Open Access Article
Open Access ArticleSynergistic modulation of ABC transporter-mediated multidrug resistance by Kalanchoe laciniata (L.) DC. phytochemicals: integrative phytochemical profiling, network pharmacology, and molecular docking insights
Anish Ruban S.,
Francis Jegan Raj,
Vetri Velavan Sundararajan and
Parimelazhagan Thangaraj *
*
Bioprospecting Laboratory, Department of Botany, Bharathiar University, Coimbatore 641046, Tamil Nadu, India. E-mail: drparimel@gmail.com
First published on 13th February 2026
Abstract
Kalanchoe laciniata (L.) DC. is an underexplored medicinal plant with promising anticancer potential. This study evaluated the phytochemical profile, antioxidant activity, and multidrug resistance (MDR) modulating effects of its methanolic extract (KLM), with emphasis on ABC transporter regulation. Soxhlet methanol extraction yielded the highest phenolic and tannin contents and exhibited strong antioxidant activity. GC-MS and LC-MS analyses identified bioactive flavonoids and phytosterols, including quercetin and kaempferol derivatives. Network pharmacology revealed key interactions with ABC transporters and hub genes such as AKT1 and TP53. Molecular docking and dynamics simulations demonstrated stable binding of KLM-derived flavonoids with BCRP, MRP1, and AKT1, supporting their role in MDR modulation. Functionally, KLM exhibited dose-dependent cytotoxicity against MDA-MB-231 breast cancer cells and showed strong synergy with doxorubicin. qPCR analysis further confirmed significant downregulation of AKT1, BCRP, and MRP1 expression following KLM treatment. Collectively, these findings indicate that K. laciniata phytochemicals may act as effective chemosensitizing agents to overcome ABC transporter-mediated drug resistance in breast cancer.
Introduction
The genus Kalanchoe is a hidden repository of underutilized plants with multiple useful secondary metabolites with significant antioxidant potential. Kalanchoe laciniata, commonly referred to as the “Christmas Tree Plant” is a succulent perennial plant belonging to the family Crassulaceae. Known for its strikingly lobed leaves and yellow to orange flowers, K. laciniata has gained recognition not only as an ornamental plant but also for its traditional medicinal uses in various cultures from Brazil and Madagascar to parts of Asia.1 This genus is widely employed in ethnomedicine, where it is used to treat ailments such as inflammation, wounds, coughs, and fever.2 Its applications in folk medicine have spurred interest in scientific research, leading to investigations into its phytochemical composition and pharmacological potential.3Plant-derived natural compounds have long served as a vital source of therapeutic agents, especially in cancer treatment. These phytochemicals including alkaloids, flavonoids, terpenoids, phenolic compounds, and saponins exhibit a widespread range of biological activities, such as antioxidant, anti-inflammatory, and cytotoxic effects. Several of the most effective anticancer drugs, such as paclitaxel (Taxol) and vincristine, originate from plants, underscoring their potential to provide novel mechanisms for targeting cancer cells.4,5
In cancer therapy, either inherent or acquired drug resistance remains a significant challenge. Cancer cells often develop resistance to conventional treatments, such as chemotherapy and targeted therapies, through mechanisms like increased drug efflux, enhanced DNA repair, and evasion of apoptosis.6,7 This resistance limits treatment efficacy and contributes to disease progression. Plant-derived compounds, with their structural diversity and ability to modulate multiple biological pathways, are emerging as powerful tools in addressing drug resistance in cancer.8 Drug resistance in cancer is mediated through several mechanisms, one of the most prominent being the overexpression of ATP-binding cassette (ABC) transporters, such as BCRP and MRP1, which actively pump therapeutic drugs out of cancer cells.9 Phytochemicals like flavonoids and alkaloids have been shown to inhibit these pumps, increasing the intracellular concentration of chemotherapeutic agents.10 Although drug efflux pumps are only one way through which drug resistance is incurred in plants it plays an important role in the occurrence of drug resistance in cancer cells. In most resistant cells drug efflux transporters are always overexpressed, leading to the efflux of chemotherapeutic drugs.11
Although several Kalanchoe species, including K. pinnata and K. daigremontiana, have been reported to exhibit antioxidant, cytotoxic, and pro-apoptotic activities, these studies are largely phenomenological and rarely extend to pathway-level or target-specific mechanistic validation.12–14 In the context of multidrug resistance (MDR), flavonoids such as quercetin and kaempferol are well known to modulate ABC transporters, particularly P-glycoprotein (ABCB1), MRP1 (ABCC1), and BCRP (ABCG2), through direct binding, ATPase modulation, or transcriptional regulation; however, this evidence predominantly originates from isolated compounds or unrelated botanical sources rather than from other Kalanchoe species.15–17 Importantly, for Kalanchoe laciniata, neither the phytochemical landscape nor the systems-level mechanistic links between its endogenous metabolites and MDR-associated targets have been comprehensively investigated. The present study addresses this gap by employing an integrative, multi-scale strategy that combines phytochemical profiling with network pharmacology, molecular docking, and molecular dynamics simulations to systematically interrogate how K. laciniata derived compounds interact with key ABC transporters and cancer-associated proteins. By contextualizing known flavonoid–transporter interactions within the species-specific and combinatorial synergism of K. laciniata, this work advances beyond activity-based observations and provides mechanistic evidence for the potential modulation of ABC transporter mediated drug efflux and interconnected MDR pathways, thereby establishing a distinct mechanistic framework for this underexplored species.
Materials and methods
Chemical and reagents
Methanol (67-56-1), petroleum ether (8032-32-4), and ethyl acetate (141-78-6) were procured from Merck (Darmstadt, Germany). Dimethyl sulfoxide (DMSO; 67-68-5), Folin–Ciocalteu's reagent (8024-55-5), DPPH (2,2-diphenyl-1-picrylhydrazyl; 1898-66-4), aluminum chloride (7446-70-0), sodium carbonate (497-19-8), potassium acetate (127-08-2), and all positive controls were obtained from Sigma-Aldrich Chemical Co. (St. Louis, MO, USA).Sample collection
Kalanchoe laciniata plant samples were gathered from the Western Ghats in the Theni district. The collected plants were thoroughly cleaned, shade-dried separately, and then ground. The sample was authenticated and certified by the Botanical Survey of India, Southern Regional Centre, Coimbatore (BSI/SRC/5/23/2023-24/Tech/58). The plant material was shade-dried for six weeks, after which it was ground into a fine powder. The literature study was conducted for different parts of Kalanchoe laciniata and the entire plant part was chosen for its maximum antioxidant and phytochemical potential.Phytochemical extraction
Plant material (50 g) was extracted using petroleum ether, ethyl acetate, methanol, and water through two methods: cold maceration and Soxhlet extraction. For cold maceration, the powdered sample was soaked in 300 mL of each solvent for 48 hours at room temperature, after which the extracts were filtered and concentrated under reduced pressure at 50 °C. For Soxhlet extraction, 50 g of plant material was continuously extracted with 300 mL of each solvent at their respective boiling points for 4–8 hours to ensure efficient recovery of phytochemicals. All extracts were concentrated under vacuum and stored for further analysis.18,19Secondary metabolite quantification
The total phenolic content (TPC), total flavonoid content (TFC), and total tannin content (TTC) of the plant extracts were quantified using five different solvent extracts. TPC and TTC were determined using the Folin–Ciocalteu method. For TPC, extract solutions (1 mg per mL stock) were reacted with the reagent, and absorbance was recorded at 725 nm, with results expressed as mg gallic acid equivalents (GAE)/g dry weight. TTC was measured similarly, with tannins removed using PVPP to obtain non-tannin phenolics. Tannin content was calculated as:| Tannins (g) = total phenolics (g) − non-tannin phenolics (g). |
TFC was assessed using the aluminum chloride colorimetric method, with absorbance measured at 510 nm and results expressed as mg rutin equivalents (RUE)/g dry weight.20,21
In vitro antioxidant assays
The antioxidant potential of the plant extracts was evaluated using multiple in vitro assays. DPPH radical scavenging activity was determined by reacting various extract concentrations (final volume 100 µL) with DPPH solution, using BHT and rutin as positive controls. The decrease in absorbance at 517 nm indicated radical scavenging, and IC50 values were calculated.22 Ferric Reducing Antioxidant Power (FRAP) was measured by the reduction of the Fe3+–TPTZ complex to Fe2+, forming a violet chromophore with absorbance recorded at 593 nm, expressed as Fe(II) equivalents.23For phosphomolybdenum activity, extracts reduced Mo(VI) to Mo(V), generating a green complex measured at 695 nm, with results expressed as ascorbic acid equivalents (AAE).24 ABTS scavenging activity was quantified by reacting extracts with ABTS+ solutions for 30 minutes and measuring absorbance at 734 nm. Antioxidant capacity was expressed as Trolox equivalence, and IC50 values were calculated.25 Superoxide radical scavenging activity was assessed using the riboflavin-light-NBT system, where inhibition of formazan formation was measured at 590 nm and expressed as percentage inhibition compared to control.26
Characterization of secondary metabolites
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50) and separated on an Agilent DB-35 MS column using helium (1 mL min−1). Mass spectra were acquired from 50–650 Da at −70 eV. Compounds were identified by comparison with NIST4 and WILEY9 libraries, considering matches with spectral fit values > 700.27
50) and separated on an Agilent DB-35 MS column using helium (1 mL min−1). Mass spectra were acquired from 50–650 Da at −70 eV. Compounds were identified by comparison with NIST4 and WILEY9 libraries, considering matches with spectral fit values > 700.27Network pharmocology
Receptor grids were then generated for each target using the Receptor Grid Generation tool, with the active site defined based on the co-crystallized ligand or known binding region. Grid parameters were optimized through the receptor, site, constraints, and rotatable groups tabs to ensure accurate docking. Prepared ligands were subsequently docked into the generated grids using glide in extra precision (XP) mode. Docking scores and binding interactions were analyzed, and protein–ligand complexes were visualized using both 2D and 3D interaction diagrams.33
Maestro was used to prepare protein–ligand complexes and refine side-chain residues before MD simulation (Schrödinger, LLC, 2019-2). Desmond added missing atoms and applied the TIP3P water model for solvation. An orthorhombic box (10 × 10 × 10 Å) was created around the complex, and Na+ and Cl− ions were added to neutralize the system. To mimic physiological conditions, 0.15 M NaCl was included during a 100 ns NPT simulation at 310 K and 1 bar, with data recorded every 10 ps. Systems were energy-minimized using Desmond's relaxation module (1 kcal per mol convergence, 2000 iterations) and pre-equilibrated before simulation. The OPLS_2005 force field and SPC solvent model were employed throughout.35
Cytotoxicity of KLM was assessed using the MTT assay after 24 h treatment. RIN5F cells were exposed to 100–600 µg mL−1, and MDA-MB-231 cells to 10–100 µg per mL extract. Doxorubicin (1.5 µM) served as the positive control. The MTT assay was performed following a reagent-based colorimetric protocol adapted from previously published literature,36 which has been routinely used to assess cytotoxic effects of bioactive compounds in cultured cells. Following treatment, MTT solution (5 mg mL−1; 20 µL per well) was added to the cultures and incubated at 37 °C for 4 h to allow for formazan crystal formation. The resulting insoluble formazan was subsequently solubilized using DMSO, and absorbance was recorded at 570 nm. Cell viability was calculated and expressed as a percentage relative to untreated control cells.
