Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Isolation and characterization of antiplasmodial and antimicrobial compounds from Tetracera alnifolia using a bioassay-guided approach
Mamadou Aliou Baldé *a,
Mohamed Sahar Traoréad,
Mamadou Saliou Telly Dialloa,
An Matheeussenc,
Camara Aïssataa,
Paul Cosc,
Louis Maesc,
Alpha Oumar Baldéa,
Mohamed Kerfala Camaraa,
Aliou Mamadou Baldea,
Emmy Tuenterb and
Kenn Foubertb
*a,
Mohamed Sahar Traoréad,
Mamadou Saliou Telly Dialloa,
An Matheeussenc,
Camara Aïssataa,
Paul Cosc,
Louis Maesc,
Alpha Oumar Baldéa,
Mohamed Kerfala Camaraa,
Aliou Mamadou Baldea,
Emmy Tuenterb and
Kenn Foubertb
aDepartment of Pharmaceutical and Biological Sciences, University Gamal Abdel Nasser of Conakry, Guinea. E-mail: baldem@ufl.edu
bNatural Products & Food Research and Analysis – Pharmaceutical Technology (NatuRAPT), Department of Pharmaceutical Sciences, University of Antwerp, Universiteitsplein 1, B-2610 Antwerp, Belgium
cLaboratory for Microbiology, Parasitology and Hygiene (LMPH), Faculty of Pharmaceutical, Biomedical and Veterinary Sciences, University of Antwerp, Universiteitsplein 1, B-2610 Antwerp, Belgium
dResearch Institute of Applied Biology of Guinea (IRBAG), Kindia, Guinea
First published on 23rd February 2026
Abstract
A preliminary biological screening demonstrated that leaf extracts from Tetracera alnifolia exhibit promising activity against Plasmodium falciparum and Candida albicans. To identify the constituents responsible for these effects, a bioassay-guided fractionation strategy was subsequently employed. Fractions were purified using flash chromatography, followed by semi-preparative HPLC-DAD-MS and LC-SPE-NMR. Structural elucidation of the isolated compounds was accomplished through comprehensive 1D and 2D NMR analyses in combination with HR-ESI-MS. The purified metabolites were subsequently evaluated for their bioactivity against Plasmodium falciparum and Candida albicans, and their cytotoxicity was assessed in MRC-5SV2 human foetal lung fibroblast cells. In total, nineteen compounds (1–19) were isolated from T. alnifolia leaves. Among these, the strongest antiplasmodial activities were observed for pheophorbide-b methyl ester (1.0 ± 0.7 µM), (1,2)-bis-nor-phytone (2.0 µM), isophytol (4.0 µM), pheophorbide-a methyl ester (2.8 ± 1.2 µM), epicatechin-3-galloyl ester (5.5 ± 2.1 µM), and phytol (6.9 ± 2.4 µM). Other compounds, including myricetin-3-O-rhamnopyranoside, α-tocopherol and cycloart-24-en-3β-yl α-linolenate, exhibited lower activity, with IC50 values ranging from 13.5 to 25 µM. None of the evaluated compounds showed antifungal activity against C. albicans. Notably, high cytotoxicity was observed for pheophorbide-b methyl ester, pheophorbide-a methyl ester, and phytol. These findings provide insight into the constituents that underlie the antiplasmodial activity of T. alnifolia leaf extracts.
1 Introduction
The discovery of new bioactive natural products as leads for therapeutic development can be inspired by ethnopharmacological knowledge or achieved by screening a collection of extracts for bioactivity, using several bioassays.1,2 The traditional workflow for studying complex plant preparations is bioactivity-guided fractionation in which bioactive extracts and subsequent fractions are chromatographically separated and retested for bioactivity until active compounds have been isolated.3,4 In the last decade, substantial developments have been made in improving extraction and separation efficiency and facilitating the isolation of minor constituents that may contribute to activity.5 Also, nowadays, the bioassay-guided fractionation workflow is still used by scientists for the discovery of new therapeutic leads from natural sources, including plant species.1The Dilleniaceae family comprises 10–14 genera and ca. 500 species, with a pantropical and subtropical distribution. Around 102 species distributed across six genera viz. Curatella, Davilla, Doliocarpus, Neodillenia, Pinzona, and Tetracera are mainly found in the Neotropical region. The Dilleniaceae family is divided into four subfamilies and the genus Tetracera belongs to the Delimoideae subfamily.6,7 The genus Tetracera contains about 50 species with a pantropical distribution, of which 20 occur in the Neotropical area. In traditional folk medicine, some species have been used for the management of various diseases and infections.8,9 Species of this genus have been reported to exhibit a broad range of pharmacological activities, including antifungal, antiplasmodial, analgesic, anti-inflammatory, antioxidant, antipyrexic, antimycobacterial, anti-HIV, anti-ulcerogenic and hepatoprotective.8,10 Several Tetracera species have previously been investigated for their phytochemical constituents, which resulted in the identification of a broad range of compounds, including flavonoids and terpenoids.10,11 Within the Guinean flora, approximately four species of the genus Tetracera have been identified, among which T. alnifolia is distributed across all regions of Guinea. T. alnifolia, a forest liana up to 15 m high, is widely used in Guinean traditional medicine for the management of several diseases including skin diseases, oral diseases, infectious diseases and malaria.12–15 This plant species is also well known in other African countries for its traditional uses in the management of inflammation, pain, cough, sexually transmitted infections and leprosy.9,16,17 Promising in vitro activities against P. falciparum and C. albicans were previously obtained in our laboratory for both methanolic and dichloromethane extracts of the leaves of T. alnifolia.18 This study aimed to isolate metabolites potentially responsible for the observed biological activities, which may serve as promising precursors for the development of novel antimicrobial and/or antiplasmodial agents. In this context, a bioguided fractionation of leaf extracts from T. alnifolia was conducted.
2 Materials and methods
2.1. Solvent and reagents
HPLC-grade acetone, acetonitrile (including far-UV grade), dichloromethane, ethanol, ethyl acetate, methanol, n-butanol, and n-hexane were sourced from Fisher Scientific (Leicestershire, UK). Formic acid, sulfuric acid and vanillin were supplied by Acros Organics (Geel, Belgium) or Sigma-Aldrich (St. Louis, MO, USA). For UPLC-MS analyses, acetonitrile and formic acid were purchased from Biosolve Chimie (Dieuze, France). Ultrapure water was produced using a Milli-Q purification system (Millipore, Bedford, MA, USA). Deuterated solvents for NMR spectroscopy (acetonitrile-d3, CDCl3, D2O, DMSO-d6, methanol-d4, and pyridine-d5) were purchased from Sigma-Aldrich.2.2. General experimental methods
Extracts and derived fractions were separated using Diaion HP-20 open-column chromatography in combination with a Reveleris X2 flash chromatography system (BUCHI Labortechnik, Breda, Netherlands) equipped with ELSD, DAD, and automated fraction collection. Chromatographic profiles of fractions collected throughout the isolation workflow were analyzed using an Agilent HPLC-DAD system (1200 Series) and/or thin-layer chromatography (TLC) on NP-F254 plates (20 cm × 20 cm; Merck, Darmstadt, Germany). TLC plates were examined under UV light at 254 and 366 nm, as well as under visible light after derivatization with vanillin sulfuric acid reagent (prepared by dissolving 5 g vanillin in 475 mL ethanol and adding 25 mL sulfuric acid). Pure compounds were isolated using a semi-preparative HPLC platform equipped with DAD and ESI-MS detection. The system comprised a 2767 sample manager, injector, and collector, a 2545 quaternary gradient module, a System Fluidics Organizer, a 515 HPLC pump, a 2998 DAD, and a Micromass Quattro TQD mass spectrometer (Waters, Milford, MA, USA). Data were processed using MassLynx version 4.1.Low yield subfractions (<100 mg) were purified using HPLC-SPE-NMR. Analyses were performed on an Agilent 1200 Series HPLC system (Agilent Technologies) equipped with a degasser, quaternary pump, autosampler, and photodiode-array (PDA) detector. Target analytes were cumulatively trapped on SPE cartridges by UV-threshold-triggered collection. Downstream of the detector, water was introduced at 3 mL min−1 via a make-up pump (Knauer K-120, Berlin, Germany) to enhance retention on the SPE cartridges. Trapping was carried out on a Bruker/Spark SPE platform fitted with HySphere Resin General Phase (GP) cartridges (poly(divinylbenzene), 5–15 µm). After the final trapping, each cartridge was dried with pressurized nitrogen gas for 40 min and eluted with 60 µL of deuterated solvent (acetonitrile-d3 or methanol-d4) into 3 mm NMR tubes with a Gilson Liquid Handler 125 (The Hague, Netherlands). Chromatographic separation and analyte trapping on SPE cartridges were controlled using Hystar v. 3.2 software (Bruker Daltonik, Bremen, Germany), whilst the elution process was controlled by Prep Gilson ST ver. 1.2 software (Bruker Biospin, Karlsruhe, Germany). Nuclear magnetic resonance (NMR) experiments, including both one and two dimensional measurements, were carried out using a Bruker DRX-400 spectrometer (Bruker BioSpin, Rheinstetten, Germany). The instrument was fitted with either a 3 mm inverse broadband (BBI) probe or a 5 mm dual 1H/13C probe, and experiments were conducted using routine Bruker pulse sequences. Data were collected at operating frequencies of 400 MHz for proton (1H) and 100 MHz for carbon (13C). All spectra were processed and interpreted using TopSpin software (version 4.0.6, Bruker BioSpin).
High-resolution mass spectrometric measurements were obtained using a Xevo G2-XS QTof instrument (Waters, Milford, MA, USA) coupled to an ACQUITY UPLC chromatographic platform and operated via MassLynx software (v4.1). Chromatographic separation was performed on a Waters Acquity UHPLC BEH Shield RP18 column (2.1 × 100 mm, 1.7 µm particle size). The elution system consisted of water supplemented with 0.1% formic acid (solvent A) and acetonitrile supplemented with 0.1% formic acid (solvent B), with a constant flow rate of 0.40 mL min−1. The gradient profile was set as follows: initial 2% B (0–1 min), increased to 100% B over 1–5 min, maintained at 100% B for 5–7 min, returned to 2% B during 7–8 min, and equilibrated at 2% B from 8–10 min. Mass detection was carried out using electrospray ionization operated in both positive and negative modes, with data acquired over an m/z interval of 50–1500 under sensitivity mode conditions (≈22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 FWHM). The capillary voltage was adjusted to +1.5 kV for positive ionization and −1.0 kV for negative ionization. Cone gas and desolvation gas flow rates were set to 50 and 1000 L h−1, respectively. The ion source temperature was maintained at 120 °C, while the desolvation temperature was set to 550 °C. Continuous mass accuracy was ensured through lock-mass correction using leucine enkephalin.
000 FWHM). The capillary voltage was adjusted to +1.5 kV for positive ionization and −1.0 kV for negative ionization. Cone gas and desolvation gas flow rates were set to 50 and 1000 L h−1, respectively. The ion source temperature was maintained at 120 °C, while the desolvation temperature was set to 550 °C. Continuous mass accuracy was ensured through lock-mass correction using leucine enkephalin.
2.3. Plant material
T. alnifolia leaves were harvested in June 2017 in Telimélimé, Republic of Guinea. Species authentication was performed by botanists at the Research and Valorization Center on Medicinal Plants (Dubréka, Guinea), where a voucher specimen (D42HK2) has been deposited. The collected material was air-dried at room temperature and subsequently ground into a fine powder.2.4. Extraction, bioguided fractionation and isolation
The air-dried and ground leaves of T. alnifolia (780 g) were initially defatted with n-hexane (5 × 2.5 L) and subsequently extracted with 80% methanol (5 × 2.5 L) at room temperature under constant magnetic stirring. For each solvent, the plant material was macerated for 24 h, with the solvent replaced daily over a total period of five days. Pooled filtrates were concentrated under reduced pressure and freeze-dried to obtain the hexane-soluble fraction (TE) (12 g) and the crude methanolic fraction (TM) (207 g) which was redissolved in water and partitioned with CH2Cl2 to give a dichloromethane soluble fraction (TD) (10 g). Based on the biological activity results, the active fractions (hexane, dichloromethane and methanol) were selected and further purified by open column chromatography, flash chromatography semi preparative HPLC-DAD-MS and LC-SPE-NMR.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9, v/v; detection at 254 nm). Fraction TE2 (2.3 g) was further purified by flash chromatography on a silica gel column (GraceResolv, 40 g) using n-hexane (A) and dichloromethane (B) as eluents at a flow rate of 40 mL min−1, affording compounds 1 (16.0 mg) and 2 (56.0 mg). The elution gradient consisted of 100% A from 0–5 min; a gradual increase of B from 5–55 min to reach 20% B; a further increase to 35% B from 55–65 min; and finally, a rise to 100% B from 65–70 min. The column was held at 100% B for an additional 5 min (70–75 min). Fractions TE3 and TE4 were purified by preparative TLC using dichloromethane/n-hexane (1
9, v/v; detection at 254 nm). Fraction TE2 (2.3 g) was further purified by flash chromatography on a silica gel column (GraceResolv, 40 g) using n-hexane (A) and dichloromethane (B) as eluents at a flow rate of 40 mL min−1, affording compounds 1 (16.0 mg) and 2 (56.0 mg). The elution gradient consisted of 100% A from 0–5 min; a gradual increase of B from 5–55 min to reach 20% B; a further increase to 35% B from 55–65 min; and finally, a rise to 100% B from 65–70 min. The column was held at 100% B for an additional 5 min (70–75 min). Fractions TE3 and TE4 were purified by preparative TLC using dichloromethane/n-hexane (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) and toluene (100%) as mobile phases, respectively, with detection at 254 nm. TE3 afforded compound 3 (7.5 mg), whereas TE4 yielded compounds 4 (3.2 mg) and 5 (8.2 mg).
1, v/v) and toluene (100%) as mobile phases, respectively, with detection at 254 nm. TE3 afforded compound 3 (7.5 mg), whereas TE4 yielded compounds 4 (3.2 mg) and 5 (8.2 mg).![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, v/v). Preparative TLC of TD1 (0.3 g) with dichloromethane/n-hexane (9
2, v/v). Preparative TLC of TD1 (0.3 g) with dichloromethane/n-hexane (9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) afforded compound 6 (13.5 mg), and TD2 yielded compounds 7 (4.2 mg) and 8 (3.7 mg). Fraction TD4 (0.77 g) was further purified by flash chromatography over a silica gel column (GraceResolv, 40 g), employing dichloromethane (A), ethyl acetate (B), and methanol (C) as mobile phases at a flow rate of 40 mL min−1. This separation yielded compounds 9 (14.6 mg), 10 (15.3 mg), and 11 (9.4 mg). The gradient was programmed as follows: 100% A (0–10 min); a linear increase of B to 100% (10–48 min); followed by the increasing amount of C to reach 20% (48–78 min). The final composition was held for 10 min (78–88 min). Compound 12 (4.5 mg) was isolated from fraction TD5 (0.3 g) by semi-preparative HPLC-MS on a C18 Kinetex column (250 × 10 mm, 5 µm; Phenomenex, Utrecht, The Netherlands), using a gradient of H2O + 0.1% formic acid (A) and far-UV acetonitrile (B) at 3 mL min−1. The gradient program was: (0–5 min) 20%, (20 min) 70% B, (35 min) 95% B, (41 min) 20% B, (41–46 min) 20% B. DAD spectra were acquired over the range 200–450 nm. Mass spectra were recorded in ESI (+) and ESI (−) modes under the following conditions: scan range m/z 100–1000; capillary voltage 3.00 kV; cone voltage 50 V; extractor voltage 3 V; source temperature 135 °C; desolvation temperature 400 °C; desolvation gas flow 750 L h−1; and cone gas flow 50 L h−1. The injection volume was 300 µL. Purification of fraction TD7 by LC-SPE-NMR allowed the isolation of compound 13 (1.3 mg). An Agilent Zorbax SB-phenyl column (4.6 × 250 mm, 5 µm) was used with a mobile phase consisting of water + 0.1% formic acid (A) and acetonitrile (B), and a flow rate of 1 mL min−1. The following gradient was used: (0–5 min) 20% B, (15 min) 60% B, (30 min) 96% B, (30–35 min) 96% B, (36 min) 20% B, (36–41 min) 20% B.
1, v/v) afforded compound 6 (13.5 mg), and TD2 yielded compounds 7 (4.2 mg) and 8 (3.7 mg). Fraction TD4 (0.77 g) was further purified by flash chromatography over a silica gel column (GraceResolv, 40 g), employing dichloromethane (A), ethyl acetate (B), and methanol (C) as mobile phases at a flow rate of 40 mL min−1. This separation yielded compounds 9 (14.6 mg), 10 (15.3 mg), and 11 (9.4 mg). The gradient was programmed as follows: 100% A (0–10 min); a linear increase of B to 100% (10–48 min); followed by the increasing amount of C to reach 20% (48–78 min). The final composition was held for 10 min (78–88 min). Compound 12 (4.5 mg) was isolated from fraction TD5 (0.3 g) by semi-preparative HPLC-MS on a C18 Kinetex column (250 × 10 mm, 5 µm; Phenomenex, Utrecht, The Netherlands), using a gradient of H2O + 0.1% formic acid (A) and far-UV acetonitrile (B) at 3 mL min−1. The gradient program was: (0–5 min) 20%, (20 min) 70% B, (35 min) 95% B, (41 min) 20% B, (41–46 min) 20% B. DAD spectra were acquired over the range 200–450 nm. Mass spectra were recorded in ESI (+) and ESI (−) modes under the following conditions: scan range m/z 100–1000; capillary voltage 3.00 kV; cone voltage 50 V; extractor voltage 3 V; source temperature 135 °C; desolvation temperature 400 °C; desolvation gas flow 750 L h−1; and cone gas flow 50 L h−1. The injection volume was 300 µL. Purification of fraction TD7 by LC-SPE-NMR allowed the isolation of compound 13 (1.3 mg). An Agilent Zorbax SB-phenyl column (4.6 × 250 mm, 5 µm) was used with a mobile phase consisting of water + 0.1% formic acid (A) and acetonitrile (B), and a flow rate of 1 mL min−1. The following gradient was used: (0–5 min) 20% B, (15 min) 60% B, (30 min) 96% B, (30–35 min) 96% B, (36 min) 20% B, (36–41 min) 20% B.The purification of fraction TM3–2 by LC-SPE-NMR yielded compound 14 (2.0 mg). The mobile phase consisted of water + 0.1% formic acid (A) and acetonitrile (B) pumped at a flow rate of 1 mL min−1. The gradient was set as follows: (0–5 min) 20% B, (15 min) 60% B, (30 min) 96% B, (30–35 min) 96% B, (36 min) 20% B, (36–41 min) 20% B. The peak detected at 280 nm was repeatedly collected on SPE cartridges during 11 consecutive runs. In turn the purification of the fraction TM9 (0.35 g) by semi-preparative HPLC-MS using a C18 Kinetex column (250 cm × 10.0 mm, particle size, 5 µm) and eluted with a mixture of water (A) and acetonitrile (B) yielded compounds 15 (8.0 mg) and 16 (7.6 mg). The gradient was set as follows: (0–5 min) 20% B, (40 min) 25% B, (50 min) 95% B, (50–55 min) 95% B, with a flow rate of 4.75 mL min−1.
Fractions TM10 and TM12 were further subjected to semi-preparative HPLC-MS analysis. Indeed, fraction TM10 (0.2 g) was purified using a gradient of 16% B (0–5 min), 30% B (40 min), and 96% B (50–55 min) at a flow rate of 4.75 mL min−1, yielding compounds 17 (5.0 mg) and 18 (24.0 mg). Fraction TM12 (0.25 g) was purified with a gradient of 20% B (0–5 min), 50% B (60 min), and 95% B (70–75 min) at a flow rate of 3.0 mL min−1, affording compound 19 (3.5 mg). All fractions were prepared at a concentration of 20 mg mL−1, with injection volumes of 300 µL for TM9 and TM10, and 400 µL for TM12.
2.5. LC-ESI-MS
LC-MS analysis was conducted under previously described metabolite profiling conditions, with minor modifications.19 The extract was initially dissolved in 80% MeOH (v/v) to 1 mg mL−1 and subsequently diluted with water to achieve a final concentration of 0.1 mg mL−1. Fraction samples were further diluted to a working concentration of 0.05 mg mL−1, whereas purified metabolites were analyzed at 0.01 mg mL−1.2.6. Biological evaluation
3 Results and discussion
Preliminary biological studies have shown that extracts from the leaves of T. alnifolia were active against P. falciparum and C. albicans. In this study, a classical activity-guided fractionation approach was used in order to characterize the active metabolites of T. alnifolia leaves.3.1. Purification and identification of isolated compounds
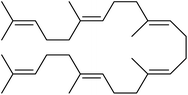
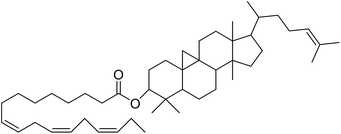
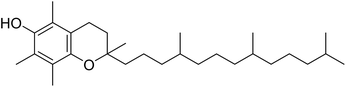
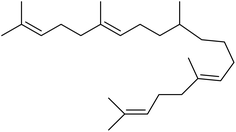
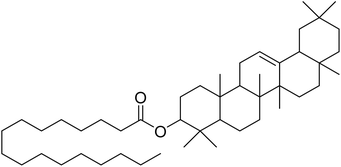
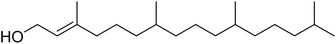
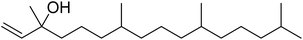
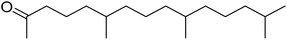
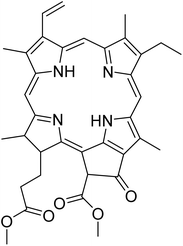
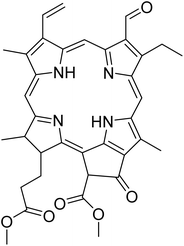
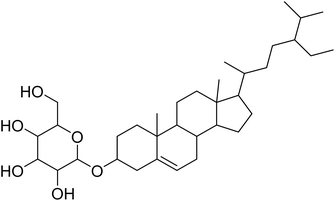
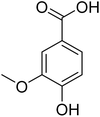
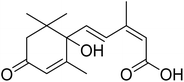
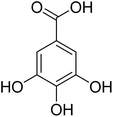
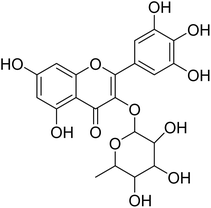
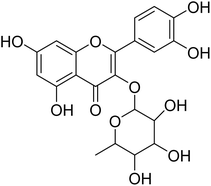
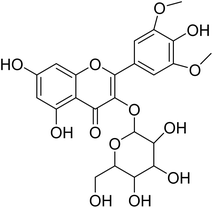
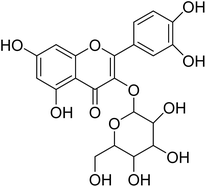
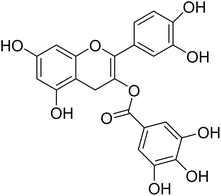
The fractions exhibiting the strongest activities (n-hexane, dichloromethane, and methanol) against P. falciparum and C. albicans (Table 1) were subjected to a multistep purification workflow comprising normal and reversed-phase flash chromatography, semi-preparative HPLC-MS, and LC-SPE-NMR. This process yielded a total of 19 distinct metabolites. Of these, five were isolated from the n-hexane fraction, eight from the dichloromethane fraction, and six from the methanol fraction. Structural characterization of all isolated metabolites was performed using HR-ESI-MS in conjunction with detailed NMR spectroscopy, employing both 1D (1H, 13C, DEPT-135, DEPT-90) and 2D (COSY, HSQC, HMBC) experiments. The spectroscopic data obtained were in full agreement with previously published assignments, allowing the identification of squalene (1),22,23 cycloart-24-en-3β-yl α-linolenate (2),24 α-tocopherol (3),25,26 trans-pentamethyl-icosa-tetraene (4),27 3-β-hydroxy-olean-12-ene-heptadecanoate (5), phytol (6),28 isophytol (7),29 (1, 2)-bis-nor-phytone (8),30 pheophorbide-a methyl ester (9),31,32 pheophorbide-b methyl ester (10),33 stigma-5-en-3-O-β-glucoside (11),34 vanillic acid (12),35 abscisic acid (13),36,37 gallic acid (14),38 myricetin-3-O-rhamnopyranoside (15),39 quercetin-3-O-rhamnopyranoside (16),40 myricetin-3′,5′-dimethylether-3-O-galactopyranoside (17),41 quercetin-3-O-β-D-galactopyranoside (18)42 and epicatechin-3-galloylester (19).43 Apart from gallic acid (14), all compounds isolated in this study are reported for the first time in T. alnifolia.| Plant name | Antimicrobial activity | Antiplasmodial activity (IC50, µg mL−1) | Cytotoxicity (CC50, µg mL−1) | Selectivity index | ||
|---|---|---|---|---|---|---|
| S. aureus | E. coli | C. albicans | Pf-K1 | MRC-5SV2 | MRC-5SV2/PfK1 | |
| TE | >64.0 | >64.0 | >64.0 | 2.2 | 32.7 | 14.8 |
| TD | >64.0 | >64.0 | 6.8 | 1.81 | 3.6 | 2.0 |
| TM | >64.0 | >64.0 | 18.3 | 46.4 | >64.0 | >1.3 |
| TM1 | >64.0 | >64.0 | >64.0 | 34.5 | >64.0 | >1.8 |
| TM2 | >64.0 | >64.0 | >64.0 | 7.1 | >64.0 | 8.92 |
| TM3 | >64.0 | >64.0 | >64.0 | 1.3 | 31.0 | 23.4 |
| TM4 | >64.0 | >64.0 | >64.0 | 8.8 | 30.4 | 3.4 |
| TM5 | >64.0 | >64.0 | >64.0 | 19.6 | >64.0 | >3.25 |
| TM6 | >64.0 | >64.0 | >64.0 | 25.2 | 30.1 | 1.2 |
| TM7 | >64.0 | >64.0 | 3.6 | 23.8 | 28.5 | 1.19 |
| TM8 | >64.0 | >64.0 | 1.7 | 16.0 | 26.3 | 1.6 |
| TM9 | >64.0 | >64.0 | 1.6 | 18.8 | 29.4 | 1.5 |
| TM10 | >64.0 | >64.0 | 5.9 ± 4.7 | 4.3 ± 0.8 | >64.0 | >13.0 |
| TM11 | >64.0 | >64.0 | 25.2 | 18.66 | >64.0 | >3.4 |
| TM12 | >64.0 | >64.0 | 3.6 ± 3.0 | 37.07 | >64.0 | >1.7 |
| Chloroquine | 0.2 ± 0.1 µM | |||||
| Doxycycline | 0.3 ± 0.2 µM | 0.6 ± 0.3 µM | ||||
| Flucytosine | 0.70 ± 0.01 µM | |||||
| Tamoxifen | 10.0 ± 1.5 µM | |||||
3.2. Antiplasmodial activity of fractions and isolated compounds
The antiplasmodial activity of most of these compounds has been evaluated and the highest activity was obtained for pheophorbide-b methyl ester (1.0 ± 0.7 µM) (10), (1,2)-bis-nor-phytone (2.0 µM) (8), isophytol (4.0 µM) (7), pheophorbide-a methyl ester (2.8 ± 1.2 µM) (9), epicatechin-3-galloylester (5.5 ± 2.1) (19) and phytol (6.9 ± 2.4 µM) (6) (Table 1). Apart from myricetin-3-O-rhamnopyranoside (15) (14.3 µM), α-tocopherol (3) (13.5 µM), cycloart-24-en-3β-yl α-linolenate (2) (25.0 µM), which showed weak antiplasmodial activity, the other tested compounds displayed no activity.Pheophorbide derivatives are chlorophyll degradation products formed through the removal of the phytol side chain and the central magnesium ion from the tetrapyrrole macrocycle of chlorophyll a.44,45 A qualitative assessment of the structure–activity relationships within this class of chlorophyll-derived metabolites highlights the influence of functional group modifications and oxidative transformations on antiplasmodial activity. Pheophorbide-a methyl ester (9) exhibited moderate activity against P. falciparum (IC50 = 2.8 ± 1.2 µM), consistent with previously reported data.46 Its potency was slightly greater than that of pheophorbide-a (IC50 = 6.7 µg mL−1) and the C-13 hydroxylated analogue (IC50 = 5.1 µg mL−1), previously isolated from Mezoneuron benthamianum.44 Taken together, these data suggest that hydroxylation at C-13, in combination with esterification of the C-17 carboxyl group, enhance the antiplasmodial activity of pheophorbide-a. Furthermore, the presence of an aldehyde moiety at C-7, as observed in pheophorbide-b methyl ester (10), resulted in a marked improvement in antiplasmodial activity (IC50 = 1.4 ± 0.4 µM). However, this structural modification was accompanied by a significant increase in cytotoxicity (CC50 = 0.7 µM), indicating that the aldehyde moiety may also augment nonspecific reactivity toward mammalian cells. Although pheophorbide-b methyl ester (10) and pheophorbide-a methyl ester (9) have previously been isolated from various plant species, including Piper penangense, Clerodendrum spp., and Garuga pinnata, this study constitutes the first report of their presence in T. alnifolia.33,47 Pheophorbide derivatives have attracted considerable interest due to their application in malaria vector control and their ability to eliminate P. falciparum through photosensitization.48,49 In particular, synthetic lipophilic pheophorbide analogues have demonstrated potent antiplasmodial activity upon exposure to red light, highlighting the therapeutic potential of photodynamic activation. Moreover, earlier studies have proposed pheophorbide-mediated photosensitization as a promising strategy for the inactivation of transfusion-transmissible parasites and viruses in blood products, further underscoring the biomedical relevance of this compound class.50
Phytol (6) and two of its derivatives (isophytol and (1,2)-bis-nor-phytone) were among the most active compounds identified in this study. Phytol, a diterpenoid alcohol that forms the phytyl side chain of chlorophyll, is ubiquitous across photosynthetic organisms.51 Its antiplasmodial activity has been previously reported against the chloroquine-sensitive P. falciparum D10 strain (IC50 = 5.60 ± 0.18 µg mL−1), and bioassay-guided fractionation study has likewise identified phytol as a potent constituent against both chloroquine-resistant P. falciparum PoW (IC50 = 8.5 µM) and chloroquine-sensitive Dd2 (IC50 = 11.5 µM) strains.52 Although phytol and its derivatives isophytol (7) and (1,2)-bis-nor-phytone (8) have been reported in other plant species such as Calotropis procera,53 Glycine hispida,54 Pellia epiphylla, Microglossa pyrifolia,55 Brassica oleracea56 and Strobilanthes crispus,57 to the best of our knowledge, this is the first report of their occurrence in T. alnifolia, and that antiplasmodial activity was reported for isophytol (7) and (1,2)-bis-nor-phytone (8). Isophytol is generally considered as a relatively short lived intermediate in the degradation of the chlorophyll phytyl side chain and has been reported in the essential oil of several terrestrial plants.58
A clear structure–activity trend is evident within this series: phytol displayed moderate activity (IC50 = 6.9 ± 2.4 µM), whereas isophytol, differing only in the position of the double bond, demonstrated improved potency (IC50 = 4.0 µM). Further oxidative modification to generate (1,2)-bis-nor-phytone yielded the most active compound in this subset (IC50 = 2.0 µM). These results indicate that both the location of unsaturation and the degree of oxidative transformation along the diterpenoid chain play critical roles in modulating biological activity. In particular, progressive oxidation and carbon-chain truncation may enhance the antiplasmodial effects. Collectively, these results suggest that electrophilic modifications on the porphyrin core, together with oxidative tailoring of the phytol side chain are key drivers of antiplasmodial activity within this chlorophyll-derived compound class.
In order to determine the specificity of the compounds' antiplasmodial activity, their cytotoxicity on MRC-5SV2 cells was evaluated. Apart from pheophorbide-a methyl ester (SI 1.7), pheophorbide-b methyl ester (SI 1.6) and phytol (SI 1.3), all other compounds did not show any cytotoxicity up to the highest concentration tested (64 µM) (Table 2). Several studies have previously reported the cytotoxicity of pheophorbide derivatives. Indeed, several of the photosensitizers that were investigated for clinical photodynamic therapy (PDT) of cancer are constituents from plants. Some examples including degradation derivatives of chlorophylls, such as pheophorbides, which are present in green leaves and are well-studied photosensitizers in photodynamic cancer therapy.47 Moreover, most of the photosensitizers clinically approved or in clinical trials are derived from naturally occurring structures based on cyclic tetrapyrroles, such as chlorophyll-based compounds from higher plants.59
Among the isolated phenolic compounds, epicatechin-3-galloyl ester (19) (IC50 = 5.5 ± 2.1 µM) and myricetin-3-O-rhamnopyranoside (4.5 µM) were the most active. Activity across this series appears to be governed by the number of free hydroxyl groups on the B-ring and the presence of a galloyl moiety. It is noteworthy that epicatechin-3-galloyl ester (19) has previously been isolated from Camellia sinensis60 and Sclerocarya birrea.43 Its antiplasmodial activity has also been documented, with IC50 values of 10.8 µM against the chloroquine-sensitive P. falciparum 3D7 strain and 7.2 µM against the chloroquine-resistant FCR-1/FVO strain.60
Myricetin-3-O-rhamnopyranoside (15) has shown moderate antiplasmodial activity (IC50 14.3 µM), while quercetin-3-O-rhamnopyranoside (16) was completely inactive at the highest concentration tested (IC50 64.0 µM). These two compounds are structurally related. However, their difference is located in position C-5′ of their aglycone moiety where myricetin bears an additional substitution with a hydroxyl group. It appears that addition of the hydroxyl group in position C-5′, as in myricetin-3-O-rhamnoside, could have a strong influence on the antiplasmodial activity. In contrast to our result, a previous study carried out by Ganesh et al.61 revealed that quercetin-3-O-rhamnoside demonstrated weak activity against fresh P. falciparum field isolates from Bangladesh (IC50 20.4 ± 14.5 µM), P. falciparum chloroquine-sensitive 3D7 (IC50 35.4 ± 6.4) and P. falciparum chloroquine-resistant K1 (IC50 12 ± 27). Quercetin and myricetin can be found both in their free form (aglycon) and in their glycosylated form (heteroside) in several plant organs. Previous studies revealed that the antiplasmodial activity found for myricetin-3-O-rhamnopyranoside and quercetin-3-O-rhamnopyranoside were much lower than the activity of their corresponding aglycons myricetin (IC50 8.9 µM) and quercetin (IC50 12.9 µM).61,62 To date, quercetin and myricetin have been found to exhibit strong inhibitory activity against three important enzymes (FabG, FabZ, and FabI) involved in the fatty acid biosynthesis of P. falciparum and these flavonoids showed in vitro activity against chloroquine-sensitive (NF54) and chloroquine-resistant (K1) P. falciparum strains. Moreover, myricetin and quercetin inhibited the intraerythrocytic growth of the chloroquine-sensitive (3D7) and chloroquine-resistant (7G8) strains of P. falciparum.63
3.3. Antibacterial and antifungal activities of fractions and isolated compounds
The fractions and isolated compounds were also tested against S. aureus and C. albicans. The methanol fractions (TM) and its subfractions TM7 to TM12 were active against C. albicans (Table 1). Apart from gallic acid (14), which showed weak activity against S. aureus, none of the other compounds showed antimicrobial activity. The antibacterial activity of gallic acid against S. aureus in suspension (MIC 2 mg mL−1) and in biofilms (MIC 4 mg mL−1) has previously been reported.64 Furthermore, a study carried out by Chanwitheesuk et al.65 described the effectiveness of gallic acid against S. aureus (MIC 1250 µg mL−1). In contrast to our results, a previous study carried out by Motlhatlego et al.66 showed that myricetin-3-O-rhamnoside isolated from Newtonia buchananii demonstrated antibacterial activity against S. aureus (MIC 62.5 µg mL−1). Previous studies have suggested that phytol exhibits broad-spectrum antimicrobial effects.51 In particular, Inoue et al.67 reported strong antimicrobial activity of this compound against S. aureus (MIC 0.15 µg mL−1).4 Conclusion
This study provide experimental support for the traditional application of T. alnifolia in treating infectious diseases, particularly malaria. Guided by bioassay results, we isolated and characterized active constituents from the leaves, thereby substantiating its ethnopharmacological basis. The n-hexane, dichloromethane, and methanol extracts were active against P. falciparum, and subsequent fractionation afforded 19 metabolites, the majority of which are reported from this species for the first time. The most potent antiplasmodial activity was obtained for pheophorbide-b methyl ester (10), (1,2)-bis-nor-phytone (8), isophytol (7), pheophorbide-a methyl ester (9) and phytol (6). In contrast, compounds such as myricetin-3-O-rhamnopyranoside (15), α-tocopherol (3) and cycloart-24-en-3β-yl α-linolenate (2) showed comparatively lower activity. Notably, pheophorbide-b methyl ester (10), pheophorbide-a methyl ester (9) and phytol (6) also demonstrated significant cytotoxicity. These findings may contribute to a better understanding of the antiplasmodial properties observed in the leaf extract of T. alnifolia.Author contributions
M. A. Baldé: conceptualization, investigation, formal analysis, data analysis, writing original draft. M. S. Traoré: data analysis, sample identification, writing review & editing. M. S. T. Diallo: sample collection, data analysis. An Matheeussen: methodology, data curation. C. Aïssata: methodology, data curation, data visualization. P. Cos: methodology, antimicrobial assay, data validation. L. Maes: methodology, antiplasmodial assay, data validation. A. O. Baldé: sample collection and preparation of raw extract. M. K. Camara: data curation and data analysis. A. M. Balde: funding acquisition, supervision, writing review and editing. E. Tuenter: data analysis and visualization, writing review and editing. K. Foubert: conceptualization, supervision, writing review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article are included within the supplementary information (SI). Supplementary information: NMR data. See DOI: https://doi.org/10.1039/d5ra09498d.Acknowledgements
This work is derived from a chapter of the author's PhD dissertation completed at the University of Antwerp. The author gratefully acknowledges Prof. Dr Luc Pieters for his dedicated supervision, insightful guidance, and valuable support throughout the development of this research.References
- L. F. Nothias, M. Nothias-Esposito, R. Da Silva, M. Wang, I. Protsyuk, Z. Zhang, A. Sarvepalli, P. Leyssen, D. Touboul, J. Costa, J. Paolini, T. Alexandrov, M. Litaudon and P. C. Dorrestein, J. Nat. Prod., 2018, 81, 758–767 Search PubMed.
- U. Grienke, C. Mair, J. Kirchmair, M. Schmidtke and J. Rollinger, Planta Med., 2018, 84, 684–695 CrossRef CAS.
- L. K. Caesar, J. J. Kellogg, O. M. Kvalheim, R. A. Cech and N. B. Cech, Planta Med., 2018, 84, 721–728 CrossRef CAS.
- H. A. Junio, A. A. Sy-Cordero, K. A. Ettefagh, J. T. Burns, K. T. Micko, T. N. Graf, S. J. Richter, R. E. Cannon, N. H. Oberlies and N. B. Cech, J. Nat. Prod., 2011, 74, 1621–1629 CrossRef CAS.
- L. K. Caesar and N. B. Cech, Nat. Prod. Rep., 2019, 36, 869–888 RSC.
- J. W. Horn, Int. J. Plant Sci., 2009, 170, 794–813 Search PubMed.
- C. C. de Lima, R. P. Lyra Lemos and L. M. Conserva, J. Appl. Pharm. Sci., 2013, 3(9), 14–1118 Search PubMed.
- M. C. Y. Fomogne-Fodjo, D. T. Ndinteh, D. K. Olivier, P. Kempgens, S. van Vuuren and R. W. M. Krause, J. Ethnopharmacol., 2017, 195, 238–245 Search PubMed.
- T. O. Lawal, K. A. Jimoh and B. A. Adeniyi, Antimicrobial Potentials of Uvaria Afzelli Scott-Elliot and Tetracera Alnifolia Willd Crude Extracts on Selected Human Pathogens: An In-Vitro Study, J. Nat. Sci. Res., 2014, 4(6), 115–119 Search PubMed.
- C. C. Lima, R. P. L. Lemos and L. M. Conserva, J. Pharmacogn. Phytochem., 2014, 3, 181–204 Search PubMed.
- C. Seo, J. E. Lee, J. A. Lee, E. K. Ahn, J. S. Kang, C. W. Hyun and S. S. Hong, Chem. Nat. Compd., 2019, 55, 386–389 Search PubMed.
- M. A. Baldé, E. Tuenter, M. S. Traoré, A. Matheeussen, P. Cos, L. Maes, A. Camara, N. L. Haba, K. Gomou, M. S. T. Diallo, E. S. Baldé, L. Pieters, A. M. Balde and K. Foubert, J. Ethnopharmacol., 2020, 263, 113232 CrossRef.
- M. S. Traore, M. A. Baldé, M. S. T. Diallo, E. S. Baldé, S. Diané, A. Camara, A. Diallo, A. Balde, A. Keïta, S. M. Keita, K. Oularé, F. B. Magassouba, I. Diakité, A. Diallo, L. Pieters and A. M. Baldé, J. Ethnopharmacol., 2013, 150, 1145–1153 CrossRef CAS PubMed.
- F. B. Magassouba, A. Diallo, M. Kouyaté, F. Mara, O. Mara, O. Bangoura, A. Camara, S. Traoré, A. K. Diallo, M. Zaoro, K. Lamah, S. Diallo, G. Camara, S. Traoré, A. Kéita, M. K. Camara, R. Barry, S. Kéita, K. Oularé, M. S. Barry, M. Donzo, K. Camara, K. Toté, D. Vanden Berghe, J. Totté, L. Pieters, A. J. Vlietinck and A. M. Baldé, J. Ethnopharmacol., 2007, 114, 44–53 Search PubMed.
- A. K. Camara, E. S. Baldé, M. S. T. Diallo, M. K. Camara, T. V. Bah, M. Condé, A. Soumah, K. Kamano, I. Tietjen and A. M. Baldé, S. Afr. J. Bot., 2025, 181, 134–140 CrossRef CAS.
- N. Nsonde, S. Boumbal, T. Gouollaly, M. C. Makambila, J. M. Ouamba and A. Abena, Research Journal of Pharmacology and Pharmacodynamics, 2018, 1, 0001–0013 Search PubMed.
- O. O. Adeyemi, I. O. Ishola, E. T. Adesanya and D. O. Alohan, J. Basic Clin. Physiol. Pharmacol., 2019, 30, 173–184 Search PubMed.
- M. A. Baldé, E. Tuenter, M. S. Traoré, A. Matheeussen, P. Cos, L. Maes, A. Camara, N. L. Haba, K. Gomou, M. S. T. Diallo, E. S. Baldé, L. Pieters, A. M. Balde and K. Foubert, J. Ethnopharmacol., 2021, 267, 113624 CrossRef PubMed.
- M. A. Baldé, E. Tuenter, A. Matheeussen, M. S. Traoré, P. Cos, L. Maes, A. Camara, M. S. T. Diallo, E. S. Baldé, A. M. Balde, L. Pieters and K. Foubert, J. Ethnopharmacol., 2021, 267, 113624 CrossRef.
- P. Cos, A. J. Vlietinck, D. Vanden Berghe and L. Maes, J. Ethnopharmacol., 2006, 106, 290–302 CrossRef CAS PubMed.
- E. Tuenter, V. Exarchou, A. Baldé, P. Cos, L. Maes, S. Apers and L. Pieters, J. Nat. Prod., 2016, 79, 1746–1751 CrossRef CAS PubMed.
- L. Pogliani, M. Ceruti, G. Ricchiardi and D. Viterbo, Chem. Phys. Lipids, 1994, 70, 21–34 CrossRef CAS PubMed.
- D. Ngnokam, G. Massiot, J.-M. Nuzillard, J. D. Connolly, E. Tsamo and C. Morin, Phytochemistry, 1993, 34, 1603–1607 CrossRef CAS.
- M. Toyota, I. Omatsu, J. Braggins and Y. Asakawa, J. Oleo Sci., 2006, 55, 579–584 Search PubMed.
- S. Ohnmacht, R. West, R. Simionescu and J. Atkinson, Magn. Reson. Chem., 2008, 46, 287–294 Search PubMed.
- J. K. Baker and C. W. Myers, Pharm. Res., 1991, 8, 763–770 Search PubMed.
- J. S. S. Damsté, S. Schouten, N. H. Van Vliet, R. Huber and J. A. J. Geenevasen, Tetrahedron Lett., 1997, 38, 6881–6884 CrossRef.
- P. J. Proteau, J. Nat. Prod., 1998, 61, 841–843 Search PubMed.
- V. U. Ahmad and M. S. Ali, Phytochemistry, 1991, 30, 4172–4174 CrossRef CAS.
- U. Langenbahn, G. Burkhardt and H. Becker, Phytochemistry, 1993, 33, 1173–1179 Search PubMed.
- H. Duan, Y. Takaishi, H. Momota, Y. Ohmoto and T. Taki, Phytochemistry, 2002, 59, 85–90 CrossRef CAS PubMed.
- S. L. Schwikkard, D. A. Mulholland and A. Hutchings, Phytochemistry, 1998, 49, 2391–2394 CrossRef CAS.
- F. Adzhar Kamarulzaman, K. Shaari, A. Siong Hock Ho, N. Haji Lajis, S. Hwang Teo and H. Boon Lee, Chem. Biodiversity, 2011, 8, 494–502 CrossRef PubMed.
- S. Faizi, M. Ali, R. Saleem, I. Irfanullah and S. Bibi, Magn. Reson. Chem., 2001, 39, 399–405 CrossRef CAS.
- A. Sakushima, M. C. O. Kun and T. Maoka, Phytochemistry, 1995, 40, 257–261 CrossRef CAS.
- R. D. Bennett, S. M. Norman and V. P. Maier, Phytochemistry, 1981, 20, 2343–2344 CrossRef CAS.
- S. R. Abrams, M. J. T. Reaney, G. D. Abrams, T. Mazurek, A. C. Shaw and L. V. Gusta, Phytochemistry, 1989, 28, 2885–2889 CrossRef CAS.
- Y. Zhang, D. L. DeWitt, S. Murugesan and M. G. Nair, Chem. Biodiversity, 2004, 1, 426–441 CrossRef CAS.
- I. I. Mahmoud, M. S. A. Marzouk, F. A. Moharram, M. R. El-Gindi and A. M. K. Hassan, Phytochemistry, 2001, 58, 1239–1244 CrossRef CAS PubMed.
- T. Fukunaga, K. Nishiya, I. Kajikawa, Y. Watanabe, N. Suzuki and K. Takeya, Chem. Pharm. Bull., 1988, 1180–1184 CrossRef CAS.
- A. Braca, A. R. Bilia, J. Mendez and I. Morelli, Fitoterapia, 2001, 72, 182–185 CrossRef CAS PubMed.
- L. Y. Foo, Y. Lu, A. L. Molan, D. R. Woodfield and W. C. McNabb, Phytochemistry, 2000, 54, 539–548 CrossRef CAS PubMed.
- J. Galvez Peralta, A. Zarzuelo, R. Busson, C. Cobbaert and P. De Witte, Planta Med., 1992, 58, 174–175 CrossRef CAS PubMed.
- O. Jansen, A. T. Tchinda, J. Loua, V. Esters, E. Cieckiewicz, A. Ledoux, P. D. Toukam, L. Angenot, M. Tits, A. M. Balde and M. Frédérich, J. Ethnopharmacol., 2017, 203, 20–26 CrossRef CAS.
- A. Pérez-Gálvez, M. Roca and Atta-ur-Rahman, in Studies in Natural Products Chemistry, Elsevier, 2017, vol. 52, pp. 159–191 Search PubMed.
- W. M. Afifi, E. A. Ragab, A.-E. I. Mohammed and A. A. El-Hela, Chemical constituents and biological activity of Ficus platypoda (Miq.) leaves, J. Biomed. Pharm. Res., 2014, 3, 21–36 Search PubMed.
- P. Wongsinkongman, A. Brossi, H. K. Wang, K. F. Bastow and K. H. Lee, Bioorg. Med. Chem., 2002, 10, 583–591 CrossRef CAS.
- G. Jori and O. Coppellotti, The Royal Society of Chemistry, 2016, pp. 573–590, 10.1039/9781782626824-00573.
- R. Vanderesse, L. Colombeau, C. Frochot and S. Acherar, in Current Topics in Malaria, InTech, 2016 Search PubMed.
- P. Grellier, R. Santus, E. Mouray, V. Agmon, J. C. Mazière, D. Rigomier, A. Dagan, S. Gatt and J. Schrével, Vox Sang., 1997, 72, 211–220 CrossRef CAS PubMed.
- M. T. Islam, E. S. Ali, S. J. Uddin, S. Shaw, M. A. Islam, M. I. Ahmed, M. Chandra Shill, U. K. Karmakar, N. S. Yarla, I. N. Khan, M. M. Billah, M. D. Pieczynska, G. Zengin, C. Malainer, F. Nicoletti, D. Gulei, I. Berindan-Neagoe, A. Apostolov, M. Banach, A. W. K. Yeung, A. El-Demerdash, J. Xiao, P. Dey, S. Yele, A. Jóźwik, N. Strzałkowska, J. Marchewka, K. R. R. Rengasamy, J. Horbańczuk, M. A. Kamal, M. S. Mubarak, S. K. Mishra, J. A. Shilpi and A. G. Atanasov, Food Chem. Toxicol., 2018, 121, 82–94 CrossRef CAS PubMed.
- M. H. Grace, C. Lategan, R. Graziose, P. J. Smith, I. Raskin and M. A. Lila, Nat. Prod. Commun., 2012, 7, 1263–1266 CrossRef CAS PubMed.
- D. Kubmarawa and I. A. Ogunwande, J. Essent. Oil Bear. Plants, 2008, 11, 75–78 CrossRef CAS.
- V. Kyslychenko, U. Karpiuk, I. Diakonova and M. S. Abu-Darwish, Adv. Environ. Biol., 2010, 4, 490–494 CAS.
- I. Köhler, K. Jenett-Siems, C. Kraft, K. Siems, D. Abbiw, U. Bienzle and E. Eich, Z. Naturforsch., C: J. Biosci., 2002, 57, 1022–1027 CrossRef.
- M. Jayalakshmi, V. Vanitha and R. Sangeetha, Asian J. Pharm. Clin. Res., 2018, 11, 133–136 CAS.
- H. Nurraihana and N. A. Norfarizan-Hanoon, Int. Food Res. J., 2013, 20, 2045–2056 Search PubMed.
- J. F. Rontani and M. A. Galeron, Org. Geochem., 2016, 97, 35–40 CrossRef CAS.
- Y. V. Tang, S. M. Phang, W. L. Chu, A. Ho, S. H. Teo and H. B. Lee, J. Appl. Phycol., 2012, 24, 783–790 CrossRef CAS.
- A. R. Sannella, L. Messori, A. Casini, F. Francesco Vincieri, A. R. Bilia, G. Majori and C. Severini, Biochem. Biophys. Res. Commun., 2007, 353, 177–181 CrossRef CAS.
- D. Ganesh, H.-P. Fuehrer, P. Starzengrüber, P. Swoboda, W. A. Khan, J. A. B. Reismann, M. S. K. Mueller, P. Chiba and H. Noedl, Parasitol. Res., 2012, 110, 2289–2295 CrossRef PubMed.
- D. Tasdemir, G. Lack, R. Brun, P. Rüedi, L. Scapozza and R. Perozzo, J. Med. Chem., 2006, 49, 3345–3353 CrossRef CAS PubMed.
- H. Jin, Z. Xu, K. Cui, T. Zhang, W. Lu and J. Huang, Fitoterapia, 2014, 94, 55–61 CrossRef CAS.
- M. Liu, X. Wu, J. Li, L. Liu, R. Zhang, D. Shao and X. Du, Food Control, 2017, 73, 613–618 CrossRef CAS.
- A. Chanwitheesuk, A. Teerawutgulrag, J. D. Kilburn and N. Rakariyatham, Food Chem., 2007, 100, 1044–1048 CrossRef CAS.
- K. E. Motlhatlego, M. A. Abdalla, C. M. Leonard, J. N. Eloff and L. J. McGaw, BMC Complementary Med. Ther., 2020, 20, 358 CrossRef CAS.
- Y. Inoue, T. Hada, A. Shiraishi, K. Hirose, H. Hamashima and S. Kobayashi, Antimicrob. Agents Chemother., 2005, 49, 1770–1774 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |