as corrosion inhibitors11 and promising PDE2A inhibitors with potential in treating Alzheimer's disease and other neurodegenerative disorders.12–14 The [1,2,4]triazolo[1,5-a]pyrimidine scaffold is a key structural motif in several clinically and pharmacologically relevant drugs, including Trapidil (a vasodilator/antiplatelet agent),15 Flumetsulam (a herbicide),16 Essramycin (an antibacterial agent),17 and GNF6702 (a broad-spectrum antiprotozoal proteasome inhibitor),18 highlighting its broad therapeutic versatility (Fig. 1).Proteins serve as primary molecular targets owing to their pivotal roles in cellular signaling and regulation and their ability to form drug–protein interactions enabling selective modulation of biochemical pathways for therapeutic effects.19 Bovine serum albumin (BSA) is a globular protein widely used in biochemical and pharmaceutical research due to its structural similarity to human serum albumin (HSA) and its ability to bind a variety of ligands.20 BSA, a type of serum albumin, plays a crucial role in maintaining blood pH, oncotic pressure and the transport of endogenous and exogenous compounds, including fatty acids, drugs, and other metabolites. BSA comprises 583 amino acids and is structured into three homologous α-helical domains.21 It contains three major drug-binding sites: site I (subdomain IIA), site II (subdomain IIIA), and site III (subdomain IB), each exhibiting distinct affinities for ligands. BSA serves as a valuable model for studying drug–protein interactions, pharmacokinetics, and drug delivery mechanisms due to its wide availability, low cost, stability, and unusual binding properties.
Two main synthetic approaches have been developed for the construction of [1,2,4]triazolo[1,5-a]pyrimidines. The first involves the annulation of a triazole ring onto a pyrimidine precursor,22 while the second and most commonly used approach involves the annulation of a pyrimidine moiety onto a triazole core.23 Literature reports the synthesis of [1,2,4]triazolo[1,5-a]pyrimidines via a cyclocondensation reaction of 3-amino-1,2,4-triazole with 1,3-dicarbonyl compounds or α,β-unsaturated carbonyl compounds.24 Multicomponent reactions (MCRs) are of significant importance in synthetic chemistry due to their operational simplicity, high atom economy, and capacity to rapidly generate structurally diverse and complex scaffolds from readily available starting materials in a single step, thereby minimizing time, resource consumption, and byproduct formation. Therefore, another efficient synthetic approach to triazolo[1,5-a]pyrimidines is Biginelli-like three-component heterocyclization of 3-amino-1,2,4-triazole with aldehyde and active methylene compounds (β-keto ester,25 cyclohexane-1,3-dione,26 β-ketonitrile,27 malononitrile,28 and acetoacetanilide29). Four-component condensation of 1,3-indanedione, aromatic aldehyde, 5-amino-4H-1,2,4-triazol-3-thiol and phenacyl bromide has also been reported to afford triazolo[1,5-a]pyrimidines.30 El-Deeb et al. reported the synthesis of 2-[(4-chlorophenyl)-7-(4-methoxyphenyl)-5-methyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidin-6-yl]phenylmethanone in 27% yield via a multicomponent reaction of 3-(4-chlorophenyl)-4H-[1,2,4]triazol-5-amine, 4-methoxybenzaldehyde, and benzoylacetone under reflux in glacial acetic acid for 36 h.31 However, to the best of our knowledge, the reaction of 3-amino-1H-1,2,4-triazole with aldehydes and unsymmetrical aryl 1,3-diketones has not yet been reported in the literature. Accordingly, this work was designed to synthesize triazolopyrimidine derivatives bearing diverse substitutions at the 5,6,7-positions by varying the aromatic aldehydes and 1,3-diketone components.
Keeping in view the biological importance of [1,2,4]triazolo[1,5-a]pyrimidines and following our ongoing work on the multicomponent synthesis of novel bioactive heterocycles,32,33 we herein report a one-pot multicomponent reaction for regioselective synthesis of 6-aroyl-7-aryl-5-methyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidine derivatives (5) using 3-amino-1H-1,2,4-triazole, aromatic aldehydes and unsymmetrical 1,3-diketones. The exact structure of the regioisomer was identified unambiguously by multinuclear 2D-NMR spectroscopy. Furthermore, the binding properties of the synthesized [1,2,4]triazolo[1,5-a]pyrimidine derivatives with BSA were investigated using an in silico molecular docking method and various spectroscopic techniques, namely UV-visible, steady-state fluorescence and competitive displacement assays. These studies are pivotal for the advancement in developing novel pharmaceutical agents and the optimization of therapeutic compounds.
2. Results and discussion
2.1. Chemistry
The synthetic strategy for functionalized [1,2,4]triazolo[1,5-a]pyrimidine derivatives through Biginelli-like condensation between 3-amino-1H-1,2,4-triazole 3, aromatic aldehyde 2 and unsymmetrical 1,3-diketones 1 is shown in Scheme 1. The presence of various nucleophilic sites on 3-amino-1H-1,2,4-triazole (3) and distinct electrophilic centers in unsymmetrical 1,3-diketones (1) could lead to the possibility of at least eight structurally diverse products (4,7-dihydrotriazolo[1,5-a]pyrimidine- and aromatized triazolo[1,5-a]pyrimidine derivatives 5–12). To study the regioselectivity pattern, a model three-component reaction of 3-amino-1H-1,2,4-triazole 3 (1.0 mmol), 1-phenylbutane-1,3-dione 1a (1.0 mmol) and benzaldehyde 2a (1.0 mmol) was initially investigated in a variety of polar protic and aprotic solvents (ethanol, water, tetrahydrofuran, acetonitrile and DMF). The results are depicted in Table 1. The reaction did not proceed at room temperature in water or DMF, even after stirring for 5 h (entries 3 and 8). Under reflux, most of the tested solvents resulted either in complex mixtures or in the formation of a gummy, non-isolable mass (entries 1, 2, 4–6, 9, and 10). Notably, the reaction in acetonitrile in the presence of I2 afforded the product 5a in a modest 30% yield (entry 7). Solvent-free conditions were also examined. No reaction occurred at room temperature (entry 11), while heating to 150 °C for 8 h led to the formation of a gummy mass instead of the desired product (entry 12).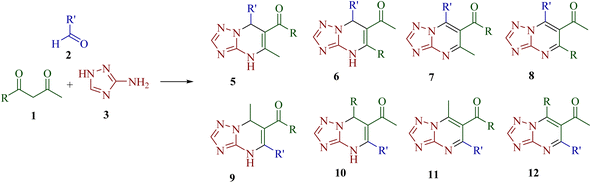 | ||
| Scheme 1 Structure of possible [1,2,4]triazolo[1,5-a]pyrimidine derivatives 5–12 via the condensation of 3-amino-1H-1,2,4-triazole, aldehydes and unsymmetrical 1,3-diketones. | ||
| Entry | Solvent | Temperature | Time | Yieldb (%) |
|---|---|---|---|---|
| a Reagents and conditions: 1-phenylbutane-1,3-dione (1a, 1.0 mmol), benzaldehyde (2a, 1.0 mmol), 3-amino-1H-1,2,4-triazole (3, 1.0 mmol) and an appropriate solvent (10.0 mL). rt = room temperature.b Isolated yield.c Complex mixture obtained.d NR = no reaction. | ||||
| 1 | EtOH | Reflux | 9 h | —c |
| 2 | EtOH | Reflux | 24 h | —c |
| 3 | Water | rt | 5 h | NRd |
| 4 | Water | Reflux | 12 h | Gummy mass |
| 5 | THF | Reflux | 9 h | —c |
| 6 | MeCN | Reflux | 9 h | —c |
| 7 | MeCN-I2 | Reflux | 9 h | 30 |
| 8 | DMF | rt | 5 h | NRd |
| 9 | DMF | Reflux | 8–9 h | Gummy mass |
| 10 | DMF-KOH | Reflux | 8–9 h | Gummy mass |
| 11 | Solvent-free | rt | 5 h | NRd |
| 12 | Solvent-free | 150 °C | 8 h | Gummy mass |
The structure of the isolated product 5a was confirmed using spectroscopic data. The IR spectrum of compound 5a displayed characteristic absorption bands at 3466 cm−1 and 1628 cm−1 corresponding to the amino (NH) group and carbonyl group stretching vibrations, respectively, indicating the product formation. The 1H NMR spectrum of 5a displayed a sharp singlet signal of three protons intensity at δ = 1.82 ppm attributed to the methyl group, three singlet signals of one proton intensity at δ = 6.42, 7.68 and 10.73 ppm corresponding to 2-H, 7-CH and NH proton, respectively, besides signals for the aromatic rings. The 13C NMR spectrum confirmed this, revealing two signals in the aliphatic region at δ = 19.56 and 61.24 ppm and one in the aromatic region at δ = 194.95 ppm assigned to methyl carbon, C-7 and the carbonyl carbon, respectively, along with a set of twelve desired carbon signals in the aromatic region. Furthermore, HRMS (ESI) m/z calculated for C19H16N4O is 316.1324 and was observed at 317.1328 for (M + H)+, confirming the successful condensation of reactants to afford targeted [1,2,4]triazolo[1,5-a]pyrimidine 5a.
To improve the yield of target product, an alternative one-pot step-wise synthetic strategy was explored. In this approach, an equimolar mixture of 1-phenylbutane-1,3-dione 1 and benzaldehyde 2 was stirred in the presence of an acetic acid–piperidine catalytic system to generate the intermediate (Z)-2-benzylidene-1-phenylbutane-1,3-dione 4, followed by the condensation of 4 with 3-amino-1H-1,2,4-triazole 3 in acetic acid under reflux for 6–7 h to afford 5-methyl-7-phenyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidin-6-yl)(phenyl)methanone 5a regioselectively in 80% yield (Scheme 2). The identity of intermediate 4 was confirmed, in the case of 4a, by comparison of its melting point and spectral data with the reported literature data.34
The scope of the described sequential multicomponent reaction was explored by using various aromatic aldehydes and unsymmetrical β-diketones (Table 2). All the substrates having electron-withdrawing or electron-donating groups on the phenyl ring of aromatic benzaldehydes and β-diketones smoothly gave a diverse range of [1,2,4]triazolo[1,5-a]pyrimidine derivatives (5b–u) with high regioselectivity in good yields (78–95%). High yields were obtained when the aldehyde contained an electron-donating substituent (4-OMe) in the aromatic ring in comparison to the electron-withdrawing group (4-NO2).
| S. No. | Diketone (1) | Aldehyde (2) | 3-Amino-1H-1,2,4-triazole | Triazolopyrimidines | Yield% |
|---|---|---|---|---|---|
| R | R′ | (3) | (5) | ||
a Reaction conditions: unsymmetrical β-diketones 1 (1.0 mmol) and aromatic aldehyde 2 (1.0 mmol) were stirred in AcOH/piperidine (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) at 0 °C for 30 min, followed by stirring at room temperature overnight. Subsequently, 3-amino-1H-1,2,4-triazole 3 (1.0 mmol) was added, and the reaction mixture was refluxed in AcOH for 6–7 h.b Isolated yields. 1) at 0 °C for 30 min, followed by stirring at room temperature overnight. Subsequently, 3-amino-1H-1,2,4-triazole 3 (1.0 mmol) was added, and the reaction mixture was refluxed in AcOH for 6–7 h.b Isolated yields. |
|||||
| 1 | Ph | H | 3 | 5a | 80 |
| 2 | 4-FPh | H | 3 | 5b | 82 |
| 3 | 4-ClPh | H | 3 | 5c | 80 |
| 4 | 4-BrPh | H | 3 | 5d | 81 |
| 5 | 4-OMePh | H | 3 | 5e | 86 |
| 6 | 3-OMePh | H | 3 | 5f | 82 |
| 7 | 2-Thienyl | H | 3 | 5g | 79 |
| 8 | Ph | 4-OMe | 3 | 5h | 86 |
| 9 | 4-FPh | 4-OMe | 3 | 5i | 92 |
| 10 | 4-ClPh | 4-OMe | 3 | 5j | 90 |
| 11 | 4-BrPh | 4-OMe | 3 | 5k | 88 |
| 12 | 4-OMePh | 4-OMe | 3 | 5l | 95 |
| 13 | 3-OMePh | 4-OMe | 3 | 5m | 92 |
| 14 | 2-Thienyl | 4-OMe | 3 | 5n | 82 |
| 15 | Ph | 4-NO2 | 3 | 5o | 80 |
| 16 | 4-FPh | 4-NO2 | 3 | 5p | 83 |
| 17 | 4-ClPh | 4-NO2 | 3 | 5q | 82 |
| 18 | 4-BrPh | 4-NO2 | 3 | 5r | 80 |
| 19 | 4-OMePh | 4-NO2 | 3 | 5s | 89 |
| 20 | 3-OMePh | 4-NO2 | 3 | 5t | 85 |
| 21 | 2-Thienyl | 4-NO2 | 3 | 5u | 78 |
The structures of the newly synthesized compounds 5 were characterized using spectroscopic techniques including IR, 1H NMR, 13C NMR, and MS spectrometry, with all spectral data made available in the SI.
To unambiguously determine the correct regioisomeric structure of the isolated product, two-dimensional NMR experiments ([1H–13C] HMBC; [1H–13C] HSQC and [1H–15N] HMBC) were conducted. The 2D NMR correlation results and 1H, 13C and 15N chemical shifts for compound 5l are shown in Fig. 2, S1–S3 and Table 3.
 | ||
| Fig. 2 1H (in red), 13C (in blue) and 15N (in green) NMR spectral characteristics (chemical shifts δ, ppm) and key correlation in the 2D NMR spectra for compound 5l. | ||
| Chemical shifts (δ in ppm) | gs-HSQC correlation | gs-HMBC correlation | Assignments |
|---|---|---|---|
| 193.00 | — | 7.54 (H2″/H6″) | CO |
| 6.35 (7-H) | |||
| 162.22 | — | 7.54 (H2″/H6″) | C4″ |
| 6.96 (H3″/H5″) | |||
| 158.68 | — | 7.09 (H2′/H6′) | C4′ |
| 6.80 (H3′/H5′) | |||
| 149.74 | 7.64 (2-H) | 147.66 (C-3a) | C2 |
| 147.66 | — | 7.64 (2-H) | C-3a |
| 140.57 | — | 1.82 (CH3) | C5 |
| 6.35 (7-H) | |||
| 133.30 | 7.09 (H2′/H6′) | C1′ | |
| 6.80 (H3′/H5′) | |||
| 6.35 (7-H) | |||
| 132.56 | 7.54 (H2″/H6″) | C1″ | |
| 6.96 (H3''/H5″) | |||
| 130.44 | 7.54 (H2''/H6″) | 193.00 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) O) |
C″/C6″ |
| 162.22 (C4″) | |||
| 132.56 (C1″) | |||
| 128.23 | 7.09 (H2′/H6′) | 158.68 (C4′) | (C2′/C6′) |
| 60.35 (C-7) | |||
| 113.77 | 6.96 (H3''/H5″) | 162.22 (C4″) | (C″/C5″) |
| 132.56 (C1″) | |||
| 113.59 | 6.80 (H3′/H5′) | 158.68 (C4′) | (C3′/C5′) |
| 133.30 (C1′) | |||
| 106.96 | 6.35 (7-H) | C6 | |
| 60.35 | 6.35 (C–H) | 133.30 (C1′) | C7 |
| 128.23 (C2′/C6′) | |||
| 140.57 (C5) | |||
| 147.66 (C-3a) | |||
| 193.00 (CO) | |||
| 106.96 (C6) | |||
| 55.31 | 3.80 (4″-OCH3) | 162.22 (C4″) | C4″-OCH3 |
| 54.90 | 3.67 (4′-OCH3) | 158.68 (C4′) | C4′-OCH3 |
| 18.86 | 1.82 (5-CH3) | 140.57 (C5) | C5–CH3 |
| 106.96 (C6) |
The (1H–13C) HMBC spectrum of 5l exhibits cross-peaks of protons of methyl group (δ = 1.82 ppm) with C-5 (δ = 140.57 ppm) and C-6 (δ = 106.96 ppm), confirming the presence of a methyl substituent at the C-5 position of the pyrimidine nucleus. Furthermore, carbonyl carbon at δ = 193.0 ppm displays cross peaks with H2″/H6″ (δ = 7.54 ppm) protons of the aryl ring and C–H at 6.35 ppm (likely H at C-7), indicating the presence of a carbonyl carbon atom with an aryl/heteroaryl ring. Similarly, the (1H–15N) HMBC spectrum also shows a cross peak of methyl proton (δ = 1.82 ppm) with N-4 (δ = −102.27), thus confirming the presence of the methyl substituent at position-5 of [1,2,4]triazolo[1,5-a]pyrimidine. Furthermore, the N8 atom (δ = −182.02 ppm) shows correlation peaks with a proton at the C-7 position (δ = 6.35 ppm) and a methine proton (δ = 7.64) at the C-2 position. Lastly, the N-1 (δ = −209.61 ppm) and N-3 (δ = −275.44 ppm) atoms display cross peaks with methine protons (δ = 7.64) at the C-2 position. The possibility of the structure 6-acetyl-5,7-diphenyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidine 6 can be ruled out, as there is no cross peak of carbonyl carbon to methyl protons in the HMBC spectrum. Thus, from the above-mentioned 2D NMR evidence, the structure of the regioisomer 5l can unequivocally be determined as (4″-methoxyphenyl)(7-(4′-methoxyphenyl)-5-methyl-4,7-dihydro-[1,2,4]triazolo[1,5-a]pyrimidin-6-yl)methanone (Fig. 2).
The plausible mechanism for the regioselective synthesis of [1,2,4]triazolo[1,5-a]pyrimidine derivatives 5 is outlined in Scheme 3. Initially, piperidine acts as a Brønsted base and abstracts a proton from the active methylene group of β-diketone 1, generating a stabilized enolate intermediate.35,36 Concurrently, the aromatic aldehyde 2 is weakly activated under acidic conditions through the protonation of the carbonyl oxygen atom by acetic acid, thereby increasing the electrophilicity of the carbonyl carbon atom. The enolate then nucleophilically attacks the aldehyde 2 to generate β-hydroxy intermediate A that is subsequently dehydrated to furnish the corresponding Knoevenagel adduct 4. Next, the endocyclic nitrogen of 3-amino-1H-1,2,4-triazole 3 undergoes a Michael-type addition onto the activated double bond of 4, leading to intermediate B. The intramolecular nucleophilic attack by the exocyclic amino group of triazole 3 on the more electrophilic and less steric hindered carbonyl carbon atom adjacent to methyl group results in the formation of cyclized intermediate C. Finally, intermediate C undergoes removal of water to furnish 4,7-dihydro[1,2,4]triazolo[1,5-a]pyrimidine 5 as the final product.
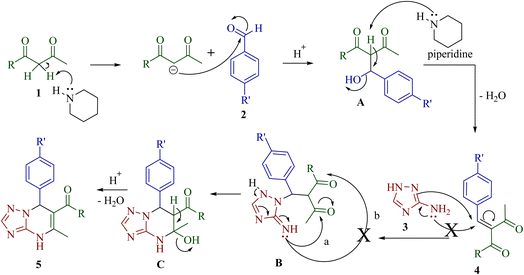 | ||
| Scheme 3 Plausible mechanism of the regioselective synthesis of the 4,7-dihydro[1,2,4]triazolo[1,5-a]pyrimidine derivatives 5. | ||
2.2 Biological studies
The molecular docking simulation study of [1,2,4]triazolo[1,5-a]pyrimidine derivatives (5a–u) with BSA revealed the binding energy scores ranging from −6.0 to −7.9 kcal mol−1, indicating moderate to strong binding interactions with the protein (Table 4). Among these, compounds 5c, 5d, and 5f, each bearing an unsubstituted phenyl ring at the 7-position of the fused heterocyclic system, exhibited the strongest affinity interaction with a binding score of −7.9 kcal mol−1, compared with ibuprofen as reference, −6.6 kcal mol−1. Specifically, compound 5c, with a 4-chlorophenyl substituent, and compound 5d, with a 4-bromophenyl group, demonstrated strong interactions with residues such as Glu125, Lys136, Leu115, Leu122, Pro117, and Phe133, through a combination of hydrogen bonding, alkyl or π-alkyl, π-anion, and hydrophobic interactions (the 2D and 3D plots are reported in Table S1). Compound 5f, containing a 3-methoxyphenyl moiety, showed an equivalent binding score of −7.9 kcal mol−1 and formed a hydrogen bond with Tyr160. Moreover, 5f interacted with key residues including Lys136, Ile181, Pro117, Leu115 (alkyl or π-alkyl) and Glu125 (π-anion), as shown in the 2D and 3D plots (Fig. 3).
| Compounds | Binding energy (kcal mol−1) | Interacted residues |
|---|---|---|
| a Hydrogen bonding.b Alkyl or π-alkyl.c π-sigma.d Amide π-stacking.e π-Anion.f π-Cation.g π-Donor hydrogen bond.h Carbon hydrogen bond.i π–π stacked.j Halogen.k Attractive charge. | ||
| 5a | −7.4 | Glu125a, Lys136b, Leu122b,c, Pro117b, Leu115b, Lys116d |
| 5b | −7.6 | Lys116a, Lys136b, Leu115b, Glu125e |
| 5c | −7.9 | Glu125a, Lys136b, Pro117b, Leu115b, Pro113b, Glu140e, Lys116f, Tyr160g, Phe133h |
| 5d | −7.9 | Lys136b, Leu115b, Pro117b, Leu122b,c, Glu125e, Lys132h |
| 5e | −7.4 | Lys136b, Leu115b, Pro117b, Tyr160g,h, Glu125h, Thr121h, Phe133i |
| 5f | −7.9 | Lys136b, Ile181b, Pro117b, Leu115b,c, Glu125e, Tyr160g,h |
| 5g | −7.1 | Lys116a, Lys136b, Pro117b, Leu115b, Leu122b, Glu125e, Thr121h |
| 5h | −7.4 | Glu125a, Pro113b, Lys136b, Phe133b, Glu140b,e,h, Leu115c, Tyr160g |
| 5i | −7.3 | Glu125a, Pro117b, Leu122b, Pro113b, Leu115c, Lys116f, Tyr160g, Glu140e,h |
| 5j | −6.4 | Asp118a, Lys136b, Leu122b, Glu140e |
| 5k | −6.5 | Lys136b, Pro117b, Leu122b, Leu115b,c, Glu125h, Phe133h,i |
| 5l | −6.4 | Asp118a, Lys136b, Leu115b, Pro113b, Leu122b, Glu140e |
| 5m | −7.0 | Lys116a,f, Lys136b, Phe133b, Pro117b, Leu115b, Glu125e,h |
| 5n | −6.0 | Asp118a, Lys136b,h, Leu122b, Tyr160h |
| 5o | −6.4 | Lys136b, Leu115c, Glu125e, Glu140e, Lys116f |
| 5p | −7.5 | Tyr160a, Lys116a, Lys136b, Leu122b, Leu115c, Glu125e, Glu140e, Phe133i,j |
| 5q | −6.6 | Tyr160a, Lys116a,f, Lys136b, Leu115b,c, Pro113b,h, Glu125e, Glu140e, Phe133h |
| 5r | −7.3 | Glu125a, Pro117b, Leu115b,c, Lys136b,k, Glu140e, Tyr160g, Arg143k, Arg144k |
| 5s | −6.2 | Pro113b, Leu115c, Glu140e, Lys116f, Thr121g,h, Asp118h, Tyr137g |
| 5t | −7.4 | Glu125a, Leu115b,c, Leu122b, Pro117b, Lys136b,k, Glu140e, Lys116f, Tyr160g, Arg143k |
| 5u | −6.5 | Glu125a, Leu115b, Lys136b, Leu122b, Phe133b, Glu140e,k, Lys116f, Tyr160g, Arg143k, Arg144k |
| PBZ | −8.1 | Lys136b, Tyr160b, Arg185b, Tyr137h |
| IBP | −6.6 | Phe133a, Tyr137a, Lys136b, Leu115b, Tyr160g |
 | ||
| Fig. 3 Types of interactions formed between the ligand and the BSA protein: (a and b) 5f, (c and d) 5m and (e and f) 5t, (g and h) phenylbutazone (PBZ) and (i and j) ibuprofen (IBP). | ||
A comparative analysis of compounds 5c, 5d, and 5f with compounds bearing a 4-methoxyphenyl group at the 7-position (5j, 5k and 5m) and those substituted with a 4-nitrophenyl group (5q, 5r and 5t) revealed that compound 5m (bearing a 3-methoxyphenyl group) and compound 5t (also bearing a 3-methoxyphenyl group) exhibited the most favorable docking scores within their respective subsets, with binding energies of −7.0 kcal mol−1 and −7.4 kcal mol−1, respectively (Table 4). Regarding 5m, the pyrimidine-NH demonstrated a hydrogen bond with Lys116, while the triazole core showed a π-cation interaction with Lys116. Alkyl or π-alkyl interaction with residues Lys136, Phe133, Pro117, Leu115 and π-anion and carbon–hydrogen bond with Glu125 also contribute to its enhanced binding affinity. In compound 5t, the pyrimidine-NH showed H-bond with amino acid Glu125. The methyl group and triazole core showed alkyl and π-alkyl interactions with Lys136 and Leu122, respectively. Fig. 3a–j display the lowest energy 2D and 3D poses of the interaction of ligands 5f, 5m, 5t, PBZ and IBP with BSA. Based on the docking results, compounds 5f, 5m, and 5t were selected for further in vitro investigation using spectroscopic techniques to evaluate their binding affinity and conformational interactions with BSA. Their superior binding energies and favorable interaction profiles with crucial BSA residues underscore their potential as effective BSA-binding agents among the studied triazolopyrimidine derivatives.
The synthesized [1,2,4]triazolo[1,5-a]pyrimidine derivatives (5a–u) possess a stereogenic center, and were obtained as racemic mixtures; therefore, the initial molecular docking studies with BSA (PDB ID: 4f5s) were performed using the racemic forms of the ligands. In order to evaluate the influence of stereochemistry on protein–ligand interactions, additional docking studies were carried out for both R- and S-enantiomers of selected representative compounds (5a, 5f, 5m, and 5t) (the 2D and 3D plots are reported in Table S2).
The docking results demonstrated a clear enantioselective preference, with the R-enantiomers exhibiting more favorable binding energies toward BSA than their corresponding S-enantiomers. This difference in binding affinity can be attributed to improved geometric complementarity and more effective interactions of the R-enantiomers within the BSA binding pocket. Notably, the racemic mixtures of these compounds displayed higher binding energies than either individual enantiomer, suggesting a cumulative or averaged interaction profile when both enantiomers are considered together.
These findings indicate that stereochemistry plays a significant role in modulating the binding affinity of the synthesized compounds toward BSA. However, the racemic form, consistent with the experimentally studied samples, provides an appropriate and reliable representation of protein–ligand interactions in the present docking study.
The stoichiometry of the BSA-ligand complex was determined using Job's method of continuous variation. Ten different solutions were prepared with different volumes of BSA and 5t fixed at a constant concentration of 15 µM. In Job's plot, the absorbance at λmax (278 nm) was measured, and absorbance versus mole fraction of ligand 5t was plotted (Fig. 4d). The maximum change in absorbance was observed at 0.5 on the axis at a mole fraction corresponding to a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding ratio, indicating the formation of a 1
1 binding ratio, indicating the formation of a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 BSA-ligand complex 5t.40
1 BSA-ligand complex 5t.40
Fluorescence spectroscopy experiments were conducted with BSA (15 × 10−6 M) and varying concentrations of compounds 5f, 5m, and 5t (0–40 µM). The emission spectra were monitored in the 300–450 nm range after excitation at 280 nm. The results revealed that the fluorescence intensity of BSA around 342 nm decreased progressively with the gradual increase in ligand concentration (Fig. 5a–c) and exhibited a slight blue shift (e.g. 4 nm in the case of 5f). The addition of ligand causes a quenching in the fluorescence spectrum of BSA, primarily arising from the conformational and dynamic changes in the protein structure.
 | ||
| Fig. 5 Emission spectra of BSA (15 µM) in the presence of increasing concentrations (0–40 µM) of compounds (a) 5f, (b) 5m, and (c) 5t. | ||
The extent of quenching depends upon the extent of ligand-BSA interaction and can be determined by calculating the quenching constant Kq using the Stern–Volmer equation41 (eqn 1):
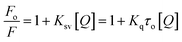 | (1) |
The plot of Fo/F versus 1/[Q] of ligand quencher with BSA revealed linear relationship, as shown in Fig. 6a. The KSV and Kq values of BSA triggered by ligands at room temperature were determined by calculating the slope of curve, and are given in Table 5. The values of quenching constant Kq at 298 K for ligands 5f, 5m and 5t are 7.94 × 1012, 9.31 × 1012 and 12.45 × 1012, respectively. The Kq values (∼1012 M−1 s−1) of ligands are greater than maximum scatter collision quenching constant42(2.0 × 1010 M−1 s−1), indicating the static quenching mechanism involving ground–state complex formation with BSA. According to the Kq values, the ability of the ligand to quench the emission intensity of BSA follows the order of 5t > 5m > 5f, suggesting that the ligand structure significantly influences the binding affinity.
 | ||
| Fig. 6 (a) Stern–Volmer plot of BSA quenching by compounds 5f, 5m and 5t. (b) Double logarithmic plot employed to determine the binding parameters for compounds 5f, 5m and 5t. | ||
| Compd | KSV × 104 (M−1) | Kq × 1012 (M−1 s−1) | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb Kb |
Kb (M−1) | n | ΔG° (kcal mol−1) |
|---|---|---|---|---|---|---|
| 5f | 7.94 ± 18 | 7.94 ± 18 | 4.97 ± 0.13 | 9.5 × 104 | 1.0 ± 0.02 | −6.83 |
| 5m | 9.31 ± 19 | 9.31 ± 19 | 5.90 ± 0.15 | 8.0 × 105 | 1.1 ± 0.02 | −11.02 |
| 5t | 12.45 ± 36 | 12.45 ± 36 | 5.93 ± 0.11 | 8.5 × 105 | 1.1 ± 0.02 | −11.72 |
2.2.3.1. Identification of the binding constant and the number of binding sites. The binding constant Kb and the number of binding sites (n) were calculated using the Scatchard equation43 (eqn 2) as follows:
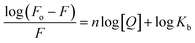 | (2) |
From the plot of log(Fo − F)/F against log[Q] (Fig. 6b), the binding constant Kb and the number of binding sites n were obtained from the intercept and slope, respectively. The calculated results are given in Table 5. The observed n value close to 1 supports the presence of a single binding site on BSA for triazolopyrimidines 5. The binding constant Kb for ligands 5f, 5m, and 5t lies within the favorable range of 104–106 M−1, indicating moderate and reversible interactions with BSA. Such binding strength is considered optimal for carrier proteins, as it allows efficient ligand transport while enabling controlled release at the target site. The results indicate that the compounds bind to BSA in the order of 5t > 5m > 5f. Furthermore, the negative value of standard Gibbs free energy change (ΔG°) from eqn 3 suggests the spontaneity of the binding process that resulted in the BSA-triazolopyrimidine complex:
ΔG° = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb Kb
| (3) |
| System | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb Kb |
Kb (M−1) |
|---|---|---|
| BSA + 5t | 5.9316 ± 0.114 | 8.54 × 105 |
| BSA + PBZ + 5t | 3.94411 ± 0.209 | 0.87 × 104 |
| BSA + IBP + 5t | 4.75972 ± 0.180 | 5.75 × 104 |
In this experiment, the fluorescence spectra of BSA and site marker-BSA (BSA-PBZ) (BSA-IBP) complexes (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) were recorded in the absence and presence of incremental addition of ligand 5t (0–40 µM) (Fig. 7a and b). The fluorescence intensity of the BSA solution decreases upon the addition of the site marker, indicating that the site marker has attached to the BSA molecule. Sequential addition of 5t to the site marker-BSA solution resulted in a gradual decrease in the fluorescence emission intensity, and the data thus obtained were utilized to plot a graph of log(Fo − F)/F against log[Q] for complex 5t in the absence and presence of site markers, as shown in Fig. 7c. The binding constant Kb for phenylbutazone and ibuprofen was found to be 0.87 × 104 and 5.75 × 104, respectively (Table 6). A decrease in the Kb and log
1) were recorded in the absence and presence of incremental addition of ligand 5t (0–40 µM) (Fig. 7a and b). The fluorescence intensity of the BSA solution decreases upon the addition of the site marker, indicating that the site marker has attached to the BSA molecule. Sequential addition of 5t to the site marker-BSA solution resulted in a gradual decrease in the fluorescence emission intensity, and the data thus obtained were utilized to plot a graph of log(Fo − F)/F against log[Q] for complex 5t in the absence and presence of site markers, as shown in Fig. 7c. The binding constant Kb for phenylbutazone and ibuprofen was found to be 0.87 × 104 and 5.75 × 104, respectively (Table 6). A decrease in the Kb and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kb values of the compound 5t with BSA in the presence of phenylbutazone and ibuprofen, compared to the Kb value in the absence of site marker, indicates that the complex and the site marker compete for the same binding site.45 As ligand 5t contains a single binding site, it can interact with either site I or site II on the BSA molecule, but not both simultaneously. Nevertheless, a significant decrease in the Kb value in the presence of phenylbutazone compared to ibuprofen was calculated, suggesting the selective binding of compound 5t to Sudlow's site I of BSA.
Kb values of the compound 5t with BSA in the presence of phenylbutazone and ibuprofen, compared to the Kb value in the absence of site marker, indicates that the complex and the site marker compete for the same binding site.45 As ligand 5t contains a single binding site, it can interact with either site I or site II on the BSA molecule, but not both simultaneously. Nevertheless, a significant decrease in the Kb value in the presence of phenylbutazone compared to ibuprofen was calculated, suggesting the selective binding of compound 5t to Sudlow's site I of BSA.
3. Conclusions
In summary, we have developed an efficient protocol for the regioselective synthesis of a series of 6-aroyl-7-aryl-5-methyl-[1,2,4]triazolo[1,5-a]pyrimidines by three-component condensation of 3-amino-1H-1,2,4-triazole with aromatic aldehydes and β-diketones. Out of the possible regioisomers, the exact one was identified using multinuclear 2D-NMR [1H–13C] HMBC; [1H–13C] HSQC and [1H–15N] HMBC spectroscopic techniques. Molecular docking studies were conducted to assess the interactions of the synthesized compounds with BSA amino acid residues, which indicated that the binding of compounds to BSA is primarily mediated by hydrogen bonding and hydrophobic interactions. Selected compounds (5f, 5m and 5t) were evaluated for their binding interaction with BSA using UV-absorption spectroscopy, fluorescence and competitive displacement assays. Fluorescence quenching studies demonstrated that the compounds interact with BSA via a static quenching mechanism, indicating the formation of a ground-state complex. Among the tested compounds, 5t exhibited the highest binding affinity and most effectively quenched the intrinsic fluorescence of BSA. Furthermore, site-marker competitive binding assay indicated that ligand 5t preferentially binds at Sudlow's site I (in subdomain IIA) of BSA.4. Experimental
4.1. General
All the chemicals and solvents used in the present study were purchased from commercial suppliers of Hi-media and Avera, India, and used without further purification. To monitor the progress of the reaction and purity of the products, TLC experiments were conducted on 0.2 mm Merck precoated silica gel 60 F254-coated aluminum plates. A mixture of ethyl acetate and petroleum ether was used as the mobile phase and spots were visualized under UV light at 254 nm. The melting points were determined using an electrical digital Melting Point Apparatus (MEPA) in an open capillary tube, which were uncorrected. IR spectra were recorded using a Buck Scientific IR M–500 instrument with KBr pellets (υmax in cm−1). 1H and 13C NMR spectra were recorded using a Bruker instrument at frequencies of 400 and 100 MHz, respectively, with DMSO-d6 as a solvent and tetramethylsilane (TMS) as an internal standard (the chemical shift in δ scale and coupling constants (J) were expressed in parts per million (ppm) and hertz, respectively). High-resolution mass spectra (HRMS) were recorded in the ESI+ mode at CIL GJU, Hisar. Furthermore, 2D correlation spectroscopy, (1H–13C) gs-HSQC and (1H–13C) gs-HMBC of the samples were carried out at Kurukshetra University, Kurukshetra. The UV-vis spectra were recorded using a UV-vis spectrophotometer 117 (Systronic, India) with a 1 cm-path-length cell. Fluorescence spectra were recorded using a Shimadzu-5301pc spectro-fluorophotometer (Kyoto, Japan).4.2. General procedure for the synthesis of 6-aroyl-7-aryl-5-methyl-4,7-dihydro- [1,2,4]triazolo[1,5-a]pyrimidines (5a–u)
A mixture of acetic acid-piperidine (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) was taken in a 100 mL round-bottomed flask and cooled in an ice bath. The appropriate diketone 1 (1.0 mmol) and aldehyde 2 (1.0 mmol) were added to the flask and stirred for half an hour under ice-cold conditions and then stirred at room temperature overnight. 3-Amino-1H-1,2,4-triazole 3 (84 mg, 1.0 mmol) and 10 mL acetic acid were added, and the mixture was heated under reflux for 6–7 h. After the completion of the reaction (TLC monitoring), the reaction mixture was allowed to cool to room temperature. The reaction mixture was neutralized with a saturated sodium bicarbonate solution (15 mL) and extracted with EtOAc (3 × 15 mL). The combined organic layers were then dried over anhydrous MgSO4 and evaporated in vacuo. The product obtained was recrystallized from ethanol to give the pure 1,2,4]triazolo[1,5-a]pyrimidines 5a–u.
1) was taken in a 100 mL round-bottomed flask and cooled in an ice bath. The appropriate diketone 1 (1.0 mmol) and aldehyde 2 (1.0 mmol) were added to the flask and stirred for half an hour under ice-cold conditions and then stirred at room temperature overnight. 3-Amino-1H-1,2,4-triazole 3 (84 mg, 1.0 mmol) and 10 mL acetic acid were added, and the mixture was heated under reflux for 6–7 h. After the completion of the reaction (TLC monitoring), the reaction mixture was allowed to cool to room temperature. The reaction mixture was neutralized with a saturated sodium bicarbonate solution (15 mL) and extracted with EtOAc (3 × 15 mL). The combined organic layers were then dried over anhydrous MgSO4 and evaporated in vacuo. The product obtained was recrystallized from ethanol to give the pure 1,2,4]triazolo[1,5-a]pyrimidines 5a–u.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3466 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.73 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.54–7.48 (m, 3H, 3″, 4″, 5″-H); 7.44–7.41 (t, 2H, J = 7.4 Hz, 2″, 6″-H); 7.29–7.26 (t, 2H, J = 7.4 Hz, 2′, 6′-H); 7.23–7.16 (m, 3H, 3′, 4′, 5′-H); 6.42 (s, 1H, C–H); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.95, 150.60, 143.25, 141.74, 141.14, 132.33, 129.17, 128.98, 128.50, 128.40, 128.35, 127.60, 107.73, 61.24, and 19.56; MS (EI) m/z: calcd for C19H16N4O: 316.1324; found: 317.1328 [M + 1]+, elemental analysis: calcd for C19H16N4O: C, 72.13%; H, 5.10%; and N, 17.71%. Found: C, 72.21%; H, 5.28%; and N, 17.59%.
O), 3466 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.73 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.54–7.48 (m, 3H, 3″, 4″, 5″-H); 7.44–7.41 (t, 2H, J = 7.4 Hz, 2″, 6″-H); 7.29–7.26 (t, 2H, J = 7.4 Hz, 2′, 6′-H); 7.23–7.16 (m, 3H, 3′, 4′, 5′-H); 6.42 (s, 1H, C–H); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.95, 150.60, 143.25, 141.74, 141.14, 132.33, 129.17, 128.98, 128.50, 128.40, 128.35, 127.60, 107.73, 61.24, and 19.56; MS (EI) m/z: calcd for C19H16N4O: 316.1324; found: 317.1328 [M + 1]+, elemental analysis: calcd for C19H16N4O: C, 72.13%; H, 5.10%; and N, 17.71%. Found: C, 72.21%; H, 5.28%; and N, 17.59%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3428 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.74 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.60–7.57 (dt, 2H, Jo = 7.2 Hz, J(m)HF = 4.6 Hz, 2″, 6″-H); 7.30–7.20 (m, 5H, 2′, 3′, 4′, 5′, 6′-H); 7.20–7.17 (dd, 2H, Jo = 7.6 Hz, J(m)HF = 4.8 Hz, 3″, 5″-H); 6.41 (s, 1H, C–H); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 192.99, 165.26–162.77 (d, 1JC–F = 249 Hz, Ph-4″-C), 149.95, 147.14, 142.54, 141.04, 136.91, 136.88, 130.74–130.65 (d, 3JC–F = 9 Hz, Ph-2″, 6″-C), 128.37, 127.90, 127.00, and 115.70–115.49 (d, 2JC–F = 21 Hz, Ph-3″, 5″-C), 106.89, 60.67, 18.97; MS (EI) m/z: calcd for C19H15FN4O: 334.1230; found: 335.1239 [M + 1]+, elemental analysis: calcd for C19H15FN4O: C, 68.25%; H, 4.52%; and N, 16.76%. Found: C, 68.13%; H, 4.38%; N, and 16.62%.
O), 3428 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.74 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.60–7.57 (dt, 2H, Jo = 7.2 Hz, J(m)HF = 4.6 Hz, 2″, 6″-H); 7.30–7.20 (m, 5H, 2′, 3′, 4′, 5′, 6′-H); 7.20–7.17 (dd, 2H, Jo = 7.6 Hz, J(m)HF = 4.8 Hz, 3″, 5″-H); 6.41 (s, 1H, C–H); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 192.99, 165.26–162.77 (d, 1JC–F = 249 Hz, Ph-4″-C), 149.95, 147.14, 142.54, 141.04, 136.91, 136.88, 130.74–130.65 (d, 3JC–F = 9 Hz, Ph-2″, 6″-C), 128.37, 127.90, 127.00, and 115.70–115.49 (d, 2JC–F = 21 Hz, Ph-3″, 5″-C), 106.89, 60.67, 18.97; MS (EI) m/z: calcd for C19H15FN4O: 334.1230; found: 335.1239 [M + 1]+, elemental analysis: calcd for C19H15FN4O: C, 68.25%; H, 4.52%; and N, 16.76%. Found: C, 68.13%; H, 4.38%; N, and 16.62%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3428 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.80 (s, 1H, NH); 7.69 (s, 1H, triazole 3-H); 7.53–7.51 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.50–7.48 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 7.29–7.26 (t, 2H, J = 7.2 Hz, 2′, 6′-H); 7.23–7.21 (m, 1H, 4′-H); 7.19–7.17 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.70, 150.60, 147.68, 143.91, 141.68, 139.79, 137.08, 130.37, 129.31, 129.00, 128.53, 127.62, 107.45, 61.18, and 19.75; MS (EI) m/z: calcd for C19H15ClN4O: 350.0934; found: 351.0981 [M + 1]+, 352.1015 [M + 2]+, and elemental analysis: calcd for C19H15ClN4O: C, 65.05%; H, 4.31%; N, 15.97%. Found: C, 65.18%; H, 4.43%; and N, 15.86%.
O), 3428 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.80 (s, 1H, NH); 7.69 (s, 1H, triazole 3-H); 7.53–7.51 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.50–7.48 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 7.29–7.26 (t, 2H, J = 7.2 Hz, 2′, 6′-H); 7.23–7.21 (m, 1H, 4′-H); 7.19–7.17 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.70, 150.60, 147.68, 143.91, 141.68, 139.79, 137.08, 130.37, 129.31, 129.00, 128.53, 127.62, 107.45, 61.18, and 19.75; MS (EI) m/z: calcd for C19H15ClN4O: 350.0934; found: 351.0981 [M + 1]+, 352.1015 [M + 2]+, and elemental analysis: calcd for C19H15ClN4O: C, 65.05%; H, 4.31%; N, 15.97%. Found: C, 65.18%; H, 4.43%; and N, 15.86%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.81 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.64–7.62 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.45–7.43 (d, 2H, J = 8.4 Hz, 3″, 5″-H); 7.29–7.26 (t, 2H, J = 7.2 Hz, 2′, 6′-H); 7.23–7.21 (m, 1H, 4′-H); 7.19–7.17 (t, 2H, J = 7.2 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.80, 150.61, 147.69, 143.99, 141.70, 140.16, 132.23, 130.51, 128.99, 128.52, 127.61, 126.06, 107.43, 61.16, and 19.78; MS (EI) m/z: calcd for C19H15BrN4O: 394.0429; found: 395.0438 [M + 1]+, and elemental analysis: calcd for C19H15BrN4O: C, 57.74%; H, 3.83%; N, 14.17%. Found: C, 57.67%; H, 3.695; and N, 14.39%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.81 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.64–7.62 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.45–7.43 (d, 2H, J = 8.4 Hz, 3″, 5″-H); 7.29–7.26 (t, 2H, J = 7.2 Hz, 2′, 6′-H); 7.23–7.21 (m, 1H, 4′-H); 7.19–7.17 (t, 2H, J = 7.2 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.80, 150.61, 147.69, 143.99, 141.70, 140.16, 132.23, 130.51, 128.99, 128.52, 127.61, 126.06, 107.43, 61.16, and 19.78; MS (EI) m/z: calcd for C19H15BrN4O: 394.0429; found: 395.0438 [M + 1]+, and elemental analysis: calcd for C19H15BrN4O: C, 57.74%; H, 3.83%; N, 14.17%. Found: C, 57.67%; H, 3.695; and N, 14.39%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.54 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.54–7.52 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.28–7.24 (t, 2H, J = 7.6 Hz, 2′, 6′-H); 7.22–7.18 (m, 1H, 4′-H); 7.16–7.14 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 6.96–6.94 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.40 (s, 1H, C–H); 3.80 (s, 3H, 4″-OCH3); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.82, 163.00, 150.52, 148.11, 141.58, 140.48, 132.96, 131.11, 128.98, 128.48, 127.54, 114.44, 107.66, 61.56, 55.96, and 19.21; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1414 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.21%; H, 5.43%; and N, 16.26%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.54 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.54–7.52 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.28–7.24 (t, 2H, J = 7.6 Hz, 2′, 6′-H); 7.22–7.18 (m, 1H, 4′-H); 7.16–7.14 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 6.96–6.94 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.40 (s, 1H, C–H); 3.80 (s, 3H, 4″-OCH3); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.82, 163.00, 150.52, 148.11, 141.58, 140.48, 132.96, 131.11, 128.98, 128.48, 127.54, 114.44, 107.66, 61.56, 55.96, and 19.21; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1414 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.21%; H, 5.43%; and N, 16.26%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.72 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.30–7.26 (t, 2H, J = 7.6 Hz, 2′, 6′-H); 7.24–7.20 (m, 1H, 4′-H); 7.18–7.16 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 7.10–7.07 (dd, 1H, J = 8.4 Hz, J = 2.8 Hz, 6″-H); 7.06–7.04 (d, 1H, J = 8 Hz, 4″-H); 6.94 (br s, 1H, 2″-H); 6.41 (s, 1H, C–H); 3.74 (s, 3H, 3″-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.69, 159.80, 150.59, 147.81, 143.31, 142.68, 142.65, 141.77, 130.36, 128.99, 128.50, 127.56, 120.71, 118.17, 112.87, 107.80, 61.20, 55.78, and 19.44; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1537 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.26%; H, 5.37%; and N, 16.48%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.72 (s, 1H, NH); 7.68 (s, 1H, triazole 3-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.30–7.26 (t, 2H, J = 7.6 Hz, 2′, 6′-H); 7.24–7.20 (m, 1H, 4′-H); 7.18–7.16 (d, 2H, J = 7.6 Hz, 3′, 5′-H); 7.10–7.07 (dd, 1H, J = 8.4 Hz, J = 2.8 Hz, 6″-H); 7.06–7.04 (d, 1H, J = 8 Hz, 4″-H); 6.94 (br s, 1H, 2″-H); 6.41 (s, 1H, C–H); 3.74 (s, 3H, 3″-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.69, 159.80, 150.59, 147.81, 143.31, 142.68, 142.65, 141.77, 130.36, 128.99, 128.50, 127.56, 120.71, 118.17, 112.87, 107.80, 61.20, 55.78, and 19.44; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1537 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.26%; H, 5.37%; and N, 16.48%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.57 (s, 1H, NH); 7.92–7.90 (dd, 1H, J1 = 4.8 Hz, J2 = 1.2 Hz, 5″-H); 7.71–7.70 (dd, 1H, J1 = 4.8 Hz, J2 = 1.2 Hz, 3″-H); 7.66 (s, 1H, triazole 3-H); 7.28–7.24 (m, 2H, 2′,6′-H); 7.22–7.18 (m, 3H, 3′, 4′, 5′-H); 7.15–7.13 (dd, 1H, J1 = 4.8 Hz, J2 = 3.6 Hz, 4″-H); 6.44 (s, 1H, C–H); 1.96 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 186.96, 150.51, 147.98, 145.29, 141.24, 140.15, 134.96, 134.09, 128.98, 128.92, 128.61, 127.75, 107.61, 61.61, and 19.19; MS (EI) m/z: calcd for C17H14N4OS: 322.0888; found: 323.0869 [M + 1]+, and elemental analysis: calcd for C17H14N4OS: C, 63.34%; H, 4.38%; and N, 17.38%. Found: C, 63.23%; H, 4.27%; and N, 17.19%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.57 (s, 1H, NH); 7.92–7.90 (dd, 1H, J1 = 4.8 Hz, J2 = 1.2 Hz, 5″-H); 7.71–7.70 (dd, 1H, J1 = 4.8 Hz, J2 = 1.2 Hz, 3″-H); 7.66 (s, 1H, triazole 3-H); 7.28–7.24 (m, 2H, 2′,6′-H); 7.22–7.18 (m, 3H, 3′, 4′, 5′-H); 7.15–7.13 (dd, 1H, J1 = 4.8 Hz, J2 = 3.6 Hz, 4″-H); 6.44 (s, 1H, C–H); 1.96 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 186.96, 150.51, 147.98, 145.29, 141.24, 140.15, 134.96, 134.09, 128.98, 128.92, 128.61, 127.75, 107.61, 61.61, and 19.19; MS (EI) m/z: calcd for C17H14N4OS: 322.0888; found: 323.0869 [M + 1]+, and elemental analysis: calcd for C17H14N4OS: C, 63.34%; H, 4.38%; and N, 17.38%. Found: C, 63.23%; H, 4.27%; and N, 17.19%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3472 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.69 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.55–7.49 (m, 3H, 3″,4″,5″-H); 7.45–7.42 (m, 2H, 2″, 6″-H); 7.13–7.10 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.38 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.83 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 195.06, 159.36, 150.46, 147.64, 142.95, 141.07, 133.88, 132.35, 129.18, 128.93, 128.42, 128.37, 114.26, 107.86, 60.69, 55.54, and 19.50; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1439 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.29%; H, 5.15%; and N, 16.42%.
O), 3472 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.69 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.55–7.49 (m, 3H, 3″,4″,5″-H); 7.45–7.42 (m, 2H, 2″, 6″-H); 7.13–7.10 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.38 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.83 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 195.06, 159.36, 150.46, 147.64, 142.95, 141.07, 133.88, 132.35, 129.18, 128.93, 128.42, 128.37, 114.26, 107.86, 60.69, 55.54, and 19.50; MS (EI) m/z: calcd for C20H18N4O2: 346.1430; found: 347.1439 [M + 1]+, and elemental analysis: calcd for C20H18N4O2: C, 69.35%; H, 5.24%; and N, 16.17%. Found: C, 69.29%; H, 5.15%; and N, 16.42%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.68 (s, 1H, NH); 7.66 (s, 1H, triazole 3-H); 7.62–7.58 (dt, 2H, J = 7.2 Hz, J = 5.6 Hz, 2″, 6″-H); 7.28–7.24 (t, 2H, J = 8.8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.73, 165.89–163.40 (d, 1JC–F = 249 Hz, Ph-4″-C), 159.39, 150.44, 147.60, 142.87, 137.47, 133.78, 131.39–131.29 (d, 3JC–F = 10 Hz, Ph-2″, 6″-C), 128.95, 116.34–116.12 (d, 2JC–F = 22 Hz, Ph-3″, 5″-C), 114.28, 107.66, 60.75, 55.55, and 19.52; MS (EI) m/z: calcd for C20H17FN4O2: 364.1336; found: 365.1372 [M + 1]+, and elemental analysis: calcd for C20H17FN4O2: C, 65.93%; H, 4.70%; and N, 15.38%. Found: C, 65.66%; H, 4.62%; and N, 15.33%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.68 (s, 1H, NH); 7.66 (s, 1H, triazole 3-H); 7.62–7.58 (dt, 2H, J = 7.2 Hz, J = 5.6 Hz, 2″, 6″-H); 7.28–7.24 (t, 2H, J = 8.8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.73, 165.89–163.40 (d, 1JC–F = 249 Hz, Ph-4″-C), 159.39, 150.44, 147.60, 142.87, 137.47, 133.78, 131.39–131.29 (d, 3JC–F = 10 Hz, Ph-2″, 6″-C), 128.95, 116.34–116.12 (d, 2JC–F = 22 Hz, Ph-3″, 5″-C), 114.28, 107.66, 60.75, 55.55, and 19.52; MS (EI) m/z: calcd for C20H17FN4O2: 364.1336; found: 365.1372 [M + 1]+, and elemental analysis: calcd for C20H17FN4O2: C, 65.93%; H, 4.70%; and N, 15.38%. Found: C, 65.66%; H, 4.62%; and N, 15.33%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.76 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.65–7.63 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.46–7.44 (d, 2H, J = 8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.88, 159.39, 150.49, 147.55, 143.63, 140.12, 133.85, 132.23, 130.52, 128.93, 126.05, 114.27 107.56, 60.62, 55.55, and 19.70; MS (EI) m/z: calcd for C20H17ClN4O2: 380.1040; found: 381.2947 [M + 1]+, 382.2982 [M + 2]+, and elemental analysis: calcd for C20H17FN4O2: C, 63.08%; H, 4.50%; and N, 14.71%. Found: C, 62.97%; H, 4.21%; and N, 14.49%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.76 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.65–7.63 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.46–7.44 (d, 2H, J = 8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.81 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.88, 159.39, 150.49, 147.55, 143.63, 140.12, 133.85, 132.23, 130.52, 128.93, 126.05, 114.27 107.56, 60.62, 55.55, and 19.70; MS (EI) m/z: calcd for C20H17ClN4O2: 380.1040; found: 381.2947 [M + 1]+, 382.2982 [M + 2]+, and elemental analysis: calcd for C20H17FN4O2: C, 63.08%; H, 4.50%; and N, 14.71%. Found: C, 62.97%; H, 4.21%; and N, 14.49%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.75 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.54–7.52 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.50–7.48 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.80 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.39 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.76, 159.39, 150.48, 147.55, 143.58, 139.76, 137.06, 133.85, 130.38, 129.30, 128.94, 114.28 107.58, 60.64, 55.55, and 19.68; MS (EI) m/z: calcd for C20H17BrN4O2: 424.0535; found: 425.0521 [M + 1]+, 426.0512 [M + 2]+, and elemental analysis: calcd for C20H17FN4O2: C, 56.48%; H, 4.03%; and N, 13.17%. Found: C, 56.03%; H, 3.98%; and N, 13.01%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.75 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.54–7.52 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.50–7.48 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 7.12–7.10 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.83–6.80 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.39 (s, 1H, C–H); 3.68 (s, 3H, 4′-OCH3); 1.86 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.76, 159.39, 150.48, 147.55, 143.58, 139.76, 137.06, 133.85, 130.38, 129.30, 128.94, 114.28 107.58, 60.64, 55.55, and 19.68; MS (EI) m/z: calcd for C20H17BrN4O2: 424.0535; found: 425.0521 [M + 1]+, 426.0512 [M + 2]+, and elemental analysis: calcd for C20H17FN4O2: C, 56.48%; H, 4.03%; and N, 13.17%. Found: C, 56.03%; H, 3.98%; and N, 13.01%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3402 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.69 (s, 1H, NH); 7.64 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.11–7.08 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.97–6.95 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.81–6.79 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.80 (s, 3H, 4″-OCH3); 3.67 (s, 3H, 4′-OCH3); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.00, 162.22, 158.68, 149.74, 147.66, 140.57, 133.30, 132.56, 130.44, 128.23, 113.77, 113.59, 106.96, 60.35, 55.31, 54.90, and 18.86; MS (EI) m/z: calcd for C21H20N4O3: 376.1535; found: 377.1422 [M + 1]+, and elemental analysis: calcd for C21H20N4O3: C, 67.01%; H, 5.36%; and N, 14.88%. Found: C, 66.91%; H, 5.08%; and N, 14.53%.
O), 3402 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.69 (s, 1H, NH); 7.64 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.11–7.08 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 6.97–6.95 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.81–6.79 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.35 (s, 1H, C–H); 3.80 (s, 3H, 4″-OCH3); 3.67 (s, 3H, 4′-OCH3); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.00, 162.22, 158.68, 149.74, 147.66, 140.57, 133.30, 132.56, 130.44, 128.23, 113.77, 113.59, 106.96, 60.35, 55.31, 54.90, and 18.86; MS (EI) m/z: calcd for C21H20N4O3: 376.1535; found: 377.1422 [M + 1]+, and elemental analysis: calcd for C21H20N4O3: C, 67.01%; H, 5.36%; and N, 14.88%. Found: C, 66.91%; H, 5.08%; and N, 14.53%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.68 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.12–7.06 (m, 4H, 2′, 6′-H, 2″, 6″-H); 6.96–6.95 (t, 1H, J = 8 Hz, 4″-H); 6.84–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.36 (s, 1H, C–H); 3.75 (s, 3H, 3″-OCH3); 3.68 (s, 3H, 4′-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.73, 159.80, 159.38, 150.48, 147.68, 143.02, 142.64, 133.97, 130.35, 128.89, 120.74, 118.16, 114.27, 112.88, 107.92, 60.65, 55.79, 55.56, and 19.41; MS (EI) m/z: calcd. For C21H20N4O3: 376.1535; found: 377.1637 [M + 1]+, and elemental analysis: calcd for C21H20N4O3: C, 67.01%; H, 5.36%; and N, 14.88%. Found: C, 67.11%; H, 5.29%; and N, 14.76%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.68 (s, 1H, NH); 7.67 (s, 1H, triazole 3-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.12–7.06 (m, 4H, 2′, 6′-H, 2″, 6″-H); 6.96–6.95 (t, 1H, J = 8 Hz, 4″-H); 6.84–6.81 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.36 (s, 1H, C–H); 3.75 (s, 3H, 3″-OCH3); 3.68 (s, 3H, 4′-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.73, 159.80, 159.38, 150.48, 147.68, 143.02, 142.64, 133.97, 130.35, 128.89, 120.74, 118.16, 114.27, 112.88, 107.92, 60.65, 55.79, 55.56, and 19.41; MS (EI) m/z: calcd. For C21H20N4O3: 376.1535; found: 377.1637 [M + 1]+, and elemental analysis: calcd for C21H20N4O3: C, 67.01%; H, 5.36%; and N, 14.88%. Found: C, 67.11%; H, 5.29%; and N, 14.76%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.53 (s, 1H, NH); 7.93–7.91 (dd, 1H, J1 = 4.8 Hz, J1 = 0.8 Hz, 5″-H); 7.74–7.73 (dd, 1H, J1 = 4 Hz, J1 = 0.8 Hz, 3″-H); 7.64 (s, 1H, triazole 3-H); 7.16–7.12 (m, 3H, 4″-H, 2′, 6′-H); 6.82–6.79 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 3.67 (s, 3H, 4′-OCH3); 1.96 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 187.03, 159.45, 150.38, 147.81, 145.34, 139.95, 134.92, 134.07, 133.39, 129.13, 128.94, 114.26, 107.74, 61.05, 55.54, and 19.16; MS (EI) m/z: calcd for C18H16 N4O2S: 352.0994; found: 353.1012 [M + 1]+, and elemental analysis: calcd for C18H16 N4O2S: C, 61.35%; H, 4.58%; and N, 15.90%. Found: C, 61.22%; H, 4.27%; and N, 15.78%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.53 (s, 1H, NH); 7.93–7.91 (dd, 1H, J1 = 4.8 Hz, J1 = 0.8 Hz, 5″-H); 7.74–7.73 (dd, 1H, J1 = 4 Hz, J1 = 0.8 Hz, 3″-H); 7.64 (s, 1H, triazole 3-H); 7.16–7.12 (m, 3H, 4″-H, 2′, 6′-H); 6.82–6.79 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 6.40 (s, 1H, C–H); 3.67 (s, 3H, 4′-OCH3); 1.96 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 187.03, 159.45, 150.38, 147.81, 145.34, 139.95, 134.92, 134.07, 133.39, 129.13, 128.94, 114.26, 107.74, 61.05, 55.54, and 19.16; MS (EI) m/z: calcd for C18H16 N4O2S: 352.0994; found: 353.1012 [M + 1]+, and elemental analysis: calcd for C18H16 N4O2S: C, 61.35%; H, 4.58%; and N, 15.90%. Found: C, 61.22%; H, 4.27%; and N, 15.78%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3450 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.95 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.52–7.50 (m, 3H, 3″, 4″,5″-H); 7.45–7.42 (t, 2H, J = 7.2 Hz, 2″,6″-H); 6.58 (s, 1H, C–H); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.66, 150.97, 148.70, 147.59, 145.05, 141.30, 132.33, 129.22, 129.19, 128.37, 124.24, 106.83, 60.54, and 19.99; MS (EI) m/z: calcd for C19H15N5O3: 361.1175; found: 362.1158 [M + 1]+, and elemental analysis: calcd for C19H15N5O3: C, 63.15%; H, 4.18%; and N, 19.38%. Found: C, 63.29%; H, 4.27%; and N, 19.23%.
O), 3450 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.95 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.52–7.50 (m, 3H, 3″, 4″,5″-H); 7.45–7.42 (t, 2H, J = 7.2 Hz, 2″,6″-H); 6.58 (s, 1H, C–H); 1.82 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.66, 150.97, 148.70, 147.59, 145.05, 141.30, 132.33, 129.22, 129.19, 128.37, 124.24, 106.83, 60.54, and 19.99; MS (EI) m/z: calcd for C19H15N5O3: 361.1175; found: 362.1158 [M + 1]+, and elemental analysis: calcd for C19H15N5O3: C, 63.15%; H, 4.18%; and N, 19.38%. Found: C, 63.29%; H, 4.27%; and N, 19.23%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 3495 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.96 (s, 1H, NH); 8.16–8.14 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.63–7.60 (dt, 2H, J = 7.2 Hz, J = 5.6 Hz, 2″, 6″-H); 7.55–7.53 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 7.28–7.24 (t, 2H, J = 8.8 Hz, 3″, 5″-H); 6.57 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.32, 165.89–163.41 (d, 1JC–F = 248 Hz, Ph-4″-C), 150.96, 148.64, 147.69, 147.61, 144.98, 137.70, 137.67, 131.36–131.27 (d, 3JC–F = 9 Hz, Ph-2″, 6″-C), 129.20, 124.24, 116.37–116.16 (d, 2JC–F = 21 Hz, Ph-3″, 5″-C), 106.63, 60.59, and 20.04; MS (EI) m/z: calcd for C19H14FN5O3: 379.1081; found: 380.1068 [M + 1]+, and elemental analysis: calcd for C19H14FN5O3: C, 60.16%; H, 3.72%; and N, 18.46%. Found: C, 60.27%; H, 3.57%; and N, 18.23%.
O), 3495 (NH); 1H NMR (400 MHz, DMSO-d6) δ: 10.96 (s, 1H, NH); 8.16–8.14 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.63–7.60 (dt, 2H, J = 7.2 Hz, J = 5.6 Hz, 2″, 6″-H); 7.55–7.53 (d, 2H, J = 8.4 Hz, 2′, 6′-H); 7.28–7.24 (t, 2H, J = 8.8 Hz, 3″, 5″-H); 6.57 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.32, 165.89–163.41 (d, 1JC–F = 248 Hz, Ph-4″-C), 150.96, 148.64, 147.69, 147.61, 144.98, 137.70, 137.67, 131.36–131.27 (d, 3JC–F = 9 Hz, Ph-2″, 6″-C), 129.20, 124.24, 116.37–116.16 (d, 2JC–F = 21 Hz, Ph-3″, 5″-C), 106.63, 60.59, and 20.04; MS (EI) m/z: calcd for C19H14FN5O3: 379.1081; found: 380.1068 [M + 1]+, and elemental analysis: calcd for C19H14FN5O3: C, 60.16%; H, 3.72%; and N, 18.46%. Found: C, 60.27%; H, 3.57%; and N, 18.23%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 11.02 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.74 (s, 1H, triazole 3-H); 7.57–7.49 (m, 6H, 2′, 6′, 2″, 6″, 3″, 5″-H); 6.58 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.45, 150.97, 148.66, 147.62, 145.67, 139.95, 137.08, 130.37, 129.35, 129.19, 124.24, 106.60, 60.46, and 20.18; MS (EI) m/z: calcd for C19H14ClN5O3: 395.0785; found: 396.0791 [M + 1]+, and elemental analysis: calcd for C19H14ClN5O3: C, 57.66%; H, 3.57%; and N, 17.69%. Found: C, 57.52%; H, 3.41%; and N, 17.75%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 11.02 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.74 (s, 1H, triazole 3-H); 7.57–7.49 (m, 6H, 2′, 6′, 2″, 6″, 3″, 5″-H); 6.58 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.45, 150.97, 148.66, 147.62, 145.67, 139.95, 137.08, 130.37, 129.35, 129.19, 124.24, 106.60, 60.46, and 20.18; MS (EI) m/z: calcd for C19H14ClN5O3: 395.0785; found: 396.0791 [M + 1]+, and elemental analysis: calcd for C19H14ClN5O3: C, 57.66%; H, 3.57%; and N, 17.69%. Found: C, 57.52%; H, 3.41%; and N, 17.75%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 11.00 (s, 1H, NH); 8.14–8.16 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.65–7.63 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.48–7.45 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.55 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 192.90, 150.35, 148.02, 146.98, 145.07, 139.69, 131.65, 129.88, 128.55, 125.44, 123.61, 105.97, 59.82, and 19.57; MS (EI) m/z: calcd for C19H14BrN5O3: 439.0280; found: 440.0268 [M + 1]+, and elemental analysis: calcd for C19H14BrN5O3: C, 51.84%; H, 3.21%; and N, 15.91%. Found: C, 51.68%; H, 3.34%; and N, 15.77%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 11.00 (s, 1H, NH); 8.14–8.16 (d, 2H, J = 8.8 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.65–7.63 (d, 2H, J = 8.4 Hz, 2″, 6″-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.48–7.45 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.55 (s, 1H, C–H); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 192.90, 150.35, 148.02, 146.98, 145.07, 139.69, 131.65, 129.88, 128.55, 125.44, 123.61, 105.97, 59.82, and 19.57; MS (EI) m/z: calcd for C19H14BrN5O3: 439.0280; found: 440.0268 [M + 1]+, and elemental analysis: calcd for C19H14BrN5O3: C, 51.84%; H, 3.21%; and N, 15.91%. Found: C, 51.68%; H, 3.34%; and N, 15.77%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.76 (s, 1H, NH); 8.16–8.13 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.72 (s, 1H, triazole 3-H); 7.57–7.55 (d, 2H, J = 9.2 Hz, 2′, 6′-H); 7.52–7.50 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.98–6.95 (d, 2H, J = 9.2 Hz, 3′, 5′-H); 6.57 (s, 1H, C–H); 3.81 (s, 3H, 4″-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.53, 163.04, 150.89, 148.53, 148.01, 147.59, 142.31, 133.06, 131.10, 129.14, 124.80, 124.23, 114.49, 106.64, 60.85, 55.96, and 19.64; MS (EI) m/z: calcd for C20H17N5O4: 391.1281; found: 392.1272 [M + 1]+, and elemental analysis: calcd for C20H17N5O4: C, 61.38%; H, 4.38%; and N, 17.89%. Found: C, 61.25%; H, 4.22%; and N, 17.75%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.76 (s, 1H, NH); 8.16–8.13 (d, 2H, J = 8.8 Hz, 2″, 6″-H); 7.72 (s, 1H, triazole 3-H); 7.57–7.55 (d, 2H, J = 9.2 Hz, 2′, 6′-H); 7.52–7.50 (d, 2H, J = 8.8 Hz, 3″, 5″-H); 6.98–6.95 (d, 2H, J = 9.2 Hz, 3′, 5′-H); 6.57 (s, 1H, C–H); 3.81 (s, 3H, 4″-OCH3); 1.84 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 193.53, 163.04, 150.89, 148.53, 148.01, 147.59, 142.31, 133.06, 131.10, 129.14, 124.80, 124.23, 114.49, 106.64, 60.85, 55.96, and 19.64; MS (EI) m/z: calcd for C20H17N5O4: 391.1281; found: 392.1272 [M + 1]+, and elemental analysis: calcd for C20H17N5O4: C, 61.38%; H, 4.38%; and N, 17.89%. Found: C, 61.25%; H, 4.22%; and N, 17.75%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.94 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.08 (m, 2H, 4″, 6″-H); 6.99–6.98 (dd, 1H, J = 2.4 Hz, J = 1.2 Hz, 2″-H); 6.57 (s, 1H, C–H); 3.76 (s, 3H, 3″-OCH3); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.33, 159.85, 150.96, 148.74, 147.69, 147.60, 145.23, 142.80, 130.40, 129.18, 124.22, 120.69, 118.16, 112.86, 106.87, 60.52, 55.82, and 19.88; MS (EI) m/z: calcd for C20H17N5O4: 391.1281; found: 392.1335 [M + 1]+, and elemental analysis: calcd for C20H17N5O4: C, 61.38%; H, 4.38%; and N, 17.89%. Found: C, 61.25%; H, 4.25%; and N, 17.72%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.94 (s, 1H, NH); 8.17–8.15 (d, 2H, J = 8.4 Hz, 3′, 5′-H); 7.73 (s, 1H, triazole 3-H); 7.55–7.53 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.36–7.32 (t, 1H, J = 8 Hz, 5″-H); 7.08 (m, 2H, 4″, 6″-H); 6.99–6.98 (dd, 1H, J = 2.4 Hz, J = 1.2 Hz, 2″-H); 6.57 (s, 1H, C–H); 3.76 (s, 3H, 3″-OCH3); 1.85 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 194.33, 159.85, 150.96, 148.74, 147.69, 147.60, 145.23, 142.80, 130.40, 129.18, 124.22, 120.69, 118.16, 112.86, 106.87, 60.52, 55.82, and 19.88; MS (EI) m/z: calcd for C20H17N5O4: 391.1281; found: 392.1335 [M + 1]+, and elemental analysis: calcd for C20H17N5O4: C, 61.38%; H, 4.38%; and N, 17.89%. Found: C, 61.25%; H, 4.25%; and N, 17.72%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); 1H NMR (400 MHz, DMSO-d6) δ: 10.79 (s, 1H, NH); 8.15–8.13 (d, 2H, J = 9.2 Hz, 3′, 5′-H); 7.94–7.93 (d, 1H, J = 6 Hz, 5″-H); 7.77–7.75 (d, 1H, J = 4.8 Hz, 3″-H); 7.71 (s, 1H, triazole 3-H); 7.53–7.51 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.16–7.14 (t, 1H, J1 = 4.8 Hz, J2 = 4 Hz, 4″-H); 6.61 (s, 1H, C–H); 1.98 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 186.63, 150.89, 148.15, 147.95, 147.69, 145.16, 141.72, 135.17, 134.34, 129.30, 128.99, 124.25, 106.57, 60.83, and 19.56; MS (EI) m/z: calcd for C17H13N5O3S: 367.0739; found: 368.0716 [M + 1]+, and elemental analysis: calcd for C17H13N5O3S: C, 55.58%; H, 3.57%; and N, 19.06%. Found: C, 55.29%; H, 3.41%; and N, 18.99%.
O); 1H NMR (400 MHz, DMSO-d6) δ: 10.79 (s, 1H, NH); 8.15–8.13 (d, 2H, J = 9.2 Hz, 3′, 5′-H); 7.94–7.93 (d, 1H, J = 6 Hz, 5″-H); 7.77–7.75 (d, 1H, J = 4.8 Hz, 3″-H); 7.71 (s, 1H, triazole 3-H); 7.53–7.51 (d, 2H, J = 8.8 Hz, 2′, 6′-H); 7.16–7.14 (t, 1H, J1 = 4.8 Hz, J2 = 4 Hz, 4″-H); 6.61 (s, 1H, C–H); 1.98 (s, 3H, CH3); 13C NMR (100 MHz, DMSO-d6) δ: 186.63, 150.89, 148.15, 147.95, 147.69, 145.16, 141.72, 135.17, 134.34, 129.30, 128.99, 124.25, 106.57, 60.83, and 19.56; MS (EI) m/z: calcd for C17H13N5O3S: 367.0739; found: 368.0716 [M + 1]+, and elemental analysis: calcd for C17H13N5O3S: C, 55.58%; H, 3.57%; and N, 19.06%. Found: C, 55.29%; H, 3.41%; and N, 18.99%.4.3. Binding studies
Fluorescence emission spectroscopy was used for site marker investigation at an excitation wavelength of 280 nm. Phenylbutazone (PBZ) and ibuprofen (IBP) were utilized as particular site-specific probes in this experiment. Solutions of each site marker (15 µM) were prepared separately in PBS. Equimolar BSA was then added to each probe solution at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) ratio, and the mixtures were maintained at 2–6 °C. The BSA + site marker binary solution was subsequently titrated with increasing concentrations of compound 5t (0–40 µM), while keeping the concentrations of BSA + IBP and BSA + PBZ constant.
1 (v/v) ratio, and the mixtures were maintained at 2–6 °C. The BSA + site marker binary solution was subsequently titrated with increasing concentrations of compound 5t (0–40 µM), while keeping the concentrations of BSA + IBP and BSA + PBZ constant.
Author contributions
Ranjana Aggarwal: supervision, writing – review & editing, resources, conceptualization, data curation. Manisha Sharma: writing – original draft, methodology, investigation, data curation. Garima Sumran: writing – original draft, methodology, formal analysis, visualization. Suresh Kumar: investigation, software, visualization. Parvin Kumar: supervision, writing – review & editing, conceptualization.Conflicts of interest
The author(s) confirm that this article's content has no conflicts of interest.Data availability
The data for this article have been included as part of the supplementary information (SI). Supplementary information: experimental data (1H, 13C, [1H–13C] HMBC, [1H–13C] HSQC, [1H–15N] HMBC and HRMS) for final compounds, as well as 2D and 3D diagrams showing the interactions of ligands 5a–u, PBZ and IBP with BSA protein. See DOI: https://doi.org/10.1039/d5ra09673a.Acknowledgements
The authors are thankful to their respective institutions for providing the required facility to complete this work. They thank the Council of Scientific and Industrial Research (CSIR), New Delhi, India, for providing financial assistance to Manisha Sharma (Grant 09/105(0274)/2018-EMR-I) as SRF.References
- S. R. Merugu, S. Cherukupalli and R. Karpoormath, An overview on synthetic and medicinal perspectives of [1,2,4]triazolo[1,5-a]pyrimidine scaffold, Chem. Biodivers., 2022, 19(9), e202200291, DOI:10.1002/cbdv.202200291.
- R. Aggarwal and G. Sumran, An insight on medicinal attributes of 1,2,4-triazoles, Eur. J. Med. Chem., 2020, 205, 112652, DOI:10.1016/j.ejmech.2020.112652.
- K. Oukoloff, B. Lucero, K. R. Francisco, K. R. Brunden and C. Ballatore, 1,2,4-Triazolo[1,5-a]pyrimidines in drug design, Eur. J. Med. Chem., 2019, 165, 332–346, DOI:10.1016/j.ejmech.2019.01.027.
- B. Huang, D. Kang, Y. Tian, D. Daelemans, E. D. Clercq, C. Pannecouque, P. Zhan and X. Liu, Design, synthesis, and biological evaluation of piperidinyl-substituted [1,2,4]triazolo[1,5-a]pyrimidine derivatives as potential anti-HIV-1 agents with reduced cytotoxicity, Chem. Biol. Drug Des., 2021, 97(1), 67–76, DOI:10.1111/cbdd.13760.
- L. H. Al-Wahaibi, S. M. Rabea, M. A. Mahmoud, B. G. M. Youssif, S. Bräse and S. A. Abdel-Aziz, Synthesis and antimicrobial evaluation of new 1,2,4-triazolo[1,5-a]pyrimidine-based derivatives as dual inhibitors of bacterial DNA Gyrase and DHFR, ACS Omega, 2024, 9(47), 47261–47273, DOI:10.1021/acsomega.4c08365.
- R. H. Abd El-Aleam, R. F. George, G. S. Hassan and H. M. Abdel-Rahman, Synthesis of 1,2,4-triazolo[1,5-a]pyrimidine derivatives: Antimicrobial activity, DNA Gyrase inhibition and molecular docking, Bioorg. Chem., 2020, 94, 103411, DOI:10.1016/j.bioorg.2019.103411.
- S. Boutaleb-Charki, C. Marin and C. Maldonado, et al., Copper (II) complexes of [1,2,4]triazolo [1,5-a]pyrimidine derivatives as potential anti-parasitic agents, Drug Metab. Lett., 2009, 3(1), 35–44, DOI:10.2174/187231209787176317.
- J.-L. Huo, S. Wang, X.-H. Yuan, B. Yu, W. Zhao and H.-M. Liu, Discovery of [1,2,4]triazolo[1,5-a]pyrimidines derivatives as potential anticancer agents, Eur. J. Med. Chem., 2021, 211, 113108, DOI:10.1016/j.ejmech.2020.113108.
- T.-Y. Zhang, C.-S. Li, P. Li, X.-Q. Bai, S.-Y. Guo, Y. Jin and S.-J. Piao, Synthesis and evaluation of ursolic acid-based 1,2,4-triazolo[1,5-a]pyrimidines derivatives as anti-inflammatory agents, Mol. Divers., 2022, 26(1), 27–38, DOI:10.1007/s11030-020-10154-7.
- L. Huang, J. Ding and M. Li, et al., Discovery of [1,2,4]-triazolo[1,5-a]pyrimidine-7(4H)-one derivatives as positive modulators of GABAA1 receptor with potent anticonvulsant activity and low toxicity, Eur. J. Med. Chem., 2020, 185, 111824, DOI:10.1016/j.ejmech.2019.111824.
- A. A. Kruzhilin, D. V. Lyapun, D. S. Shevtsov, O. A. Kozaderov, A. Y. U. Potapov, I. D. Zartsyn, C. H. Prabhakar and K. H. S. Shikhaliev, New [1,2,4]triazolo[1,5-a]pyrimidine-7-one corrosion inhibitors for copper in chloride environments, Int. J. Corros. Scale Inhib., 2021, 10(4), 1474–1492, DOI:10.17675/2305-6894-2021-10-4-7.
- G. Tresadern, I. Velter and A. A. Trabanco, et al., [1,2,4]Triazolo[1,5-a]pyrimidine phosphodiesterase 2A inhibitors: Structure and free-energy perturbation-guided exploration, J. Med. Chem., 2020, 63(21), 12887–12910, DOI:10.1021/acs.jmedchem.0c01272.
- L. Zribi, I. Pachòn-Angona and Ò. M. Bautista-Aguilera, et al., Triazolopyridopyrimidine: A new scaffold for dual-target small molecules for Alzheimer's disease therapy, Molecules, 2020, 25(14), 3190, DOI:10.3390/molecules25143190.
- T. Alle, C. Varricchio and Y. Yao, et al., Microtubule-stabilizing 1,2,4-triazolo[1,5-a ]pyrimidines as candidate therapeutics for neurodegenerative disease: Matched molecular pair analyses and computational studies reveal new structure–activity insights, J. Med. Chem., 2023, 66(1), 435–459, DOI:10.1021/acs.jmedchem.2c01411.
- Y. Yuan, E. Takashi, P. Hou, A. Kamijo, D. Miura and H. Ten, Topical skin application of small-molecule antiplatelet agent against pressure injury in rat models, Int. J. Mol. Sci., 2024, 25(3), 1639, DOI:10.3390/ijms25031639.
- M. Zhong, T. Wang and J. Hu, Dissipation kinetics and residues of triazolopyrimidine herbicides flumetsulam and florasulam in corn ecosystem, Environ. Monit. Assess., 2015, 187(6), 390, DOI:10.1007/s10661-015-4593-4.
- M. M. A. El-Gendy, M. Shaaban, K. A. Shaaban, A. M. El-Bondkly and H. Laatsch, Essramycin: A first triazolopyrimidine antibiotic isolated from nature, J. Antibiot., 2008, 61(3), 149–157, DOI:10.1038/ja.2008.124.
- H. Zhang and G. Lin, Microbial proteasomes as drug targets, PLoS Pathog., 2021, 17(12), e1010058, DOI:10.1371/journal.ppat.1010058.
- H. Nada, Y. Choi, S. Kim, K. S. Jeong, N. A. Meanwell and K. Lee, New insights into protein–protein interaction modulators in drug discovery and therapeutic advance, Signal Transduction Targeted Ther., 2024, 9, 341, DOI:10.1038/s41392-024-02036-3.
- M. A. Abdelaziz, M. Shaldam, R. A. El-Domany and F. Belal, Multi-Spectroscopic, thermodynamic and molecular dynamic simulation studies for investigation of interaction of dapagliflozin with bovine serum albumin, Spectrochim. Acta, Part A, 2022, 264, 120298, DOI:10.1016/j.saa.2021.120298.
- D. A. Belinskaia, P. A. Voronina, V. I. Shmurak, R. O. Jenkins and N. V. Goncharov, Serum albumin in health and disease: Esterase, antioxidant, transporting and signaling properties, Int. J. Mol. Sci., 2021, 22(19), 10318, DOI:10.3390/ijms221910318.
- L. Song, X. Tian, Z. Lv, E. Li, J. Wu, Y. Liu, W. Yu and J. Chang, I2/KI-Mediated Oxidative N–N bond formation for the synthesis of 1,5-fused 1,2,4-triazoles from N-aryl amidines, J. Org. Chem., 2015, 80(14), 7219–7225, DOI:10.1021/acs.joc.5b01183.
- A. Hibot, A. Oumessaoud, A. Hafid, M. Khouili and M. D. Pujol, Different synthetic methods for the preparation of triazolopyrimidines and their biological profile, ChemistrySelect, 2023, 8(23), e202301654, DOI:10.1002/slct.202301654.
- L. Souza, J. Santos and M. Mittersteiner, et al., Synthetic versatility of β-alkoxyvinyl trichloromethyl ketones for obtaining [1,2,4]triazolo[1,5-a]pyrimidines, Synthesis, 2018, 50(18), 3686–3695, DOI:10.1055/s-0037-1610191.
- J. Liu, M. Lei and L. Hu, Thiamine hydrochloride (VB1): an efficient promoter for the one-pot synthesis of benzo[4,5]imidazo[1,2-a]pyrimidine and [1,2,4]triazolo[1,5-a]pyrimidine derivatives in water medium, Green Chem., 2012, 14(3), 840, 10.1039/c2gc16499j.
- F. Yang, L.-Z. Yu, P.-C. Diao, X.-E. Jian, M.-F. Zhou, C.-S. Jiang, W.-W. You, W.-F. Ma and P.-L. Zhao, Novel [1,2,4]triazolo[1,5-a]pyrimidine derivatives as potent antitubulin agents: Design, multicomponent synthesis and antiproliferative activities, Bioorg. Chem., 2019, 92, 103260, DOI:10.1016/j.bioorg.2019.103260.
- M. A. A. Radwan, F. M. Alminderej, H. E. M. Tolan and H. M. Awad, One-pot three-component synthesis of new triazolopyrimidine derivatives bearing indole moiety as antiproliferative agents, J. Appl. Pharm. Sci., 2020, 10(9), 12–22, DOI:10.7324/JAPS.2020.10902.
- K. Ablajan, W. Kamil, A. Tuoheti and S. Wan-Fu, An efficient three component one-pot synthesis of 5-amino-7-aryl-7,8-dihydro-[1,2,4]triazolo[4,3-a]-pyrimidine-6-carbonitriles, Molecules, 2012, 17(2), 1860–1869, DOI:10.3390/molecules17021860.
- A. R. Moosavi-Zare, R. Najafi and H. Goudarziafshar, The preparation of [1,2,4]triazolo[1,5-a ]pyrimidines catalyzed by Schiff base zinc(II) complex supported on magnetite nanoparticles under mild conditions, RSC Adv., 2024, 14(27), 19167–19173, 10.1039/D4RA02339K.
- P. Venkatesham, M. Kalonia, A. A. Sangolkar, A. Mudiraj, P. P. Babu and R. R. Vedula, Efficient four-component synthesis of 1,2,4-triazolo[1,5-a]pyrimidines and their in vitro antiproliferative studies, ChemistrySelect, 2023, 8(39), e202302461, DOI:10.1002/slct.202302461.
- A. Y. Hassan, M. T. Sarg, A. H. Bayoumi and M. A. El-Deeb, Synthesis and anticancer evaluation of some novel 5-amino[1,2,4]triazole derivatives, J. Heterocycl. Chem., 2018, 55(6), 1450–1478, DOI:10.1002/jhet.3184.
- R. Aggarwal, S. Kumar, G. Sumran, V. Kumar and R. Sadana, Multicomponent regioselective synthesis of 7-aryl-5-methyl- and 5-aryl-7-trifluoromethyl-2-amino-3-(4′-arylazo)-pyrazolo[1,5-a]pyrimidines and their cytotoxic evaluation, J. Chem. Sci., 2023, 135(4), 102, DOI:10.1007/s12039-023-02219-5.
- R. Aggarwal, M. Sharma, G. Sumran and P. Kumar, Multicomponent catalyst-free regioselective synthesis and binding studies of 3-aroyl-2-methylimidazo[1,2-a ]pyrimidines with BSA using biophysical and computational techniques, RSC Adv., 2025, 15(20), 15999–16014, 10.1039/D5RA01795E.
- Y. Yang, M. Gao, W.-M. Shu, L.-M. Wu, D.-X. Zhang and A.-X. Wu, Synthesis of α-iodoketals from methyl ketones via sustainable and orthogonal tandem catalysis, Org. Biomol. Chem., 2013, 11(7), 1226, 10.1039/c2ob27028e.
- E. V. Dalessandro, H. P. Collin, L. G. L. Guimarães, M. S. Valle and J. R. Pliego, Mechanism of the piperidine-catalyzed Knoevenagel condensation reaction in methanol: The role of iminium and enolate Ions, J. Phys. Chem. B, 2017, 121(20), 5300–5307, DOI:10.1021/acs.jpcb.7b03191.
- I. Tchoffo Tchoupou, M. H. Manyeruke and S. A. Salami, et al., An overview of the synthesis of coumarins via Knoevenagel condensation and their biological properties, Results Chem., 2025, 15, 102181, DOI:10.1016/j.rechem.2025.102181.
- T. Göktürk, E. Sakallı Çetin, T. Hökelek, H. Pekel, O. Sensoy, E. N. Aksu and R. Gup, Synthesis, structural investigations, DNA/BSA interactions, molecular docking studies, and anticancer activity of a new 1,4-disubstituted 1,2,3-triazole derivative, ACS Omega, 2023, 8(35), 31839–31856, DOI:10.1021/acsomega.3c03355.
- R. Patel, N. Maurya, P. M. ud din, N. Farooq, A. Siddique, K. L. Verma and N. Dohare, Esterase activity and conformational changes of bovine serum albumin toward interaction with mephedrone: Spectroscopic and computational studies, J. Mol. Recognit., 2018, 31(11), e2734, DOI:10.1002/jmr.2734.
- A. Anish Babu, K. Karthick, R. Subramanian and K. Swarnalatha, Exploring the structural interaction of BSA with amine functionalized ruthenium(II) metal complex, J. Biomol. Struct. Dyn., 2020, 38(13), 4032–4039, DOI:10.1080/07391102.2019.1683071.
- B. Pastrello, G. C. dos Santos, L. C. D. A. Silva-Filho, A. R. de Souza, N. H. Morgon and V. F. Ximenes, Novel aminoquinoline-based solvatochromic fluorescence probe: Interaction with albumin, lysozyme and characterization of amyloid fibrils, Dyes Pigments, 2020, 173, 107874, DOI:10.1016/j.dyepig.2019.107874.
- M. H. Gehlen, The centenary of the Stern-Volmer equation of fluorescence quenching: From the single line plot to the SV quenching map, J. Photochem. Photobiol. C Photochem. Rev., 2020, 42, 100338, DOI:10.1016/j.jphotochemrev.2019.100338.
- G. Rabbani, M. H. Baig, E. J. Lee, W.-K. Cho, J. Y. Ma and I. Choi, Biophysical study on the interaction between eperisone hydrochloride and human serum albumin using spectroscopic, calorimetric, and molecular docking analyses, Mol. Pharm., 2017, 14(5), 1656–1665, DOI:10.1021/acs.molpharmaceut.6b01124.
- K. Paliwal, P. Haldar, P. K. S. Antharjanam and M. Kumar, Mixed ligand mononuclear copper(II) complex as a promising anticancer agent: Interaction studies with DNA/HSA, molecular docking, and in vitro cytotoxicity studies, ACS Omega, 2022, 7(25), 21961–21977, DOI:10.1021/acsomega.2c02354.
- S. Perontsis, A. G. Hatzidimitriou and G. Psomas, Coordination compounds of cobalt(II) with carboxylate non-steroidal anti-inflammatory drugs: structure and biological profile, Dalton Trans., 2024, 53(36), 15215–15235, 10.1039/D4DT01846J.
- R. Ma, H. Pan, T. Shen, P. Li, Y. Chen, Z. Li, X. Di and S. Wang, Interaction of flavonoids from Woodwardia unigemmata with bovine serum albumin (BSA): Application of spectroscopic techniques and molecular modeling methods, Molecules, 2017, 22(8), 1317, DOI:10.3390/molecules22081317.
| This journal is © The Royal Society of Chemistry 2026 |