For cell death visualization, propidium iodide (PI) staining was performed under identical treatment conditions. Following exposure, cells were washed with phosphate-buffered saline (PBS), incubated with PI solution (10 µg mL−1) for 5–10 min at room temperature in the dark, and immediately visualized under an inverted EVOS fl AMG. In addition, bright-field images of RIN5F cells were acquired using the same microscope. The same conditions were followed for RNA extraction from extract-treated cells and further used for RT-qPCR analysis.
Cells were treated with various concentrations of K. laciniata extract and DOX individually, as well as in fixed-ratio combinations derived from their respective IC50 values. After 24 hours of exposure, cell viability was assessed using the MTT assay as described above, and the percentage inhibition values were used for CI calculation.
The combination index (CI) was calculated using the equation:
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g for 15 min at 4 °C. The aqueous phase was transferred to a fresh tube, and RNA was precipitated with isopropanol, followed by centrifugation at 12
000 × g for 15 min at 4 °C. The aqueous phase was transferred to a fresh tube, and RNA was precipitated with isopropanol, followed by centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g for 10 min at 4 °C. The RNA pellet was washed with 75% ethanol, air-dried, dissolved in RNase-free water, and quantified using a NanoDrop spectrophotometer. RNA samples were stored at −80 °C until use. cDNA was synthesized from 1 µg total RNA using the Bio-Rad iScript™ cDNA Synthesis Kit in a 20 µL reaction volume following the manufacturer's instructions. Reverse transcription was performed at 25 °C for 5 min, 46 °C for 20 min, and 95 °C for 1 min. The resulting cDNA was stored at 20 °C. Quantitative real-time PCR was performed using iTaq™ Universal SYBR® Green Supermix (Bio-Rad, USA) on a Bio-Rad CFX Opus system. Each 20 µL reaction contained SYBR Green master mix, gene-specific primers, and diluted cDNA. The primers used were β-actin – forward: 5′-AGAGCTACGAGCTGCCTGAC-3′, reverse: 5′-AGCACTGTGTTGGCGTACAG-3′; AKT1 – forward: 5′-GCTGCTGCTGAGGAGAAACT-3′, reverse: 5′-TGAGGACAGGATGAGGAGGA-3′; BCRP – forward: 5′-CAGTGGAGAGGAGGACATCA-3′, reverse: 5′-TCTCCTGCTGCTGATGGTAA-3′; PGP – forward: 5′-ATGGTGCTGGAAGGAGACAA-3′, reverse: 5′-CTCAGGGTGGTGATGGTGTA-3′; and MRP1 forward: 5′-TGCTGGAGCTGTTTGATGAA-3′, reverse: 5′-TTTGGTGGTGGTGTCTGTCT-3′. Reactions were run in duplicate under the following conditions: 95 °C for 3 min, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Melt curve analysis was performed to confirm amplification specificity. Relative gene expression was normalized to β-actin and calculated using the 2−ΔΔCt method, with results expressed as fold change relative to the control.36,38
000 × g for 10 min at 4 °C. The RNA pellet was washed with 75% ethanol, air-dried, dissolved in RNase-free water, and quantified using a NanoDrop spectrophotometer. RNA samples were stored at −80 °C until use. cDNA was synthesized from 1 µg total RNA using the Bio-Rad iScript™ cDNA Synthesis Kit in a 20 µL reaction volume following the manufacturer's instructions. Reverse transcription was performed at 25 °C for 5 min, 46 °C for 20 min, and 95 °C for 1 min. The resulting cDNA was stored at 20 °C. Quantitative real-time PCR was performed using iTaq™ Universal SYBR® Green Supermix (Bio-Rad, USA) on a Bio-Rad CFX Opus system. Each 20 µL reaction contained SYBR Green master mix, gene-specific primers, and diluted cDNA. The primers used were β-actin – forward: 5′-AGAGCTACGAGCTGCCTGAC-3′, reverse: 5′-AGCACTGTGTTGGCGTACAG-3′; AKT1 – forward: 5′-GCTGCTGCTGAGGAGAAACT-3′, reverse: 5′-TGAGGACAGGATGAGGAGGA-3′; BCRP – forward: 5′-CAGTGGAGAGGAGGACATCA-3′, reverse: 5′-TCTCCTGCTGCTGATGGTAA-3′; PGP – forward: 5′-ATGGTGCTGGAAGGAGACAA-3′, reverse: 5′-CTCAGGGTGGTGATGGTGTA-3′; and MRP1 forward: 5′-TGCTGGAGCTGTTTGATGAA-3′, reverse: 5′-TTTGGTGGTGGTGTCTGTCT-3′. Reactions were run in duplicate under the following conditions: 95 °C for 3 min, followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s. Melt curve analysis was performed to confirm amplification specificity. Relative gene expression was normalized to β-actin and calculated using the 2−ΔΔCt method, with results expressed as fold change relative to the control.36,38Results
Extraction and quantification of secondary metabolites
The powdered plant (50 g) was extracted using four different solvents using two different extraction methods, Soxhlet and cold maceration. Non-polar to polar solvents were used (petroleum ether, ethyl acetate, methanol, and water) (300 mL). The extract was concentrated to dryness under reduced pressure in a rotary evaporator to yield dried extracts. Further, yield extract was calculated and the dried extracts were used for the follow-up studies.Secondary metabolites, including TPC, TFC, and TTC contents of Kalanchoe laciniata extracts, were analyzed. The total phenolic content, assessed using the Folin–Ciocalteu method and expressed as gallic acid equivalents (GAE), was highest in the Soxhlet methanol extract (278.98 ± 0.65a mg GAE per g extract). Similarly, the total flavonoid content, determined using the aluminum chloride method and expressed as rutin equivalents (RE), peaked in the macerated methanol extract (371.32 ± 45.73a mg RE/g extract). Total tannin content, calculated by subtracting non-tannin phenolics from the total phenolics and expressed as GAE, was also highest in the Soxhlet methanol extract (55.39 ± 1.46a mg GAE/g extract). These findings suggest that Soxhlet extraction is the most effective method, and methanol is the most suitable solvent for isolating secondary metabolites from K. laciniata Table 1.
| Method used | Solvent used | Total phenolic content (GAE/g extract) | Total flavonoid content (RE/g extract) | Total tannin content (GAE/g extract) |
|---|---|---|---|---|
| Different letters within one column indicate a significant difference between treatments n = 3, (a < b < c < d < e ≤ 0.05) on DMRT analysis. Values consisted of mean ± SD. | ||||
| Soxhlet | Petroleum ether | 13.54 ± 1.57e | 134.91 ± 10.43c | 3.54 ± 0.51d |
| Ethyl acetate | 81.91 ± 1.94c | 258.60 ± 13.26b | 9.34 ± 2.97b | |
| Methanol | 278.9815 ±0.65a | 182.28 ± 5.41c | 55.39 ±1.46a | |
| Water | 46.89 ± 5.40d | 109.91 ± 0.62c | 10.92 ± 1.71b | |
| Macerated | Petroleum ether | 10.31 ± 3.21e | 258.16 ± 67.68b | 1.53 ± 0.19c |
| Ethyl acetate | 18.64 ± 1.90e | 371.32 ±45.73a | 6.23 ± 2.37c | |
| Methanol | 244.41 ± 31.03b | 181.40 ± 30.20c | 51.97 ± 1.02a | |
| Water | 55.99 ± 3.08d | 109.91 ± 1.64c | 12.89 ± 0.25b | |
Characterization of phytocompounds
| S. no. | Retention time | CAS no. | Name | Molecular formula | Molecular weight | Probability% | Peak area |
|---|---|---|---|---|---|---|---|
| 1 | 7.14 | 150-86-7 | Phytol | C20H40O | 296 | 35.45 | 4.54 |
| 2 | 9.03 | 112-72-1 | 1-Tetradecanol | C14H30O | 214 | 19.64 | 0.67 |
| 3 | 10.98 | 2462-85-3 | 9,12-Octadecadienoic acid, methyl ester | C19H34O2 | 294.5 | 79.45 | 27.67 |
| 4 | 12.66 | 519-96-0 | Patuletin | C16H12O | 332 | 17.37 | 0.53 |
| 5 | 14.05 | 88-32-4 | 3-tert-Butyl-4-methoxyphenol | C11H16O2 | 180 | 36.78 | 4.91 |
| 6 | 14.61 | 818-23-5 | 8-Pentadecanone | C15H30O | 226.39 | 65.57 | 19.15 |
| 7 | 15.53 | 628-97-7 | Hexadecanoic acid, ethyl ester | C18H36O2 | 284.47 | 31.46 | 1.89 |
| 8 | 15.96 | 83-48-7 | Stigmasterol | C29H48O | 412.69 | 23.41 | 2.05 |
| 9 | 17.24 | 2363-71-5 | Heneicosanoic acid | C21H42O2 | 326.55 | 40.15 | 5.31 |
| 10 | 19.5 | 7132-64-1 | Pentadecanoic acid methyl ester | C16H32O2 | 256 | 46.78 | 6.56 |
| 11 | 22.09 | 629-94-7 | Heneicosane | C21H44 | 296 | 27.48 | 0.71 |
| 12 | 24.22 | 57-10-3 | Palmitic acid | C16H32O2 | 256 | 37.69 | 3.84 |
| 13 | 24.98 | 112-61-8 | Stearic acid methyl ester | C19H38O2 | 298 | 56.74 | 14.89 |
| 14 | 27.11 | 6068-80-0 | Quercetin 7,3′,4′-trimethyl ether | C18H16O7 | 344 | 17.84 | 0.59 |
| 15 | 29.68 | 331-39-5 | Caffeic acid | C9H8O4 | 180 | 11.65 | 0.08 |
| 16 | 32.02 | 111-02-4 | Squalene | C30H50 | 410 | 20.47 | 0.31 |
| 17 | 35.28 | 2631-37-0 | Phenol, 3-methyl-5-(1-methylethyl)-, methylcarbamate | C12H17NO2 | 207 | 16.78 | 0.09 |
| 18 | 37.16 | 459-80-3 | Geranic acid | C10H16O2 | 168 | 24.74 | 0.21 |
| 19 | 37.93 | 124-07-2 | Octanoic acid | C8H16O2 | 144 | 39.74 | 2.87 |
| 20 | 38.55 | 82304-66-3 | 7,9-Di-tert-butyl-1-oxaspiro(4.5)deca-6,9-diene-2,8-dione | C17H24O3 | 276 | 19.43 | 1.93 |
| 21 | 40.08 | 28564-83-2 | 2,3-Dihydro-3,5-dihydroxy-6-methyl-4H-pyran-4-one | C6H8O4 | 144.12 | 19.61 | 0.53 |
| 22 | 41.23 | 33983-40-3 | (+)-Medicarpin | C16H14O4 | 270.28 | 22.78 | 0.61 |
Among the identified compounds, octadecadienoic acid, known for its role as a fatty acid with potential health benefits, exhibited the highest peak area percentage of 27.67% at a retention time of 10.98. Additionally, stearic acid, a saturated fatty acid recognized for its moisturizing properties, and pentadecanone, a ketone that may possess antimicrobial activity, were also prominent in the extract. Furthermore, the analysis revealed the presence of quercetin derivatives, which are known for their antioxidant properties, as well as various phytol groups and other phenolic compounds that contribute to the plant's bioactivity.
The compounds detected in the GC-MS analysis are associated with significant biological and medicinal properties, although many of their specific activities remain to be fully explored. This analysis underscores the potential therapeutic applications of K. laciniata based on its rich profile of biactive compounds.
| S. no. | Pubchem ID | Molecular formula | Molecular weight | Compound name |
|---|---|---|---|---|
| 1 | 49798774 | C30H42O10 | 562.6 | Kalanchoside C |
| 2 | 90658606 | C27H29O16− | 609.5 | Kaempferol 3-O-(2′-O-D-glucopyranosyl)-beta-D-galactopyranoside |
| 3 | 74124649 | C27H30O15 | 594.5 | Kaempferol-3-O-glucoside-3″-rhamnoside |
| 4 | 44258959 | C25H24O12 | 516.4 | Kaempferol 3-(3″,4″-diacetylrhamnoside) |
| 5 | 11994609 | C30H42O10 | 562.6 | Kalanchoside B |
| 6 | 14238625 | C27H30O16 | 610.5 | Quercetin 3-(2-glucosylrhamnoside) |
| 7 | 5369484 | C31H40O15 | 652.6 | Cinnamic acid (compound NP-002173) |
| 8 | 14189384 | C36H58O8 | 618.8 | Oleanolic acid beta-D-glucopyranosyl ester |
| 9 | 5274585 | C21H18O13 | 478.4 | Miquelianin |
| 10 | 129889798 | C18H16O9 | 376.3 | Tri-(methoxy)-kaempferol |
| 11 | 5280805 | C27H30O16 | 610.5 | Rutin |
| 12 | 16218542 | C27H36O19 | 664.6 | Rutin trihydrate |
| 13 | 12366 | C18H36O2 | 284.5 | Ethyl palmitate |
| 14 | 5274586 | C26H28O16 | 596.5 | Quercetin 3-O-beta-D-xylopyranosyl-(1→2)-beta-D-galactopyranoside |
| 15 | 10349838 | C32H36O18 | 708.6 | Kalambroside-A |
Network pharmacology-based analysis
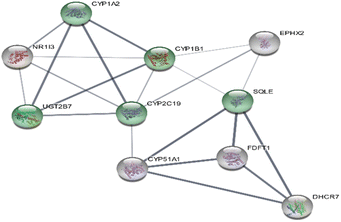
In parallel, ABC transporter-related target genes were retrieved from publicly available databases, including GeneCards and OMIM. After removal of duplicate entries, a total of 2123 ABC transporter targets were obtained. The intersection between the predicted ligand targets and ABC transporter-associated genes was performed using the online bioinformatics tool Venny, which identified 185 common genes as potential therapeutic targets, as presented in Fig. S3. The ligand–target interaction network consisted of 474 nodes and 1500 edges. The network exhibited a centralization value of 0.199, an average number of neighbors of 6.329, and a characteristic path length of 3.254, indicating a moderately connected biological network.
The different pathways enriched were identified through KEGG pathway enrichment analysis. We used DAVID GO for enrichment analysis to identify the pathways and obtained 204 signaling pathways. Among the top 20 pathways, sorted based on p-values from smallest to largest, the results are plotted in the form of a bar graph as shown in Fig. S6. The analysis revealed that key targets are enriched in pathways such as pathways in cancer, PI3K–AKT signaling pathway, estrogen signaling pathway, PD-1 checkpoint pathways in cancer, IL-17 signaling pathway, and Th17 cell differentiation. KEGG analysis shows that most of the common genes are involved in multiple pathways associated with cancer, providing valuable insights into the anti-cancer potential of K. laciniata compounds through ABC transporter interaction. Fig. S7.
 | ||
| Fig. 2 Interaction profiles of ABC transporter proteins and Hub genes with high-affinity KLM phytocompounds. | ||
These results underscore the importance of quercetin and kaempferol derivatives in mediating the drug resistance effects associated with ABC transporters. Overall, the findings from this study provide a solid basis for further investigation into the pharmacological applications of quercetin and kaempferol derivatives derived from Kalanchoe laciniata in multi-drug resistance treatment.
Dynamics simulation
Root mean square fluctuation (RMSF) analysis revealed that C-terminal regions of MRP1, P-gp, and AKT1 displayed higher flexibility compared to the N-terminal regions, which remained relatively stable, suggesting greater structural rigidity at the N-terminus. P-gp showed a pronounced peak around residue 600, indicating a highly flexible loop or exposed region, while AKT1 displayed notable fluctuations near residues 50, 250, and 300, reflecting dynamic loop regions. Protein–ligand interaction profiling further demonstrated that hydrogen bonding and hydrophobic interactions were the dominant stabilizing forces across complexes, with AKT1 and BCRP showing strong interaction profiles, including water bridges and ionic interactions. Two-dimensional interaction maps confirmed persistent contacts between key residues and ligands, supporting the stability and functional relevance of these complexes during simulation.
For comparison, the standard drug doxorubicin (positive control) exhibited a strong cytotoxic response. The results revealed a concentration-dependent increase in cytotoxicity, with percentage inhibitions of approximately 5.63%, 24.10%, 31.52%, 42.29%, 52.39%, 69.32%, and 92.89% at 0, 1, 2, 3, 4, 5, and 10 µg mL−1, respectively. The calculated IC50 value of doxorubicin was 4.10 µg mL−1, indicating potent cytotoxic activity against MDA-MB-231 cells Fig. 4B. The methanolic extract of Kalanchoe laciniata showed concentration-dependent cytotoxicity in normal RIN-5F cells (IC50 = 242.17 µg mL−1), which was substantially higher than that observed in MDA-MB-231 breast cancer cells (IC50 = 60.55 µg mL−1), indicating a favorable degree of cancer-selective toxicity (Fig. S12).
Isobologram analysis of KLM–doxorubicin combinations demonstrated a concentration- and effect-level-dependent interaction pattern. At IC30 and IC50, the experimental combination points were positioned below their respective additivity lines, indicative of synergistic interactions (Fig. 5). Consistent with the Chou–Talalay median-effect principle, this positioning corresponds to Combination Index (CI) values < 1, reflecting achievement of equivalent cytotoxic effects at lower-than-predicted doses relative to dose additivity. Such synergism suggests that KLM enhances doxorubicin efficacy at low to moderate effect levels, supporting a chemosensitizing role that may involve modulation of drug-resistance-associated mechanisms. In contrast, the IC70 combination point was located above the additivity line, corresponding to CI values > 1 and indicating an antagonistic interaction at higher effect levels (Table 5). This attenuation of synergy at elevated concentrations may be attributed to target saturation or activation of compensatory cellular responses, a phenomenon frequently reported in phytochemical-based combination studies.41,42 Collectively, the concordance between isobologram positioning and Chou–Talalay CI interpretation indicates that KLM exhibits optimal synergistic potential with doxorubicin at IC30–IC50 levels, supporting its role as a dose-modulating adjuvant rather than a high-dose cytotoxic agent.
| % Inhibition | CA (µg ml−1) | CB (µg ml−1) | ICA (µg ml−1) | ICB (µg ml−1) | Combination index | Synergy or antagonism |
|---|---|---|---|---|---|---|
| 29.979 | 10 | 1 | 28.79443 | 1.76507 | 0.913837 | Synergy |
| 50.844 | 20 | 2 | 61.88815 | 4.201118 | 0.799231 | Synergy |
| 60.486 | 40 | 3 | 77.17934 | 5.32685 | 1.081454 | Additive |
| 70.866 | 60 | 4 | 93.64249 | 6.538744 | 1.252469 | Antagonism |
| 83.603 | 80 | 5 | 113.8447 | 8.025826 | 1.325701 | Antagonism |
| 95.320 | 100 | 10 | 132.4285 | 9.393819 | 1.819656 | Antagonism |
Quantitative real-time PCR analysis demonstrated that KLM treatment significantly altered the mRNA expression of AKT1 and ABC transporter genes, including BCRP, P-gp, and MRP1, compared with the untreated control group (Fig. 4D). Notably, KLM treatment was associated with a significant downregulation of AKT1, BCRP, and MRP1 transcript levels. In contrast, the expression of P-gp did not show a statistically significant increase under the experimental conditions employed. These results indicate that KLM influences the transcriptional levels of selected genes implicated in cell survival and drug resistance pathways. These findings suggest that KLM modulates AKT signaling and selectively influences the expression of multidrug resistance transporters, indicating its potential role in regulating cancer cell survival and drug resistance mechanisms.
Discussion and conclusion
Discussion
Multidrug resistance (MDR) mediated by ATP-binding cassette (ABC) transporters remains a major challenge in effective cancer chemotherapy. In this study, Kalanchoe laciniata was investigated as a potential source of MDR-modulating phytochemicals through an integrated approach combining phytochemical profiling, network pharmacology, molecular docking, and experimental validation. Our findings provide mechanistic evidence that flavonoid glycosides from K. laciniata, particularly quercetin and kaempferol derivatives, target key regulators of drug resistance and cancer cell survival43Phytochemical analyses consistently identified quercetin- and kaempferol-based glycosides as major constituents of the KLM. Network pharmacology analysis revealed that these compounds converge on ABC transporters, including P-glycoprotein (P-gp), breast cancer resistance protein (BCRP), and multidrug resistance protein-1 (MRP1), as well as survival-associated hub genes such as AKT1 and TP53. Enrichment of the PI3K–AKT signaling pathway and drug resistance-related pathways highlight a coordinated mechanism through which K. laciniata phytochemicals may regulate both transporter activity and intracellular survival signaling. This multi-target action aligns with current concepts that MDR arises from interconnected molecular networks rather than single-gene alterations.44,45
Molecular docking and molecular dynamics simulations further supported these findings by demonstrating stable interactions between quercetin and kaempferol derivatives and ABC transporters, particularly BCRP and MRP1. These interactions involved key residues within the transporter binding pockets, suggesting potential competitive or allosteric inhibition of efflux activity. Molecular dynamics simulations (100 ns) revealed that the BCRP–rutin, MRP1–quercetin 3-(2-glucosylrhamnoside), and AKT1–quercetin 3-O-β-D-xylopyranosyl-(1→2)-β-D-galactopyranoside complexes rapidly achieved structural stability within the first 10 ns and remained stable throughout the simulation (RMSD 0.5–4 Å), whereas the P-gp–quercetin 3-(2-glucosylrhamnoside) complex exhibited higher RMSD fluctuations, indicating weak and unstable binding.46 Notably, strong interactions with BCRP are of particular interest, as BCRP-mediated drug efflux contributes significantly to resistance against several clinically used chemotherapeutics and remains less extensively explored than P-gp.15
Additionally, interactions with AKT1 suggest that these flavonoids may indirectly regulate transporter expression through suppression of pro-survival signaling pathways.43 It is also important to note that in silico prediction tools are biased toward well-characterized targets and pathways, which may limit identification of less-studied or context-specific interactions. Accordingly, the predicted networks should be interpreted as hypothesis-generating. To partially address this limitation, experimental validation was performed using qPCR analysis of selected key target genes, providing preliminary biological support for the network pharmacology predictions.
Quantitative real-time PCR analysis demonstrated that KLM treatment significantly downregulated the mRNA expression of AKT1, BCRP, and MRP1, providing experimental support for selected targets predicted through network pharmacology and molecular docking.47 Suppression of AKT1 is particularly noteworthy given its established role in promoting cell survival, inhibiting apoptosis, and regulating ABC transporter expression. In contrast, P-gp (ABCB1) expression did not show significant transcriptional modulation, which is consistent with the relatively lower stability observed for the P-gp–ligand complex during molecular dynamics simulations. Taken together, these findings suggest that KLM-mediated MDR modulation may preferentially involve AKT signaling, BCRP, and MRP1, rather than broad or nonspecific inhibition of all ABC transporters. Accordingly, the in silico and qPCR data were interpreted in a complementary and cautious manner, highlighting target selectivity rather than uniform ABC transporter inhibition. While qPCR analysis provided initial experimental support for the in silico predicted modulation of ABC transporter related genes, the absence of protein expression and functional activity assays represents a limitation of the present study. Future investigations incorporating transporter activity assays and protein-level validation will be essential to confirm the functional relevance of these findings.
Functionally, KLM exhibited selective cytotoxicity toward MDA-MB-231 breast cancer cells compared with normal RIN-5F cells, indicating a favorable in vitro selectivity profile. In addition, the synergistic interaction observed between KLM and doxorubicin at defined effect levels suggests enhanced cytotoxic efficacy when used in combination. While the observed synergy is consistent with reduced ABC transporter-mediated drug efflux, confirmation of direct transporter activity modulation will require dedicated efflux assays in future studies. Taken together, the cytotoxicity and combination data provide preliminary biological context for the molecular and transcriptional observations, supporting a hypothesis of selective MDR modulation by KLM under in vitro conditions.
Conclusion
Collectively, this study provides integrative evidence that Kalanchoe laciniata exerts a multi-targeted influence on MDR-associated pathways, involving modulation of ABC transporter-related genes and survival signaling at the in silico and transcriptional levels. The combined computational and in vitro findings identify quercetin and kaempferol derived constituents as potential chemosensitizing candidates, rather than confirmed therapeutic agents. While these results highlight the promise of K. laciniata as an adjunct in combination chemotherapy, comprehensive validation through functional efflux assays, protein-level analysis, and in vivo pharmacokinetic and efficacy studies will be essential before proceeding to translational and clinical studies.Author contributions
Anish Ruban S.: writing – original draft, conceptualization, methodology, investigation; Francis Jegan Raj: formal analysis, data curation; Vetri Velavan Sundararajan: validation, resources, writing – review & editing; Parimelazhagan Thangaraj: writing – review, supervision.Conflicts of interest
The authors declare no conflict of interest.Data availability
No software or code have been included and all data generated has been included in either the manuscript or supplementary data.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra09649a.
Acknowledgements
We thank CSIR-UGC, and the Department of Botany, Bharathiar University, Coimbatore, for the support provided to undertake this study. This work was funded by CSIR UGC (NTA Ref. No: 211610197248), and the Department of Botany, Bharathiar University, Coimbatore.References
- J. M. Fernandes, L. M. Cunha, E. P. Azevedo, E. M. G. Lourenço, M. F. Fernandes-Pedrosa and S. M. Zucolotto, Rev. Bras. Farmacogn., 2019, 29, 529–558 CrossRef CAS.
- S. El Abdellaoui, E. Destandau, A. Toribio, C. Elfakir, M. Lafosse, I. Renimel, P. André, P. Cancellieri and L. Landemarre, Anal. Bioanal. Chem., 2010, 398, 1329–1338 CrossRef CAS PubMed.
- E. R. D. de Araújo, G. C. B. Guerra, D. F. de Souza Araújo, A. A. de Araújo, J. M. Fernandes, R. F. de Araújo Júnior, V. C. da Silva, T. G. de Carvalho, L. de Santis Ferreira and S. M. Zucolotto, Int. J. Mol. Sci., 2018, 19(5), 1265 CrossRef PubMed.
- A. Rudzińska, P. Juchaniuk, J. Oberda, J. Wiśniewska, W. Wojdan, K. Szklener and S. Mańdziuk, Nutrients, 2023, 15, 1896 CrossRef PubMed.
- D. M. Kopustinskiene, V. Jakstas, A. Savickas and J. Bernatoniene, Nutrients, 2020, 12(2), 457 CrossRef CAS PubMed.
- H. Patel, Z. X. Wu, Y. Chen, L. Bo and Z. S. Chen, Mol. Biomed., 2021, 2, 27 CrossRef PubMed.
- X. Wang, H. Zhang and X. Chen, Cancer Drug Resist., 2019, 2, 141–160 Search PubMed.
- A. E. van Herwaarden and A. H. Schinkel, Trends Pharmacol. Sci., 2006, 27(1), 10–16 CrossRef CAS PubMed.
- R. W. Robey, K. M. Pluchino, M. D. Hall, A. T. Fojo, S. E. Bates and M. M. Gottesman, Nat. Rev. Cancer, 2018, 18(7), 452–464 CrossRef CAS PubMed.
- K. Katayama, K. Masuyama, S. Yoshioka, H. Hasegawa, J. Mitsuhashi and Y. Sugimoto, Cancer Chemother. Pharmacol., 2007, 60, 789–797 CrossRef CAS PubMed.
- H. Xiao, Y. Zheng, L. Ma, L. Tian and Q. Sun, Front. Pharmacol, 2021, 12, 648407 CrossRef CAS PubMed.
- M. E. Hernández-Caballero, J. A. Sierra-Ramírez, R. Villalobos-Valencia and E. Seseña-Méndez, Molecules, 2022, 27, 6425 CrossRef PubMed.
- D. C. Faustino, N. N. das Chagas Lima, K. J. Allahdadi and L. C. Pinto, RPS Pharm. Pharmacol. Rep., 2022, 1, rqac009 CrossRef.
- F. B. Barlas, J. Radiat. Res. Appl. Sci., 2023, 16, 100612 CAS.
- A. Singh, S. K. Patel, P. Kumar, K. C. Das, D. Verma, R. Sharma, T. Tripathi, R. Giri, N. Martins and N. Garg, J. Biomol. Struct. Dyn., 2022, 40, 4507–4515 CrossRef CAS PubMed.
- G. An, J. Gallegos and M. E. Morris, Drug Metab. Dispos., 2011, 39, 426–432 CrossRef CAS PubMed.
- S. A. Ruban, F. J. Raj and P. Thangaraj, Biochim. Biophys. Acta, Rev. Cancer, 2025, 1880, 189349 CrossRef PubMed.
- A. A. Jovanović, V. B. Đorđević, G. M. Zdunić, D. S. Pljevljakušić, K. P. Šavikin, D. M. Gođevac and B. M. Bugarski, Sep. Purif. Technol., 2017, 179, 369–380 CrossRef.
- F. J. Raj, G. Jagadeesan, B. M. Paul, P. Thangaraj and R. Kilimas, Appl. Biochem. Biotechnol., 2023, 195, 6790–6808 CrossRef CAS PubMed.
- K. Gurning, H. A. Simanjuntak, H. Purba, R. F. R. Situmorang, L. Barus and S. Silaban, J. Phys., Conf. Ser., 2021, 1811, 012121 CrossRef CAS.
- A. Mehmood, S. Javid, M. F. Khan, K. S. Ahmad and A. Mustafa, BMC Chem., 2022, 16, 1–10 Search PubMed.
- İ. Gulcin and S. H. Alwasel, Processes, 2023, 11, 2248 CrossRef.
- A. Mehta, Vegetos, 2023, 36, 1570–1575 CrossRef CAS.
- H. Aziz, A. Saeed, F. Jabeen, M. A. Khan, A. Ur Rehman, M. Q. Khan and M. Saleem, J. Mol. Struct., 2023, 1278, 134924 CrossRef CAS.
- J. Rumpf, R. Burger and M. Schulze, Int. J. Biol. Macromol., 2023, 233, 123470 CrossRef CAS PubMed.
- R. Apak, A. Calokerinos, S. Gorinstein, M. A. Segundo, D. B. Hibbert, I. Gülçin, S. D. Çekiç, K. Güçlü, M. Özyürek, S. E. Çelik, L. M. Magalhães and P. Arancibia-Avila, Pure Appl. Chem., 2022, 94, 87–144 CrossRef CAS.
- F. P. Nonglang, A. Khale and S. Bhan, Future J. Pharmaceut. Sci., 2022, 8, 1–12 Search PubMed.
- J. M. Dinore, H. S. Patil, B. S. Dobhal and M. Farooqui, Nat. Prod. Res., 2022, 36, 5631–5637 CrossRef CAS PubMed.
- L. Zhao, H. Zhang, N. Li, J. Chen, H. Xu, Y. Wang and Q. Liang, J. Ethnopharmacol., 2023, 309, 116306 CrossRef CAS PubMed.
- M. Kohl, S. Wiese and B. Warscheid, Methods Mol. Biol., 2011, 696, 291–303 CrossRef CAS PubMed.
- SRplot-Science and Research online plot, https://www.bioinformatics.com.cn/srplot, accessed 31 May 2025 Search PubMed.
- RCSB PDB: Homepage, https://www.rcsb.org/, accessed 31 May 2025 Search PubMed.
- A. Lakshmanan, B. Balasubramanian, V. Maluventhen, A. Malaisamy, R. Baskaran, W. C. Liu and M. Arumugam, Appl. Sci., 2022, 12, 13010 CrossRef CAS.
- S. Menaga and N. Geetha, Next Res., 2025, 2, 100500 CrossRef.
- S. K. Mandal, B. K. Kumar, P. K. Sharma, S. Murugesan and P. R. Deepa, Comput. Biol. Med., 2022, 147, 105796 Search PubMed.
- N. S. Natesh, M. Arumugam and G. Karanam, Mol. Biol. Rep., 2018, 45(6), 2641–2651 CrossRef PubMed.
- T. C. Chou, Cancer Res., 2010, 70, 440–446 CrossRef CAS PubMed.
- G. Karanam and M. K. Arumugam, Mol. Biol. Rep., 2020, 47(5), 3347–3359 CrossRef CAS PubMed.
- M. Suriyakanthan and G. Natesan, RSC Adv., 2025, 15, 49399–49417 RSC.
- S. Menaga and N. Geetha, Next Re., 2025, 100500 CrossRef.
- L. K. Caesar and N. B. Cech, Nat. Prod. Rep., 2019, 36, 869–888 RSC.
- N. Chaachouay, Drugs Drug Candidates, 2025, 4, 4 CrossRef.
- N. S. Yarla, J. Mar. Sci. Res. Dev., 2013, 4(1) DOI:10.4172/2155-9910.1000e123.
- R. Liu, Y. Chen, G. Liu, C. Li, Y. Song, Z. Cao, W. Li, J. Hu, C. Lu and Y. Liu, Cell Death Dis., 2020, 11, 1–12 Search PubMed.
- S. A. Mirzaei, F. Dinmohammadi, A. Alizadeh and F. Elahian, Life Sci., 2019, 235, 116825 CrossRef CAS PubMed.
- S. A. Hollingsworth and R. O. Dror, Neuron, 2018, 99(6), 1129–1143 CrossRef CAS PubMed.
- G. Ramarajyam, R. Murugan and S. Rajendiran, Hum. Gene, 2024, 42, 201328 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |