 Open Access Article
Open Access ArticleCholesterol tunes lipid bilayer interactions
Wei Liua,
Jinwei Zhonga,
Pui Wo Felix Yeung b,
Xiahui Xiaoa,
Zuwei Zhao
b,
Xiahui Xiaoa,
Zuwei Zhao a,
Hang Jiang
a,
Hang Jiang a,
Yuwei Zhu
a,
Yuwei Zhu *a and
To Ngai
*a and
To Ngai *b
*b
aThe Key Laboratory of Synthetic and Biological Colloids, Ministry of Education & School of Chemical and Material Engineering, Jiangnan University, Wuxi 214122, P.R. China. E-mail: uvyzhu@gmail.com
bDepartment of Chemistry, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong 999077, P.R. China. E-mail: tongai@cuhk.edu.hk
First published on 13th January 2026
Abstract
In this study, we prepared SLBs on polystyrene colloids (CSLBs) and planar glass substrates with tunable cholesterol content. Using a fixed DPPC![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DOPC molar ratio of 3
DOPC molar ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to promote the phase separation, we observed pronounced heterogeneity in CSLB diffusion near planar membranes. Notably, by profiling interbilayer interaction potentials, we found that cholesterol concentrations of ∼11 and ∼30 mol% significantly enhance bilayer adhesion—likely through membrane fusion intermediates. These findings highlight cholesterol's critical role in modulating interbilayer mechanics.
1 to promote the phase separation, we observed pronounced heterogeneity in CSLB diffusion near planar membranes. Notably, by profiling interbilayer interaction potentials, we found that cholesterol concentrations of ∼11 and ∼30 mol% significantly enhance bilayer adhesion—likely through membrane fusion intermediates. These findings highlight cholesterol's critical role in modulating interbilayer mechanics.
1 Introduction
Accumulating evidence highlights the pivotal role of cholesterol in regulating the biophysical properties of cellular membranes, particularly in processes involving membrane fusion.1–3 Cholesterol primarily reduces membrane fluidity and permeability by rigidifying the fluid phase of lipid bilayers.4,5 In addition, it enhances membrane stability and increases the bending modulus, promoting negative curvature that serves as a structural precursor to fusion events.6,7 Through these effects, cholesterol is thought to modulate a range of critical cellular processes, including immune signalling,8 host–pathogen interactions,9 cancer development and progression,10 and cardiovascular disease.11Cellular membranes exhibit intrinsic spatiotemporal dynamics and are characterized by lateral and temporal heterogeneity at sub-microscopic scales. This heterogeneity, and its physiological relevance, has been closely associated with the lipid raft hypothesis,12 which posits the emergence of liquid–liquid phase separation (LLPS) within lipid bilayers. According to this framework, affinity–driven interactions and preferential associations between cholesterol and saturated lipids facilitate the formation of relatively ordered microdomains with reduced fluidity. These liquid-ordered (Lo) phases, enriched in hydrophobic components, promote key cellular processes such as signalling and trafficking, particularly via lipidated proteins and glycosylphosphatidylinositol (GPI)-anchored proteins.13 A previous study reported that approximately 35% of plasma membrane proteins localize within the Lo phase.14 Given the critical role of these phase-separated raft nanodomains (typically 10–200 nm) that have the potential to form microdomains (>300 nm) in mediating lipid–lipid and lipid–protein interactions, a range of biophysical model systems—including lipid monolayers, supported lipid bilayers (SLBs), micro/nanoscopic liposomes, and giant unilamellar vesicles (GUVs)—have been employed to investigate LLPS and the underlying principles of lipid raft formation.15–18
State-of-the-art techniques—including super-resolution optical microscopy (e.g., PALM, STED, and STORM),19–21 single-molecule fluorescence tracking,22–28 fluorescence correlation spectroscopy,29–34 Förster resonance energy transfer (FRET),35–39 and quartz crystal microbalance with dissipation monitoring (QCM-D)40,41—have been employed to probe nanoscale structures, membrane heterogeneity, and lateral diffusion of membrane components. These methodologies have provided compelling evidence for the structural and dynamic characteristics of cholesterol-enriched domains. In addition, simulations serve as powerful tools for probing biomolecular diffusivity, phase separation, membrane fusion, and vesicle–membrane interactions.2,7,42–54 Recent systematic molecular dynamics studies have revealed that membrane fusion is facilitated when lipid bilayers are enriched in cholesterol, whereas fusion is markedly reduced when both the vesicle and planar membrane are cholesterol-rich.7 Another study further demonstrated that elevating the cholesterol content of exosome membranes enhances their cellular uptake via fusion with the plasma membrane.2 Collectively, these theoretical investigations highlight the critical role of cholesterol in regulating the biofunctions of cellular membranes.
However, direct quantification of cholesterol's impact on interbilayer interactions remains a fundamental challenge. Neutron and X-ray reflectivity experiments have been widely employed to probe the attractive interactions between fully charged lipid bilayers, as well as to quantify key physical properties such as bending modulus, orientational order parameters, and membrane curvature during fusion events.6,55–57 In addition, Bevan and co-workers utilized total internal reflection microscopy (TIRM) to directly measure the kT-scale interactions between silica colloids modified with SLBs of varying compositions (e.g., PEGylated vs. non-PEGylated).58 Their findings demonstrated that colloid size and the extent of SLB PEGylation jointly dictate the balance between colloidal aggregation and SLB fusion, thereby highlighting the interplay between colloidal surface chemistry and bilayer stability. A deeper understanding of the underlying physical principles governing membrane–membrane interactions could pave the way for the rational design of lipid vesicle-based drug delivery systems and next-generation gene therapies.
In this study, we aimed to answer, at least in part, how cholesterol tunes the inter-bilayer interactions by a direct measurement using TIRM (Fig. 1). We prepared the SLBs on 3 µm polystyrene colloids and planar glass substrates with tunable cholesterol content. The bilayers consisted of saturated DPPC and unsaturated DOPC at a fixed molar ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. This composition was specifically selected to promote LLPS across a broad cholesterol concentration range (approximately 15–50 mol%), as supported by previous literature.59 We demonstrated that cholesterol content can significantly modulate the diffusivity of colloidal-supported lipid bilayers near planar membranes, with notably enhanced interfacial adhesion observed at approximately 11 and 30 mol% cholesterol.
1. This composition was specifically selected to promote LLPS across a broad cholesterol concentration range (approximately 15–50 mol%), as supported by previous literature.59 We demonstrated that cholesterol content can significantly modulate the diffusivity of colloidal-supported lipid bilayers near planar membranes, with notably enhanced interfacial adhesion observed at approximately 11 and 30 mol% cholesterol.
 | ||
| Fig. 1 Schematic illustration of the confined motion of colloidal-supported lipid bilayers near a bio-interface by evanescent scattering with a penetration depth of dp ∼100 nm. | ||
2 Experimental
2.1. Preparation of liposomes
Liposomes were prepared using saturated 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC, 99%, Macklin), unsaturated 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC, >99%, Aladdin) and cholesterol (≥99%, Macklin) based on an established protocol.60 In typical experiments, cholesterol (0.3867 g) was dissolved in 1 mL of a mixed organic solvent (cyclohexane/ethanol = 19/1, v/v) to prepare a stock solution at a concentration of 1.0 mmol mL−1. 0.01 g DPPC and 0.0035 g DOPC were dissolved in the mixed solvent to prepare a solution at desired concentrations. Lipid samples were dissolved with ultrasonic assistance to yield liposome solutions at a final concentration of 2 mM. To remove impurities, each sample was filtered three times through a 0.22 µm Nylon membrane filter.2.2. Cleaning of glass plates
Glass slide (25 × 75 × 1 mm, Fisher Scientific) was immersed in piranha solution (H2SO4/H2O2 = 3/1, v/v) for 2 h and then washed with D. I. H2O for 10 times to remove the excess piranha solution. Subsequently, the slide was sequentially washed with D. I. H2O, absolute ethanol, and D. I. H2O, each for 15 min through ultrasonication. The cleaned slide was treated with plasma (PDC-002, Harrick Plasma, USA) for 3 min to obtain a superhydrophilic surface.2.3. Preparation of planar supported lipid bilayers
Supported lipid bilayers (SLBs) were prepared through the Langmuir–Blodgett (LB) technique (Fig. 2).61 Lipid stock solutions were spread at the air–water interface of a LB trough (Microtrough G2, Kibron, Finland), making the surface pressure to be 1 mN m−1 after calibration, and the interface was allowed to equilibrate for 30 min. The lipid monolayer was compressed at 10 mm min−1 until the surface pressure reached ∼40 mN m−1, and then transferred to the glass substrate by vertical deposition at a pulling rate of 2 mm min−1. The second monolayer was spread and compressed in the same way as the one for the first layer, then deposited on the coated surface by dipping the substrate into the subphase at a rate of 2 mm min−1. The lipid bilayer-coated glass was transferred using a polytetrafluoroethylene (PTFE) mold, during which the lipid bilayer was maintained in an aqueous environment.2.4. Preparation of colloidal supported lipid bilayers
12 µL lipid solution was dried under nitrogen to form a thin film. After drying, 1 mM NaCl solution was added, and the sample was sonicated for 5 min to disperse the lipids. The mixture was incubated in a 60 °C water bath for 1 h to form vesicles. Polystyrene (PS) particles (8 w/w%, 3.1 µm, Thermo Fisher Scientific) was added into the solution. After 30 min of incubation, the sample was shaken for 30 min to yield lipid bilayer-coated particle surfaces (colloidal-supported lipid bilayers, CSLBs). Above the phase transition temperature, lipid molecules become more fluid-like and loosely packed, facilitating bilayer formation on the particle surface.2.5. Observation of lipid monolayer and bilayer
