 Open Access Article
Open Access ArticlePolarity-determined triple partition of dual binding pockets unlocks mechanistic insights and a novel antagonist design of NMDA receptors: a combined MD/DFT study
Zihan Yu abc,
Bingkun Chen
abc,
Bingkun Chen abc,
Shan Yangabd,
Pengfei Songabd,
Zhuo Zhangabd,
Yuanjun Zhuabc,
Chao Ma*abd and
Yanjuan Wang*abc
abc,
Shan Yangabd,
Pengfei Songabd,
Zhuo Zhangabd,
Yuanjun Zhuabc,
Chao Ma*abd and
Yanjuan Wang*abc
aKey Laboratory of Structure-Based Drug Design & Discovery of Ministry of Education, Shenyang Pharmaceutical University, Shenyang 110016, People's Republic of China. E-mail: machao_syphu@163.com; yanjuan00836@163.com; Tel: +8613478369377
bKey Laboratory of Intelligent Drug Design and New Drug Discovery of Liaoning Province, Shenyang Pharmaceutical University, Shenyang 110016, People's Republic of China
cSchool of Pharmacy, Shenyang Pharmaceutical University, Shenyang 110016, People's Republic of China
dSchool of Pharmaceutical Engineering, Shenyang Pharmaceutical University, Shenyang 110016, People's Republic of China
First published on 9th March 2026
Abstract
N-Methyl-D-aspartate (NMDA) receptors are key glutamate-gated ion channels regulating synaptic plasticity and cognition, which are critically associated with neurodegenerative diseases through dysregulation-induced excitotoxicity, yet their existing antagonists suffer from poor binding affinity, unfavorable safety and low selectivity. Current research studies on NMDA receptors focus mainly on the primary binding pocket, with the secondary binding pocket remaining largely unexplored. The focus of this study is to gain mechanistic insights into the dual binding pockets and establish a universal binding frame and ligand modification strategies, thereby designing novel ligands occupying both pockets with enhanced binding affinity and selectivity through a combination of molecular dynamics (MD) simulations, ABFE/RBFE calculations and quantum calculations. 500 ns MD simulations were performed on 22 NMDA protein–ligand complexes, and 24 groups of key interactions were quantified via DFT calculations. As a result, a polarity-determined triple partition hypothesis was proposed based on the comprehensive analysis of key residues within dual binding pockets, with three termini (A, N, and G) featuring their distinct regions. Novel mechanistic insights including polarity selectivity and an elongation strategy were applied in the modification and design of new antagonists with the guidance of the above-mentioned hypothesis. The newly designed ligand ANG01 integrated the complementary binding advantages of both pockets, demonstrating a significantly enhanced binding affinity.
1 Introduction
Neurodegenerative diseases represent a group of irreversible neurological disorders, including conditions such as stroke, epilepsy, Alzheimer's disease, Parkinson's disease, and Huntington's disease.1 According to a comprehensive analysis from the Global Burden of Disease Study, these conditions stand as a leading cause of disability and mortality among the elderly worldwide, placing an enormous and growing burden on healthcare systems, economies, and societies. The development of novel targeted therapeutic agents has become a critical imperative.2N-Methyl-D-aspartate (NMDA) receptors belong to the ionotropic glutamate receptor family, which is essential for synaptic plasticity and memory,3 mediating most excitatory synaptic transmissions in the mammalian brain.4 Both hyper- and hypo-activation of NMDA receptors can lead to neurological disorders, causing neurodegenerative diseases.5 Previous studies have reported that the neurological disorders mediated by NMDA receptors in the central nervous system can be attributed to NMDA receptors in astrocytes to a large extent, for the astrocytic NMDA receptors were shown to stimulate the secretion of pro-inflammatory cytokines, which interferes with the homeostatic mechanism of astrocytes and participates in the progression of neurodegenerative diseases.6 Studies have shown that the overactivation of N-methyl-D-aspartate receptors (NMDARs) is a well-established driver of excitotoxicity and neuronal loss in neurodegenerative pathologies, making the NMDAR a potential drug target for the treatment of neurodegenerative diseases.7 The inhibition of NMDA receptors significantly prevents neurological disorders,8 and the relevant antagonists show great activity in treating neurodegenerative diseases.9
GluN2B is one of the six subtypes of NMDA receptors, namely, GluN1, GluN2(A-D), and GluN3. The extrasynaptic NMDA GluN2B receptor might be significantly beneficial for treatment because it prevents neuronal cell death without disrupting the protective pathways.10 Studies examining the role of GluN2-containing NMDAR subtypes in neuronal death generally suggest that GluN2B-containing NMDARs play a dominant role in mediating neuronal toxicity by coupling to multiple neuronal death signaling complexes.11 Further research found that pathological overactivity of GluN2B-containing NMDARs is associated with neuronal death.12 Based on the aforesaid studies, GluN2B became the focus of NMDAR research.
Currently, most NMDA antagonists are still in pre-clinical trial stage. Memantine was approved by the FDA in 2003 for the treatment of moderate and severe Alzheimer's disease, while its low competitiveness and affinity cause a number of side effects including hallucinations, confusion, high blood pressure and restlessness.13 Ifenprodil had been studied as a cardiovascular modulator agent until the 1990s,14 when it was found to be able to block NMDA receptors with different mechanisms of action from other NMDAR blockers. Recent study has shown that Ifenprodil binds to the GluN1–GluN2B ATD interface, stabilizing a closed-cleft conformation that prevents the relative rotation of GluN1 and GluN2B subunits (about 15°), thereby inhibiting the coordinated movement of LBD and TMD and the activation of the ion channel.15 Despite their promising neuroprotective efficacy in preclinical studies, GluN2B-selective NMDAR antagonists like Ifenprodil have been hampered by substantial limitations including poor bioavailability, suboptimal selectivity and dose-limiting psychotomimetic side effects, underscoring the urgent need for novel compounds with optimized pharmacological profiles.16
Structural and functional studies have established the existence of two distinct allosteric pockets at the GluN1/GluN2B interface: a primary pocket targeted by Ifenprodil-like ligands and a secondary pocket bound by chemotypes such as EVT-101.4 Recent work had further delineated these cavities and functionally validated their roles through comparative analysis and mutagenesis.17 These two pockets are situated at the interface of the two protein subunits (chain A and chain B). They partially overlap at one end and extend in opposite directions at the other, collectively forming a V-shaped dual-pocket binding region (Fig. 1). Although a critical structural framework has been provided, the detailed intermolecular forces, particularly the residue-specific polarity contributions and dynamic interaction patterns that govern ligand selectivity and stability across both pockets, remain to be fully quantified and integrated. These questions have been addressed in our research by conducting comprehensive molecular dynamics simulations on a series of molecules, including initial lead compounds and novel candidates that were screened out or modified, complemented via the ABFE/RBFE method, MM/PBSA and DFT calculations.
2 Materials and methods
2.1. Software
Schrödinger suite 2021,18 GROMACS 2025.4,19 AmberTools25 and AMBER24,20 Discovery Studio4.5,21 ChemDraw20.0,22 PyMOL3.1.6.1,23 VMD1.9.4a53,24 R studio 2024,25 Gaussian09 Revision E.01,26 Orca 6.1.0![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 27 and Multiwfn 3.8
27 and Multiwfn 3.8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 28,29 were employed in this study.
28,29 were employed in this study.
2.2. Ligand and protein preparation
The structures of proteins and ligands were imported into the Maestro 12.8 software in Schrödinger suite 2021. The Ligprep module was performed to optimize all the ligands involved in this study, generating states at a pH of 7.0 ± 2.0. Force field OPLS4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 was utilized to ensure proper structural initialization and charge assignment for subsequent simulations. The crystal structure of NMDA receptor subunits GluN1 and GluN2B (PDB ID: 5EWJ31) was downloaded from the RCSB Protein Data Bank (PDB, https://www.rcsb.org/), which was co-crystallized with Ifenprodil and used to define the ligand-binding site. The downloaded protein file was optimized by the Protein Preparation Wizard module. Constructing missing side-chains and atoms, adding hydrogen atoms, removing waters beyond 5 Å from het groups and optimizing the protein conformation were applied. Protonation states of histidine, aspartic acid and glutamic acid were predicted by optimizing the hydrogen bonding network. Force field OPLS4 was utilized for minimization and relaxing steric clashes. Chain C and chain D of the protein were deleted as the whole binding pocket was situated at the interface of chain A and chain B, which were retained during molecular docking and MD simulations.
30 was utilized to ensure proper structural initialization and charge assignment for subsequent simulations. The crystal structure of NMDA receptor subunits GluN1 and GluN2B (PDB ID: 5EWJ31) was downloaded from the RCSB Protein Data Bank (PDB, https://www.rcsb.org/), which was co-crystallized with Ifenprodil and used to define the ligand-binding site. The downloaded protein file was optimized by the Protein Preparation Wizard module. Constructing missing side-chains and atoms, adding hydrogen atoms, removing waters beyond 5 Å from het groups and optimizing the protein conformation were applied. Protonation states of histidine, aspartic acid and glutamic acid were predicted by optimizing the hydrogen bonding network. Force field OPLS4 was utilized for minimization and relaxing steric clashes. Chain C and chain D of the protein were deleted as the whole binding pocket was situated at the interface of chain A and chain B, which were retained during molecular docking and MD simulations.
2.3. Molecular docking study
As a modern technology, molecular docking was used to explore the interactions between ligands and receptors and calculate the corresponding binding energy. Glide module in the Schrödinger software package was employed for molecular docking.32 The receptor box for docking was generated around the centrally located binding pocket at the interface of the chain A–chain B protein–protein complex, with a 10 Å diameter centered on the ligand coordinates to fully encompass both the ligand and key residues of the binding site. All subsequent docking procedures were performed based on this defined coordinate system. The Glide SP (Standard Precision) module was used for virtual screening, and the remaining parameters kept their default values. Lead compounds from the Binding Database33 (https://www.bindingdb.org/), 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 742
742![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 020 molecules from the Pharmit Database34 (https://pharmit.csb.pitt.edu/) and molecules that were developed or modified were applied for molecular docking, with the Glide docking scores and Glide energy evaluated to assess their strength of binding. The docking score provided a comprehensive measure that integrates all relevant energy terms to predict the binding affinity between NMDAR-GluN2B and ligands, and the Glide energy specifically represented the adjusted Coulomb–van der Waals interaction energy. Each docking process was repeated twice in verifying the reliability.
020 molecules from the Pharmit Database34 (https://pharmit.csb.pitt.edu/) and molecules that were developed or modified were applied for molecular docking, with the Glide docking scores and Glide energy evaluated to assess their strength of binding. The docking score provided a comprehensive measure that integrates all relevant energy terms to predict the binding affinity between NMDAR-GluN2B and ligands, and the Glide energy specifically represented the adjusted Coulomb–van der Waals interaction energy. Each docking process was repeated twice in verifying the reliability.
2.4. Molecular dynamics simulations
All the NMDA complex systems were subjected to molecular dynamics (MD) simulations via Desmond (v4.3) for the optimal conformation, comprehensive stability assessment of the ligand–protein complexes and verification of their interaction relationships. To more accurately simulate the ligand binding under physiological conditions, the protein structures were preprocessed using the Orientations of Proteins in Membranes (OPM) database35 to optimize their membrane positioning. The processed structures were imported into the Desmond software in Maestro 12.8 for subsequent processing.36 An orthorhombic box of 10 Å side length was applied to include all the system, with simple point charge (SPC) water as the solvent model. The OPLS4 force field was applied to position the complex within the simulation box properly. The system was neutralized with an appropriate number of counterions, with the physiological solvent containing both positively charged Na+ and negatively charged Cl− ions added. The system was energy-minimized using the Desmond module with the steepest descent and limited-memory Broyden–Fletcher–Goldfarb–Shanno (L-BFGS) algorithms.37 The minimization was considered converged when the maximum force per atom reached a threshold of 1.0 kcal mol−1 Å−1. Following minimization, the system was subjected to a standard multistage heating and equilibration protocol. This included a gradual heating phase in the NVT ensemble, comprising 12 ps short simulations with restraints on solute heavy atoms to bring the system to the target temperature of 300 K. Subsequently, the system was equilibrated in the isothermal–isobaric (NPT) ensemble through three steps including 12 ps and 24 ps simulations, where restraints were progressively relaxed to stabilize the system density. The fully equilibrated system from the final step above was used to initiate the production molecular dynamics simulation. A total amount of 500 ns MD simulation was employed for each of the complex systems under the NPT system at 300 K and 1.01325 bar, recording trajectory every 500 ps and system energy every 1.2 ps. Simulation interaction diagrams were constructed to elucidate key interactions within the complex throughout MD simulations.2.5. MD simulation trajectory analysis
Parameters were obtained for the trajectory analysis including the trajectory data of root mean square deviation (RMSD) and root mean square fluctuation (RMSF). The RMSD was calculated for all frames in the trajectory of the entire protein–ligand system to assess the equilibrium during simulation, while the RMSF played a key role in characterizing local variations of the protein. The two parameters can be expressed as follows:
 | (1) |
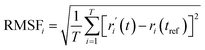 | (2) |
2.6. Molecular dynamics convergence analysis
The stability and convergence of protein–ligand simulating systems for virtual screening were rigorously validated through three complementary metrics. Global structural stability was validated by backbone RMSD and RMSF (Fig. S1–S4), binding free energy convergence was verified via MM-PBSA and ABFE/RBFE calculations yielding consistent binding affinities, and robust interaction relationships given by interaction diagrams indicated binding processivity. All these analyses were collectively utilized to confirm that a state of structural equilibrium, energetic convergence and interaction pattern stabilization has been achieved within 500 ns.382.7. Ligand-based pharmacophore modeling
The ligand-based pharmacophore model delineated the essential structural features required for binding, representing their spatial arrangement through key chemical characteristics, thereby elucidating interaction patterns and supporting the rational design of targeted ligands. The Maestro Phase package was employed for three-dimensional pharmacophore modeling via a multi-ligand strategy,39 and pharmacophore hypotheses were generated on the basis of a set of possible common pharmacophore features identified through individual ligands. For the dual binding pockets, key chemical features including hydrogen bond acceptors, hydrogen bond donors, hydrophobic groups, negative or positive ionic centers and aromatic centers were identified.2.8. Binding free energy calculation via MMPBSA − TΔS
The molecular mechanics–Poisson Boltzmann surface area (MM/PBSA) method was employed to calculate the binding free energy of complexes within dual binding pockets via gmx_MMPBSA (v1.6.4).40 Utilizing the Interaction Entropy method for entropic correction and solvation treatment, the MM/PBSA calculations implemented in this study have been shown to provide qualitatively reliable rankings of protein–ligand binding affinities.41 The structures of three independent snapshots from the equilibrated portion of 500 ns Desmond MD simulations were extracted at regular intervals as starting structures for GROMACS simulations using the CHARMM-GUI website, as the Desmond-derived MD trajectory does not support the MM/PBSA calculation.42 Three independent 100 ns simulations were conducted for each complex, given that the system had been adequately equilibrated in Desmond MD simulations, with the CHARMM36m force field43 applied. Each protein–ligand complex was embedded in a periodic water box using the TIP3P model, maintaining a minimum distance of 15 Å between the solute and box boundaries. Counterions Na+ and Cl− were added to achieve a concentration of 0.15 M using the Monte Carlo displacement. Periodic boundary conditions (PBC) were applied and non-bonded interactions were truncated at 12 Å for Lennard-Jones and Coulomb potentials, with long-range electrostatics handled by the Particle Mesh Ewald (PME) method. All bonds were constrained with the linear constraint solver (LINCS) algorithm.44 Each system was energy-minimized using the steepest descent algorithm with a max step of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and a convergence threshold of 10.0 kJ mol−1 nm−1. Subsequent equilibration was carried out using canonical (NVT) and isothermal–isobaric (NPT) ensembles,45 conducting temperature equilibration at 300 K using a Nosé–Hoover thermostat46 for 500 ps and pressure equilibration at 300 K using the Parrinello–Rahman barostat47 for another 500 ps. The binding free energy was calculated using the following equation:
000 and a convergence threshold of 10.0 kJ mol−1 nm−1. Subsequent equilibration was carried out using canonical (NVT) and isothermal–isobaric (NPT) ensembles,45 conducting temperature equilibration at 300 K using a Nosé–Hoover thermostat46 for 500 ps and pressure equilibration at 300 K using the Parrinello–Rahman barostat47 for another 500 ps. The binding free energy was calculated using the following equation:| ΔGeff = ΔEMM + ΔGsol − TΔSMM | (3) |
| ΔEMM = ΔEele + ΔEvdW | (4) |
| ΔGsol = ΔGele,sol + ΔGnonpolar,sol | (5) |
The binding free energies obtained were statistically processed, and the mean value was calculated using the standard averaging equation as the final result:
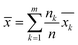 | (6) |
The MM/PBSA calculations were performed for three independent 100 ns MD simulations to ensure a robust and reproducible result. Alanine scanning mutagenesis of key residues was conducted along with the MM/PBSA calculation.49 Therefore, a highly positive ΔG value indicates the significance of the residue in the protein–ligand binding process.50 Dielectric constant ε = 4 was assigned to the charged residues ASP133, ASP136 and ASP138.
2.9. Binding free energy calculation via ABFE and RBFE
The binding free energy of selected protein–ligand complexes were rigorously quantified using alchemical free energy calculations performed with the BAT2.py program,51 with simulations executed using the AMBER24 molecular dynamics engine. Absolute binding free energy (ABFE) and relative binding free energy (RBFE) calculations were employed for the direct determination of binding affinities and for precise comparison between ligand analogues, respectively. These methods have been extensively validated and demonstrate benchmarked accuracy in predicting protein–ligand binding free energies.52 For all calculations, the calc_type = rank was utilized.53 BAT2.py automatically identified and utilized three key protein atoms as the anchor points for the alchemical transformations. The proteins and ligands were described using the ff14SB54 and GAFF55 force fields, respectively, with ligand partial charges derived at pH 7.4. Each system was solvated in a rectangular box of TIP3P water. A padding of 12 Å from the solute to the box edge was applied in each dimension for protein–ligand complexes, while a 15 Å padding was used for ligand-only systems. Systems were neutralized and brought to a physiological ion concentration of 0.15 M NaCl. Long-range electrostatics was handled using Particle Mesh Ewald with a 9 Å direct space cutoff.56 Constant temperature (298 K) was maintained using a Langevin thermostat with a collision frequency (gamma_ln) of 1.0 ps−1, and a constant pressure of 1 atm was regulated using a Monte Carlo barostat.57 Thermodynamic Integration (TI)58 was employed to estimate the free energy changes along the alchemical pathway, using 12 λ windows: [0.00921968, 0.04794137, 0.11504866, 0.20634102, 0.31608425, 0.43738330, 0.56261670, 0.68391575, 0.79365898, 0.88495134, 0.95205863, 0.99078032].The alchemical transformation for each ligand was decomposed into four distinct free energy components (m, c, e, and v), and each component was simulated independently across the aforementioned 12 λ windows. The sampling time for each λ window was 1.2 ns for the m and c components, 4.4 ns for the e component, and 4.0 ns for the v component, ensuring adequate convergence for each interaction type. Following simulation, a block data analysis procedure was applied to assess the robustness and convergence of the free energy estimates.59 The standard deviation across blocks as a conservative estimate of uncertainty was reported:
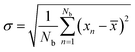 | (7) |
The production trajectories were divided into 10 sequential blocks (b1–b10). The binding free energy was calculated independently from each block, and the final reported value represented the mean of these 10 block estimates. The variance between blocks provided a measure of the statistical uncertainty and serves as a key metric for demonstrating the convergence of the calculations.
The ABFE calculations utilized the simultaneous decoupling and recoupling (SDR) method with fe_type = express.53 The binding free energy was computed as follows:
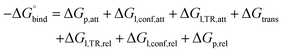 | (8) |
| ΔGtrans-SDR = ΔGelec + ΔGLJ | (9) |
The SDR transformation was performed with the ligand copy displaced by sdr_dist = 60.0 Å along the Z-axis to eliminate the protein–ligand interactions in the bulk state. The RBFE calculations employed the separate topologies (SepTop) approach with fe_type = relative.53 This method computed the difference in binding affinity, ΔGbind, between a reference ligand 1 and a target ligand 2. The computational protocol and parameters for RBFE calculations were identical to those used for the above-described ABFE calculations. The soft-core potentials for non-bonded interactions utilized the default, optimized parameters within BAT2.py. Four systems (complexes of Ifenprodil, IF03, EVT01, and ANG01) and two pairs (complexes of Ifenprodil and IF03, EVT-101 and EVT01) were strategically chosen for ABFE and RBFE calculations, respectively.
2.10. Quantum chemistry calculation for key interactions
Key interactions including hydrogen bonds and π–π stacking were quantified via Orca (v6.1.0) for the binding mechanism study. The accurate quantification of hydrogen bond (HB) energies is essential for rational drug optimization, as even subtle changes in HB strength can significantly alter the pharmacological activity.60 Density functional theory (DFT) provided atomic-level insights into these interactions, enabling the prediction and further refinement of ligand binding modes based on accurate calculation.61The use of MD-derived snapshots for single-point energy calculations emphasizes the statistical and thermodynamic behaviors of hydrogen bonds, which reflects the dynamic interactions observed in simulations. Maestro Desmond trajectory clustering in the Desmond software was employed to extract the five most representative frames from 1000 frames obtained through 500 ns MD simulations based on clustering analysis. The final interaction strength was determined by averaging the energy derived from these five representative frames.62 The geometric optimization procedure was intentionally omitted in energy calculations to preserve the physiologically relevant configurations sampled by MD simulations. The wB97M-V/def2-QZVP hybrid functional63 was applied. The SMD solvation model64 was incorporated to account for van der Waals interactions and solvent effects, respectively. To account for the basis set superposition error (BSSE), the counterpoise correction method was applied to all interaction energy calculations.65 Single-point energy calculations for the residue–ligand complexes, isolated ligands and residues were performed at the same theoretical level. The strength of a single interaction was calculated as follows:
| ΔEinteraction = Ecomplex − (Eresidue + Eligand) | (10) |
2.11. ADMET prediction
ADMET (absorption, distribution, metabolism, excretion, and toxicity), as pharmacokinetic features, are important considerations for the perspective medicines during pharmacological investigation and development phase.66 The SwissADME server (https://www.swissadme.ch/)67 and ADMETlab (https://admetmesh.scbdd.com/)68 were utilized. General characteristics including molecular weight, polarity, topological polar surface area (TPSA), flexibility, lipophilicity and water solubility of a molecule were provided. The safety profile characteristics and drug-likeness based on Lipinski's “rule of five” were also anticipated. Lipinski's “rule of five” is most well-known and widely used for ADMET evaluation, including molecular weight (less than 500), log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (less than 5), hydrogen bond donors (less than 5) and hydrogen bond acceptors (less than 10). hERG blockers, hepatotoxicity (H-HT) and AMES toxicity were estimated to give the value between 0 and 1, indicating their degrees of possibility.
P (less than 5), hydrogen bond donors (less than 5) and hydrogen bond acceptors (less than 10). hERG blockers, hepatotoxicity (H-HT) and AMES toxicity were estimated to give the value between 0 and 1, indicating their degrees of possibility.
3 Results and discussion
3.1. Integrated mechanistic analysis on action modes of two binding pockets
Interactions were observed merely on one side of Ifenprodil, Compound A and Compound C complexes. Residue GLU236 contributed to hydrogen bond formation in all complexes and exhibited a lifetime exceeding 60%. π–Cation interactions formed with ARG115 were highly stable in Ifenprodil (50%) and Compound A (59%) complexes. π–π stacking was consistently observed between PHE176 and aromatic rings on the same side of all these ligands (52% in Ifenprodil, 79% in Compound A and 40% in Compound B complexes). GLN110 served as a strong hydrogen bond acceptor in Ifenprodil (98%) and Compound C (15%) complexes, and SER132 hydrogen bonded with Compound A (100%), underscoring their critical importance. Notably, a unique π–π stacking with PHE114 on the opposite side of binding pocket was exclusive to Compound B (18%), and the formation of a strong hydrogen bond to LEU135 was distinctive to Compound C (85%).
Despite divergent core scaffolds of the four lead compounds, they shared a conserved architectural feature that aromatic rings positioned at both termini of central carbon chains. Each of them consistently formed a stable hydrogen bond with residue GLU236 at one single end (designated as G-end), while exhibiting minimal interactions at the opposite end (designated as N-end, indicating “None”). Three-dimensional binding diagrams revealed that the residue PHE114 at N-end was distal to the ligand center, hindering interactions with shorter ligands. We proposed that the elongated tetracyclic scaffold of Compound B facilitated the unique π–π stacking between its N-terminal aromatic ring and PHE114. Such interaction mode at both termini further stabilized the complex, compensating for the absence of strong hydrogen bonds. Furthermore, the fused ring system of Compound A provided superior complementarity to the G-end binding pocket, enabling a stronger π–π stacking with the G-terminal aromatic residue PHE176. Compound C exclusively interacted with LEU135 and showed weak interaction with crucial residues GLN110 or SER132. Such preferential interaction with specific residues was possibly dictated by the group polarity. It was easily recognized from 3D binding diagrams that LEU135 resided on the opposite side of the ligand to GLN110 and SER132. The polar carbonyl groups in Ifenprodil and Compound A preferentially formed strong hydrogen bonds with polar residues GLN110 and SER132. Although energy barrier associated with molecular torsion might be necessary, the process was thermodynamically favorable, resulting in a new complex with lower energy. In contrast, the sulfonyl group with weaker polarity in Compound C preferentially formed a hydrogen bond with the aliphatic residue LEU135. Therefore, properly improving the strength and number of polar groups of ligands possibly strengthened interactions with such key amino acids.
Lead compounds targeting the secondary binding pocket exhibited structural diversity, yet several common features were observed through interaction relationships. Aromatic residues (PHE113, PHE114, TYR109, and HIS134) within the N-terminal region significantly contributed to ligand binding via π–π stacking. Four aspartate residues dominated the protein–ligand binding mode on the opposite end (designated as A-end, indicating aspartate), further stabilizing complexes through hydrogen bonds and salt bridges with polar groups of ligands. The results of RMSD curves (Fig. S3) and interaction analysis from MD simulations indicated that ligand binding through the secondary pocket generally exhibited weaker binding affinity than those targeting the primary pocket.
3.2. Identification of novel potential ligands for the two binding pockets via virtual screening and lead compound modification
Five key residues (GLU236, GLN110, SER132, PHE176, and ARG115) of the primary binding pocket and four key residues (TYR109, ASP113, ASP136, and ASP138) of the secondary binding pocket were determined according to 2D interaction diagrams. The virtual screening of the Pharmit database was carried out for the validation of the dual-pocket binding modes and exploration of related mechanisms. Twelve molecules with the absolute value of Glide docking scores higher than 10 were screened out (Table S2). Predicted poses of all these molecules were subjected to 500 ns MD simulations (Fig. S2 and S61–S68), and four of them (IF01, IF03, IF08, and IF09) with stable and anticipated strong interactions were advanced for further study of primary binding pocket. It is noteworthy that the 3D binding pocket schematic revealed that all hit compounds identified through virtual screening occupied only the primary binding pocket, which directly highlighted the necessity of an in-depth investigation into the secondary binding site.Additionally, since initial virtual screening failed to identify potential ligands for the secondary binding pocket, Compound E and F were modified to EVT01 and EVT02 guided by analysis of the ligand–residue interaction characteristics within the secondary binding pocket, aiming to verify the binding features.
3.3. Molecular dynamics simulations study of identified ligands
Results from 500 ns MD simulations of the identified ligands (IF01, IF03, IF08, IF09, EVT01, and EVT02) were employed to further validate the identified binding modes and investigate potentially undiscovered mechanisms. MD trajectories, RMSD, RMSF values (Fig. S2 and S4) and protein–ligand interaction diagrams (Fig. 5) were included.3.3.1.1. Protein–ligand interaction analysis in the ligand-targeting primary binding pocket. Fig. 5A–D depict the dynamic interactions between the identified ligands and the key residues within the primary binding pocket throughout the 500 ns MD simulation. The residue GLU236 formed strong hydrogen bonds with all the ligands (>65%), demonstrating its decisive role in stabilizing complexes within the primary binding pocket. PHE176 formed strong π–π stacking with three ligands, particularly IF01 (83%), and ARG115 engaged in π–cation interactions with the G-terminal benzene ring of IF03 (13%) and IF08 (48%). The consistent presence of polar groups in all ligands enhanced their interactions with the residue GLN110, resulting in strong hydrogen bonds in IF01, 03, and 09 complexes (>75%) and two water bridges in the IF08 complex (34% and 42%). Notably, GLN110 formed strong hydrogen bonds with three polar groups of IF03 (>90%), respectively, validating our previous conclusion regarding the preference for polar interactions with such amino acids. Unlike the other ligands, IF03 exhibited unique π–π stacking with the N-terminal aromatic residue PHE114 (50%), resembling the behavior of Compound B. π–Cation interactions with PHE114 (42%) and PHE113 (70%) were found in the IF08 complex due to its lack of N-terminal benzene ring. Overall, the strong conservation of binding modes between these potential ligands and lead compounds of the primary binding pocket validated the mechanistic insights we proposed, suggesting their promising activity.
3.3.1.2. Protein–ligand interaction analysis in the ligand-targeting secondary binding pocket. Fig. 5E and F depict the dynamic interactions between the modified ligands and key residues within the secondary binding pocket throughout 500 ns MD simulations. Modified ligands EVT01 and EVT02 exhibited pronounced interactions with the residue TYR109 (52% and 60%), confirming its critical role in the secondary binding pocket. Two newly introduced amino groups located at the A-end of EVT01 formed hydrogen bonds with ASP138 (38%), and the N-terminal aromatic ring engaged in π–π stacking with PHE114 (24%). The amide group introduced at the A-end of EVT02 played a major role in interactions with surrounding aspartates at this end of the pocket. Notably, the central amidine group of EVT02 formed a strong hydrogen bond with the polar residue GLN110 (79%). Such polar interactions were mediated via water bridges in the MD simulations of Compound F before modification. Their difference in binding modes possibly arose from the introduced electron-withdrawing substituents in EVT02, which reduced the electron density of the benzene ring and weakened its conjugative electron-donating ability. The enhanced amidine polarity significantly promoted a stronger interaction with the surrounding polar amino acids, which was in consistence with the polarity selectivity of GLN110 that we observed.
Few ligands were estimated to interact with N-terminal residues across all evaluated complexes except π–π stacking between PHE114 and Compound B, π–π stacking between PHE114 and IF03, and π–cation interactions in IF08. Three-dimensional binding diagrams demonstrated that these residues were situated at a considerable distance from the ligands. The more extended carbon chain owned by Compound B, IF03 and IF08 successfully bridged this gap, and such filling effect facilitated interactions between the terminal aromatic rings of the ligands and the aromatic residues at the distal end (particularly N-end) of the binding pocket. Additionally, the modification of the terminal aromatic ring into a fused system similar to Compound A further strengthened interactions within the terminal region via enhancing the protein–ligand complementarity.
Based on the MD simulation study and pharmacophore modeling, we visualized key residues commonly involved in the formation of complexes within the dual binding pockets (Fig. 7C). The secondary and primary binding pockets featured an A-terminal polar interaction zone (aspartates) and a G-terminal polar/hydrophobic hybrid interaction zone (GLU236 and PHE176), respectively, and they overlapped at the N-terminal hydrophobic interaction zone (PHE114 and TYR109). This pronounced that the regionalization of interaction characteristics dividing the whole binding pockets into three parts was clearly demonstrated in Fig. 8. The distinct region-specific interaction mode, dictated by the binding pockets, revealed the essence of protein–ligand binding mechanism for the NMDA receptor and provided a universal binding framework, thereby directly guiding the discovery and modification of potential active compounds.
3.4. Binding free energy calculations
The residues GLU236, GLN110, PHE176, TYR109, PHE114, and ASP138 were mutated to alanine in alanine scanning mutagenesis conducted together with MM/PBSA (Table 1). The ΔG contribution from GLN110 was optimal for the IF03 complex (5.56 ± 0.79 kcal mol−1), which was consistent with MD simulation conclusions. TYR109 exhibited significantly higher ΔG values than other residues within the secondary binding pocket, demonstrating its dominant contribution. ΔG contributions from the residues of secondary binding pocket were smaller than those of the primary pocket, which was consistent with the overall binding free energy results.
| Index | Complex system | Wild type | ΔG (kcal mol−1) |
|---|---|---|---|
| 1 | 5EWJ/IF01 | GLU236 | 5.83 ± 1.74 |
| 2 | 5EWJ/IF01 | GLN110 | 4.82 ± 1.30 |
| 3 | 5EWJ/IF01 | PHE176 | 2.56 ± 1.58 |
| 4 | 5EWJ/IF03 | GLU236 | 6.08 ± 1.34 |
| 5 | 5EWJ/IF03 | GLN110 | 5.56 ± 0.79 |
| 6 | 5EWJ/IF03 | PHE176 | 2.90 ± 2.84 |
| 7 | 5EWJ/IF08 | GLU236 | 4.28 ± 1.42 |
| 8 | 5EWJ/IF08 | GLN110 | 3.11 ± 1.00 |
| 9 | 5EWJ/IF08 | PHE176 | 0.15 ± 1.59 |
| 10 | 5EWJ/IF09 | GLU236 | 5.81 ± 1.56 |
| 11 | 5EWJ/IF09 | GLN110 | 4.28 ± 2.18 |
| 12 | 5EWJ/IF09 | PHE176 | 2.81 ± 1.48 |
| 13 | 5EWJ/EVT01 | TYR109 | 5.25 ± 0.92 |
| 14 | 5EWJ/EVT01 | PHE114 | 1.70 ± 1.42 |
| 15 | 5EWJ/EVT01 | ASP138 | 0.83 ± 0.55 |
| 16 | 5EWJ/EVT02 | TYR109 | 8.08 ± 1.02 |
| 17 | 5EWJ/EVT02 | PHE114 | 2.42 ± 1.53 |
| 18 | 5EWJ/EVT02 | ASP138 | 0.59 ± 0.57 |
The mean values and standard deviations derived from ten simulation blocks for each free energy component and the total binding free energy across the four systems are presented in Table 2. Despite statistical fluctuations, the significant differences in binding free energy among the ligands robustly substantiated the conclusion of this study.76 EVT01, which occupied the secondary pocket, exhibited the weakest binding affinity (−5.12 ± 1.47 kcal mol−1), thereby validating the conclusions from MD simulations regarding suboptimal interactions in this pocket. IF03 (−14.33 ± 2.33 kcal mol−1) demonstrated a stronger binding mode than Ifenprodil (−10.94 ± 0.93 kcal mol−1), confirming the effectiveness of introducing polar groups and extending the ligand scaffold within the primary pocket. The consistency between the ABFE results and MM/PBSA calculations strongly supported the conclusions regarding dual-pocket binding hypothesis proposed in this study.
| Index | Compound | ΔGattach | ΔGelec | ΔGLJ | ΔGbind |
|---|---|---|---|---|---|
| 1 | Ifenprodil | 7.94 ± 0.10 | 9.55 ± 0.78 | 15.49 ± 0.18 | −10.94 ± 0.93 |
| 2 | IF03 | 10.83 ± 0.55 | −3.16 ± 0.53 | 29.46 ± 2.16 | −14.33 ± 2.33 |
| 3 | EVT01 | 10.78 ± 0.77 | −8.35 ± 0.35 | 21.14 ± 1.19 | −5.12 ± 1.47 |
| 4 | ANG01 | 11.51 ± 0.55 | 4.99 ± 1.05 | 22.75 ± 1.10 | −16.17 ± 1.62 |
The RBFE calculations were performed on two ligand pairs (Ifenprodil and IF03, and EVT-101 and EVT01) to separately assess the thermodynamic impact of chemical modifications targeting primary and secondary binding pockets on complex stability (Table 3). The RBFE for IF03 relative to Ifenprodil yielded a value of −0.82 kcal mol−1, which demonstrated that extending the ligand length and introducing additional polar groups enhanced binding interactions within the primary pocket. Notably, a substantial RBFE of −6.03 kcal mol−1 was obtained for EVT01 compared to the lead compound EVT-101. This pronounced change directly confirmed the success of our ligand modifications targeting secondary pocket and validated the critical contribution of the terminal aspartate residues to overall complex stability.
| Group | Id | Compound | Attach_all | Ele-TI | LJ-TI | Release-TR | Release-CF | ΔG | ΔGBAT-x1→2 |
|---|---|---|---|---|---|---|---|---|---|
| a | a-1 | Ifenprodil | 9.58 | −1.57 | 4.71 | −10.38 | −10.39 | 8.05 | −0.82 |
| a | a-2 | IF03 | 10.86 | 4.62 | 0.43 | −11.40 | −11.73 | 7.23 | |
| b | b-1 | EVT-101 | 10.32 | −4.48 | −0.95 | −11.48 | −12.62 | 19.20 | −6.03 |
| b | b-2 | EVT01 | 8.34 | −4.66 | 2.93 | −11.31 | −8.46 | 13.17 |
3.5. Quantum chemistry calculation for key interactions
Key interactions including hydrogen bonds, halogen bonds and π–π stacking were systematically investigated utilizing DFT calculations. Our comprehensive interaction calculations of the eight ligands (Ifenprodil, IF03, IF09, Compound A, Compound F, EVT-101, EVT02, and ANG01) within dual binding pockets established a quantitative foundation for the proposed polarity-determined triple partition hypothesis.| Index | Interaction system | Energy | Index | Interaction system | Energy |
|---|---|---|---|---|---|
| 1 | Ifenprodil/GLN110 | −7.61 | 13 | Compound A/ARG115 | −1.28 |
| 2 | Ifenprodil/GLU236 | −6.28 | 14 | Compound A/PHE176 | −5.40 |
| 3 | Ifenprodil/SER132 | −1.70 | 15 | IF03/PHE114 | −2.64 |
| 4 | IF03/GLN110 | −6.55 | 16 | EVT-101/TYR109 | −6.92 |
| 5 | IF03/GLU236 | −4.67 | 17 | EVT-101/ASP138 | −2.66 |
| 6 | IF03/SER132 | −2.60 | 18 | Compound F/ASP136 | −1.89 |
| 7 | IF09/GLN110 | −7.67 | 19 | Compound F/ASP138 | −1.48 |
| 8 | IF09/GLU236 | −5.56 | 20 | EVT02/GLN110 | −8.63 |
| 9 | IF09/SER132 | −3.01 | 21 | ANG01/GLU236 | −5.64 |
| 10 | Ifenprodil/ARG115 | −1.20 | 22 | ANG01/GLN110 | −6.48 |
| 11 | Ifenprodil/PHE114 | −1.92 | 23 | ANG01/PHE114 | −2.87 |
| 12 | Ifenprodil/PHE176 | −5.09 | 24 | ANG01/ASP113 | −2.61 |
The general exceptional hydrogen bonds with GLN110 underscored its essential role in complex binding. For the active reference Ifenprodil, the strong and stable HB interaction with GLN110 (−7.61 kcal mol−1) dominated its binding mode, and interaction with GLU236 failed to keep stable ranging from −3.96 to −7.20 kcal mol−1. Variations in HB strength across simulation frames likely stemmed from thermal fluctuations altering donor–acceptor angles and distances,77 which discredited the dominant binding contribution of HB. IF09 exhibited a stronger HB with GLN110 (7.67 kcal mol−1) and a more stable HB with GLU236 (ranging from −5.21 to −5.99 kcal mol−1), indicating its stronger binding affinity.
Compound A featured a fused-ring at one terminus and IF03 was characterized by an extended scaffold. Strong π–π stacking was observed between PHE176 and these two ligands, which was consistent with our determination of key amino acids. Hydrophobic interactions between Compound A and ARG115 (−1.28 kcal mol−1), Compound A and PHE176 (−5.40 kcal mol−1), and IF03 and PHE114 (−2.64 kcal mol−1) were stronger than corresponding interactions observed in Ifenprodil. The overall enhancement of π–π stacking exerted superior binding affinity we observed on IF03 and Compound A, corroborating our elongation strategies to strengthen π–π stacking with aromatic residues at the ends of the pocket.
3.6. Design of novel ligands occupying dual binding pockets
The scaffold of pre-ANG01 was extended to enhance the interactions with key residues at both ends, and the number of hydrogen bond donors and acceptors was minimized to maintain favorable drug-likeness. The resulting molecule pre-ANG02 achieved anticipated interactions within a 20 ns MD simulation, including GLN-110 (77%), PHE-114 (23%) and GLU-236 (134%). Subsequent structural extension toward the secondary binding pocket was initiated from the N-terminal aromatic ring. Guided by polarity–selectivity hypothesis, a polar group was introduced to enhance interactions with surrounding polar residues. Ligand pre-ANG03 simultaneously formed a hydrogen bond (84%) and a salt bridge (30%) with ASP-113. To accommodate the polar residues and fill the cavity of A-terminal region, a phenol group was introduced in the fourth step, yielding the final compound ANG01. Our modification process allowed stepwise optimization based on changes in key residual interactions surrounding the ligand, ensuring both scientific rigor and maximal interactions of the resulting molecule.
The ABFE calculation further confirmed the superior stability of the ANG01 complex (−16.17 ± 1.62 kcal mol−1), which exhibited a lower ΔGbind value than the ligands targeting one single pocket (Table 2). The result demonstrated that simultaneous engagement of both binding pockets could significantly enhance the ligand binding affinity. Furthermore, the key interaction energy calculated for each group (Table 4, index 21–24) was comparable to those of the previously studied ligands within the single pocket, suggesting that the cooperative interactions with residues from both pockets further stabilized the complex system compared to binding within a single pocket. Meanwhile, the improved structural complementarity achieved through this binding mode would possibly contribute to a higher selectivity toward the NMDA receptor.78 ADME predictions demonstrated high GI absorption and bioavailability of ANG01, as well as full compliance with Lipinski, Pfizer rule and Golden-triangle. The results of our comprehensive analysis positioned ANG01 as a promising lead compound for the development of novel NMDA receptor antagonists with a higher binding affinity and enhanced selectivity. Subsequent structural optimizations and pharmacological tests are pursued with the goal of minimizing the potential toxicity of such molecules.
4 Conclusions
Although numerous NMDA receptor antagonists with known activity have been reported experimentally, the dual-pocket binding mechanism, particularly regarding the secondary binding pocket and their integrated exploration, has received limited attention and remained unexplored. In this study, integrated computational methods were applied to gain mechanistic insights into the primary and secondary binding sites, including 500 ns MD simulations of 22 complex systems, ABFE/RBFE calculations for six complex groups and DFT calculations for 20 interactions. Residues GLU236, GLN110, SER132, PHE176, ARG115, TYR109, PHE114, ASP136, and ASP138 were demonstrated as key amino acids in the dual-pocket binding process. Polarity-determined triple partition hypothesis of dual binding pockets was proposed as a universal binding frame, dividing the whole binding region into polar zone, hydrophobic zone and polar/hydrophobic hybrid zone, featuring three termini (A, N, and G), unlocking the essence of the NMDAR-ligand binding mode. Novel mechanistic insights including polarity selectivity and elongated complementarity were defined as the decoration of this universal frame. The new ligand ANG01 occupying both binding pockets was designed based on the above hypothesis and modification strategies, which exhibited significantly improved binding affinity and favorable ADME properties. Additionally, two potential NMDAR antagonists IF03 and IF09 (cid: 121503195 and 71751913) were discovered via virtual screening. In the future study, pharmacological evaluation and further structural optimization of ANG01 are warranted to eliminate the potential toxicity, advancing such compounds towards highly efficacious NMDAR antagonists with improved selectivity and safety.Author contributions
Zihan Yu: writing – original draft, writing – review & editing, visualization, validation, supervision, software, project administration, methodology, investigation, formal analysis, data curation, conceptualization. Bingkun Chen: writing – review & editing, software, methodology, formal analysis, data curation. Pengfei Song: writing – review & editing, software, methodology, formal analysis. Zhuo Zhang: writing – review & editing, resources, methodology, conceptualization. Yuanjun Zhu: writing – review & editing, visualization. Chao Ma: writing – review & editing, supervision, resources, methodology, data curation, conceptualization.Conflicts of interest
The authors declare no financial or personal relationships with any individuals or entities that could inappropriately influence (bias) this work. There are no professional or personal interests of any nature or kind, direct or indirect, in products, services, or companies that might affect the objectivity or interpretation of the research.Data availability
The data supporting this article are included in the supplementary information (SI), which encompass molecular docking scores, MD simulation-derived parameters and diagrams (RMSD, RMSF, ligand properties, and histograms), input files for ABFE/RBFE calculations, and original documents related to DFT calculations. Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra08979d.Acknowledgements
This research was funded by the National Natural Science Foundation of China (22477086) and the General Scientific Research Project of the Education Department of Liaoning Province, China (JYTMS20231359). The authors gratefully acknowledge Prof. Chao Ma's research group for their technical guidance and valuable discussions during the experimental design phase. We also extend our appreciation for their provision of critical computational resources and analytical tools, which significantly contributed to the completion of this study. Zihan Yu would like to extend special thanks to his family for providing him with confidence and unwavering courage.References
- F. T. Zindo, S. F. Malan, S. I. Omoruyi, A. B. Enogieru, O. E. Ekpo and J. Joubert, Design, synthesis and evaluation of pentacycloundecane and hexacycloundecane propargylamine derivatives as multifunctional neuroprotective agents, Eur. J. Med. Chem., 2019, 163, 83–94, DOI:10.1016/j.ejmech.2018.11.051.
- C. Zhang, X. Yang, D. Wan, Q. Ma, P. Yin, M. Zhou and J. H. J. Med, Burden of neurological disorders in China and its provinces, 1990–2021: Findings from the global burden of disease study 2021, Med, 2025, 6(8), 100692, DOI:10.1016/j.medj.2025.100692.
- V. Vyklicky, C. Stanley, C. Habrian and E. Y. Isacoff, Conformational rearrangement of the NMDA receptor amino-terminal domain during activation and allosteric modulation, Nat. Commun., 2021, 12(1), 2694, DOI:10.1038/s41467-021-23024-z.
- E. Karakas and H. Furukawa, Crystal structure of a heterotetrameric NMDA receptor ion channel, Science, 2014, 344(6187), 992–997, DOI:10.1126/science.1251915.
- T. H. Chou, M. Epstein, R. G. Fritzemeier, N. S. Akins, S. Paladugu, E. Z. Ullman, D. C. Liotta, S. F. Traynelis and H. Furukawa, Molecular mechanism of ligand gating and opening of NMDA receptor, Nature, 2024, 632(8023), 209–217, DOI:10.1038/s41586-024-07742-0.
- A. Verkhratsky and A. Chvatal, NMDA Receptors in Astrocytes, Neurochem. Res., 2020, 45(1), 122–133, DOI:10.1007/s11064-019-02750-3.
- K. Zhang, M. Wen, X. Nan, S. Zhao, H. Li, Y. Ai and H. Zhu, NMDA receptors in neurodegenerative diseases: mechanisms and emerging therapeutic strategies, Front. Aging Neurosci., 2025, 17, 1604378, DOI:10.3389/fnagi.2025.1604378.
- M. R. Wilcox, A. Nigam, N. G. Glasgow, C. Narangoda, M. B. Phillips, D. S. Patel, S. Mesbahi-Vasey, A. L. Turcu, S. Vázquez, M. G. Kurnikova and J. W. Johnson, Inhibition of NMDA receptors through a membrane-to-channel path, Nat. Commun., 2022, 13(1), 4114, DOI:10.1038/s41467-022-31817-z.
- J. P. Dupuis, O. Nicole and L. Groc, NMDA receptor functions in health and disease Old actor, new dimensions, Neuron, 2023, 111(15), 2312–2328, DOI:10.1016/j.neuron.2023.05.002.
- K. M. Grochowska, P. A. Yuanxiang, J. Bär, R. Raman, G. Brugal, G. Sahu, M. Schweizer, A. Bikbaev, S. Schilling, H. U. Demuth and M. R. Kreutz, Posttranslational modification impact on the mechanism by which amyloid-β induces synaptic dysfunction, EMBO Rep., 2017, 18(6), 962–981, DOI:10.15252/embr.201643519.
- Q. J. Wu and M. Tymianski, Targeting NMDA receptors in stroke: new hope in neuroprotection, Mol. Brain, 2018, 11(1), 15, DOI:10.1186/s13041-018-0357-8.
- Y. Ge, W. L. Chen, P. Axerio-Cilies and Y. T. Wang, NMDARs in Cell Survival and Death: Implications in Stroke Pathogenesis and Treatment, Trends Mol. Med., 2020, 26(6), 533–551, DOI:10.1016/j.molmed.2020.03.001.
- B.-C. Tang, Y.-T. Wang and J. Ren, Basic information about memantine and its treatment of Alzheimer's disease and other clinical applications, Ibrain, 2023, 9(3), 340–348, DOI:10.1002/ibra.12098.
- S. J. Zhu and P. Paoletti, Allosteric modulators of NMDA receptors: multiple sites and mechanisms, Curr. Opin. Pharmacol., 2015, 20, 14–23, DOI:10.1016/j.coph.2014.10.009.
- N. Tajima, E. Karakas, T. Grant, N. Simorowski, R. Diaz-Avalos, N. Grigorieff and H. Furukawa, Activation of NMDA receptors and the mechanism of inhibition by ifenprodil, Nature, 2016, 534(7605), 63, DOI:10.1038/nature17679.
- J. E. Hanson, H. Yuan, R. E. Perszyk, T. G. Banke, H. Xing, M. C. Tsai, F. S. Menniti and S. F. Traynelis, Therapeutic potential of N-methyl-D-aspartate receptor modulators in psychiatry, Neuropsychopharmacology, 2024, 49(1), 51–66, DOI:10.1038/s41386-023-01614-3.
- D. Stroebel, D. L. Buhl, J. D. Knafels, P. K. Chanda, M. Green, S. Sciabola, L. Mony, P. Paoletti and J. Pandit, A Novel Binding Mode Reveals Two Distinct Classes of NMDA Receptor GluN2B-selective Antagonists, Mol. Pharmacol., 2016, 89(5), 541–551, DOI:10.1124/mol.115.103036.
- R. A. Friesner, J. L. Banks, R. B. Murphy, T. A. Halgren, J. J. Klicic, D. T. Mainz, M. P. Repasky, E. H. Knoll, M. Shelley, J. K. Perry, D. E. Shaw, P. Francis and P. S. Shenkin, Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy, J. Med. Chem., 2004, 47(7), 1739–1749, DOI:10.1021/jm0306430.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX, 2015, 1–2, 19–25, DOI:10.1016/j.softx.2015.06.001.
- R. Salomon-Ferrer, D. A. Case, D. R. Roe and R. C. Walker, Amber 2024, University of California, San Francisco: San Francisco, California, 2024 Search PubMed.
- BIOVIA, BIOVIA Discovery Studio, Version 4.5, Dassault Systèmes, San Diego, CA, USA, 2021 Search PubMed.
- PerkinElmer, ChemDraw Professional, Version 20.0, PerkinElmer Informatics, Waltham, MA, USA, 2015 Search PubMed.
- L. Schrödinger, The PyMOL Molecular Graphics System, Version 2.0, 2015 Search PubMed.
- W. Humphrey, A. Dalke and K. Schulten, VMD: visual molecular dynamics, J. Mol. Graphics, 1996, 14(1), 27–28, DOI:10.1016/0263-7855(96)00018-5.
- R Core Team, R: A Language and Environment for Statistical Computing, 2014, vol. 1 Search PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16 Rev. C.01, Wallingford, CT, 2016 Search PubMed.
- F. Neese, The ORCA program system, WIREs Comput. Mol. Sci., 2012, 2(1), 73–78, DOI:10.1002/wcms.81.
- T. Lu, A comprehensive electron wavefunction analysis toolbox for chemists, Multiwfn, J. Chem. Phys., 2024, 161(8), 082503, DOI:10.1063/5.0216272.
- T. Lu and F. W. Chen, Multiwfn: A multifunctional wavefunction analyzer, J. Comput. Chem., 2012, 33(5), 580–592, DOI:10.1002/jcc.22885.
- C. Lu, C. Wu, D. Ghoreishi, W. Chen, L. Wang, W. Damm, G. A. Ross, M. K. Dahlgren, E. Russell, C. D. Von Bargen, R. Abel, R. A. Friesner and E. D. Harder, OPLS4: Improving Force Field Accuracy on Challenging Regimes of Chemical Space, J. Chem. Theory Comput., 2021, 17(7), 4291–4300, DOI:10.1021/acs.jctc.1c00302.
- D. Stroebel, D. L. Buhl, J. D. Knafels, P. K. Chanda, M. Green, S. Sciabola, L. Mony, P. Paoletti and J. Pandit, A Novel Binding Mode Reveals Two Distinct Classes of NMDA Receptor GluN2B-selective Antagonists, Mol. Pharmacol., 2016, 89(5), 541–551, DOI:10.1124/mol.115.103036.
- R. A. Friesner, R. B. Murphy, M. P. Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin and D. T. Mainz, Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes, J. Med. Chem., 2006, 49(21), 6177–6196, DOI:10.1021/jm051256o.
- T. Liu, Y. Lin, X. Wen, R. N. Jorissen and M. K. Gilson, BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities, Nucleic Acids Res., 2007, 35, D198–D201, DOI:10.1093/nar/gkl999.
- J. Sunseri and D. R. Koes, Pharmit: interactive exploration of chemical space, Nucleic Acids Res., 2016, 44(W1), W442–W448, DOI:10.1093/nar/gkw287.
- M. A. Lomize, I. D. Pogozheva, H. Joo, H. I. Mosberg and A. L. Lomize, OPM database and PPM web server: resources for positioning of proteins in membranes, Nucleic Acids Res., 2011, 40(D1), D370–D376, DOI:10.1093/nar/gkr703.
- D. E. Shaw, M. M. Deneroff, R. O. Dror, J. S. Kuskin, R. H. Larson, J. K. Salmon, C. Young, B. Batson, K. J. Bowers, J. C. Chao, M. P. Eastwood, J. Gagliardo, J. P. Grossman, C. R. Ho, D. J. Ierardi, I. Kolossváry, J. L. Klepeis, T. Layman, C. McLeavey, M. A. Moraes, R. Mueller, E. C. Priest, Y. B. Shan, J. Spengler, M. Theobald, B. Towles and S. C. Wang, Anton, a special-purpose machine for molecular dynamics simulation, Commun. ACM, 2008, 51(7), 91–97, DOI:10.1145/1364782.1364802.
- K. Bowers, E. Chow, H. Xu, R. Dror, M. Eastwood, B. Gregersen, J. Klepeis, I. Kolossváry, M. Moraes, F. Sacerdoti, J. Salmon, Y. Shan and D. Shaw, Molecular dynamics⋯Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters, 2006, p. 84 Search PubMed.
- A. T. McNutt, F. Bisiriyu, S. Song, A. Vyas, G. R. Hutchison and D. R. Koes, Conformer Generation for Structure-Based Drug Design: How Many and How Good?, J. Chem. Inf. Model., 2023, 63(21), 6598–6607, DOI:10.1021/acs.jcim.3c01245.
- S. L. Dixon, A. M. Smondyrev, E. H. Knoll, S. N. Rao, D. E. Shaw and R. A. Friesner, PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results, J. Comput.-Aided Mol. Des., 2006, 20(10), 647–671, DOI:10.1007/s10822-006-9087-6.
- M. S. Valdés-Tresanco, M. E. Valdés-Tresanco, P. A. Valiente and E. Moreno, gmx_MMPBSA: A New Tool to Perform End-State Free Energy Calculations with GROMACS, J. Chem. Theory Comput., 2021, 17(10), 6281–6291, DOI:10.1021/acs.jctc.1c00645.
- L. Duan, X. Liu and J. Z. Zhang, Interaction Entropy: A New Paradigm for Highly Efficient and Reliable Computation of Protein-Ligand Binding Free Energy, J. Am. Chem. Soc., 2016, 138(17), 5722–5728, DOI:10.1021/jacs.6b02682.
- S. Jo, T. Kim, V. G. Iyer and W. Im, CHARMM-GUI: a web-based graphical user interface for CHARMM, J. Comput. Chem., 2008, 29(11), 1859–1865, DOI:10.1002/jcc.20945.
- J. Huang, S. Rauscher, G. Nawrocki, T. Ran, M. Feig, B. L. de Groot, H. Grubmüller and A. D. MacKerell, CHARMM36m: an improved force field for folded and intrinsically disordered proteins, Nat. Methods, 2017, 14(1), 71–73, DOI:10.1038/nmeth.4067.
- B. Hess, P-LINCS: A Parallel Linear Constraint Solver for Molecular Simulation, J. Chem. Theory Comput., 2008, 4(1), 116–122, DOI:10.1021/ct700200b.
- M. Oubahmane, I. Hdoufane, C. Delaite, A. Sayede, D. Cherqaoui and A. El Allali, Design of Potent Inhibitors Targeting the Main Protease of SARS-CoV-2 Using QSAR Modeling, Molecular Docking, and Molecular Dynamics Simulations, Pharmaceuticals, 2023, 16(4), 608, DOI:10.3390/ph16040608.
- S. Nosé, A Unified Formulation of the Constant Temperature Molecular Dynamics Methods, J. Chem. Phys., 1984, 81(1), 511–519, DOI:10.1063/1.447334.
- M. Parrinello and A. Rahman, Polymorphic transitions in single crystals: A new molecular dynamics method, J. Appl. Phys., 1981, 52, 7182–7190, DOI:10.1063/1.328693.
- M. Aldeghi, M. J. Bodkin, S. Knapp and P. C. Biggin, Statistical Analysis on the Performance of Molecular Mechanics Poisson–Boltzmann Surface Area versus Absolute Binding Free Energy Calculations: Bromodomains as a Case Study, J. Chem. Inf. Model., 2017, 57(9), 2203–2221, DOI:10.1021/acs.jcim.7b00347.
- X. Liu, L. Peng, Y. Zhou, Y. Zhang and J. Z. H. Zhang, Computational Alanine Scanning with Interaction Entropy for Protein-Ligand Binding Free Energies, J. Chem. Theory Comput., 2018, 14(3), 1772–1780, DOI:10.1021/acs.jctc.7b01295.
- P. Lagarias, K. Barkan, E. Tzortzini, M. Stampelou, E. Vrontaki, G. Ladds and A. Kolocouris, Insights to the Binding of a Selective Adenosine A(3) Receptor Antagonist Using Molecular Dynamic Simulations, MM-PBSA and MM-GBSA Free Energy Calculations, and Mutagenesis, J. Chem. Inf. Model., 2019, 59(12), 5183–5197, DOI:10.1021/acs.jcim.9b00751.
- G. Heinzelmann and M. K. Gilson, Automation of absolute protein-ligand binding free energy calculations for docking refinement and compound evaluation, Sci. Rep., 2021, 11(1), 1116, DOI:10.1038/s41598-020-80769-1.
- L. Wang, Y. Wu, Y. Deng, B. Kim, L. Pierce, G. Krilov, D. Lupyan, S. Robinson, M. K. Dahlgren, J. Greenwood, D. L. Romero, C. Masse, J. L. Knight, T. Steinbrecher, T. Beuming, W. Damm, E. Harder, W. Sherman, M. Brewer, R. Wester, M. Murcko, L. Frye, R. Farid, T. Lin, D. L. Mobley, W. L. Jorgensen, B. J. Berne, R. A. Friesner and R. Abel, Accurate and reliable prediction of relative ligand binding potency in prospective drug discovery by way of a modern free-energy calculation protocol and force field, J. Am. Chem. Soc., 2015, 137(7), 2695–2703, DOI:10.1021/ja512751q.
- G. Heinzelmann, D. J. Huggins and M. K. Gilson, BAT2: an Open-Source Tool for Flexible, Automated, and Low Cost Absolute Binding Free Energy Calculations, J. Chem. Theory Comput., 2024, 20(15), 6518–6530, DOI:10.1021/acs.jctc.4c00205.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB, J. Chem. Theory Comput., 2015, 11(8), 3696–3713, DOI:10.1021/acs.jctc.5b00255.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, Development and testing of a general amber force field, J. Comput. Chem., 2004, 25(9), 1157–1174, DOI:10.1002/jcc.20035.
- T. Darden, D. York and L. Pedersen, Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems, J. Chem. Phys., 1993, 98(12), 10089–10092, DOI:10.1063/1.464397.
- J. Åqvist, P. Wennerström, M. Nervall, S. Bjelic and B. O. Brandsdal, Molecular dynamics simulations of water and biomolecules with a Monte Carlo constant pressure algorithm, Chem. Phys. Lett., 2004, 384(4), 288–294, DOI:10.1016/j.cplett.2003.12.039.
- C. D. Christ, A. E. Mark and W. F. van Gunsteren, Basic ingredients of free energy calculations: a review, J. Comput. Chem., 2010, 31(8), 1569–1582, DOI:10.1002/jcc.21450.
- J. D. Chodera, W. C. Swope, J. W. Pitera, C. Seok and K. A. Dill, Use of the Weighted Histogram Analysis Method for the Analysis of Simulated and Parallel Tempering Simulations, J. Chem. Theory Comput., 2007, 3(1), 26–41, DOI:10.1021/ct0502864.
- S. J. Marrink, H. J. Risselada, S. Yefimov, D. P. Tieleman and A. H. de Vries, The MARTINI force field: coarse grained model for biomolecular simulations, J. Phys. Chem. B, 2007, 111(27), 7812–7824, DOI:10.1021/jp071097f.
- W. Kohn and L. J. Sham, Self-Consistent Equations Including Exchange and Correlation Effects, Phys. Rev., 1965, 140(4A), A1133–A1138, DOI:10.1103/PhysRev.140.A1133.
- B. Chen, B. Hu, Y. Zong, Y. Wang, L. Jing, Z. Yu, H. Xue, M. Hou and X. Jia, Spatial and electronic features driving SGLT1/2 selectivity: a combined molecular dynamics and quantum mechanics study, Phys. Chem. Chem. Phys., 2025, 27(36), 18978–18996, 10.1039/d5cp00708a.
- F. Weigend and R. Ahlrichs, Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy, Phys. Chem. Chem. Phys., 2005, 7(18), 3297–3305, 10.1039/b508541a.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, Universal Solvation Model Based on Solute Electron Density and on a Continuum Model of the Solvent Defined by the Bulk Dielectric Constant and Atomic Surface Tensions, J. Phys. Chem. B, 2009, 113(18), 6378–6396, DOI:10.1021/jp810292n.
- A. J. Misquitta and K. Szalewicz, Symmetry-adapted perturbation-theory calculations of intermolecular forces employing density-functional description of monomers, J. Chem. Phys., 2005, 122(21), 214109, DOI:10.1063/1.1924593.
- P. J. Eddershaw, A. P. Beresford and M. K. Bayliss, ADME/PK as part of a rational approach to drug discovery, Drug Discovery Today, 2000, 5(9), 409–414, DOI:10.1016/s1359-6446(00)01540-3.
- A. Daina, O. Michielin and V. Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Sci. Rep., 2017, 7, 42717, DOI:10.1038/srep42717.
- G. Xiong, Z. Wu, J. Yi, L. Fu, Z. Yang, C. Hsieh, M. Yin, X. Zeng, C. Wu, A. Lu, X. Chen, T. Hou and D. Cao, ADMETlab 2.0: an integrated online platform for accurate and comprehensive predictions of ADMET properties, Nucleic Acids Res., 2021, 49(W1), W5–w14, DOI:10.1093/nar/gkab255.
- D. T. Monaghan, M. W. Irvine, B. M. Costa, G. Fang and D. E. Jane, Pharmacological modulation of NMDA receptor activity and the advent of negative and positive allosteric modulators, Neurochem. Int., 2012, 61(4), 581–592, DOI:10.1016/j.neuint.2012.01.004.
- I. Borza, S. Kolok, A. Gere, J. Nagy, L. Fodor, K. Galgóczy, J. Fetter, F. Bertha, B. Agai, C. Horváth, S. Farkas and G. Domány, Benzimidazole-2-carboxamides as novel NR2B selective NMDA receptor antagonists, Bioorg. Med. Chem. Lett., 2006, 16(17), 4638–4640, DOI:10.1016/j.bmcl.2006.06.002.
- L. R. Marcin, J. Warrier, S. Thangathirupathy, J. Shi, G. N. Karageorge, B. C. Pearce, A. Ng, H. Park, J. Kempson, J. Li, H. Zhang, A. Mathur, A. B. Reddy, G. Nagaraju, G. Tonukunuru, G. Gupta, M. Kamble, R. Mannoori, S. Cheruku, S. Jogi, J. Gulia, T. Bastia, C. Sanmathi, J. Aher, R. Kallem, B. N. Srikumar, K. K. Vijaya, P. S. Naidu, M. Paschapur, N. Kalidindi, R. Vikramadithyan, M. Ramarao, R. Denton, T. Molski, E. Shields, M. Subramanian, X. Zhuo, M. Nophsker, J. Simmermacher, M. Sinz, C. Albright, L. J. Bristow, I. Islam, J. J. Bronson, R. E. Olson, D. King, L. A. Thompson and J. E. Macor, BMS-986163, a Negative Allosteric Modulator of GluN2B with Potential Utility in Major Depressive Disorder, ACS Med. Chem. Lett., 2018, 9(5), 472–477, DOI:10.1021/acsmedchemlett.8b00080.
- G. Barta-Szalai, I. Borza, E. Bozó, C. Kiss, B. Agai, A. Proszenyák, G. M. Keseru, A. Gere, S. Kolok, K. Galgóczy, C. Horváth, S. Farkas and G. Domány, Oxamides as novel NR2B selective NMDA receptor antagonists, Bioorg. Med. Chem. Lett., 2004, 14(15), 3953–3956, DOI:10.1016/j.bmcl.2004.05.053.
- S. T. Wilkinson and G. Sanacora, A new generation of antidepressants: an update on the pharmaceutical pipeline for novel and rapid-acting therapeutics in mood disorders based on glutamate/GABA neurotransmitter systems, Drug Discovery Today, 2019, 24(2), 606–615, DOI:10.1016/j.drudis.2018.11.007.
- L. Zhang, T. Bu, X. Bao, T. Liang, Y. Ge, Y. Xu and Q. Zhu, Design, synthesis and biological evaluation of novel 3H-imidazole [4,5-b] pyridine derivatives as selective mTOR inhibitors, Bioorg. Med. Chem. Lett., 2017, 27(15), 3395–3398, DOI:10.1016/j.bmcl.2017.06.010.
- C. F. Claiborne, J. A. McCauley, B. E. Libby, N. R. Curtis, H. J. Diggle, J. J. Kulagowski, S. R. Michelson, K. D. Anderson, D. A. Claremon, R. M. Freidinger, R. A. Bednar, S. D. Mosser, S. L. Gaul, T. M. Connolly, C. L. Condra, B. Bednar, G. L. Stump, J. J. Lynch, A. Macaulay, K. A. Wafford, K. S. Koblan and N. J. Liverton, Orally efficacious NR2B-selective NMDA receptor antagonists, Bioorg. Med. Chem. Lett., 2003, 13(4), 697–700, DOI:10.1016/s0960-894x(02)01061-2.
- M. M. Ghahremanpour, A. Saar, J. Tirado-Rives and W. L. Jorgensen, Computation of Absolute Binding Free Energies for Noncovalent Inhibitors with SARS-CoV-2 Main Protease, J. Chem. Inf. Model., 2023, 63(16), 5309–5318, DOI:10.1021/acs.jcim.3c00874.
- W. L. Jorgensen, The many roles of computation in drug discovery, Science, 2004, 303(5665), 1813–1818, DOI:10.1126/science.1096361.
- G. Scapin, Structural biology in drug design: selective protein kinase inhibitors, Drug Discovery Today, 2002, 7(11), 601–611, DOI:10.1016/S1359-6446(02)02290-0.
| This journal is © The Royal Society of Chemistry 2026 |












