Open Access Article
Open Access ArticleExploring the chemical reactivity of 2,6-dimethylchromone toward heterocyclic hydrazines and enamines: experimental and theoretical investigations
Mohamed Abdel-Megida,
Najla A. Alshayeb,
Al-Shimaa Badran *c and
Magdy A. Ibrahimc
*c and
Magdy A. Ibrahimc
aDepartment of Chemistry, College of Science, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh, Saudi Arabia. E-mail: moabmohamed@imamu.edu.sa
bDepartment of Chemistry, College of Science, Princess Nourah Bint Abdulrahman University, P.O. Box 84428, Riyadh 11671, Saudi Arabia. E-mail: naalshaye@pnu.edu.sa
cDepartment of Chemistry, Faculty of Education, Ain Shams University, Roxy, Cairo 11711, Egypt. E-mail: badran.shimaa@yahoo.com; elshimaabadran@edu.asu.edu.eg; Fax: +20 022581243; Tel: +20 01011444940
First published on 30th March 2026
Abstract
The current study aimed to investigate the chemical behavior of 2,6-dimethylchromone (1) towards some heterocyclic hydrazines and cyclic enamines. Reaction of compound 1 with 7-chloro-4-hydrazinylquinoline and 3-hydrazinyl-5,6-diphenyl-1,2,4-triazine gave pyrazole derivatives 4 and 5. Treatment of compound 1 with 5-amino-3-methyl-1H-pyrazole, 6-aminouracil, 6-aminothiouracil and 1,3-dimethyl-6-aminouracil produced pyrazolo[3,4-b]pyridine 6 and pyrido[2,3-d]pyrimidines 7–9, respectively. The biological efficiency of the synthesized compounds against hepatocellular carcinoma (HepG-2) cell lines was investigated through a combination of experimental and theoretical approaches, including molecular docking with the CDK1 protein (PDB ID: 4Y72). Among the tested molecules, compound 5 exhibited the most potent anticancer activity, with an IC50 value of 6.57 µM L−1, surpassing the reference drug cis-platin. For theoretical studies, all compounds were geometrically optimized using DFT/B3LYP functional with the 6-311G++(d,p) basis set. Structural parameters and global reactivity descriptors were calculated to predict the compounds' chemical reactivity and kinetic stability. Molecular Electrostatic Potential (MEP) surfaces were employed to visualize charge distribution and identify potential reactive sites within the molecules. Likewise, simulated infrared (IR) and NMR spectra from DFT calculations were consistent with their experimental counterparts. Additionally, experimental infrared (IR) and nuclear magnetic resonance (NMR) spectra were compared with their corresponding simulated spectra derived from DFT calculations, demonstrating strong consistency. In addition, the non-linear optical (NLO) properties of the compounds were assessed and found to exceed those of urea, a standard reference compound. SwissADME analysis was also performed to evaluate the pharmacokinetic profiles and drug-likeness of the synthesized molecules. Finally, molecular docking studies targeting cyclin-dependent kinases CDK1 (PDB ID: 4Y72) were conducted to explore the binding interactions of the synthesized compounds. The docking results showed a strong correlation with their observed anticancer activities, supporting that these compounds are promising candidates for HepG-2 cell lines.
1. Introduction
Naturally occurring chromones are mostly isolated from numerous families of plants including Cucurbitaceae, Rutaceae and Gramineae1,2 Chromones are widely used in the medical industry.3,4 They have a variety of biological applications including anti-Alzheimer's,5 antimicrobial,6 anticancer7 anti-inflammatory,8 antimalarial,9 antioxidant,10 as well as α-glucosidase inhibitors.11 Chromones represent a versatile class of oxygen-containing heterocycles and serve as key precursors in the construction of diverse bioactive heterocyclic frameworks. Their synthetic utility arises primarily from the electrophilic character of the γ-pyrone ring system, which facilitates nucleophilic addition and subsequent heterocyclization reactions.12–16 The electron-deficient C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–C
C–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O motif within the chromone core provides an activated platform for ring activation and transformation under both basic and nucleophilic conditions. Previous studies by Ibrahim et al.17 comprehensively outlined the physicochemical properties and reactivity patterns of 2-methylchromones, highlighting the presence of two principal electrophilic centers; C-2 and C-4. Among these, the C-2 position is significantly more susceptible to nucleophilic attack owing to π-electron withdrawal by the adjacent carbonyl group. Such nucleophilic addition at C-2 is typically followed by opening of the γ-pyrone ring, generating reactive intermediates capable of undergoing intramolecular cyclization or further condensation steps to afford structurally diverse heterocycles.18 These distinctive reactivity characteristics make chromones highly versatile synthetic intermediates for constructing nitrogen containing heterocyclic scaffolds with significant pharmacological and biological relevance.
O motif within the chromone core provides an activated platform for ring activation and transformation under both basic and nucleophilic conditions. Previous studies by Ibrahim et al.17 comprehensively outlined the physicochemical properties and reactivity patterns of 2-methylchromones, highlighting the presence of two principal electrophilic centers; C-2 and C-4. Among these, the C-2 position is significantly more susceptible to nucleophilic attack owing to π-electron withdrawal by the adjacent carbonyl group. Such nucleophilic addition at C-2 is typically followed by opening of the γ-pyrone ring, generating reactive intermediates capable of undergoing intramolecular cyclization or further condensation steps to afford structurally diverse heterocycles.18 These distinctive reactivity characteristics make chromones highly versatile synthetic intermediates for constructing nitrogen containing heterocyclic scaffolds with significant pharmacological and biological relevance.
On the other hand, molecular modeling using Density Functional Theory (DFT) approaches attracted a lot of attention in recent years.19–21 Where, it aims to explore how a molecule's chemical and physical properties are connected to its chemical composition and three-dimensional (3D) structure.22–24
This study investigates the reactivity of 2,6-dimethylchromone (1) toward selected heterocyclic hydrazines and cyclic enamines to construct novel pyrazoles and fused pyrimidines, followed by evaluation of their cytotoxic effects against HepG-2 liver cancer cells which represent one of the most common cancers worldwide and a leading cause of cancer-related mortality.25,26 DFT calculations (B3LYP/6-311++G(d,p)) were performed to analyze electronic structures, global reactivity descriptors, MEP surfaces, and NLO properties. Theoretical 1H and 13C NMR (GIAO) and FT-IR data were correlated with experimental spectra, and molecular docking studies were conducted to further support the biological evaluation.
2. Experimental
Melting points were recorded on a Stuart SMP3 apparatus. Elemental analyses (C, H, N) were performed using a PerkinElmer CHN-2400 analyzer. FT-IR spectra (KBr) were obtained using a Nicolet iS10 spectrophotometer. Mass spectra were acquired on a Shimadzu GC-2010 GC-MS (EI, 70 eV). 1H (300 MHz) and 13C (75 MHz) NMR spectra were recorded on a Mercury-300BB instrument in DMSO-d6 with TMS as the internal standard. Compound purity was monitored by TLC. All chemicals including 2-hydroxyacetophenone, hydrazine hydrate, 4,7-dichloroquinoline, benzil, thiosemicarbazide, 5-amino-3-methyl-1H-pyrazole, 6-aminopyrimidine-2,4(1H,3H)-dione, 6-amino-2-thioxo-2,3-dihydropyrimidin-4(1H)-one, and 6-amino-1,3-dimethylpyrimidine-2,4(1H,3H)-dione were purchased from Merck and used without further purification.2.1. Synthesis and characterization of compounds
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1575 (C
N), 1575 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.15 (s, 3H, CH3), 2.31 (s, 3H, CH3), 6.92–7.09 (m, 2H, Ar–H), 7.24 (d, 1H, Ar–H, J = 7.5 Hz), 7.44 (d, 1H, Ar–H, J = 7.5 Hz), 7.69 (s, 1H, Ar–H), 7.78 (s, 1H, Ar–H), 8.02 (d, 1H, H-3quinoline, J = 8.1 Hz), 8.17 (s, 1H, H-4pyrrazole), 8.52 (d, 1H, H-2quinoline, J = 8.1 Hz), 10.76 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 18.5 (CH3), 22.1 (CH3), 103.2 (C-4pyrazole), 117.5 (C-3quinoline), 120.9 (Ar–C), 123.6 (Ar–C), 124.7 (Ar–C), 125.2 (Ar–C), 125.8 (Ar–C), 127.6 (Ar–C), 128.4 (Ar–C), 129.1 (Ar–C), 130.5 (Ar–C), 133.4 (Ar–C), 134.7 (Ar–C), 142.0 (C-3pyrazole), 143.5 (C-2quinoline), 144.3 (C-8a quinoline), 145.9 (C-5pyrazole), 151.7 (C–OH). Mass spectrum (m/z, I %): 349/351 (M+/M + 2; 100/33), 319/321 (21/7), 387/389 (48/16), 162/164 (45/15), 107 (44), 94 (57), 77 (31), 65 (16). Anal. Calcd for C20H16ClN3O (349.81): C, 68.67; H, 4.61; N, 12.01%. Found: C, C, 68.48; H, 4.52; N, 11.87%.
C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.15 (s, 3H, CH3), 2.31 (s, 3H, CH3), 6.92–7.09 (m, 2H, Ar–H), 7.24 (d, 1H, Ar–H, J = 7.5 Hz), 7.44 (d, 1H, Ar–H, J = 7.5 Hz), 7.69 (s, 1H, Ar–H), 7.78 (s, 1H, Ar–H), 8.02 (d, 1H, H-3quinoline, J = 8.1 Hz), 8.17 (s, 1H, H-4pyrrazole), 8.52 (d, 1H, H-2quinoline, J = 8.1 Hz), 10.76 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 18.5 (CH3), 22.1 (CH3), 103.2 (C-4pyrazole), 117.5 (C-3quinoline), 120.9 (Ar–C), 123.6 (Ar–C), 124.7 (Ar–C), 125.2 (Ar–C), 125.8 (Ar–C), 127.6 (Ar–C), 128.4 (Ar–C), 129.1 (Ar–C), 130.5 (Ar–C), 133.4 (Ar–C), 134.7 (Ar–C), 142.0 (C-3pyrazole), 143.5 (C-2quinoline), 144.3 (C-8a quinoline), 145.9 (C-5pyrazole), 151.7 (C–OH). Mass spectrum (m/z, I %): 349/351 (M+/M + 2; 100/33), 319/321 (21/7), 387/389 (48/16), 162/164 (45/15), 107 (44), 94 (57), 77 (31), 65 (16). Anal. Calcd for C20H16ClN3O (349.81): C, 68.67; H, 4.61; N, 12.01%. Found: C, C, 68.48; H, 4.52; N, 11.87%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1585 (C
N), 1585 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.18 (s, 3H, CH3), 2.30 (s, 3H, CH3), 7.04 (d, 1H, Ar–H, J = 7.5 Hz), 7.16–7.20 (m, 5H, Ar–H), 7.36–7.41 (m, 5H, Ar–H), 7.56 (d, 1H, Ar–H, J = 7.5 Hz), 7.84 (s, 1H, Ar–H), 8.29 (s, 1H, H-4pyrrazole), 10.96 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 17.8 (CH3), 22.3 (CH3), 104.1 (C-4pyrazole), 121.4 (Ar–C), 124.1 (Ar–C), 124.6 (Ar–C), 125.2 (Ar–C), 125.7 (Ar–C), 126.2 (Ar–C), 126.7 (Ar–C), 127.1 (Ar–C), 127.5 (Ar–C), 127.9 (Ar–C), 128.3 (Ar–C), 128.6 (Ar–C), 129.2 (Ar–C), 129.5 (Ar–C), 130.1 (Ar–C), 130.5 (Ar–C), 131.9 (Ar–C), 141.4 (C-3pyrazole), 142.2 (C-6triazine), 142.7 (C-5 triazine), 143.8 (C-3triazine), 144.1 (C-5pyrazole), 150.8 (C–OH).Mass spectrum (m/z, I %): 419 (67), 241 (45), 213 (25), 178 (100), 132 (26), 108 (13), 91 (56), 77 (39), 65 (15). Anal. Calcd for C26H21N5O (419.48): C, 74.44; H, 5.05; N, 16.70%. Found: C, 74.25; H, 4.96; N, 16.54%.
C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.18 (s, 3H, CH3), 2.30 (s, 3H, CH3), 7.04 (d, 1H, Ar–H, J = 7.5 Hz), 7.16–7.20 (m, 5H, Ar–H), 7.36–7.41 (m, 5H, Ar–H), 7.56 (d, 1H, Ar–H, J = 7.5 Hz), 7.84 (s, 1H, Ar–H), 8.29 (s, 1H, H-4pyrrazole), 10.96 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 17.8 (CH3), 22.3 (CH3), 104.1 (C-4pyrazole), 121.4 (Ar–C), 124.1 (Ar–C), 124.6 (Ar–C), 125.2 (Ar–C), 125.7 (Ar–C), 126.2 (Ar–C), 126.7 (Ar–C), 127.1 (Ar–C), 127.5 (Ar–C), 127.9 (Ar–C), 128.3 (Ar–C), 128.6 (Ar–C), 129.2 (Ar–C), 129.5 (Ar–C), 130.1 (Ar–C), 130.5 (Ar–C), 131.9 (Ar–C), 141.4 (C-3pyrazole), 142.2 (C-6triazine), 142.7 (C-5 triazine), 143.8 (C-3triazine), 144.1 (C-5pyrazole), 150.8 (C–OH).Mass spectrum (m/z, I %): 419 (67), 241 (45), 213 (25), 178 (100), 132 (26), 108 (13), 91 (56), 77 (39), 65 (15). Anal. Calcd for C26H21N5O (419.48): C, 74.44; H, 5.05; N, 16.70%. Found: C, 74.25; H, 4.96; N, 16.54%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1566 (C
N), 1566 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.11 (s, 3H, CH3), 2.27 (s, 3H, CH3), 2.40 (s, 3H, CH3), 7.19 (d, 1H, Ar–H, J = 7.2 Hz), 7.44 (d, 1H, Ar–H, J = 7.2 Hz), 7.72 (s, 1H, Ar–H), 8.27 (s, 1H, H-3pyridine), 10.82 (bs, 1H, OH exchangeable with D2O), 11.13 (bs, 1H, NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.7 (CH3), 17.5 (CH3), 22.6 (CH3), 107.1 (C-3a), 122.4 (Ar–C), 124.3 (Ar–C), 126.1 (Ar–C), 128.3 (Ar–C), 130.4 (Ar–C), 132.3 (C-5), 140.0 (C-4), 142.3 (C-3), 144.7 (C-6), 148.3 (C-7a), 151.4 (C–OH). Mass spectrum (m/z, I %): 253 (57), 212 (32), 168 (42), 132 (21), 108 (100), 94 (62), 91 (43), 77 (51), 65 (31). Anal. Calcd for C15H15N3O (253.29): C, 71.13; H, 5.97; N, 16.59%. Found: C, 71.00; H, 5.85; N, 16.38%.
C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.11 (s, 3H, CH3), 2.27 (s, 3H, CH3), 2.40 (s, 3H, CH3), 7.19 (d, 1H, Ar–H, J = 7.2 Hz), 7.44 (d, 1H, Ar–H, J = 7.2 Hz), 7.72 (s, 1H, Ar–H), 8.27 (s, 1H, H-3pyridine), 10.82 (bs, 1H, OH exchangeable with D2O), 11.13 (bs, 1H, NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.7 (CH3), 17.5 (CH3), 22.6 (CH3), 107.1 (C-3a), 122.4 (Ar–C), 124.3 (Ar–C), 126.1 (Ar–C), 128.3 (Ar–C), 130.4 (Ar–C), 132.3 (C-5), 140.0 (C-4), 142.3 (C-3), 144.7 (C-6), 148.3 (C-7a), 151.4 (C–OH). Mass spectrum (m/z, I %): 253 (57), 212 (32), 168 (42), 132 (21), 108 (100), 94 (62), 91 (43), 77 (51), 65 (31). Anal. Calcd for C15H15N3O (253.29): C, 71.13; H, 5.97; N, 16.59%. Found: C, 71.00; H, 5.85; N, 16.38%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine), 1620 (C
Opyrimidine), 1620 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1590 (C
N), 1590 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.24 (s, 3H, CH3), 2.38 (s, 3H, CH3), 7.08 (d, 1H, Ar–H, J = 7.2 Hz), 7.44 (d, 1H, Ar–H, J = 7.2 Hz), 7.74 (s, 1H, Ar–H), 8.13 (s, 1H, H-3pyridine), 10.78 (bs, 1H, OH exchangeable with D2O), 11.40 (bs, 2H, 2NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 17.6 (CH3), 21.3 (CH3), 109.5 (C-4a), 121.3 (Ar–C), 124.9 (Ar–C), 126.8 (Ar–C), 129.7 (Ar–C), 130.3 (Ar–C), 132.3 (C-6), 141.2 (C-5), 146.0 (C-7), 147.3 (C-8a), 151.2 (C–OH), 166.0, 169.1 (2C
C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.24 (s, 3H, CH3), 2.38 (s, 3H, CH3), 7.08 (d, 1H, Ar–H, J = 7.2 Hz), 7.44 (d, 1H, Ar–H, J = 7.2 Hz), 7.74 (s, 1H, Ar–H), 8.13 (s, 1H, H-3pyridine), 10.78 (bs, 1H, OH exchangeable with D2O), 11.40 (bs, 2H, 2NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 17.6 (CH3), 21.3 (CH3), 109.5 (C-4a), 121.3 (Ar–C), 124.9 (Ar–C), 126.8 (Ar–C), 129.7 (Ar–C), 130.3 (Ar–C), 132.3 (C-6), 141.2 (C-5), 146.0 (C-7), 147.3 (C-8a), 151.2 (C–OH), 166.0, 169.1 (2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine). Mass spectrum (m/z, I %): 283 (100), 255 (46), 226 (29), 198 (21), 158 (33), 131 (19), 107 (68), 91 (50), 77 (28), 65 (17). Anal. Calcd for C15H13N3O3 (283.28): C, 63.60; H, 4.63; N, 14.83%. Found: C, 63.46; H, 4.50; N, 14.69%.
Opyrimidine). Mass spectrum (m/z, I %): 283 (100), 255 (46), 226 (29), 198 (21), 158 (33), 131 (19), 107 (68), 91 (50), 77 (28), 65 (17). Anal. Calcd for C15H13N3O3 (283.28): C, 63.60; H, 4.63; N, 14.83%. Found: C, 63.46; H, 4.50; N, 14.69%.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine), 1615 (C
Opyrimidine), 1615 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1593 (C
N), 1593 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 1252 (C
C), 1252 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.28 (s, 3H, CH3), 2.40 (s, 3H, CH3), 7.43 (d, 1H, Ar–H, J = 7.8 Hz), 7.62 (d, 1H, Ar–H, J = 7.8 Hz), 7.86 (s, 1H, Ar–H), 8.28 (s, 1H, H-3pyridine), 11.09 (bs, 1H, OH exchangeable with D2O), 11.33 (bs, 2H, 2NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.5 (CH3), 21.3 (CH3), 109.5 (C-4a), 122.1 (Ar–C), 125.0 (Ar–C), 126.2 (Ar–C), 128.6 (Ar–C), 130.5 (Ar–C), 131.8 (C-6), 141.1 (C-5), 146.5 (C-7), 147.7 (C-8a), 150.4 (C–OH), 168.3 (C
S). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.28 (s, 3H, CH3), 2.40 (s, 3H, CH3), 7.43 (d, 1H, Ar–H, J = 7.8 Hz), 7.62 (d, 1H, Ar–H, J = 7.8 Hz), 7.86 (s, 1H, Ar–H), 8.28 (s, 1H, H-3pyridine), 11.09 (bs, 1H, OH exchangeable with D2O), 11.33 (bs, 2H, 2NH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.5 (CH3), 21.3 (CH3), 109.5 (C-4a), 122.1 (Ar–C), 125.0 (Ar–C), 126.2 (Ar–C), 128.6 (Ar–C), 130.5 (Ar–C), 131.8 (C-6), 141.1 (C-5), 146.5 (C-7), 147.7 (C-8a), 150.4 (C–OH), 168.3 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine), 186.2 (C
Opyrimidine), 186.2 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S). Mass spectrum (m/z, I %): 299 (M+; 100), 285 (41), 271 (26), 227 (42), 214 (19), 193 (36), 164 (25), 148 (19), 132 (17), 117 (21), 108 (79), 93 (62), 77 (34), 65 (17). Anal. Calcd for C15H13N3O2S (299.35): C, 60.18; H, 4.38; N, 14.04; S, 10.71%. Found: C, 60.03; H, 4.35; N, 13.86; S, 10.55%.
S). Mass spectrum (m/z, I %): 299 (M+; 100), 285 (41), 271 (26), 227 (42), 214 (19), 193 (36), 164 (25), 148 (19), 132 (17), 117 (21), 108 (79), 93 (62), 77 (34), 65 (17). Anal. Calcd for C15H13N3O2S (299.35): C, 60.18; H, 4.38; N, 14.04; S, 10.71%. Found: C, 60.03; H, 4.35; N, 13.86; S, 10.55%.
2.1.5.1 7-(2-Hydroxy-5-methylphenyl)-1,3,5-trimethylpyrido[2,3-d]pyrimidine-2,4(1H,3H)-dione (9). A mixture of 2,6-dimethylchromone (1) (0.35 g, 2 mmol) and 6-amino-1,3-dimethylpyrimidine-2,4(1H,3H)-dione (0.31 g, 2 mmol) in sodium ethoxide (prepared by adding 0.2 g sodium in 15 mL absolute ethanol) was heated under reflux for 2 h. After cooling, the reaction mixture was poured onto crushed ice (50 g) and neutralized with conc. HCl. The solid obtained filtered and crystallized from DMF/H2O, mp > 300 °C, yield (0.44 g, 71%). IR (KBr, cm−1): 3415 (OH), 3036 (CHarom), 2963, 2934 (CHaliph), 1682, 1663 (2C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine), 1617 (C
Opyrimidine), 1617 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1586 (C
N), 1586 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.20 (s, 3H, CH3), 2.35 (s, 3H, CH3), 3.26 (s, 3H, NCH3), 3.52 (s, 3H, NCH3), 7.53 (d, 1H, Ar–H, J = 8.1 Hz), 7.73 (d, 1H, Ar–H, J = 8.1 Hz), 7.91 (s, 1H, Ar–H), 8.43 (s, 1H, H-3pyridine), 11.54 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.2 (CH3), 20.8 (CH3), 28.4 (NCH3), 30.1 (NCH3), 109.3 (C-4a), 122.4 (Ar–C), 124.2 (Ar–C), 126.5 (Ar–C), 129.8 (Ar–C), 130.9 (Ar–C), 132.5 (C-6), 140.8 (C-5), 146.3 (C-7), 147.4 (C-8a), 150.9 (C–OH), 167.6, 170.3 (2C
C). 1H-NMR (DMSO-d6, δ, 300 MHz): 2.20 (s, 3H, CH3), 2.35 (s, 3H, CH3), 3.26 (s, 3H, NCH3), 3.52 (s, 3H, NCH3), 7.53 (d, 1H, Ar–H, J = 8.1 Hz), 7.73 (d, 1H, Ar–H, J = 8.1 Hz), 7.91 (s, 1H, Ar–H), 8.43 (s, 1H, H-3pyridine), 11.54 (bs, 1H, OH exchangeable with D2O). 13C-NMR (DMSO-d6, δ, 75 MHz): 16.2 (CH3), 20.8 (CH3), 28.4 (NCH3), 30.1 (NCH3), 109.3 (C-4a), 122.4 (Ar–C), 124.2 (Ar–C), 126.5 (Ar–C), 129.8 (Ar–C), 130.9 (Ar–C), 132.5 (C-6), 140.8 (C-5), 146.3 (C-7), 147.4 (C-8a), 150.9 (C–OH), 167.6, 170.3 (2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine). Mass spectrum (m/z, I %): 311 (M+; 100), 283 (60), 269 (41), 241 (28), 226 (33), 205 (29), 177 (53), 150 (21), 121 (34), 108 (17), 94 (51), 77 (52), 64 (21). Anal. Calcd for C17H17N3O3 (311.34): C, 65.58; H, 5.50; N, 13.50%. Found: C, 65.52; H, 5.43; N, 13.39%.
Opyrimidine). Mass spectrum (m/z, I %): 311 (M+; 100), 283 (60), 269 (41), 241 (28), 226 (33), 205 (29), 177 (53), 150 (21), 121 (34), 108 (17), 94 (51), 77 (52), 64 (21). Anal. Calcd for C17H17N3O3 (311.34): C, 65.58; H, 5.50; N, 13.50%. Found: C, 65.52; H, 5.43; N, 13.39%.
2.2. Computational details
Density functional theory (DFT) calculations were performed to obtain reliable insights into the structural and electronic properties of the synthesized compounds. All calculations were carried out using the B3LYP functional in combination with the 6-311++G(d,p) basis set as implemented in GAUSSIAN 09W, a well-established package for quantum chemical simulations.27–29 Geometry optimizations were conducted without imposing symmetry constraints, and the resulting structures were confirmed as true minima by the absence of imaginary frequencies. Molecular visualization and analysis were performed using GaussView 5.0.30,31A series of quantum-chemical descriptors were investigated, including optimized geometries, molecular electrostatic potential (MEP) maps, and frontier molecular orbital (HOMO–LUMO) energies. In addition, vibrational frequency calculations were carried out to support experimental IR spectra. 1H and 13C NMR chemical shifts were predicted using the GIAO approach with the same basis set, and the computed values were compared with the corresponding experimental data to verify structural assignments.32 Furthermore, key nonlinear-optical (NLO) parameters, such as dipole moment, polarizability, and first hyperpolarizability, were evaluated to provide deeper insight into the electronic characteristics of the molecules.
2.3. Biological evaluation
3. Results and discussion
3.1. Characterization of the synthesized compounds
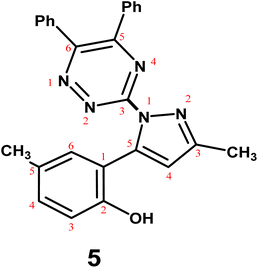
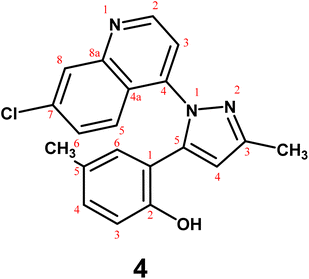
The present study aimed to investigate the reactivity of 2,6-dimethylchromone (1) with selected heterocyclic hydrazines and cyclic enamines, and to identify the resulting products using comprehensive spectroscopic and computational analyses. Hence, reaction of substrate 1 with 7-chloro-4-hydrazinylquinoline (2)47 in sodium ethoxide solution, afforded quinolinylpyrazole derivative 4; in 73% yield (Scheme 1). The reaction proceeds through a nucleophilic attack at the C-2 position of the chromone ring, followed by γ-pyrone ring opening to generate intermediate A, which subsequently undergoes cyclization to afford the pyrazole moiety. Similarly, triazinylpyrazole 5 was synthesized from reaction of compound 1 with 3-hydrazinyl-5,6-diphenyl-1,2,4-triazine (3)48 (Scheme 1). The mass spectra of quinolinylpyrazole 4 and triazinylpyrrazole 5 presented the molecular formula weights at m/z 349 and 419, supporting the proposed molecular formulae C20H16ClN3O and C26H21N5O, respectively. The IR spectra of pyrazoles 4 and 5 showed typical absorption bands assigned to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N at ṽ 1613 and 1608 cm−1, respectively. The pyrazole ring proton was seen as specific singlet in the 1H-NMR spectra for compounds 4 and 5 at δ 8.17 and 8.29, respectively. The spectrum of compound 4 presented two definite doublets due to H-3quinoline and H-2quinoline at δ 8.02 and 8.52, respectively.
N at ṽ 1613 and 1608 cm−1, respectively. The pyrazole ring proton was seen as specific singlet in the 1H-NMR spectra for compounds 4 and 5 at δ 8.17 and 8.29, respectively. The spectrum of compound 4 presented two definite doublets due to H-3quinoline and H-2quinoline at δ 8.02 and 8.52, respectively.
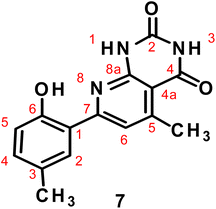
On the other hand, reaction of compound 1 with 5-amino-3-methyl-1H-pyrazole in sodium ethoxide under reflux afforded pyrazolo[3,4-b]pyridine 6 (Scheme 2).49 This reaction occurs through γ-pyrone ring opening by the activated C-4pyrrazole (intermediate B) with concomitant cyclization via intermediate C to produce the final product 6 (Scheme 2). In the same manner, boiling compound 1 with 6-aminopyrimidine-2,4(1H,3H)-dione in sodium ethoxide yielded pyrido[2,3-d]pyrimidine 7 (Scheme 2).47 The mass spectra of compound 6 and 7 displayed the parent ion peaks at m/z 253 and 283; supporting the proposed molecular formulas C15H15N3O and C15H13N3O3, respectively. The 1H NMR spectrum of compound 6 recorded three upfield singlet signals assigned to three methyl groups at δ 2.11, 2.27 and 2.40, as well as singlet signal due to H-3pyridine at δ 8.27. The spectrum also displayed D2O exchangeable signal at δ 10.82 (OH) and 11.13 (NH). The 1H NMR spectrum of compound 7 recorded two upfield singlet signals assigned to two methyl groups at δ 2.24 and 2.38, in addition to characteristic singlet due to H-3pyridine at δ 8.13. D2O exchangeable signals were observed at δ 10.78 (OH) and 11.40 (2NH). The 13C-NMR spectrum of compound 7 displayed specific signals at δ 17.6 (CH3), 21.3 (CH3), 151.2 (C–OH), 166.0 and 169.1 (2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine).
Opyrimidine).
Further, treatment of substrate 1 with 6-aminothiouracil and 1,3-dimethyl-6-aminouracil, in boiling ethanol containing sodium ethoxide, furnished the novel pyrido[2,3-d]pyrimidines 8 and 9, respectively. Structures 8 and 9 were supported by the mass spectra which presented the parent ion peaks, as the base peaks, at m/z 299 and 311. Singlet signals attributed to H-3pyridine were seen in the 1H NMR spectra of compounds 8 and 9 at δ 8.28 and 8.43, respectively. The 2NCH3 protons in compound 9 were observed at δ 3.26 and 3.52 (Scheme 3).
3.2. Theoretical studies
Additionally, various molecular properties, including chemical potential (Pi), electronegativity (χ), chemical hardness (η), softness (S), electrophilicity (ω), nucleophilicity (ε), and maximum additional electronic charge (ΔN), were calculated using DFT at the B3LYP/6-311++G(d,p) level, as outlined in the following equations:52–55
| I = −EHOMO | (1) |
| A = −ELUMO | (2) |
 | (3) |
 | (4) |
 | (5) |
 | (6) |
 | (7) |
 | (8) |
 | (9) |
Table 1 summarizes the calculated frontier molecular orbital (FMO) parameters for the synthesized compounds The HOMO energy (EHOMO) indicates a molecule's ability to donate electrons and is correlated with its ionization potential, while the LUMO energy (ELUMO) reflects its electron-accepting capacity and is directly related to electron affinity.56 The HOMO–LUMO energy gap is a key factor influencing a molecule's kinetic stability and chemical reactivity.57,58 Where, a smaller frontier orbital gap typically indicates higher chemical reactivity and lower kinetic stability, as less energy is needed to promote an electron from the HOMO to the LUMO.58 The energy gap (ΔE) for all compounds was found to range between 3.534 and 4.894 eV.
| Compound no. | ET (au) | HOMO (au) | LUMO (au) | E (HOMO) | E (LUMO) | I (eV) | A (eV) | ΔE (eV) | χ (eV) | Pi (eV)< | η (eV) | S (eV−1) | ω (eV) | ε (eV−1) | ΔN |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | −575.81 | −0.245 | −0.065 | −6.655 | −1.761 | 6.655 | 1.761 | 4.894 | 4.208 | −4.208 | 2.447 | 0.409 | 3.618 | 0.276 | 1.720 |
| 4 | −1471.71 | −0.229 | −0.077 | −6.226 | −2.094 | 6.226 | 2.094 | 4.132 | 4.160 | −4.160 | 2.066 | 0.484 | 4.188 | 0.239 | 2.013 |
| 5 | −1352.68 | −0.215 | −0.085 | −5.851 | −2.317 | 5.851 | 2.317 | 3.534 | 4.084 | −4.084 | 1.767 | 0.566 | 4.721 | 0.212 | 2.312 |
| 6 | −820.32 | −0.217 | −0.053 | −5.894 | −1.444 | 5.894 | 1.444 | 4.450 | 3.669 | −3.669 | 2.225 | 0.449 | 3.025 | 0.331 | 1.649 |
| 7 | −969.69 | −0.224 | −0.087 | −6.096 | −2.365 | 6.096 | 2.365 | 3.731 | 4.230 | −4.230 | 1.865 | 0.536 | 4.796 | 0.208 | 2.268 |
| 8 | −1292.65 | −0.228 | −0.093 | −6.192 | −2.535 | 6.192 | 2.535 | 3.657 | 4.363 | −4.363 | 1.828 | 0.547 | 5.207 | 0.192 | 2.386 |
| 9 | −1048.33 | −0.221 | −0.082 | −5.998 | −2.229 | 5.998 | 2.229 | 3.768 | 4.113 | −4.113 | 1.884 | 0.531 | 4.490 | 0.223 | 2.183 |
Furthermore, a reduced HOMO–LUMO separation is typically associated with enhanced molecular softness and higher polarizability,59 whereas larger energy gaps correspond to increased hardness and greater thermodynamic stability. In agreement with this principle, compound 5 shows the highest softness (S = 0.566 eV−1), while compound 1 demonstrates the maximum hardness (η = 2.447 eV), reflecting its greater resistance toward electronic deformation.
Additionally, the chemical potential (Pi) reflects the tendency of an electron to escape from a chemical system,60 whereas electronegativity (χ) quantifies a molecule's affinity for acquiring electrons.61 The calculated chemical potential values for the investigated compounds range from −3.669 to −4.363 eV (Table 1), indicating limited propensity for electron loss and a favorable tendency toward electron acceptance. Notably, compound 8 exhibits the highest electronegativity (χ = 4.363 eV), identifying it as the most efficient electron-accepting species in the series.
Further, the electrophilicity index (ω), introduced by Parr et al.,62 quantifies the energy stabilization upon accepting additional charge (ΔN). It servs as a measure of a molecule's electrophilic character.63 High values of ω and ΔN suggest strong electrophilic behavior; thus, compound 8 (ω = 5.207 eV, ΔN = 2.386) stands out as the most electrophilic and electronegative. Conversely, strong nucleophiles are characterized by low electrophilicity (ω) and high nucleophilicity (ε), making compound 6 (ε = 0.331 eV−1) the most nucleophilic among the current compounds. Fig. 2 represents a summary of the global reactivity indices for the present compounds 1 and 4–9.
As illustrated in Fig. 3, S3 and S4, the blue regions are predominantly localized around hydrogen atoms, particularly those attached to oxygen and nitrogen atoms, as well as certain carbon atoms influenced by the electron withdrawing inductive or mesomeric effects of adjacent oxygen or nitrogen atoms—highlighting electrophilic sites. In contrast, the red regions are primarily distributed around electronegative oxygen and nitrogen atoms, along with carbon atoms affected by the electron-donating hydroxyl group on the benzene ring in the product after ring opening—indicating nucleophilic centers.
| Compound 4 | Compound 5 | Compound 6 | ||||||
|---|---|---|---|---|---|---|---|---|
| υexp (cm−1) | υthe (cm−1) | Assignment | υexp (cm−1) | υthe (cm−1) | Assignment | υexp (cm−1) | υthe (cm−1) | Assignment |
| 3328 | 3379 | OH | 3346 | 3397 | OH | 3424 | 3444 | OH |
| 3058 | 3096 | CHarom | 2937 | 2992 | CHaliph | 3218 | 3272, 3256 | 2NH |
| 2945 | 3024 | CHaliph | 1608 | 1624 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
2918 | 3016 | CHaliph |
| 1613 | 1616 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
1585 | 1590 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
1616 | 1624 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
| 1575 | 1595 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
1566 | 1600 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
|||
| Compound 7 | Compound 8 | Compound 9 | ||||||
|---|---|---|---|---|---|---|---|---|
| υexp (cm−1) | υthe (cm−1) | Assignment | υexp (cm−1) | υthe (cm−1) | Assignment | υexp (cm−1) | υthe (cm−1) | Assignment |
| 3409 | 3430 | OH | 3420 | 3456 | OH | 3415 | 3428 | OH |
| 3149 | 3205, 3181 | 2NH | 3234, 3198 | 3260, 3192 | 2NH | 3036 | 3108 | CHarom |
| 3051 | 3067 | CHarom | 3065 | 3096 | CHarom | 2963, 2934 | 3044, 2980 | CHaliph |
| 2977 | 3010 | CHaliph | 2975, 2949 | 3016, 2992 | CHaliph | 1682, 1663 | 1700, 1676 | 2C = Opyrimidine |
| 1685 | 1705, 1690 | 2C = Opyrimidine | 1667 | 1696, 1680 | 2C = Opyrimidine | 1617 | 1620 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
| 1620 | 1624 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
1615 | 1620 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N N |
1586 | 1598 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
| 1590 | 1609 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
1593 | 1600 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C C |
|||
| 1252 | 1264 | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S S |
||||||
General features were observed in the IR spectra of the synthesized compounds 4–9. The phenolic O–H stretching vibrations (νO–H) appeared experimentally in the range 3328–3420 cm−1, while the corresponding calculated values were found at 3379–3456 cm−1. Additionally, the N–H stretching vibrations (νN–H) were recorded experimentally in the range 3149–3234 cm−1, with the calculated wavenumbers appearing at 3181–3272 cm−1. The aromatic C–H stretching vibrations (νC–H) were observed experimentally at 3036–3065 cm−1, whereas the computed values ranged from 3067–3108 cm−1. Similarly, the aliphatic C–H stretching vibrations were detected experimentally in the range 2918–2977 cm−1, while the corresponding calculated values were located at 2980–3044 cm−1.
Moreover, the azomethine C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching vibrations (νC
N stretching vibrations (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) were observed experimentally in the range 1608–1620 cm−1, showing good agreement with the calculated values of 1616–1624 cm−1. The absorption bands attributed to C
N) were observed experimentally in the range 1608–1620 cm−1, showing good agreement with the calculated values of 1616–1624 cm−1. The absorption bands attributed to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretching vibrations (νC
C stretching vibrations (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) were recorded experimentally at 1566–1693 cm−1, while the computed values were found in the range 1590–1609 cm−1. For compounds 7–9, the carbonyl stretching vibrations of the pyrimidine moiety (νC
C) were recorded experimentally at 1566–1693 cm−1, while the computed values were found in the range 1590–1609 cm−1. For compounds 7–9, the carbonyl stretching vibrations of the pyrimidine moiety (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) were experimentally observed at 1663–1685 cm−1, with the corresponding calculated values ranging from 1676–1705 cm−1. Furthermore, the C
O) were experimentally observed at 1663–1685 cm−1, with the corresponding calculated values ranging from 1676–1705 cm−1. Furthermore, the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S stretching vibration (νC
S stretching vibration (νC![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S) of the pyrimidine moiety in compound 8 was detected experimentally at 1252 cm−1, in good agreement with the calculated value of 1264 cm−1.
S) of the pyrimidine moiety in compound 8 was detected experimentally at 1252 cm−1, in good agreement with the calculated value of 1264 cm−1.
To evaluate the agreement between experimentally observed and theoretically calculated wavenumbers of the functional groups, correlation graphs are presented in Fig. S5–S7. These graphs exhibit a high correlation coefficient (R2 = 0.99), indicating excellent agreement between experimental and theoretical values, and confirming the reliability of the computational prediction technique.
| Compound 4 | Compound 5 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 33-H | CH3 | 1.312539 | 2.15 | 19-H | CH3 | 1.614114 | 2.18 |
| 34-H | CH3 | 2.016742 | 2.15 | 21-H | CH3 | 2.291107 | 2.18 |
| 35-H | CH3 | 2.083407 | 2.15 | 20-H | CH3 | 2.292911 | 2.18 |
| 39-H | CH3 | 2.294007 | 2.31 | 25-H | CH3 | 2.355586 | 2.30 |
| 38-H | CH3 | 2.437161 | 2.31 | 23-H | CH3 | 2.458817 | 2.30 |
| 37-H | CH3 | 2.485912 | 2.31 | 24-H | CH3 | 2.463472 | 2.30 |
| 31-H | Ar–H | 7.121756 | 6.92 | 17-H | Ar–H | 7.174151 | 7.04 |
| 30-H | Ar–H | 7.306446 | 7.09 | 15-H | Ar–H | 6.847266 | 7.16 |
| 25-H | Ar–H | 7.605370 | 7.24 | 49-H | Ar–H | 7.267616 | 7.17 |
| 21-H | Ar–H | 7.646732 | 7.44 | 41-H | Ar–H | 7.331977 | 7.18 |
| 29-H | Ar–H | 7.733079 | 7.69 | 52-H | Ar–H | 7.351159 | 7.19 |
| 26-H | Ar–H | 8.011164 | 7.78 | 38-H | Ar–H | 7.386973 | 7.20 |
| 19-H | H-3quinoline | 8.080045 | 8.02 | 16-H | Ar–H | 7.414939 | 7.36 |
| 41-H | H-4pyrrazole | 8.200833 | 8.17 | 53-H | Ar–H | 7.650476 | 7.37 |
| 23-H | H-2quinoline | 8.780292 | 8.52 | 42-H | Ar–H | 7.720835 | 7.38 |
| 40-H | OH | 10.67650 | 10.76 | 51-H | Ar–H | 7.774681 | 7.39 |
| 40-H | Ar–H | 7.839250 | 7.41 | ||||
| 47-H | Ar–H | 8.065967 | 7.56 | ||||
| 36-H | Ar–H | 8.121905 | 7.84 | ||||
| 27-H | H-4pyrrazole | 8.607195 | 8.29 | ||||
| 26-H | OH | 11.36753 | 10.96 | ||||
| Compound 6 | Compound 7 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 29-H | CH3 | 2.138881 | 2.11 | 28-H | CH3 | 1.990108 | 2.24 |
| 27-H | CH3 | 2.142194 | 2.11 | 27-H | CH3 | 2.435136 | 2.24 |
| 28-H | CH3 | 2.458724 | 2.11 | 29-H | CH3 | 2.435368 | 2.24 |
| 19-H | CH3 | 2.566401 | 2.27 | 33-H | CH3 | 2.255059 | 2.38 |
| 20-H | CH3 | 2.568626 | 2.27 | 32-H | CH3 | 2.937239 | 2.38 |
| 21-H | CH3 | 2.585603 | 2.27 | 31-H | CH3 | 2.937305 | 2.38 |
| 34-H | CH3 | 2.153130 | 2.40 | 23-H | Ar–H | 7.009666 | 7.08 |
| 32-H | CH3 | 2.785887 | 2.40 | 24-H | Ar–H | 7.456872 | 7.44 |
| 33-H | CH3 | 2.837816 | 2.40 | 25-H | Ar–H | 7.935175 | 7.74 |
| 25-H | Ar–H | 6.946737 | 7.19 | 34-H | H-3pyridine | 8.267729 | 8.13 |
| 24-H | Ar–H | 7.068655 | 7.44 | 22-H | OH | 10.90081 | 10.78 |
| 23-H | Ar–H | 7.821711 | 7.72 | 21-H | NH | 11.86255 | 11.4 |
| 30-H | H-3pyridine | 8.037436 | 8.27 | 14-H | NH | 11.91355 | 11.4 |
| 22-H | OH | 11.11657 | 10.82 | ||||
| 14-H | NH | 11.83394 | 11.13 | ||||
| Compound 8 | Compound 9 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 27-H | CH3 | 1.97523 | 2.28 | 26-H | CH3 | 2.003243 | 2.20 |
| 28-H | CH3 | 2.423421 | 2.28 | 31-H | CH3 | 2.291672 | 2.35 |
| 26-H | CH3 | 2.423452 | 2.28 | 27-H | CH3 | 2.449094 | 2.20 |
| 32-H | CH3 | 2.291102 | 2.40 | 25-H | CH3 | 2.449112 | 2.20 |
| 30-H | CH3 | 2.914005 | 2.40 | 35-H | NCH3 | 2.920509 | 3.26 |
| 31-H | CH3 | 2.914115 | 2.40 | 34-H | NCH3 | 2.927599 | 3.26 |
| 22-H | Ar–H | 7.019014 | 7.43 | 30-H | CH3 | 3.009217 | 2.35 |
| 23-H | Ar–H | 7.462446 | 7.62 | 29-H | CH3 | 3.009511 | 2.35 |
| 24-H | Ar–H | 7.931906 | 7.86 | 38-H | NCH3 | 3.055131 | 3.52 |
| 33-H | H-3pyridine | 8.588581 | 8.28 | 39-H | NCH3 | 3.05569 | 3.52 |
| 21-H | OH | 10.62159 | 11.09 | 36-H | NCH3 | 4.280042 | 3.26 |
| 20-H | NH | 11.08248 | 11.33 | 40-H | NCH3 | 4.636198 | 3.52 |
| 14-H | NH | 11.23007 | 11.33 | 21-H | Ar–H | 7.042978 | 7.53 |
| 22-H | Ar–H | 7.461108 | 7.73 | ||||
| 23-H | Ar–H | 7.964887 | 7.91 | ||||
| 32-H | H-3pyridine | 8.717135 | 8.43 | ||||
| 20-H | OH | 10.93315 | 11.54 | ||||
For the synthesized compounds 4–9, the methyl protons attached to the benzene rings appeared in the experimental 1H NMR spectra at chemical shifts (δ) ranging from 2.11 to 2.28 ppm, which showed good agreement with the computed values of δ 1.31–2.46 ppm. Similarly, the CH3 pyrazole in compounds 4–6 were observed experimentally at δ 2.27–2.31 ppm, with corresponding calculated values between δ 2.29 and 2.56 ppm. The CH3 group on the pyridine ring in compounds 6–9 exhibited experimental signals at δ 2.35–2.40 ppm, which closely matching the calculated range of δ 2.26–3.01 ppm. Furthermore, the two methyl groups on the pyrimidine moiety in compound 9 were observed at δ 3.26 and 3.52 ppm, which presented good agreement with the computed values ranged between δ 2.92–4.28 and 3.06–4.64 ppm, respectively.
Notably, in the experimental 1H NMR spectra, each methyl group appeared as a singlet, indicating that the three protons occupy equivalent magnetic environments. Meanwhile, the theoretical spectra predicted three separate signals for each methyl group. This discrepancy likely arises from the distinct spatial orientations of the methyl protons, which result in varied local chemical environments and differences in electron density distribution. These findings are further supported by the molecular electrostatic potential (MEP) maps (Fig. 3 and S3–S4), which reveal variable electron densities surrounding the methyl protons. As a result, each proton is subject to a slightly different magnetic field, accounting for the observed splitting in the computed spectra.
Furthermore, the aromatic (benzo) protons appeared in their characteristic region in the experimental spectra, between δ 6.92 and 7.91 ppm. These signals showed excellent agreement with the theoretically predicted chemical shifts, which ranged from δ 6.84 to 8.12 ppm.
In the 1H NMR spectra of compounds 4 and 5, the H-4 proton of the pyrazole ring was experimentally observed at δ 8.17 and 8.29 ppm, which corresponding theoretical values at δ 8.20 and 8.61 ppm, respectively. For compounds 6–9, the H-3 proton of the pyridine moiety appeared in the experimental range of δ 8.13–8.43 ppm, which closely matched the computed range of δ 8.04–8.72 ppm. Additionally, the hydroxyl (OH) protons were recorded in the experimental spectra at δ 10.76–11.54 ppm, showing good agreement with the calculated chemical shifts ranging from δ 10.62 to 11.37 ppm.
The 13C NMR spectra of compounds 6–9 are presented in Fig. S14, S18, S22, S26, S30 and S34. The experimental and theoretical chemical shift values (δ, ppm) for these compounds are summarized in Tables 7–9.
| Compound 4 | Compound 5 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 36-C | CH3 | 16.04123 | 18.5 | 22-C | CH3 | 16.37035 | 17.8 |
| 32-C | CH3 | 20.03835 | 22.1 | 18-C | CH3 | 20.36041 | 22.3 |
| 8-C | C-4pyrazole | 109.601 | 103.2 | 8-C | C-4pyrazole | 115.0985 | 104.1 |
| 15-C | C-3quinoline | 118.3876 | 117.5 | 5-C | Ar–C | 122.7267 | 121.4 |
| 5-C | Ar–C | 120.4593 | 120.9 | 3-C | Ar–C | 126.7115 | 124.1 |
| 3-C | Ar–C | 122.7618 | 123.6 | 37-C | Ar–C | 127.4954 | 124.6 |
| 14-C | Ar–C | 127.1452 | 124.7 | 48-C | Ar–C | 128.7039 | 125.2 |
| 16-C | Ar–C | 128.5288 | 125.2 | 46-C | Ar–C | 128.8355 | 125.7 |
| 20-C | Ar–C | 129.8325 | 125.8 | 35-C | Ar–C | 129.1542 | 126.2 |
| 22-C | Ar–C | 130.6145 | 127.6 | 44-C | Ar–C | 129.869 | 126.7 |
| 2-C | Ar–C | 131.3425 | 128.4 | 2-C | Ar–C | 130.0925 | 127.1 |
| 1-C | Ar–C | 132.7052 | 129.1 | 50-C | Ar–C | 130.9401 | 127.5 |
| 6-C | Ar–C | 132.7319 | 130.5 | 1-C | Ar–C | 131.4779 | 127.9 |
| 24-C | Ar–C | 135.6543 | 133.4 | 45-C | Ar–C | 132.597 | 128.3 |
| 13-C | Ar–C | 136.2726 | 134.7 | 34-C | Ar–C | 133.0551 | 128.6 |
| 11-C | C-3pyrazole | 143.3956 | 142 | 6-C | Ar–C | 134.3273 | 129.2 |
| 18-C | C-2quinoline | 144.9399 | 143.5 | 33-C | Ar–C | 134.9496 | 129.5 |
| 17-C | C-8a quinoline | 146.6616 | 144.3 | 39-C | Ar–C | 135.4928 | 130.1 |
| 7-C | C-5pyrazole | 147.2444 | 145.9 | 32-C | Ar–C | 136.0035 | 130.5 |
| 4-C | C–OH | 155.364 | 151.7 | 43-C | Ar–C | 136.9156 | 131.9 |
| 11-C | C-3pyrazole | 142.8217 | 141.4 | ||||
| 14-C | C-6triazine | 145.2531 | 142.7 | ||||
| 30-C | C-5 triazine | 145.3566 | 142.2 | ||||
| 13-C | C-3triazine | 146.7012 | 143.8 | ||||
| 7-C | C-5pyrazole | 146.8773 | 144.1 | ||||
| 4-C | C–OH | 154.665 | 150.8 | ||||
| Compound 6 | Compound 7 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 18-C | CH3 | 16.2857 | 16.7 | 26-C | CH3 | 20.79757 | 17.6 |
| 31-C | CH3 | 20.7996 | 17.5 | 30-C | CH3 | 24.20442 | 21.3 |
| 26-C | CH3 | 20.8586 | 22.6 | 13-C | C-4a | 108.7055 | 109.5 |
| 13-C | C-3a | 115.0560 | 107.1 | 3-C | Ar–C | 118.7887 | 121.3 |
| 5-C | Ar–C | 121.5214 | 122.0 | 1-C | Ar–C | 121.8902 | 124.9 |
| 3-C | Ar–C | 123.3304 | 124.3 | 4-C | Ar–C | 130.7422 | 126.8 |
| 6-C | Ar–C | 127.4543 | 126.1 | 5-C | Ar–C | 132.6759 | 129.7 |
| 1-C | Ar–C | 129.2197 | 128.3 | 6-C | Ar–C | 133.2412 | 130.3 |
| 2-C | Ar–C | 130.5930 | 130.4 | 8-C | C-6 | 135.3256 | 132.3 |
| 8-C | C-5 | 131.3823 | 132.3 | 10-C | C-5 | 140.8247 | 141.2 |
| 10-C | C-4 | 142.1141 | 140.0 | 7-C | C-7 | 144.7709 | 146.0 |
| 17-C | C-3 | 144.8144 | 142.3 | 11-C | C-8a | 149.0900 | 147.3 |
| 7-C | C-6 | 145.7319 | 144.7 | 2-C | C–OH | 152.6519 | 151.2 |
| 11-C | C-7a | 150.8757 | 148.3 | 17-C | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine Opyrimidine |
165.5737 | 166.0 |
| 4-C | C–OH | 158.9102 | 151.4 | 18-C | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine Opyrimidine |
170.9304 | 169.1 |
| Compound 8 | Compound 9 | ||||||
|---|---|---|---|---|---|---|---|
| Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental | Atoms (theoretical) | Atoms (experimental) | Calculated | Experimental |
| 25-C | CH3 | 20.61305 | 16.5 | 24-C | CH3 | 20.78461 | 16.2 |
| 29-C | CH3 | 24.11115 | 21.3 | 28-C | CH3 | 25.01277 | 20.8 |
| 13-C | C-4a | 110.5799 | 109.5 | 33-C | NCH3 | 29.18944 | 28.4 |
| 3-C | Ar–C | 118.5432 | 122.1 | 37-C | NCH3 | 30.12316 | 30.1 |
| 1-C | Ar–C | 122.0142 | 125.0 | 13-C | C-4a | 109.6181 | 109.3 |
| 4-C | Ar–C | 128.1293 | 126.2 | 3-C | Ar–C | 119.2506 | 122.4 |
| 5-C | Ar–C | 130.1193 | 128.6 | 1-C | Ar–C | 121.7722 | 124.2 |
| 6-C | Ar–C | 131.9459 | 130.5 | 4-C | Ar–C | 130.6600 | 126.5 |
| 8-C | C-6 | 132.0892 | 131.8 | 5-C | Ar–C | 131.7313 | 129.8 |
| 10-C | C-5 | 140.6713 | 141.1 | 6-C | Ar–C | 133.9255 | 130.9 |
| 7-C | C-7 | 145.4148 | 146.5 | 8-C | C-6 | 135.8494 | 132.5 |
| 11-C | C-8a | 149.6338 | 147.7 | 10-C | C-5 | 143.6066 | 140.8 |
| 2-C | C–OH | 151.7754 | 150.4 | 7-C | C-7 | 146.5615 | 146.3 |
| 18-C | C=Opyrimidine | 169.7040 | 168.3 | 11-C | C-8a | 149.4658 | 147.4 |
| 17-C | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S S |
180.8169 | 186.2 | 2-C | C–OH | 153.2882 | 150.9 |
| 16-C | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine Opyrimidine |
164.0081 | 167.6 | ||||
| 17-C | C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine Opyrimidine |
165.1212 | 170.3 | ||||
The methyl carbons attached to the benzene ring in compounds 6–9 were observed experimentally in the range of δ 16.2–17.6 ppm, while the corresponding computed values ranged from δ 16.3 to 20.8 ppm. For the methyl groups on the pyridine rings, experimental chemical shifts appeared between δ 20.8 and 22.6 ppm, closely matching the calculated range of δ 20.9–25.0 ppm. In compound 6, the methyl carbon on the pyrazole ring was recorded experimentally at δ 17.5 ppm, which agrees well with the computed value of δ 20.8 ppm. The relatively higher δ values of the methyl group on the pyridine ring compared to other methyl groups may be attributed to its attachment at the C-4 position of the pyridine moiety. This position is more deshielded due to the electron-withdrawing effect of the nitrogen atom, resulting in a downfield shift (higher δ value) and hence experiencing a lower magnetic field relative to other methyl groups. This was further supported by the relatively high δ value of C-4pyridine as compared with other benzo carbons, where the experimental δ values ranged between δ 140.0–141.2 ppm which matched with the calculated values at δ 140.7–143.6 ppm.
The experimental δ values for the C–OH carbons were observed in the range of 150.4–151.4 ppm, while the corresponding computed values ranged from δ 151.8 to 158.9 ppm. The relatively high chemical shifts of these carbons can be attributed to the deshielding effect induced by the electron-withdrawing inductive influence of the adjacent hydroxyl atom.
Furthermore, for compounds 7–9, the experimental δ values for the C-8a carbon were recorded in the range of 147.3–147.7 ppm, which are in good agreement with the computed values of 149.0–149.6 ppm. The relatively high chemical shifts can be attributed to deshielding effects arising from both mesomeric and inductive influences of the adjacent nitrogen atoms.
The experimental δ values for the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Opyrimidine of compounds 7–9 were recorded in the range of 166.0–170.3 ppm, which correspond well with the calculated values ranging from 164.0 to 169.7 ppm. Additionally, the chemical shift for the C
Opyrimidine of compounds 7–9 were recorded in the range of 166.0–170.3 ppm, which correspond well with the calculated values ranging from 164.0 to 169.7 ppm. Additionally, the chemical shift for the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) S group was observed experimentally at δ 186.2 ppm, closely matching the computed value of δ 180.0 ppm.
S group was observed experimentally at δ 186.2 ppm, closely matching the computed value of δ 180.0 ppm.
Finally, the chemical shift values for the remaining carbon atoms in the experimental spectra show good agreement with the corresponding values in the computed spectra, as presented in Tables 7–9.
Based on the above results, the observed chemical shift values showed excellent correlation with the calculated values, with correlation coefficients (R2) ranging from 0.98 to 0.99, as illustrated in Fig. 4, 5 and S8–S11.
 | ||
| Fig. 4 The correlation relationships of the experimental versus calculated 1H NMR chemical shifts of compounds 4 and 5. | ||
| µ = (µx2 + µy2 + µz2)1/2 | (10) |
| α = (αxx + αyy + αzz)/3 | (11) |
| Δα = (2)−0.5[(αxx − αyy)2 + (αyy − αzz)2 + (αzz − αxx)2 + 6(αyz)2 + 6(αxy)2 + 6(αxz)2]0.5 | (12) |
| βtot = [(βxxx + βxyy + βxzz)2 + (βyyy + βyzz + βyxx)2 + (βzzz + βzxx + βzyy)2]0.5 | (13) |
Elevated dipole moment, polarizability, and hyperpolarizability values are generally associated with enhanced nonlinear optical (NLO) performance. The calculated parameters for the investigated compounds, which reflect their potential NLO activity, are summarized in Table 10. According to the results, the dipole moments (µ) ranged from 2.861 to 5.471 D, while the polarizability (α) values fell within 1.710 × 10−23 to 2.569 × 10−23. Additionally, all the molecules exhibit total hyperpolarizability (βtot) values that are 1.28 to 5 times greater than that of urea (0.3728 × 10−30 esu).73 These enhanced NLO parameters suggest that the synthesized compounds hold strong potential for NLO-based applications.
| Compound no. | µx | µy | µz | µtotal | 〈α〉 (au) | 〈α〉 (esu) × 10−23 | Δα (au) | Δα (esu) × 10−24 | βtotal (au) | βtotal (esu) × 10−30 |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 0.240 | −4.099 | 0.001 | 4.106 | 115.40 | 1.710 | 133.34 | 19.760 | 55.44 | 0.479 |
| 4 | −3.021 | 0.788 | −1.287 | 3.377 | 152.43 | 2.259 | 21.03 | 3.117 | 101.21 | 0.874 |
| 5 | −3.279 | 0.901 | 1.691 | 3.798 | 173.32 | 2.569 | 18.33 | 2.716 | 105.21 | 0.909 |
| 6 | −1.782 | 2.374 | 0.298 | 2.983 | 104.13 | 1.543 | 16.06 | 2.381 | 112.43 | 0.971 |
| 7 | −3.049 | 2.483 | 0 | 3.932 | 124.58 | 1.846 | 3.40 | 0.504 | 120.94 | 1.045 |
| 8 | 4.521 | 3.083 | 0 | 5.471 | 134.25 | 1.990 | 3.42 | 0.507 | 216.94 | 1.874 |
| 9 | −2.225 | 1.799 | −0.001 | 2.861 | 134.73 | 1.997 | 8.09 | 1.110 | 67.97 | 0.587 |
3.3. Biological evaluation
 | ||
| Fig. 6 Relation between cell viability and concentration of all synthesized compounds on the proliferation of HepG2 cell line (cis-platin is the standard drug). | ||
The IC50 values of the synthesized compounds, as summarized in Fig. 7 and Table 11, were compared to that of the reference drug cis-platin. The findings revealed that all compounds exhibited moderate to strong anticancer activity. Notably, compound 5 showed the highest potency with an IC50 of 6.57 µM L−1, surpassing cis-platin (IC50 = 17.86 µM L−1) in effectiveness. This enhanced activity is likely due to the incorporation of a triazine chromophore within the pyrazole ring.
 | ||
| Fig. 7 Virtual IC50 values of the target compounds and cis-platin as a standard drug against HepG-2 cell lines. | ||
| Compounds no. | (HepG-2 cells) IC50 (µM L−1) |
|---|---|
| 4 | 22.78 ± 1.07 |
| 5 | 6.57 ± 0.59 |
| 6 | 27.76 ± 1.12 |
| 7 | 11.30 ± 0.84 |
| 8 | 8.86 ± 0.71 |
| 9 | 16.54 ± 0.97 |
| Cisplatin | 17.86 ± 0.93 |
Additionally, compounds 7–9, with IC50 values of 11.30, 8.86, and 16.54 µM, respectively, also demonstrated greater antiproliferative activity against HepG-2 cell lines than cis-platin. This improved performance is likely attributed to the presence of the pyrido[2,3-d]pyrimidine core. Additionally, the substitution of oxygen atom in the pyrimidine ring of compound 7 with a sulfur atom (compound 8) markedly improved the biological activity.
Moreover, compounds 4 and 6, with IC50 values of 22.78 and 27.76 µM L−1, respectively, displayed moderate inhibitory effects compared to the standard drug. These findings suggest that certain synthesized compounds could serve as promising candidates for the treatment of human hepatocellular carcinoma.
Linear regression analysis was conducted using the experimental inhibitory activity (IC50) as the dependent variable and the DFT-derived global reactivity descriptors in the gas phase as independent variables (Table 1). The key findings from this analysis are summarized as follows:
1 The positive slope of the linear correlation equation, IC50 = −75.37 + 23.46 ΔE/eV (r = 0.97, n = 6), indicates that a reduction in the energy gap (ΔE) of the compounds is associated with a decrease in IC50 values. This suggests that compounds with lower energy gaps tend to exhibit stronger anticancer activity by requiring lower concentrations to inhibit 50% of cell viability (Fig. 8).
2 Further, the positive slope of the correlation between IC50 and hardness, expressed as IC50 = −75.32 + 49.90 η/eV (r = 0.97, n = 6), indicates that lower hardness values are associated with reduced IC50, reflecting enhanced inhibitory activity (Fig. 8). Conversely, an inverse relationship was observed between IC50 and softness, as shown by the negative slope in the equation IC50 = 112.19–186.10 S eV−1 (r = 0.90, n = 8), further supporting that increased molecular softness contributes to stronger anticancer effects (Fig. 8).
The drug-likeness of the synthesized compounds was evaluated using the SwissADME online tool.36 Compounds that violate more than one of the Lipinski,76 Veber,77 Egan78 or Ghose79 rules are generally considered unsuitable for oral administration. According to Lipinski's rule of five, a compound should have a molecular weight (MW) below 500 g mol−1, a criterion met by all compounds. Additionally, drug-like molecules should have no more than five hydrogen bond donors and ten hydrogen bond acceptors. The MLogP value, which reflects lipophilicity, should also be below 5. The calculated MLogP values for the synthesized compounds range from 1.14 to 3.99, indicating acceptable lipophilic behavior.
According to Veber's rule, the number of rotatable bonds should not exceed ten—a condition met by all the synthesized compounds. Another important parameter is the Total Polar Surface Area (TPSA), which influences a compound's ability to cross biological membranes. Veber's rule recommends a TPSA value below 140 Å2, and all compounds comply with this limit. Moreover, Egan's rule further specifies two criteria: TPSA should be ≤131 Å2 and WLOGP (a measure of lipophilicity) should be ≤5.88. Further, Ghose's rule outlines additional requirements: molecular weight between 160 and 480, WLOGP between −0.4 and 5.6, molar refractivity between 40 and 130, and a total atom count between 20 and 70. The relevant ADME data for all tested compounds are summarized in Table 12.
| Compound no. | MW | MLOGP | HBA | HBD | TPSA | nRotB | WLOGP | XLOGP3 | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S S |
Fraction Csp3 | MR | Lipinski violations | Veber violations | Egan violations | Ghose violations |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 160.17 | 1.14 | 2 | 0 | 30.21 | 0 | 2.10 | 2.27 | −2.88 | 0.10 | 47.45 | 0 | 0 | 0 | 0 |
| 4 | 349.81 | 3.46 | 3 | 1 | 50.94 | 2 | 5.06 | 4.86 | −5.56 | 0.10 | 101.27 | 0 | 0 | 0 | 0 |
| 5 | 419.48 | 3.99 | 5 | 1 | 76.72 | 4 | 5.38 | 4.89 | −5.93 | 0.08 | 125.21 | 0 | 0 | 0 | 0 |
| 6 | 253.30 | 2.41 | 3 | 2 | 61.80 | 1 | 3.26 | 3.25 | −3.98 | 0.20 | 76.25 | 0 | 0 | 0 | 0 |
| 7 | 283.28 | 1.80 | 4 | 3 | 98.84 | 1 | 1.60 | 1.74 | −3.19 | 0.13 | 80.38 | 0 | 0 | 0 | 0 |
| 8 | 299.35 | 1.80 | 3 | 3 | 113.86 | 1 | 2.97 | 2.34 | −3.67 | 0.13 | 84.94 | 0 | 0 | 0 | 0 |
| 9 | 311.34 | 1.89 | 4 | 1 | 77.12 | 1 | 1.62 | 2.11 | −3.55 | 0.24 | 90.18 | 0 | 0 | 0 | 0 |
Furthermore, the bioavailability radar offers assessment of a molecule's drug-likeness by evaluating six key physicochemical properties. The optimal range for each property is indicated by the pink area on the radar and includes: (1) LIPO – lipophilicity (XLOGP3 between −0.7 and +5.0), (2) SIZE – molecular weight between 150 and 500 g mol−1, (3) POLAR – polarity, with a total polar surface area (TPSA) between 20 and 130 Å2, (4) INSOLU – solubility, where log S should not exceed 6, (5) INSATU – saturation, with a fraction of Csp3 between 0.25 and 1, and (6) FLEX – flexibility, defined as having no more than 9 rotatable bonds. As illustrated in Fig. 9 and S36, all evaluated parameters for the compounds fall within the ideal range, except for INSATU.
As a result, the studied compounds obey the above rules and oral bioavailability radar. Therefore, the studied compounds are orally active because they adhere to these rules with zero. Theoretically, they would not have problems with oral bioavailability.
The docking methodology was validated by re-docking the co-crystallized ligand, [(2,6-difluorophenyl)carbonyl]amino)-N-(4-fluorophenyl)-1H-pyrazole-3-carboxamide, and the binding pose closely matched the native structure (Fig. S37), confirming the reliability of the docking protocol.
Docking results for compounds 4–9 are summarized in Table 13, S1, Fig. 10 and S38. These results show the predicted binding modes and interactions with amino acid residues. For example, compound 4 shows Pi-Sigma and Pi–Alkyl interactions with ALA31, LYS89, and ILE10, while compound 5 exhibits hydrogen bonding with ASN133 and GLN132, along with additional alkyl interactions. Compound 8 displays the most favorable predicted binding energy (−9.10 kcal mol−1) due to Pi-Anion and Pi-Lone Pair interactions. Full interaction details are provided in Table 13 and S1.
| Compound 5 (binding energy −9.10 kcal mol−1) | Compound 7 (binding energy −8.70 kcal mol−1) | ||||
|---|---|---|---|---|---|
| Receptor | Distance | Interaction type | Receptor | Distance | Interaction type |
| ASN133 | 3.05 | H-bond | VAL18 | 2.40 | H-bond |
| GLN132 | 2.94 | Pi-lone pair | ILE10 | 2.71 | H-bond |
| LYS88 | 3.94 | Alkyl | ILE10 | 2.18 | H-bond |
| LYS89 | 4.07 | Alkyl | ASP86 | 2.50 | Pi-anion |
| TYR15 | 4.96 | Pi–alkyl | LYS88 | 3.83 | Alkyl |
| LEU135 | 5.41 | Pi–alkyl | LYS89 | 4.54 | Alkyl |
| LYS88 | 4.67 | Pi–alkyl | VAL18 | 4.54 | Pi–alkyl |
| LYS89 | 4.59 | Pi–alkyl | LYS89 | 4.69 | Pi–alkyl |
| VAL18 | 4.84 | Pi–alkyl | |||
| Compound 8 (binding energy −9.10 kcal mol−1) | Co-crystallized ligand (binding energy −10.60 kcal mol−1) | ||||
|---|---|---|---|---|---|
| ASP86 | 2.34 | Pi-anion | LEU83 | 2.1229 | H-bond |
| GLY11 | 3.68 | Pi-sigma | GLU81 | 1.66193 | H-bond |
| ILE10 | 2.73 | Pi-lone pair | ASP146 | 4.75511 | Pi-anion |
| LYS88 | 3.24 | Alkyl | VAL18 | 3.99529 | Pi-sigma |
| LYS89 | 5.11 | Alkyl | LEU135 | 3.50724 | Pi-sigma |
| TYR15 | 4.94 | Pi–alkyl | ILE10 | 4.07235 | Pi–alkyl |
| VAL18 | 5.10 | Pi–alkyl | ALA31 | 3.38275 | Pi–alkyl |
| ILE10 | 5.10 | Pi–alkyl | VAL64 | 5.26978 | Pi–alkyl |
| LYS89 | 4.79 | Pi–alkyl | LEU83 | 5.45921 | Pi–alkyl |
| ALA145 | 5.06422 | Pi–alkyl | |||
 | ||
| Fig. 10 3D representation of the hydrogen bonding between the studied compounds (5, 7, 8) with the amino acids residues of the target protein (PDB ID: 4Y72). | ||
Although these docking results provide insights into potential binding interactions, they are predictive and qualitative in nature. They suggest trends in binding affinity but do not confirm target inhibition. As such, the correlation between docking energies and in vitro IC50 values against HepG-2 cells (Table 13 and S1) should be interpreted with caution. Compounds 5 and 8, which show favorable predicted binding energies, also exhibit lower IC50 values, while compounds 4 and 6 show weaker binding energies and higher IC50 values. These observations indicate a possible association between predicted interactions and cytotoxicity, but further biochemical validation would be required to confirm any mechanistic link.
Overall, molecular docking provides supportive information that complements the in vitro cytotoxicity data, helping to rationalize observed trends without implying direct target inhibition.
4. Conclusions
A series of novel heterocyclic compounds including pyrazole derivatives (4 and 5), pyrazolo[3,4-b]pyridine 6, and pyrido[2,3-d]pyrimidines 7–9, were synthesized via ring opening-ring closure (RORC) reactions of 2,6-dimethylchromone (1) with various heterocyclic hydrazines and cyclic enamines. The structures of the synthesized compounds were confirmed by elemental analysis and various spectroscopic techniques. Biological evaluation against HepG-2 cell lines revealed that compound 5 exhibited the most potent anticancer activity, with IC50 of 6.57 µM, making it the most promising candidate among the studied compounds. The molecular geometries of the current compounds were optimized using DFT with the B3LYP functional and the 6-311++G(d,p) basis set. Frontier molecular orbital (FMO) analysis indicated that compound 1 had the largest energy gap (ΔE = 4.894 eV) and was the hardest compound (η = 2.447 eV), reflecting high chemical stability. that of urea, indicating potential for nonlinear optical (NLO) applications. In contrast, compound 5 exhibited the smallest HOMO–LUMO energy gap (ΔE = 3.534 eV), indicating enhanced reactivity. It also exhibited the highest softness value (S = 0.566 eV−1), further confirming that it is the most reactive compound among the studied molecules. Additionally, all synthesized compounds exhibited hyperpolarizability (βtot) values greater than urea. To support the experimental NMR data, GIAO calculations were employed to simulate 1H and 13C NMR chemical shifts, showing strong agreement with the experimental results. SwissADME analysis confirmed that the compounds possess drug-like properties and are potentially suitable for oral bioavailability. Finally, molecular docking simulations against cyclin-dependent kinase CDK1 (PDB ID: 4Y72) demonstrated favorable binding profiles, notably for compounds 5, 7, and 8, whose strong binding affinities correlate well with their in vitro anticancer activities.Author contributions
Mohamed Abdel-Megid: investigation, formal analysis, writing – review & editing. Najla A. Alshaye: investigation, formal analysis, writing – review & editing. Al-Shimaa Badran: supervision, conceptualization, investigation, methodology, formal analysis, writing – review & editing. Magdy A. Ibrahim: conceptualization, investigation, methodology, formal analysis, writing – review & editing.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
Data are available upon request from the authors.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra08430j.
Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).References
- S. Zhang, Y. Xie, L. Song, Y. Wang, H. Qiu, Y. Yang, C. Li, Z. Wang, Z. Han and L. Yang, Seven new 2-(2-phenylethyl)chromone derivatives from agarwood of Aquilaria agallocha with inhibitory effects on nitric oxide production, Fitoterapia, 2022, 159, 105177, DOI:10.1016/j.fitote.2022.105177.
- M. Fan, W. Yang, M. He, Y. Li, Z. Peng and G. Wang, Occurrence, synthesis and biological activity of 2-(2-phenyethyl)chromones, Eur. J. Med. Chem., 2022, 237, 114397, DOI:10.1016/j.ejmech.2022.114397.
- R. S. Keri, S. Budagumpi, R. K. Pai and R. G. Balakrishna, Chromones as a privileged scaffold in drug discovery: A review, Eur. J. Med. Chem., 2014, 78, 340–374, DOI:10.1016/j.ejmech.2014.03.047.
- V. M. Patil, N. Masand, S. Verma and V. Masand, Chromones: Privileged scaffold in anticancer drug discovery, Chem. Biol. Drug Des., 2021, 98, 943–953, DOI:10.1111/cbdd.13951.
- S. Abdpour, L. Baleh, H. Nadri, H. Forootanfar, S. N. A. Bukhari, A. Ramazani, S. E. S. Ebrahimi, A. Foroumadi and M. Khoobi, Chromone derivatives bearing pyridinium moiety as multi-target-directed ligands against alzheimer’s disease, Bioorg. Chem., 2021, 110, 104750, DOI:10.1016/j.bioorg.2021.104750.
- M. A. Ibrahim and T. E. Ali, Synthesis and antimicrobial activity of chromone-linked 2-pyridone fused with 1,2,4-triazoles, 1,2,4-triazines and 1,2,4-triazepines ring systems, J. Braz. Chem. Soc., 2010, 21, 1007–1016, DOI:10.1590/S0103-50532010000600010.
- S. Khursheed, M. R. Wani, G. G. H. A. Shadab, S. Tabassum and F. Arjmand, Synthesis, structure elucidation by multi-spectroscopic techniques and single-crystal X-ray diffraction of promising fluoro/bromo-substituted-chromone(bpy)copper(II) anticancer drug entities, Inorg. Chim. Acta, 2022, 538, 120967, DOI:10.1016/j.ica.2022.120967.
- V. Kumar, M. Gupta, S. G. Gandhi, S. S. Bharate, A. Kumar, R. A. Vishwakarma and S. B. Bharate, Anti-inflammatory chromone alkaloids and glycoside from Dysoxylum binectariferum, Tetrahedron Lett., 2017, 58, 3974–3978, DOI:10.1016/j.tetlet.2017.09.005.
- A. K. Narula, C. S. Azad and L. M. Nainwal, New dimensions in the field of antimalarial research against malaria resurgence, Eur. J. Med. Chem., 2019, 181, 111353, DOI:10.1016/j.ejmech.2019.05.043.
- X. Ouyang, X. Li, W. Lu, D. Zhao and A. Null B-Ring, Improves the Antioxidant Levels of Flavonol: A Comparative Study between Galangin and 3,5,7-trihydroxychromone, Molecules, 2018, 23, 3083–3095, DOI:10.3390/molecules23123083.
- A. N. Mpitimpiti, J. P. Petzer, A. Petzer, J. H. L. Jordaan and A. C. U. Lourens, Synthesis and evaluation of chromone derivatives as inhibitors of monoamine oxidase, Mol. Divers., 2019, 23, 897–913, DOI:10.1007/s11030-019-09917-8.
- N. A. Alshaye and M. A. Ibrahim, Design, synthesis, characterization, and antimicrobial evaluation of the novel 2-heteroarylchromeno[2,3-d]pyrimidines, Synth. Commun., 2023, 53, 744–754, DOI:10.1080/00397911.2023.2196025.
- M. A. Ibrahim and A. Badran, Novel Heteroannulated Chromeno[2,3-b]pyridines and Related Compounds Using 6-Methylchromone-3-carbonitrile, Heterocycles, 2022, 104, 707–722, DOI:10.3987/COM-21-14607.
- A. Badran, Z. Hussain, N. S. Abdelshafi, S. S. Ibrahim, A. Ahmed, M. Abdel-Megid and M. A. Ibrahim, Ring opening and recyclization reactions with 3-functionalized chromones: Recent synthetic approaches for five, six and seven membered heterocycles, Synth. Commun., 2025, 55(10), 693–716, DOI:10.1080/00397911.2025.2463603.
- A. Badran, M. A. Ibrahim and A. Ahmed, Nucleophilic reactions with the novel condensation product derived from 3-formylchromone and 4-hydroxycoumarin, Synth. Commun., 2021, 51(12), 1868–1881, DOI:10.1080/00397911.2021.1910961.
- S. Abdel Halim and M. A. Ibrahim, Synthesis, structure investigation, DFT analysis, optical, and photoelectrical properties of 9-bromo-3-hydroxychromeno[4,3-b]pyrazolo[4,3-e]pyridin-5 (1H)-one (BHCPP), Results Chem., 2022, 4, 100572, DOI:10.1016/j.rechem.2022.100572.
- M. A. Ibrahim, T. E. Ali, Y. A. Gabr and Y. A. Alnamer, Synthesis and chemical reactivity of 2-methylchromones, Arkivoc, 2010, 1, 98–135, DOI:10.3998/ark.5550190.0011.103.
- S. S. Ibrahim, H. M. El-Shaaer and A. Hassan, Phosphorus Sulfur Silicon Relat. Elem., 2002, 177, 151–172, DOI:10.1080/10426500210228.
- L. Ouksel, R. Bourzami, N. Hamdouni and A. Boudjada, Synthesis, supramolecular structure, spectral properties and correlation between nonlinear optic, thermochemistry and thermal behavior of an α-Hydroxyphosphonate acid ester, dual experimental and DFT approaches, J. Mol. Struct., 2021, 1229, 129792, DOI:10.1016/j.molstruc.2020.129792.
- M. Belhocine, R. Bourzami, F. Dergal, L. Ouksel, A. Ammari, Z. Benladghem, A. Haouzi and S. Bouktab, Physical, chemical and antibacterial properties of 1-methyl-3-(4-vinylbenzyl) imidazole-3-ium chloride ionic liquid: Experimental and ab-initio analysis, J. Mol. Struct., 2023, 1271, 133955, DOI:10.1016/j.molstruc.2022.133955.
- M. A. Ibrahim, H. M. Hassanin, Y. Gabr and Y. A. Alnamer, Synthesis and Chemical Reactivity of the Novel 4-Hydroxy-6-methyl-2,5-dioxo-5,6-dihydro-2H-pyrano[3,2-c]quinoline-3-carboxaldehyde, J. Heterocycl. Chem., 2018, 55(12), 2834–2844, DOI:10.1002/jhet.3353.
- M. Medimagh, C. B. Mleh, N. Issaoui, A. S. Kazachenko, T. Roisnel, O. M. Al-Dossary, H. Marouani and L. G. Bousiakoug, DFT and molecular docking study of the effect of a green solvent (water and DMSO) on the structure, MEP, and FMOs of the 1-ethylpiperazine-1,4-diium bis(hydrogenoxalate) compound, J. Mol. Liq., 2023, 369, 120851, DOI:10.1016/j.molliq.2022.120851.
- D. M. Aziz, S. A. Hassan, D. M. Mamand and K. Qurbani, New azo-azomethine derivatives: Synthesis, characterization, computational, solvatochromic UV–Vis absorption and antibacterial studies, J. Mol. Struct., 2023, 1284, 135451, DOI:10.1016/j.molstruc.2023.135451.
- S. Sukumaran, A. Zochedh, T. M. Viswanathan, A. B. Sultan and T. Kathiresan, Theoretical investigation of 5-fluorouracil and tamoxifen complex–structural, spectrum, DFT, ADMET and docking simulation, Polycycl. Aroma, Compds, 2023, 43(10), 9443–9460, DOI:10.1080/10406638.2022.2164018.
- J.-K. Hou, Y. Huang, W. He, Z. W. Yan, L. Fan, M.-H. Liu, W.-L. Xiao, H.-D. Sun and G.-Q. Chen, Adenanthin targets peroxiredoxin I/II to kill hepatocellular carcinoma cells, Cell Death Dis., 2014, 5, e1400, DOI:10.1038/cddis.2014.345.
- X. Yang, X. Xie, Y. F. Xiao, R. Xie, C. J. Hu, B. Tang, B. S. Li and S. M. Yang, The emergence of long non-coding RNAs in the tumorigenesis of hepatocellular carcinoma, Cancer Lett., 2015, 360, 119–124, DOI:10.1016/j.canlet.2015.02.035.
- C. Peng, P. Y. Ayala, H. B. Schlegel and M. J. Frisch, Using redundant internal coordinates to optimize equilibrium geometries and transition states, J. Comput. Chem., 1996, 17(1), 49–56, DOI:10.1002/(SICI)1096-987X(19960115.
- P. J. Stephens, F. J. Devlin, C. F. Chablowski and M. J. Frisch, Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields, J. Phys. Chem., 1994, 98(45), 11623–11627, DOI:10.1021/j100096a001.
- M. J. Frisch, J. A. Pople and J. S. Binkley, Self-consistent molecular orbital methods 25. supplementary functions for gaussian basis sets, J. Chem. Phys., 1984, 80, 3265–3269, DOI:10.1063/1.447079.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, J. A. Montgomery, J. E. Peralta, F. Ogliaro, M. Bearpark, J. J. Heyd, B. E, K. N. Kudin, V. N. Staroverov, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. J. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, D. 0109, Revision D. 01, Gaussian, Inc., Wallingford, CT. ( 2009 Search PubMed.
- R. D. Dennington, T. A. Keith and J. M. Millam, Gauss View 5.0.8, Gaussian Inc. ( 2008 Search PubMed.
- K. Wolinski, J. F. Hinton and P. Pulay, Efficient implementation of the gauge-independent atomic orbital method for NMR chemical shift calculations, J. Am. Chem. Soc., 1990, 112(23), 8251–8260, DOI:10.1021/ja00179a005.
- T. Mosmann, Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays, J. Immunol. Methods, 1983, 65(1–2), 55–63, DOI:10.1016/0022-1759(83)90303-4.
- A. Sabt, W. M. Eldehna, T. Al-Warhi, O. J. Alotaibi, M. M. Elaasser, H. Suliman and H. A. Abdel-Aziz, Discovery of 3,6-disubstituted pyridazines as a novel class of anticancer agents targeting cyclin-dependent kinase 2: synthesis, biological evaluation and in silico insights, J. Enzyme Inhib. Med. Chem., 2020, 35(1), 1616–1630, DOI:10.1080/14756366.2020.1806259.
- S. M. Gomha, S. M. Riyadh, E. A. Mahmmoud and M. M. Elaasser, Synthesis and anticancer activities of thiazoles, 1,3-thiazines, and thiazolidine using chitosan-grafted-poly(vinyl pyridine) as basic catalyst, Heterocycles, 2015, 91(6), 1227–1243, DOI:10.1002/chin.201538150.
- A. Daina, O. Michielin and V. Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Sci. Rep., 2017, 7, 42717, DOI:10.1038/srep42717.
- V. Pogaku, K. Gangarapu, S. Basavoju, K. K. Tatapudi and S. B. Katragadda, Design, synthesis, molecular modelling, ADME prediction and anti-hyperglycemic evaluation of new pyrazole-triazolopyrimidine hybrids as potent α-glucosidase inhibitors, Bioorg. Chem., 2019, 93, 103307, DOI:10.1016/j.bioorg.2019.103307.
- O. Trott and A. J. Olson, AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 2010, 31(2), 455–461, DOI:10.1002/jcc.21334.
- N. R. Brown, S. Korolchuk, M. P. Martin, W. A. Stanley, R. Moukhametzianov, M. E. Noble and J. A. Endicott, CDK1 structures reveal conserved and unique features of the essential cell cycle CDK, Nat. Commun., 2015, 6(1), 6769, DOI:10.1038/ncomms7769.
- S. Narra and N. Munirathinam, Synthesis, structure analysis, Hirshfeld surface studies, molecular docking studies against cyclin-dependent kinases (CDKs) of sulfonamide decorated N6-benzyl aminopurines for cancer treatment, Results Chem., 2024, 8, 101592, DOI:10.1016/j.rechem.2024.101592.
- P. Jayavel, V. Ramasamy, N. Amaladoss, V. Renganathan and V. I. Shupeniuk, A facile synthesis, characterization, DFT, ADMET and in-silico molecular docking analysis of novel 4-ethyl acridine-1,3,9 (2,4,10H)-trione, Chem. Phys. Impact, 2024, 8, 100476, DOI:10.1016/j.chphi.2024.100476.
- A. Zahirović, S. Fetahović, M. Feizi-Dehnayebi, R. Bešta-Gajević, M. Dizdar, J. Ostojić and S. Roca, Substituent effect in salicylaldehyde 2-furoic acid hydrazones: Theoretical and experimental insights into DNA/BSA affinity modulation, antimicrobial and antioxidant activity, J. Mol. Struct., 2024, 1312, 138628, DOI:10.1016/j.molstruc.2024.138628.
- B. Parveen, S. Shahzadi, S. Ali, M. Feizi-Dehnayebi, K. S. Munawar, M. Ashfaq and M. N. Tahir, Synthesis, spectral characterizations, computational studies and biological investigation of 4-(4-(2-hydroxyethyl)phenylamino)-4-oxobutanoic acid and its trimethyltin(IV) complex, J. Mol. Struct., 2024, 1315, 138851, DOI:10.1016/j.molstruc.2024.138851.
- D. Sahin, R. A. Kepekci, M. Feizi-Dehnayebi, S. Akkoc, J. V. Cuevas-Vicario and N. Micale, Biological Activities, DFT Calculations, and Molecular Docking Simulation of Thymol-Based Compounds, ChemistrySelect, 2024, 9(23), e202304572, DOI:10.1002/slct.202304572.
- R. K. Mohapatra, A. Mahal, A. Ansari, M. Kumar, J. P. Guru, A. K. Sarangi, A. Abdou, S. Mishra, M. Aljeldah and B. M. AlShehail, Comparison of the binding energies of approved mpox drugs and phytochemicals through molecular docking, molecular dynamics simulation, and ADMET studies: An in silico approach, J. Biosaf. Biosecur., 2023, 5(3), 118–132, DOI:10.1016/j.jobb.2023.09.001.
- A. M. El-Saghier, A. Abdou, M. A. Mohamed, H. M. Abd El-Lateef and A. M. Kadry, Novel 2-acetamido-2-ylidene-4-imidazole derivatives (El-Saghier reaction): Green synthesis, biological assessment, and molecular docking, ACS Omega, 2023, 8(33), 30519–30531, DOI:10.1021/acsomega.3c03767.
- N. H. Al-Shaalan, Synthesis, Characterization and biological Activities of Cu(II), Co(II), Mn(II), Fe(II), and UO2(VI) complexes with a new schiff base hydrazone: o-hydroxyacetophenone-7-chloro-4-quinoline hydrazone, Molecules, 2011, 16(10), 8629–8645, DOI:10.3390/molecules16108629.
- P. V. Laakso, R. Robinson and H. P. Vandrewala, Studies in the triazine series including a new synthesis of 1:2:4-triazines, Tetrahedron, 1957, 1, 103–118, DOI:10.1016/0040-4020(57)85014-5.
- S. Abdel Halim and M. A. Ibrahim, Synthesis, spectral analysis, quantum studies, NLO, and thermodynamic properties of the novel 5-(6-hydroxy-4-methoxy-1-benzofuran-5-ylcarbonyl)-6-amino-3-methyl-1H-pyrazolo[3,4-b]pyridine (HMBPP), RSC Adv., 2022, 12, 13135–13153, 10.1039/d2ra01469f.
- M. J. Mphahlele, G. K. More, J. K. Nkoana, Y. S. Choong and A. A. Elhenawy, Design, synthesis and evaluation of the 2′-hydroxy-3′-iodo-5′-nitrochalcones for cytotoxicity (MCF-7 & A549) and potential to inhibit tyrosine kinase (VEGFR-2) Activity, J. Mol. Struct., 2024, 1305, 137785, DOI:10.1016/j.molstruc.2024.137785.
- M. M. Maluleka, R. S. Segodi, M. J. Mphahlele, V. G. Mbazima, A. A. Elhenawy and B. A. Monchusi, Synthesis, structural characterization, and quantum chemical study of the 7-acetyl-5-nitrobenzofurans as anticancer agents with antioxidant properties, J. Mol. Struct., 2024, 1311, 138398, DOI:10.1016/j.molstruc.2024.138398.
- R. G. Pearson, Absolute electronegativity and hardness: applications to organic chemistry, J. Org. Chem., 1989, 54(6), 1423–1430, DOI:10.1021/jo00267a034.
- C. Kucuk, S. Celik, S. Yurdakul and E. Coteli, A new Ag(I)-complex of 5-chloroquinolin-8-ol ligand: Synthesis, spectroscopic characterization, and DFT investigations, in vitro antioxidant (DPPH and ABTS), α-glucosidase, α-amylase inhibitory activities with protein-binding analysis, J. Mol. Struct., 2025, 1325, 141285, DOI:10.1016/j.molstruc.2024.141285.
- N. A. Alshaye, M. A. Ibrahim and A. Badran, Nucleophilic transformation of 3-substituted-6,8-dimethylchromones with phenylhydrazine under various reaction conditions: Theoretical, Spectroscopic characterization and in silico ADME studies, J. Mol. Struct., 2025, 1297, 37006, DOI:10.1016/j.molstruc.2023.137006.
- K. Periyasamy, P. Sakthivel, G. Venkatesh, V. Palanisamy and Y. S. Mary, Synthesis and design of carbazole-based organic sensitizers for DSSCs applications: experimental and theoretical approaches, Chem. Pap., 2024, 78, 447–461, DOI:10.1007/s11696-023-03101-x.
- Y. Zheng, M. Zheng, X. Ling, Y. Liu, Y. Xue, L. An, N. Gu and M. Jin, Design, synthesis, quantum chemical studies and biological activity evaluation of pyrazole–benzimidazole derivatives as potent Aurora A/B kinase inhibitors, Bioorg. Med. Chem. Lett., 2013, 23, 3523–3530, DOI:10.1016/j.bmcl.2013.05.084.
- M. Uzzaman and M. Jabedul Hoque, Physiochemical, molecular docking, and pharmacokinetic studies of Naproxen and its modified derivatives based on DFT, Int. j. sci. res. manag., 2018, 6(9), 12–19, DOI:10.18535/ijsrm/v6i9.c01.
- H. N. Banavath, O. P. Sharma, M. S. Kumar and R. Baskaran, Identification of novel tyrosine kinase inhibitors for drug resistant T315I mutant BCR-ABL: a virtual screening and molecular dynamics simulations study, Sci. Rep., 2014, 4, 6948, DOI:10.1038/srep06948.
- L. Touafri, A. Hellal, S. Chafaa, A. Khelifa and A. Kadri, Synthesis, characterisation and DFT studies of three Schiff bases derived from histamine, J. Mol. Struct., 2017, 1149, 750–760, DOI:10.1016/j.molstruc.2017.08.052.
- R. P. Iczkowski and J. L. Margrave, Electronegativity, J. Am. Chem. Soc., 1961, 83(17), 3547–3551, DOI:10.1021/ja01478a001.
- L. Pauling, The nature of the chemical bond. iv. the energy of single bonds and the relative electronegativity of atoms, J. Am. Chem. Soc., 1932, 54(9), 3570–3582, DOI:10.1021/ja01348a011.
- R. G. Parr, L. V. Szentpaly and S. Liu, Electrophilicity index, J. Am. Chem. Soc., 1999, 121(9), 1922–1924, DOI:10.1021/ja983494x.
- A. Ali, M. Khalid, M. A. Rehman, F. Anwar, H. Z. U. Aabidin, M. N. Akhtar, M. U. Khan, A. A. C. Braga, M. A. Assiri and M. Imran, An Experimental and computational exploration on the electronic, spectroscopic, and reactivity properties of novel halo-functionalized hydrazones, ACS Omega, 2020, 5(30), 18907–18918, DOI:10.1021/acsomega.0c02128.
- V. Dixit and Y. Ra, Experimental IR and Raman spectroscopy and DFT methods-based material characterization and data analysis of 2-Nitrophenol, Biochem. Pharmacol., 2015, 4(4), 100018, DOI:10.4173/2167-0501.1000183.
- M. J. Alam and S. Ahmad, Quantum chemical and spectroscopic investigations of 3-methyladenine, Spectrochim. Acta, Part A, 2014, 128, 653–664, DOI:10.1016/j.saa.2014.02.170.
- F. M. Bickelhaupt, Essentials of computational chemistry theories and models. by Christopher J. cramer, Angew. Chem., Int. Ed., 2003, 42(4), 381–382, DOI:10.1002/anie.200390122.
- R. A. Omer, K. M. Ahmed, K. A. Omar, W. M. Hamad and D. M. Mamad, N, N-Bis (2,4-dihydroxybenzaldehyde)benzidine: Synthesis, characterization, DFT, and theoretical corrosion study, J. Mol. Struct., 2024, 1300, 137279, DOI:10.1016/j.molstruc.2023.137279.
- D. R. Durai, S. Sowrirajan, N. Elangovan, S. Y. Alomar, T. Sankar Ganesan, C. G. Priya, B. R. Venkatraman and A. Nawaz, Local energy decomposition (through methanol), biological activity prediction, solvents interaction (DFT) and molecular docking simulation studies of Schiff base: synthesis, characterization and fluorescence properties, Results Chem., 2023, 6, 101030, DOI:10.1016/j.rechem.2023.101030.
- R. Suja, A. Rathika, V. S. J. Reeda, A. A. kumar and P. Divya, Synthesis, spectroscopic analysis (FT-IR, FT-Raman, UV, NMR), non-covalent interactions (RDG, IGM) and dynamic simulation on Bis (8-hydroxyquinoline)salicylate salicylic acid, J. Mol. Struct., 2024, 1310, 138231, DOI:10.1016/j.molstruc.2024.138231.
- M. Irfan, H. A. Khan, S. Bibi, G. Wu, A. Ali, S. G. Khan, N. Alhokbany, F. Rasool and K. Chen, Exploration of nonlinear optical properties of 4-methyl-4H-1,2,4-triazol-3-yl)thio)-N-phenylpropanamide based derivatives: experimental and DFT approach, Sci. Rep., 2024, 14, 2732, DOI:10.1038/s41598-024-51788-z.
- V. Kakekochi, U. Kumar D, P. P. Nikhil and K. Chandrasekharan, An investigation on photophysical and third– order nonlinear optical properties of novel thermally–stable thiophene–imidazo[2,1-b][1,3,4]thiadiazole based azomethines, Dyes Pigm., 2019, 167, 216–224, DOI:10.1016/j.dyepig.2019.04.033.
- T. Rajamani and S. Muthu, Electronic absorption, vibrational spectra, non-linear optical properties, NBO analysis and thermodynamic properties of 9-[(2-hydroxyethoxy)methyl] guanine molecule by density functional method, Solid State Sci., 2013, 16, 90–101, DOI:10.1016/j.solidstatesciences.2012.10.023.
- M. Dupuis, C. Adant and J. L. Bredas, Ab initio study of the nonlinear optical properties of urea: electron correlation and dispersion effects, Int. J. Quant. Chem., 1995, 56(S29), 497–507, DOI:10.1002/qua.560560853.
- S. A. Hassan, D. M. Aziz, M. N. Abdullah, A. R. Bhat, R. S. Dongre, T. B. Hadda, F. A. Almalki, S. M. A. Kawsar, A. K. Rahiman, S. Ahmed, M. H. Abdellattif, M. Berredjem, S. A. Sheikh and J. Jamalis, In vitro and in vivo evaluation of the antimicrobial, antioxidant, cytotoxic, hemolytic activities and in silico POM/DFT/DNA-binding and pharmacokinetic analyses of new sulfonamide bearing thiazolidin-4-ones, J. Biomol. Struct. Dyn., 2023, 42(7), 3747–3763, DOI:10.1080/07391102.2023.2226713.
- N. T. P. Nyoni, N. B. Ncube, M. X. Kubheka, N. P. Mkhwanazi, S. Senzani, T. Singh and M. Tukulula, Synthesis, characterization, in vitro antimycobacterial and cytotoxicity evaluation, DFT calculations, molecular docking and ADME studies of new isomeric benzimidazole-1,2,3-triazole-quinoline hybrid mixtures, Bioorg. Chem., 2023, 141, 106904, DOI:10.1016/j.bioorg.2023.106904.52.
- C. A. Lipinski, Drug-like properties and the causes of poor solubility and poor permeability, J. Pharmacol. Toxicol. Methods, 2000, 44(1), 235–249, DOI:10.1016/s1056-8719(00)00107-6.
- D. F. Veber, S. R. Johnson, H. Y. Cheng, B. R. Smith, K. W. Ward and K. D. Kopple, Molecular properties that influence the oral bioavailability of drug candidates, J. Med. Chem., 2002, 45(12), 2615–2623, DOI:10.1021/jm020017n.
- W. J. Egan, K. M. Merz and J. J. Baldwin, Prediction of drug absorption using multivariate statistics, J. Med. Chem., 2000, 43, 3867–3877, DOI:10.1021/jm000292e.
- A. K. Ghose, V. N. Viswanadhan and J. J. Wendoloski, A knowledge-based approach in designing combinatorial or medicinal chemistry libraries for drug discovery. 1. A qualitative and quantitative characterization of known drug databases, J. Comb. Chem., 1999, 1(1), 55–68, DOI:10.1021/cc9800071.
- T. Lanez, M. Feizi-Dehnayebi and E. Lanez, Assessment of the electrostatic binding of ferrocenylmethyl-nitroaniline derivatives to DNA: A combined experimental and theoretical study, J. Mol. Struct., 2024, 1308, 138386, DOI:10.1016/j.molstruc.2024.138386.
- A. M. Abu-Dief, O. A. Omran, M. Feizi-Dehnayebi, A. Alqurashi, I. Omar, D. Alhashmialameer and A. D. M. Mohamad, Fabrication, structural elucidation, and DFT calculation of some new hydrophilic metal chelates based on N N′-(1-methyl-2-oxoindolin-3-ylidene)benzohydrazide ligand: Pharmaceutical studies and molecular docking approach, Appl. Organomet. Chem., 2024, 38(9), e7593, DOI:10.1002/aoc.7593.
- H. M. Abd El-Lateef, A. M. Ali, M. M. Khalaf and A. Abdou, New iron (III), cobalt (II), nickel (II), copper (II), zinc (II) mixed-ligand complexes: Synthesis, structural, DFT, molecular docking and antimicrobial analysis, Bull. Chem. Soc. Ethiop., 2024, 38(1), 147–166, DOI:10.4314/bcse.v38i1.12.
- A. M. El-Saghier, S. S. Enaili, A. M. Kadry, A. Abdou and M. A. Gad, Green synthesis, biological and molecular docking of some novel sulfonamide thiadiazole derivatives as potential insecticidal against Spodoptera littoralis, Sci. Rep., 2023, 13(1), 19142, DOI:10.1038/s41598-023-46602-1.
- M. Malumbres and M. Barbacid, Cell cycle, CDKs and cancer: a changing paradigm, Nat. Rev. Cancer, 2009, 9(3), 153–166, DOI:10.1038/nrc2602.
| This journal is © The Royal Society of Chemistry 2026 |