Open Access Article
Open Access ArticleDiscovery of thieno[3,2-c]pyran-based leads for liver, lung and glioma cancer: synthesis, docking-guided optimization and apoptotic profiling
Abhishek Jauharia,
Poojab,
Lakshay Tanejac,
Ajay Kumar Yadavc,
Ismail Althagafid,
Ramendra Pratap *e and
Dharmendra Kumar Yadav
*e and
Dharmendra Kumar Yadav *f
*f
aDepartment of Neurological Surgery, University of Pittsburgh, 15213, USA
bDepartment of Chemistry, Sri Venkateshwara College, Delhi, 110021, India
cDr B. R. Ambedkar Center for Biomedical Research, University of Delhi, North Campus, Delhi, 110007, India
dDepartment of Chemistry, Faculty of Science, Umm Al-Qura University, Makkah, 21955, Saudi Arabia
eDepartment of Chemistry, University of Delhi, North Campus, Delhi, 110007, India. E-mail: rpratap@chemistry.du.ac.in; ramendrapratap@gmail.com
fDepartment of Biologics, College of Pharmacy, Gachon University, Hambakmoeiro 191, Yeonsu-gu, Incheon 21924, Republic of Korea
First published on 20th April 2026
Abstract
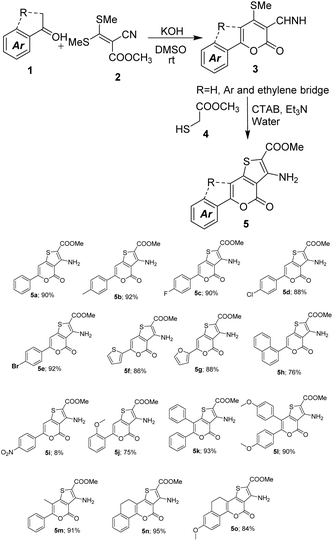
Thieno[3,2-c]pyran is a versatile heterocyclic framework recognised as a core scaffold in numerous synthetic anticancer agents and naturally occurring alkaloids exhibiting potent antitumor activity across various malignancies, including lung, liver, breast, and prostate cancers. In this study, we report the synthesis of a series of substituted thieno[3,2-c]pyran derivatives and their biological evaluation against human hepatocellular carcinoma (HepG2) and lung adenocarcinoma (A549) cell lines. The in vitro cytotoxicity assays revealed promising IC50 values: 5a exhibited 13.65 ± 0.32 µM (HepG2) and 12.35 ± 0.30 µM (A549), while 5n showed 13.26 ± 0.32 µM (HepG2) and 12.36 ± 0.30 µM (A549). Apoptosis-induction studies confirmed the anticancer potential of these compounds. However, compound 5e works well against Glioma cancer with a promising IC50 value of 26 µM (LN229). We have also performed the testing of selected compounds against the non-cancerous cell line HEK293 under the same conditions and concentration, and no cytotoxicity. Among the synthesized compounds, 5a and 5n demonstrated significant binding affinities in molecular docking studies, with docking scores of −8.834 and −8.994 kcal mol−1 against liver cancer targets, and −7.925 and −7.871 kcal mol−1 against lung cancer targets, respectively. To further confirm the EGFR inhibition, selected compounds were tested agaist EGFR overexpressed glioma cell line, and good cytotoxicity confirms the proposed pathway. These findings suggest that compounds 5a, 5e and 5n warrant further investigation as lead molecules for the development of targeted therapies for liver, glioma and lung cancers.
1 Introduction
Cancer, characterized by the abnormal growth of cells in the body, is a major global health concern. According to the American Cancer Society (ACS), it is estimated that cancer causes approximately 1700 deaths daily, contributing to around 15% of all human deaths worldwide. Cancer cells often evade apoptosis, leading to unchecked growth and angiogenesis—hallmarks of all cancer types.1–4 Apoptosis is essential for preventing cancer, and targeting this process is a highly effective cancer treatment strategy. Many anticancer drugs focus on various stages of the intrinsic and extrinsic apoptotic pathways.5,6Lung cancer is one of the most prevalent cancers, with the highest rates of morbidity and mortality globally. Despite existing chemotherapeutic treatments, many are inadequate due to serious side effects and drug resistance.7 Therefore, developing safe and effective new treatments for lung cancer is critical. According to global cancer statistics from 2018, lung cancer accounts for 11.6% of all diagnosed cancer cases and is the leading cause of cancer death, responsible for 18.4% of all cancer-related deaths. Lung cancer is primarily classified into two histological groups: Small Cell Lung Cancer (SCLC) and Non-Small Cell Lung Cancer (NSCLC), with NSCLC accounting for approximately 85% of lung cancers. NSCLC is further divided into adenocarcinoma, squamous cell carcinoma, and large cell carcinoma.
A significant challenge in treating NSCLC is that when one of the targeted pathways in signal transduction is blocked, others can serve as escape mechanisms for cancer cells.8–11
Thienopyran is a prominent heterocyclic structure that serves as the foundation for many pharmacologically significant compounds, particularly those with anticancer properties.11 Fused pyran12–19 derivatives have shown promising results in cancer treatment by targeting cellular pathways that inhibit cancer cell growth, induce apoptosis, and disrupt vital processes necessary for tumour progression. These compounds also exhibit a range of pharmacological activities, including antimicrobial, antitubercular, antioxidant, analgesic, antileishmanial, antiplatelet, FPR1 antagonist, and anticonvulsant properties.20–26 Among these, the chromene analogues EPC2407 (Crolibulin, I) stand out as a promising agent for disrupting blood vessels in tumours and inducing apoptosis in advanced solid tumours. It has shown effectiveness in phase I clinical trials for patients with advanced thoracic and abdominal tumours and is currently in phase I/II trials in combination with cisplatin for anaplastic thyroid cancer (ATC). Similarly, MX58151 (II) has been identified as a caspase activator and tubulin inhibitor, while SP-6-27 (III) demonstrates high potency against glioma, melanoma, and prostate cancer cell lines due to its antiproliferative activity.18,19 Other chromene analogues, such as LY290181 (IV), fluorine-containing pyrano-chromenes (V) and (VI), and indole-substituted tetrahydro-chromene (VII) [Fig. 1], have also been reported as potent anticancer agents.27,28 Despite progress in developing chemotherapeutic agents, the development of new, target-specific anticancer drugs with minimal side effects to address drug resistance is always required.29–31 Various thiophene-fused heterocycles are explored for anticancer activity and show very good activity32 using different targets39 (Table 1). The thieno[3,2-c]pyran nucleus is well explored for its photophysical properties, role as a metal sensor, such turn on sensor for Zn2+.11d Additionally, this class of compounds are also studied for their synergistic antibacterial properties.11e The anticancer properties of this compound were not explored, so we decided to explore the anticancer activity, as it contains both thiophene and pyran rings fused in the same nucleus.
| No. | Thieno-fused molecule39 | Biological target and/or activity |
|---|---|---|
| 1 | Thieno[3,2-d]pyrimidine | CDK inhibition; apoptosis induction |
| 2 | Thieno[2,3-d]pyrimidine | EGFR and DHFR inhibition and anticancer cytotoxicity |
| 3 | Thieno[2,3-d][1,2,4]triazolo-pyrimidine hybrids | Anticancer activity via kinase inhibition |
| 4 | Oxadiazole-incorporated thieno heterocycles | Cytotoxic activity against A549 lung cancer cells |
| 5 | Thieno[3,2-e]pyrrolo[1,2-a]pyrimidine | Antitumor activity in pulmonary metastatic models |
| 6 | Thieno[3,2-c]pyran | Antibacterial activity, anticancer (this work) |
The structural flexibility of fused pyran derivatives allows the synthesis of compounds with customised properties, optimising their anticancer efficacy and minimising off-target effects.32,33
By introducing diverse functional groups and modifying specific regions of the fused pyran scaffold, researchers have improved potency and pharmacokinetics, resulting in better bioavailability and target specificity. Consequently, a series of functionalized thieno[3,2-c]pyran derivatives has been synthesized to evaluate their anticancer efficacy against various cancer cell lines, study their mechanisms of action, and conduct in silico ADMET predictions to assess their drug-likeness properties.18,19
2 Result and discussion
2.1 Chemistry
We used the precursor and synthesized the thieno[3,2-c]pyran36,37 using triethylamine as a base under a green approach. We synthesized various 6-aryl/5,6-diaryl-thieno[3,2-c]pyrans 5a–l, 6-aryl-5-methyl-thieno[3,2-c]pyrans 5m and methyl 1-amino-11-oxo-5,11-dihydro-4H-benzo[h]thieno[3,2-c]chromene-2-carboxylates 5n and 5o by treatment of synthesized precursor with methyl thioglycolate. All reactions were carried out in water using CTAB as surfactant and Et3N as a base at 85 °C. The reaction was heated at the given temperature for 2–4 hours, depending on the derivatives. After completion of the reaction, the obtained precipitate was filtered and washed with water. The compounds were dried and recrystallized from ethanol and further tested.
Considering the structure-activity relationship (SAR), we synthesized a series of thienopyran derivatives with various substitutions on the thiophene and pyran rings. We have incorporated different groups such as –NO2, alkyl, alkoxy, esters, amines and halides to study their outcome on activity.
Additionally, this study represents the first exploration of these compounds as potential anti-cancer agents. We introduced substitutions at the 6th position of the pyran ring with an aromatic group and at the 4th and 5th positions of the thiophene ring with –NH2 and –COOMe, respectively (Fig. 2). It was found that –NH2 and –COOMe substitutions provided basic binding sites that enhanced biological activity. Modifications to the pyran ring revealed that compounds with an aromatic group alone at the 6th position demonstrated effective IC50 values, as illustrated by 5a and 5n.
2.2 Biological study
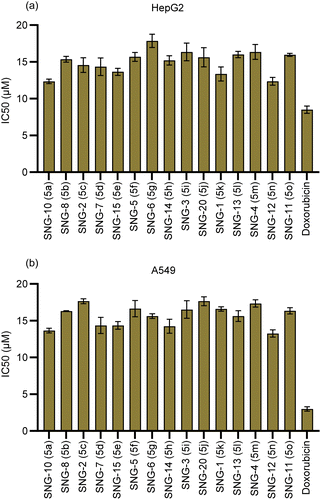 | ||
| Fig. 3 (a) Cytotoxic effects of synthesized compounds (5a–o) on HepG2; (b) cytotoxic effects of synthesized compounds (5a–o) on A549 cells. | ||
Cell viability was assessed after 48 h of treatment with the indicated compounds at varying concentrations. The percentage of viable cells was calculated relative to untreated controls. IC50 values (concentration causing 50% inhibition of cell viability) were determined. Data represent mean ± SD of three independent experiments performed in triplicate. Compounds 5a and 5n exhibited the highest cytotoxicity, with the lowest IC50 values in both (a) HepG2 and (b) A549 cells.
In summary, compounds 5a and 5n exhibited the strongest cytotoxic effects among the synthesized compounds with low micromolar IC50 values in both HepG2 and A549 cells. Cytotoxicity is likely mediated by G1 phase cell cycle arrest, with apoptosis contributing partially to the antiproliferative effect, particularly for compound 5a. These findings identify 5a as a promising lead for further mechanistic and therapeutic studies.
Quantification of cell viability expressed as a percentage of DMSO control is presented in Figure B. Treatment with all compounds at 26 µM resulted in a significant reduction in cell viability, with values ranging from approximately 40% to 60% relative to the vehicle control. Statistical analysis confirmed that all compound-treated groups showed highly significant reductions in cell viability compared to DMSO (***p < 0.001), indicating potent cytotoxic activity against LN229 cells at this concentration.
Among the PBS dissolved compounds, 5d and 5e (PBS pre-incubated formulations) demonstrated the most pronounced cytotoxicity, reducing cell viability to approximately 40–43%, suggesting near-maximal growth inhibition at 26 µM. In contrast, compounds 5a and 5n show relatively moderate cytotoxicity, with viability values in the range of 55–60%, though still significantly different from the control.
A similar trend was observed when compound 5a was pre-incubated in fetal calf serum (FCS)-containing media under identical conditions (FCS-48 and FCS-72). The FCS-pre-incubated samples likewise demonstrated statistically significant cytotoxicity compared to the DMSO control (p < 0.001), with a comparable 5–10% attenuation in antiproliferative activity relative to the freshly dissolved compound. Collectively, these results demonstrate that compound 5a exerts significant cytotoxic activity against EGFR-expressing LN229 glioblastoma cells at 13 µM, and that this activity is largely preserved following pre-incubation under both aqueous and serum-containing conditions for up to 72 hours, supporting the stability and translational potential of the compound under physiological conditions.
The stability of compounds was also confirmed by mass spectrometry. We have recoded the chromatogram of compound 5a after treatment with PBS for 48 and 72 h at 26 °C. Each chromatogram shows no decomposition of compounds and suggests their stability during testing against the cell line (see SI; Fig. S2–S4).
 | ||
| Fig. 8 HEK293 Cell images taken after incubation with MTT reagent at 565 nm, cell number is the same, but the intensity of crystal formation is higher in cells treated with 5a. | ||
Overall, the data indicate that compounds 5a, 5d, 5e, and 5n are non-toxic to HEK293 cells under the specified conditions and may promote cell proliferation or increased metabolic activity.
Compound 5a exhibited favourable binding to EGFR, with a docking score of −7.925. A hydrogen bond (3.25 Å) was observed with Cys773, a polar residue. The binding pocket consisted of hydrophobic residues (Ala719, Leu764, Leu820, Leu834, and Val702), basic residues (Lys721, Gly772), acidic residues (Asp86), and polar uncharged residues (Thr766). These interactions indicate a stable and specific binding conformation, illustrated in Fig. 9(a) and (b).
Similarly, compound 5n demonstrated a high docking score of −7.871 with EGFR. It formed a hydrogen bond (2.0 Å) via a bridging water molecule, enhancing binding specificity. The residues surrounding the binding site included Asp831 (acidic), Thr830 (polar), Lys721, Gly772 (basic), Val702, Leu794, Leu768, Leu820 (hydrophobic), and Met769 (nonpolar). This configuration supports a robust interaction profile, detailed in Fig. 10(c) and (d).
Compound 5a displayed a strong binding affinity toward VEGFR-2, yielding a Glide docking score of −8.834. It formed a key hydrogen bond (2.84 Å) with the thiol group of Cys919, a polar uncharged residue. The residues within 3 Å of the ligand included a combination of aromatic (Phe918), hydrophobic (Val916, Val899, Val848, Ala866, Leu840, Leu1035), basic (Lys920, Lys868, Gly922), acidic (Glu917, Asp1046), and polar amide residues (Cys1045). This network of hydrophobic and polar interactions contributed to the stable binding conformation observed in Fig. 10 (a) and (b).
In comparison, compound 5n showed an even higher affinity with a docking score of −8.994. A notable π–π stacking interaction (2.7 Å) was observed with Phe918, further stabilizing the complex. The surrounding pocket was similarly composed of hydrophobic (Val916, Val899, Val848, Ala866, Leu840, Leu1035), basic (Lys920, Gly922), acidic (Glu917), and polar (Cys919) residues, facilitating strong hydrophobic interactions and potential functional activity, as depicted in Fig. 10(c) and (d).
As reported, compounds 5a and 5n exhibited the most potent antiproliferative effects among the tested series, with very similar IC50 values in both HepG2 and A549 cell lines. In HepG2 cells, compound 5a showed an IC50 of 13.65 ± 0.32 µM, while 5n exhibited an IC50 of 13.26 ± 0.32 µM. Likewise, in A549 cells, the IC50 values were 12.35 ± 0.30 µM for 5a and 12.36 ± 0.30 µM for 5n. These differences are minimal and fall within experimental variability, indicating that both compounds possess comparable cytotoxic potency at the cellular level.
Although compound 5n demonstrated a slightly better VEGFR-2 docking score, cytotoxic activity reflects an integrated cellular response rather than binding affinity alone. The comparable IC50 values suggest that both compounds achieve similar levels of growth inhibition, while differences in apoptotic induction may arise from factors beyond receptor binding, such as intracellular uptake, metabolic stability, or differential engagement of downstream signalling pathways. Importantly, compound 5a showed consistent and robust cytotoxicity across both cell lines, which may explain its relatively stronger apoptotic response despite a marginally lower docking score.
The standard drug Doxorubicin, used as a control, displayed comparatively lower binding affinities with VEGFR-2 and EGFR, scoring −7.157 and −5.252, respectively. These results underscore the enhanced binding potential of the novel compounds 5a and 5n over the reference compound, reinforcing their candidacy for further biological evaluation.
3 Experimental section
3.1 Materials
Commercially available reagents and solvents from TCI Chemicals, Sigma Aldrich, and Alfa Aesar were used without further purification. Reactions were monitored by analytical thin-layer chromatography (TLC) and were conducted in dry glassware. Infrared (IR) spectra were recorded using a PerkinElmer AX-1 spectrophotometer, with stretching frequencies reported as wave numbers (cm−1). 1H and 13C NMR spectra were acquired on 400 MHz and 100 MHz NMR spectrometers, respectively. CDCl3 (from Eurisotop) was used as the solvent to obtain the NMR data and chemical shifts reported in parts per million (ppm) relative to CDCl3, where δ 7.26 ppm and δ 77.00 ppm served as internal standards for 1H and 13C NMR, respectively. Additionally, DMSO-d6 (from Eurisotop) was used as a solvent, with chemical shifts referenced in ppm relative to DMSO-d6, where δ 2.50 ppm and δ 39.52 ppm were used as internal standards for 1H and 13C NMR, respectively. NMR signal patterns are designated as s (singlet), d (doublet), dd (doublet of doublets), t (triplet), m (multiplet), and bs (broad singlet). Extraction and chromatography were performed with reagent-grade solvents, and yields are reported for chromatographically pure materials.3.2 Synthesis of compounds
3.3 Cell line study
Dose–response curves and half-maximal cytotoxic concentration (IC50) values were determined by nonlinear regression (log[inhibitor] vs. response − variable slope) using GraphPad Prism. Data are presented as ± SEM from three independent experiments.
3.4 Cell cycle and apoptosis assays
3.5 MTT cell viability assay against LN229 and HEK293 cell lines40
Cell viability was assessed using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) colorimetric assay. LN229 and HEK293 cells were harvested and seeded in 96-well flat-bottom tissue culture plates at a density of 5 × 103 cells per well in a final volume of 100 µL complete DMEM per well. Cells were allowed to adhere and equilibrate overnight at 37 °C with 5% CO2 before treatment. Following overnight incubation, the culture medium was replaced with 100 µL of fresh serum-free DMEM containing dilutions of the compounds at concentrations 26 µM, 13 µM, 6.5 µM. Test compounds were prepared as stock solutions in dimethyl sulfoxide (DMSO) and further diluted in DMEM. After 48 hours, the medium was carefully aspirated from each well without disturbing the cell monolayer. Subsequently, 10 µL of the MTT stock solution (5 mg mL−1) was added to each well along with 90 µL of serum-free DMEM, to a final MTT concentration of 1 mg mL−1 per well. Plates were incubated at 37 °C for 4 hours in dark. The insoluble purple formazan crystals formed in each well were dissolved by adding 200 µL of DMSO to each well. Plates were placed 37 °C for 20 minutes. The absorbance was measured at 565 nm in ELISA plate reader (TECAN).3.6 Molecular docking
The chemical structures of the lead compounds, 5a and 5n, were initially constructed in ChemDraw (.cdx format) and subsequently converted to the Maestro-compatible .maegz format using Maestro version 11.8 (Schrödinger, LLC). Molecular docking studies were conducted using the Glide module. High-resolution crystal structures relevant to lung (PDB ID: 4HJO)40,41 and liver cancer (PDB ID: 4ASD)42 were retrieved from the Protein Data Bank. These protein models were pre-processed using the Protein Preparation Wizard in Maestro, which included hydrogen addition, assignment of appropriate protonation states, removal of water molecules, and energy minimization utilizing the OPLS-2005 force field. Ligand preparation was carried out using the LigPrep tool, with Epik used to generate ionization states at a physiological pH of 7.0, applying the same OPLS-2005 force field. Docking simulations were executed using Glide in Extra Precision (XP) mode to ensure accurate binding predictions. The resulting docked complexes were visualized and analysed with PyMOL version 2.5.434 Conclusions
In this study, we synthesized a series of compounds (5a–o) and evaluated their cytotoxic potential against human hepatocellular carcinoma (HepG2) and lung carcinoma (A549) cell lines. Among all tested compounds, 5a and 5n exhibited the best antiproliferative activity, with IC50 values in the low micromolar range. The comparable activity in both HepG2 and A549 cells suggests that these compounds possess broad cytotoxic effects against distinct cancer types, highlighting their potential as lead molecules for further anticancer development. Additionally, compound 5e also exhibits good activity against Glioma cancer with an IC50 of 26 µM. Further study of these compounds is going on.The mechanism underlying the observed cytotoxicity was explored through cell cycle analysis. Both 5a and 5n induced significant G1 phase arrest in HepG2 cells, indicating that these compounds interfere with the normal progression of the cell cycle. G1 arrest is often associated with inhibition of DNA synthesis and the activation of cellular checkpoints, which can slow proliferation and sensitize cells to apoptotic signals. Our findings suggest that disruption of cell cycle machinery is a key contributor to the growth-inhibitory effects of these compounds.
Further investigation into apoptosis induction revealed a differential effect between the two compounds. While 5a induced apoptosis in approximately 12–14% of cells, 5n showed a relatively modest apoptotic effect (6–7%). These results indicate that, although both compounds can arrest the cell cycle, compound 5a more effectively triggers programmed cell death, which may explain its slightly superior antiproliferative activity. The partial contribution of apoptosis to overall cytotoxicity suggests that G1 arrest and apoptosis act in concert to inhibit cell growth, with cell cycle arrest likely being the primary mechanism.
Further, the testing of selected compounds against the normal cell line HEK293 under the same conditions and concentration shows no cytotoxicity. We also confirmed the role of EGFR inhibition in the proposed study. The selected compounds were tested against the EGFR-expressing glioma cell line, and good cytotoxicity confirms that the proposed pathway is probable.
In conclusion, this study demonstrates that compounds 5a and 5n are effective cytotoxic agents, with compound 5a showing the most promising activity. The ability of these compounds to inhibit proliferation through cell cycle arrest and apoptosis provides a strong basis for further optimization and development as potential anticancer therapeutics.
Author contributions
A.J. was involved in an apoptotic study and D.K.Y. was involved in a molecular docking study. R.P. was involved in designing the concept, the synthetic protocols and writing the manuscript. Pooja and IA were involved in the synthesis and writing of the manuscript. L.T. and A.K.Y. was involved in further study of compounds on normal and glioma cell lines and further biological study. They are also involved in manuscript preparation.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Data for these compounds are reported in Biomedicine & Pharmacotherapy, 2021, 142, 112084. RSC Advances 2016, 6, 85515–85520. RSC Adv., 2025, 15, 12117–12124. Supplementary information: 1H and 13C spectral data and spectra of compound 5a–o, LN229 cell images taken after incubation with MTT, dose response curves and IC50 plots for all the synthesized compounds. See DOI: https://doi.org/10.1039/d5ra09341d.Acknowledgements
Pooja thanks ANRF for providing the research funding (EEQ/2023/000121) and Sri Venkateswara College, University of Delhi, for providing research lab space and facility. RP thanks CSIR, New Delhi, for research funding via project number 02/0469/23/EMR II. RP thanks AvH Foundation, Germany, for providing funding under the alumni visit program. RP thanks IOE for providing research funding during 2025–26.Notes and references
- R. L. Siegel, K. D. Miller and A. Jemal, Ca-Cancer J. Clin., 2019, 69, 7 CrossRef PubMed.
- R. L. Siegel, K. D. Miller and A. Jemal, Ca-Cancer J. Clin., 2016, 66, 7 CrossRef PubMed.
- H. Sung, J. Ferlay, R. L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal and F. Bray, Ca-Cancer J. Clin., 2021, 71, 209 CrossRef PubMed.
- A. Bistrović Popov, R. Vianelo, P. Grbčić, M. Sedić, S. K. Pavelić, K. Pavelić and S. Raić-Malić, Molecules, 2021, 26, 3334 CrossRef PubMed.
- C. Pfeffer and A. Singh, Int. J. Mol. Sci., 2018, 19, 448 CrossRef.
- A. Kamal, A. Mallareddy and P. Suresh, Heterocyclics as Inducers of Apoptosis, Springer, 2012, p. 45 Search PubMed.
- H. Zhou, S. Wu, S. Zhai, A. Liu, Y. Sun, R. Li, Y. Zhang, S. Ekins, P. W. Swaan, O. B. Fang, B. Zhang and B. Yan, J. Med. Chem., 2008, 51, 1242 CrossRef CAS PubMed.
- W. G. Kaelin, Nat. Rev. Cancer, 2005, 5, 689 CrossRef CAS.
- A. T. Fojo, K. Ueda, D. J. Slamon, D. G. Poplack, M. M. Gottesman and I. Pastan, Proc. Natl. Acad. Sci. U. S. A., 1987, 84, 265 CrossRef CAS.
- M. M. Gottesman, T. Fojo and S. E. Bates, Nat. Rev. Cancer, 2002, 2, 48 CrossRef CAS.
- (a) N. Kartner, J. R. Riordan and V. Ling, Science, 1983, 221, 1285 CrossRef CAS PubMed; (b) S. N. Sahu, M. K. Gupta, T. Jadhav, P. Yadav, S. Singh, R. Misra and R. Pratap, RSC Adv., 2014, 4, 56779–56783 RSC; (c) S. N. Sahu, S. Singh, R. Shaw, V. J. Ram and R. Pratap, RSC Adv., 2016, 6, 85515–85520 RSC; (d) D. Singhal, I. Althagafi, A. Kumar, S. Yadav, A. K. Prasad and R. Pratap, New J. Chem., 2020, 44, 12019–12026 RSC; (e) G. R. Dwivedi, R. Rai, R. Pratap, K. Singh, S. Pati, S. N. Sahu, R. Kant, M. P. Darokar and D. K. Yadav, Biomed. Pharmacother., 2021, 142, 112084 CrossRef CAS PubMed.
- K. Fabitha, A. Kallingal, N. Maciejewska, C. G. Arya, M. Chandrakanth, N. M. Thomas, Y. Li, R. Gondru, M. Munikumar and J. Banothu, New J. Chem., 2024, 48, 8038 RSC.
- F. M. Dean, Naturally Occurring Oxygen Ring Compounds, Butterworth-Heinemann, 1963, p. 176 Search PubMed.
- S. A. Patil, R. Patil, L. M. Pfeffer and D. D. Miller, Future Med. Chem., 2013, 5, 1647 CrossRef CAS.
- V. Raj and J. Lee, Front. Chem., 2020, 8, 1 Search PubMed.
- M. K. Katiyar, G. K. Dhakad, Shivani, S. Arora, S. Bhagat, T. Arora and R. Kumar, J. Mol. Struct., 2022, 1263, 133012 CrossRef CAS.
- B. Borah, K. D. Dwivedi and L. R. Chowhan, Polycyclic Aromat. Compd., 2022, 42, 5893 CrossRef CAS.
- M. S. Malik, H. Ather, S. M. Asif Ansari, A. Siddiqua, Q. M. S. Jamal, A. H. Alharbi, M. M. Al-Rooqi, R. S. Jassas, E. M. Hussein, Z. Moussa, R. J. Obaid and S. A. Ahmed, Pharmaceuticals, 2023, 16, 333 CrossRef CAS.
- W. Kemnitzer, J. Drewe, S. Jiang, H. Zhang, C. Crogan-Grundy, D. Labreque, M. Bubenick, G. Attardo, R. Denis, S. Lamothe, H. Gourdeau, B. Tseng, S. Kasibhatla and X. C. Sui, J. Med. Chem., 2008, 51, 417 CrossRef CAS PubMed.
- N. R. Kamdar, D. D. Haveliwala, P. T. Mistry and S. K. Patel, Med. Chem. Res., 2011, 20, 854 CrossRef CAS.
- G. Singh, A. Sharma, H. Kaur and M. P. S. Ishar, Chem. Biodiversity, 2016, 87, 213 CAS.
- C. Bingi, E. Narender Reddy, M. Chennapuram, Y. Poornachandra, C. G. Kumar, N. Jagadeesh Babu and K. Atmakur, Bioorg. Med. Chem. Lett., 2015, 25, 1915 CrossRef CAS PubMed.
- N. D. Vala, H. H. Jardosh and M. P. Patel, Chin. Chem. Lett., 2016, 27, 168 CrossRef CAS.
- E. Rajanarendar, M. Nagi Reddy, S. Rama Krishna, K. Rama Murthy, Y. N. Reddy and M. V. Rajam, Eur. J. Med. Chem., 2012, 55, 273 CrossRef CAS PubMed.
- N. C. Lazzara, R. J. Rosano, P. P. Vagadia, M. T. Giovine, M. W. Bezpalko, N. A. Piro, W. S. Kassel, W. J. Boyko, D. L. Zubris, K. K. Schrader, D. E. Wedge, S. O. Duke and R. M. Giuliano, J. Org. Chem., 2019, 84, 666 CrossRef CAS PubMed.
- J. H. Roireau, R. J. Rosano, N. C. Lazzara, T. Chen, J. Bajsa-Hirschel, K. K. Schrader, S. O. Duke, D. Wykoff and R. M. Giuliano, J. Agric. Food Chem., 2020, 68, 9906 CrossRef CAS PubMed.
- D. Rajguru, B. S. Keshwal and S. Jain, Med. Chem. Res., 2013, 22, 5934 CrossRef CAS.
- K. Parthasarathy, C. Praveen, C. Balachandran, P. Senthil Kumar, S. Ignacimuthu and P. T. Perumal, Bioorg. Med. Chem. Lett., 2013, 23, 2708 CrossRef CAS.
- L. Braconi, E. Teodori, C. Riganti, M. Coronnello, A. Nocentini, G. Bartolucci, M. Pallecchi, M. Contino, D. Manetti, M. N. Romanelli, C. T. Supuran and S. Dei, J. Med. Chem., 2022, 65, 14655 CrossRef CAS PubMed.
- S. Wang, S. Q. Wang, Q. Teng, L. Yang, Z. Lei, X. Yuan, J. Huo, X. Chen, M. Wang, B. Yu, Z. Chen and H. Liu, J. Med. Chem., 2020, 63, 15979 CrossRef CAS PubMed.
- D. J. Hong, S. H. Jung, J. Kim, D. Jung, Y. G. Ahn, K. H. Suh and K. H. Min, J. Enzyme Inhib. Med. Chem., 2020, 35, 227 CrossRef CAS.
- M. M. Heravi and V. Zadsirjan, RSC Adv., 2020, 10, 44247 RSC.
- (a) N. Kerru, L. Gummidi, S. Maddila, K. K. Gangu and S. B. Jonnalagadda, Molecules, 2020, 25, 1909 CrossRef CAS PubMed; (b) S. N. Sahu, R. Shaw, S. Yadav, I. Althagafi, M. K. Upadhyay and R. Pratap, RSC Adv., 2025, 15, 12117–12124 RSC.
- S. Singh, I. Althagafi, P. Yadav, R. Panwar, A. Kumar and R. Pratap, Tetrahedron, 2014, 70, 8879 CrossRef CAS.
- V. J. Ram, M. Nath, P. Srivastava, S. Sarkhel and P. R. Maulik, J. Chem. Soc., Perkin Trans. 1, 2000, 3719 RSC.
- S. N. Sahu, M. K. Gupta, T. Jadhav, P. Yadav, S. Singh, R. Misra and R. Pratap, RSC Adv., 2014, 4, 56779 RSC.
- D. Singhal, I. Althagafi, A. Kumar, S. Yadav, A. K. Prasad and R. Pratap, New J. Chem., 2020, 44, 12019 RSC.
- A. Jauhari, et al., Mol. Neurobiol., 2018, 55, 936 CrossRef CAS PubMed.
- (a) A. Pandey, et al., Toxicol. Res., 2015, 4, 1578 CrossRef CAS; (b) Z. Ruzi, A. Buronov, L. Nie, A. Nasrullaev, Z. Murtazaeva, R. Kuryazov, J. Zhao, T. Efferth, H. Akber and K. Bozorov, Int. J. Mol. Sci., 2025, 26(17), 8528 CrossRef CAS PubMed; (c) M. M. Fouad, H. A. Ghabbour, I. A. Shehata and M. B. El-Ashmawy, Bioorg. Chem., 2024, 148, 107401 CrossRef CAS PubMed; (d) E. S. M. Elsenbawy, Z. S. Alshehri, N. A. Babteen, A. A.-H. Abdel-Rahman, M. A. El-Manawaty, E. S. Nossier, R. K. Arafa and N. A. Hassan, Molecules, 2024, 29(5), 1067 CrossRef CAS; (e) A. Rogova, I. A. Gorbunova, T. E. Karpov, R. Y. Sidorov, A. E. Rubtsov, D. A. Shipilovskikh, A. R. Muslimov, M. V. Zyuzin, A. S. Timin and S. A. Shipilovskikh, Eur. J. Med. Chem., 2023, 254, 115325 CrossRef CAS; (f) R. Vasamsetti, N. B. Gatchakayala, P. V. Kumar, C. Praveen, S. V. G. V. A. Prasad and B. Madhav, Results Chem., 2024, 101823 CrossRef CAS.
- (a) J. H. Park, Y. Liu, M. A. Lemmon and R. Radhakrishnan, Biochem. J., 2012, 448, 417 CrossRef CAS PubMed; (b) S. Bhardwaj and A. K. Yadav, J. Neuro Oncol., 2025, 171, 47–63 CrossRef CAS PubMed.
- E. A. Sobh, M. A. Dahab, E. B. Elkaeed, A. A. Alsfouk, I. M. Ibrahim, A. M. Metwaly and I. H. Eissa, J. Enzyme Inhib. Med. Chem., 2023, 38, 2220579 CrossRef.
- P. Limpachayaporn, S. Nuchpun, J. Sirirak, P. Charoensuksai, P. Wongprayoon, N. Chuaypen, P. Tangkijvanich and A. Suksamrarn, Bioorg. Med. Chem., 2022, 74, 117048 CrossRef CAS PubMed.
- D. K. Yadav, S. Kumar, Saloni, S. Misra, L. Yadav, M. Teli, P. Sharma, S. Chaudhary, N. Kumar, E. H. Choi, H. S. Kim and M. H. Kim, Sci. Rep., 2018, 8, 4777 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |