 Open Access Article
Open Access ArticleCPB@PLH NPs accelerate wound healing by antibacterial and anti-inflammatory effects
Yilong Li†
ac,
Aidi Tong†a,
Pengxiang Xu†a,
Li Chang ab,
Jialong Fand,
Zhoubing Shana,
Chunyi Tong
ab,
Jialong Fand,
Zhoubing Shana,
Chunyi Tong *a and
Bin Liu
*a and
Bin Liu *a
*a
aCollege of Biology, Hunan University, Changsha, 410082, PR China. E-mail: sw_tcy@hnu.edu.cn; binliu2001@hotmail.com
bInstitute of Bast Fiber Crop, Chinese Academy of Agriculture Sciences, Changsha, 410304, China
cHunan Zhongsheng Xingxiu Biotech Co., Ltd, Changsha, China
dHunan Provincial Key Laboratory of the Research and Development of Novel Pharmaceutical Preparations, Changsha Medical University, Changsha, 410219, China
First published on 2nd March 2026
Abstract
Diabetic wounds are a severe complication of diabetes, with their non-healing nature closely associated with infection, oxidative stress, and the inflammatory microenvironment. To address the limited therapeutic efficacy of conventional antibiotics, which often lead to drug resistance and offer functionality, this study constructed a multifunctional composite nano-platform (CPB@PLH NPs) based on carboxylated Prussian blue nanoparticles (CPB NPs) and a novel antimicrobial peptide of PLH054001130. This platform integrates the broad-spectrum antimicrobial and biofilm-disrupting capabilities of PLH with the antioxidant and anti-inflammatory functions of CPB NPs, enabling synergistic “anti-bacteria–anti-inflammation” therapy. In vitro assay demonstrated the over 99% antibacterial rate of CPB@PLH NPs against methicillin-resistant Staphylococcus aureus (MRSA), effective reactive oxygen species (ROS) scavenging, inflammation suppression, and endothelial cell migration acceleration. In a mouse model of MRSA-infected wounds in the context of streptozotocin (STZ)-induced diabetes, CPB@PLH NPs significantly promoted wound closure by enhancing collagen deposition, stimulating angiogenesis, and modulating the expression of inflammatory cytokines. This study presents an innovative approach for comprehensively treating chronically infected diabetic wounds, offering excellent biocompatibility.
1 Introduction
As one of the most severe complications of diabetes, diabetic wounds are chronic and refractory injuries resulting from diabetic neuropathy, vascular disease, and bacterial infections.1,2 The pathogenesis of diabetic wounds involves the combined effects of peripheral sensory neuropathy and microvascular dysfunction, leading to full-thickness skin defects. This triggers local tissue ischemia and metabolic disorders, ultimately manifesting as ulcer formation, secondary infection, and even tissue necrosis.3 The physiological healing of wounds follows a well-established sequence of four phases: hemostasis, inflammation, proliferation, and remodeling. However, in diabetic wounds, the persistent hyperglycemia and locally hypoxic pathological microenvironment significantly impair tissue repair capacity and markedly increase the risk of wound infection.4 On this basis, secondary bacterial infections trigger excessive recruitment of neutrophils and macrophages. During pathogen clearance, these immune cells produce large quantities of ROS, leading to oxidative stress imbalance. This exacerbates cellular damage and ultimately results in a pathologically prolonged inflammatory response, hindering the healing process.5,6 The standard clinical treatment protocol for diabetic wounds usually involves removing necrotic tissue, performing skin grafts, applying topical antibiotics, using negative pressure wound therapy and hyperbaric oxygen therapy, and applying wound dressings.7 Among these, antibiotics are crucial for controlling infections, but their efficacy has certain limitations. For example, over 60% of Gram-positive bacteria exhibit resistance to penicillin and erythromycin. Additionally, approximately 31% of diabetic wound-associated strains possess the ability to form biofilms.8 The extracellular polymers (EPS) within these biofilms physically impede antibiotic penetration, enhancing bacterial survival by nearly a hundredfold and constituting one of the primary obstacles to clinical cure.9,10Antimicrobial peptides (AMPs) are molecules that act as natural immune defenses found in all living things. Typically composed of 12 to 50 amino acids, they exhibit broad-spectrum antimicrobial activity alongside multiple functions, including immune modulation and wound healing promotion.11,12 AMPs possess multi-target bactericidal mechanisms, including disruption of bacterial membranes leading to intracellular leakage; intracellular entry to inhibit nucleic acid and protein synthesis and interfere with metabolic pathways; degradation of extracellular polysaccharides; and suppression of quorum sensing to disintegrate biofilm structures.13,14 However, AMPs are rapidly degraded by proteases in the complex wound microenvironment, limiting their clinical application.15 Recent studies indicated that nanotechnology strategies not only significantly enhance the stability of AMPs but also endow them with the ability to address multiple pathological pathways in diabetic wounds, such as oxidative stress and inflammatory dysregulation, thereby overcoming their functional limitations.16
Prussian blue nanoparticles (PB NPs), as an FDA-approved metal–organic framework (MOF) material with high stability and excellent biocompatibility,17,18 serve not only as nanocarriers for drug delivery but also possess multiple bioactivities such as antioxidant and anti-inflammatory properties. These characteristics confer unique advantages for synergistically promoting wound healing.19 Specifically, PB NPs mitigate oxidative stress damage in tissues by scavenging excess ROS.20 In addition to suppressing LPS-induced inflammatory responses, PB NPs promote the repolarization of macrophages from the pro-inflammatory M1 phenotype toward the anti-inflammatory M2 phenotype, thereby effectively improving the inflammatory microenvironment at wound sites.21,22
In this study, we screened a novel antimicrobial peptide, PLH054001130 (amino acid sequence: GKWGFYTKVFRLKKWIKKVICTFGC, molecular weight: 3034.78 Da), and loaded it onto carboxylated Prussian blue nanoparticles (CPB NPs) via electrostatic adsorption, thereby constructing a novel antimicrobial nanomedicine. Within this composite system, PLH exerts potent antibacterial activity while CPB NPs provide anti-inflammatory and antioxidant support. Their synergistic interaction achieves dual-function coordinated therapy—combining antibacterial and anti-inflammatory effects for MRSA-infected wounds in diabetic mice (Scheme 1).
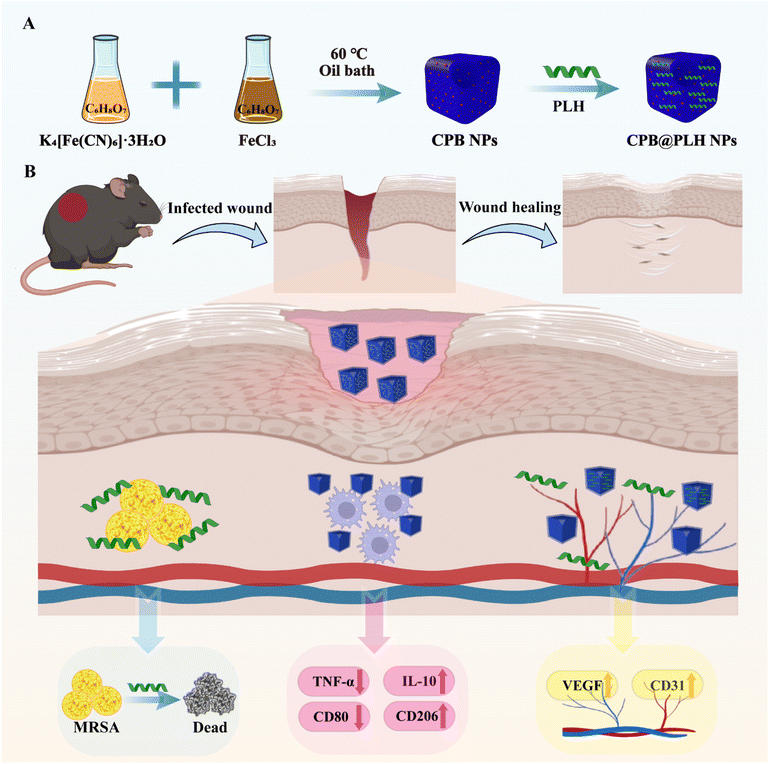 | ||
| Scheme 1 (A) Schematic diagram of CPB@PLH NPs synthesis. (B) The mechanism of wound healing promotion by CPB@PLH NPs. | ||
2 Experimental methods
2.1 Chemicals and reagents
K4[Fe(CN)6]·3H2O was supplied by Fuchen Chemical Reagent Co., Ltd. Citric acid was purchased from Shanghai Hushi Co., Ltd. FeCl3·6H2O was sourced from Aladdin Biochemical Technology Co., Ltd. The antimicrobial peptide PLH054001130 (sequence: GKWGFYTKVFRLKKWIKKVICTFGC) was synthesized by Hunan Zhongsheng Xingxiu Biotech Co., Ltd. The DAMO/PI assay kit was purchased from Biyuntian Biotechnology Research Institute. The DCFH-DA probe was acquired from Solarbio Science & Technology Co., Ltd. The H&E and Masson kits were obtained from Biosharp Co., Ltd and Beijing Solarbio Co., Ltd, respectively. TNF-α, IL-10, and CD31 antibodies were obtained from Wuhan Sanying Biotechnology Co., Ltd. The VEGF antibody was purchased from Beyotime Biotechnology Institute.2.2 Strains, cells, and laboratory animals
The MRSA strain was acquired from Beijing Beina Biotechnology and grown in LB liquid medium under incubation at 37 °C with 200 rpm shaking until reaching the logarithmic growth phase for subsequent experiments. We received the RAW264.7 and HUVEC cell lines from the Xiangya Laboratory of Central South University. All cultures were sustained using high-glucose DMEM. The medium contained 10% fetal bovine serum (FBS, SiJiQing Co., Ltd) and 1% (m/v) penicillin/streptomycin, and cells were grown at 37 °C under 5% CO2. We obtained healthy, 6–8-week-old male C57BL/6 mice from Hunan SJA Laboratory Animal Co., Ltd. All animal procedures adhered to the principles of the Declaration of Helsinki and were reviewed and approved by the Animal Ethics Committee of Hunan University (Approval No. SYXK-2023-0010).2.3 Preparation and characterization of CPB@PLH NPs
CPB NPs were synthesized according to a previously reported method.17 Briefly, a citric acid solution (200 mL) containing K4[Fe(CN)6]·3H2O (Solution A) and another citric acid solution (200 mL) containing FeCl3·6H2O (Solution B) were preheated to 60 °C separately. Solution B was then added to Solution A, with stirring continuing throughout, at a temperature of 60 °C for a duration of 30 min. After completion, the resulting bright blue solution was cooled to room temperature with continuous stirring. The product was finally subjected to a centrifuge process at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for a duration of 30 min. This resulted in the collection of a precipitate, which was then utilised to obtain CPB NPs.
000 rpm for a duration of 30 min. This resulted in the collection of a precipitate, which was then utilised to obtain CPB NPs.
The loading capacity of CPB NPs for PLH was evaluated using three mass feeding ratios (CPB NPs![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PLH = 1
PLH = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, 1
2, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). The mixtures were subjected to continuous stirring at 800 rpm over a period of 12 h to achieve complete binding. Subsequently, the solutions were centrifuged at 15
4). The mixtures were subjected to continuous stirring at 800 rpm over a period of 12 h to achieve complete binding. Subsequently, the solutions were centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 10 min to collect the composite nanoparticles to obtain CPB@PLH NPs.
000 rpm for 10 min to collect the composite nanoparticles to obtain CPB@PLH NPs.
For the preparation of CPB@PLH@Gel, a Tris buffer solution (0.5 mL, 10 mM, pH 7.4) composed of 1.3% (v/v) glycerol was preheated to 56 °C, Then, 18.5 mg of hydroxyethyl cellulose (HEC) was added. The mixture was stirred until a homogeneous gel matrix formed. Subsequently, the as-prepared CPB@PLH NPs were dispersed in the same Tris buffer and then uniformly mixed with the HEC gel, ultimately yielding the CPB@PLH@Gel composite gel formulation.
Transmission electron microscopy (TEM) (JEOL, operating at 200 kV) was used to characterize the morphologies of CPB NPs and CPB@PLH NPs. A Zetasizer Nano ZS (Malvern Instruments) was employed to determine the size distribution and zeta potential of the nanoparticles. UV-Vis absorption spectra of PLH, CPB NPs, and CPB@PLH NPs were recorded using a spectrophotometer from Beckman Coulter Inc. (USA).
2.4 In vitro anti-MRSA activity of CPB@PLH NPs
Minimum bactericidal concentration (MBC) determination: to evaluate the antibacterial activity of CPB@PLH NPs, the MBC was determined using the microbroth dilution method. Different concentrations of PLH and CPB@PLH NPs were co-incubated with approximately 105 CFU mL−1 of MRSA and E. coli bacterial suspension for 24 h. Each bacterial suspension was plated onto LB agar and then incubated at 37 °C for 24 h, after which colonies were enumerated.Scanning electron microscopy (SEM) morphological observation: the MRSA bacterial suspensions treated with PBS, PLH, CPB NPs, and CPB@PLH NPs were collected and centrifuged at 6000 rpm for 5 min. The bacterial pellets were fixed with 2.5% glutaraldehyde solution at 4 °C for 4 h. Subsequently, the samples were dehydrated through a graded ethanol series, with each step lasting 10 min. The bacterial morphology was then observed and imaged using a scanning electron microscope.
MRSA crystal violet biofilm assay: to form mature biofilms, MRSA was seeded into 96-well plates at a density of 5 × 108 CFU mL−1 and cultured at 37 °C for 72 h. After supernatant removal, the MRSA biofilms were challenged with PBS, PLH, CPB NPs, or CPB@PLH NPs and incubated at 37 °C for 24 h. The biofilms were then fixed with 100 µL methanol (15 min), stained with 100 µL of 0.1% crystal violet (30 min), and imaged. Finally, the stained biofilms were dissolved in absolute ethanol, and the absorbance at 590 nm was measured with a microplate reader for quantification.
2.5 Cytotoxicity evaluation of CPB@PLH NPs
The biosafety of the materials was evaluated by assessing their cytotoxicity on RAW264.7 cells and HUVECs using the MTT assay. Briefly, cells were seeded in 96-well plates at a density of 1 × 104 cells per well and cultured for 24 h at 37 °C. After this initial culture, they were exposed to various sample formulations for an additional 24 h, after which cell viability was measured.2.6 ROS scavenging capacity of CPB@PLH NPs in RAW264.7 cells
To ensure cell adherence, RAW264.7 cells were plated onto glass coverslips placed in 24-well plates and cultured for 24 h. The cells were then treated with different sample solutions and co-incubated for 2 h, followed by stimulation with 500 ng per mL LPS for 4 h to induce oxidative stress. Subsequently, the cells were incubated with 10 µM DCFH-DA probe at 37 °C for 30 min to protect from light. After washing with PBS, the fluorescence intensity reflecting intracellular ROS levels was observed and recorded using fluorescence microscopy.2.7 Analysis of HUVEC migration capacity via the scratch assay
The cells were plated in 12-well plates at 2 × 104 cells per well and grown for 24 h until a confluent monolayer formed. Uniform scratches were created in the cell monolayer using a sterile pipette tip, followed by the addition of culture media containing different sample components. The same locations were imaged under an inverted microscope at 0, 24, and 48 h, and the wound closure rate was analyzed using ImageJ software to evaluate cell migration ability.2.8 Optimized screening of bacterial load in a wound infection model of diabetic mice
The male C57BL/6J mice (8 weeks old, 20 g) under specific pathogen-free (SPF) conditions were selected and acclimatized under standard housing conditions. When their body weight reached 25 g, an infectious wound model was established. After anesthesia, the dorsal hair of the mice was shaved, and a full-thickness skin defect wound with a diameter of 10 mm was created. The wounds were inoculated with different concentrations and types of bacteria. Beginning the following day, the wound area was measured regularly, and wound tissue samples were collected for quantitative bacterial culture to assess the degree of infection and healing progression, thereby determining the appropriate bacterial load for subsequent experiments.2.9 PLH in vivo therapeutic concentration screening
Specific pathogen-free (SPF) male C57BL/6J mice (8 weeks old, initial weight 20 g) were fed a high-fat, high-sucrose diet for 2 weeks until their body weight reached approximately 25 g. Diabetes was then induced by intraperitoneal injection of streptozotocin (STZ, 60 mg kg−1) for five consecutive days. Fasting blood glucose levels were monitored regularly, and diabetes model establishment was confirmed when blood glucose levels stabilized at ≥20 mmol L−1. Subsequently, the infectious wound phase was initiated. After anesthesia, a 10 mm full-thickness wound was created on the dorsal skin of each mouse.19 Except for the blank control group, all other groups were inoculated with 1 × 108 CFU mL−1 of MRSA bacterial suspension.23 Following successful infection, mice were treated with different concentrations of PLH via topical application once daily. Wound healing was monitored by capturing standardized digital photographs with a reference scale at defined intervals. The wound area was quantified through digital planimetry using ImageJ software, and the percentage of wound closure was calculated. Concurrently, and quantitative bacterial analysis of wound tissues was performed to evaluate the therapeutic efficacy of PLH and determine its optimal concentration.242.10 Antimicrobial and wound-healing effects of CPB@PLH@Gel on MRSA wound infections in diabetic mice
Diabetic models were established in SPF male C57BL/6J mice (8 weeks, 20 g) by a high-fat/sucrose diet combined with five consecutive daily intraperitoneal injections of STZ (60 mg kg−1). Mice with sustained blood glucose levels ≥20 mmol L−1 were included in the study. To establish the infection model, a 10 mm full-thickness wound was made on the dorsal skin under anesthesia and inoculated with 1 × 108 CFU per mL MRSA. After successful infection, the mice were randomly assigned to different groups that received specific daily treatments. Wound healing was monitored regularly, and quantitative bacterial analysis was performed. Nine days after the start of treatment, skin tissue samples were collected, fixed in 4% paraformaldehyde for 24 h, and then dehydrated using a graded ethanol series. The samples were then cleared in xylene and embedded in paraffin wax for sectioning. The sections were then stained with H&E and Masson's trichrome to evaluate tissue morphology and collagen deposition. Meanwhile, immunohistochemistry was performed to detect VEGF and CD31 expression for assessing angiogenesis, while TNF-α and IL-10 staining were used to analyze the extent of inflammatory cell infiltration. Additionally, the polarization of macrophages within the wound was assessed by immunohistochemical staining for the M1 marker CD80 and the M2 marker CD206.2.11 Statistical analysis
Experiments were carried out with at least three independent replicates. Data are presented as mean ± standard deviation and were analyzed by one-way ANOVA followed by Tukey's post hoc test, with statistical significance denoted as *P < 0.05, **P < 0.01, and ***P < 0.001.3 Results
3.1 Characterization of CPB@PLH NPs
CPB NPs were synthesized via a hydrothermal method.17 Subsequently, PLH and CPB NPs were dissolved in a solvent, stirred uniformly, and centrifuged to obtain CPB@PLH NPs. To further optimize the loading conditions, the encapsulation efficiency and drug loading capacity were investigated at different mass ratios (CPB![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PLH = 1
PLH = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, 1
2, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). The hydrodynamic diameter and surface potential were measured using dynamic light scattering (DLS) and zeta potential analysis, respectively, to systematically characterize the physicochemical properties. As shown in Fig. 1A, dynamic light scattering (DLS) analysis revealed the hydrodynamic diameters of the nanoparticles. The pristine CPB NPs exhibited a size of ∼220 nm. After coating with PLH at different mass ratios, the CPB@PLH NPs showed sizes of ∼192 nm (1
4). The hydrodynamic diameter and surface potential were measured using dynamic light scattering (DLS) and zeta potential analysis, respectively, to systematically characterize the physicochemical properties. As shown in Fig. 1A, dynamic light scattering (DLS) analysis revealed the hydrodynamic diameters of the nanoparticles. The pristine CPB NPs exhibited a size of ∼220 nm. After coating with PLH at different mass ratios, the CPB@PLH NPs showed sizes of ∼192 nm (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), ∼190 nm (1
1), ∼190 nm (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2), and ∼160 nm (1
2), and ∼160 nm (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). Notably, at the CPB
4). Notably, at the CPB![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PLH ratio of 1
PLH ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, the particle size reached a minimum of 160 nm, and the Zeta potential was optimized to +15.63 mV (Fig. 1B), indicating successful coating of PLH and effective reversal of surface charge. The results showed that as the PLH ratios increased, the encapsulation efficiency decreased while the drug loading capacity increased. The absolute mass of PLH loaded onto a fixed amount of CPB rose from 57.9 µg (1
4, the particle size reached a minimum of 160 nm, and the Zeta potential was optimized to +15.63 mV (Fig. 1B), indicating successful coating of PLH and effective reversal of surface charge. The results showed that as the PLH ratios increased, the encapsulation efficiency decreased while the drug loading capacity increased. The absolute mass of PLH loaded onto a fixed amount of CPB rose from 57.9 µg (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to 87.3 µg (1
1) to 87.3 µg (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) and 148.8 µg (1
2) and 148.8 µg (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4). Consequently, the drug loading capacity reached a maximum of 74.40% at the 1
4). Consequently, the drug loading capacity reached a maximum of 74.40% at the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 ratio, which was therefore selected as the optimal formulation for all subsequent experiments (Fig. 1C). Transmission electron microscopy images revealed that CPB NPs exhibited a regular cubic morphology with a size of 50 ± 5 nm. After PLH adsorption, the size of CPB@PLH NPs increased to 80 ± 10 nm, with uniform particle morphology and good dispersity (Fig. 1D and E). UV-Vis absorption spectra showed that CPB NPs exhibited a characteristic absorption peak at 710 nm, while PLH showed a characteristic peak at 280 nm. For CPB@PLH NPs, the absorption at both 280 nm and 710 nm was significantly enhanced, confirming the successful adsorption of PLH onto the surface of CPB NPs (Fig. 1F).
4 ratio, which was therefore selected as the optimal formulation for all subsequent experiments (Fig. 1C). Transmission electron microscopy images revealed that CPB NPs exhibited a regular cubic morphology with a size of 50 ± 5 nm. After PLH adsorption, the size of CPB@PLH NPs increased to 80 ± 10 nm, with uniform particle morphology and good dispersity (Fig. 1D and E). UV-Vis absorption spectra showed that CPB NPs exhibited a characteristic absorption peak at 710 nm, while PLH showed a characteristic peak at 280 nm. For CPB@PLH NPs, the absorption at both 280 nm and 710 nm was significantly enhanced, confirming the successful adsorption of PLH onto the surface of CPB NPs (Fig. 1F).
3.2 Multi-enzyme activity of CPB@PLH NPs
To verify the catalase-like activity of CPB@PLH NPs, an oxygen production experiment was conducted. After the addition of hydrogen peroxide, evident bubble formation was observed in both the CPB NPs and CPB@PLH NPs solutions, whereas no such phenomenon occurred in the control and PLH groups. This confirmed that the CPB NPs and CPB@PLH NPs possess catalase-like activity and can catalyze the decomposition of hydrogen peroxide to produce oxygen (Fig. 2A). As shown in Fig. 2B, after the reaction of CPB@PLH NPs with the TMB substrate, the solution exhibited a distinct blue color change, visually indicating the successful oxidation of TMB. UV-Vis spectroscopy revealed a characteristic absorption peak near 650 nm, consistent with the chromogenic reaction of oxidized TMB, demonstrating that the nanomaterial can mimic peroxidase activity by catalyzing the colorimetric reaction of TMB (Fig. 2C). Furthermore, the DPPH radical scavenging assay showed that 25 and 50 µg per mL CPB@PLH NPs caused noticeable fading of the purple color in the solution (Fig. 2D). Fig. 2E indicated a decrease in absorbance at 510 nm, and the OD 510 values significantly decreased with increasing concentration, confirming that CPB@PLH NPs possess efficient and concentration-dependent DPPH radical scavenging ability (Fig. 2F).3.3 In vitro antibacterial efficacy of CPB@PLH NPs
We employed the dilution spot assay to determine the MBC of CPB@PLH NPs against MRSA and E. coli. As shown in Fig. 3A and B, PLH and CPB@PLH NPs exhibited significant antibacterial activity, achieving over 99% bactericidal efficiency at 6 µg mL−1 of PLH against MRSA and 4 µg mL−1 against E. coli (Fig. S1A and B). This result demonstrated that CPB NPs did not significantly compromise the inherent antibacterial activity of PLH. In addition, we assessed the efficacy under oxidative stress and against enzymatic degradation. The antibacterial efficacy of PLH and CPB@PLH NPs against MRSA remained high, achieving over 99% bacterial reduction even in the presence of H2O2 (Fig. 3C and D). More importantly, CPB@PLH NPs demonstrated strong anti-trypsin hydrolysis, as evidenced by the preserved bactericidal effect (Fig. 3E and S2). These findings indicate that the CPB NPs can protect PLH digestion to enhance its functional resilience in the complex microenvironment in vivo. We utilized DMAO/PI double staining to further validate the bactericidal effect. Fluorescence images revealed that the Control and CPB NPs groups exhibited only strong green fluorescence with a weak red signal. In contrast, the PLH and CPB@PLH NPs groups showed a significant increase in red fluorescence, indicating a markedly higher bacterial mortality rate (Fig. 3F). Consistent with this, intracellular ROS levels in MRSA were significantly elevated following treatment with both PLH and CPB@PLH NPs (Fig. 3G), suggesting ROS generation is part of the antimicrobial mechanism. Furthermore, scanning electron microscopy (SEM) images indicated that MRSA cells appeared intact and spherical with smooth surfaces in the PBS control group, the CPB NPs group showed some particles attached to the bacterial surface. In contrast, PLH treatment caused obvious damage to the bacterial membrane structure, including rupture, pore formation, and collapse. In the CPB@PLH NPs group, nanoparticles were observed accumulating on the bacterial surface, leading to membrane folding, deformation, and disruption (Fig. 3H), providing structural evidence for the membrane-damaging effects. Biofilm is a complex structure to protects the microenvironment for bacteria and severely impedes the wound healing process.25 Crystal violet staining assay indicated a significant reduction of the stained area in the PLH and CPB@PLH NPs treatment groups. Statistical analysis revealed biofilm clearance rates of 68.65% and 64.61% for PLH and CPB@PLH NPs, respectively (Fig. 3I and S3), demonstrating the potent anti-biofilm activity.3.4 The effects of CPB@PLH NPs on cell migration and proliferation capabilities
HUVECs play a critical role in angiogenesis and contribute to improved oxygen and nutrient supply in the wound microenvironment.26 RAW264.7 cells, as central regulators of inflammation and repair processes, participate in multiple stages of wound healing.27 Therefore, evaluating the activity of CPB@PLH NPs on the two cell types is of significant importance. In contrast to the control group, Fig. 4A and B demonstrated that cell viability decreased after treatment with PLH alone, while the CPB NPs group showed an increase in cell number to approximately 112%, indicating a degree of biocompatibility. The CPB@PLH NPs group did not induce significant cytotoxicity, suggesting that CPB NPs effectively mitigated the potential damage caused by PLH. Furthermore, high levels of ROS can impede the repolarization of M1 macrophages toward a reparative phenotype,28 the ability to scavenge ROS serves as a crucial indicator for assessing the anti-inflammatory potential of materials. DCFH-DA probe staining indicated strong green fluorescence signal, which reflected high ROS levels in the LPS-induced M1-type RAW264.7 model. However, CPB NPs and CPB@PLH NPs treatment markedly decreased the fluorescence signal, which reflected the reduction of ROS levels triggered by LPS (Fig. 4C). Subsequently, using a co-culture system to further evaluate the regulatory effect of CPB@PLH NPs on the migration ability of HUVECs, it was found that both CPB NPs and CPB@PLH NPs significantly promoted the migration of HUVECs. The migration rates reached 72.0% and 73.8% after 12 h of culture, respectively, and further increased to 83.1% and 87.3% after 24 h (Fig. 4D–F). These results suggested the ability of CPB@PLH NPs for enhancing wound healing.3.5 Screening of bacterial load in wound-infected mice
Using normal mice as subjects, full-thickness skin defect wounds were used to screen for an appropriate bacterial load to significantly delay the healing process. Fig. 5A showed that all experimental groups essentially reached the healing endpoint by day 11. Among them, the group infected with 1 × 108 CFU mL−1 of MRSA exhibited the most delayed healing, showing the slowest wound healing progression. By day 11, 99.3% of patients in the control group had healed, compared to 89.7% of patients in the MRSA-infected group (Fig. 5B). Further quantitative analysis of wound bacterial loads revealed that the group infected with 1 × 108 CFU mL−1 of MRSA had significantly higher bacterial loads on days 1, 3, 5, and 7 compared to all other groups (Fig. 5C). In the remaining groups, the wound bacterial colony counts showed a gradient decline with the initial inoculum decrease (Fig. 5D and E). In summary, 1 × 108 CFU mL−1 of MRSA maintained a high bacterial burden throughout the observation period and effectively delayed the wound healing process. Therefore, this concentration was selected as the appropriate bacterial strain and inoculation concentration for the subsequent diabetic mouse wound infection model.3.6 The effect of PLH therapy on MRSA infection in diabetic mice
We created a mouse model of diabetes using streptozotocin (STZ) and full-thickness skin wounds. Research indicates that hydrogels, as wound dressings, offer significant advantages in skin wound healing due to their ability to provide a moist environment and promote tissue regeneration.29 Hydroxyethyl cellulose (HEC) was selected as the gel matrix for its multifunctional role in the therapeutic design. As a natural cellulose derivative, this material with excellent biocompatibility provides a moist environment essential for tissue regeneration. Furthermore, it can act as a sustained-release reservoir to localize and prolong the antibacterial action of the PLH at the wound site.30,31 The therapeutic effects of HEC-based functional dressings loaded with PLH (PLH@Gel) on MRSA-infected wounds were systematically evaluated. Results showed that all investigated mice achieved FBG levels exceeding 11.1 mmol L−1, meeting the criteria for the diabetic model (Fig. 6A). By day 20 of the treatment period, the body weight of the mice in the different groups had decreased. Followed by a slow increasing trend, with no significant differences observed between groups (Fig. 6B). To screen the optimal therapeutic concentration of PLH@Gel, three concentrations were used for treatment: 0.25% PLH@Gel, 0.5% PLH@Gel, and 1% PLH@Gel, with silver sulfadiazine (AgSD) as the positive control.32 With the exception of the blank control group, circular skin wounds measuring 1 cm in diameter with 1 × 108 CFU mL−1 of MRSA suspension and treated with PBS, AgSD, or PLH@Gel, respectively. Results indicated the reduction of wound areas in all infected groups on day 9 due to the self-repair capacity of mice. The remaining wound areas in the Control, Model, AgSD, 0.25% PLH@Gel, 0.5% PLH@Gel, and 1% PLH@Gel groups decreased to 6.70%, 17.77%, 15.49%, 7.84%, 5.87%, and 5.06%, respectively, on day 13 (Fig. 6C–E). Compared to the Model group, both AgSD and PLH@Gel promoted wound healing, and the healing effect in the PLH@Gel group was superior to that of AgSD group. Within the PLH@Gel groups, the wound healing promotion at 0.5% and 1% concentrations was significantly higher than that at 0.25%. The body weight of mice remained stable across all groups throughout the treatment period (Fig. 6F). Quantitative analysis of the bacterial loads in wound tissues served to further evaluate the antibacterial efficacy. On day 3 of treatment, the antibacterial rates of AgSD, 0.25% PLH@Gel, 0.5% PLH@Gel, and 1% PLH@Gel reached 80.67%, 23.59%, 96.23%, and 99.99%, respectively (Fig. 6G and H). Wound skin tissues collected on day 13 of treatment were subjected to H&E staining. The results showed that the Control group exhibited intact epidermal structure with visible hair follicle regeneration. The Model group displayed severe disruption of the epithelial structure, accompanied by massive inflammatory cell infiltration. The AgSD and 0.25% PLH@Gel groups showed larger wound areas with erythematous edges and residual inflammation. In contrast, the 0.5% PLH@Gel and 1% PLH@Gel groups demonstrated complete epidermal regeneration, essentially closed wounds, and observable hair follicle regeneration. As evidenced by Masson's trichrome staining, the wound beds in the Model group were predominantly composed of red-stained fibroblasts and inflammatory cells, with sparse collagen deposition. In comparison, the 0.5% and 1% PLH@Gel groups exhibited abundant blue-stained collagen fibers and red myofibers, indicating nearly complete tissue repair (Fig. 6I). In summary, 0.5% PLH@Gel demonstrated optimal therapeutic efficacy in the diabetic mouse group, not only efficiently eliminating pathogenic bacteria but also significantly promoting wound healing and tissue regeneration.3.7 CPB@PLH@Gel can enhance healing-promoting efficacy in diabetic mice with MRSA-infected wounds
We evaluated the potential of CPB@PLH@Gel as a functional wound dressing for MRSA-infected wounds in diabetic mice. As shown in Fig. 7A, by day 18 of modeling, the FBG levels in the mice reached above 20 mM, meeting the criteria for a severe hyperglycemia model and exhibiting characteristic impaired chronic wound healing features.33,34 These mice were subsequently used for infectious wound treatment. No significant fluctuations in body weight were observed among the groups during the study (Fig. 7B). Following 13 days of treatment, wound area analysis revealed that the remaining wound areas in the Model, CPB@Gel, PLH@Gel, and CPB@PLH@Gel groups were reduced to 46.05%, 21.56%, 14.13%, and 4.26%, respectively (Fig. 7C–E). Body weights remained stable throughout the treatment period across all groups (Fig. 7F). The results demonstrated that CPB@PLH@Gel possessed the optimal wound-healing promotion efficacy. To further assess the in vivo antibacterial effect, wound bacterial samples were collected on days 5 and 7 of treatment, and bacterial colonies were counted using the dilution spot assay. The results indicated that the CPB@PLH@Gel group cleared approximately 99% of MRSA, significantly outperforming the other groups (Fig. 7G), confirming the strong antibacterial activity in vivo. H&E staining showed that exhibited severe epidermal necrosis, loose structure, and extensive inflammatory cell infiltration in the wounds of Model group. The CPB@Gel group showed superficial gaps but fewer underlying inflammatory cells, indicating incomplete tissue recovery. The PLH@Gel group showed an intact epidermal structure, with only minor infiltration of inflammatory cells. In contrast, the CPB@PLH@Gel group showed significant epidermal regeneration, with new hair follicles on both sides of the skin, reduced inflammation, and compact tissue arrangement. As demonstrated by Masson staining, the most substantial collagen deposition was observed in the CPB@PLH@Gel group (Fig. 7H), effectively promoting structural reconstruction of the dermal layer. In summary, CPB@PLH@Gel significantly accelerated the healing of diabetic infected wounds through anti-bacterial and tissue regeneration promotion.3.8 Immunohistochemical assessment of inflammation and angiogenesis
Angiogenesis is a critical process in wound healing, which can significantly accelerate tissue repair by delivering oxygen, growth factors, and immune cells to the wound site.35,36 To evaluate the effects of CPB@PLH@Gel on angiogenesis, immunohistochemistry (IHC) was employed to determine the expression levels of VEGF and CD31. The results showed that the CPB@PLH@Gel group exhibited higher levels of both VEGF and CD31 compared to the Model and PLH@Gel groups (Fig. 8A and S4). This result suggested that CPB@PLH@Gel effectively promotes neovascularization in the wound tissue, thereby accelerating wound healing. IHC analysis revealed a significantly lower area of TNF-α staining in the CPB@PLH@Gel group compared to others, while the anti-inflammatory cytokine IL-10 was markedly upregulated (Fig. 8B). This shift indicates effective inflammation suppression and a transition of the wound microenvironment toward repair.37 To further investigate the immunomodulatory effect at the cellular level, macrophage polarization was assessed by performing immunohistochemical staining of M1 marker (CD80) and M2 marker (CD206). The results showed that CPB@PLH@Gel treatment promoted a phenotypic shift from the pro-inflammatory M1 state towards the pro-healing M2 state in wound macrophages (Fig. 8C). In summary, CPB@PLH@Gel synergistically promoted angiogenesis by upregulating VEGF and CD31 to accelerate neovascularization, while simultaneously modulated the immune response by reducing TNF-α, elevating IL-10, and facilitating the transition of macrophages from M1 to M2 phenotype. These actions collectively alleviated excessive inflammation and facilitated the repair process of diabetic chronic wounds.4 Conclusion
This study successfully constructed a composite nano-platform (CPB@PLH NPs) based on a novel antimicrobial peptide and carboxylated Prussian blue nanoparticles for the treatment of MRSA-infected wounds in diabetic mice. The composite nanomedicine, prepared via a straightforward process, exhibited favorable stability and dispersity. In vivo results demonstrated that CPB@PLH NPs possess significant dual functionalities of antibacterial and anti-inflammatory effects. The rapid bacteria clearance and wound microenvironment modulation by alleviating inflammation and oxidative stress synergistically promoted wound healing in diabetic mice. In summary, this composite nanosystem demonstrates promising potential for the treatment of chronically infected wounds in diabetes.Author contributions
Yilong Li: validation, funding acquisition, Aidi Tong: conceptualization, investigation, writing original draft. Pengxiang Xu: conceptualization, methodology, resources. Li Chang: validation, data curation. Jialong Fan: data analysis. Zhoubing Shan: formal analysis, validation. Chunyi Tong: resources methodology, funding acquisition. Bin Liu: writing – review & editing, funding acquisition, project administration.Conflicts of interest
The authors declare that there is no conflict of interest between authors. These authors contributed to the work equally and should be regarded as co-first authors.Data availability
The authors declare that all data relevant to the study are included in the article.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra09475e.
Acknowledgements
This work was financially supported by the Natural Science Foundation of Hunan Province (2025JJ50075).References
- K. McDermott, M. Fang, A. J. M. Boulton, E. Selvin and C. W. Hicks, Diabetes Care, 2023, 46, 209–221 CrossRef PubMed.
- A. Staniszewska, F. Game, J. Nixon, D. Russell, D. G. Armstrong, C. Ashmore, S. A. Bus, J. Chung, V. Chuter, K. Dhatariya, G. Dovell, M. Edmonds, R. Fitridge, C. Gooday, E. J. Hamilton, A. Jones, V. Kavarthapu, L. A. Lavery, J. L. Mills, M. Monteiro-Soares, M. Osborne-Grinter, E. J. G. Peters, J. Shalhoub, J. van Netten, D. K. Wukich and R. J. Hinchliffe, Diabetes Care, 2024, 47, 1958–1968 CrossRef PubMed.
- Z. Zheng, J. Yang, W. Zheng, Z. Chu, W. Wang, H. Qian and L. Xu, J. Contr. Release, 2025, 385, 114058 CrossRef CAS PubMed.
- X. Wang, S. Liu, S. Jing, S. Zhang, Y. Jin, W. Zhang, Q. Ruan and W. Li, Adv. Sci., 2025, e12127 CrossRef CAS PubMed.
- W. Luo, J. Liu, Y. Jiang, T. Wang, N. Du, S. A. Algharib and S. Xie, Mater. Today Bio, 2025, 33, 102094 CrossRef CAS PubMed.
- Z. Yu, M. Li, L. Yang, H. Liu, G. Ding, S. Ma, L. Liu and S. Dong, Nano Today, 2024, 57, 102358 CrossRef CAS.
- Y. Chen, X. Wang, S. Tao, Q. Wang, P. Q. Ma, Z. B. Li, Y. L. Wu and D. W. Li, Mil. Med. Res., 2023, 10, 37 Search PubMed.
- M. Cui, J. Zhang, P. Han, L. Shi, X. Li, Z. Zhang, H. Bao, Y. Ma, Z. Tao, X. Dong, L. Fu and Y. Wu, Mater. Today Bio, 2024, 28, 101186 CrossRef CAS PubMed.
- M. Chang and T. T. Nguyen, Acc. Chem. Res., 2021, 54, 1080–1093 CrossRef CAS PubMed.
- W. Cai, Y. Song, Q. Xie, S. Wang, D. Yin, S. Wang, S. Wang, R. Zhang, M. Lee, J. Duan and X. Zhang, J. Contr. Release, 2024, 375, 627–642 CrossRef CAS PubMed.
- R. Yang, X. Ma, F. Peng, J. Wen, L. W. Allahou, G. R. Williams, J. C. Knowles and A. Poma, Biotechnol. Adv., 2025, 81, 108570 CrossRef CAS PubMed.
- G. Li, Z. Lai and A. Shan, Adv. Sci., 2023, 10, e2206602 CrossRef PubMed.
- C. A. Roque-Borda, L. M. D. G. Primo, K. P. Medina-Alarcón, I. C. Campos, C. F. Nascimento, M. M. S. Saraiva, A. Berchieri Junior, A. M. Fusco-Almeida, M. J. S. Mendes-Giannini, J. Perdigão, F. R. Pavan and F. Albericio, Adv. Sci., 2025, 12, e2410893 CrossRef PubMed.
- L. Wang, L. Qu, S. Lin, Q. Yang, X. Zhang, L. Jin, H. Dong and D. Sun, Curr. Protein Pept. Sci., 2022, 23, 226–247 CrossRef.
- X. Ma, N. Yang, R. Mao, Y. Hao, Y. Li, Y. Guo, D. Teng, Y. Huang and J. Wang, J. Nanobiotechnol., 2024, 22, 668 CrossRef CAS PubMed.
- Z. Liao, J. Li, W. Ni, R. Zhan and X. Xu, Carbohydr. Polym., 2025, 348, 122873 CrossRef CAS PubMed.
- J. Fan, Y. Qin, W. Qiu, J. Liang, C. Xiao, Q. Xie, C. Tong, L. Yuan, Y. Long and B. Liu, Biomaterials, 2025, 314, 122851 CrossRef CAS.
- J. Liang, X. Tian, M. Zhou, F. Yan, J. Fan, Y. Qin, B. Chen, X. Huo, Z. Yu, Y. Tian, S. Deng, Y. Peng, Y. Wang, B. Liu and X. Ma, Biomaterials, 2024, 309, 122608 CrossRef CAS PubMed.
- A. Tong, C. Tong, J. Fan, J. Shen, C. Yin, Z. Wu, J. Zhang and B. Liu, Biomater. Sci., 2023, 11, 6342–6356 RSC.
- H. Zhou, P. You, H. Liu, J. Fan, C. Tong, A. Yang, Y. Jiang and B. Liu, J. Contr. Release, 2022, 341, 828–843 CrossRef CAS PubMed.
- H. Y. Kwon, Y. Jung, H. Jeon and H. S. Han, Bioact. Mater., 2025, 48, 71–99 Search PubMed.
- Z. Wu, L. Chang, C. Li, P. Xu, L. Liu, A. Tong, Y. Li, C. Tong and B. Liu, Adv. Healthcare Mater., 2025, 14, e2405100 CrossRef PubMed.
- C. Tong, X. Zhong, Y. Yang, X. Liu, G. Zhong, C. Xiao, B. Liu, W. Wang and X. Yang, Biomaterials, 2020, 243, 119936 CrossRef CAS PubMed.
- R. Cong, C. Deng, P. Li, Y. Tang, J. Hou, J. Zhao, Q. Wang, Y. Chen, J. Tu, X. Jiang and L. Jiang, Nat. Commun., 2025, 16, 5797 CrossRef PubMed.
- X. Lv, L. Wang, A. Mei, Y. Xu, X. Ruan, W. Wang, J. Shao, D. Yang and X. Dong, Small, 2023, 19, e2206220 Search PubMed.
- Z. Li, Y. Bai, H. Wu, Y. Feng, X. Wang, C. Zhao and X. Wang, Mater. Today Bio, 2025, 32, 101651 CrossRef CAS PubMed.
- X. Wei, C. Liu, Z. Li, Z. Gu, J. Yang and K. Luo, Carbohydr. Polym., 2024, 331, 121873 CrossRef CAS PubMed.
- C. Liu, R. Zhou, B. Chen, X. Yan, L. Guo, Y. Tang, X. Zuo, X. Guo, H. Yu, J. Chen, Z. Guo, F. Wang and C. Xu, Mater. Today Bio, 2025, 31, 101622 CrossRef CAS PubMed.
- L. Xu, J. Zhang, J. Luo, Y. Cui, J. Chen, B. Zeng, Z. Deng and L. Shao, J. Nanobiotechnol., 2025, 23, 387 CrossRef PubMed.
- P. Luo, L. Liu, W. Xu, L. Fan and M. Nie, Carbohydr. Polym., 2018, 199, 170–177 CrossRef CAS PubMed.
- F. Song, A. Ye, L. Jiang, Y. Lu, Y. Feng, R. Huang, S. Du, X. Dong, T. Huang, P. Li, L. Yang, J. Zhang, M. Xu, L. Cheng and J. Xiao, Mater. Today Bio, 2024, 30, 101439 CrossRef PubMed.
- S. J. Lee, Z. Wu, M. Huang, C. Liang, Z. Huang, S. Chen, V. Rajasekar, M. M. Abdalla, H. Nah, D. N. Heo, I. K. Kwon, M. J. Cho, S. J. Kim, S. Sohn, S. H. Kim, R. Sugimura and C. K. Y. Yiu, Mater. Today Bio, 2025, 31, 101451 Search PubMed.
- A. Couturier, C. Calissi, J. L. Cracowski, D. Sigaudo-Roussel, C. Khouri and M. Roustit, EBioMedicine, 2023, 98, 104856 CrossRef PubMed.
- X. Li, Y. Yang, Z. Li, Y. Wang, J. Qiao and Z. Chen, Diabetologia, 2023, 66, 1084–1096 CrossRef CAS.
- Y. Hu, R. Tao, L. Chen, Y. Xiong, H. Xue, L. Hu, C. Yan, X. Xie, Z. Lin, A. C. Panayi, B. Mi and G. Liu, J. Nanobiotechnol., 2021, 19, 150 CrossRef CAS PubMed.
- L. Sheng, Z. Zhang, Y. Zhang, E. Wang, B. Ma, Q. Xu, L. Ma, M. Zhang, G. Pei and J. Chang, Biomaterials, 2021, 264, 120414 CrossRef CAS.
- C. H. Kuan, L. Chang, C. Y. Ho, C. H. Tsai, Y. C. Liu, W. Y. Huang, Y. N. Wang, W. H. Wang and T. W. Wang, Biomaterials, 2025, 314, 122848 CrossRef CAS PubMed.
Footnote |
| † Co-first authors. |
| This journal is © The Royal Society of Chemistry 2026 |








