Open Access Article
Open Access ArticleExploration of the aromatic abietane diterpenoid scaffold in the development of ArnT-mediated colistin resistance inhibitors
Silvia Cammarone†
 a,
Valentina Pastore†b,
Mariya Ryzhuka,
Martina Cristoferib,
Arianna Speroni
a,
Valentina Pastore†b,
Mariya Ryzhuka,
Martina Cristoferib,
Arianna Speroni a,
Francesco Imperic,
Davide Corinti
a,
Francesco Imperic,
Davide Corinti a,
Maria Carmela Bonaccorsi di Patti
a,
Maria Carmela Bonaccorsi di Patti d,
Bruno Botta
d,
Bruno Botta a,
Deborah Quaglio
a,
Deborah Quaglio *a,
Mattia Mori
*a,
Mattia Mori *e,
Fiorentina Ascenzionib and
Francesca Ghirga
*e,
Fiorentina Ascenzionib and
Francesca Ghirga a
a
aDepartment of Chemistry and Technology of Drugs, Sapienza University of Rome, Italy. E-mail: deborah.quaglio@uniroma1.it
bDepartment of Biology and Biotechnology Charles Darwin, Sapienza University of Rome, Italy
cDepartment of Science, Roma Tre University, Italy
dDepartment of Biochemical Sciences “A. Rossi Fanelli” Sapienza University of Rome, Italy
eDepartment of Biotechnology, Chemistry and Pharmacy, University of Siena, Italy. E-mail: mattia.mori@unisi.it
First published on 25th February 2026
Abstract
Colistin is a last-resort antibiotic for treating multidrug-resistant Gram-negative bacterial infections. However, some bacteria, such as Pseudomonas aeruginosa, can develop resistance to colistin thanks to the modification of the lipid A component of the outer membrane catalyzed by the 4-amino-4-deoxy-L-arabinose transferase ArnT. Here, building on our previous identification of the natural ent-beyerene diterpene FDO as an ArnT inhibitor, we report a rational procedure to simplify the ent-beyerene scaffold into drug-like synthetic ArnT inhibitors. Starting from the aromatic abietane scaffold of podocarpic acid 1, a function-oriented synthesis (FOS) approach guided the preparation of twenty-four semisynthetic oxygen and nitrogen abietane-based derivatives. Biological evaluation against a colistin-resistant P. aeruginosa strain revealed that four compounds (2, 13, 18, and 30) significantly restore the activity of sub-inhibitory colistin, with compound 18 showing the lowest IC90 (182.5 µM). Structure–activity relationship (SAR) studies revealed that positions C-12 and C-16 are critical for activity, while molecular docking suggested that the selected analogs can engage in interactions with conserved residues within the ArnT catalytic site. Microbiological assays highlighted key structural features of the aromatic abietane skeleton in restoring colistin sensitivity in drug-resistant P. aeruginosa, and rationalized biological activity data to corroborate the quality of the selected scaffold in the development of cost-effective colistin resistance breakers.
Introduction
Global resistance to most available antibiotics is threatening human health and making antibiotic choices for infection control limited and more expensive.1 In particular, the emergence of multidrug-resistant (MDR) pathogens such as Pseudomonas aeruginosa, Acinetobacter baumannii, and Enterobacteriaceae is alarming.2 Hence, in 2017, the World Health Organization (WHO) included these pathogens in the list of bacteria for which antibiotics are urgently needed as a priority.3Although new treatment options such as ceftazidime/avibactam, ceftolozane/tazobactam (Avycaz®), (Zerbaxa®), plazomicin (Zemdri®), and eravacycline (Xerava®) have become available for the treatment of Gram-negative bacteria, resistance to these treatments has already started to emerge.4 Thus, the medical community re-evaluated the use of old antibiotics, such as polymyxins, including colistin and polymyxin B.5–7 The reintroduction of colistin in clinical practice has inevitably led to the emergence and spread of colistin-resistant isolates, as evidenced by numerous case reports.8 Colistin targets the lipopolysaccharide (LPS) of the outer membrane (OM) of Gram-negative bacteria by electrostatic interaction, which in turn displaces the OM stabilizing cations, Mg2+ and Ca2+, causing OM destabilization, leakage of cell contents, and ultimately cell death. More recently, it has been proposed that colistin binding to newly synthesized LPS in the cytoplasmic membrane (CM) also contributes to its bactericidal activity.9 Accordingly, accumulation of LPS in the CM increases the susceptibility of P. aeruginosa to colistin.9 Therefore, to counteract the antimicrobial activity of colistin, bacteria have developed mechanisms to decrease the overall charge of LPS, which in turn reduces the binding affinity of the cationic lipopeptide and its bactericidal activity.7 Although different mechanisms of LPS modification have been described, including the addition of phosphoethanoamine (pEtN) or 4-amino-4-deoxy-L-arabinose (L-Ara4N) to the lipid A component of LPS,10–12 in P. aeruginosa the latter is the major colistin resistance mechanism.10,13 This mechanism, which is activated by mutations in two-component regulatory systems, such as PmrAB and PhoPQ, is mediated by the arn operon, and requires multiple steps, the final one being the transfer of L-Ara4N to lipid A by the integral CM enzyme ArnT (undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase).7 ArnT catalyzes the transfer of L-Ara4N, provided by the lipid carrier undecaprenyl phosphate, to lipid A phosphate groups.14
Enhancing the effectiveness of existing antimicrobial compounds is a promising strategy for addressing the current antibiotic resistance crisis. Specifically, inhibitors of key enzymes involved in antibiotic resistance offer an alternative approach to combating this threat. Remarkably, only a few examples of colistin adjuvants have been reported to date.15,16 By combining these inhibitors with clinically relevant antibiotics, we can potentially prolong the lifespan of these antibacterial drugs and reduce the impact of resistance. Notwithstanding, identifying novel classes of these life-saving drugs has become more challenging, requiring new paradigms. In the current scenario, exploring the biologically relevant chemical space of natural products (NPs) remains a promising approach for discovering novel antimicrobial or antibiotic adjuvants.17–22 Indeed, several design strategies were employed to develop novel and fascinating architectures by skillfully manipulating the NP's core scaffold.20,21,23
In a previous work, we have identified the ent-beyerene diterpene FDO (formerly known as BBN149), isolated from the leaves of Fabiana densa var. ramulosa, a native shrub of Chile, as a potent ArnT inhibitor capable of restoring colistin efficacy in resistant P. aeruginosa strains without significantly affecting colistin-susceptible strains (Fig. 1a).24–27 These results suggest that FDO prevents lipid A aminoarabinosylation, likely by inhibiting ArnT enzymatic activity, thus restoring colistin efficacy.25 A large variety of chemical analogs was produced for SAR studies to validate the potential of the diterpene scaffold as a key platform for the further development of ArnT-mediated colistin resistance inhibitors with improved activity.27 However, notable challenges are associated with this type of diterpene, including the low concentration of the compound in the Chilean plant F. densa var. ramulosa, the need for multistep purification of the raw material, the absence of chromophore groups and low solubility. The latter was recently addressed by the development of liposomal carriers for the co-delivery of colistin and diterpenoid ArnT inhibitors.28
 | ||
| Fig. 1 Chemical structures of the ent-beyerene diterpene FDO (a) and aromatic abietane podocarpic acid (1) (b). | ||
Here, to address these issues, we considered rational procedures that simplify the ent-beyerene complex structure into drug-like synthetic molecules, thereby minimizing the pharmacophore structures that are crucial for biological activity. Therefore, other naturally related diterpene scaffolds were evaluated as starting materials to prepare a novel generation of ArnT inhibitors. In particular, we investigated the aromatic abietane scaffold, which lacks ring D compared to the ent-beyerene one and features an aromatic ring C and varying degrees of oxygenation at several positions.29 Several synthetic studies that aim to modify the abietane skeleton to obtain novel biologically active compounds have been reported. To achieve this, naturally occurring terpenoids, which are available in large quantities, were utilized as enantiomerically pure starting materials for the synthesis of bioactive abietanes.30 In this study, we report the design, synthesis, and biological evaluation of a series of novel abietane-based derivatives derived from podocarpic acid 1 (Fig. 1b), conceived as function-oriented analogs of the natural ent-beyerene hit FDO (Fig. 1a).
By exploiting the structural modularity of the abietane scaffold and guided by in silico modeling, we aimed to identify simplified and synthetically accessible compounds capable of selectively inhibiting ArnT and restoring colistin efficacy against resistant P. aeruginosa strains. This work highlights the potential of aromatic diterpenoids as a viable chemical platform and FOS as a successful design strategy for the development of next-generation colistin resistance inhibitors.
Results and discussion
Rational design strategy
Recently, we demonstrated that an ent-beyerene scaffold bearing an oxalate-like group at C-18/C-19 or a sugar residue at C-19 is an essential requirement for efficient inhibition of bacterial growth in the presence of colistin, likely resulting from a more efficient inhibition of the ArnT activity.27 Here, we simplified the complex structure of ent-beyerene through a function-oriented synthesis (FOS) approach to optimize the hit compound by minimizing the crucial pharmacophores necessary for biological activity.31,32 In particular, this study investigates the aromatic abietane scaffold of 1, which lacks the ring D compared to the ent-beyerene one, maintains the same configuration at C-4 concerning FDO, and features an aromatic ring C and a hydroxyl group at the C-12 position. Based on a function-oriented design, compound 4, in which the abietane scaffold of 1 incorporates the key recognition feature of the hit FDO, the oxalyl group at C-16, was designed (Fig. 2) and evaluated in silico towards the predicted structure of P. aeruginosa ArnT by adapting from the computational protocol already established and validated in previous works.25,27 | ||
| Fig. 2 The hit compound FDO and the simplified aromatic analog 4. The key binding element is highlighted in blue. | ||
The abietane scaffold of 1 was prioritized for further experimental investigations due to its potential interaction with the target receptor as well as functionalization strategies. We synthesized a focused library of twenty-four derivatives of 1, combining the aromatic abietane scaffold with key binding elements essential for efficient inhibition of ArnT activity, as previously investigated in SAR studies of the ent-beyerene derivatives.27 In particular, different derivatives of 1 were synthesized to investigate the role of (i) the oxalyl group at C-16, an essential requirement for the ArnT activity inhibition of the hit compound FDO; (ii) the length and flexibility of the alkyl chain of the functional group at C-16; (iii) the presence of a sugar unit to mimic L-Ara4N; (iv) the substitution of the oxygen with nitrogen heteroatom in the functional group at C-16 on the biological properties of the original abietane scaffold; and (v) the role of the phenolic group at C-12.
Chemistry
Commercially available 1 was used as the starting material for preparing derivatives of the oxygen series (2–8, 12–18) and the nitrogen series (19–26, 30–31).Oxygen (O) series
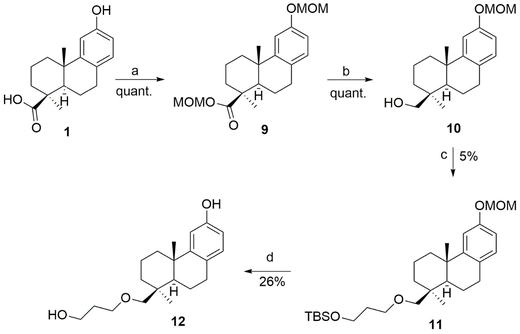
According to the SAR analysis previously performed on the ent-beyerene scaffold, we designed and synthesized a small focus library of 1 O-derivatives bearing an alcohol or an oxalate-like group or a sugar residue at C-16 (2–8) (Scheme 1).To specifically assess whether the removal of the carbonyl groups at C-16 could affect the colistin adjuvant activity, alcohol 2 was prepared in quantitative yield through a synthetic approach that involved a first activation of carboxylic acid to the corresponding acyl chloride, followed by the reduction of the carbonyl group of 1 using lithium aluminum hydride (LiAlH4). This compound was then used to synthesize derivatives 3–8 by chemical modifications of the alcohol group at C-16. In particular, to investigate the role of the distance between the lipophilic portion of the terpene scaffold and the polar function, the alcohol group linked at position C-16, compound 3 was obtained in 40% yield via condensation reaction of 2 with commercially available 3-[(tert-butyldimethylsilyl)oxy]propanoic acid previously activated to acyl chloride (Scheme S1). Indeed, to evaluate the length and flexibility of the alkyl chain of the functional group at C-16, the corresponding oxalyl 4, malonyl 5, and succinyl 6 esters were obtained by treating diterpene 2 with the corresponding acyl chloride in Et2O at room temperature for 30 min, in quantitative, 67%, and 20% yields, respectively. Furthermore, to assess the influence of a sugar unit to mimic L-Ara4 N at C-16 position, the glycosidic derivative 7 was prepared by adding the alcohol 2 to a solution of acetobromo α-D-glucose and zinc chloride, according to the Koenigs–Knorr reaction.33 Most commonly, the glycosylation reaction follows an unimolecular SN1 mechanism, forming the resonance-stabilized oxocarbenium ion. As a result, the nucleophilic attack could occur almost equally from either the top (to give the 1,2-trans isomer or β) or the bottom face (to give the 1,2-cis or α isomer) of the ring. However, the anomeric effect promotes the α-glycosylation.34 In particular, the formation of the alpha isomer was confirmed by 1H-NMR analysis observing a coupling constant (J) of 3.36 Hz for the douplet signal centered at 5.03 ppm, related to the proton H1′ (the one on anomeric carbon). This value is characteristic of an eq-ax coupling, as typically observed between the protons H1′ and H2′.35 Additionally, deacetylation of sugar hydroxyl groups via alkaline hydrolysis with triethylamine produced 8 in 27% yield. Lastly, to evaluate the effect of introducing a more flexible alkyl chain that enhances the distance between the lipophilic portion and the polar function of the abietane scaffold, alcohol 10 was prepared by first protecting 1 with chloromethyl methyl ether (MOMCl), followed by reduction using LiAlH4 to obtain it in a quantitative yield (Scheme 2).
Protecting the phenolic group at position 12 as a methoxy-methyl ether was necessary to prevent its etherification during the subsequent nucleophilic substitution reaction. Intermediate 10, formed after the reduction with LiAlH4, was then deprotonated with sodium hydride (NaH) to allow nucleophilic substitution with 3-bromo-1-propanol previously protected with tert-butyldimethylsilyl (TBS) (Scheme S1). Finally, a double deprotection of MOM and TBS was performed using trifluoroacetic acid (TFA) and trifluoroethanol (TFE) (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio) in dichloromethane (DCM), resulting in the desired derivative 12 in 26% yield.
1 ratio) in dichloromethane (DCM), resulting in the desired derivative 12 in 26% yield.
According to the docking study, an additional binding interaction at the aromatic C ring of the abietane scaffold of 1 may provide advantages in binding affinity compared to the reference molecule FDO. Therefore, to investigate the role of the phenolic group at C-12, the dimethylated ester 13 was prepared by treating 1 with methyl iodide (CH3I) and potassium carbonate (K2CO3) as base, resulting in a 70% yield. The methyl ester 14 was also obtained in 30% yield. Compound 13 was further used for synthesizing derivatives 15–16. In particular, hydrolysis of 13 led to the formation of acid 15 with an 80% yield. Compound 16 was prepared by reducing 13 with LiAlH4 to the corresponding alcohol in a quantitative yield. Then, the hydroxyl group of 13 was esterified with the desired acyl chloride, resulting in the oxalyl ester 17 in quantitative yield and the succinyl ester 18 in 80% yield (Scheme 3).
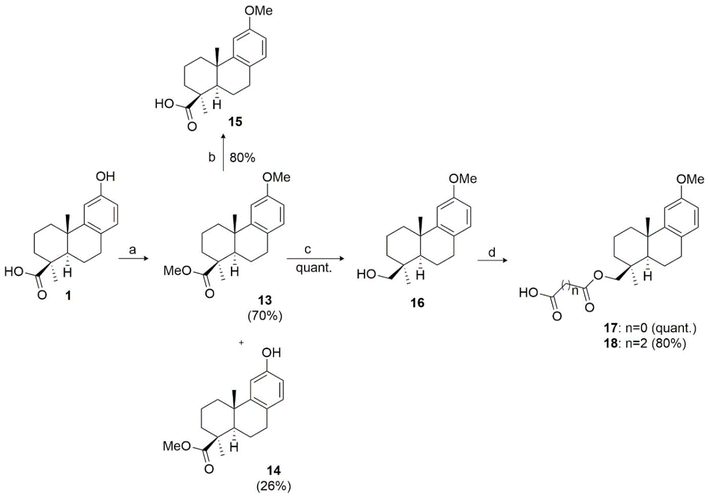 | ||
| Scheme 3 Reagents and conditions. (a) CH3I, K2CO3, acetone, 56 °C, 24 h; (b) t-BuOK, DMSO, 100 °C, 72 h; (c) LiAlH4, dry THF, 0 °C to r.t. 24 h; (d) ClCO(CH2)nCOCl, Et2O, 0 °C to r.t., 30 min. | ||
Thus, derivatives 13, 15, 16, 17 and 18 differ from the structurally correlated ones 14, 1, 2, 4 and 6, respectively, only due to the methoxyl group at C-12.
Nitrogen (N) series
A series of N-derivatives of compound 1, featuring a primary or secondary amide or amine at the C-16 position, was designed according to bioisosterism. The aim was to assess whether replacing the oxygen atom with a nitrogen atom could affect the chemical stability and the effectiveness of inhibition of ArnT. Indeed, this chemical modification could facilitate compound entry and its accumulation according to the “eNTRy rules”.36 Accordingly, an efficient strategy for the preparation of the primary amide 19 was developed, involving the use of N,N-diisopropylethylamine (DIPEA), hexafluorophosphate azabenzotriazole tetramethyl uranium (HATU) as a coupling agent, and ammonium chloride (NH4Cl).37 This strategy, reported in the literature for amidation reactions, enabled the production of 19 in a single step with quantitative yield (Scheme 4). | ||
| Scheme 4 Reagents and conditions. (a) DIPEA, HATU, NH4Cl, dry DMF, r.t., 16 h; (b) LiAlH4 2M in THF, dry THF, 0 °C to r.t., 16 h; (c) succinic anhydride, DMAP, DCE, 0 °C to r.t., 30 min. | ||
Furthermore, reducing the carbonyl group of 19 using LiAlH4 resulted in the corresponding primary amine 20 in 50% yield. Compound 21, featuring an oxalate-like group at C-16, an essential requirement for the ArnT activity inhibition of the hit compound FDO, was obtained by amidation reaction of 20 with succinic anhydride and 4-dimethylaminopyridine (DMAP) as base in quantitative yield. This derivative differs from the structurally correlated one, 6, only due to the substitution of the oxygen with a nitrogen atom at C-16. According to docking analysis, the introduction of an alkyl chain to enhance the distance between the lipophilic portion and the polar function (alcohol group) of the abietane scaffold was also investigated in this series. In particular, the secondary amide 22 was prepared using the same amidation reaction employed for 19, by coupling 1 with β-alanine in the presence of HATU and DIPEA (Scheme 5).
 | ||
| Scheme 5 Reagents and conditions. (a) DIPEA, HATU, NH2(CH2)nOH or NH2(CH2)2COOH, dry DMF; (b) LiAlH4 2M in THF, dry THF, 0 °C to r.t., 32 h. | ||
Similarly, secondary amides 23–25 were synthesized by coupling 1 with the suitable linear amino alcohol in the presence of HATU and DIPEA, resulting in the corresponding N-hydroxypropyl, butyl, and pentyl amides 23–25 in 90%, quantitative and 85% yields, respectively (Scheme 5). Furthermore, amide 23 was reduced with LiAlH4 to the corresponding secondary amine 26 in low yield (12% yield), whereas amides 24 and 25 showed no reactivity under the same reduction conditions. For this reason, a different procedure via reductive amination became necessary to obtain amines 30 and 31 (Scheme 6).
 | ||
| Scheme 6 Reagents and conditions. (a) DMP, NaHCO3, dry CH2Cl2, 0 °C to r.t., 2 h; (b) NH2(CH2)nOH, AcOH 1%, NaCNBH3, dry CH2Cl2, r.t., 24 h; (c) TFA, TFE, CH2Cl2, 0 °C to r.t. | ||
In particular, oxidation under mild conditions using Dess–Martin Periodinane (DMP) as the oxidizing agent of alcohol 11, previously obtained as shown in Scheme 2, led to the aldehyde 27 in 65% yield.38 This intermediate underwent a reductive amidation reaction with the suitable amino-alcohol,39 followed by deprotection with TFA and TFE in DCM, to provide the corresponding N-hydroxybutyl and penthyl amines 30 and 31 in 55% and 15% yields, respectively.
Analysis of the colistin adjuvant activity of the abietane diterpenoids
The colistin adjuvant activity of the compounds was analyzed by determining their effect on the growth of the P. aeruginosa colistin resistant strain PA14 ColR5.10 As colistin resistance in PA14 ColR5 depends on the activity of the ArnT transferase, it is expected to be reduced by the compounds.10 Of note, compounds targeting ArnT are not expected to possess antibacterial activity per se, as demonstrated by the comparable growth profile of P. aeruginosa parental strains and their isogenic Δarn mutants.10 To obtain measurable effects from the combination compound-colistin on the bacterial growth, the sublethal dose of 8 µg mL−1 of colistin, which reduces the PA14 ColR5 growth to 50% (Fig. S1), was selected.40 Based on this, pairwise interaction compound-colistin was analyzed by combining each compound, in a range of concentrations from 16 to 250 µM, with 8 µg mL−1 of colistin, the resulting effect was compared to the effect of the individual compounds. First, the O-derivatives (2–8) of compound 1, which differed for the residue at C-16, were tested. These compounds showed a broad spectrum of adjuvant activity, from no effect to complete growth inhibition (Fig. 3A).In particular, compounds 1, 4, and 8 did not reduce the growth of PA14 ColR5, with compound 4 showing the opposite effect, namely, enhanced growth. Compounds 7 and 12 induced a marginal reduction in growth, while compounds 3, 5, and 6 caused a dose-dependent reduction in growth, ranging from 80 to 60%. Finally, compound 2 led to an almost complete inhibition of PA14 ColR5 growth. The promising result obtained with 2 was further corroborated by repeating the assay with additional samples containing the compound-only. As expected, the growth reduction caused by compound 2 was greatest when combined with colistin and significantly less in the absence of colistin (Fig. 3B). This data supports the specificity of compound 2, which differs from 1 for the presence of an alcohol group at C16, as its antimicrobial activity is approximately 20-fold higher when combined with colistin than in its absence. Differently, compounds 3, 5, and 6, enhancing the distance between the acid function at C16 and the lipophilic portion of the abietane scaffold, maintained the capacity to reduce the growth of PA14 ColR5, although at a reduced level compared to compound 2. Otherwise, increasing the distance between the alcohol group at C16 and the lipophilic portion of the abietane scaffold appeared to completely abrogate the binding to ArnT, as demonstrated by the absence of colistin adjuvant activity of compound 12 (compare compounds 12 and 2 in Fig. 3A).
Finally, the possible effect of the C ring in mediating the interaction of the abietane scaffold with ArnT was studied by analyzing the antimicrobial activity of compounds 13–18 (Fig. 4).
Among these, compound 13 was the most active, with complete abrogation of PA14 ColR5 growth at 250 µM. Compounds 14, 15, and 18 only partially reduced bacterial growth at the same concentration, from approximately 60% for 14 and 15 to 80% for 18. Compound 18 maintained high activity even at 125 µM. While compounds 13, 14, and 18 appeared to be specific colistin adjuvants, as no growth inhibition was detected in the absence of colistin, compound 15 showed similar antibacterial activity irrespective of colistin (Fig. S2). Compounds 16 and 17 were shown to be the least active, exhibiting little to no growth inhibition (Fig. 4 and S3). Further, direct comparison of the activity of compounds 15–18 and 13 with that of the correlated ones 1, 2, 4, 6 and 14, respectively, showed that the methoxyl group at C-12 is associated with higher colistin adjuvant activity, except for compound 2, in which the highest activity was associated with the hydroxyl group (Fig. S3).
The colistin adjuvant activity of the N-derivatives of compound 1 was determined using the same approach as reported above. Compounds 19, 21, and 22 showed only marginal growth inhibitory activity, while 20 inhibited the growth of PA14 ColR5 by 70% (Fig. 5A). To assess the effect of enhancing the distance between the lipophilic portion and the polar function (alcohol group) of the abietane scaffold, amides 23–25 were tested (Fig. 5B).
Only compounds 24 and 25 showed some activity, with growth inhibition of approximately 60% and 80%, respectively, at the highest dose. Their specificity was demonstrated by negligible activity in the absence of colistin (Fig. S4). Similarly, the corresponding amines 30 and 31 exhibited a dose-dependent activity, while compound 26 showed no effect on PA14 ColR5 growth (Fig. 5C). Additionally, the inhibitory activity on bacterial growth of compound 30 appeared significantly higher with colistin than in its absence at the concentration ranging from 63 to 250 µM (Fig. 6A).
Differently, the antimicrobial activity of compound 31 appeared to be independent of colistin, except at the highest concentration (Fig. 6B), possibly due to off-target effects. Finally, for a more direct comparison of the compounds showing colistin-dependent inhibition of P. aeruginosa growth, ranging from 50 to 99% at 250 µM, the growth inhibitory concentrations, IC50 and IC90, were determined as reported in Materials and methods (Table 1). Based on the IC50, the compounds that showed the highest activity (lowest IC50 values) were 18, 13, 30, and 2, all but 30 belonging to the oxygen derivative group. Among these, compound 18 also showed the lowest IC90.
| Compound | IC50 | IC90 | R2* |
|---|---|---|---|
| 18 | 73.91 | 182.49 | 0.87 |
| 13 | 80.79 | 267.01 | 0.82 |
| 30 | 96.36 | 217.10 | 0.80 |
| 2 | 130.50 | 225.94 | 0.89 |
| 6 | 130.70 | 276.74 | 0.86 |
| 5 | 156.20 | 311.44 | 0.85 |
| 20 | 166.70 | 341.57 | 0.77 |
| 31 | 171.90 | 432.90 | 0.80 |
| 25 | 225.10 | 259.22 | 0.70 |
| 24 | 227.00 | 365.42 | 0.74 |
We confirmed the compound-colistin synergy for 18, 13, 30, and 2 with chequerboard assays. As shown in Fig. 7, all compounds caused a dose-dependent reduction of PA14 ColR5 growth. Compound 2 reduced the colistin MIC to 4 µg mL−1 at 250 µM, while compounds 13, 18, and 30 to 8 µg mL−1 at the same concentration.
Overall, this analysis confirmed that compounds 18, 13, 30, and 2 act as bona fide nonantibiotic adjuvants of colistin as they potentiate colistin activity by targeting its arn-dependent resistant mechanism. Accordingly, these compounds, although showing no antimicrobial activity, reduce the MIC of colistin by 8- or 16-fold in the combined treatments. Finally, taking into consideration lung infection, the cytocompatibility of these compounds was evaluated in the bronchial epithelial cells 16HBE. For this, cells were treated with increasing concentration of the compounds and at two timepoints (3 and 18 h) cell viability was analyzed by the MTT assay. No significant changes in cell viability were observed under all conditions tested (Fig. 8).
Binding studies on the most active ArnT inhibitors
The possible binding mode of the most active compounds 2, 13, 18, and 30 to the ArnT protein from P. aeruginosa PA14 was investigated by molecular docking simulations. The previously developed protocol, which has been validated against the X-ray crystallography structure of Cupriavidus metallidurans ArnT, was adopted herein.14,25,27 Given the lack of experimental structure of ArnT from P. aeruginosa, its three-dimensional structure was computationally built by AlphaFold2 using the Uniprot sequence coded by ID Q02R27.41,42 Docking simulations were carried out with the FRED docking program (OpenEye Cadence Molecular Sciences) (Fig. 9).43While 2, 18, and 30 bind in a similar manner, 13 was found to bind with an opposite polarity, having the aromatic ring projected towards basic residues that bind the undecaprenylphosphate in the experimental ArnT structure.14 In detail, 2 is H-bonded to the backbone of Ile29 through the phenolic OH group, while the aromatic ring is in a parallel displaced π–π stacking conformation to the side chain of Tyr131; the primary alcohol group is H-bonded to the side chain of Asp135 near the sugar binding pocket of the ArnT catalytic site. The aromatic ring of 13 is π–π stacked with the side chain of Tyr35, whereas the methoxyl moiety is H-bonded to Arg34; the methylester group at C4 is H-bonded to the side chain of Gln128. 18 binds similarly to 2, although the functionalization inserted on the alkyl chain exploits additional interactions within the ArnT sugar binding pocket as well as with basic residues involved in the interaction with the undecaprenylphosphate substrate. Besides stacking to Tyr131, 18 establishes H-bonds with the side chain of Arg34 and Arg245. Finally, 30 shares binding hotspots with 2 and 18, although the protonated amino group establishes a charge-assisted H-bond with the side chain of Asp135, and the primary alcohol group is H-bonded to Arg245 and Glu60. Docking scores, calculated with the Chemgauss4 function implemented in the FRED docking program, suggested that the different binding polarity of 13 might account for a weaker binding affinity for the ArnT compared to 2, 18 and 30 (Table 2).
| Compound | FRED Chemgauss4 score (dimensionless) |
|---|---|
| 2 | −6.18 |
| 13 | −5.48 |
| 18 | −6.87 |
| 30 | −7.02 |
Overall, docking results show that tested compounds are able to fit within the lipid A binding pocket of the ArnT catalytic site. The qualitative agreement between docking parameters and bioactivity data supports the proposed binding poses, allowing further rational optimization studies.
Notably, residues contacted by these molecules are highly conserved in the ArnT orthologs from multiple P. aeruginosa strains, as highlighted by sequence similarity analysis (Fig. S5).
In order to confirm that ArnT is the target of the compounds, a binding assay was performed. Because of the high fluorescence due to the aromatic ring of the PD derivatives, the FDO and the most active derivative FDO-H (compound 15 in Quaglio et al., 2020),27 which differs from the hit FDO by the absence of the double bond between C-15 and C-16,25,27 were employed to assess binding to recombinant P. aeruginosa ArnT by intrinsic tryptophan fluorescence spectroscopy. The lead compound FDO was initially assayed, but it showed high background, making analysis problematic; however, even in these conditions, a KD of about 10 µM could be estimated. To obtain more reliable data, the FDO-H derivative was therefore employed. The choice of FDO-H is supported by previous molecular docking data, which show similar positioning of the PD compounds in the substrate binding site of ArnT.27 Titration of ArnT with FDO-H caused quenching of tryptophan fluorescence (Fig. 10, upper panel). Analysis of the data yielded a KD value close to 10 µM (Fig. 10, lower panel), substantiating the result obtained for FDO. This value is substantially lower than KD 31.76 µM for isostevic acid,28 in line with the higher potency of the lead compound FDO-H.
Structure–activity relationship
The naturally occurring aromatic abietane 1 and its newly synthesized twenty-four chemical derivatives highlighted that derivatives 2, 13, 18, and 30, belonging to the oxygen and nitrogen series, respectively, show the highest colistin adjuvant activity, causing an almost complete reduction of bacterial growth at 250 µM. At the same time, no effects were observed in the absence of colistin. Based on the overall outcomes of microbiological and computational investigations, a preliminary SAR was elaborated at two structural levels: the C-16 position at ring A and the C-12 position at ring C, and is graphically depicted in Fig. 11.Concerning ring A, the reduction of the carbonyl group at C-16 and the presence of an alcohol group at the same position led to a high colistin adjuvant activity (2), causing an almost complete reduction of bacterial growth at 250 µM and 37% reduction at 125 µM, suggesting that an sp3 hybridization favors the interaction of the compound with the catalytic site of ArnT.
An increased distance between the lipophilic portion of the terpene scaffold and the polar function, the alcohol group linked at position C-16, maintains (3, 5, 6) the ability to reduce the growth of PA14 ColR5. However, this effect is observed at a lower level compared to compound 2. The introduction of an oxalyl group (4) completely abrogates the effect, probably due to the more rigid and planar conformation. Similarly, when a hindered group, such as a sugar moiety at C-16 (7, 8), is present, the effect is completely nullified.
The substitution of oxygen with a nitrogen heteroatom in the functional group at C-16 of 1 shows a good colistin adjuvant activity at 250 µM, and only when the distance between the lipophilic portion of the scaffold and the polar function is enhanced (24, 25). The reduction of the carbonyl group and the presence of a primary amine group at C-16 (20) partially inhibited the growth of PA14 ColR5. The presence of a secondary amine featuring a 4-hydroxybutyl chain (30) resulted in a notable dose-dependent growth inhibition, which was significantly enhanced in the presence of colistin.
Concerning ring C, the methylation of the phenol group at the C-12 position led to a higher colistin adjuvant activity only in the presence of an acid or an ester group at the C-16 (13, 18) position, compared to the phenol analogs (14, 6). This trend, consistently observed across our experimental series, supports an empirical structure–activity relationship (SAR) suggesting that methylation at C-12 enhances the interaction of the aromatic scaffold with the biological target only when combined with specific polar functionalities at C-16. Although the mechanistic basis of this synergistic effect remains to be elucidated, it provides a valuable guideline for further optimization and mechanistic investigation.
Remarkably, microbiological assays coupled with molecular modeling indicated that for more efficient colistin adjuvant activity, likely resulting from the inhibition of ArnT activity by the selected compounds and therefore from their interaction with the catalytic site of ArnT, hydrogen bonding acceptor and donor groups at C-16 are required. The substitution of oxygen with a nitrogen heteroatom in the functional group at C-16 increases the biological activity in the amine series. Taken together, these findings outline a preliminary SAR profile centered on the abietane scaffold, highlighting the significance of specific hydrogen-bonding functionalities at C-16 and phenol methylation at C-12 in modulating colistin adjuvant activity. These insights provide a rational framework for the further optimization of abietane-based ArnT inhibitors as next-generation colistin potentiators.
Conclusions
This study demonstrates the successful application of function-oriented synthesis to the design of abietane-based derivatives as colistin resistance breakers targeting the P. aeruginosa ArnT enzyme, with the final aim of systematic reduction and optimization of a complex natural scaffold toward pharmacologically relevant minimal structures. By leveraging the abietane scaffold of 1 as a rationally simplified mimic of ent-beyerene, we generated a library of analogues that retained or restored adjuvant activity against colistin-resistant P. aeruginosa. The most effective compounds, particularly 2, 13, 18, and 30, were able to restore colistin activity in resistant bacterial strains in a dose-dependent and colistin-dependent manner. Molecular docking supported the proposed binding mode of ArnT inhibitors and highlighted specific interactions at the catalytic site, notably involving the C-16 substituent and, in some cases, the methoxylated C-12 phenolic group. SAR analysis underscored the importance of hydrogen-bonding features, flexibility, and linker length between the lipophilic scaffold and polar functional groups. Moreover, nitrogen-based analogs such as compound 30 demonstrated effective colistin adjuvant activity, suggesting a viable strategy for further chemical development. Even if the relatively high IC50/IC90 values of the most active compounds imply that these molecules require further optimization to improve their potency, these results establish the aromatic abietane scaffold as a valuable platform and FOS as a powerful design strategy for the development of next-generation colistin resistance breakers to overcome resistance in Gram-negative pathogens.Author contributions
Silvia Cammarone and Valentina Pastore contributed equally to this work. CRediT: Silvia Cammarone and Valentina Pastore: validation, investigation, methodology, data curation, writing – original draft. Mariya Ryzhuk, Martina Cristoferi, Arianna Speroni and Davide Corinti: visualization, investigation, data curation. Francesco Imperi: visualization, writing – review & editing. Maria Carmala Bonaccorsi di Patti: validation, investigation, methodology. Bruno Botta: validation, writing – review & editing. Deborah Quaglio and Mattia Mori: conceptualization, methodology, investigation, supervision, funding acquisition, writing – original draft. Fiorentina Ascenzioni and Francesca Ghirga: conceptualization, methodology, investigation, resources, funding acquisition, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting the findings of this study are available within the article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra09142j.Acknowledgements
This research was supported by: EU funding within the MUR PNRR Extended partnership initiative on Emerging Infectious Diseases (Project no. PE00000007, INF-ACT) to MM; Progetto ECS 0000024 Rome Technopole, – CUP B83C22002820006, PNRR Missione 4 Componente 2 Investimento 1.5, finanziato dall’Unione europea – NextGenerationEU to SC, FG and DQ; PROGETTI MEDI ATENEO 2023 – RM123188F706283C to FG; Sapienza University (Medie Attrezzature Scientifiche_MA32218167926358) to FG; Sapienza, project 2023 – RG123188B49A8CB3 to FA. Mariya Ryzhuk was supported by the PNRR PhD scholarship funded by the Rome Technopole project. We thank OpenEye Cadence Molecular Sciences for their free academic license. This article is based upon work from COST Action EURESTOP, CA21145, supported by COST (European Cooperation in Science and Technology). We gratefully acknowledge Dr Giulia Mazzoccanti for her technical assistance with HPLC analyses.References
- B. Li and T. J. Webster, Bacteria antibiotic resistance: New challenges and opportunities for implant-associated orthopedic infections, J. Orthop. Res., 2018, 36, 22–32 CrossRef PubMed.
- V. H. Krishnaprasad and S. Kumar, Antimicrobial Resistance: An Ultimate Challenge for 21st Century Scientists, Healthcare Professionals, and Policymakers to Save Future Generations, J. Med. Chem., 2024, 67, 15927–15930 Search PubMed.
- W. H. Organization, WHO publishes list of bacteria for which new antibiotics are urgently needed, https://www.who.int/en/news-room/detail/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed.
- H. Wright, R. A. Bonomo and D. L. Paterson, New agents for the treatment of infections with Gram-negative bacteria: restoring the miracle or false dawn?, Clin. Microbiol. Infect., 2017, 23, 704–712 Search PubMed.
- H. Giamarellou, Epidemiology of infections caused by polymyxin-resistant pathogens, Int. J. Antimicrob. Agents, 2016, 48, 614–621 CrossRef CAS PubMed.
- D. J. Payne, Desperately seeking new antibiotics, Science, 2008, 321, 1644–1645 Search PubMed.
- L. Poirel, A. Jayol and P. Nordmann, Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes, Clin. Microbiol. Rev., 2017, 30, 557–596 Search PubMed.
- F. A. Gogry, M. T. Siddiqui, I. Sultan and Q. M. R. Haq, Current update on intrinsic and acquired colistin resistance mechanisms in bacteria, Front. Med., 2021, 8, 677720 Search PubMed.
- A. Sabnis, K. L. H. Hagart, A. Klöckner, M. Becce, L. E. Evans, R. C. D. Furniss, D. A. I. Mavridou, R. Murphy, M. M. Stevens and J. C. Davies, Colistin kills bacteria by targeting lipopolysaccharide in the cytoplasmic membrane, eLife, 2021, 10, e65836 CrossRef CAS PubMed.
- A. Lo Sciuto and F. Imperi, Aminoarabinosylation of lipid A is critical for the development of colistin resistance in Pseudomonas aeruginosa, Antimicrob. Agents Chemother., 2018, 62, 10–1128 Search PubMed.
- E. M. Nowicki, J. P. O'Brien, J. S. Brodbelt and M. S. Trent, Extracellular zinc induces phosphoethanolamine addition to Pseudomonas aeruginosa lipid A via the ColRS two-component system, Mol. Microbiol., 2015, 97, 166–178 Search PubMed.
- T. Pedersen, J. O. Sekyere, U. Govinden, K. Moodley, A. Sivertsen, Ø. Samuelsen, S. Y. Essack and A. Sundsfjord, Spread of plasmid-encoded NDM-1 and GES-5 carbapenemases among extensively drug-resistant and pandrug-resistant clinical Enterobacteriaceae in Durban, South Africa, Antimicrob. Agents Chemother., 2018, 62, 10–1128 CrossRef PubMed.
- M. Cervoni M, D. Sposato, A. Lo Sciuto and F. Imperi, Regulatory Landscape of the Pseudomonas aeruginosa Phosphoethanolamine Transferase Gene eptA in the Context of Colistin Resistance, Antibiotics, 2023, 12, 200 Search PubMed.
- V. I. H. Petrou, C. M. Schultz, K. M. Clarke, O. B. Vendome, J. Tomasek, S. R. D. Banerjee, K. R. Belcher Dufrisne and M. Kloss, Structures of 51 aminoarabinose transferase ArnT suggest a molecular basis for lipid A glycosylation, Science, 2016, 351, 608–612 Search PubMed.
- W. T. Barker, A. M. Nemeth, S. M. Brackett, A. K. Basak, C. E. Chandler, L. A. Jania, W. J. Zuercher, R. J. Melander, B. H. Koller and R. K. Ernst, Repurposing eukaryotic kinase inhibitors as colistin adjuvants in Gram-negative bacteria, ACS Infect. Dis., 2019, 5, 1764–1771 Search PubMed.
- A. M. Nemeth, A. K. Basak, A. W. Weig, S. A. Marrujo, W. T. Barker, L. A. Jania, T. A. Hendricks, A. E. Sullivan, P. M. O'Connor and R. J. Melander, Structure–Function Studies on IMD-0354 Identifies Highly Active Colistin Adjuvants, ChemMedChem, 2020, 15, 210–218 CrossRef CAS PubMed.
- F. Cappiello, M. R. Loffredo, C. Del Plato, S. Cammarone, B. Casciaro, D. Quaglio, M. L. Mangoni, B. Botta and F. Ghirga, The revaluation of plant-derived terpenes to fight antibiotic-resistant infections, Antibiotics, 2020, 9, 325 Search PubMed.
- B. Casciaro, L. Mangiardi, F. Cappiello, I. Romeo, M. R. Loffredo, A. Iazzetti, A. Calcaterra, A. Goggiamani, F. Ghirga and M. L. Mangoni, Naturally-occurring alkaloids of plant origin as potential antimicrobials against antibiotic-resistant infections, Molecules, 2020, 25, 3619 CrossRef CAS PubMed.
- N. Jubair, M. Rajagopal, S. Chinnappan, N. B. Abdullah and A. Fatima, Review on the antibacterial mechanism of plant-derived compounds against multidrug-resistant bacteria (MDR), Evidence-Based Complementary and Alternative Medicine, 2021, 2021, 3663315 Search PubMed.
- D. J. Newman and G. M. Cragg, Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019, J. Nat. Prod., 2020, 83, 770–803 CrossRef CAS PubMed.
- G. D. Wright, Opportunities for natural products in 21 st century antibiotic discovery, Nat. Prod. Rep., 2017, 34, 694–701 RSC.
- S. E. Rossiter, M. H. Fletcher and W. M. Wuest, Natural products as platforms to overcome antibiotic resistance, Chem. Rev., 2017, 117, 12415–12474 CrossRef CAS PubMed.
- H. Lachance, S. Wetzel, K. Kumar and H. Waldmann, Charting, navigating, and populating natural product chemical space for drug discovery, J. Med. Chem., 2012, 55, 5989–6001 CrossRef CAS PubMed.
- S. Erazo, M. Zaldívar, C. Delporte, N. Backhouse, P. Tapia, E. Belmonte, F. Delle Monarche and R. Negrete, Antibacterial diterpenoids from Fabiana densa var. ramulosa, Planta Med., 2002, 68, 361–363 Search PubMed.
- F. Ghirga, R. Stefanelli, L. Cavinato, A. Lo Sciuto, S. Corradi, D. Quaglio, A. Calcaterra, B. Casciaro, M. R. Loffredo and F. Cappiello, A novel colistin adjuvant identified by virtual screening for ArnT inhibitors, J. Antimicrob. Chemother., 2020, 75, 2564–2572 CrossRef CAS PubMed.
- D. Quaglio, S. Corradi, S. Erazo, V. Vergine, S. Berardozzi, F. Sciubba, F. Cappiello, M. E. Crestoni, F. Ascenzioni and F. Imperi, Structural elucidation and antimicrobial characterization of novel diterpenoids from Fabiana densa var. ramulosa, ACS Med. Chem. Lett., 2020, 11, 760–765 CrossRef CAS PubMed.
- D. Quaglio, M. L. Mangoni, R. Stefanelli, S. Corradi, B. Casciaro, V. Vergine, F. Lucantoni, L. Cavinato, S. Cammarone and M. R. Loffredo, Ent-Beyerane diterpenes as a key platform for the development of ArnT-mediated colistin resistance inhibitors, J. Org. Chem., 2020, 85, 10891–10901 CrossRef CAS PubMed.
- V. Pastore, J. Frison, C. Pesce, M. Ryzhuk, M. Garofalo, M. Cristoferi, S. Cammarone, G. Fabrizio, M. C. B. Di Patti and D. Quaglio, Development of nanovehicles for co-delivery of colistin and ArnT inhibitors, Int. J. Pharm., 2025, 675, 125515 CrossRef CAS PubMed.
- M. A. González, Synthetic derivatives of aromatic abietane diterpenoids and their biological activities, Eur. J. Med. Chem., 2014, 87, 834–842 CrossRef PubMed.
- M. A. González, Aromatic abietane diterpenoids: Their biological activity and synthesis, Nat. Prod. Rep., 2015, 32, 684–704 Search PubMed.
- S. Ichikawa, Function-Oriented Synthesis: How to Design Simplified Analogues of Antibacterial Nucleoside Natural Products?, Chem. Rec., 2016, 16, 1106–1115 CrossRef CAS PubMed.
- P. A. Wender, V. A. Verma, T. J. Paxton and T. H. Pillow, Function-oriented synthesis, step economy, and drug design, Acc. Chem. Res., 2008, 41, 40–49 CrossRef CAS PubMed.
- I. Y. Strobykina, O. V. Andreeva, B. F. Garifullin, R. R. Sharipova and V. E. Kataev, First conjugate of glucuronic acid with triterpenoid dihydrobetulin, Russ. J. Gen. Chem., 2017, 87, 579–582 Search PubMed.
- A. V. Demchenko, Handbook of Chemical Glycosylation: Advances in Stereoselectivity and Therapeutic Relevance, John Wiley & Sons, 2008 Search PubMed.
- I. Y. Strobykina, B. F. Garifullin, R. R. Sharipova, A. D. Voloshina, A. S. Strobykina, A. B. Dobrynin and V. E. Kataev, Synthesis and antimicrobial activity of dihydrobetulin N-acetylglucosaminides, Chem. Nat. Compd., 2017, 53, 1101–1106 Search PubMed.
- K. A. Muñoz and P. J. Hergenrother, Facilitating compound entry as a means to discover antibiotics for gram-negative bacteria, Acc. Chem. Res., 2021, 54, 1322–1333 CrossRef PubMed.
- S. Hummelgaard, J. P. Vilstrup, C. Gustafsen, S. Glerup and K. Weyer, Targeting PCSK9 to tackle cardiovascular disease, Pharmacol. Ther., 2023, 249, 108480 CrossRef CAS PubMed.
- D. B. M. Dess and J. C. Martin, Readily accessible 12-I-5 oxidant for the conversion of primary and secondary alcohols to aldehydes and ketones, J. Org. Chem., 1983, 48, 4155–4156 CrossRef CAS.
- O. I. K. Afanasyev, E. Kuchuk, D. L. Usanov and D. Chusov, Reductive amination in the synthesis of pharmaceuticals, Chem. Rev., 2019, 119, 11857–11911 CrossRef CAS PubMed.
- P. Yeh, A. I. Tschumi and R. Kishony, Functional classification of drugs by properties of their pairwise interactions, Nat. Genet., 2006, 38, 489–494 Search PubMed.
- UniProt Consortium, Uniprot: the universal protein knowledgebase in 2025, Nucleic Acids Res., 2025, 53, D609–D617 Search PubMed.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek and A. Potapenko, Highly accurate protein structure prediction with AlphaFold, Nature, 2021, 596, 583–589 CrossRef CAS PubMed.
- M. McGann, FRED pose prediction and virtual screening accuracy, J. Chem. Inf. Model., 2011, 51, 578–596 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |