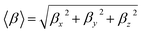Open Access Article
Open Access ArticleImidazolium versus phenylalaninium hybrid salts: supramolecular control of electronic, nonlinear optical and bioactive properties
Sandra Walhaab,
Rageh K. Husseinc,
Thierry Roisneld and
Ali Ben Ahmed *ef
*ef
aLaboratory Physical-Chemistry of Solid State, Chemistry Department, Faculty of Sciences of Sfax, University of Sfax, BP 1171, 3000 Sfax, Tunisia
bDepartment of Chemical, Preparatory Institute for Engineering Studies of Gabes, Gabes 6029, Tunisia
cDepartment of Physics, College of Sciences, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh 11623, Saudi Arabia
dCentre National de Recherche Scientifique, Institut des Sciences Chimiques de Rennes, Université de Rennes, UMR 6226, F-35000 Rennes, France
eDepartment of Biomedical, Higher Institute of Biotechnology of Sfax, University of Sfax, Route Sokra km 4, BP 1175, 3000 Sfax, Tunisia
fLaboratory of Applied Physics, Department of Physics, Faculty of Sciences of Sfax, University of Sfax, Route Soukra km 3.5, BP 1171, 3000, Sfax, Tunisia. E-mail: ali.benahmed@isbs.usf.tn
First published on 20th February 2026
Abstract
Two new amino-acid-based hybrid salts, L-phenylalaninium chloride (1) and L-histidinium iodide (2), were synthesized and comprehensively characterized to elucidate how cation identity, halide nature, and supramolecular architecture govern their optical, electronic, and biological properties. Single-crystal X-ray diffraction revealed that both compounds crystallize in the non-centrosymmetric P212121 space group but exhibit distinct packing motifs: a compact chloride-stabilized network in (1) and a more open, highly polarized iodide-based framework in (2). DFT calculations show that compound (2) possesses a significantly reduced HOMO–LUMO band gap, higher molecular softness, and enhanced intramolecular charge transfer, leading to a first hyperpolarizability nearly three times larger than that of (1). FT-IR and RDG analyses confirm stronger charge-assisted N–H⋯I interactions in (2), consistent with its superior electronic polarization. Biological assays further demonstrate that compound (2) exhibits markedly enhanced antioxidant and antibacterial activities, attributed to its imidazolium-based charge distribution and iodide polarizability. Together, these findings establish clear structure–property relationships and highlight the effectiveness of combining heterocyclic amino acids with highly polarizable halides to design multifunctional organic–inorganic materials with promising NLO and bioactive potential.
1. Introduction
Organic–inorganic hybrid salts derived from amino acids have emerged as versatile functional materials due to their unique combination of structural flexibility, intrinsic chirality, and biological compatibility.1,2 These molecular systems often exhibit rich supramolecular architectures governed by directional hydrogen bonding and tunable electrostatic interactions, making them highly attractive for nonlinear optical (NLO) applications, optoelectronic devices, and bioactive materials design. Among naturally occurring amino acids, L-phenylalanine and L-histidine occupy a central place owing to the distinct chemical characteristics of their side chains: an aromatic phenyl ring in L-phenylalanine that promotes π-mediated interactions and non-centrosymmetric packing, and an imidazole heterocycle in L-histidine capable of proton transfer, metal coordination, and strong charge-assisted hydrogen bonding. These features endow their salts with advantageous optical, electronic, and biological properties. Recent studies have shown that amino-acid-based crystals can exhibit remarkable second harmonic generation (SHG) efficiencies, enhanced charge-transfer characteristics, and promising antimicrobial or antioxidant activities, positioning them as candidates for bio-inspired multifunctional materials.3–6Biologically, phenylalaninium salts exhibit moderate antimicrobial activity and can influence membrane interactions through the presence of halide counter-ions. This effect is often observed in model systems where the phenyl ring enhances hydrophobic interactions with lipid bilayers.7,8 For example, R-bis homo-phenylalanine has been shown to mildly disrupt bacterial membrane integrity, although its activity remains significantly weaker than that of heterocycle-based antimicrobial agents.9 Despite these properties, the biomedical relevance of phenylalaninium derivatives remains limited compared with imidazole-containing systems. Consequently, phenylalaninium complexes are more frequently investigated for their contributions to advanced photonic and optoelectronic materials, such as nonlinear optical crystals or proton-transfer chromophores, rather than for therapeutic applications. For instance, phenylalaninium bromide salts have been used as benchmark materials for second-harmonic generation (SHG) studies, where the aromatic backbone facilitates efficient π–π stacking in the crystal lattice.10
In contrast, L-histidinium, featuring the highly versatile imidazole heterocycle, demonstrates strong biological relevance. Histidine-derived salts are widely studied owing to the imidazole ring's capacity to participate in proton transfer, metal coordination, and acid–base buffering. A well-known example is the role of histidine residues in zinc-binding domains of serum proteins, where the imidazole nitrogen coordinates Zn2+ with high affinity, stabilizing catalytic or regulatory sites.11,12 This inherent biological functionality translates into potent antimicrobial activity against a broad spectrum of pathogens, as the imidazole moiety can disrupt key biochemical pathways. Moreover, L-histidinium exhibits a remarkable antioxidant capability, particularly through the scavenging of Reactive Oxygen Species (ROS). Studies have demonstrated that histidinium-based salts can neutralize hydroxyl radicals and superoxide anions, thereby reducing oxidative stress in biological systems.13 This antioxidant behavior further enhances their pharmaceutical relevance, especially in formulations targeting inflammation or oxidative damage. Therefore, while phenylalaninium complexes tend to be prioritized for their structural versatility and optical functionality supporting developments in material science L-histidinium complexes are valued for their direct therapeutic potential, ranging from antimicrobial to antioxidant applications. The comparative study of these two salt families is thus essential for bridging the gap between structural chemistry, optical physics, and pharmaceutical science.
Despite this growing interest, several key gaps remain in the literature.14–16 First, systematic investigations correlating the halide identity with the crystal packing, electronic structure, and functional performance of amino-acid salts are still rare, although halide substitution is known to profoundly influence polarizability, hydrogen-bonding topology, and NLO response. Second, while multiple reports describe either structural or biological aspects of such salts, integrated studies combining crystallography, Hirshfeld surface analysis, DFT calculations, NLO descriptors, and biological assays within the same molecular framework are still lacking. Finally, comparative structure–property relationships involving phenylalaninium versus histidinium systems two families with fundamentally different supramolecular behaviors and electronic signatures remain insufficiently explored. Addressing these gaps is crucial for establishing predictive design principles in amino-acid-based photonic and biofunctional materials.
In this context, the present work focuses on two new protonated amino-acid salts: L-phenylalaninium chloride (C9H12NO3·Cl, compound 1) and L-histidinium iodide (C6H10N3O2·I, compound 2). Both compounds were synthesized by slow evaporation and structurally characterized by single-crystal X-ray diffraction, revealing non-centrosymmetric orthorhombic lattices (P212121) stabilized by extensive hydrogen-bond networks and halide-dependent supramolecular interactions. To complement the crystallographic study, Hirshfeld surface analysis, density functional theory (DFT) calculations, frontier molecular orbital (FMO) analysis, reduced-density-gradient (RDG)/NCI mapping, and electrostatic potential (MEP) mapping were employed to elucidate the role of intermolecular forces and electronic polarization in determining the functional properties. The optical relevance of the two salts was assessed through IR spectroscopy and nonlinear optical (NLO) calculations, while their biological activities were evaluated via antimicrobial assays against clinically relevant strains and antioxidant tests (DPPH, ABTS, FRAP).
By combining experimental characterization, theoretical modelling, and functional testing, this study provides a comprehensive structure–property analysis of two amino-acid-derived hybrid salts. Special emphasis is placed on understanding how the distinct chemical nature of phenylalaninium versus histidinium cations, combined with halide identity (Cl− vs. I−), governs their supramolecular organization, electronic distribution, nonlinear optical response, and biological activities. These insights help establish design principles for the development of bio-inspired hybrid materials for photonic, optoelectronic, and biomedical applications.
2. Experimental and theoretical methods
2.1. Materials and synthesis
All reagents used in this study, hydrochloric acid (HCl, 37%), hydroiodic acid (57%), L-phenylalanine (C9H11NO2, 98%), and L-histidine (C6H9N3O2, 99%) were purchased from Sigma-Aldrich and employed as received, without any additional purification. The two hybrid salts, (1) and (2), were prepared using a slow-evaporation approach. For the synthesis of compound (1), L-phenylalanine (1 mmol; 0.165 g) and hydrochloric acid (1 mmol; five drops) in water. The reaction mixture was magnetically stirred until complete dissolution of the components was achieved. After one-week, well-defined needle-shaped crystals suitable for analysis were obtained, while for compound (2), L-histidine (2 mmol; 0.310 g) and hydriodic acid (1 mmol; approximately five drops) were dissolved in 50 mL of double-distilled water and stirred at 293 K for 4 h. The solution was filtered and allowed to dry at room temperature by slow evaporation. After three weeks, pale-yellow block-shaped crystals were obtained in a yield of 95%.2.2. Single-crystal X-ray diffraction
Single-crystal X-ray diffraction data for compounds (1) and (2) were collected at 293 K using a Kappa CCD diffractometer with Mo-Kα radiation. Both structures were found to crystallize in the orthorhombic system with the space group P212121. Structure solution was carried out with SHELXT,17 followed by full-matrix least-squares refinement on F2 using SHELXL.18 All non-hydrogen atoms were refined anisotropically. Hydrogen atoms bonded to nitrogen were located from Fourier difference maps, while the remaining H atoms were placed in their idealized geometrical positions and refined using a riding model. Molecular graphics of the asymmetric units were generated using the MERCURY19 and DIAMOND20 programs. A summary of the crystallographic parameters, data collection conditions, and refinement statistics for salts (1) and (2) is provided in Table 1.| Structural data | (1) | (2) |
|---|---|---|
| Empirical formula | C9H12NO3·Cl | C6H10N3O2·I |
| Formula weight (g mol−1) | 201.65 | 283.07 |
| Temperature (K) | 293(2) | 293(2) |
| Wavelength (Å) | MoKα 0.71073 | MoKα 0.71073 |
| Crystal system | Orthorhombic | Orthorhombic |
| Space group | P212121 | P212121 |
| a (Å) | 5.3830(17) | 7.8399(8) |
| b (Å) | 7.045(2) | 9.4995(8) |
| c (Å) | 27.746(10) | 24.376(2) |
| α (°) | 90 | 90 |
| β (°) | 90 | 90 |
| γ (°) | 90 | 90 |
| Volume (Å3) | 1052.2(6) | 1815.4(3) |
| Space group Z | 4 | 4 |
| CCDC number | 2504992 | 1400108 |
2.3. Hirshfeld surface and supramolecular analysis
Hirshfeld surface (HS) analysis was conducted using CrystalExplorer,21 to examine intermolecular contacts and quantify contributions from hydrogen bonds, halide interactions, and van der Waals forces. Surfaces were mapped over dnorm, and 2D fingerprint plots were generated to elucidate individual interaction types. The analysis provided complementary insight into crystal packing and supramolecular stabilization.2.4. FT-IR spectroscopy
Infrared spectra were recorded at room temperature using a Bruker FT-IR spectrometer in the range 400–4000 cm−1, employing the KBr pellet method. Experimental assignments were supported by theoretical vibrational calculations, allowing reliable identification of functional groups and confirmation of protonation states.2.5. Computational method
Density Functional Theory (DFT) calculations were performed using Gaussian 16.22 The ground-state geometries of the investigated compounds were optimized by searching for the minimum on the potential energy surface, ensuring that the total electronic energy reached a stable equilibrium configuration. These quantum-chemical calculations were further employed to support the analysis of the experimental spectroscopic features through simulated vibrational and electronic data. All computations were performed with the wB97XD functional23 in combination with the 6-311++G(d,p)24 basis set. All calculations were performed within the static electric field (finite-field) approach at the same DFT level as the geometry optimizations, ensuring methodological consistency. Optimized geometries corresponded to true minima, confirmed by the absence of imaginary frequencies. Simulated IR spectra, frontier molecular orbitals (FMOs), global reactivity descriptors, molecular electrostatic potential (MEP) maps, and reduced density gradient (RDG)/NCI analyses were computed to correlate experimental observations with electronic structure properties. Molecular visualizations were generated using GaussView interface.25 The hyperpolarizability tensor components were extracted from the Gaussian output using the standard definition:26Since the values of the first hyperpolarizability of Gaussian 16 output are reported in atomic units (a.u.), the calculated values have been converted into electrostatic units (e.s.u.): βtot: 1 a.u. = 8.639418 × 10−33 e.s.u.
Urea was selected as a reference for hyperpolarizability calculations due to its well-established NLO properties, providing a consistent benchmark for relative comparison across molecular systems. Urea is widely recognized as a standard reference compound for first-order hyperpolarizability (β) calculations because of its well-characterized NLO properties and extensive use in theoretical and experimental studies. While we acknowledge that urea is chemically distinct from amino-acid-based hybrid salts and that its solid-state packing differs, the purpose of using urea in our study is to provide a consistent and reproducible reference point for comparing relative β values. By calculating the hyperpolarizability of urea under identical computational conditions (same level of theory, basis set, and electric field), we can reliably express the NLO response of the studied hybrid salts in relative units, which is a standard and accepted practice for screening and comparative studies of molecular NLO behavior.
2.6. Antimicrobial activity
In the DPPH assay, 100 µL of sample solution was added to 3.9 mL of 0.1 mM DPPH solution in methanol, and the mixture was incubated in the dark at room temperature for 30 min. Absorbance was measured at 517 nm, and the percentage inhibition was calculated relative to a blank. For the ABTS assay, the ABTS radical cation (ABTS˙+) was generated by reacting 7 mM ABTS with 2.45 mM potassium persulfate for 12–16 h in the dark. The radical solution was diluted to an absorbance of approximately 0.70 at 734 nm, and 100 µL of sample was mixed with 3.9 mL of ABTS˙+ solution. After 6 min incubation at room temperature, absorbance was recorded at 734 nm. The scavenging activity was calculated as a percentage of inhibition.
The FRAP assay was performed by mixing 100 µL of sample with 3 mL of freshly prepared FRAP reagent (300 mM acetate buffer, pH 3.6; 10 mM TPTZ in 40 mM HCl; 20 mM FeCl3·6H2O) and incubating at 37 °C for 30 min. The formation of the Fe2+–TPTZ complex was monitored at 593 nm, and results were expressed in µM Fe(II) equivalents per mg of compound. Ascorbic acid was used as a positive control in all assays. All experiments were performed in triplicate, and IC50 values for DPPH and ABTS assays were determined by plotting percentage inhibition against sample concentration using non-linear regression.
3. Results and discussion
3.1. Structural analysis
 | ||
| Fig. 1 The asymmetric unit of compound (1), showing the atom-labeling scheme. Displacement ellipsoids are drawn at the 50% probability level. The dashed line represents the possible hydrogen bond. | ||
The cations and anions entities are interconnected through an extensive system of hydrogen bonds and short intermolecular contacts. These interactions, predominantly of the N–H⋯O, N–H⋯Cl, and O/C–H⋯Cl types, give rise to a robust three-dimensional framework (Fig. 2).
This bonded network plays a decisive role in stabilizing the overall crystal architecture and governing the packing efficiency observed in the structure. The crystal structure can be described as an alternation of cationic and anionic layers along the [011] direction. The C–C bond lengths range from 1.320(4) Å to 1.534(5) Å, yielding an average value of 〈C–C〉 = 1.426 Å. The C–C–C bond angles range from 114.3(3)° to 121.2(5)°. These geometric parameters fall within expected limits and closely match those reported for aromatic frameworks,27,28 within the interval 106.1(6) to 125.2(8)°.
The crystal structure of compound L-phenylalaninium bromide,29 differs markedly from that of compound (1). While compound (1) crystallizes in the orthorhombic P212121 space group, L-phenylalaninium bromide adopts the monoclinic and noncentrosymmetric space group P21, resulting in a distinct molecular arrangement and symmetry environment. The substitution of chloride by the larger and more polarizable bromide anion modifies the hydrogen-bonding pattern. Although the geometry of the L-phenylalaninium cation remains comparable in both structures, slight variations in C–C distances and C–C–C angles reflect the influence of the different anionic environment. Overall, the comparison highlights how the change in halide and the adoption of the noncentrosymmetric space group P21 significantly affect the packing efficiency and supramolecular organization of the phenylalaninium salt.
 | ||
| Fig. 3 The asymmetric unit of (2) with atom labelling scheme. Displacement ellipsoids are drawn at the 50% probability level. | ||
The crystal arrangement may also be viewed as a mixed cationic and anionic layer propagated along the [010] direction (Fig. 4). The presence of two cation–anion pairs in the asymmetric unit enhances the complexity and richness of the supramolecular assembly. The structural cohesion arises from an extensive network of N–H⋯I hydrogen bonds linking the protonated amine groups to the iodide ions. These interactions serve as the dominant organizing forces and play a crucial role in stabilizing the non-centrosymmetric lattice. The N–H⋯I contact are particularly strong due to the high polarizability and size of the iodide anion, which facilitates directional binding and creates extended chains of electrostatic interactions throughout the crystal. These hydrogen-bonded motifs induce an ordered alignment of the histidinium cations, supporting cooperative dipole orientation a critical requirement for efficient second-harmonic generation (SHG). Overall, the supramolecular arrangement highlights how the combination of protonation, hydrogen bonding, and halide polarizability collectively drives the formation of an optically active structure.
 | ||
| Fig. 4 Projection of the crystal structure of (2) in the ac-plane showing the N–H⋯I and C–H⋯I hydrogen bonds. | ||
A comparison between L-histidinium iodide (2) and L-histidine chloride monohydrate30 reveals clear structural distinctions driven by the nature of the halide and the presence of lattice water. Compound (2) crystallizes in the non-centrosymmetric space group P212121 and develops a three-dimensional network dominated by strong N–H⋯I hydrogen bonds that generate channel-like supramolecular motifs. In contrast, L-histidine chloride monohydrate adopts a more compact and isotropic packing, where the chloride anion and the coordinated water molecule form an extensive set of N–H⋯Cl, O–H⋯Cl, and O–H⋯O interactions. While the histidinium cation maintains comparable internal geometry in both materials, the higher polarizability of I− and the absence of hydration in (2) promote a more open framework relative to the denser architecture of the chloride monohydrate.
 | ||
| Fig. 5 (A and B) View of the Hirshfeld surface of (1) and (2) mapped over dnorm, respectively. The 2D finger prints plots of all interactions. | ||
The dnorm values are projected onto the Hirshfeld surface using a red-white-blue gradient: red areas correspond to closer than van der Waals contacts (negative dnorm), blue areas indicate contacts longer than the sum of van der Waals radii (positive dnorm), and white regions reflect contacts equal to the van der Waals separation (dnorm ≈ 0). Hydrogen-bond donor acceptor interactions are clearly identifiable on the dnorm surface. Deep red circular depressions highlight strong hydrogen-bonding contacts, while smaller red or white spots correspond to H⋯H interactions, as depicted in Fig. 6(A) and (B).
In compound (1), the HS fingerprint plots reveal that H⋯H contacts constitute the largest contribution to the surface, accounting for 41.1% of the interactions, reflecting the high proportion of hydrogen atoms within the organic moieties. Significant H⋯Cl/Cl⋯H interactions (23.2%) and notable H⋯C/C⋯H contacts (16.0%) further stabilize the packing, while O⋯H/H⋯O interactions contribute 16.2%, indicating the presence of directional hydrogen bonding.
For compound (2), H⋯H contacts again dominate (31.1%), followed by strong O⋯H/H⋯O interactions (30.9%), highlighting their key role in forming hydrogen-bond networks. The presence of H⋯I/I⋯H contacts (23.8%) reflects the influence of the halide anion on the crystal cohesion, whereas C⋯H/H⋯C interactions (6.4%) play a minor but non-negligible role. Thus, the HS analysis confirms that a combination of hydrogen bonding and halogen-related contacts shapes the supramolecular architecture of both structures.
3.2. Vibrational analysis
Based on IR spectroscopic studies reported for compounds structurally related to our organic cation C9H12NO2+,31,32 the main absorption bands observed at room temperature for the L-phenylalaninium chloride salt can be reliably assigned. In the IR spectrum of the compound (1) (Fig. 7), the band at approximately 3385 cm−1 corresponds to the asymmetric stretching vibration of the NH3+ group. The absorptions near 2989 cm−1 and 2900 cm−1 are attributed to the symmetric and asymmetric stretching modes of CH2, respectively. A strong band appearing at 1737 cm−1 is assigned to the stretching of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O group. The absorption around 1587 cm−1 is associated with the asymmetric bending of NH3+.
O group. The absorption around 1587 cm−1 is associated with the asymmetric bending of NH3+.
The two characteristic bands at 1491 cm−1 and 1409 cm−1 correspond to the asymmetric and symmetric stretching vibrations of the carboxylate (COO−) group. The deformation mode δ(O–H) and CH2 twisting vibrations appear near 1340 cm−1. The group of absorptions in the 780–1129 cm−1 frequency region is attributed to stretching vibrations of C–N, C–C, and C–C–N bonds [ν(C–N), ν(C–C), ν(C–C–N)], as well as in-plane C–H bending.
The infrared (experimental and theoretical) spectra of the compound (2) are showing in Fig. 8. At the higher energy region there is an intense peak at 3439 cm−1 showing the presence of NH2 asymmetric stretching and the peak at 3226 cm−1 is due to NH2 symmetric stretching. C–H stretching is assigned at 3058 cm−1. The peak at 3013 cm−1 is due to strong CH2 asymmetric stretching. The corresponding strong CH2 symmetric stretching is positioned at 2951 cm−1. The weak C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching is represented by the peak at 1746 cm−1. The corresponding strong NH3+ asymmetric bending is positioned at 1643 cm−1 and the peak at 1616 cm−1 is due to symmetric bending.
O stretching is represented by the peak at 1746 cm−1. The corresponding strong NH3+ asymmetric bending is positioned at 1643 cm−1 and the peak at 1616 cm−1 is due to symmetric bending.
A direct comparison of the two compounds reveals the pronounced effect of halide identity on vibrational features. NHx stretching frequencies are broader and redder-more shifted in compound (2), reflecting stronger, more polarizable N–H⋯I hydrogen bonding compared to N–H⋯Cl. C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching is slightly higher in compound (2), consistent with a less compact packing environment and reduced O⋯H hydrogen-bond strength. C–N stretching is more intense and slightly shifted in compound (1), attributable to tighter N–H⋯Cl interactions that polarize the cation framework. The imidazolium ring in (2) introduces additional characteristic modes (C
O stretching is slightly higher in compound (2), consistent with a less compact packing environment and reduced O⋯H hydrogen-bond strength. C–N stretching is more intense and slightly shifted in compound (1), attributable to tighter N–H⋯Cl interactions that polarize the cation framework. The imidazolium ring in (2) introduces additional characteristic modes (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, N–H ring protonation) absent in (1), contributing to a richer vibrational profile.
N, N–H ring protonation) absent in (1), contributing to a richer vibrational profile.
These distinctions support the structural conclusions derived from SCXRD and Hirshfeld analysis. Compound (1) has a more compact, chloride-dominated hydrogen-bond network. Compound (2) exhibits stronger charge-assisted interactions with iodide and greater electronic delocalization.
3.3. Noncovalent interaction analysis
Reduced Density Gradient (RDG) and Non-Covalent Interaction (NCI) analyses provide further insight into the supramolecular architecture. The RDG plots of compound (1) (Fig. 9) reveal the presence of broad green isosurfaces corresponding to van der Waals interactions, in addition to sharp blue regions characteristic of strong hydrogen bonds. These weak but widespread interactions create a compact three-dimensional network that enhances crystal cohesion while preventing electron localization, a combination typically beneficial for maintaining optical transparency and enabling NLO responses.The NCI maps thus confirm that the crystal packing is dominated by a cooperative system of hydrogen-bonding and dispersive interactions that collectively stabilize the noncentrosymmetric framework required for second-harmonic generation (SHG). NCI analysis provides additional insight into the intermolecular forces that consolidate the packing.
For compound (2), the NCI maps (Fig. 10) display both strong blue isosurfaces corresponding to classical N–H⋯I hydrogen bonds and widespread green regions indicating van der Waals and dispersion interactions. These diffuse contacts help regulate molecular spacing and enhance the mechanical stability of the lattice while maintaining the electronic environment necessary for delocalization. This balanced network of strong and weak interactions ensures that the crystal remains rigid enough to maintain optical coherence but flexible enough to minimize internal strain, a combination highly favorable for NLO crystals.
3.4. Molecular electrostatic potential maps
The MEP maps (Fig. 11) further elucidate the charge distribution within the protonated L-phenylalaninium moiety. The strongly negative regions localized around the chloride anion and carboxylate oxygen atoms indicate potential sites for electrophilic attack, while the positive potentials concentrated around the ammonium protons correspond to regions susceptible to nucleophilic interaction. This polarization confirms a significant intramolecular and intermolecular charge separation, which is an essential factor in enhancing dipole moment and, consequently, the nonlinear optical response. The visualization thus validates the electronic asymmetry inferred from the crystallographic analysis.The MEP maps of compound (2) (see Fig. 12) reveal pronounced regions of negative potential around the iodide anions, as expected from their high electron density and polarizability. Conversely, positive potentials cluster around the protonated nitrogen sites of the histidinium cations. This well-defined charge polarization supports strong cation–anion electrostatic interactions and confirms the presence of a significant dipole moment at the molecular scale. Such polarized charge distribution is directly associated with enhanced hyperpolarizability and is consistent with the excellent NLO performance observed for this compound.
3.5. Frontier molecular orbitals maps and global chemical reactivity descriptors
Frontier molecular orbital (FMO) calculations provide quantitative insight into the electronic structure. As can be see Fig. 13, the HOMO is located predominantly on the L-phenylalaninium cation, while the LUMO is more delocalized, indicating possible intramolecular charge-transfer (ICT) transitions. The large HOMO–LUMO energy gap (9.780 eV) suggests high chemical stability and low intrinsic reactivity, meaning that the compound is resistant to spontaneous electron transfer but can exhibit stable optical activity under external excitation.The global reactivity descriptors align with this interpretation (see Table 2), the high hardness (η = 4.890 eV) and low softness (S = 0.204 eV−1) confirm a robust electronic framework, while the moderate electrophilicity index (ω = 1.917 eV) indicates controlled charge-accepting capacity. Collectively, these values reveal that compound (1) is electronically stable and structurally rigid conditions favorable for consistent and reliable NLO performance.
| Global reactivity descriptors | (1) | (2) |
|---|---|---|
| Energy HOMO (EH) | −9.220 | −7.760 |
| Energy LUMO (EL) | 0.560 | −0.380 |
| Energy gap (ΔEL−H) | 9.780 | 7.380 |
| Ionization potential (IP) | 9.220 | 7.760 |
| Electron affinity (EA) | −0.560 | 0.380 |
| Electronegativity (χ) | 4.330 | 4.070 |
| Chemical potential (µ) | −4.330 | −4.070 |
| Global hardness (η) | 4.890 | 3.690 |
| Global softness (S) | 0.204 | 0.271 |
| Electrophilicity index (ω) | 1.917 | 2.244 |
In Fig. 14, the FMO analysis shows a HOMO–LUMO energy gap of 7.38 eV, significantly smaller than that of compound (1). This reduced gap indicates increased charge-transfer capability and higher electronic polarizability, both of which are essential for strong nonlinear optical responses. The HOMO is mainly localized on the iodide anions and partially on the imidazolium ring, while the LUMO is distributed over the cationic framework, suggesting a favorable ICT pathway.
Global reactivity descriptors further support this interpretation, the relatively low hardness (η = 3.690 eV) and high softness (S = 0.271 eV−1) indicate an easily deformable electron cloud, which enhances the compound's ability to respond to external electric fields. The electrophilicity index (ω = 2.254 eV) is markedly higher than that of compound (1), reflecting stronger electron-accepting capability and suggesting greater reactivity in polarization-driven processes.
3.6. Biologic activities
| Bacterial strain | (1) | (2) | Standard drug |
|---|---|---|---|
| Staphylococcus aureus | 11 ± 0.5 | 16 ± 0.7 | 22 ± 0.5 |
| Bacillus subtilis | 10 ± 0.4 | 15 ± 0.6 | 20 ± 0.4 |
| Enterococcus faecalis | 9 ± 0.3 | 14 ± 0.5 | 19 ± 0.6 |
| Listeria monocytogenes | 10 ± 0.5 | 15 ± 0.6 | 21 ± 0.5 |
| Escherichia coli | 8 ± 0.3 | 13 ± 0.5 | 18 ± 0.6 |
| Pseudomonas aeruginosa | 7 ± 0.4 | 12 ± 0.5 | 17 ± 0.5 |
| Klebsiella pneumoniae | 8 ± 0.4 | 14 ± 0.6 | 19 ± 0.6 |
| Salmonella typhimurium | 9 ± 0.4 | 15 ± 0.6 | 20 ± 0.5 |
| Shigella flexneri | 8 ± 0.3 | 13 ± 0.5 | 18 ± 0.5 |
| Proteus vulgaris | 7 ± 0.3 | 12 ± 0.5 | 17 ± 0.5 |
 | ||
| Fig. 15 Proposed antibacterial mechanisms of L-phenylalaninium chloride (1) and L-histidinium iodide (2) against Gram-negative and Gram-positive bacteria. | ||
Conversely, Gram-negative bacteria (E. coli, P. aeruginosa, K. pneumoniae, S. typhimurium) showed smaller inhibition zones; nevertheless, L-histidinium iodide maintained measurable activity against these strains, suggesting abroader antibacterial spectrum. The observed increase in antibacterial activity may be tentatively correlated with the higher chemical softness and polarizability of the histidinium-based salt. In related ionic and imidazolium-derived systems, enhanced polarizability has been associated with stronger interactions with bacterial lipid membranes, resulting in membrane permeabilization and loss of cellular integrity37–39 (Epand and Epand, 2011). However, it should be emphasized that, in the present study, this interpretation is based on inhibition zone measurements and literature comparisons rather than direct mechanistic evidence. Overall, these findings indicate that L-histidinium iodide possesses promising antibacterial properties compared to L-phenylalaninium chloride, in agreement with previous reports on amino-acid-derived ionic species and imidazolium-based bioactive materials.40,41
| Assay | Concentration (mg mL−1) | (1) | (2) | Standard (ascorbic acid) |
|---|---|---|---|---|
| DPPH | 0.0625 | 18 ± 1.2 | 12 ± 0.9 | 25 ± 1.1 |
| 0.125 | 31 ± 1.5 | 22 ± 1.3 | 45 ± 1.5 | |
| 0.25 | 52 ± 1.7 | 38 ± 1.6 | 70 ± 2.0 | |
| 0.5 | 71 ± 2.0 | 57 ± 1.8 | 88 ± 2.1 | |
| 1.0 | 88 ± 2.1 | 73 ± 2.0 | 96 ± 1.9 | |
| IC50 (µg mL−1) | 245 ± 5 | 375 ± 6 | 180 ± 4 | |
| ABTS | 0.0625 | 22 ± 1.1 | 15 ± 1.0 | 30 ± 1.2 |
| 0.125 | 36 ± 1.4 | 27 ± 1.3 | 50 ± 1.4 | |
| 0.25 | 55 ± 1.6 | 42 ± 1.5 | 72 ± 2.0 | |
| 0.5 | 74 ± 2.0 | 60 ± 1.7 | 88 ± 2.0 | |
| 1.0 | 89 ± 2.1 | 77 ± 1.9 | 97 ± 1.5 | |
| IC50 (µg mL−1) | 230 ± 5 | 360 ± 6 | 170 ± 3 | |
| FRAP | — | 320 ± 8 µM Fe(II) per mg | 240 ± 7 µM Fe(II) per mg | 450 ± 10 µM Fe(II) per mg |
The FRAP results further support this trend, revealing higher reducing power for (1) (320 ± 8 µM Fe(II) per mg) relative to (2) (240 ± 7 µM Fe(II) per mg), yet both remain below the activity of ascorbic acid (450 ± 10 µM Fe(II) per mg), as previously reported for non-conjugated organic salts with moderate reduction capacity.43 Overall, the combined results show that (1) displays stronger antioxidant performance than (2), while both compounds demonstrate meaningful but moderate radical-scavenging and reducing abilities. This result confirms the higher electron-transfer capability of compound (2), consistent with its lower HOMO–LUMO gap and greater molecular softness observed in the DFT analysis.
Taken together, the antioxidant results indicate that the redox activity of the two salts is strongly governed by their intrinsic structural and electronic features. In compound (1), the redox response is mainly influenced by the phenyl moiety, leading to moderate single-electron transfer reactivity. In contrast, compound (2) benefits from stronger resonance-assisted stabilization within the imidazolium ring, combined with the high polarizability of the iodide anion, which collectively enhance its electron- and proton-transfer capabilities and result in superior antioxidant properties. These experimental trends are fully consistent with the theoretical descriptors namely the lower HOMO–LUMO band gap and higher molecular softness of compound (2) demonstrating excellent agreement between computational predictions and observed biological behaviour.
3.7. Nonlinear optical parameters
The nonlinear optical (NLO) calculations demonstrate the functional relevance of the structural and electronic findings. As showing Table 5 The total dipole moment (µtot = 7.45 D) is significantly higher than that of urea, a standard reference NLO material. The polarizability (αtot = 12.53 × 10−24 e.s.u.) and hyperpolarizability (βtot = 8.83 × 10−31 e.s.u.) also substantially exceed those of urea (βtot = 1.3 × 10−31 e.s.u.), translating to an SHG efficiency 6.8 times larger. These enhanced optical parameters result directly from the non-centrosymmetric arrangement, hydrogen-bond-supported molecular orientation, and the strong dipolar nature imposed by protonation. The synergy between crystal symmetry, intermolecular interactions, and electronic polarization thus positions compound (1) as a promising candidate for photonic and optoelectronic applications, including frequency doubling and electro-optical modulation.| (1) | (2) | Urea | |
|---|---|---|---|
| µtot (D) | 7.45 | 15.94 | — |
| αtot (×10−24 e.s.u.) | 12.64 | 13.80 | — |
| βtot (×10−31 e.s.u.) | 8.83 | 42.30 | 1.3 |
| Efficiency for SHG | 6.80 | 32.54 | 1 |
The nonlinear optical response of compound (2) is remarkably strong. The total dipole moment (µtot = 15.94 D) is significant, although slightly lower than that of compound (1). However, this is compensated by an exceptionally high total polarizability (αtot = 13.80 × 10−24 e.s.u.) and hyperpolarizability (βtot = 42.30 × 10−31 e.s.u.), which exceed those of urea by more than twenty-six times. The calculated SHG efficiency 32.54 times that of urea clearly demonstrates the excellent NLO potential of this material. These outstanding values can be directly linked to the presence of the highly polarizable iodide anion, the robust hydrogen-bonding network that aligns molecular dipoles, and the reduced HOMO–LUMO gap enabling effective charge-transfer under optical excitation.
Taken together, the structural, electronic, and optical results converge to show that compound (2) possesses a highly optimized arrangement for nonlinear optical applications. The synergy between the noncentrosymmetric crystal symmetry, strong cation–anion charge separation, intramolecular and intermolecular charge-transfer pathways, and the influence of the iodide anion leads to a material with exceptional NLO efficiency, making it a promising candidate for optical modulation devices, frequency-doubling components, and photonic technologies.
3.8. Structure–property relationships
Understanding how subtle structural variations in organic–inorganic hybrid salts dictate their optical, electronic, and biological behaviour is central to rational material design. The two amino-acid-based salts investigated in this work L-phenylalaninium chloride (1) and L-histidinium iodide (2) provide an excellent platform to elucidate structure–property relationships driven by cation identity, halide nature, hydrogen-bond topology, and supramolecular architecture. The key correlations derived from experimental and computational data are summarized below.In contrast, compound (2) features an expanded iodide-stabilized network in which the highly polarizable I− ion forms strong charge-assisted N–H⋯I interactions and permits greater structural flexibility. Coupled with the π-rich imidazolium ring, this environment enhances electronic delocalization, lowers the band gap, increases softness and electrophilicity, and strengthens the molecular polarization response under an applied field, ultimately leading to superior optical and biological performance. Overall, the results clearly demonstrate that halide size and polarizability (Cl− < I−) play a decisive role in modulating crystal packing and governing the electronic reactivity profile of these hybrid salts.
In compound (1), HOMO localization on the phenyl ring restricts intramolecular charge transfer (ICT), resulting in modest hyperpolarizability.45
In compound (2), however, HOMO delocalization across the imidazolium ring and an expanded LUMO facilitate a more efficient ICT mechanism, producing a first hyperpolarizability nearly three times higher. This pronounced enhancement reflects the combined influence of imidazolium protonation and the high polarizability of iodide, which together amplify electronic flexibility.
The electronic trends fully coincide with the supramolecular openness and halide-driven polarization observed crystallographically, establishing a strong and coherent structure–property correlation.
Compound (1) displays sharper ν(NH3+) and ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) absorptions, consistent with more localized and well-defined hydrogen bonds involving the chloride anion within its compact supramolecular framework.
O) absorptions, consistent with more localized and well-defined hydrogen bonds involving the chloride anion within its compact supramolecular framework.
In contrast, compound (2) exhibits broader NH stretching bands, attributable to stronger charge-assisted N–H⋯I interactions and the greater delocalization of electron density across the imidazolium ring.
These vibrational contrasts closely mirror the electronic properties of the materials, with compound (2) showing enhanced polarizability and compound (1) maintaining a more rigid structural arrangement, thereby demonstrating that vibrational fingerprints reliably reflect deeper electronic and supramolecular trends.
Compound (1), characterized by higher global hardness and lower softness, exhibits reduced chemical reactivity; its hydrophobic phenyl ring limits interactions with radical species, resulting in only moderate antioxidant and antibacterial performance, likely due to weaker interactions with microbial membranes.
In contrast, compound (2) displays lower hardness and higher softness, features that enhance its ability to participate in electron and proton transfer processes. The combination of a highly polarizable iodide anion and a protonated imidazolium ring promotes stronger electrostatic and hydrogen-bonding interactions with biological membranes, leading to markedly superior antioxidant and antibacterial activities. Overall, the electron-rich and polarizable nature of the imidazolium-iodide framework confers enhanced redox reactivity and facilitates membrane-disruptive mechanisms, thereby underpinning the enhanced biological properties of compound (2).
Although global electronic softness (η−1) is often associated with enhanced redox reactivity, the present results indicate that antioxidant activity in the studied hybrid salts cannot be rationalized solely on the basis of the HOMO–LUMO energy gap or global softness descriptors (see Table 2). Instead, a more meaningful correlation emerges when local electronic reactivity and structural accessibility are considered simultaneously.
DFT calculations reveal that the frontier orbitals of compound (1) are preferentially localized on chemically active donor sites involved in hydrogen atom or electron transfer processes, favoring efficient radical scavenging despite its larger band gap. In contrast, compound (2), although characterized by higher global softness, exhibits stronger intermolecular interactions and denser supramolecular packing, as evidenced by Hirshfeld surface and NCI/RDG analyses. This structural confinement partially limits the accessibility and effective reactivity of redox-active sites, reducing the biological response.
Furthermore, electrophilicity-related descriptors indicate a greater stabilization of the radical species formed after charge or hydrogen donation in compound (1), which compensates for its lower intrinsic softness. Consequently, the antioxidant activity correlates more strongly with localized frontier orbital density and electrophilicity than with the global HOMO–LUMO gap, highlighting the necessity of combining electronic descriptors with solid-state structural factors to accurately interpret biological activity.48–50
| Structural feature | Effect on property (1) | Effect on property (2) |
|---|---|---|
| Halide size | Compact packing | Expanded, polarized packing |
| Cation type | Phenyl (limited ICT) | Imidazolium (strong ICT) |
| Hydrogen bonding | Localized (Cl−) | Charge-assisted (I−) |
| Band gap | High | Low |
| NLO response | Moderate | High |
| Antioxidant/antibacterial | Moderate | Strong |
4. Conclusion
In this work, two new amino-acid-based hybrid salts L-phenylalaninium chloride (1) and L-histidinium iodide (2) were synthesized, structurally characterized, and investigated through a combined experimental and theoretical approach. Single-crystal X-ray diffraction revealed that both compounds crystallize in the non-centrosymmetric P212121 space group, yet their supramolecular architectures differ markedly due to the distinct chemical nature of their cations and counterions. Compound (1) forms a compact, chloride-stabilized hydrogen-bond network, while compound (2) exhibits a more open and highly polarized framework dominated by charge-assisted N–H⋯I interactions and enhanced π-delocalization from the imidazolium ring. These structural features directly govern their electronic and optical responses.DFT and TD-DFT analyses show that compound (2) displays a significantly lower HOMO–LUMO band gap, higher molecular softness, and stronger intramolecular charge-transfer pathways than compound (1). These characteristics translate into a markedly enhanced nonlinear optical response, with β values for compound (2) nearly three times higher. Vibrational data corroborate the crystallographic findings, with broader and more red-shifted NH stretching bands confirming the stronger hydrogen bonding and increased electronic polarization in compound (2).
Biological assays further highlight the influence of structural and electronic factors on functional behaviour. Compound (2) exhibits superior antioxidant (DPPH, ABTS, FRAP) and antibacterial activities, which are attributed to the high polarizability of the iodide ion, the redox-active imidazolium ring, and the greater propensity for electron/proton transfer. In contrast, compound (1) shows moderate activity, consistent with its more rigid supramolecular framework and larger band gap.
Overall, this study establishes clear structure–property relationships linking crystal packing, electronic descriptors, vibrational behaviour, optical nonlinearity, and biological performance. The results demonstrate that combining protonated heterocyclic amino acids with highly polarizable halides is an effective strategy for enhancing both NLO and biological properties in organic–inorganic hybrid salts. These findings provide valuable design principles for the development of next-generation multifunctional materials with potential applications in photonics, optoelectronics, and bioactive systems.
Author contributions
Sandra Walha: investigation, software. Rageh K. Hussein: formal analysis, writing – original draft, investigation. Thierry Roisnel: methodology, conceptualization, validation, and Ali Ben Ahmed: supervision, writing, review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
Publicly available datasets were analyzed in this study.CCDC 2504992 (1) and 1400108 (2) contain the supplementary crystallographic data for this paper.53a,b
Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2602).References
- V. V Ghazaryan, M. Fleck and A. M. Petrosyan, J. Mol. Struct., 2012, 1026, 140–144 CrossRef.
- M. Periasamy, M. Seenivasaperumal and V. D. Rao, Tetrahedron: Asymmetry, 2004, 15, 3847–3852 CrossRef CAS.
- G. Albano, G. Pescitelli and L. Di Bari, Chem. Rev., 2020, 120, 10145–10243 CrossRef CAS PubMed.
- R. Bano, M. Asghar, K. Ayub, T. Mahmood and J. Iqbal, Front. Mater., 2021, 8, 1–24 Search PubMed.
- R. M. F. Baptista, D. Santos, N. F. Cunha, M. C. R. Castro, P. V. Rodrigues, A. V. Machado, M. S. Belsley, E. de and M. Gomes, Novel Benzothiazole Boc-Phe-Phe-Bz Derivative Dipeptide Forming Fluorescent and Nonlinear Optical Self-Assembled Structures, Molecules, 2025, 30, 942–956 CrossRef CAS PubMed.
- Y. Fu, Z. Liu, S. Yue, K. Zhang, R. Wang and Z. Zhang, Nanomaterials, 2024, 14, 1–36 Search PubMed.
- J. A. Doolan, J. S. Fossey, B. T. Goult, J. R. Hiscock and J. S. Fossey, Chem. Soc. Rev., 2022, 51, 8696–8755 RSC.
- N. Joondan, S. Jhaumeer-laulloo and P. Caumul, Microbiol. Res., 2014, 1–11 Search PubMed.
- K. Je, Bioorg. Med. Chem., 2017, 25, 4265–4276 CrossRef PubMed.
- G. Zhao, M. Tian, R. Wang, Y. Deng, Y. Zhao, S. Luo and L. Chen, Sci. China:Chem., 2024, 4–11 Search PubMed.
- J. Wa, A. Hecel, M. Rowin and H. Kozłowski, Inorg. Chim. Acta, 2017, 12, 1–8 Search PubMed.
- L. Zhou, S. Li, Y. Su, X. Yi, A. Zheng and F. Deng, J. Phys. Chem. B, 2013, 8954–8965 CrossRef CAS PubMed.
- J. Lemire, Y. Milandu, C. Auger, A. Bignucolo, V. P. Appanna and V. D. Appanna, FEMS Microbiol. Lett., 2010, 309, 170–177 CAS.
- S. Walha, N. Zammel, M. Horchani, M. Bourwina, N. Mhadhbi, H. Wahbi, B. Basha, F. Aouaini and A. Tahiri, Polyhedron, 2025, 282, 117782 CrossRef CAS.
- S. Walha, A. Guesmi, N. Mhadhbi, S. Antonio, A. Ben Ahmed, L. Khezami, N. Ben Hamadi and H. Naïli, J. Mol. Struct., 2026, 1349, 1–13 CrossRef.
- S. Walha, N. Mhadhbi, N. Ktari, F. Aouaini, B. Basha, H. Wahbi, R. Ben Salah, F. Saadi, A. Ben Ahmed and H. Naïli, J. Mol. Struct., 2025, 1325, 140984 CrossRef CAS.
- G. M. Sheldrick, SHELXT – Integrated space-group and crystal-structure determination, Found. Adv., 2014, 71, 3–5 CrossRef PubMed.
- G. M. Sheldrick, Acta Crystallogr., Sect. A: Found. Crystallogr., 2008, 64, 112–122 CrossRef CAS PubMed.
- C. F. Macrae, I. J. Bruno, J. A. Chisholm, P. R. Edgington, P. McCabe, E. Pidcock, L. Rodriguez-Monge, R. Taylor, J. Van De Streek and P. A. Wood, J. Appl. Crystallogr., 2008, 41, 466–470 CrossRef CAS.
- H. Putz and K. Brandenburg, Sci. Comput. World, 2002, 63, 19–21 Search PubMed.
- P. R. Spackman, M. J. Turner, J. J. McKinnon, S. K. Wolff and D. J. Grimwood, J. Appl. Crystallogr., 2021, 54, 1006–1011 CrossRef CAS PubMed.
- J. B. Frisch, M. J., G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman; G. Scalmani; V. Barone; G. A. Petersson; H. Nakatsuji; X. Li; M. Caricato; A. V. Marenich; J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci and H. P. Hratchian, Gaussian 16, 2016, https://gaussian.com/gaussian16/ Search PubMed.
- A. D. Becke, J. Chem. Phys., 1996, 104, 1040–1046 CrossRef CAS.
- A. V. Mitin, J. Comput. Chem., 2013, 34, 2014–2019, DOI:10.1002/jcc.23353.
- R. Dennington, T. Keith, J. Millam, Semichem Inc., Shawnee Mission, 2009.
- A. Moumad, A. Bouhaoui, M. Eddahmi, A. Hafid, L. R. Domingo and L. Bouissane, ChemistrySelect, 2023, 8(14), e202300669–e202300679, DOI:10.1002/slct.202300669.
- M. Rahmani, A. Ghasemi, F. Moattar and A. Reza, Ind. Crops Prod., 2019, 137, 1–8 CrossRef.
- M. Prakash, D. Geetha and M. L. Caroline, Spectrochim. Acta, Part A, 2011, 81, 48–52 CrossRef CAS PubMed.
- R. Surekha, R. Gunaseelan, P. Sagayaraj and K. Ambujam, CrystEngComm, 2014, 7979–7989 RSC.
- A. Ben Ahmed, H. Feki, Y. Abid, H. Boughzala and C. Minot, Spectrochim. Acta, Part A, 2010, 75, 293–298 CrossRef CAS PubMed.
- I. Stroganova, S. Bakels and A. M. Rijs, Molecules, 2022, 27(7), 2367–2385, DOI:10.3390/molecules27072367.
- B. G. Carvalho, L. J. Raniero, A. A. Martin and P. P. Favero, Spectrochim. Acta, Part A, 2013, 106, 73–79 CrossRef CAS PubMed.
- A. Hollmann, M. Martinez, P. Maturana and L. C. Semorile, Front. Chem., 2018, 6, 1–13 Search PubMed.
- Q. Y. Zhang, Z. Bin Yan, Y. M. Meng, X. Y. Hong, G. Shao, J. J. Ma and X. R. Cheng, Mil. Med. Res., 2021, 1–25 Search PubMed.
- G. Bacteria, A. Gauba and K. M. Rahman, Antibiotics, 2023, 12, 1590–1605 CrossRef PubMed.
- O. Oancea, C. N. Ciurea, A. D. Mare, A. Man, R. Stefanescu and A. Rusu, Antibiotics, 2025, 14, 415–440 CrossRef CAS PubMed.
- J. Pernak, K. Sobaszkiewicz and I. Mirska, Green Chem., 2003, 5, 52–56 RSC.
- K. M. Docherty and C. F. Kulpa Jr., Green Chem., 2005, 7, 185 RSC.
- R. M. Epand and R. F. Epand, J. Pept. Sci., 2011, 17, 298–305 CrossRef CAS PubMed.
- H. Akram, B. Arshad, M. Imtiaz, W. Ahmed, M. Shafiq, Z. Ul-haq, R. Khan, A. Bahadar, B. Ahmad and M. Iqbal, BMC Chemistry, 2025, 19, 220 CrossRef CAS PubMed.
- L. P. Hu, Chunli Liu and Jianggao, Inorg. Chem. Front., 2025, 12, 4306–4311 RSC.
- J. Hu, J. Wang, S. Li, B. Yang, M. Gong, X. Li, L. Zhang and J. Tian, J. Appl. Bot. Food Qual., 2016, 89, 150–155 CAS.
- F. Miccichè, The Combination of Ascorbic Acid Derivatives/Iron Salts as Catalyst for the Oxidative Drying of Alkyd-Based Paints: A Biomimetic Approach, 2005 Search PubMed.
- J. Bojarska, K. Łyczko and A. Mieczkowski, Crystals, 2024, 14, 133 CrossRef CAS.
- P. Long, C. Hu, X. Liu and J. Mao, Inorg. Chem. Front., 2025, 12, 4306–4311 RSC.
- J. Bojarska, K. Łyczko and A. Mieczkowski, Crystals, 2024, 14, 497 CrossRef CAS.
- E. V. Ivanov, E. Y. Lebedeva, V. V. Baranov and A. N. Kravchenko, J. Mol. Liq., 2023, 384, 122234 CrossRef CAS.
- A. Sarkar, T. R. Middya and A. D. Jana, J. Mol. Model., 2012, 18, 2621–2631 CrossRef CAS PubMed.
- A. H. Bakheit, T. A. Wani, A. A. Al-Majed, H. M. Alkahtani, M. M. Alanazi, F. R. Alqahtani and S. Zargar, Front. Chem., 2024, 12, 1443718–1443734, DOI:10.3389/fchem.2024.1443718.
- Y. Chemam, S. Benayache, A. Bouzina, E. Marchioni, O. Sekiou, H. Bentoumi, M. Zhao, Z. Bouslama, N.-E. Aouf and F. Benayache, RSC Adv., 2024, 14, 22209–22228 RSC.
- S. Hafeez, Z. Rasool, S. Hafeez, R. Z. Paracha, M. Iqbal, D. Khan and F. Adnan, Heliyon, 2024, 10, e39829 CrossRef CAS PubMed.
- Y. Cheng, F. Luo, Z. Zeng, L. Wen, Z. Xiao, H. Bu, F. Lv, Z. Xu and Q. Lin, Struct. Chem., 2015, 26, 739–747 CrossRef CAS.
- (a) CCDC 2504992: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2q2n6p; (b) CCDC 1400108: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc1hzxs5.
| This journal is © The Royal Society of Chemistry 2026 |