To visualize lipid monolayer/bilayer under confocal fluorescent microscopy, 1 wt% fluorescent lipid, NBD PC (Avanti) was added in the lipid solution. The lipid monolayer and bilayer were prepared on the glass substrate through LB technique and on particle surfaces as aforementioned. The as-prepared lipid monolayer and bilayer were observed using a confocal laser fluorescent microscope (Eclipse Ti2, Nikon Inc., JP) at the exciting wavelength of 488 nm (Fig. 3). The lipid bilayer was maintained in an aqueous environment during the experiment.2.6. Total internal reflection microscopy
To form an evanescent field at the glass/water interface, a HeNe laser with a wavelength λ = 632.8 nm (HeNe Laser, Red, 633 nm, Newport Corporation, USA) was employed in our setup. The intensity of scattering light I exponentially decays with a separation distance h from the interface, expressed as62| I(h) = I0e−h/dp | (1) |
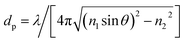 | (2) |
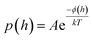 | (3) |
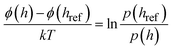 | (4) |
2.7. Image acquisition and data processing
A high-speed back-illuminated sCMOS camera (Kinetix22, 2400 × 2400 6.5 µm pixels, Teledyne Photometrics Inc., Tucson, US) in liquid-cooling mode was utilized to capture the TIR-scattering image sequences. A 40× objective (CFI Super Plan Fluor ELWD, NA/WD: 0.6/3.6 mm) mounted on an upright microscope (ECLIPSE FN1, Nikon Inc., JP) was used for imaging. The effective pixel size, lpix, was determined to be ∼0.16 µm. Typically, an image sequence of N = 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames in size of 256 × 256 pixels in 16 bits was captured at an exposure time of 10 ms, referring to a frame rate of 100 fps.
000 frames in size of 256 × 256 pixels in 16 bits was captured at an exposure time of 10 ms, referring to a frame rate of 100 fps.
Before TIRM measurements, the background intensity and the maximum intensity I0 corresponding the zero height for 3.1 µm PS letax microspheres were determined. The setup was calibrated by standard NaCl solutions to ensure the accuracy. In the experiments, two microscopic glass plates sandwiched by a silicone O-ring were employed to construct a flowing cell with 1 mm chamber height. Solutions of CSLBs were injected into the flowing cell through a micro-injection pump (1 mL min−1, SPLab02, Tansoole). CSLBs were allowed to equilibrate for ∼20 min before image capturing. The images were processed based on the radial symmetry method under particle-tracking framework using previously reported MATLAB program.65
2.8. The diExp elastic model
For particles stuck on the substrate surface, non-DLVO forces—such as hydrophobic interaction and hydration force—turn to be dominated in the intersurface potentials, thus the DLVO theory is not applicable. Accordingly, we use a modified Hookean spring model in an asymmetric shape to describe the potential wells, namely, the diExp elastic model, which is expressed as66
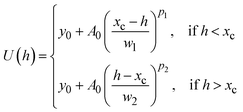 | (5) |
| keff ∼ (w1p1w2p2)−1/2 | (6) |
3 Results and discussion
3.1. Near-wall confined motion of CSLBs
Using total internal reflection microscopy, we first observed pronounced heterogeneity in the diffusive behavior of CSLBs near planar membrane surfaces (Fig. 4). This technique leverages evanescent scattering to enable high-precision tracking of kT-scale interactions between colloids and flat interfaces along the z-axis (normal to the surface). Detailed methodology is provided in our recent publication and other literature.66,67 Quantitative analysis revealed that the diffusion range of CSLBs initially increases with cholesterol content, reaching a maximum at approximately 27 mol% (x = 1.5), followed by a sharp decline (Fig. 5a). Coincidentally, the diffusion range under strictly-confined motion (e.g., x = 0, 0.5, 1.75 and 2) corresponded to the size of the phase-separated raft domains (∼200–300 nm). To further characterize this behavior, we performed time-averaged mean square displacement (MSD) measurements in three dimensions44
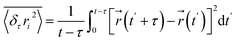 | (7) |
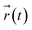 denotes the position of the tracked CSLBs and τ ≪ t is the lag time. After correcting for particle positional errors, a linear fit to the MSD yielded the diffusion coefficient D. Notably, two distinct minima in the z-directional diffusion coefficient (D ≈ 1.5–2 nm2 s−1) were observed at ∼11 and 30 mol% cholesterol, indicating strong confinement (Fig. 5b). Consistent with this, we detected extremely weak fluctuations in the corresponding z-directional displacements (data not shown). Furthermore, we confirmed that the observed low diffusivity was not solely attributable to near-wall hindrance effects (see height profile in Fig. 7a). These findings suggest that high-curvature lipid-stalk or hemifusion intermediates may form under these conditions, contributing to the observed confinement.
denotes the position of the tracked CSLBs and τ ≪ t is the lag time. After correcting for particle positional errors, a linear fit to the MSD yielded the diffusion coefficient D. Notably, two distinct minima in the z-directional diffusion coefficient (D ≈ 1.5–2 nm2 s−1) were observed at ∼11 and 30 mol% cholesterol, indicating strong confinement (Fig. 5b). Consistent with this, we detected extremely weak fluctuations in the corresponding z-directional displacements (data not shown). Furthermore, we confirmed that the observed low diffusivity was not solely attributable to near-wall hindrance effects (see height profile in Fig. 7a). These findings suggest that high-curvature lipid-stalk or hemifusion intermediates may form under these conditions, contributing to the observed confinement.
 | ||
Fig. 4 Representative lateral trajectories of colloidal-supported lipid bilayers (CSLBs) diffusing on planar membranes composed of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DPPC/DOPC, with varying cholesterol content. 1 DPPC/DOPC, with varying cholesterol content. | ||
3.2. Anomalous diffusion of CSLBs
Substantial heterogeneity in the diffusivity of individual CSLBs was revealed through the broad distributions of MSD and the power-law index α, where MSD scales as ∼τα) (Fig. 5c and d). In classical diffusion, MSD exhibits a linear dependence on lag time, i.e., MSD ∼ 2Dτ, with the diffusion coefficient D determined solely by the viscosity of a homogeneous environment. However, anomalous or subdiffusive behavior is commonly observed in biological systems, such as the lateral diffusion of transmembrane proteins on cellular membranes.44 Analogously, we observed widespread transient subdiffusion of CSLBs on membrane surfaces across a broad range of cholesterol content (x ∼ 0–3), as indicated by MSD analysis. Interestingly, the power-law index α derived from short lag times exhibited a strong positive correlation with D, whereas the index obtained from longer lag times (αlag) showed an inverse trend with D as a function of x. Compared to long-time measurements, short-time diffusivity is inherently more variable, reflecting dynamic heterogeneity in the local environment and transient interactions.68 This inverse relationship between short- and long-time diffusivity suggests a breakdown of ergodicity in the time-dependent diffusion process of CSLBs on membranes.693.3. Interbilayer interactions
The observed diffusive properties revealed abnormal adhesion behavior—manifested as extremely low diffusivity—of CSLBs on bilayer membranes at specific cholesterol concentrations (∼11 and 30 mol%). To further investigate how cholesterol content influences interbilayer interactions, we applied Boltzmann statistics to derive the intersurface potential energy U(h) by transforming the positional distribution of individual fluctuating particles along the z-axis. Fig. 6 presents the interbilayer potential energy profiles, with the equilibrium height h0 selected as the reference position. The most confined diffusion corresponds to the narrowest potential energy distribution, indicating strong adhesion with a confinement scale of <10 nm at U = 6kT. Consistent with our previous observation of asymmetric potential profiles—where single coronated particles diffused on SLBs66—we also detected asymmetry in the mechanical coupling of CSLBs. To further interpret this behavior, we applied the diExp elastic model (see Experimental), which yielded potential energy profiles in good agreement with non-DLVO theoretical predictions.To quantify the adhesion strength between bilayers, we calculated the effective elastic coefficient, keff, which serves as an estimate of the intersurface interaction for particles exhibiting confined motion under soft boundary conditions. A bimodal distribution of keff as a function of cholesterol content x was observed (Fig. 7b), consistent with the two minima in the diffusion coefficient. Additionally, Fig. 7c reveals a convergent distribution of the elastic exponents around a value of 2, indicative of Hookean spring-like behavior, regardless of whether the motion is directed toward or away from the wall.
 | ||
| Fig. 7 Quantitative analysis of interbilayer interactions. (a) Distribution of equilibrium height h0. (b) Effective elastic coefficient keff. (c) Corresponding elastic exponents for the potential wells shown in Fig. 6. | ||
Notably, the peak value of keff approaches ∼50 mN m−1, approximately three orders of magnitude greater than the adhesion strength observed between protein-coronated particles and SLBs.66 Such pronounced sticking behavior is unlikely to arise from purely near-field interactions without physical contact.
Therefore, we propose that high-curvature lipid-stalk or hemifusion intermediates70 may form between the bilayers at the two specific cholesterol contents (i.e., ∼11 and 30 mol%), which lie near the phase separation boundaries in the ternary phase diagram59,71 of the DPPC:DOPC:cholesterol system (see Fig. 6, top panel). At a relatively low cholesterol content (x ∼ 0–1), an uplifted and flexural fold structure with high internal fluidity is expected accordingly. As cholesterol increases beyond 20 mol%, the bilayer becomes more elastic and structurally stable. Under these conditions, the hydration layer between bilayers may act as a lubricating boundary, facilitating enhanced lateral diffusion at x ∼1.5. Further increasing cholesterol content (∼30 mol%, x ∼1.75–2) likely promotes the formation of cholesterol-rich microdomains (lipid rafts), potentially triggering hemifusion or full fusion events. At extremely high cholesterol levels (e.g., >40 mol%), the membranes become significantly rigidified, suppressing lipid exchange between bilayers; instead, interbilayer bridging via lipid rafts may become the dominant interaction mechanism.
4 Conclusions and discussion
Cholesterol is widely recognized as a key regulator of membrane fusion, where its cone-shaped molecular geometry introduces intrinsic negative curvature that promotes stalk formation.70 For fusion to occur, however, lipid bilayers must first overcome the hydration repulsion barrier imposed by ordered water molecules at the lipid–water interface.72,73 Recent studies suggest that cholesterol not only contributes negative curvature but also reduces hydration forces, thereby facilitating close membrane apposition and lowering the energetic cost of stalk formation.74,75 Importantly, the impact of cholesterol is not universal: while it can soften unsaturated lipid membranes, it stiffens saturated bilayers, underscoring its complex role in modulating membrane mobility, curvature sensing, and elasticity.4,76In our system, peak-like adhesion enhancement at ∼11 mol% and ∼30 mol% cholesterol concentrations in DPPC![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) DOPC (3
DOPC (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) supported bilayers reveals a nonlinear regulatory effect that resonates with established phase diagram studies of model membranes. Prieto and colleagues have shown that cholesterol drives liquid-ordered (Lo) and liquid-disordered (Ld) phase segregation within specific compositional windows, while suppressing domain formation outside these ranges.77–82 Such nonlinear behavior is attributed to cholesterol's wedge-like geometry, its modulation of local curvature, and preferential interactions with sphingolipids. We propose that the enhanced adhesion observed at intermediate cholesterol concentrations reflects the stabilization of fusion intermediates through curvature changes in local membrane domains. These findings extend the physicochemical framework of cholesterol action beyond classical DLVO-type interactions, highlighting the combined roles of hydrophobic and hydration forces in shaping membrane mechanics. By situating our results within this broader context, we provide a mechanistic basis for the unexpected nonlinear pattern and open avenues for confirmatory experiments using curvature-sensitive probes or molecular simulations.
1) supported bilayers reveals a nonlinear regulatory effect that resonates with established phase diagram studies of model membranes. Prieto and colleagues have shown that cholesterol drives liquid-ordered (Lo) and liquid-disordered (Ld) phase segregation within specific compositional windows, while suppressing domain formation outside these ranges.77–82 Such nonlinear behavior is attributed to cholesterol's wedge-like geometry, its modulation of local curvature, and preferential interactions with sphingolipids. We propose that the enhanced adhesion observed at intermediate cholesterol concentrations reflects the stabilization of fusion intermediates through curvature changes in local membrane domains. These findings extend the physicochemical framework of cholesterol action beyond classical DLVO-type interactions, highlighting the combined roles of hydrophobic and hydration forces in shaping membrane mechanics. By situating our results within this broader context, we provide a mechanistic basis for the unexpected nonlinear pattern and open avenues for confirmatory experiments using curvature-sensitive probes or molecular simulations.
In summary, we used total internal reflection microscopy to directly image and measure the near-wall confined diffusion and the kT-scale interaction between supported lipid bilayer-modified colloids and planar surfaces. We demonstrated that cholesterol content of approximately 11 and 30 mol% significantly enhance adhesion between lipid bilayers, likely through mechanisms involving membrane phase separation and fusion. These findings highlight the sophisticated role of cholesterol in modulating interbilayer interactions—an open question that remains unresolved. Future work will benefit from direct experimental observations such as FRET, single-particle tracking, Neutron or X-ray scattering and theoretical simulations to further validate the proposed framework, particularly within the DPPC:DOPC:cholesterol system.
Author contributions
Wei Liu: conceptualization, methodology, formal analysis, visualization, supervision, project administration, funding acquisition, writing – original draft. Jinwei Zhong: software, validation, formal analysis, investigation, data curation. Pui Wo Felix Yeung: software, validation, formal analysis. Xiahui Xiao: validation, investigation. Zuwei Zhao: validation, investigation. Hang Jiang: writing – review & editing. Yuwei Zhu: funding acquisition, writing – review & editing. To Ngai: conceptualization, project administration, writing – review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
Data will be made available on reasonable request.Acknowledgements
This work was supported by the National Natural Science Foundation of China (22303033) and the Fundamental Research Funds for the Central Universities, China (JUSRP202501136).Notes and references
- E. Sezgin, I. Levental, S. Mayor and C. Eggeling, Nat. Rev. Mol. Cell Biol., 2017, 18, 361–374 CrossRef CAS PubMed.
- Y. Zhuo, Z. Luo, Z. Zhu, J. Wang, X. Li, Z. Zhang, C. Guo, B. Wang, D. Nie and Y. Gan, Nat. Nanotechnol., 2024, 19, 1858–1868 CrossRef CAS PubMed.
- A. Mangiarotti, E. Sabri, K. V. Schmidt, C. Hoffmann, D. Milovanovic, R. Lipowsky and R. Dimova, Nat. Commun., 2025, 16, 1–20 Search PubMed.
- M. Pöhnl, M. F. Trollmann and R. A. Böckmann, Nat. Commun., 2023, 14, 8038 CrossRef PubMed.
- S. Chakraborty, M. Doktorova, T. R. Molugu, F. A. Heberle, H. L. Scott, B. Dzikovski, M. Nagao, L.-R. Stingaciu, R. F. Standaert and F. N. Barrera, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 21896–21905 CrossRef CAS PubMed.
- A. Ivankin and D. Gidalevitz, Biophys. J., 2012, 102, 293a CrossRef.
- C.-P. Chng, K. J. Hsia and C. Huang, Soft Matter, 2022, 18, 7752–7761 RSC.
- A. R. Tall and L. Yvan-Charvet, Nat. Rev. Immunol., 2015, 15, 104–116 CrossRef CAS PubMed.
- G. A. Kumar, M. Jafurulla and A. Chattopadhyay, Chem. Phys. Lipids, 2016, 199, 179–185 CrossRef CAS.
- O. F. Kuzu, M. A. Noory and G. P. Robertson, Cancer Res., 2016, 76, 2063–2070 CrossRef CAS.
- G. N. Levine, J. F. Keaney Jr and J. A. Vita, N. Engl. J. Med., 1995, 332, 512–521 CrossRef CAS PubMed.
- K. Simons and E. Ikonen, Nature, 1997, 387, 569–572 Search PubMed.
- M. G. Paulick and C. R. Bertozzi, Biochem., 2008, 47, 6991–7000 CrossRef CAS.
- I. Levental, D. Lingwood, M. Grzybek, Ü. Coskun and K. Simons, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 22050–22054 CrossRef CAS.
- J. A. Jackman and N.-J. Cho, Langmuir, 2020, 36, 1387–1400 CrossRef CAS.
- J. R. Silvius, Biochim. Biophys. Acta Biomembr., 2003, 1610, 174–183 CrossRef CAS PubMed.
- T. Baumgart, A. T. Hammond, P. Sengupta, S. T. Hess, D. A. Holowka, B. A. Baird and W. W. Webb, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 3165–3170 Search PubMed.
- S.-T. Yang, A. J. Kreutzberger, J. Lee, V. Kiessling and L. K. Tamm, Chem. Phys. Lipids, 2016, 199, 136–143 CrossRef CAS PubMed.
- K. Zhanghao, W. Liu, M. Li, Z. Wu, X. Wang, X. Chen, C. Shan, H. Wang, X. Chen and Q. Dai, Nat. Commun., 2020, 11, 5890 CrossRef CAS PubMed.
- M. B. Stone, S. A. Shelby and S. L. Veatch, Chem. Rev., 2017, 117, 7457–7477 CrossRef CAS PubMed.
- E. Sezgin, J. Phys. Condens. Matter, 2017, 29, 273001 CrossRef PubMed.
- L. Gascoigne, R. P. Tas, P. G. Moerman and I. K. Voets, Soft Matter, 2025, 21, 3058–3066 RSC.
- G. I. Mashanov, T. A. Nenasheva, A. Mashanova, R. Lape, N. J. Birdsall, L. Sivilotti and J. E. Molloy, Faraday Discuss., 2021, 232, 358–374 RSC.
- A. Kusumi, Y. M. Shirai, I. Koyama-Honda, K. G. Suzuki and T. K. Fujiwara, FEBS Lett., 2010, 584, 1814–1823 CrossRef CAS PubMed.
- S. Wieser and G. J. Schütz, Methods, 2008, 46, 131–140 CrossRef CAS PubMed.
- V. Kiessling, J. M. Crane and L. K. Tamm, Biophys. J., 2006, 91, 3313–3326 Search PubMed.
- K. Ritchie, X.-Y. Shan, J. Kondo, K. Iwasawa, T. Fujiwara and A. Kusumi, Biophys. J., 2005, 88, 2266–2277 CrossRef CAS.
- A. Kusumi, C. Nakada, K. Ritchie, K. Murase, K. Suzuki, H. Murakoshi, R. S. Kasai, J. Kondo and T. Fujiwara, Annu. Rev. Biophys. Biomol. Struct., 2005, 34, 351–378 CrossRef CAS PubMed.
- G.-S. Yang, A. Wagenknecht-Wiesner, B. Yin, P. Suresh, E. London, B. A. Baird and N. Bag, Biophys. J., 2024, 123, 2256–2270 CrossRef CAS.
- S. Chiantia, J. Ries and P. Schwille, Biochim. Biophys. Acta, Biomembr., 2009, 1788, 225–233 Search PubMed.
- J. Ries and P. Schwille, Phys. Chem. Chem. Phys., 2008, 10, 3487–3497 RSC.
- A. J. García-Sáez and P. Schwille, Methods, 2008, 46, 116–122 CrossRef.
- A. E. Hac, H. M. Seeger, M. Fidorra and T. Heimburg, Biophys. J., 2005, 88, 317–333 CrossRef CAS.
- J. Korlach, P. Schwille, W. W. Webb and G. W. Feigenson, Proc. Natl. Acad. Sci. U. S. A., 1999, 96, 8461–8466 CrossRef CAS.
- P. Sharma, R. Varma, R. Sarasij, K. Gousset, G. Krishnamoorthy, M. Rao and S. Mayor, Cell, 2004, 116, 577–589 CrossRef CAS.
- J. Škerle, J. Humpolíčková, N. Johnson, P. Rampírová, E. Poláchová, M. Fliegl, J. Dohnálek, A. Suchánková, D. Jakubec and K. Strisovsky, Biophys. J., 2020, 118, 1861–1875 CrossRef PubMed.
- D. F. Ma, C. H. Xu, W. Q. Hou, C. Y. Zhao, J. B. Ma, X. Y. Huang, Q. Jia, L. Ma, J. Diao and C. Liu, Angew. Chem., Int. Ed., 2019, 58, 5577–5581 CrossRef CAS.
- Y. Ma, Y. Yamamoto, P. R. Nicovich, J. Goyette, J. Rossy, J. J. Gooding and K. Gaus, Nat. Biotechnol., 2017, 35, 363–370 CrossRef CAS PubMed.
- M. You, E. Li, W. C. Wimley and K. Hristova, Anal. Biochem., 2005, 340, 154–164 CrossRef CAS PubMed.
- M. Hanzawa, H. Sugasawa, T. Ogura, K.-i. Iimura and T. Misono, Phys. Chem. Chem. Phys., 2025, 27, 6858–6866 RSC.
- F. Ruggeri, F. Zhang, T. Lind, E. D. Bruce, B. L. Lau and M. Cárdenas, Soft Matter, 2013, 9, 4219–4226 RSC.
- G. Meher, S. Sinha, G. P. Pattnaik, S. Ghosh Dastidar and H. Chakraborty, J. Phys. Chem. B, 2019, 123, 7113–7122 Search PubMed.
- M. D. Elola and J. Rodriguez, J. Phys. Chem. B, 2018, 122, 5897–5907 CrossRef CAS.
- E. Yamamoto, T. Akimoto, A. C. Kalli, K. Yasuoka and M. S. Sansom, Sci. Adv., 2017, 3, e1601871 Search PubMed.
- J. Yang, J. Martí and C. Calero, Soft Matter, 2016, 12, 4557–4561 Search PubMed.
- M. M. Kozlov and L. V. Chernomordik, Curr. Opin. Struct. Biol., 2015, 33, 61–67 CrossRef CAS PubMed.
- Y. G. Smirnova, S.-J. Marrink, R. Lipowsky and V. Knecht, J. Am. Chem. Soc., 2010, 132, 6710–6718 Search PubMed.
- F. de Meyer and B. Smit, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 3654–3658 Search PubMed.
- L. V. Chernomordik and M. M. Kozlov, Nat. Struct. Biol., 2008, 15, 675–683 CrossRef CAS PubMed.
- V. Knecht and S.-J. Marrink, Biophys. J., 2007, 92, 4254–4261 Search PubMed.
- A. Grafmüller, J. Shillcock and R. Lipowsky, Phys. Rev. Lett., 2007, 98, 218101 CrossRef.
- P. M. Kasson, N. W. Kelley, N. Singhal, M. Vrljic, A. T. Brunger and V. S. Pande, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 11916–11921 CrossRef CAS.
- R. Blumenthal, M. J. Clague, S. R. Durell and R. M. Epand, Chem. Rev., 2003, 103, 53–70 CrossRef CAS PubMed.
- V. S. Markin and J. P. Albanesi, Biophys. J., 2002, 82, 693–712 CrossRef CAS.
- T. Mukhina, A. Hemmerle, V. Rondelli, Y. Gerelli, G. Fragneto, J. Daillant and T. Charitat, J. Phys. Chem. Lett., 2019, 10, 7195–7199 CrossRef CAS.
- A. Hemmerle, L. Malaquin, T. Charitat, S. Lecuyer, G. Fragneto and J. Daillant, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 19938–19942 CrossRef CAS PubMed.
- J. Pan, T. T. Mills, S. Tristram-Nagle and J. F. Nagle, Phys. Rev. Lett., 2008, 100, 198103 CrossRef.
- W. N. Everett and M. A. Bevan, Soft Matter, 2014, 10, 332–342 Search PubMed.
- S. L. Veatch, I. Polozov, K. Gawrisch and S. L. Keller, Biophys. J., 2004, 86, 2910–2922 Search PubMed.
- F. Szoka Jr and D. Papahadjopoulos, Annu. Rev. Biophys., 1980, 9, 467–508 Search PubMed.
- J. Yuan, C. Hao, M. Chen, P. Berini and S. Zou, Langmuir, 2013, 29, 221–227 CrossRef CAS PubMed.
- D. C. Prieve and N. A. Frej, Langmuir, 1990, 6, 396–403 CrossRef CAS.
- M. A. Bevan and D. C. Prieve, J. Chem. Phys., 2000, 113, 1228–1236 CrossRef CAS.
- H.-J. Wu and M. A. Bevan, Langmuir, 2005, 21, 1244–1254 CrossRef CAS.
- F. Cao, J. Wu, Y. Li and T. Ngai, Langmuir, 2019, 35, 8910–8920 CrossRef CAS PubMed.
- W. Liu, Z. Zhao, J. Zhong, P. W. F. Yeung, J. Wu, Y. Li, H. Jiang, Y. Zhu and T. Ngai, Chem. Sci., 2025, 16, 16625–16637 RSC.
- J. Wu, W. Liu and T. Ngai, Soft Matter, 2023, 19, 4611–4627 RSC.
- B. Wang, S. M. Anthony, S. C. Bae and S. Granick, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 15160–15164 Search PubMed.
- C. Manzo, J. A. Torreno-Pina, P. Massignan, G. J. Lapeyre Jr, M. Lewenstein and M. F. Garcia Parajo, Phys. Rev. X, 2015, 5, 011021 Search PubMed.
- G. P. Pattnaik and H. Chakraborty, Vitamins and hormones, Elsevier, 2021, vol. 117, pp. 133–155 Search PubMed.
- H. Himeno, N. Shimokawa, S. Komura, D. Andelman, T. Hamada and M. Takagi, Soft Matter, 2014, 10, 7959–7967 RSC.
- T. J. McIntosh and S. A. Simon, Annu. Rev. Biophys. Biomol. Struct., 1994, 23, 27–51 Search PubMed.
- R. Rand and V. Parsegian, Biochim. Biophys. Acta Rev. Biomembr., 1989, 988, 351–376 CrossRef CAS.
- C. S. Poojari, K. C. Scherer and J. S. Hub, Nat. Commun., 2021, 12, 6594 Search PubMed.
- S. Aeffner, T. Reusch, B. Weinhausen and T. Salditt, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, E1609–E1618 Search PubMed.
- T. Kumarage, S. Gupta, N. B. Morris, F. T. Doole, H. L. Scott, L.-R. Stingaciu, S. V. Pingali, J. Katsaras, G. Khelashvili and M. Doktorova, Nat. Commun., 2025, 16, 7024 Search PubMed.
- R. F. de Almeida, L. M. Loura and M. Prieto, Chem. Phys. Lipids, 2009, 157, 61–77 CrossRef CAS.
- R. F. De Almeida, J. Borst, A. Fedorov, M. Prieto and A. J. Visser, Biophys. J., 2007, 93, 539–553 CrossRef PubMed.
- L. s. M. Loura, A. Fedorov and M. Prieto, Biochim. Biophys. Acta Biomembr., 2001, 1511, 236–243 CrossRef CAS PubMed.
- F. M. Goñi, A. Alonso, L. A. Bagatolli, R. E. Brown, D. Marsh, M. Prieto and J. L. Thewalt, Biochim. Biophys. Acta Mol. Cell Biol. Lipids, 2008, 1781, 665–684 CrossRef.
- R. F. De Almeida, A. Fedorov and M. Prieto, Biophys. J., 2003, 85, 2406–2416 Search PubMed.
- R. F. De Almeida, L. M. Loura, A. Fedorov and M. Prieto, J. Mol. Biol., 2005, 346, 1109–1120 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |




