Open Access Article
Open Access ArticleDirected C–H activation of 13α-estrone: a pathway to promising AKR1C inhibitors via docking and biological studies
Erzsébet Mernyák *a,
Marija Gjorgoškab,
Masa Sinreihb,
Aljaž Kotnikb,
Zala Zanoškarb,
Ajda Godecb,
Marko Jukičcd,
Urban Brencde,
Rebeka Ignácza,
Kornél Szőria,
Zoltán Kelef,
Attila Hunyadi
*a,
Marija Gjorgoškab,
Masa Sinreihb,
Aljaž Kotnikb,
Zala Zanoškarb,
Ajda Godecb,
Marko Jukičcd,
Urban Brencde,
Rebeka Ignácza,
Kornél Szőria,
Zoltán Kelef,
Attila Hunyadi ag and
Tea Lanišnik Rižner*b
ag and
Tea Lanišnik Rižner*b
aInstitute of Pharmacognosy, University of Szeged, Eötvös u. 6, H-6720 Szeged, Hungary. E-mail: mernyak.erzsebet@szte.hu
bInstitute of Biochemistry and Molecular Genetics, Faculty of Medicine, University of Ljubljana, Vrazov trg 2, 1000 Ljubljana, Slovenia. E-mail: tea.lanisnik-rizner@mf.uni-lj.si
cLaboratory of Physical Chemistry and Chemical Thermodynamics, Faculty of Chemistry and Chemical Engineering, University of Maribor, Smetanova ulica 17, SI-2000 Maribor, Slovenia
dFaculty of Mathematics, Natural Sciences and Information Technologies, University of Primorska, Glagoljaška ulica 8, SI-6000 Koper, Slovenia
eInstitute of Environmental Protection and Sensors, Beloruska ulica 7, SI-2000 Maribor, Slovenia
fDepartment of Medicinal Chemistry, University of Szeged, Dóm tér 8, H-6720 Szeged, Hungary
gHUN-REN-SZTE Biologically Active Natural Products Research Group, Eötvös u. 6, H-6720 Szeged, Hungary
First published on 27th February 2026
Abstract
The aldo-keto reductase isoenzymes AKR1C1–3 regulate local steroid hormone availability through the interconversion of active and inactive ligands, thereby modulating prereceptor signaling. This regulatory mechanism has been implicated in the progression of hormone-dependent malignancies, highlighting AKR1C enzymes as attractive therapeutic targets for endocrine-related cancers. The AKR1C family is also known to mediate resistance to multiple classes of chemotherapeutic agents through various mechanisms. Inhibition of AKR1C enzymes may therefore potentiate the cytotoxic effects of chemotherapeutic agents. Building on our recent work describing potent A-ring halogenated 13α-estrone-based AKR1C inhibitors, we now report further structural modifications via directed C–H activation on the same scaffold. Following the introduction of a directing group, hydroxylation or acetoxylation was performed at the C-2 position. The newly synthesized compounds were evaluated against recombinant AKR1C1–3 enzymes. Notably, two new derivatives (4 and 6a) exhibited low micromolar, isoform-selective inhibitory activity against AKR1C2. Moreover, using in silico molecular docking, we postulated the binding conformations of active pyridyloxy derivative (6a), triazinyl derivative (7) and aryl carbamate (4) within the AKR1C2 binding site, with all of them showing key interactions with Trp86, Val128, Ile129 and Trp227. The AKR1C2 inhibitors identified in this study represent promising starting points for the development of novel therapeutic agents, limiting metastatic dissemination, particularly in certain aggressive tumor types. Given that AKR1C1–3 isoenzymes often catalyze overlapping biochemical transformations, inhibition of one member may be compensated by another. Thus, while selective AKR1C inhibitors remain valuable, the development of pan-inhibitors also represents a promising therapeutic strategy.
1. Introduction
Human AKR1C1–4 enzymes are cytosolic enzymes broadly expressed across tissues, but most abundant in the liver, lung, kidney, and steroid hormone–responsive tissues such as the breast, uterus, prostate, and ovary, where they play a key role in steroid hormone metabolism and detoxification of aldehydes and xenobiotics.1–3 AKR1C enzymes have high amino-acid sequence similarity, especially AKR1C1 and AKR1C2, which differ only by seven amino-acid residues. These enzymes use nicotinamide adenine dinucleotides as cofactors. AKR1C3, otherwise known as 17β-hydroxysteroid dehydrogenase type 5 (17β-HSD5) plays the main role in all pathways leading to potent androgens,4–6 and is also involved in the stereospecific reduction of estrone to 17β-estradiol (E2).3 AKR1C1 and AKR1C2 act as 3-ketosteroid reductases, but with different stereoselectivity (3β- or 3α-diol is formed, respectively). Nevertheless, the ability of these enzymes to catalyze reduction of ketones is not limited to the steroidal compounds. AKR1C enzymes act on other substrates, including prostaglandins, isoprenoids and retinoids.7,8 Additionally, the AKR1C enzymes act as phase I metabolism enzymes and metabolize different xenobiotics.7The AKR1C family is also known to mediate resistance to multiple classes of chemotherapeutic agents through various mechanisms.9 In the first mechanism, inactivation of chemotherapeutic agents occurs by the reduction of their carbonyl to hydroxyl groups. The second mechanism involves the reduction of certain quinones to hydroquinones with the simultaneous production of ROS, leading to elevated AKR expression by a feed-forward mechanism. The third mechanism is characterized by AKR upregulation, which results in elimination of cellular stress induced by drug exposure. Specifically, AKR1C3 catalyzes the inactivation of certain chemotherapeutic agents, including doxorubicin, daunorubicin and idarubicin.10,11 Additionally, overexpression of AKR1C1–AKR1C3 is proposed to be responsible for the resistance to platinum based drugs12 and AKR1C2 plays an important role in promoting metastasis.13,14
We recently reported the development of AKR1C1–3 inhibitors based on a 13α-estrone (1, Fig. 1) scaffold.15 The core-modified 13-epimer of natural estrone, characterized by its low affinity for nuclear estrogen receptors, provides a promising framework for the design of selective estrone-based enzyme inhibitors.16,17 The most active A-ring–halogenated derivatives (2 and 3) exhibited low or submicromolar IC50 values against one or more AKR1C isoforms (Fig. 1). It is hypothesized that both the size and electronegativity of the halogen atoms influence inhibitory potency and selectivity.
Given that electron-withdrawing halogens on the A-ring generally enhanced activity, the incorporation of oxygen-containing groups may also prove beneficial. Further exploration of modifications at the C-3 hydroxy group is encouraged, as these may impact AKR1C inhibitory activity.
The introduction of oxygen-containing substituents onto phenolic aromatic rings can be effectively accomplished via a directed C–H activation-substitution strategy.18 Several nitrogen-containing DGs have been reported in the literature that enable regioselective substitution not only at the ortho but also at the meta position.18 Literature sources identify carbamoyl, pyridyl, and triazinyl DGs as effective moieties for facilitating directed C–H activations on small-molecular phenolic substrates. Introduction of such nitrogen-containing groups into biologically active molecules may provide advantages from both chemical and biological perspectives. Brožič et al. have described a series of moderate to potent AKR1C inhibitors bearing carbamoyl or pyridyl groups.19 Additionally, Savić et al. reported potent androstane-based AKR1C3 inhibitors, wherein fusion of a pyridine ring to the A-ring resulted in potency comparable to that of ibuprofen.20
Yang et al. demonstrated that the introduction of a carbamoyl DG enables ruthenium(II)-catalyzed ortho C–H hydroxylation of phenolic substrates.21 This methodology was later adapted by Ba et al. for the regioselective hydroxylation of estradiol at C2.22 Regioselective ortho-acetoxylation of 2-aryloxypyridines using PhI(OAc)2 as the oxidant and the acetate source might be achieved via palladium catalysis, following the introduction of a 2-pyridyl DG.23 The facile removal of both the acetyl and pyridyl moieties enables the synthesis of structurally diverse di- or polyhydroxylated phenol derivatives. 1,2-Dihydroxybenzene derivatives, also known as catechols represent a frequent structural motif present in numerous natural products and pharmaceutical agents.24 Peng et al. employed the 4,6-dimethoxy-1,3,5-triazin-2-yloxy group as a modifiable and readily cleavable DG to achieve ortho C–H acetoxylation of phenols.25
Tetralone and its derivatives are widely utilized scaffolds in medicinal chemistry due to their broad range of biological activities.26 Natural and semi-synthetic tetralone derivatives have been increasingly investigated for their anticancer, antibacterial, anti-inflammatory, and antiviral properties. Certain nonsteroidal inhibitors of steroidogenic enzymes are derived from structural motifs that mimic key substructures of endogenous steroidal compounds. Notably, 6-hydroxy-1-tetralone, a commercially available compound, has recently been employed in the development of Vepdegestrant (ARV-471, Arvinas, Phase 2) a first-in-class, orally bioavailable proteolysis-targeting chimera (PROTAC) estrogen receptor degrader, intended for the treatment of endocrine-resistant breast cancer.27
Based on these considerations, the present study aimed to develop novel A-ring modified 13α-estrone derivatives as potential AKR1C inhibitors. Following the introduction of nitrogen-containing DGs, catalytic and regioselective ortho-hydroxylation or acetoxylation reactions were planned. Additionally, the 6-hydroxy-1-tetralone scaffold was involved into selected structural modifications. A comparative evaluation of the inhibitory activity of the newly synthesized compounds against the AKR1C1–3 isoforms was also designed. To gain insight into the interaction patterns of the synthesized A-ring modified 13α-estrone derivatives, structure-based molecular docking experiments were planned. In silico studies were designed to identify key protein-ligand interactions, and provide a structural rationale for the observed activity.
2. Materials and methods
2.1. Chemistry
Microwave-assisted syntheses were performed in the Anton-Paar Monowave 400 reactor. Melting points (Mp) were determined with a Kofler hot-stage apparatus and are uncorrected. Thin-layer chromatography was performed on silica gel 60 F254; (layer thickness 0.2 mm, (Merck KGaA, Darmstadt, Germany)); eluent: a: 30% ethyl acetate/70% hexanes, b: 50% ethyl acetate/70% hexanes, c: 20% ethyl acetate/80% hexanes. The spots were detected with I2 or UV (365 nm) after spraying with 5% phosphomolybdic acid in 50% aqueous phosphoric acid and heating at 100–120 °C for 10 min. Flash chromatography was performed on silica gel 60, 40–63 µm (Merck KGaA, Darmstadt, Germany). 1H NMR spectra were recorded in DMSO-d6 or CDCl3 solution with a Bruker DRX-500 instrument (Bruker BioSpin GmbH, Rheinstetten, Germany) at 500 MHz. 13C NMR spectra were recorded with the same instrument at 125 MHz under the same conditions. Mass spectrometry: full scan mass spectra of the newly synthesized compounds were acquired in the range of 100 to 1100 m/z with a Q Exactive Plus quadrupole-orbitrap mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) equipped with a heated electrospray (HESI). Analyses were performed in positive or negative ion mode by flow injection mass spectrometry with a mobile phase of 50% aqueous acetonitrile containing 0.1 v/v% formic acid (0.3 mL min−1 flow rate). Aliquots of 5 µL of samples were injected into the flow. The ESI capillary was adjusted to 3.5 kV and N2 was used as a nebulizer gas. The analytical HPLC measurements were performed on a Jasco HPLC instrument (Jasco International Co., Ltd, Hachioji, Tokyo, Japan) equipped with an MD-2010 Plus PDA detector to collect data in a range of 210–410 nm. The measurements were performed on a Kinetex Biphenyl 5 µm, 250 mm 4.6 mm column (Phenomenex Inc.) equipped with a C-18 guard column using gradient elution. Mobile phase A was water (Sigma-Aldrich Ltd, Budapest, Hungary), while mobile phase B was acetonitrile (Merck Ltd, Budapest, Hungary). A linear gradient was applied from 20% B to 100% B in 30 min and finally the column was re-equilibrated for 20 min. The flow rate was set to 1 mL min−1.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Carbamate 4 was obtained as a white solid (317 mg, 93%), which is identical with the compound described in the literature.28 Mp.: 135.9–136.7 °C; Rf: 0.15a; 1H NMR (500 MHz, CDCl3) δ ppm: 1.05 (s, 3H, 13-CH3); 2.83 (m, 2H, 6-H2); 2.99 and 3.07 (2 × s, 2 × 3H, 2 × N-CH3); 6.81 (d, 1H, J = 2.5 Hz, 4-H); 6.86 (dd, 1H, J = 8.5 Hz, J = 2.5 Hz, 2-H); 7.23 (d, 1H, J = 8.5 Hz, 1-H). Purity from HPLC: 98.6%.
1 (v/v) as eluent. Carbamate 4 was obtained as a white solid (317 mg, 93%), which is identical with the compound described in the literature.28 Mp.: 135.9–136.7 °C; Rf: 0.15a; 1H NMR (500 MHz, CDCl3) δ ppm: 1.05 (s, 3H, 13-CH3); 2.83 (m, 2H, 6-H2); 2.99 and 3.07 (2 × s, 2 × 3H, 2 × N-CH3); 6.81 (d, 1H, J = 2.5 Hz, 4-H); 6.86 (dd, 1H, J = 8.5 Hz, J = 2.5 Hz, 2-H); 7.23 (d, 1H, J = 8.5 Hz, 1-H). Purity from HPLC: 98.6%.Carbamate 13 was obtained as a white solid (186 mg, 80%). Mp.: 110.1–112.7 °C; Rf: 0.34a;
1H NMR (CDCl3) δ [ppm]: 2.03 (m, 2H); 2.54 (m, 2H); 2.86 (m, 2H, 4-H2); 2.92 and 3.01 (2 × s, 2 × 3H, N(CH3)2); 6.95 (overlapping multiplets, 2H, 5-H and 7-H); 7.94 (d, 1H, J = 8.5 Hz, 8-H);. 13C NMR (CDCl3) δ [ppm]: 23.0 (C-3); 29.6 (C-4); 36.3 and 36.5 (N(CH3)2); 38.7 (C-2); 119.9 (C-7); 121.2 (C-5); 128.7 (C-8); 129.6 (C-8a); 146.0 (C-4a); 153.8 (NCO); 155.2 (C-6); 197.0 (C-1). ESI-HRMS: m/z: 234.1123 [M + H]+ (C13H15NO3 + H+ requires 234.1125).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) were added into a 10 mL microwave vessel. The mixture was stirred at 90 °C for 1 h and cooled to rt. The reaction mixture was poured onto water (10 mL) and extracted with CH2Cl2 (3 × 15 mL). The organic layer was dried over Na2SO4, concentrated in vacuo and the resulting residue was purified by column chromatography, using hexanes/EtOAc = 3
1) were added into a 10 mL microwave vessel. The mixture was stirred at 90 °C for 1 h and cooled to rt. The reaction mixture was poured onto water (10 mL) and extracted with CH2Cl2 (3 × 15 mL). The organic layer was dried over Na2SO4, concentrated in vacuo and the resulting residue was purified by column chromatography, using hexanes/EtOAc = 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 5 was obtained as a white solid (37 mg, 71%). Mp.: 149.9–150.7 °C; Rf: 0.39b.
1 (v/v) as eluent. Compound 5 was obtained as a white solid (37 mg, 71%). Mp.: 149.9–150.7 °C; Rf: 0.39b.1H NMR (DMSO-d6) δ [ppm]: 0.97 (s, 3H, 18-H3); 2.64 (m, 2H, 6-H2); 2.86 and 3.00 (2 × s, 2 × 3H, N(CH3)2); 6.62 and 6.75 (2 × s, 2 × 1H, 1-H and 4-H); 9.05 (s, 1H, OH). 13C NMR (DMSO-d6) δ [ppm]: 20.4 (C-15); 24.4 (C-18); 27.8 and 28.0: C-7 and C-11; 28.5 (C-6); 31.6 (C-12); 32.8 (C-16); 40.0 (2C, N(CH3)2); 40.5 (C-8); 40.9 (C-9); 48.5 (C-14); 49.4 (C-13); 113.6 (C-4); 122.8 (C-1); 126.9 (C-10); 136.8 (C-5); 137.0 (C-2); 146.8 (C-3); 154.0 (NCO); 220.5 (C-17). ESI-HRMS: m/z: 358.2015 [M + H]+ (C21H27NO4 + H+ requires 358.2013). Purity from HPLC: 95.5%.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 6a was obtained as a white solid (278 mg, 80%). Mp.: 130.2–131.7 °C; Rf: 0.47a.
1 (v/v) as eluent. Compound 6a was obtained as a white solid (278 mg, 80%). Mp.: 130.2–131.7 °C; Rf: 0.47a.1H NMR (DMSO-d6) δ [ppm]: 0.99 (s, 3H, 18-H3); 2.78 (m, 2H, 6-H2); 6.79 (d, 1H, J = 2.2 Hz, 4 H); 6.84 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 6.95 (d, 1H, J = 8.3 Hz, CH); 7.09 (dd, 1H, J = 7.1 Hz, J = 5.0 Hz, CH); 7.29 (d, 1H, J = 8.6 Hz, 1 H); 7.81 (dd, 1H, J = 5.0 Hz, J = 2.0 Hz, CH); 8.12 (dd, 1H, J = 5.0 Hz, J = 1.9 Hz, CH). 13C NMR (DMSO-d6) δ [ppm]: 20.4 (C-15); 24.4 (C-18); 27.4 and 27.8: C-7 and C-11; 29.4 (C-6); 31.5 (C-12); 32.8 (C-16); 40.5 (C-8); 40.9 (C-9); 48.5 (C-14); 49.4 (C-13); 111.2 (C-3′); 118.4 (C-5′); 118.7 (C-2); 120.7 (C-4); 126.9 (C-1); 135.7 (C-10); 138.2 (C-5); 139.9 (C-4′); 147.3 (C-6′); 151.5 (C-3); 163.2 (C-2′); 220.5 (C-17). ESI-HRMS: m/z: 348.1954 [M + H]+ (C23H25NO2 + H+ requires 348.1958). Purity from HPLC: 95.1%.
Compound 14 was obtained as a white solid (184 mg, 77%). Mp.: 120.1–121.0 °C; Rf: 0.46b.
1H NMR (DMSO-d6) δ [ppm]: 2.04 (m, 2H); 2.59 (m, 2H); 2.93 (m, 2H, 4-H2); 7.04–7.07 (overlapping multiplets, 2H, 2 × CH); 7.12 (d, 1H, J = 8.3 Hz, CH); 7.20 (dd, 1H, J = 7.0 Hz, J = 5.0 Hz, CH); 7.89–7.93 (overlapping multiplets, 2H, 2 × CH); 8.20 (dd, 1H, J = 5.0 Hz, J = 1.1 Hz, CH). 13C NMR (DMSO-d6) δ [ppm]: 22.7 (C-3); 28.9 (C-4); 38.3 (C-2); 112.3 (C-3′); 118.8 (C-7); 119.8 (C-5′); 119.9 (C-5); 128.4 (C-8); 128.5 (C-8a); 140.4 (C-4′); 146.9 (C-4a); 147.6 (C-6′); 158.0 (C-6); 162.0 (C-2′); 196.3 (C-1). ESI-HRMS: m/z: 240.1014 [M + H]+ (C15H13NO2 + H+ requires 240.1019).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 8 was obtained as a white solid (38.5 mg, 66%). Mp.: 190.9–191.7 °C; Rf: 0.38a.
1 (v/v) as eluent. Compound 8 was obtained as a white solid (38.5 mg, 66%). Mp.: 190.9–191.7 °C; Rf: 0.38a.1H NMR (CDCl3) δ [ppm]: 1.05 (s, 3H, 18-H3); 2.02 (s, 3H, Ac-CH3); 2.82 (m, 2H, 6-H2); 6.90 (d, 1H, J = 8.3 Hz, CH); 6.92 (s, 1H, CH); 6.97 (dd, 1H, J = 8.3 Hz, J = 5.0 Hz, CH); 7.06 (s, 1H, CH); 7.66 (ddd, 1H, J = 8.4 Hz, J = 7.3 Hz, J = 2.0 Hz, CH); 8.16 (dd, 1H, J = 5.0 Hz, J = 1.5 Hz, CH). 13C NMR (CDCl3) δ [ppm]: 20.4 (Ac-CH3); 21.0 (C-15); 25.1 (C-18); 28.0 and 28.1: C-7 and C-11; 29.6 (C-6); 32.0 (C-12); 33.4 (C-16); 40.8 (C-8); 41.6 (C-9); 49.2 (C-14); 50.1 (C-13); 110.9 (C-3′); 118.4 (C-5′); 120.9 (C-1); 122.8 (C-4); 135.7 (C-10); 137.3 (C-5); 139.4 (C-4′); 140.3 (C-3); 142.8 (C-2); 147.7 (C-6′); 163.2 (C-2′); 168.8 (Ac-CO); 221.4 (C-17). ESI-HRMS: m/z: 406.2009 [M + H]+ (C25H27NO4 + H+ requires 406.2013).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 7 was obtained as a white solid (340 mg, 83%). Mp.: 281.9–282.8 °C; Rf: 0.42a.
1 (v/v) as eluent. Compound 7 was obtained as a white solid (340 mg, 83%). Mp.: 281.9–282.8 °C; Rf: 0.42a.1H NMR (DMSO-d6) δ [ppm]: 0.99 (s, 3H, 18-H3); 2.80 (m, 2H, 6-H2); 3.90 (s, 2 × 3H, 2 × OCH3); 6.90 (d, 1H, J = 2.2 Hz, 4-H); 6.95 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 7.32 (d, 1H, J = 8.6 Hz, 1-H). 13C NMR (DMSO-d6) δ [ppm]: 20.4 (C-15); 24.4 (C-18); 27.4 and 27.8: C-7 and C-11; 29.4 (C-6); 31.5 (C-12); 32.8 (C-16); 40.3 (C-8); 40.9 (C-9); 48.4 (C-14); 49.3 (C-13); 55.1 (2 × C, 2 × OCH3); 118.6 (C-2); 120.8 (C-4); 126.8 (C-1); 137.1 (C-10); 138.2 (C-5); 149.2 (C-3); 172.7 (C-2′); 173.2 (2C, C-4′ and C-6′); 220.5 (C-17). ESI-HRMS: m/z: 410.2077 [M + H]+ (C23H27N3O4 + H+ requires 410.2074). Purity from HPLC: 97.5%.
Compound 15 was obtained as a white solid (229 mg, 76%). Mp.: 115.9–116.7 °C; Rf: 0.45b.
1H NMR (DMSO-d6) δ [ppm]: 2.06 (m, 2H); 2.60 (m, 2H); 2.96 (m, 2H, 4-H2); 3.91 (s, 2 × 3H, 2 × OCH3); 7.22 (dd, 1H, J = 8.5 Hz, J = 2.3 Hz, 7-H); 7.25 (d, 1H, J = 2.3, 5-H); 7.93 (d, 1H, J = 8.5, 8-H). 13C NMR (DMSO-d6) δ [ppm]: 22.6 (C-3); 28.8 (C-4); 38.3 (C-2); 55.3 (2C, 2 × OCH3); 120.1 (C-7); 121.3 (C-5); 128.1 (C-8); 129.9 (C-8a); 146.7 (C-4a); 154.9 (C-6); 172.2 (C-2′); 173.2 (2C, C-4′ and C-6′); 196.4 (C-1). ESI-HRMS: m/z: 302.1132 [M + H]+ (C15H15N3O4 + H+ requires 302.1135).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 9 was obtained as a white solid (39 mg, 68%). Mp.: 170.9–172.0 °C; Rf: 0.15a.
1 (v/v) as eluent. Compound 9 was obtained as a white solid (39 mg, 68%). Mp.: 170.9–172.0 °C; Rf: 0.15a.1H NMR (DMSO-d6) δ [ppm]: 0.98 (s, 3H, 18-H3); 2.09 (s, 3H, Ac-CH3); 2.80 (m, 2H, 6-H2); 3.90 (s, 2 × 3H, 2 × OCH3); 7.04 (s, 1H, CH); 7.16 (s, 1H, CH). 13C NMR (DMSO-d6) δ [ppm]: 20.2 (Ac-CH3); 20.4 (C-15); 24.4 (C-18); 27.4 and 27.7: C-7 and C-11; 28.8 (C-6); 31.4 (C-12); 32.8 (C-16); 40.0 (C-8); 40.9 (C-9); 48.4 (C-14); 49.3 (C-13); 55.3 (2 × C, 2 × OCH3); 120.8 (C-1); 122.4 (C-4); 135.4 (C-10); 138.3 (C-5); 139.5 (C-2); 140.5 (C-3); 168.2 (AcCO); 172.3 (C-2′); 173.2 (2C, C-4′ and C-6′); 220.5 (C-17). ESI-HRMS: m/z: 468.2138 [M + H]+ (C25H29N3O6 + H+ requires 468.2129).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent. Compound 11 was obtained as a white solid (12 mg, 58%). Mp.: 244.9–246.2 °C; Rf: 0.40b.
1 (v/v) as eluent. Compound 11 was obtained as a white solid (12 mg, 58%). Mp.: 244.9–246.2 °C; Rf: 0.40b.1H NMR (DMSO-d6) δ [ppm]: 0.96 (s, 3H, 18-H3); 2.58 (m, 2H, 6-H2); 6.38 and 6.60 (2 × s, 2 × 1H, 1-H and 4-H); 8.44 and 8.50 (2 × s, 2 × 1H, 2 × OH). 13C NMR (DMSO-d6) δ [ppm]: 20.4 (C-15); 24.5 (C-18); 28.0 and 28.2: C-7 and C-11; 28.8 (C-6); 31.6 (C-12); 32.8 (C-16); 40.8 (C-8); 40.9 (C-9); 48.4 (C-14); 49.3 (C-13); 112.8 (C-4); 115.3 (C-1); 126.8 (C-5); 129.8 (C-10); 143.0 (2C, C-2 and C-3); 220.6 (C-17). ESI-HRMS: m/z: 285.1480 [M + H]+ (C18H22O3 − H+ requires 285.1491).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) as eluent.
1 (v/v) as eluent.
2.1.8.1. Synthesis of 3-(3-trifluoromethyl-pyridyn-2-yloxy)-13α-estra-1,3,5(10)-triene-17-one (6b). Compound 6b was obtained as a white solid (77 mg, 78%). Mp.: 135.0–136.7 °C; Rf: 0.29c.
1H NMR (CDCl3) δ [ppm]: 1.07 (s, 3H, 18-H3); 2.86 (m, 2H, 6-H2); 6.87 (d, 1H, J = 2.0, 4-H); 6.92 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 7.05 (dd, 1H, J = 7.5 Hz, J = 5.1 Hz, 5′-H); 7.30 (d, 1H, J = 8.6 Hz, 1 H); 7.96 (dd, 1H, J = 7.5 Hz, J = 1.1 Hz) and 8.29 (dd, 1H, J = 4.8 Hz, J = 1.1 Hz): 4′-H and 6′-H. 13C NMR (CDCl3) δ [ppm]: 21.1 (C-15); 25.1 (C-18); 28.2 and 28.3: C-7 and C-11; 30.2 (C-6); 32.2 (C-12); 33.5 (C-16); 41.3 (C-8); 41.8 (C-9); 49.5 (C-14); 50.1 (C-13); 114.2 (q, 2JC–F = 33.0 Hz, C-3′); 117.5 (C-5′); 118.9 (C-2); 121.3 (C-4); 122.8 (q, 1JC–F = 273.7 Hz, CF3); 127.1 (C-1); 136.8 (C-10); 136.9 (q, 3JC–F = 4.5 Hz, C-4′); 138.5 (C-5); 150.9 (C-6′); 151.0 (C-3); 160.6 (C-2′); 221.2 (C-17). ESI-HRMS: m/z: 416.1835 [M + H]+ (C24H24F3NO2 + H+ requires 416.1832). Purity from HPLC: 98.4%.
2.1.8.2. Synthesis of 3-(4-trifluoromethyl-pyridyn-2-yloxy)-13α-estra-1,3,5(10)-triene-17-one (6c). Compound 6c was obtained as a white solid (84 mg, 81%). Mp.: 163.9–164.7 °C; Rf: 0.42c.
1H NMR (CDCl3) δ [ppm]: 1.07 (s, 3H, 18-H3); 2.86 (m, 2H, 6-H2); 6.85 (d, 1H, J = 2.0, 4-H); 6.90 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 7.12 (s, 1H, 3′-H); 7.17 (d, 1H, J = 5.2 Hz, 5′-H); 7.31 (d, 1H, J = 8.6 Hz, 1-H); 8.32 (d, 1H, J = 5.2 Hz, 6′-H). 13C NMR (CDCl3) δ [ppm]: 21.1 (C-15); 25.1 (C-18); 28.1 and 28.2: C-7 and C-11; 30.2 (C-6); 32.1 (C-12); 33.4 (C-16); 41.2 (C-8); 41.7 (C-9); 49.4 (C-14); 50.1 (C-13); 107.9 (q, 3JC–F = 3.6 Hz) and 113.7 (q, 3JC–F = 2.9 Hz): C-3′ and C-5′; 118.7 (C-2); 121.1 (C-4); 122.5 (q, 1JC–F = 273.0 Hz, CF3); 127.3 (C-1); 136.9 (C-10); 138.7 (C-5); 141.6 (q, 2JC–F = 33.9 Hz, C-4′); 149.0 (C-6′); 151.1 (C-3); 164.4 (C-2′); 221.2 (C-17). ESI-HRMS: m/z: 416.1837 [M + H]+ (C24H24F3NO2 + H+ requires 416.1832). Purity from HPLC: 97.9%.
2.1.8.3. Synthesis of 3-(5-trifluoromethyl-pyridyn-2-yloxy)-13α-estra-1,3,5(10)-triene-17-one (6d). Compound 6d was obtained as a white solid (86 mg, 83%). Mp.: 118.7–119.9 °C; Rf: 0.45c.
1H NMR (CDCl3) δ [ppm]: 1.07 (s, 3H, 18-H3); 2.86 (m, 2H, 6-H2); 6.85 (d, 1H, J = 2.0, 4 H); 6.90 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 6.97 (d, 1H, J = 8.7 Hz, 3′-H); 7.31 (d, 1H, J = 8.6 Hz, 1-H); 7.87 (dd, 1H, J = 8.7 Hz, J = 2.5 Hz, 4′-H); 8.44 (d, 1H, J = 2.5 Hz, 6′-H). 13C NMR (CDCl3) δ [ppm]: 21.0 (C-15); 25.0 (C-18); 28.1 (2C, C-7 and C-11); 30.1 (C-6); 32.1 (C-12); 33.4 (C-16); 41.2 (C-8); 41.7 (C-9); 49.3 (C-14); 50.1 (C-13); 111.2 (C-3′); 118.7 (C-2); 121.2 (C-4); 121.3 (q, 2JC–F = 32.9 Hz, C-5′); 123.7 (q, 1JC–F = 271.3 Hz, CF3); 127.3 (C-1); 136.6 (q, 3JC–F = 3.2 Hz, C-4′); 137.0 (C-10); 138.7 (C-5); 145.5 (q, 2JC–F = 4.4 Hz, C-6′); 150.9 (C-3); 166.0 (C-2′); 221.4 (C-17). ESI-HRMS: m/z: 416.1835 [M + H]+ (C24H24F3NO2 + H+ requires 416.1832). Purity from HPLC: 96.8%.
2.1.8.4. Synthesis of 3-(3-bromo-5-trifluoromethyl-pyridyn-2-yloxy)-13α-estra-1,3,5(10)-triene-17-one (6f). Compound 6f was obtained as a white solid (96 mg, 78%). Mp.: 158.6–160.0 °C; Rf: 0.53c.
1H NMR (CDCl3) δ [ppm]: 1.07 (s, 3H, 18-H3); 2.86 (m, 2H, 6-H2); 6.87 (d, 1H, J = 2.0, 4-H); 6.91 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 7.32 (d, 1H, J = 8.6 Hz, 1-H); 8.12 (d, 1H, J = 1.7 Hz) and 8.31 (d, 1H, J = 1.7 Hz): 4′-H and 6′-H. 13C NMR (CDCl3) δ [ppm]: 21.1 (C-15); 25.1 (C-18); 28.1 (2C, C-7 and C-11); 30.1 (C-6); 32.1 (C-12); 33.4 (C-16); 41.2 (C-8); 41.7 (C-9); 49.3 (C-14); 50.1 (C-13); 107.6 (C-3′); 118.8 (C-2); 121.2 (C-4); 122.4 (q, 2JC–F = 33.6 Hz, C-5′); 122.7 (q, 1JC–F = 272.0 Hz, CF3); 127.2 (C-1); 137.3 (C-10); 138.7 (C-5); 139.5 (q, 3JC–F = 3.2 Hz) and 143.5 (q, 3JC–F = 4.2 Hz): C-4′ and C-6′; 150.7 (C-3); 162.2 (C-2′); 221.4 (C-17). ESI-HRMS: m/z: 494.0945 [M + H]+ (C24H23BrF3NO2 + H+ requires 494.0937). Purity from HPLC: 99.3%.
2.1.8.5. Synthesis of 3-(6-trifluoromethyl-pyridyn-2-yloxy)-13α-estra-1,3,5(10)-triene-17-one (6e). Compound 6e was obtained as a white solid (83 mg, 80%). Mp.: 173.2–174.9 °C; Rf: 0.39c.
1H NMR (CDCl3) δ [ppm]: 1.07 (s, 3H, 18-H3); 2.84 (m, 2H, 6-H2); 6.88 (d, 1H, J = 2.0, 4-H); 6.93 (dd, 1H, J = 8.6 Hz, J = 2.0 Hz, 2-H); 6.96 (d, 1H, J = 8.7 Hz, CH); 7.28 (d, 1H, J = 8.6 Hz, 1-H); 7.36 (d, 1H, J = 7.5 Hz, CH); 7.78 (t, 1H, J = 7.9 Hz, CH). 13C NMR (CDCl3) δ [ppm]: 21.0 (C-15); 25.1 (C-18); 28.2 (2C, C-7 and C-11); 30.1 (C-6); 32.1 (C-12); 33.4 (C-16); 41.2 (C-8); 41.7 (C-9); 49.4 (C-14); 50.1 (C-13); 114.0 (C-3′); 114.7 (q, 3JC–F = 3.1 Hz, C-5′); 118.3 (C-2); 120.7 (C-4); 121.0 (d, 1JC–F = 274.5 Hz, CF3); 127.2 (C-1); 136.6 (C-10); 138.6 (C-5); 140.4 (C-4′); 146.5 (q, 2JC–F = 34.2 Hz, C-6′); 151.3 (C-3); 163.7 (C-2′); 221.5 (C-17). ESI-HRMS: m/z: 416.1840 [M + H]+ (C24H24F3NO2 + H+ requires 416.1832). Purity from HPLC: 96.0%.
2.2. In silico investigations
In a structure-based manner, we investigated the available X-ray data in RCSB PDB database firstly for AKR1C1 (UNIPROT ID: Q04828; DDH, DDH1) and examined (PDB IDs): 1MRQ, 3C3U, 3NTY, 4YVP, 6 A7A and 6IJX. 1MRQ system with the resolution of 1.59 Å chain A with co-factor NAP and STR co-crystallized ligand (progesterone) was chosen for further studies.29 Secondly, AKR1C2 (UNIPROT ID: P52895; DDH2) PDB IDs: 1IHI, 1J96, 1XJB, 2HDJ, 4JQ1, 4JQ2, 4JQ3, 4JQ4, 4JQA, 4JTQ, 4JTR, 4L1W, 4L1X, 4XO6, 4XO7 systems were collected, only to select 1J96 chain A with the lowest resolution of 1.25 Å, NAP co-factor and a co-crystallized ligand TES (testosterone30). Lastly, AKR1C3 (UNIPROT ID: P42330; DDH1, HSD17B5, KIAA0119, PGF) systems 1RY0, 1RY8, 1ZQ5, 4DBS, 4DBU, 4DBW, 4YVV, 4YVX, 4ZFC and PDB ID: 6A7B were studied and 1ZQ5 (chain A) was selected for molecular docking with an excellent resolution of 1.30 Å, NAP co-factor and a co-crystallized E04 ligand (3-carboxamido-1,3,5(10)-estratrien-17(r)-spiro-2′(5′,5′-dimethyl-6′oxo)tetrahydropyran)31 (SI, Table: target selection). All the targets were prepared using the protein preparation module of Schrödinger Small-Molecule Discovery Suite (Release Schrödinger 2024-2, Schrödinger, LLC, New York, NY, United States, 2025). Missing hydrogen atoms were added, the H-bond network was optimized using the PROPKA tool at pH 7.4, waters were removed, and restrained minimization was performed with convergence of heavy atoms toward 0.3 Å. The cofactor NADP (or modelled NADPH) were retained in the binding site. The molecular docking receptors were generated with the docking package CmDock (https://gitlab.com/Jukic/cmdock/; v. libcmdock.so/0.2.1) using Cmcavity software (v. libcmdock.so/0.2.1). The reference ligand method was used to calculate the molecular docking receptor, where we used STR, TES and E04 ligands for AKR1C1–C3 systems, respectively. A sphere of 10 Å around the ligand heavy atoms was used to calculate the docking volumes (parameters: min_volume 100, gridstep 0.5 Å).The in house developed CmDock molecular docking software (v. 0.2.1; https://gitlab.com/Jukic/cmdock; viewed on 5th January 2026) was used for molecular docking with rDOCK-SF3 scoring function, with 100 runs using DOCK.prm settings.32 The sampling and scoring performed better than comparable open-source tools33 and were verified against protein34 and RNA32,35 (SI, Table: scoring) targets. The input small molecules were precalculated by LigPrep (Release Schrödinger 2024-2, Schrödinger, LLC, New York, NY, United States, 2025) (SI, Table, compound reference).
Beforehand, a redocking experiment was performed in which we successfully re-docked reference ligands (RMSD < 1.5 Å), and then the docking experiments were performed. Potential binding energies for the hit compounds were calculated in-place using the AMBER14 force field for the solute, GAFF2 and AM1BCC for ligands using YASARA Structure TWINSET bioinformatics package (v. 25.9.17). 2D Ligand explicit interaction projections from docked/crystal poses were generated using LigPlot+ software (v. 2.3.1).
2.3. Inhibition assays for the AKR1C enzymes
Recombinant AKR1C1–3 enzymes were prepared as described previously.36 Enzymatic activity was measured spectrophotometrically at 340 nm by monitoring NAD+ reduction to NADH (ελ340 = 6220 M−1 cm−1) in the presence of the substrate 1-acenaphthenol. Assays (300 µL reaction mixtures) were carried out in 100 mM potassium phosphate buffer (pH 9.0), 0.005% Triton X-114, 0.05% DMSO, 2.3 mM NAD+, and substrate (30 µM for AKR1C1, 60 µM for AKR1C2, 100 µM for AKR1C3).37 Test compounds (5 µL in DMSO) or DMSO (control) were added, followed by a 5 min incubation at 37 °C. Background absorbance was recorded, then reactions were initiated with enzyme (0.1 µM AKR1C1, 0.3 µM AKR1C2, 1.5 µM AKR1C3 in PBS). Absorbance at 340 nm was monitored every 8 s for 3 min.Experiments were performed in duplicate and repeated independently. Data analysis involved background correction and calculation of activity from the slope of NADH formation. Residual activity was expressed relative to control, and IC50 values were derived from residual activity vs. log10 (inhibitor concentration) plots using GraphPad Prism 10.0.
3. Results and discussion
3.1. Synthesis
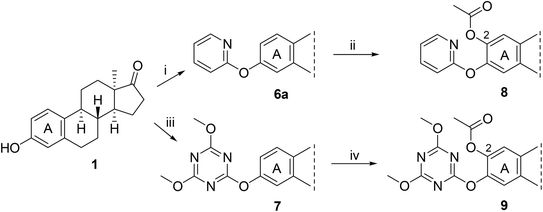
The dimethylcarbamoyl DG was first introduced onto the phenolic hydroxy group of 13α-estrone (Scheme 1). In our previous work, we described a convenient method for synthesizing aryl carbamates using sodium hydride (NaH) as a strong base.28 To avoid the use of this highly reactive and difficult-to-handle reagent, here we adopted a greener alternative, cesium carbonate (Cs2CO3), in accordance with sustainable chemistry principles.38 Due to the lower basicity of Cs2CO3, the reaction was conducted under microwave irradiation at 100 °C in toluene as the solvent, furnishing the desired compound 4 in higher yield than earlier.28 To enhance efficiency and reduce reaction times, thermally driven transformations were often performed in a microwave reactor rather than under conventional heating conditions.The ruthenium-catalyzed C–H hydroxylation of aryl carbamate (4) employed [RuCl2(p-cymene)]2 as the catalyst and K2S2O8 as the oxidant, with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of trifluoroacetic acid (TFA) and trifluoroacetic anhydride (TFAA) as the reaction medium. Microwave-assisted heating at 90 °C for 1 h furnished the desired 2-monohydroxylated product (5) in high yield and with excellent regioselectivity. By applying microwave irradiation, the reaction times reported in the literature (Ba et al., 2019)—typically requiring 24 hours—were successfully reduced to 1 h.
1 mixture of trifluoroacetic acid (TFA) and trifluoroacetic anhydride (TFAA) as the reaction medium. Microwave-assisted heating at 90 °C for 1 h furnished the desired 2-monohydroxylated product (5) in high yield and with excellent regioselectivity. By applying microwave irradiation, the reaction times reported in the literature (Ba et al., 2019)—typically requiring 24 hours—were successfully reduced to 1 h.
The present results are consistent with our previous findings on the regioselectivity of ortho-arylations on the 13α-estrone scaffold via C–H activation, which predominantly occur at the C-2 position.28 Substitution at the C-4 position was not observed, either previously or in the present study, likely due to steric hindrance imposed by the adjacent B-ring.
Subsequently, our focus turned to the introduction of two heterocyclic DGs to prepare suitable substrates for ortho-acetoxylation reactions. It was hypothesized that the structural and polarity differences between hydroxy and acetoxy groups could yield valuable insights into structure–activity relationships. Furthermore, the presence of pyridyl and triazinyl moieties at the C-3-O position may significantly influence the biological activity of the resulting compounds.
The introduction of heterocyclic DGs was carried out under distinct conditions, tailored to the substrate type (Scheme 2). The pyridyloxy derivative (6a) was obtained by O-arylation of the phenolic compound with 2-bromopyridine in the presence of a copper(I) catalyst, using potassium phosphate (K3PO4) as a base and 2-carboxypyridine as an additive. The reaction was carried out under microwave irradiation in DMSO as a solvent. The introduction of a pyridyl DG could be achieved in high yield.
The triazinyl DG was introduced by reacting 13α-estrone (1) with 2-chloro-4,6-dimethoxy-1,3,5-triazine under basic conditions, using Cs2CO3 as a base (Scheme 2). The reaction proceeded efficiently in toluene solvent, yielding the desired product (7) in good yield.
Following the introduction of the heterocyclic DGs, we proceeded with the acetoxylation reactions (Scheme 2). The acetoxylation of the (2-pyridyl)oxy derivative (6a) was investigated under both conventional heating (reflux) and microwave-assisted conditions at 90 °C. To avoid difunctionalization, only 1.5 equivalents of PhI(OAc)2 (PIDA) were employed as the acetoxy source, along with 5 mol% of Pd(OAc)2 as catalyst. As expected, the reaction rate was significantly enhanced under microwave conditions, with completion in 30 min compared to 60 min under conventional heating. Similar to the hydroxylation reactions discussed above, acetoxylation occurred selectively at the C-2 position, resulting in compound 8 in high yield.
The next transformation was the acetoxylation of the triazinyl derivative (7). Instead of conventional heating, we performed the reaction in a microwave reactor. Reaction in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (v/v) mixture of acetic acid and acetic anhydride at 100 °C for 1.5 h, using 2 equivalents of PhI(OAc)2 and 10 mol% Pd(OAc)2, afforded the acetoxylated triazine (9) in good yield.
1 (v/v) mixture of acetic acid and acetic anhydride at 100 °C for 1.5 h, using 2 equivalents of PhI(OAc)2 and 10 mol% Pd(OAc)2, afforded the acetoxylated triazine (9) in good yield.
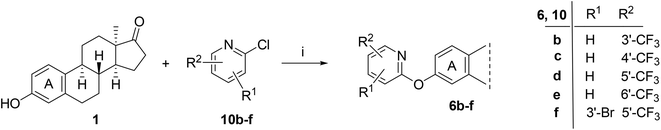
Based on the promising AKR1C2 inhibitory activity (Table 1) observed for compound 6a, a series of trifluoromethyl-substituted analogues (6b–f) were designed. Incorporation of a trifluoromethyl group into biologically active molecules is known to enhance their potency, lipophilicity, and metabolic stability.39,40 Adeniji et al. developed five classes of flufenamic acid derivatives, specifically 2-{[3-(trifluoromethyl)phenyl]amino}-benzoic acids, as potent AKR1C3 inhibitors.41 The removal of the trifluoromethyl group from certain inhibitors led to a significant reduction in their AKR1C3 inhibitory potency.41 These literature findings inspired the present synthesis of trifluoromethylated derivatives of compound 6a (Scheme 3). Various regioisomers were prepared starting from 13α-estrone (1) and the corresponding 2-chloropyridine derivatives (10b–f, Scheme 3). The Cu(I)-catalyzed etherification reaction was performed using K3PO4 as a base in the presence of 2-carboxypyridine under microwave irradiation at 100 °C for 2 h. The desired products (6b–f) were obtained in high yields.
| Compd. number | Structure | AKR1C1 | AKR1C2 | AKR1C3 |
|---|---|---|---|---|
| Inhibition (%) (IC50 (95% CI)) | Inhibition (%) (IC50 (95% CI)) | Inhibition (%) (IC50 (95% CI)) | ||
| 10 µM | 10 µM | 10 µM | ||
| a Data taken from the previous publication.15b Positive control for AKR1C1, medroxyprogesterone acetate (10 µM); positive control for AKR1C2, ursodeoxycholic acid (10 µM), positive control for AKR1C3, medroxyprogesterone acetate (1 µM). NI, no inhibition. IC50 values were calculated in GraphPad Prism using non-linear regression curve fitting with four parameters. | ||||
| 1 | 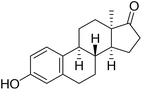 |
7.5a | 3.5a | 22.5a |
| 4 | 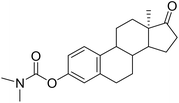 |
NI | 56.25 (IC50 = 7.32 µM (4.21–15.68 µM)) | 30.35 |
| 5 | 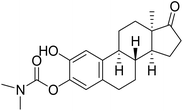 |
32.1 | 19.5 | 24.4 |
| 6a | 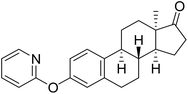 |
30.5 | 47.8 (IC50 = 4.64 µM (2.98–16.96 µM)) | 41.6 |
| 6b | 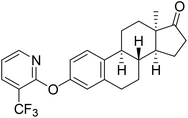 |
NI | 6.48 | 27.20 |
| 6c | 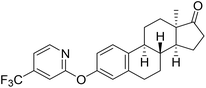 |
NI | NI | 24.95 |
| 6d | 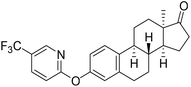 |
NI | 14.56 | 16.87 |
| 6e | 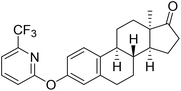 |
NI | NI | 23.00 |
| 6f | 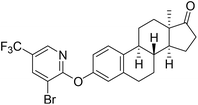 |
7.47 | NI | 18.55 |
| 7 | 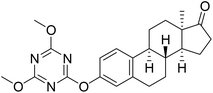 |
NI | 29.27 (IC50 = 21.31 µM (9.35–32.35 µM)) | 14.56 |
| 8 | 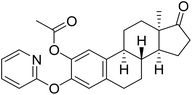 |
15.4 | 42.42 (IC50 = 16.67 µM (11.26–47.18 µM)) | 25.6 |
| 9 | 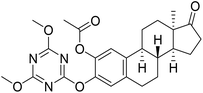 |
17.7 | 17.3 | 10.6 |
| 11 | 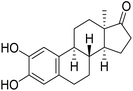 |
30.19 | 43.70 | 33.45 |
| 13 | 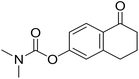 |
NI | NI | 7.1 |
| 14 | 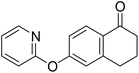 |
40.3 | NI | NI |
| 15 | 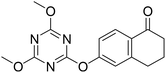 |
28.4 | 5.7 | 27.2 |
| Positive controlb | 93.4 | 94.2 | 95.0 | |
In the final stage of our steroid syntheses, a DG removal was carried out in order to obtain the 2,3-dihydroxy derivative (11, Scheme 4). The carbamoyl group could be efficiently cleaved under microwave-assisted heating, providing the desired compound (11) in good yield. This mild and rapid deprotection protocol proceeds under neutral microwave-assisted conditions, avoiding the use of acid or base catalysts and enabling efficient cleavage of the carbamate functionality. Such a methodology may find broad application in the transformation of complex, polycyclic, and multifunctional bioactive molecules, where the development of cleavage procedures with high functional-group tolerance remains a significant challenge. Due to the presence of ortho-positioned hydroxy groups, the newly and efficiently synthesized catechol derivative (11) possesses considerable potential for applications in medicinal chemistry. In addition to its possible direct bioactivity, it may also serve as a key intermediate, notably as a starting material in ring-closing reactions leading to potentially biologically active compounds.
 | ||
| Scheme 4 Synthesis of the 2,3-dihydroxy catechol derivative of 13α-estrone (11). Reagents and conditions: (i) toluene, MW, 120 °C, 30 min. | ||
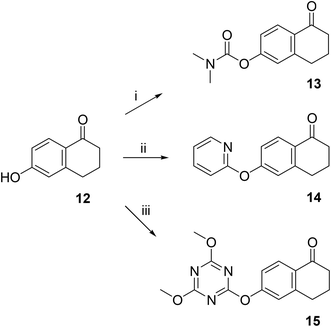
Following the synthesis of the steroidal compounds, DGs were also introduced onto the phenolic hydroxy function of 6-hydroxy-1-tetralone (12, Scheme 5). The target compounds (13–15) were efficiently synthesized using the above established protocols. Overall, the tetralone derivatives were obtained in slightly lower yields. The results of the biological studies (Table 1) suggest, that our newly synthesized tetralone derivatives (13–15) did not prove to be promising AKR1C inhibitors; therefore, we did not pursue further post-functionalization of these compounds.
The structures of the newly synthesized compounds were confirmed by 1H and 13C NMR spectroscopy (the 1H and 13C NMR spectra are listed in the supporting material). In assigning the signals, the NMR spectra of previously synthesized 13α-estrone derivatives were taken into account, particularly the characteristic coupling patterns of the protons in the A-ring.16,42,43 In the following section, several key correlations observed in the 1H NMR spectra are highlighted, which provide structural insight. However, these correlations alone are not conclusive; rather, they support the success of the transformations when evaluated alongside other NMR data.
The successful introduction of the three different DGs was confirmed by the appearance of distinct proton signals corresponding to the respective functional groups. The presence of the N,N-dimethylcarbamoyl group at the phenolic hydroxy position (in 1H NMR spectra of compounds 4 and 5) was evidenced by two singlets with triple integral value of each, around 3 ppm, consistent with previously reported data.28 The characteristic proton signals of the two heterocyclic DGs further confirmed the incorporation of these substituents. In the 1H NMR spectrum of compound 6a, four additional multiplets appeared in the region above 6 ppm, indicating the presence of a pyridyl ring. In contrast, the spectrum of compound 7 displayed a singlet (with six-fold integral value) at approximately 3.8 ppm, corresponding to the two methoxy groups of the triazine moiety. In the 1H NMR spectra of compounds 8, 9, and 11, the characteristic splitting pattern observed in those of their starting materials (5, 6a, and 7) due to their three aromatic protons is absent. The remaining two protons are too far apart to exhibit spin–spin coupling and therefore appear as singlets at different chemical shifts, depending on the nature of the C-3-O substituent. This observation supports the presence of the new ortho-substituent at the C-2 position (in compounds 8, 9, and 11). If the substituent had been introduced at the alternative ortho position (C-4), protons at positions C-1 and C-2 would have appeared as two doublets. Acetoxylation was confirmed by the appearance of a singlet at around 2 ppm, attributable to the three protons of the acetyl methyl group. Formation of the 2,3-dihydroxy derivative (11) was evidenced by four singlets in the 6–9 ppm region, corresponding to protons at positions C-1 and C-4, as well as the two hydroxy protons (2-OH and 3-OH).
The carbon atoms of the trifluoromethyl 2-pyridil rings were assigned using JMOD 13C NMR and HMBC spectra (in compounds 6b–f). Characteristic splittings related to this structural moiety appeared in the 13C spectra. Because of the C–F couplings, which typically occur over three bonds, quartets are observed in the JMOD spectra, with coupling constants decreasing by approximately one order of magnitude as the number of bonds between coupled nuclei increases. The HMBC spectra also assisted in the assignment of carbon signals of these compounds (a selected HMBC spectral region is shown in the SI, figure SHMBC).
3.2. Molecular docking
Using in silico molecular docking, the binding conformations of the identified active ligands within the AKR1C2 binding site were explored. Compound 6a (GFF_potential= −40.9 kcal mol−1) displayed predominantly hydrophobic interactions with residues Tyr24, Val54, Tyr55, Trp86, Trp227, and Leu308. Notably, in addition to hydrophobic contacts, Trp227—positioned at the entrance of the binding pocket—formed a hydrogen bond with the nitrogen atom of the terminal pyridine moiety of 6a, yielding a docking score of −4.08 kcal mol−1.Comparison with the reference ligand testosterone (TES) in the AKR1C2 crystal structure (PDB ID: 1J96, chain A) revealed hydrophobic interactions with Trp86, Val128, and Trp227 at the pocket entrance. The principal distinction between TES and 6a lies in the overall orientation of the steroid scaffold. In the crystal structure, TES adopts a binding mode wherein the C-3 carbonyl group positions deeply into the pocket, oriented towards the NAP co-factor. In contrast, due to steric hindrance imposed by the bulky substituent at C-3, compound 6a assumes an inverted steroid orientation, positioning its C-17 carbonyl toward the NAP co-factor, occupying the space typically occupied by TES C-3 carbonyl group (Fig. 2a). Consequently, the pyridine moiety at the C-3 position of 6a extends toward the pocket entrance, establishing an additional hydrogen bond with Trp227. This binding pose remained stable upon inclusion of the explicit NADPH cofactor during docking.
For compound 7 (GFF_potential= −30.4 kcal mol−1), a comparable scaffold inversion was observed. The presence of the methoxy-substituted 1,3,5-triazine terminal group enabled additional interactions, reflected in a docking score of −6.76 kcal mol−1 (Fig. 2b). The binding mode of 7 included hydrophobic contacts with Val54, Tyr55, Trp86, Ile129, Trp227 (π-stacking), and Leu308, alongside hydrogen bonds involving Tyr55, His117, Val128, Ile129, and Lys131. Compound 4 (GFF_potential = −36.5 kcal mol−1) exhibited a similar binding orientation relative to TES, engaging hydrophobic contacts with Tyr24, Tyr55, Trp227, and Leu306 (Fig. 2c). Its C-3 dimethylcarbamate substituent projected toward the pocket entrance, consistent with a docking score of −4.69 kcal mol−1.
The AKR1C2 binding site is defined by the residues Tyr24, Ala27, Val54, Tyr55, Trp86, Val128, Ile129, His222, Glu224, Pro226, Trp227, Leu306, Leu308, and Phe311. The highly homologous AKR1C1 (PDB ID: 1MRQ) shares an essentially identical binding site, comprising Tyr24, Ala27, Leu54, Tyr55, Trp86, Val128, Ile129, His222, Glu224, Pro226, Trp227, Leu306, Leu308, and Phe311, with only minor conformational differences (Fig. 2d). AKR1C3 (PDB ID: 1ZQ5) exhibits a largely conserved binding site architecture as well, with the key variation being Arg226 (in place of Pro226 in AKR1C1 and AKR1C2) located at the entrance of the binding pocket. The full AKR1C3 binding site includes Tyr24, Pro27, Leu54, Tyr55, Trp86, Leu128, Ser129, Gln222, Asp224, Arg226, Trp227, Phe306, Ser308, and Phe311.
Docking studies provided several noteworthy observations. Firstly, they underscore the importance of considering modifications on both the A- and D-rings of core-modified estrone derivatives, as these compounds may adopt two distinct binding orientations within the binding pocket. Both the chemical nature and steric bulk of the substituents are likely to substantially influence the binding mode and interaction strength. Furthermore, particular attention should be directed toward the pocket entrance, where interactions may play a critical role in determining binding affinity. In the present study, the side chain of Trp227, located at the entrance of the AKR1C2 binding site, was identified as a key residue warranting further investigation. We postulate that interactions with Trp227 and neighboring residues at the binding site entrance may represent important determinants of isoform selectivity and inhibitor binding affinity. This finding is particularly relevant given the high sequence and structural conservation among AKR1C isoforms, which complicates the straightforward rationalization of ligand selectivity based solely on static docking analyses. Nevertheless, pronounced differences at the pocket entrance—most notably the substitution of Pro226 with Arg226 in AKR1C3, involving a change from a cis- to a trans-peptide bond—may contribute to isoform-specific interactions. Explicit ligand protein interactions as LigPlot+ 2D projections are presented in Fig. 3a for the benefit of the reader. Further investigations employing molecular dynamics simulations may provide valuable insights into the roles of conformational flexibility and transient interactions in ligand binding and selectivity.
3.3. Evaluation of biological activity
The aldo-keto reductase isoenzymes AKR1C1, AKR1C2, and AKR1C3 regulate the local availability of ligands capable of binding to steroid hormone receptors. This prereceptor modulation of steroid hormone action has been implicated in the progression of hormone-dependent malignancies.3 Consequently, AKR1C inhibitors are of considerable interest for the treatment of hormone-dependent cancers and related endocrine disorders. Owing to their capacity to reduce carbonyl-containing compounds, these enzymes can also highly contribute to the development of chemoresistance. Inhibition of AKR1C enzymes may therefore potentiate the cytotoxic effects of chemotherapeutic agents. Given that these isoenzymes often catalyze overlapping biochemical transformations, inhibition of one member may be compensated by another. Thus, while selective AKR1C inhibitors remain valuable, the development of pan-inhibitors also represents a promising therapeutic strategy.The AKR1C1–3 inhibitory activity of newly synthesized steroidal compounds bearing three distinct DGs (4, 6a, and 7), along with their C-2-substituted analogs (5, 8, and 9) was initially evaluated. All compounds were tested at a concentration of 10 µM (Table 1) using 1-acenaphthenol as substrate (see Section 3.3. for substrate concentrations for each enzyme). Among them, compounds 4, 6a, and 8 exhibited inhibitory activity of approximately 40% or greater, exclusively against the AKR1C2 isozyme (SI, figure SDR). Of these, the pyridyl-substituted derivative (6a) demonstrated the most potent and broad-spectrum inhibition across all three AKR1C isozymes, with the highest effect observed for AKR1C2.
Following these results, the IC50 value of compound 6a was determined for AKR1C2, yielding a value of 4.64 µM. Motivated by this promising activity and supported by favorable literature precedents,39–41 we synthesized a series of trifluoromethyl-substituted derivatives of 6a (compounds 6b–e), where the CF3 group was present at different positions of the pyridyl ring. Additionally, one analog (6f) incorporated a bromine atom. However, enzymatic assays revealed that none of these modified compounds exhibited significant inhibition toward the AKR1C isozymes, suggesting that the introduction of a CF3 group may negatively affect enzyme inhibition.
A derivative lacking a DG (compound 11), which featured hydroxy substituents at C-2 and C-3, displayed moderate inhibitory activity (>30%) against all three AKR1C isozymes, with the greatest effect again observed against AKR1C2.
In comparison to the parent compound 13α-estrone (1), the semi-synthetic derivatives demonstrated an overall increase in inhibitory potency. Specifically, for AKR1C1, the combined modification via 3-O-carbamoylation and hydroxylation at C-2 (compound 5) increased the inhibition from 7.5% (compound 1) to 32.1%. For AKR1C2, the introduction of a pyridyl moiety resulted in a more than 50% increase in inhibitory activity. Similarly, incorporation of the pyridyl group led to an approximately 20% improvement in AKR1C3 inhibition.
These findings suggest that etherification of the phenolic 3-hydroxy group with a pyridyl ring, or the introduction of an additional hydroxy group to the ortho-position relative to the existing 3-OH, may be beneficial for AKR1C isozyme inhibition. Conversely, substitution at the C-2 position was generally detrimental to inhibitory activity in all DG-containing steroid derivatives.
Consistent with our previous publication,15 the present results also suggest that modifications of the A-ring of the studied estrone derivatives significantly influence their affinity toward AKR1C1–3 enzymes. Both substitution of the aromatic ring and derivatization of the phenolic hydroxy group through arylation or acylation were found to affect enzyme inhibition. To establish more precise structure–activity relationships, further synthesis and evaluation of additional derivatives will be required.
In parallel with the steroid series, tetralone analogs bearing the same three DGs (compounds 13–15) were also evaluated at a test concentration of 10 µM. These compounds did not display notable inhibitory effects, except for the pyridyl derivative (14), which selectively inhibited AKR1C1 by over 40%. Due to the overall low activity of the tetralone scaffold, ortho-functionalizations were not pursued further in this series.
4. Conclusions
Directed C–H activation-based functionalizations were successfully performed on the 13α-estrone (1) scaffold. Following the introduction of directing groups (DGs), such as N,N-dimethylcarbamoyl, 2-pyridyl, and 4,6-dimethoxy-1,3,5-triazin-2-yl moieties, regioselective ortho-hydroxylation and acetoxylation reactions were accomplished at the C-2 position. The C–H activation methodologies, previously developed primarily for small molecules, were effectively extended to estrane-based scaffolds. Most reactions were performed under microwave-assisted conditions, which significantly reduced reaction times and thereby rendered the processes more sustainable. Furthermore, an efficient and environmentally friendly carbamate cleavage protocol was developed, proceeding under neutral, microwave-assisted conditions with mild heating. This method may play an important role in the selective transformation of multifunctional drug-like molecules. The DGs were also introduced onto the tetralone scaffold (12); however, C–H activation reactions were not pursued owing to the modest biological performance of the resulting compounds (13–15). The inhibitory activities of the synthesized compounds against aldo-keto reductase isoenzymes AKR1C1–3 were evaluated. Among them, three compounds (4, 6a, and 7) exhibited potent inhibition of AKR1C2. To the best of our knowledge, these inhibitors are the first 13α-estrone derivatives featuring a nitrogen-containing substituent on the A-ring reported to date. To gain insight into the molecular basis of inhibition, docking studies were performed. These revealed that compound 6a adopts an inverted binding orientation on AKR1C2 isoform compared to the reference ligand testosterone (TES), with its 17-carbonyl group directed toward the NAP cofactor. Notably, the pyridyl moiety of 6a engages in a hydrogen-bonding interaction with Trp227 at the entrance of the binding pocket. This interaction, in combination with additional contacts, is proposed to contribute significantly to both binding affinity and isoform selectivity.The AKR1C2 inhibitors identified in this study could serve as promising starting points for the development of valuable therapeutic agents. Considering the central role of AKR1C2 as a 3-ketosteroid reductase, its inhibition could represent a therapeutic approach for androgen insufficiency syndromes.9 AKR1C2 has also emerged as a potential target for the suppression of metastatic dissemination. Particular attention should be paid to triple-negative breast cancer (TNBC), which displays an elevated risk of distant metastasis compared with other breast cancer subtypes.44 AKR1C2 serves as a key regulator of tumor growth and metastasis in TNBC cells, modulating androgen receptor (AR) expression and being associated with the luminal androgen receptor (LAR) subtype.
The identified estrone-based AKR1C isoform-selective or pan-inhibitors represent promising candidates for combination therapies against drug-resistant tumors. Considering the growing clinical relevance of therapy-resistant cancers, further evaluation of the most potent inhibitors for their ability to overcome resistance mechanisms is strongly justified.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra08903d.Acknowledgements
This work was supported by National Research, Development and Innovation Office-NKFIH through project SNN 139323 to EM. The work was supported by the Slovenian Agency for Research and Innovations grants no. N1-0234 and P3-0449 to TLR. This work was supported by the Ministry of Innovation and Technology of Hungary (TKP2021-EGA-32). The work was supported by Slovenian Research and Innovation Agency ARIS grant no. J7-50043 to UB. This work was supported by the NTP-NFTÖ-25-0797 to IR.References
- T. L. Rižner and T. M. Penning, Steroids, 2014, 79, 49–63 CrossRef PubMed.
- T. M. Penning, M. E. Burczynski, J. M. Jez, C. F. Hung, H. K. Lin, M. Ma, M. Moore, K. Ratnam and J. Rogash, Biochem. J., 2000, 351, 67–77 CrossRef CAS PubMed.
- T. M. Penning, P. Wangtrakuldee and R. J. Auchus, Endocr. Rev., 2019, 40, 447–475 CrossRef PubMed.
- K. K. Sharma, A. Lindqvist, X. J. Zhou, R. J. Auchus, T. M. Penning and S. Andersson, Mol. Cell. Endocrinol., 2006, 248, 79–86 CrossRef CAS PubMed.
- M. C. Byrns, S. Steckelbroeck and T. M. Penning, Biochem. Pharmacol., 2008, 75, 484–493 CrossRef CAS PubMed.
- R. J. Auchus, Trends Endocrinol. Metab., 2004, 15, 432–438 CrossRef CAS PubMed.
- T. L. Rižner, Front. Pharmacol, 2012, 3, 34 Search PubMed.
- T. Suzuki-Yamamoto, M. Nishizawa, M. Fukui, E. Okuda-Ashitaka, T. Nakajima, S. Ito and K. Watanabe, FEBS Lett., 1999, 462, 335–340 CrossRef CAS PubMed.
- T. M. Penning, S. Jonnalagadda, P. C. Trippier and T. L. Rižner, Pharmacol. Rev., 2021, 73, 1150–1171 CrossRef CAS PubMed.
- T. Zhong, F. Xu, J. Xu, L. Liu and Y. Chen, Biomed. Pharmacother., 2015, 69, 317–325 CrossRef CAS PubMed.
- J. Hofman, B. Malcekova, A. Skarka, E. Novotna and V. Wsol, Toxicol. Appl. Pharmacol., 2014, 278, 238–248 CrossRef CAS PubMed.
- H. B. Deng, H. K. Parekh, K. C. Chow and H. Simpkins, J. Biol. Chem., 2002, 277, 15035–15043 CrossRef CAS PubMed.
- C. Li, X. Wu, W. Zhang, J. Li, H. Liu, M. Hao, T. Han, C. Xie, G. Zhao and X. Cheng, J. Biomol. Screening, 2016, 21, 101–107 CrossRef CAS PubMed.
- S. Li, W. Lee, W. Heo, H. Y. Son, Y. Her, J. I. Kim and H. G. Moon, J. Breast Cancer, 2023, 26, 60–76 CrossRef PubMed.
- M. Sinreih, R. Jójárt, Z. Kele, T. Büdefeld, G. Paragi, E. Mernyák and T. L. Rižner, J. Enzyme Inhib. Med. Chem., 2021, 36, 1500–1508 Search PubMed.
- B. Schonecker, C. Lange, M. Kotteritzsch, W. Gunther, J. Weston, E. Anders and H. Görls, J. Org. Chem., 2000, 65, 5487–5497 CrossRef PubMed.
- D. Ayan, J. Roy, R. Maltais and D. Poirier, J. Steroid Biochem. Mol. Biol., 2011, 127, 324–330 CrossRef CAS PubMed.
- Y. Huang, A. Bhan, M. Sharma and R. Gounder, ACS Catal., 2019, 9, 521–555 CrossRef.
- P. Brožič, S. Turk, A. O. Adeniji, J. Konc, D. Janežič, T. M. Penning, T. L. Rižner and S. Gobec, J. Med. Chem., 2012, 55, 7417–7424 CrossRef PubMed.
- M. P. Savić, J. J. Ajduković, J. J. Plavša, S. S. Bekić, A. S. Ćelić, O. R. Klisurić, D. S. Jakimov, E. T. Petri and E. A. Djurendić, Med. Chem. Commun., 2018, 9, 969–981 RSC.
- X. Yang, Y. Sun, Z. Chen and Y. Rao, Adv. Synth. Catal., 2014, 356, 2137–2142 Search PubMed.
- M. Y. Ba, L. W. Xia, H. L. Li, Y. G. Wang, Y. N. Chu, Q. Zhao, C. P. Hu, X. T. He, T. X. Li, K. Y. Liang, Y. H. Zhang, L. Yang, W. H. Xie, H. Yang and M. R. Sun, Steroids, 2019, 146, 99–103 CrossRef CAS PubMed.
- L. Wang, L. Pan, Y. Huang, Q. Chen and M. He, Eur. J. Org Chem., 2016, 2016, 2530–2535 Search PubMed.
- J. H. P. Tyman, Synthetic and Natural Phenols, Elsevier, New York, 1996 Search PubMed.
- Z. Peng, Z. Yu, T. Li, N. Li, Y. Wang, L. Song and C. Jiang, Organometallics, 2017, 36, 1884–1891 CrossRef.
- K. Sheng, Y. Song, F. Lei, W. Zhao, L. Fan, L. Wu, Y. Liu, S. Wu and Y. Zhang, Eur. J. Med. Chem., 2022, 227, 113964 CrossRef CAS PubMed.
- R. K. Rej, J. E. Thomas Jr, R. K. Acharyya, J. M. Rae and S. Wang, J. Med. Chem., 2023, 66, 8339–8381 CrossRef CAS PubMed.
- P. Traj, A. H. Abdolkhaliq, A. Németh, S. T. Dajcs, F. Tömösi, T. L. Rižner, I. Zupkó and E. Mernyák, J. Enzyme Inhib. Med. Chem., 2021, 36, 895–902 CrossRef CAS PubMed.
- J. F. Couture, P. Legrand, L. Cantin, V. Luu-The, F. Labrie and R. Breton, J. Mol. Biol., 2003, 331, 593–604 CrossRef CAS PubMed.
- V. Nahoum, A. Gangloff, P. Legrand, D. W. Zhu, L. Cantin, B. S. Zhorov, V. Luu-The, F. Labrie, R. Breton and S. X. Lin, J. Biol. Chem., 2001, 276, 42091–42098 CrossRef CAS PubMed.
- W. Qiu, M. Zhou, M. Mazumdar, A. Azzi, D. Ghanmi, V. Luu-The, F. Labrie and S. X. Lin, J. Biol. Chem., 2007, 282, 8368–8379 CrossRef CAS PubMed.
- S. Ruiz-Carmona, D. Alvarez-Garcia, N. Foloppe, A. B. Garmendia-Doval, S. Juhos, P. Schmidtke, X. Barril, R. E. Hubbard and S. D. Morley, PLoS Comput. Biol., 2014, 10, e1003571 CrossRef PubMed.
- D. Soler, Y. Westermaier and R. Soliva, J. Comput.-Aided Mol. Des., 2019, 33, 613–626 CrossRef CAS PubMed.
- M. J. Hartshorn, M. L. Verdonk, G. Chessari, S. C. Brewerton, W. T. M. Mooij, P. N. Mortenson and C. W. Murray, J. Med. Chem., 2007, 50, 726–741 CrossRef CAS PubMed.
- S. D. Morley and M. Afshar, J. Comput.-Aided Mol. Des., 2004, 18, 189–208 CrossRef CAS PubMed.
- P. Brozic, N. Beranič, S. Gobec and T. L. Rižner, Mol. Cell. Endocrinol., 2006, 259, 30–42 CrossRef CAS PubMed.
- N. Beranič, P. Brozic and T. L. Rižner, Chem. Biol. Interact., 2013, 202, 204–209 CrossRef PubMed.
- R. K. Henderson, A. P. Hill, A. M. Redman and H. F. Sneddon, Green Chem., 2015, 17, 945–949 RSC.
- A. S. Nair, A. K. Singh, A. Kumar, S. Kumar, S. Sukumaran, V. P. Koyiparambath, L. K. Pappachen, T. M. Rangarajan, H. Kim and B. Mathew, Processes, 2022, 10, 2054 CrossRef CAS.
- C. C. Tseng, G. Baillie, G. Donvito, M. A. Mustafa, S. E. Juola, C. Zanato, C. Massarenti, S. Dall'Angelo, W. T. A. Harrison, A. H. Lichtman, R. A. Ross, M. Zanda and I. R. Greig, J. Med. Chem., 2019, 62, 5049–5062 CrossRef CAS PubMed.
- A. O. Adeniji, B. M. Twenter, M. C. Byrns, Y. Jin, M. Chen, J. D. Winkler and T. M. Penning, J. Med. Chem., 2012, 55, 2311–2323 CrossRef CAS PubMed.
- P. Ciuffreda, P. Ferraboschi, E. Verza and A. Manzocchini, Magn. Reson. Chem., 2001, 39, 648–650 CrossRef CAS.
- I. Bacsa, R. Jójárt, G. Schneider, J. Wölfling, P. Maróti, B. E. Herman, M. Szécsi and E. Mernyák, Steroids, 2015, 104, 230–236 CrossRef CAS PubMed.
- J. Li, D. He, Y. Bi and S. Liu, Breast Cancer, 2023, 15, 825–840 CAS.
| This journal is © The Royal Society of Chemistry 2026 |