 Open Access Article
Open Access ArticleComment on “Insight into the interaction between tannin acid and bovine serum albumin from a spectroscopic and molecular docking perspective” by W. Xu, Y. Ning, S. Cao, G. Wu, H. Sun, L. Chai, S. Wu, J. Li and D. Luo, RSC Adv., 2023, 13, 10592
Jonghoon Kang
Department of Biology, Valdosta State University, Valdosta, Georgia 31698, USA. E-mail: jkang@valdosta.edu
First published on 18th February 2026
Abstract
In an article [W. Xu et al., RSC Adv., 2023, 13(16), 10592–10599], the authors characterized the interaction between tannic acid (TA) and bovine serum albumin (BSA) using a variety of biophysical techniques, including thermodynamic analysis. Upon examining their reported thermodynamic parameters, we identified an internal inconsistency: the enthalpy change (ΔH) was treated as temperature-independent, whereas the entropy change (ΔS) was reported as temperature-dependent, a combination that contradicts fundamental principles of equilibrium thermodynamics. In this Comment, we clarify the source of this inconsistency by reanalyzing the reported Gibbs free energy (ΔG) values and applying the appropriate thermodynamic relationships. Our analysis yields self-consistent values of ΔH and ΔS and underscores the limitations imposed by the small number of temperature data points. We further discuss the conditions under which heat-capacity effects (ΔC) can be meaningfully evaluated and recommend future experiments that incorporate additional temperature measurements. These refinements will enable a complete and internally consistent thermodynamic characterization of the TA–BSA interaction.
The above-cited paper (hereafter referred to as ‘the original paper’) was published in RSC Advances by Xu et al.1 The objective of the original paper was to characterize the interaction between tannic acid (TA) and bovine serum albumin (BSA) using a suite of biophysical approaches, including spectroscopic, thermodynamic, and computational techniques. Their study provides valuable insights into the molecular basis of TA–protein interactions, particularly considering the proposed health benefits of TA, such as gastroprotection through mitigation of oxidative and inflammatory stress.2
Upon examining the reported thermodynamic data, we identified an internal inconsistency in the analysis of the thermodynamic parameters. As we and others emphasized, the correct treatment and interpretation of thermodynamic data are essential for ensuring conceptual clarity and scientific reproducibility.3,4 In this Comment, we clarify a theoretical issue that may strengthen the original work and prevent conceptual misunderstandings among readers.
The authors reported three parameters describing the binding of TA to BSA: the changes in enthalpy (ΔH), entropy (ΔS), and Gibbs free energy (ΔG). The inconsistency arises from reporting ΔH as temperature-independent while ΔS is reported as temperature-dependent. This contradicts a fundamental theorem of equilibrium thermodynamics: if ΔS varies with temperature, the heat-capacity change ΔC must be nonzero; and if ΔC ≠ 0, then ΔH must also vary with temperature.3,5,6
We first verified the ΔG values using the reported equilibrium constant (K) via eqn (1):
ΔG = −RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) K K
| (1) |
To obtain self-consistent thermodynamic parameters, we applied linear regression to the relationship between ΔG and T according to eqn (2):
| ΔG = ΔH − TΔS | (2) |
 | ||
| Fig. 1 Gibbs free energy (ΔG) for the binding of tannic acid to bovine serum albumin at three temperatures. The solid line represents the linear fit based on eqn (2). Data visualization and regression analysis were performed using SigmaPlot (version 15, Grafiti LLC, Palo Alto, CA). | ||
The relatively modest coefficient of determination (R2 = 0.756) suggests that ΔH and ΔS may themselves be temperature-dependent. In such cases, the appropriate expression relating ΔG and T is:
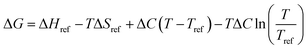 | (3) |
In summary, we clarify an internal inconsistency in the thermodynamic analysis conducted in the original paper. This Comment is intended not as criticism but as a constructive refinement to ensure accurate thermodynamic interpretation. We emphasize the scientific value of the work by Xu et al.1 and hope our clarification aids readers in avoiding conceptual misconceptions. We further suggest measuring binding affinities at additional temperatures, which would permit determination of ΔC (if nonzero) and enable a more complete thermodynamic characterization of the TA–BSA interaction.
Author contributions
JK is the sole author of this paper and is solely responsible for its content.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.References
- W. Xu, Y. Ning, S. Cao, G. Wu, H. Sun, L. Chai, S. Wu, J. Li and D. Luo, RSC Adv., 2023, 13, 10592–10599, 10.1039/d3ra00375b.
- B. O. de Veras, M. V. da Silva and P. P. Cabral Ribeiro, Food Chem. Toxicol., 2021, 156, 112482, DOI:10.1016/j.fct.2021.112482.
- J. Kang and A. M. Kang, J. Mater. Chem. B, 2020, 8, 10205–10208, 10.1039/d0tb01160f.
- J. C. Bollinger, H. N. Tran and E. C. Lima, RSC Adv., 2022, 12, 5769–5771, 10.1039/d1ra08212d.
- R. Chang, Physical Chemistry for the Chemical and Biological Sciences, University Science Books, Sausalito, CA, 2000 Search PubMed.
- D. Klostermeier and M. G. Rudolph, Biophysical Chemistry, CRC Press, Boca Raton, FL, 2017 Search PubMed.
- J. R. Harris and J. Kang, Mol. Immunol., 2022, 141, 338–339, DOI:10.1016/j.molimm.2021.12.005.
| This journal is © The Royal Society of Chemistry 2026 |
