Open Access Article
Open Access ArticleAnalysis of metabolic characteristics of epristeride in zebrafish based on LC-Q-TOF MS and its potential applications in doping control
Yirang Wang†
a,
Zhongquan Li†a,
Jiahui Cheng† a,
Tingyuan Zheng†a,
Peijie Chenab,
Xiaojun Denga,
Pei-chao Zhang*c and
Bing Liu
a,
Tingyuan Zheng†a,
Peijie Chenab,
Xiaojun Denga,
Pei-chao Zhang*c and
Bing Liu *a
*a
aResearch Institute for Doping Control, Shanghai University of Sport, Shanghai 200438, China. E-mail: liubing2019@sus.edu.cn
bSchool of Exercise and Health, Shanghai University of Sport, Shanghai 200438, China
cThe Center for Basic Research and Innovation of Medicine and Pharmacy (MOE), School of Pharmacy, Second Military Medical University (Naval Medical University), Shanghai, P. R. China. E-mail: pczhang@smmu.edu.cn
First published on 11th March 2026
Abstract
5α Reductase inhibitors (finasteride and dutasteride) have been classified as monitored substances by the World Anti-Doping Agency (WADA) due to their potential abuse as masking agents for anabolic-androgenic steroids. As a member of the same class of inhibitors, epristeride shares a similar pharmacological mechanism of action and thus poses an inherent risk of abuse. However, current research on the metabolic characteristics of epristeride remains limited, which hinders its effective monitoring in anti-doping practices. In this study, a zebrafish model combined with liquid chromatography-quadrupole-time-of-flight mass spectrometry (LC-Q-TOF MS) was employed to systematically investigate the metabolic transformation of epristeride in vivo. Additionally, omics analysis was utilized to elucidate the effects of epristeride administration on endogenous metabolic pathways. A total of 11 epristeride metabolites were successfully identified, including 4 phase I metabolites and 7 phase II metabolites, primarily involving metabolic reactions such as oxidation, methylation, and glucuronide conjugation. Omics analysis revealed that N,N-dimethyldecylamine oxide (DDAO) with an AUC value of 0.80 could serve as a potential biomarker. Pathway enrichment analysis indicated that epristeride significantly perturbed purine metabolism and aromatic amino acid biosynthesis. These findings provide a more precise understanding of the metabolic fate of epristeride in organisms, offering a scientific basis for the development of accurate detection methods.
Introduction
Steroidal stimulants have long been a widely abused class of substances in competitive sports due to their significant effects on promoting muscle growth and enhancing athletic performance.1,2 Currently, the World Anti-Doping Agency (WADA) has adopted strict detection methods to monitor the abuse of anabolic-androgenic steroids (AAS), with urine and blood testing as the primary approaches. These methods focus on the quantitative analysis of steroid metabolites and the determination of carbon isotope ratios (δ13C) to distinguish exogenous from endogenous steroids.3However, the use of masking agents has emerged as a potential strategy for athletes to evade detection in recent years. In particular, 5α reductase inhibitors may alter the metabolism of endogenous androgens (e.g., inhibiting the conversion of testosterone to dihydrotestosterone, DHT), thereby affecting the ratios of steroid metabolites in urine (e.g., 5αAdiol/5βAdiol). This reduces the sensitivity and reliability of WADA's existing detection strategies.4–6 Significant differences exist in the pharmacokinetic profiles of different 5α reductase inhibitors.7 For example, finasteride has a half-life of 4–6 hours and is mainly metabolized in the liver to hydroxyl derivatives,8 while dutasteride has an extremely long half-life of 3–5 weeks, inhibits both type I and type II 5α reductase, and exhibits prolonged metabolic persistence.9 Both have been classified as monitored substances by WADA. As a type II 5α reductase inhibitor, epristeride displays inhibitory activity comparable to finasteride in reducing DHT levels but is less potent than the broad-spectrum inhibitor dutasteride. The differences in efficacy among the three agents primarily stem from their varying selectivities for enzyme subtypes.10–12 Notably, although epristeride biotransformation has been explored in vitro, its vivo metabolic behavior and the resulting biological context under whole-organism exposure remain insufficiently characterized, particularly with respect to Phase II conjugation patterns and potential markers relevant to doping control. Accordingly, a systematic in vivo assessment is still needed to define the metabolite spectrum and to evaluate whether epristeride produces distinctive metabolites that could inform anti-doping monitoring.
Zebrafish hold significant value in drug metabolism research, primarily attributed to their high conservation in genetic composition and metabolic functions with humans.13–16 Studies have demonstrated that zebrafish share approximately 70% of their genes with humans, and the metabolic functions of their organs (such as the liver and pancreas) are similar to those of humans. Notably, the drug metabolic pathways mediated by cytochrome P450 (CYP) enzymes are highly comparable.17–19 For instance, zebrafish CYP3A65 exhibits functional similarity to human CYP3A4 and is commonly used to predict drug–drug interactions.20 Furthermore, zebrafish possess distinct technical advantages: their small size and rapid reproduction make them suitable for high-throughput drug screening, significantly improving research efficiency and reducing costs.21,22 Examples include antitumor drugs,23 antiepileptic drugs24 and therapeutic agents for metabolic diseases.25 The transparency of embryos enables real-time in vivo imaging, allowing direct visualization of drug distribution and metabolic processes, such as the clearance of β-amyloid by gold nanoparticles.26 Meanwhile, the convenient application of gene-editing technologies like CRISPR/Cas9 facilitates the rapid and efficient construction of models with specific metabolic enzyme deficiencies or disease models, such as amyotrophic lateral sclerosis (ALS),27 autism spectrum disorder (ASD),28 and mucopolysaccharidosis (MPS),29 which are utilized for mechanism research and drug discovery.30 In terms of ethics and regulations, zebrafish embryos at specific developmental stages are not classified as laboratory animals, thus complying with the “3R” principles (Replacement, Reduction, Refinement) and providing an important alternative for drug safety evaluation.31–39 These advantages establish zebrafish as a key model bridging basic research and preclinical evaluation.
Therefore, this study employed a zebrafish model combined with liquid chromatography-quadrupole-time-of-flight mass spectrometry (LC-Q-TOF MS) technology to systematically investigate the in vivo metabolic characteristics of epristeride. Informed by prior in vitro findings, we aimed to provide in vivo characterization of epristeride biotransformation, including Phase I and Phase II metabolite profiling under whole-organism exposure, and to link exposure to endogenous metabolic responses using untargeted metabolomics Based on the similarity in steroid metabolic pathways between zebrafish and humans,40,41 the advantages of zebrafish, including simple operation, short experimental cycle, and high throughput, were exploited to rapidly screen metabolites and elucidate pharmacokinetic profiles, thereby providing a foundation for subsequent research. Furthermore, metabolomic approaches (multivariate statistical analysis) were integrated to analyze the effects of epristeride administration on in vivo metabolic pathways, which not only offers data support for optimizing existing detection strategies but also lays a basis for future clinical studies.
Materials and methods
Reagents and solutions
Methanol (HPLC grade), acetonitrile (HPLC grade), and screw-cap test tubes (10 mL) were purchased from ThermoFisher Scientific. DMSO (HPLC grade), formic acid (HPLC grade), and ammonium formate (99%) were obtained from Aladdin. Centrifuge tubes (50 mL) were purchased from Corning. The C18-PFP liquid chromatography column (150 mm × 2.1 mm, 3 µm) was acquired from ACE. Epristeride (98% analytical grade) was purchased from TargetMol. Organic PTFE microfiltration membranes (0.22 µm) and EP tubes (2 mL) were obtained from Beijing Lanjieke Technology Co., Ltd. Culture tanks (4 L) were purchased from Beijing Aisheng Technology Development Co., Ltd. Disposable syringes (1 mL) were obtained from Kefu Medical Technology Co., Ltd. Filter paper was purchased from Cytiva.Study subjects
The animal experiments in this study have been approved by the Institutional Animal Care and Use Committee (IACUC Approval No.: IACUC-2024-10452-01). A total of 9 adult wild-type AB strain zebrafish (9 males, body length: 2.5–3.0 cm) at 3 months post-fertilization were used. All zebrafish were reared in fish culture water maintained at 28 °C (water quality parameters: 200 mg of instant sea salt added to 1 L of reverse osmosis water, conductivity: 450–550 µS cm; pH: 6.5–8.5; hardness: 50–100 mg L−1 CaCO3) and were provided by the company's fish breeding center. The Laboratory Animal Use License No. is SYXK (Zhe) 2022-0004, and the rearing and management procedures comply with the requirements of the international AAALAC accreditation (Accreditation No.: 001458). All animal experiments were conducted at the company, and the processed samples were stored frozen for subsequent detection and analysis.Experimental protocol
Prior to the experiment, zebrafish were fasted for 24 hours to avoid the interference of food residues on drug absorption and metabolism, thereby standardizing their basal metabolic levels. A total of 4 mg of epristeride powder was dissolved in 100 µL of DMSO and then diluted to 2 L with standard dilution water to achieve a final concentration of 2 µg mL−1. Nine zebrafish were randomly divided into two groups: 3 fish in the control group and 6 fish in the experimental group (further divided into two subgroups, each containing 1 L of standard dilution water). All rearing conditions except for the drug immersion concentration were maintained consistent between the experimental and control groups.Sample preparation
After 2 hours of immersion treatment in the experimental group, 3 zebrafish were retrieved, transferred to clean water to rinse off residual surface drug, and then euthanized via ice-water bath. The surface moisture of the zebrafish was blotted dry with filter paper, and each individual was transferred to a 2 mL EP tube containing 1 mL of acetonitrile solution. Tissue homogenization was performed using a cryogenic grinder (30 s grinding followed by 5 s pause per cycle) with an initial setting of 5 minutes, until the fish tissues were completely pulverized with no visible tissue residues to the naked eye. Subsequently, after 12 hours of immersion treatment, the remaining 3 zebrafish were processed following the identical protocol. For the control group, zebrafish were directly retrieved, euthanized via ice-water bath, and processed according to the afore mentioned steps. The ground EP tubes were centrifuged in a refrigerated centrifuge at 4 °C and 5000 rpm for 10 minutes. The supernatant was collected using a disposable syringe and filtered through a 0.22 µm organic PTFE microfiltration membrane. The filtrate was concentrated to dryness under nitrogen flow using a dry nitrogen evaporator at 40 °C for 4 hours. The dried residue was reconstituted by vortex mixing with methanol–water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v), and the reconstituted solution was transferred to a liner-equipped injection vial for instrumental analysis. A pooled QC sample was prepared by mixing equal aliquots of all zebrafish extracts and was injected at a regular interval throughout the analytical sequence, Three QC injections were run at the beginning of the sequence for system conditioning, followed by one pooled QC injection after every 10 study-sample injections throughout the run.
1, v/v), and the reconstituted solution was transferred to a liner-equipped injection vial for instrumental analysis. A pooled QC sample was prepared by mixing equal aliquots of all zebrafish extracts and was injected at a regular interval throughout the analytical sequence, Three QC injections were run at the beginning of the sequence for system conditioning, followed by one pooled QC injection after every 10 study-sample injections throughout the run.
Liquid chromatography quadrupole time-of-flight mass spectrometry parameters
High-resolution untargeted metabolomic analysis was performed using an AB SCIEX TripleTOF 7600 LC-Q-TOF MS instrument. Mobile phases A and B consisted of 5 mM aqueous 0.05% formic acid and methanol, respectively. The column temperature was set at 30 °C, and the gradient elution program was as follows: 2% B from 0 to 1.5 min, 2% to 50% B from 1.5 to 3.0 min, 50% to 74% B from 3.0 to 9.0 min, 74% to 95% B from 9.0 to 12.0 min, 95% B from 12.0 to 15.0 min, and 2% B from 15.1 to 21.0 min. The flow rate was 0.3 mL min−1, and the sample injection volume was 5 µL.Mass spectrometric analysis was conducted using an electrospray ionization (ESI) source with alternating positive and negative ion modes to cover a more comprehensive range of metabolite information. The instrument was operated in TOF MS-IDA-TOF MS/MS scanning mode with the Zeno fragmentation function enabled. The mass scan range was set to 50–800 Da for both TOF MS and TOF MS/MS. The optimized ion source parameters were as follows: curtain gas (CUR) at 35 psi, collision-activated dissociation (CAD) at 8 (relative units), nebulizer gas (GS1) at 50 psi, and auxiliary gas (GS2) at 55 psi. The ion source voltage was 5500 V in positive ion mode and −4500 V in negative ion mode, with the ion source temperature maintained at 500 °C. The declustering potential (DP) was 80 V in positive ion mode and −60 V in negative ion mode. The collision energy (CE) was set to 40 ± 20 eV in positive ion mode and −40 ± 20 eV in negative ion mode to ensure optimal fragmentation efficiency.
Metabolite identification
For epristeride-related metabolite identification, results were primarily interpreted from the positive ion mode because the parent compound and its related metabolite signals were markedly weaker in the negative ion mode under these conditions, leading to limited MS/MS signal quality for confident annotation. In contrast, untargeted metabolomics processing and statistical analyses were performed using both positive and negative ion datasets to capture a broader range of endogenous metabolites.Data processing and metabolite identification were performed using AB SCIEX Molecule Profiler software. Candidate epristeride-related metabolites were first screened based on accurate mass matching (mass error within 10 ppm) and chromatographic peak quality, followed by MS/MS-based structural annotation. Metabolite assignments were supported by characteristic fragment ions and/or diagnostic neutral losses consistent with expected Phase I and Phase II biotransformations, together with retention-time behavior (polarity changes) across related metabolite pairs. Database searches (e.g., METLIN and HMDB) were used as auxiliary references for chemical class and fragment interpretation. Key identification information for each metabolite (precursor m/z, mass error, major product ions, and proposed transformation) is summarized in Table 1 to ensure transparency and reproducibility.
| Compound | Biotransformation | RT (min) | Formula | ESI mode | Theoretical mass (m/z, Da) | Accurate mass (m/z, Da) | Error (ppm) | Production (m/z, Da) |
|---|---|---|---|---|---|---|---|---|
| M | — | 14.82 | C25H37NO3 | [M + H] | 400.2846 | 400.2886 | 10.0 | 344.2246, 72.0427 |
| M1 | Oxidation | 13.64 | C25H37NO4 | [M + H] | 416.2795 | 416.2792 | −0.7 | 360.2170, 119.0857 91.0528, 72.0427 |
| M2 | Oxidation | 13.99 | C25H37NO4 | [M + H] | 416.2795 | 416.2801 | 1.4 | 360.2177, 344.2199![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 299.2003, 189.0907 299.2003, 189.0907 |
| M3 | Glucuronide conjugation | 12.84 | C31H45NO9 | [M + H] | 576.3167 | 576.3159 | −1.4 | 520.2545, 400.2841![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 344.2218, 326.2115 344.2218, 326.2115 |
| M4 | Loss of C4H8 | 14.82 | C21H29NO3 | [M + H] | 344.2220 | 344.2225 | 1.5 | 309.1840, 273.1842![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 157.1006, 72.0443 157.1006, 72.0443 |
| M5 | Methylation | 14.82 | C26H39NO3 | [M + H] | 414.3003 | 414.2997 | −1.4 | 339.2679, 201.1636![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 175.1480 175.1480 |
| M6 | Oxidation and taurine conjugation | 11.10 | C27H42N2O6S | [M + H] | 523.2836 | 523.2843 | 1.3 | 406.2060, 398.2699![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 281.1903, 253.1953 281.1903, 253.1953 |
| M7 | Loss of C4H8 and glucuronidation | 12.84 | C27H37NO9 | [M + H] | 520.2541 | 520.2543 | 0.4 | 344.2225, 326.2108![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 281.1970 281.1970 |
| M8 | Loss of C4H8 and taurine conjugation | 12.67 | C23H34N2O5S | [M + H] | 451.2261 | 451.2282 | 4.7 | 326.2125, 281.1912![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 253.1936 253.1936 |
| M9 | Cysteine conjugation | 13.61 | C28H42N2O4S | [M + H] | 503.2938 | 503.2938 | 0.0 | 382.2743, 326.2116 |
| M10 | Taurine conjugation | 12.66 | C27H42N2O5S | [M + H] | 507.2887 | 507.2913 | 5.1 | 382.2753, 326.2125![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 281.1904 281.1904 |
| M11 | Oxidation and glucuronide conjugation | 11.96 | C31H45NO10 | [M + H] | 592.3116 | 592.3131 | 2.5 | 416.2800, 360.2172![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 342.2075 342.2075 |
Multivariate statistical analysis was conducted using SIMCA-P software, including principal component analysis (PCA) and orthogonal partial least squares-discriminant analysis (OPLS-DA), to identify metabolic signature differences between the experimental group and the control group, and to further screen for significantly altered metabolites. Additionally, heatmap analysis, cluster analysis, and other methods were utilized to explore metabolic patterns among different samples and inter-individual metabolic variations.
Results and discussion
Metabolite identification
In this study, systematic identification and structural elucidation of the metabolites of epristeride in zebrafish were performed using a LC-Q-TOF high-resolution mass spectrometry platform, combined with MS/MS fragmentation information. To simulate the in vivo metabolic process of epristeride, a drug exposure experiment was conducted on zebrafish. After the obtained samples were subjected to protein precipitation, centrifugation, and supernatant extraction, high-resolution untargeted metabolomic analysis was performed to screen for potential long-acting metabolites.The metabolic products of epristeride were identified by comparing retention times (RT) in chromatograms and characteristic fragment ions in mass spectra via LC-Q-TOF mass spectrometry analysis. A total of 11 metabolites of epristeride were detected in the zebrafish model, including 4 phase I metabolites (M1–M2, M4–M5) and 7 phase II metabolites (M3, M7–M11). Representative extracted-ion chromatograms (XICs) and full MS/MS spectra for epristeride and all proposed metabolites (M1–M11) are provided in the SI (see Fig. S1 and S2). The mass deviations between the measured values and theoretical values were all within 10 ppm, with the exception of M10. Among which metabolites M1–M3 were consistent with the previously reported results in vitro liver microsomal metabolism42 (Table 1). No parent drug or metabolites were detected in the blank control group. The metabolic transformation pathways of epristeride are illustrated in Fig. 1, mainly involving various metabolic reactions such as oxidation, methylation, dealkylation (loss of C4H8), as well as conjugation with glucuronide, cysteine, and taurine.
The selection of the 2 h and 12 h time points allowed for the identification of both early Phase I products and late stage Phase II conjugates of epristeride. However, the absence of intermediate samples such as those at 4 h or 6 h as well as observations beyond 12 h is recognized as a study limitation. While these two time points provided sufficient information for the initial characterization of the metabolic profile, future investigations should employ a more frequent sampling strategy to define the complete metabolic time course and determine the specific detection window for epristeride more accurately.
Among the identified metabolites, M1 and M2 are both monohydroxylated metabolites, M5 is a methylated metabolite, and M4 is a dealkylated metabolite. These metabolic forms are relatively common in the biotransformation of steroids and 5α reductase inhibitors.43–45 M3 and M7–M11 are mainly involved in phase II metabolic processes such as glucuronide conjugation, taurine conjugation, and cysteine conjugation. These metabolic reactions play a crucial role in the biotransformation and excretion of drugs.46
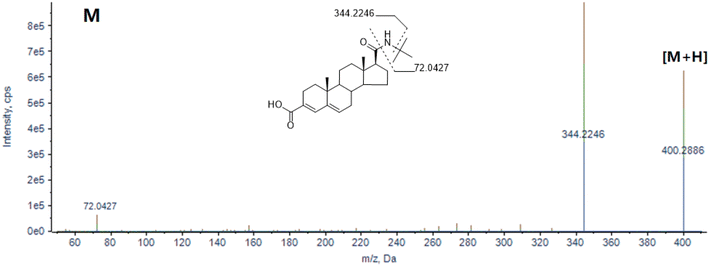
Epristeride (M), as the parent compound in this study, exhibited typical steroid structural characteristics under high-resolution UPLC-Q-TOF-MS analysis. Fig. 2 shows the MS/MS spectrum of its parent ion, with the protonated molecular ion peak at m/z 400.2848, corresponding to the [M + H]+ form of epristeride, indicating a molecular weight of approximately 399.28 Da. This parent ion displayed a stable mass-to-charge ratio (m/z) in positive ion mode. As inferred from its chemical structure, the molecule possesses a tetracyclic steroid core and a side chain containing a tert-butyl group. Epristeride is a non-competitive type II 5α reductase inhibitor, featuring a typical unsaturated open-ring steroid skeleton and a substituted side chain in its structure. The side chain contains a tert-butyl group and a carbamoyl moiety, endowing it with excellent receptor-binding specificity and enzymatic inhibitory stability. The lipophilic groups in its structure also confer strong tissue affinity in vivo, facilitating its metabolic conversion into various structural derivatives. In its MS/MS spectrum, a typical secondary fragment ion peak at m/z 344.2246 was also observed. This ion is presumably generated by the cleavage of the tert-butyl side chain at the amide position, representing a product that retains the main steroid backbone. This fragmentation pattern is commonly observed in the MS/MS analysis of steroid compounds,47 particularly the cleavage of carbon–nitrogen or carbon–carbon bonds leading to the loss of small-molecule moieties. This indicates that the parent drug possesses certain stability and characteristic fragmentation pathways. Unlike other steroid drugs, epristeride exhibits fewer high-mass peaks in its mass spectrum due to the substitution of its tert-butyl group. This stability also provides a reference basis for the structural elucidation of subsequent metabolites.
M1 and M2 were tentatively assigned as mono-oxidation metabolites of the parent drug formed at different positions. The protonated molecular ion of M1 was observed at m/z 416.2792, which is 16 Da higher than that of the parent compound, consistent with the introduction of one oxygen atom. In the MS/MS spectrum of M1, the fragment ion at m/z 360.2170 showed a marked change in relative abundance compared with the parent compound, suggesting that the oxidation likely occurs on the substructure contributing to this fragment. However, because MS/MS data alone may not distinguish positional isomers, the oxidation site of M1 should be considered putative and requires further confirmation by additional independent evidence. For M2, fragments such as m/z 327.1913 and m/z 361.2203 were observed in its mass spectrum, suggesting that hydroxylation occurred in the side chain region linked to the D-ring. This hydroxylation resulted in the retention of strong steroid skeleton characteristics even after side chain cleavage, which is consistent with the results of in vitro incubation metabolism.42 M4 is formed by dealkylation of the parent structure at the C17 side chain, with a molecular ion peak at m/z 344.2223. It can be inferred that this metabolite is generated by the loss of a C4H8 side chain from the parent drug, indicating a typical β-cleavage reaction. The molecular ion peak of M5 is m/z 414.2997, which is 14 Da higher than that of the parent drug, presumably formed by dehydrogenation accompanied by methylation. Typical signals such as m/z 321.2575 and m/z 339.2679 were detected in its MS/MS spectrum. Such modifications are uncommon in steroid drugs, suggesting that they may be caused by microbial metabolism or specific enzymatic modifications (Fig. 3).
Seven phase II metabolites were successively analyzed, as shown in Fig. 4. M3 is a typical phase II metabolite with a molecular ion peak at m/z 576.3159, which is 176 Da higher than that of the parent drug, corresponding to a glucuronic acid moiety. In its MS/MS spectrum, m/z 344.2218 corresponds to the parent-related backbone fragment, while m/z 400.2841, m/z 273.1853, and other fragments are characteristic ions of the parent steroid skeleton. Based on comprehensive analysis of the fragmentation data, the glucuronic acid conjugation site of M3 is presumably located at the C3 carboxyl group, forming an acyl glucuronide conjugate, which is consistent with previous literature reports.42 M3 exhibited a high detection signal intensity and long duration, making it a potential long-acting metabolite of focus in this study. M6 (m/z 523.2843) is a typical taurine conjugate; the characteristic fragments m/z 326.2109 and m/z 398.2699 support that it is derived from M2 via taurine conjugation. M7 (m/z 520.2543) is the glucuronide conjugate of M4; the presence of m/z 344.2225 and m/z 321.2108 in the fragments indicates the form after glycosidic bond cleavage, while m/z 281.1970 corresponds to the form after cleavage of the C17 amide functional group. The molecular ion of M8 is m/z 451.2282, with fragmentation characteristics such as m/z 281.1912 and m/z 145.1015 observed in its MS/MS spectrum, suggesting that it is a phase II metabolite formed by the conjugation of M4 with taurine. The molecular ion peak of M9 is m/z 503.2938, which is presumably the product of the parent drug conjugated with a cysteine moiety via dehydration. The presence of characteristic fragments such as m/z 382.2743 and 326.2116 in its MS/MS spectrum indicates that the parent steroid skeleton is largely retained, with cleavage occurring in the side chain region. The molecular ion peak of M10 is m/z 507.2913, suggesting that it may be the product of the parent drug conjugated with a taurine moiety via dehydration. The molecular ion peak of M11 is m/z 592.3131, which is the glucuronide conjugate of M1; the fragment m/z 360.2172 and the parent ion m/z 592.3131 provide evidence for structural confirmation. Multiply conjugated metabolites exhibit high polarity and excretion efficiency, and are commonly observed in the late metabolic stages of steroid drugs. Importantly, compared with prior in vitro work that mainly described biotransformation pathways, our current study provides in vivo evidence for these Phase II conjugation patterns under physiological exposure, which helps to prioritize metabolites with potential relevance to persistence and detection.
To further interpret our findings, we compared the metabolism of epristeride with that reported for other 5α reductase inhibitors, particularly finasteride and dutasteride. Overall, these inhibitors share common biotransformation themes, with oxidative Phase I reactions typically followed by Phase II conjugation to increase polarity and facilitate elimination. In our zebrafish in vivo study, epristeride generated multiple Phase I products and prominent Phase II conjugates, indicating an active secondary metabolism under whole-organism exposure. Differences in chemical structure and isoform selectivity may contribute to variation in metabolic routes and persistence among these inhibitors, which is directly relevant for anti-doping applications. Specifically, while class-wide metabolic patterns are shared, the metabolite profile observed here suggests that epristeride may require an inhibitor-specific marker panel and time-window evaluation rather than relying on markers established for finasteride or dutasteride. Further validation in mammalian models and human matrices will be needed to confirm translational relevance.
The results of OPLS-DA model analysis showed that the samples were clearly separated in metabolic patterns before and after administration, suggesting that epristeride intake was associated with changes in the body metabolic pathways (Fig. 5a). In the model validation analysis, the R2 value (>0.72) indicated a good model fitting degree, while the relatively low Q2 predictive ability indicates limited predictive generalization and suggests that inter-individual metabolic variation may affect model stability (Fig. 5b). Accordingly, we treat the OPLS-DA results as exploratory and use them for feature prioritization rather than as evidence of strong predictive performance. Specific information of differential metabolites was confirmed based on statistical analysis results, as shown in Tables 2 and S1. A total of 60 common compounds were identified, among which 21 metabolites exhibited significant differences, including 6 downregulated and 15 upregulated metabolites. To evaluate the group discrimination ability of differential metabolites and clarify their potential application value as potential biomarkers, ROC curve plotting and AUC value calculation were performed for each of the significantly differential metabolites. The results revealed obvious variations in the discrimination ability among different differential metabolites, with 3 metabolites showing better performance (Fig. 6). The upregulated metabolite Hexanoyl-L-carnitine had an AUC value of 0.71, which was classified as “fair” according to AUC grading standards. This indicates that it has certain discriminative potential for the two groups of samples, but its discrimination efficiency is limited when used alone, possibly due to overlapping metabolic phenotypes in some samples. The downregulated metabolite 11-Hydroxydocosahexaenoic acid (11-HDoHE) had an AUC value of 0.74, with slightly better discrimination ability than Hexanoyl-L-carnitine, also belonging to the “fair” grade. Its expression difference between the two groups was more significant, leading to relatively higher reliability in group discrimination. In contrast, the downregulated metabolite N,N-dimethyldecylamine oxide achieved an AUC value of 0.80, which exactly reached the critical threshold for “good” discrimination ability. This demonstrates that this metabolite shows good discrimination in the current dataset; however, given the modest Q2 and the limited sample size, biomarker screening is considered exploratory and requires independent validation. In this study, we employed the AUC to evaluate the classification performance of the model. Although the AUC is a widely recognized metric that can intuitively reflect the model's ability to distinguish between different groups, we are well aware of its limitations in small sample data. We plan to further validate the results in future studies by increasing the sample size and employing more sophisticated data analysis methods.
 | ||
| Fig. 5 (a) OPLS-DA score plot of zebrafish before and after administration; (b) OPLS-DA model evaluation plot of zebrafish before and after administration. | ||
| Var ID (primary) | VIP_pred | Fold change | Variation trend | ROC (AUC) |
|---|---|---|---|---|
| Hexanoy-L-carnitine | 2.475 | 1.951 | ↑ | 0.71 |
| 11-HDoHE | 2.054 | 2.084 | ↓ | 0.74 |
| Decanoylcarnitine | 2.023 | 1.857 | ↑ | 0.62 |
Lysophosphatidylcholine (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) 2) |
1.790 | 2.166 | ↑ | 0.63 |
| Phenylethanolamine | 1.727 | 2.236 | ↑ | 0.59 |
| Guanosine | 1.716 | 1.929 | ↑ | 0.63 |
| N,N-Dimethyldecylamine oxide | 1.592 | 0.510 | ↓ | 0.80 |
| Taurine | 1.467 | 1.781 | ↑ | 0.69 |
Lysophosphatidylcholine (18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) 1) |
1.430 | 1.560 | ↑ | 0.62 |
| Xanthine | 1.427 | 1.694 | ↑ | 0.52 |
| 4-Methyl-5-thiazoleethanol | 1.341 | 1.744 | ↑ | 0.57 |
| Phenylalanine | 1.326 | 2.113 | ↑ | 0.51 |
| Vitamin B5 (pantothenic acid) | 1.261 | 1.446 | ↑ | 0.63 |
Linoleic acid (C18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2) 2) |
1.211 | 1.947 | ↑ | 0.66 |
| Lauroylcarnitine | 1.198 | 1.054 | ↑ | 0.57 |
| Eudesmin | 1.151 | 0.744 | ↓ | 0.63 |
| Palmitoylcarnitine | 1.124 | 0.935 | ↓ | 0.53 |
| Guanine | 1.107 | 1.447 | ↑ | 0.57 |
| Inosine | 1.061 | 1.293 | ↑ | 0.51 |
| Tetradecanoylcarnitine | 1.031 | 0.986 | ↓ | 0.54 |
Given the high AUC value observed for DDAO, we further considered its biochemical properties and physiological context. Chemically, DDAO is an amine oxide that can be formed by oxidation of the corresponding tertiary amine. Such N-oxidation reactions are commonly catalyzed by flavin-containing monooxygenases and, in some cases, by specific cytochrome P450 enzymes. In this study, DDAO abundance in zebrafish was significantly reduced after epristeride exposure, suggesting a shift in amine-related oxidative metabolism under the drug challenge. Several non-exclusive mechanisms may explain this observation, including changes in hepatic oxidative capacity, altered availability of endogenous amine substrates, or indirect effects arising from broader metabolic adjustments. Importantly, the current data do not demonstrate whether the decrease in DDAO reflects direct inhibition of a specific enzyme. Therefore, DDAO is presented as a candidate marker that requires further investigation. Future work should verify its formation pathway and assess its robustness using targeted LC-MS analysis, including evaluation in mammalian models and human urine.
Subsequently, KEGG pathway enrichment analysis was performed on the differential metabolites, and the results are shown in Fig. 7. The pathways of the differential metabolites are mainly involved in two pathways: purine metabolism, and biosynthesis of phenylalanine, tyrosine and tryptophan. Combined with the metabolic pathway annotations of the three metabolites (e.g., hexanoyl-L-carnitine is involved in carnitine metabolism, and 11-HDoHE participates in the fatty acid oxidation pathway), this study can further reveal the metabolic regulatory mechanisms underlying the formation of differential phenotypes between the two groups of samples, thereby providing directions for subsequent functional verification.
Purine metabolism is essential for nucleotide synthesis and cellular energy balance. In this study, the alterations observed in purine metabolism are discussed as an exposure-related metabolic fingerprint following epristeride administration. From an anti-doping perspective, such untargeted endogenous alterations can inform screening by providing orthogonal, system-level signals of exposure that complement direct detection of epristeride-related metabolites. While drug-derived Phase I and II metabolites offer high specificity for confirmatory identification, endogenous pathway shifts (e.g., purine metabolism) may help flag samples when parent drug signals are low or when the compound is used as a potential masking agent. Furthermore, the significant impact on phenylalanine, tyrosine and tryptophan biosynthesis is interpreted as part of a broader metabolic disturbance pattern. This coordinated shift may be leveraged for anti-doping screening, for example by supporting the development of multi-marker panels and/or multivariate classifiers to improve screening sensitivity under real-world conditions. Accordingly, we view these pathway-level changes primarily as potential screening indicators of exposure, rather than as evidence to support clinical interpretation of adverse effects. However, these observations remain preliminary and further work is needed to evaluate their reproducibility. Further validation is also required to assess their applicability across species and matrices, ideally through expanded time-course experiments and targeted verification in mammalian models and human specimens.
Conclusion
This study employed LC-Q-TOF MS technology to analyze the metabolic characteristics of the 5α reductase inhibitor epristeride in a zebrafish model. Upon identification, a total of 11 metabolites of epristeride were detected, including 4 Phase I metabolites and 7 Phase II metabolites, primarily involving metabolic reactions such as oxidation, methylation, and glucuronidation. These metabolic reactions are relatively common in the biotransformation of steroids and 5α reductase inhibitors. Further metabolomic analysis revealed that 20 metabolites exhibited significant differences, with three metabolites standing out prominently. Among them, the AUC (Area Under the Curve) value of N,N-dimethyldecylamine oxide reached 0.80, making it a potential biomarker. The metabolic changes in zebrafish were mainly concentrated in purine metabolism and the biosynthesis of aromatic amino acids (phenylalanine, tyrosine, and tryptophan synthesis), suggesting that epristeride may alter energy metabolism and amino acid synthesis in organisms by influencing these key pathways. This study clarified the metabolic characteristics of epristeride in the zebrafish model, and the identified metabolite profile can provide references for drug monitoring, individual metabolic research, and the improvement of doping detection methods for epristeride.Author contributions
Yirang Wang: research concept, literature review, statistical analysis, writing – draft. Zhongquan Li: conceptualization, research concept, project administration, writing – review & editing; Yirang Wang: data collection, data analysis and interpretation, visualization; Jiahui Cheng: data analysis and interpretation, visualization, writing – draft; Tingyuan Zheng: data analysis and interpretation, visualization, writing – draft; Peijie Chen: supervision, writing – review; Xiaojun Deng: supervision, writing – review; Pei-chao Zhang: supervision, reviewing the draft; Bing Liu: supervision, writing – review, project administration. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
Data will be made available on request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra09094f.
Acknowledgements
The authors acknowledge the financial support of research project of Shanghai University of Sport (2025STD004).References
- P. Oleksak, E. Nepovimova, M. Valko, S. Alwasel, S. Alomar and K. Kuca, Comprehensive analysis of prohibited substances and methods in sports: Unveiling trends, pharmacokinetics, and WADA evolution, Environ. Toxicol. Pharmacol., 2024, 108, 104447 CrossRef CAS PubMed
.
- Z. Wenbo and Z. Yan, The Uses of Anabolic Androgenic Steroids Among Athletes; Its Positive and Negative Aspects- A Literature Review, J. Multidiscip. Healthc., 2023, 16, 4293–4305 CrossRef PubMed
.
- Z. Li, J. Cheng, C. Zhao, B. Wang, S. Yu, W. Qian, R. Wang, H. Zhang, Z. Fang and F. Guan, Advances in steroid purification for novel techniques in carbon isotope ratio mass spectrometry of doping control, RSC Adv., 2025, 15, 17548–17561 RSC
.
- U. Mareck, H. Geyer, G. Opfermann, M. Thevis and W. Schänzer, Factors influencing the steroid profile in doping control analysis, J. Mass Spectrom., 2008, 43, 877–891 CrossRef CAS
.
- T. Kuuranne, M. Saugy and N. Baume, Confounding factors and genetic polymorphism in the evaluation of individual steroid profiling, Br. J. Sports Med., 2014, 48, 848–855 CrossRef PubMed
.
- V. Rosa and S. Jordi, Masking and manipulation, Handb. Exp. Pharmacol., 2010, 195, 327–354 Search PubMed
.
- A. K. Gupta, M. Talukder and G. Williams, Emerging and traditional 5-α reductase inhibitors and androgen receptor antagonists for male androgenetic alopecia, Expert Opin. Emerg. Drugs, 2024, 29, 251–261 CrossRef CAS PubMed
.
- A. K. Gupta, M. Venkataraman, M. Talukder, D. M. Mainashah, A. G. Nusbaum, R. G. Loria and J. J. Jimenez, Finasteride for hair loss: a review, J. Dermatolog. Treat., 2022, 33, 1938–1946 CrossRef CAS PubMed
.
- J. F. M. Andrade, A. Verbinnen, A. W. Bakst, M. Cunha-Filho, G. M. Gelfuso and T. Gratieri, Topical dutasteride for androgenic alopecia: current state and prospects, Ther. Deliv., 2025, 16, 271–283 CrossRef CAS PubMed
.
- A. Shrivastava and V. B. Gupta, Various treatment options for benign prostatic hyperplasia: A current update, J. Midlife Health, 2012, 3, 10–19 Search PubMed
.
- F. P. Posastiuc, N. T. Constantin, G. Domain, A. Van Soom, A. I. Diaconescu and M. D. Codreanu, A Systematic Review of Medical Treatments for Benign Prostatic Hyperplasia in Dogs: Evaluating Strategies for Reproductive Function Preservation, Vet. Sci., 2025, 12, 70 Search PubMed
.
- A. Courtney, D. Triwongwarant, I. Chim, S. Eisman and R. Sinclair, Evaluating 5-alpha reductase inhibitors for the treatment of male androgenic alopecia, Expert Opin. Pharmacother., 2023, 24, 1919–1922 Search PubMed
.
- L. Wagmann, T. M. Gampfer and M. R. Meyer, Recent trends in drugs of abuse metabolism studies for mass spectrometry-based analytical screening procedures, Anal. Bioanal. Chem., 2021, 413, 5551–5559 Search PubMed
.
- S. Rasheed, F. Fries, R. Müller and J. Herrmann, Zebrafish: An Attractive Model to Study Staphylococcus aureus Infection and Its Use as a Drug Discovery Tool, Pharmaceuticals, 2021, 14, 594 CrossRef CAS PubMed
.
- T. M. Gampfer, V. Schütz, P. Schippers, S. Rasheed, J. Baumann, L. Wagmann, B. Pulver, F. Westphal, V. Flockerzi, R. Müller and M. R. Meyer, Metabolism and cytotoxicity studies of the two hallucinogens 1cP-LSD and 4-AcO-DET in human liver and zebrafish larvae models using LC-HRMS/MS and a high-content screening assay, J. Pharm. Biomed. Anal., 2024, 245, 116187 Search PubMed
.
- C. de Souza Anselmo, V. F. Sardela, B. F. Matias, A. C. G. de Castro, C. V. Maurer-Morelli, A. A. de Oliveira, F. W. A. Silva and D. C. Oliveira Jr, Is zebrafish (Danio rerio) a tool for human-like metabolism study, Drug Test. Anal., 2017, 9, 1685–1694 Search PubMed
.
- L. Wagmann and M. R. Meyer, Reviewing toxicokinetics with a focus on metabolism of new psychoactive substances in the zebrafish (larvae) model, WIREs Forensic Sci., 2022, 4, e1454 CrossRef
.
- M. Adhish and I. Manjubala, Effectiveness of zebrafish models in understanding human diseases-A review of models, Heliyon, 2023, 9, e14557 Search PubMed
.
- K. Howe, M. D. Clark, C. F. Torroja, J. Torrance, C. Berthelot, M. Muffato, J. E. Collins, S. Humphray, K. McLaren and L. Matthews, et al., The zebrafish reference genome sequence and its relationship to the human genome, Nature, 2013, 496, 498–503 CrossRef CAS PubMed
.
- H.-P. Tseng, T.-H. Hseu, D. R. Buhler, W. D. Wang and C. H. Hu, Constitutive and xenobiotics-induced expression of a novel CYP3A gene from zebrafish larva, Toxicol. Appl. Pharmacol., 2005, 205, 247–258 CrossRef CAS PubMed
.
- L. Costalonga Rodrigues, N. Cristine Bueno Faria dos Santos and I. Locilento Denkena, et al., A dive into the new psychoactive substances: a review of the use of zebrafish (Danio rerio) as an in vivo model, Drug Chem. Toxicol., 2025, 1–29 Search PubMed
.
- A. Małkowska, K. Ługowska, K. Grucza, W. Małkowska and D. Kwiatkowska, Ethyl glucuronide and ethyl sulfate in the zebrafish after ethanol exposure, Alcohol, 2024, 115, 33–39 CrossRef PubMed
.
- J. Mandelbaum, I. A. Shestopalov, R. E. Henderson, N. G. Chau, B. Knoechel, M. J. Wick and L. I. Zon, Zebrafish blastomere screen identifies retinoic acid suppression of MYB in adenoid cystic carcinoma, J. Exp. Med., 2018, 215, 2673–2685 Search PubMed
.
- M. Moog and S. C. Baraban, Clemizole and trazodone are effective antiseizure treatments in a zebrafish model of STXBP1 disorder, Epilepsia Open, 2022, 7, 504–511 Search PubMed
.
- Y. Zhang, J. Ear, Z. Yang, K. Morimoto, B. Zhang and S. Lin, Defects of protein production in erythroid cells revealed in a zebrafish Diamond-Blackfan anemia model for mutation in RPS19, Cell Death Dis., 2014, 5, e1352 CrossRef CAS PubMed
.
- I. Javed, G. Peng, Y. Xing, T. Yu, M. Zhao, A. Kakinen, A. Faridi, C. L. Parish, F. Ding, T. P. Davis, P. C. Ke and S. Lin, Inhibition of amyloid beta toxicity in zebrafish with a chaperone-gold nanoparticle dual strategy, Nat. Commun., 2019, 10, 3780 Search PubMed
.
- L. Lescouzères and S. A. Patten, Promising animal models for amyotrophic lateral sclerosis drug discovery: a comprehensive update, Expert Opin. Drug Discov., 2024, 19, 1213–1233 CrossRef PubMed
.
- D. Chivchibashi-Pavlova and K. Bratoeva, Animal models of autism spectrum disorder: Insights into genetic, structural and environmental models, Vet. Med., 2025, 70, 227–241 CrossRef PubMed
.
- R. Manzoli, L. Badenetti, M. Bruzzone, M. C. Macario, M. Rubin, M. Dal Maschio, A. Roveri and E. Moro, Mucopolysaccharidosis type II zebrafish model exhibits early impaired proteasomal-mediated degradation of the axon guidance receptor Dcc, Cell Death Dis., 2024, 15, 269 Search PubMed
.
- L. Sanxiong, Utilization of Zebrafish as a Model System in Medical Research, BIO Integr., 2022, 3, 188–192 Search PubMed
.
- P. García-Atienza, E. Sancho, M. D. Ferrando and S. Armenta, Danio rerio embryo as in vivo model for the evaluation of the toxicity and metabolism of pyrovalerone cathinones, Ecotoxicol. Environ. Saf., 2024, 286, 117174 CrossRef PubMed
.
- A. Morales-Noé, F. A. Esteve-Turrillas and S. Armenta, Metabolism of third generation synthetic cannabinoids using zebrafish larvae, Drug Test. Anal., 2022, 14, 594–603 CrossRef PubMed
.
- M. Murari, S. Pesavento, F. Greco, A. Vettori, F. Tagliaro and R. Gottardo, Study of metabolism and potential toxicity of nine synthetic opioid analogs using the zebrafish larvae model, Drug Test. Anal., 2023, 16, 629–637 Search PubMed
.
- L. Wagmann, F. Frankenfeld, Y. M. Park, J. Baumann, B. Pulver, F. Westphal, R. Maurer, R. Müller and M. R. Meyer, How to Study the Metabolism of New Psychoactive Substances for the Purpose of Toxicological Screenings-A Follow-Up Study Comparing Pooled Human Liver S9, HepaRG Cells, and Zebrafish Larvae, Front. Chem., 2020, 8, 539 Search PubMed
.
- J. Lachowicz, A. Szopa, K. Ignatiuk, K. Świąder, A. Serefko, E. Poleszak and G. Florek-Łuszczki, Zebrafish as an Animal Model in Cannabinoid Research, Int. J. Mol. Sci., 2023, 24, 10455 Search PubMed
.
- T. Cooman, S. A. Bergeron, R. Coltogirone, J. S. Bailey, C. L. Kerrigan and J. M. Rice, Evaluation of fentanyl toxicity and metabolism using a zebrafish model, J. Appl. Toxicol., 2022, 42, 706–714 CrossRef CAS PubMed
.
- K. T. Kirla, C. Erhart, K. J. Groh, J. Stadnicka-Michalak, R. I. L. Eggen, K. Schirmer and T. Kraemer, Zebrafish early life stages as alternative model to study designer drugs: Concordance with mammals in response to opioids, Toxicol. Appl. Pharmacol., 2021, 419, 115483 CrossRef CAS PubMed
.
- P. I. Racz, M. Wildwater, M. Rooseboom, E. Kerkhof, R. Pieters, E. S. Yebra-Pimentel, R. P. Dirks, H. P. Spaink, C. Smulders and G. F. Whale, Application of Caenorhabditis elegans (nematode) and Danio rerio embryo (zebrafish) as model systems to screen for developmental and reproductive toxicity of Piperazine compounds, Toxicol. Vitro, 2017, 44, 11–16 CrossRef CAS PubMed
.
- Z. Jie, S. Qin, W. Zhang, J. Wang, J. Lu, G. Qin, X. Hou and P. Xu, Metabolic Profile Analysis of Designer Benzodiazepine Etizolam in Zebrafish and Human Liver Microsomes, Metabolites, 2023, 13, 699 CrossRef CAS PubMed
.
- A. L. D. de Araujo, I. K. da C. Nunes, V. F. Sardela, H. M. G. Pereira, L. M. Cabral and C. de S. Anselmo, Is zebrafish (Danio rerio) water tank model applicable for the assessment of glucocorticoids metabolism The budesonide assessment, J. Chromatogr. B, 2021, 1179, 122826 CrossRef CAS PubMed
.
- R. R. Matos, C. de Souza Anselmo, V. F. Sardela and H. M. G. Pereira, Phase II stanozolol metabolism study using the zebrafish water tank (ZWT) model, J. Pharm. Biomed. Anal., 2021, 195, 113886 Search PubMed
.
- Z. Li, B. Liu, Y. Wang, J. Cheng, R. Aguilera, X. Deng, Q. Chen and P. Chen, Identification of Metabolites for the Novel 5α-Reductase Inhibitor Epristeride In Vitro and Its Potential Impact on Doping Testing, Drug Test. Anal., 2025, 1–11 Search PubMed
.
- L. Schiffer, L. Barnard, E. S. Baranowski, L. C. Gilligan, A. Taylor, W. Arlt, C. Shackleton and K.-H. Storbeck, Human steroid biosynthesis, metabolism and excretion are differentially reflected by serum and urine steroid metabolomes: A comprehensive review, J. Steroid Biochem. Mol. Biol., 2019, 194, 105439 Search PubMed
.
- A. Lundahl, A. Tevell Åberg, U. Bondesson, H. Lennernäs and M. Hedeland, High-resolution mass spectrometric investigation of the phase I and II metabolites of finasteride in pig plasma, urine and bile, Xenobiotica, 2014, 44, 498–510 Search PubMed
.
- A. Lundahl, H. Lennernäs, L. Knutson, U. Bondesson and M. Hedeland, Identification of finasteride metabolites in human bile and urine by high-performance liquid chromatography/tandem mass spectrometry, Drug Metab. Dispos., 2009, 37, 2008–2017 Search PubMed
.
- O. A. Almazroo, M. K. Miah and R. Venkataramanan, Drug Metabolism in the Liver, Clin. Liver Dis., 2017, 21, 1–20 CrossRef PubMed
.
- T. A. Gloriozova, V. V. Poroikov and V. M. Dembitsky, Synthetic steroids containing a tertiarybutyl group and their biological activities, Pharma Innov., 2017, 6, 84–92 CAS
.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |