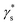DOI:
10.1039/D5RA08276E
(Paper)
RSC Adv., 2026,
16, 14806-14821
Bioprocess optimization and purification of DLA produced using food and fishery waste
Received
28th October 2025
, Accepted 3rd March 2026
First published on 17th March 2026
Abstract
Food waste is a sustainable and attractive waste biomass which can be utilized as a substrate for the production of value-added bioproducts. In this study, Y. lipolytica engineered for D-lactic acid (DLA) production was optimised for food waste hydrolysate (FWH) and fish protein hydrolysate (FPH). The substrate inhibition studies using FWH showed that after a 50 (g L−1) concentration of glucose, there was a decline in both specific growth rate and DLA yield, and the Luong model with an R2 of 0.933 gave a better model fit. Nitrogen screening studies revealed that fish protein hydrolysate (FPH) could be an economical replacement for yeast extract. The Placket–Burman (PB) screening evaluation revealed that FWH, pH, and KH2PO4 were the key factors affecting DLA production. Furthermore, in central composite design (CCD) studies with optimal parameter levels, an 8.7% increase in DLA production was observed. Additionally, using an Artificial Neural Network (ANN)-linked Genetic Algorithm (GA), optimised parameter levels of FWH 49.98 (g L−1), pH 8.52, and KH2PO4 3.93 (g L−1) were obtained, enhancing DLA production by 12.6% compared with the Plackett–Burman studies. Bioreactor studies with FPH as the only nitrogen source and FWH as the carbon source exhibited 0.94 (g g−1) DLA yields, which were similar to the GA prediction. Kinetic modelling studies using MATLAB/Simulink showed that DLA production followed a mixed growth-dependent product-formation pattern. DLA purification using a butanol and ammonium sulfate solvent system yielded 92.7% recovery efficiency with glucose as the carbon source, and 83.51% recovery efficiency with FWH as the carbon source. Furthermore, characterization with HPLC, FTIR, and NMR reiterated the presence of DLA.
1. Introduction
With the exponential increase in global demand for fossil-based fuels, the demand for commodity chemicals derived from them “is also raising rapidly”. According to studies, 140 million tons of petrochemical polymers are produced globally each year, and their waste streams are also released into the environment.1 It is estimated that the continued production of petrochemical-based polymers could result in the accumulation of 850 million tonnes of plastic waste by 2050. Studies have reported that 150 million tonnes of plastic are seen on the ocean's surface as floating plastic waste.2 To address these environmental concerns and reduce dependence on fossil-based plastic production, there is a growing interest in bioplastics. Globally, total bioplastic production is estimated at 2.42 million tonnes, of which poly lactic acid (PLA) is the most preferred bioplastic, accounting for 20% of the global bioplastic market.3 It is projected that global PLA production will increase by 0.75 million tonnes, reflecting the demand for its unique and beneficial properties in biomedical and food packaging applications.3 For efficient PLA polymer production, a pure, economical lactic acid monomer is essential. In nature, lactic acid exists in two optical isomers, D and L. DLA exhibits distinctive thermal and biomedical properties when used in PLA preparation, leading to a high demand for DLA in PDLA production.4 DLA is an important platform chemical attracting interest in numerous applications, such as the pharmaceutical, biomedical, food, and leather industries, and serves as a precursor for caproic acid, butyric acid, etc.5 Globally lactic acid market size is estimated at 2.6 billion USD in the year 2021 and expected to grow to 5.02 billion by 2028.6 According to a report, annually 270![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tons of lactic acid (LA) are produced, and demand for LA is increasing by 10% annually.6 LA can be produced by both chemical and biological methods, with 90% of LA production occurring via fermentation.7 The cost of lactic acid depends on its applications, which range from 3.0 to 4.0 $ per kg, which was 1.5–3.0 $ per kg as of 2011.8 Recent market trends reported by the Tridge group indicate that in 2024, the global lactic price ranged from $2.74 per kg to $13.68 per kg. Also, there was a significant decline in the price from $ 1.06 per kg to $6.58 per kg. The reasons are sudden shifts in demand and supply. Annually, 3.3 × 105 tons of lactic acid are produced through the biochemical route using glucose derived from food-based materials as the substrate, contributing to the ongoing food-versus-fuel debate.9 Producing LA primarily from first-generation feedstocks may increase production costs. Further exploiting second- and third-generation renewable feedstocks derived from agro-industrial waste may substantially reduce lactic acid production costs.10 According to the Food Wastage Index Report 2021, globally, 1.3 billion tonnes of food waste are generated annually.11 Of the overall food waste generated, a minor portion is redistributed, composted, or treated by other methods; the majority ends up in landfills.12 Food waste is complex, making it difficult for microbes to hydrolyse it due to their limited metabolic capabilities.13 Food waste contains valuable nutrients and can be effectively used to produce fungal enzymes, such as glucoamylase and proteases, which efficiently break it down.14 Several studies are reporting the utilisation of food waste, either directly or after treatment, to produce bioethanol, biodiesel, organic acids, pigments, antibiotics, enzymes, etc.13,15,16
000 tons of lactic acid (LA) are produced, and demand for LA is increasing by 10% annually.6 LA can be produced by both chemical and biological methods, with 90% of LA production occurring via fermentation.7 The cost of lactic acid depends on its applications, which range from 3.0 to 4.0 $ per kg, which was 1.5–3.0 $ per kg as of 2011.8 Recent market trends reported by the Tridge group indicate that in 2024, the global lactic price ranged from $2.74 per kg to $13.68 per kg. Also, there was a significant decline in the price from $ 1.06 per kg to $6.58 per kg. The reasons are sudden shifts in demand and supply. Annually, 3.3 × 105 tons of lactic acid are produced through the biochemical route using glucose derived from food-based materials as the substrate, contributing to the ongoing food-versus-fuel debate.9 Producing LA primarily from first-generation feedstocks may increase production costs. Further exploiting second- and third-generation renewable feedstocks derived from agro-industrial waste may substantially reduce lactic acid production costs.10 According to the Food Wastage Index Report 2021, globally, 1.3 billion tonnes of food waste are generated annually.11 Of the overall food waste generated, a minor portion is redistributed, composted, or treated by other methods; the majority ends up in landfills.12 Food waste is complex, making it difficult for microbes to hydrolyse it due to their limited metabolic capabilities.13 Food waste contains valuable nutrients and can be effectively used to produce fungal enzymes, such as glucoamylase and proteases, which efficiently break it down.14 Several studies are reporting the utilisation of food waste, either directly or after treatment, to produce bioethanol, biodiesel, organic acids, pigments, antibiotics, enzymes, etc.13,15,16
In lactic acid production, the LA yield is directly influenced by glucose concentration and is affected by the composition of the nitrogen source.17 According to a study, 38% of the process cost for lactic acid fermentation is taken by the nitrogen source itself.18 Several studies also reported that yeast extract is the preferred nitrogen source for lactic acid production.18,19 The cost of yeast extract is a challenging parameter to replace with an economical and efficient nitrogen source, making lactic acid affordable for PLA.20 An extensive research has been conducted to find a better substitute for yeast extract. One such better alternative could be a nitrogen source from fish waste. In addition to agriculture, India leads in aquaculture, ranking 2nd globally in fish production for human consumption and contributing 1.07% to GDP.21 It is estimated that by 2028, the share of fish for human consumption from aquaculture could increase to 58%, up from 53% in 2016–18.22 Similarly, according to a report on aquaculture, 185.4 million tons of waste have been generated, accounting for 35% of total fish production.23 So the acid or enzymatic hydrolysis of fish waste can yield a large amount of water-soluble protein, essential polypeptides, small peptides, and amino acids.24 Studies have also shown that acid-hydrolysed proteins, such as casein hydrolysate, fish hydrolysate, soybean hydrolysate, and ram horn hydrolysate, can serve as effective nitrogen sources in plant nutrition, PET nutrition products, and industrial applications, such as lactic acid production.24 Optimisation is the inevitable part of any fermentation-based studies for industrial applications. At industrial scale, complex media can affect the economics of product formation, so utilising minimal media can reduce the cost of lactic acid production. Machine learning and metaheuristic-based optimisation tools were explored to attain global optimisation of lactic acid production.25
In the present study, an attempt was made to produce high-titer DLA using minimal media. The media was optimised for a renewable carbon source (FWH) and a nitrogen source (FPH). Furthermore, ANN-linked GA optimised the response (DLA) obtained from Placket–Burman (PB) & Response Surface Methodology (RSM) studies. Further, kinetic modelling studies supported experimental results in explaining the parameter optimisation. This study is one of the limited reports focusing on the purification and characterization of DLA produced from the fermentation broth of a genetically engineered yeast strain.
2. Materials and methods
2.1. Materials
All chemicals used in this study were of analytical grade and procured from Himedia Laboratories Pvt. Ltd, Mumbai. Mixed Food Waste (MFW) was collected from the NIT Warangal Hostel mess. The collected MFW was mixed in a kitchen blender and autoclaved. Furthermore, the autoclaved MFW was stored in zip-lock bags and used for subsequent studies. Fish protein hydrolysate was procured from Janatha Agro Products, Udupi, Karnataka, India. FPH is used directly after procurement, without any additional pretreatment. The composition of fish protein hydrolysate is given in Table S6. All studies in the present study were performed in triplicate.
2.2. Microorganisms
The engineered Yarrowia lipolytica (YDl-Acs) strain expressing the ldhA and ACS2 genes was used to optimise processing parameters to enhance DLA production.26 The YDl-Acs yeast strain was maintained on YEPD (in (g L−1) – Yeast extract-10, Peptone-20 and Dextrose-20) as glycerol stocks at (−80 °C). Aspergillus terreus BPD01 (Accession Number-OQ603456.1), the fungal strain used in the present study, was isolated from a soil sample. Further information on the isolation, identification and production of glucoamylase is mentioned in our previous study, Chandukishore et al. (2023).25 The Aspergillus fungal strain cultures were stored as glycerol stocks at −80 °C and maintained on potato dextrose medium.
2.3. Food waste hydrolysate preparation
Food waste hydrolysate was prepared using A. terreus, following the protocol described by Chandukishore et al. (2023).25 A. terreus BPD01 was cultured on PDA agar plates for 4 days until complete spore formation on PDA plates. Later, spores were harvested using sterile water and inoculated on 10 g of MFW at a concentration of 2.5 × 105 (Spores per mL). Further, the inoculated MFW was incubated at 30 °C for 5 days in a solid-state fermentation to produce fungal mash containing glucoamylase. Later, fresh MFW was treated with fungal mash at a ratio of (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4) in a submerged hydrolysis at 55 °C, 150 rpm. Further samples were taken at regular intervals until the FWH was recorded, the optimum glucose concentration, which was analysed using the GOD-POD assay. The obtained FWH was concentrated at 60 °C using a rotary evaporator by vacuum evaporation.27
4) in a submerged hydrolysis at 55 °C, 150 rpm. Further samples were taken at regular intervals until the FWH was recorded, the optimum glucose concentration, which was analysed using the GOD-POD assay. The obtained FWH was concentrated at 60 °C using a rotary evaporator by vacuum evaporation.27
2.4. Substrate inhibition studies
Further substrate inhibition studies were performed by varying the glucose concentration in the FWH from 3% to 8% in a semi-synthetic medium using the protocol given by Chandukishore et al. (2025).26 Additionally, semi-synthetic medium was composed based on the media composition reported by Narisetty et al. (2022) with the following modifications in (g L−1): yeast extract-10, yeast nitrogen base-1.3, ammonium chloride-1.5 in 0.5 M potassium phosphate buffer pH (6.8).28 The seed culture was incubated in 10 mL YEPD medium at 30 °C and 180 rpm for 24 hours. Later, active cells were harvested and washed twice in sterile water and inoculated into the semi-synthetic medium at (0.1 OD). Incubated at 30 °C, 100 rpm.28 Later samples were taken at regular intervals to estimate lactic acid, acetic acid, biomass and residual glucose concentration.
2.5. Substrate inhibition models
Substrate inhibition models were analysed using the protocol followed by Dutta et al., (2015).29 Substrate inhibition models are based on Monod's growth model, with modifications and extensions by various scientists, providing insight into the inhibition and affinity constants. In the Monod model, microbial growth kinetics depend on substrate concentration. In the case of the Aiba model, the substrate inhibition constant is an exponential term that also accounts for product inhibition when predicting kinetics. In the case of the Andrew model, the inhibition constant is a significant factor; this model is preferable at high substrate concentrations. In the Haldane model, an increase in substrate concentration provides insights into the substrate affinity constant and the inhibition constant. The uniqueness of the Luong model is the maximum substrate concentration (Sm) constant above which it monitors a decline in the sharpness of the growth rate. In the case of the Moser model, it yields the Monod model (n = 1) without substrate inhibition and can also provide information on the lag phase in fermentation studies. The value in the Moser model helps incorporate experimental data into bioreactor studies. The Yano model was designed to analyse the specific growth rate during continuous fermentation at high substrate concentrations, incorporating a substrate inhibition constant. The equations for all the models studied are listed in Table S1. The detailed sums of squares (SS), degrees of freedom (df), regression coefficient (R2), root mean square error (RMSE), and other constants for all the models were calculated using MATLAB 7.1.
2.6. Screening of nitrogenous sources
The screening of nitrogen sources for DLA production was performed based on the protocol of Wongsirichot et al. (2022).27 The nitrogen source plays a crucial role in fermentation processes, particularly influencing lactate dehydrogenase enzyme activity and biomass formation. Additionally, both the type and concentration of the nitrogen source directly affect lactic acid production. To understand this, various organic and inorganic nitrogen sources were selected and screened to assess their effects on DLA yield and economic viability. For optimal microbial lactic acid production, the nitrogen concentration was typically maintained at 0.2 to 0.5 g L−1.30 With the above information, the following nitrogen sources were considered for the study: Yeast Extract (YE), Casein hydrolysate, ammonium sulfate, Fish Protein Hydrolysate (FPH), and urea. The fermentation media were composed of (g L−1): Glucose-50, YNB-1.3, and the nitrogen source was varied by type; overall, a nitrogen concentration of 0.5 (g L−1) was kept constant. The pre-inoculum was prepared as specified in the section above, and samples were collected at regular intervals for metabolite analysis.
2.7. Placket Burman design (PB)
After one-factor-at-a-time optimisation of food-waste hydrolysate concentration and screening of nitrogen sources, nine parameters were further screened using a PB design with two levels and three centre points, affecting DLA production (Table S2). All runs of the Plackett–Burman design were performed in a 100 mL conical flask containing 30 mL of medium. Further samples were collected at 12-hour intervals and analysed for DLA, acetic acid, biomass and glucose concentration.
2.8. Central composite design (CCD) for parameter optimisation
Based on the results of the Plackett–Burman studies, three factors have been finalised for further analysis of their effects on DLA production and for optimisation using a central composite design. The following experiments were set up in 20 runs, including 6 centre points, and were conducted in a 100 mL conical flask containing 30 mL of fermentation medium with the following composition (in g L−1): yeast extract-10, FPH-15, K2HPO4-4, YNB-1.5, ammonium chloride-1.5, and inoculum size-1.0 OD. The parameters and levels used for the central composite design are shown in Table S3. The CCD model is represented by (eqn (1)), where Yg is predicted DLA yield in (g L−1), β0 is the intercept, β1, β2, and β3 are linear coefficients; β11, β22 and β33 are the quadratic coefficients; β12,β13 and β23 are the interactive coefficients. The experimental design was created using Minitab.| | |
Yg = β0 + β1X1 + β2X2 + β3X3 + β11X12 +β22X22 + β33X12 + β12X1X2 + β13X1X3 +β23X2X3
| (1) |
2.9. Artificial neural networks
Artificial neural network and genetic algorithm studies were performed according to the protocol followed by Chandukishore et al. (2023).25 An artificial neural network (ANN) model is a computational model that imitates the functioning of the human neural system. It consists of input signals at interconnected nodes, yielding dynamic responses as outputs. The efficiency of the generated neural model is measured using mean squared error (MSE). The neural model performs best when its training dataset mean squared error (MSE) is at a minimum. In the present study, the Levenberg–Marquardt (LM) method has been incorporated to optimise DLA production using data sets generated from CCD studies. In this study, a neural model is developed to train data sets from CCD modelling experiments to further optimise the parameters for DLA production. In our research, ANN follows a feed-forward back-propagation neural model, which consists of three layers: input, hidden layer and output. The input layer includes parameters such as FWH, KH2PO4, and pH; in the output layer, the DLA titer is considered a process variable.
2.10. Genetic algorithm (GA)
A genetic algorithm (GA) is a metaheuristic optimisation tool based on Darwin's evolutionary algorithm.25 In ANN-based prediction, only local values are used for optimisation. In contrast, GA predicts input parameter levels by fixing the maximum output or response using a global optimisation approach. A genetic algorithm is defined by elements such as gene input parameters, a set of gene interactions (a chromosome), and fitness, which is the response to the gene interactions. In the present study, the focus is on DLA yield. Offspring with low fitness undergo mutation and crossover to interact with the previously better chromosome, thereby achieving the best global optimisation. The above interactions are repeated until the best fitness is achieved. Fitness is calculated by average error values between the expected DLA yield and predicted values, and is given by the equation where fD is the fitness value for the DLA yield, Yij is given by the predicted ith DLA yield of jth chromosome, Yi* is given by the expected DLA yield, and N is the number of estimated experiments. The minimal fitness error is the best optimal model.| |
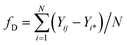 | (2) |
2.11. Validation studies
The final optimal parameter levels obtained from CCD and ANN-linked GA studies were used for shake flask validation. The optimal semi-synthetic media was composed of (in (g L−1)-FWH-50, FPH-20, YNB-2, ammonium chloride-2, K2HPO4-4, KH2PO4-4). Pre-inoculum was cultured in 10 mL YEPD medium, incubated at 30 °C, 180 rpm for 24 hours. Later, the active cells were harvested and washed twice in sterile H2O and inoculated into the semi-synthetic medium at an optical density (OD) of 1. Later fermentation was carried out at 30 °C, 100 rpm at pH 8.5. Samples were taken at regular intervals to analyse organic acids, biomass and glucose concentration.
2.12. Batch bioreactor studies
Bioreactor-based scale-up studies were performed according to the protocol given by Chandukishore et al. (2025),.26 An in situ-sterilised stirred-tank bioreactor (Biojenik Engineering, India) with a capacity of 3.5 L and a working volume of 2 L was used for batch fermentation. Initially, pre-inoculum was prepared in a 500 mL Erlenmeyer flask with medium composed of (in (g L−1)-FWH-20, FPH-20, YNB-2, ammonium chloride-2, K2HPO4-4, KH2PO4-4 and trace elements) and incubated for 24 hours at 30 °C, 200 rpm. The inoculum size was maintained at 1 OD in the fermentation medium. The fermentation medium composition was the same as that of the pre-inoculum, except for a change in the FWH concentration from 20 (g L−1) in the pre-inoculum to 50 (g L−1) in the fermentation medium. Batch fermentation conditions were maintained at pH 8.5 using 5 M NaOH and 5 M HCl. A two-level dissolved oxygen (DO) strategy of (>30%) was maintained using the impeller speed of 500 rpm, and 1 (L minute−1) air sparged using a 0.22 µm air filter up to 24 hours. Later, a micro-aerobic condition was maintained by reducing the speed to 200 rpm, and air sparging was stopped to favor LA production.
2.13. Kinetic modelling of bioreactor studies
2.13.2. Product formation kinetics. Lactic acid production kinetics were studied using the Luedeking–Piret equation. The kinetics of lactic acid production were studied as a function of biomass concentration (X) and growth rate (dX/dt).| |
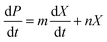 | (6) |
In eqn (6), m and n are kinetic constants for product formation. ‘n’ can be calculated by the stationery phase data of batch fermentation studies, at which biomass concentration (dX/dt = 0) and (X = Xm) to rearrange the equation with the above assumptions to give ‘n’| |
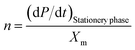 | (7) |
Eqn (6) is rearranged as follows P as a function of time (t)
The eqn (9) on product formation is derived by integrating eqn (8) utilising the eqn (4)
| |
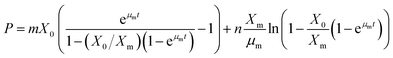 | (9) |
Eqn (9) is written in simple form as
where
| |
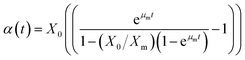 | (11) |
| |
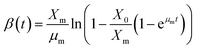 | (12) |
In
eqn (10), ‘
m’ is the growth associated constant determined by plotting (
P −
nβ(
t))
versus α(
t), and ‘
n’ is the non-growth rate constant estimated by
eqn (7).
2.13.3. Substrate utilization kinetics. Glucose is also used as the carbon source for lactic acid formation and biomass accumulation. Substrate utilisation kinetics is usually represented by eqn (13).| |
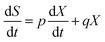 | (13) |
In eqn (13) (p = 1/Yx/s) and q is the maintenance coefficient (ms). The procedure followed for product formation kinetics is also applied to substrate utilization kinetics. The maintenance coefficient can be estimated by the following equation at the stationary phase (dX/dt = 0; X = Xm).| |
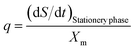 | (14) |
The eqn (14) is rearranged as follows to analyse ‘p’
| | |
−dS = pdX + q∫X(t)dt
| (15) |
Further integration of eqn (15) by substituting eqn (4) with initial conditions (S = S0; t = 0)
| |
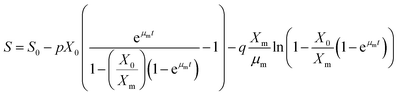 | (16) |
The eqn (16) is written in the other form to estimate ‘p’
| | |
S = S0 − pγ(t) − qδ(t)
| (17) |
| |
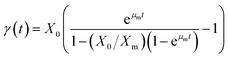 | (18) |
| |
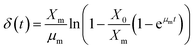 | (19) |
The value of ‘p’ can be determined from the slope of the plot between (S0 − Sq − δ(t)) and γ(t).
2.13.4. Estimation of the kinetic model. Fundamental microbial growth kinetics may follow a linear expression. The microbial growth pattern involves complex parameters that are difficult to explain using linear models. The complex parameters involved in the biomass (X), product (P), and substrate (S) kinetics can be described more effectively by nonlinear least-squares regression using nonlinear expressions. These models reduce the error that may arise from inaccurate variable estimates in linear models. Further, in the present study, biomass kinetics (X), product kinetics (P), and substrate kinetics (S) for the batch bioreactor studies were estimated using the MATLAB command ‘lsqcurvefit’ with non-linear expressions and iterative nonlinear least-squares regression. The MATLAB Simulink tool was used to compare the mathematical and predicted values across several constants and variables. The accuracy of the models was finally analysed using the root mean square error and correlation regression coefficient.| |
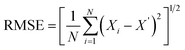 | (20) |
| |
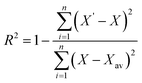 | (21) |
where X, X′ and Xav are experimental, predicted and average biomass values, respectively.
2.14. Purification of DLA
The fermentation broth was initially centrifuged at 8000 rpm to remove the biomass, and the supernatant was further filtered under vacuum. The filtered broth is analysed by HPLC (High Pressure Liquid Chromatography) to estimate the initial DLA concentration. Purification of fermentation broth for DLA concentration was performed as the protocol of Kumar et al., (2020),31 with the following modifications. Liquid–liquid extraction forming an aqueous two-phase system (ATPS) with butanol and ammonium sulphate (ATPS) was used for phase separation of DLA and other organic impurities. In a 100 mL conical flask, 10 mL filtered fermentation broth and 5 g (50% w/v) ammonium sulphate were added, and the mixture was mixed until the salt was completely dissolved. For the following reaction mixture, 15 mL of butanol was transferred & vortexed on a rotary shaker at 150 rpm for 2 hours until a clear phase separation formed. The aqueous phase was separated, mixed with fresh butanol, and the process was repeated three times to recover DLA. The organic phase was separated with DLA with rotary evaporation at controlled pressure. The DLA obtained was estimated and characterized accordingly. Fourier Transform Infrared Spectroscopy (FTIR) characterization was analysed using Bruker Alpha II, a compact FTIR characterization of Unpurified DLA, Purified DLA and Standard DLA. All the samples were tested using the Attenuated Total Reflectance (ATR) method. The spectra were recorded on a Nicolet iS50 spectrometer (Thermo Fisher Scientific Inc., Waltham, MA, USA) in the 400–4000 cm−1 wavelength range. Nuclear Magnetic Resonance (NMR) characterisation was performed on an Avance III HD 400 MHz instrument using 1H NMR with CDCl3 as the solvent. A total of three samples were selected: unpurified DLA (UDLA), purified DLA (P-DLA), and standard DLA (DLA).
2.15. Analytical methods
1 mL of samples was collected from each fermentation study and centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm and 25 °C. The latter supernatant was filtered using a 0.22 µm syringe filter and used for organic acid and glucose analysis. A cell pellet was used to estimate biomass concentration according to the protocol described by Chandukishore et al. (2025).26 Lactic acid & acetic acid were analysed through HPLC using Rezex ROA-Organic Acid H+ (8%) ion exclusion column (7.8 × 150 mm, Phenomenex) with a flow rate of 0.4 (mL minute−1) and oven temperature of 40 °C with 5 mM H2SO4 as the mobile phase equipped with a photodiode detector at 210 nm wavelength. Glucose was estimated using the GOD-POD kit (Excel Diagnostics) at a wavelength of 505 nm.
000 rpm and 25 °C. The latter supernatant was filtered using a 0.22 µm syringe filter and used for organic acid and glucose analysis. A cell pellet was used to estimate biomass concentration according to the protocol described by Chandukishore et al. (2025).26 Lactic acid & acetic acid were analysed through HPLC using Rezex ROA-Organic Acid H+ (8%) ion exclusion column (7.8 × 150 mm, Phenomenex) with a flow rate of 0.4 (mL minute−1) and oven temperature of 40 °C with 5 mM H2SO4 as the mobile phase equipped with a photodiode detector at 210 nm wavelength. Glucose was estimated using the GOD-POD kit (Excel Diagnostics) at a wavelength of 505 nm.
3. Results and discussions
In our previous study, the Ydl-Acs (Y. lipolytica engineered with the ldhA gene and overexpressing the Acetyl CoA synthase (ACS2) gene from Saccharomyces cerevisiae (S288C) strain was cultured in YEPD medium, showing 0.7 (g g−1) of DLA production).26 The complex YEPD medium poses an economic challenge for the industrial production of DLA. The YDl-Acs strain was cultured in minimal medium to analyse and optimise the individual effects of the components involved in DLA production.
3.1. Substrate inhibition studies
FWH obtained from fungal hydrolysis was composed of 47.5 (g L−1) of glucose with a yield of 0.49 (g g−1), free amino nitrogen of 125 (mg L−1) and 155 (mg L−1) of inorganic phosphate. So, after concentrating FWH with a rotary evaporator, the glucose concentration was increased to 93.4 (g L−1). The preliminary control studies with minimal medium composed of 20 (g L−1) glucose and devoid of external nitrogen sources produced lower DLA yield (data not shown).
The concentration of the carbon source is a limiting factor in any lactic acid fermentation studies.5 The DLA yield increased linearly with glucose concentration in FWH up to 5%, but further increases resulted in potent inhibition at 8% glucose. It can be observed in Fig. 1A that at 8% glucose concentration, lactic acid yield was as low as 0.164 (g g−1) with a titer of 13.45 (g L−1), and the highest lactic acid yield of 0.567 (g g−1) was observed at 5% glucose with a titer of 28.65 (g L−1). A possible reason for the growth inhibition could be a high glucose concentration, which may lead to the formation of toxic substances, such as organic acids and ethanol.32 Higher glucose concentrations increase the cell's oxidation–reduction potential (ORP) and shift the stable ORP across the cell membrane, leading to transcriptional imbalance and deleterious effects on the cell, as well as lactic acid formation.33,34 When glucose concentration exceeds the threshold, it can lead to accumulation of carboxylic acids in the intracellular space, increasing the acidity of the cells & fermentation medium.35 In Y. lipolytica, high glucose promotes rapid glycolysis and leads to byproducts such as erythritol, arabitol, and mannitol, rather than fermentation-derived lactate.36 Even a minimal medium composition with a low nitrogen concentration could be a possible reason for the low lactic acid yield37 (Fig. 1A).
 |
| | Fig. 1 (A) Substrate inhibition studies for glucose in FWH (B) effect of nitrogen sources on biomass and DLA production; ammonium sulphate (AS), Casein hydrolysate (CH), fish protein hydrolysate (FPH), urea (U) and yeast extract (YE). | |
3.2. Effect of nitrogen sources on lactic acid production
Nitrogen sources play a vital role in fermentation by providing amino acids, coenzymes, enzymes, and nucleotides required for cell growth and metabolite formation. Incorporation of organic nitrogen in fermentation increases the fermentation rate and reduces the overall fermentation period.38,39 Additionally, organic nitrogen influences the types and concentrations of carboxylic acids produced by yeast.39 In lactic acid fermentation, although glucose concentration is the rate-limiting factor, yeast extract concentration as a nitrogen source is also a key constraint.18 Studies have reported that inorganic ammonium salts are less effective than organic ammonium salts in maintaining yeast viability and promoting product formation. Studies have reiterated the role of yeast extract in the growth and production of Yarrowia lipolytica. In the present study, attempts to partially or fully replace yeast extract with other nitrogen sources showed a preliminary success when FWH was used as the carbon source. The initial nitrogen concentrations of all the nitrogen sources studied were kept constant. Fig. 1B shows that FPH (17.05 g L−1) was a better nitrogen source than the other tested alternatives; however, its yield was still lower than that of yeast extract (19.33 g L−1). Compared to FPH, urea was also competitive, but FPH's organic nature could make it a better choice for lactic acid production. Studies have reported that utilising urea as the sole nitrogen source can't provide essential amino acids and peptides.40 Fig. 1B also showed that DLA yields varied widely with the nitrogen source, whereas biomass production varied only slightly. This could be due to nitrogen supporting only biomass formation, while the bioactive compounds and micronutrients required for lactic acid production were provided only by yeast extract and FPH.24,41 Studies reported that the type of nitrogen source significantly affects the optical purity of DLA.42 Fish protein hydrolysate produced by acid hydrolysis contains essential amino acids and bioactive peptides, making it a significant nitrogen source for lactic acid production.43 The preliminary study showed that FPH may be an attractive option but needs to be analysed for effects across varying concentrations to fully replace yeast extract. With respect to approximation, considering the market trends, yeast extract costs around ($1000–7000 per tonne).44 Relatively, fish waste costs around ($0–200 per tonne) and fish waste hydrolysate costs around ($200–500 per tonne).45 Although processing fish waste to produce an enriched hydrolysate adds cost to fish-waste studies, it is often more economical than YE. Typically, the nutrient composition of YE dominates marginally, but several studies also prove that fish protein waste could be a good replacement for YE in lactic acid production.24 In general, a minimalist YE could cost around $ 5 per kg, and fish protein hydrolysate costs around $0.5 per kg. This can reduce the medium cost by 25–35% in the lactic acid production cost.44 Y. lipolytica is also known for assimilation of complex carbon and nitrogen sources, which helps in the ease of selection of waste biomass and reducing the overall cost.46
3.3. Substrate inhibition model
The Monod model explains the utilization of substrate for the growth of the organism. It also has a few limitations: the model can't explain specific growth rates at high substrate concentrations, and at low concentrations, specific growth is more dependent on substrate concentration. It also can't explain the inhibition caused by toxic substrates in waste biomass, such as FWH and wastewater, when used as substrates.47 The Monod model also can't explain the substrate's contribution to cell maintenance during the organism's lag and death phases.48 Unstructured substrate inhibition models are developed to address the limitations of the Monod model. The Luong (R2-0.933) model showed the highest regression coefficient (R2) out of several models analysed. Though a few models fit to a certain extent, other models couldn't fit and explain the present study conditions (Table 1). The Luong model is an extension of Monod kinetics that includes an additional term 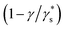 , which explains the lower, upper, and threshold limits of the optimal substrate concentration. The Haldane model is the preferred model to explain substrate inhibition at high substrate concentrations, and inhibition is explained by a toxic substrate with an inhibition constant (KI). The Yano model with two positive constants explained the substrate inhibition with both toxic and non-toxic substrates.29 The other inhibition models did not satisfactorily explain the inhibition kinetics of the lactic acid from FWH. Optimization of industrial production of biochemicals with fermentation poses several challenges in bioprocess optimization. Substrate inhibition models address many such problems by aiding bioreactor design for high-density fermentation and fermentation at high substrate concentrations.49 They also help in deciding the substrate concentration for feeding strategies in fed-batch fermentation. In continuous culture systems, they help determine reactor sizing and dilution rate. Even large-scale fermentation is directly affected by substrate concentration both economically and technically.48 In the Luong model, the critical substrate concentration (Sm) also has practical importance, helping to understand the threshold concentration beyond which microbial growth ceases completely.50 The model's compatibility was calculated using the Akaike information criterion (AIC) as defined in eqn (7). The Haldane and Luong models were chosen for AIC analysis because they achieved the Highest R2. The AIC values for the two models are tabulated in Table 2. The ΔAIC values were analysed using 3- and 4-parameter models; positive ΔAIC values indicate that the simpler Haldane model can comfortably explain the substrate inhibition kinetics of glucose (Fig. S1).
, which explains the lower, upper, and threshold limits of the optimal substrate concentration. The Haldane model is the preferred model to explain substrate inhibition at high substrate concentrations, and inhibition is explained by a toxic substrate with an inhibition constant (KI). The Yano model with two positive constants explained the substrate inhibition with both toxic and non-toxic substrates.29 The other inhibition models did not satisfactorily explain the inhibition kinetics of the lactic acid from FWH. Optimization of industrial production of biochemicals with fermentation poses several challenges in bioprocess optimization. Substrate inhibition models address many such problems by aiding bioreactor design for high-density fermentation and fermentation at high substrate concentrations.49 They also help in deciding the substrate concentration for feeding strategies in fed-batch fermentation. In continuous culture systems, they help determine reactor sizing and dilution rate. Even large-scale fermentation is directly affected by substrate concentration both economically and technically.48 In the Luong model, the critical substrate concentration (Sm) also has practical importance, helping to understand the threshold concentration beyond which microbial growth ceases completely.50 The model's compatibility was calculated using the Akaike information criterion (AIC) as defined in eqn (7). The Haldane and Luong models were chosen for AIC analysis because they achieved the Highest R2. The AIC values for the two models are tabulated in Table 2. The ΔAIC values were analysed using 3- and 4-parameter models; positive ΔAIC values indicate that the simpler Haldane model can comfortably explain the substrate inhibition kinetics of glucose (Fig. S1).
Table 1 Kinetic parameters estimated for glucose substrate inhibition models
| Model |
Parameter |
Df |
µmax |
KI |
Ks |
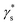
|
K |
R2 |
RMSE |
| Andrew |
3 |
3 |
0.34 |
34.17 |
34.19 |
— |
— |
0.74 |
0.0050 |
| Haldane |
3 |
2 |
1.73 |
4.035 |
370.3 |
— |
— |
0.88 |
0.0035 |
| Luong |
4 |
2 |
0.24 |
— |
40.24 |
134.9 |
— |
0.93 |
0.0025 |
| Yano |
4 |
2 |
1.59 |
269.4 |
552.9 |
|
1.28 |
0.92 |
0.0043 |
Table 2 Model compatibility test using Akaike's information
| Models |
Model specific information |
Akaike's information criterion |
| prm |
P |
SS (·10−5) |
Df |
AICc |
ΔAICc |
pAICc |
| Haldane model |
3 |
6 |
3.78 |
3 |
−61.84 |
|
1 |
| Luong model |
4 |
6 |
1.99 |
2 |
−65.69 |
3.851 |
0.12 |
3.4. Screening of parameters for DLA production
The output response, DLA, was significantly affected by 3 parameters (FWH, pH & KH2PO4) out of 9 parameters screened using a PB design. Substrate inhibition studies show that glucose concentration plays a key role in LA fermentation. Even the PB studies reasserted the importance of FWH in DLA yield, with the highest significance factor (S) among the 9 parameters, as shown in the Pareto chart (Fig. S2). pH is the second significant parameter in the screening design that affects lactic acid production. The optimal pH for D-lactate dehydrogenase from Klebsiella pneumoniae generally falls within the alkaline range and clearly affects lactic acid yield in Yarrowia lipolytica, as shown in our previous study.26 KH2PO4 plays a crucial role in lactic acid fermentation by providing essential nutrients in the form of phosphorus & potassium, and also acts as a buffering medium to maintain the pH.51 Similar results on the importance of KH2PO4 in yeast cell growth were reported in studies of Pichia pastoris producing human interferon-γ.52 In terms of nitrogen source, FPH was significantly better than yeast extract in determining lactic acid yield. Though inoculum size did not show a direct effect on lactic acid production, when OD600 was as high as 2, it reduced the DLA yield. YNB and ammonium chloride showed a minimal impact on the lactic acid production. In some yeast species, ammonium chloride cannot be assimilated as readily as amino acids, and it also adds osmotic stress due to chloride ions.53 The single-order regression eqn (1) is used to fit the Plackett–Burman design. P and t values for each parameter were calculated using analysis of variance and the Student's t-test. The parameter with a high t value and a low P value is designated as a significant factor (Fig. S2) illustrates the Pareto chart consisting of horizontal bars representing the parameters S value; parameters crossing the S value (S = 2.78) are considered significant factors. This is also evident in the normal plot of the standardised effects of the variables. The response for all variables is shown in the (Table 3) and ANOVA for variables is shown in (Table S4) with R2-0.995, Rpred2-0.732 & Radj2-0.984 which shows the ‘goodness of fit’ for the model is significant. The model couldn't explain only 0.5% of the variation in the design. ANOVA results also showed that the model is significant (P < 0.05), and the lack-of-fit value (P = 0.132) supports the model's accuracy. Based on the above results, the three factors—FWH, pH, and K2HPO4—were selected for further optimisation studies. The other parameters had only a marginal effect on the DLA yield, so they were set at a middle level for further studies.
Table 3 Placket–Burman design for screening parameters for DLA production
| Run number |
Yeast extract (g) |
FPH (g) |
FWH (g) |
YNB (W/o AA) (g) |
Ammonium chloride (g) |
K2HPO4 (g) |
KH2PO4 (g) |
Inoculum size |
pH |
DLA (g g−1) |
| 1 |
15.0 |
10.0 |
60 |
1.0 |
1.0 |
2 |
6 |
2.00 |
9 |
0.82 |
| 2 |
15.0 |
15.0 |
30 |
2.0 |
1.0 |
2 |
2 |
2.00 |
9 |
0.45 |
| 3 |
10.0 |
15.0 |
60 |
1.0 |
2.0 |
2 |
2 |
0.50 |
9 |
0.79 |
| 4 |
15.0 |
10.0 |
60 |
2.0 |
1.0 |
6 |
2 |
0.50 |
3 |
0.54 |
| 5 |
15.0 |
15.0 |
30 |
2.0 |
2.0 |
2 |
6 |
0.50 |
3 |
0.29 |
| 6 |
15.0 |
15.0 |
60 |
1.0 |
2.0 |
6 |
2 |
2.00 |
3 |
0.63 |
| 7 |
10.0 |
15.0 |
60 |
2.0 |
1.0 |
6 |
6 |
0.50 |
9 |
0.81 |
| 8 |
10.0 |
10.0 |
60 |
2.0 |
2.0 |
2 |
6 |
2.00 |
3 |
0.65 |
| 9 |
10.0 |
10.0 |
30 |
2.0 |
2.0 |
6 |
2 |
2.00 |
9 |
0.38 |
| 10 |
15.0 |
10.0 |
30 |
1.0 |
2.0 |
6 |
6 |
0.50 |
9 |
0.41 |
| 11 |
10.0 |
15.0 |
30 |
1.0 |
1.0 |
6 |
6 |
2.00 |
3 |
0.25 |
| 12 |
10.0 |
10.0 |
30 |
1.0 |
1.0 |
2 |
2 |
0.50 |
3 |
0.24 |
| 13 |
12.5 |
12.5 |
45 |
1.5 |
1.5 |
4 |
4 |
1.25 |
6 |
0.66 |
| 14 |
12.5 |
12.5 |
45 |
1.5 |
1.5 |
4 |
4 |
1.25 |
6 |
0.67 |
| 15 |
12.5 |
12.5 |
45 |
1.5 |
1.5 |
4 |
4 |
1.25 |
6 |
0.69 |
3.5. Central composite design (CCD) for optimization of lactic acid production
The CCD statistical approach was implemented to optimise the three significant factors (FWH, pH and K2HPO4) obtained from the Plackett–Burman design. The CCD design considers both independent variables & interaction variables to design a total number of runs, with 6 center points, which were designed with a polynomial expression to optimise the response DLA yield. The model-predicted responses for all 20 runs are shown in Table 4. Run 16 had the highest lactic acid yield of 0.90 (g g−1) with 40 (g L−1) glucose, pH 8.5, and KH2PO4 4 (g L−1). Additionally, run 16th also reiterated the importance of FWH as a limiting factor in lactic acid production, with pH playing a driving role in the enzymatic conversion of FWH to DLA. In CCD studies, the FPH concentration was increased by 20% relative to the yeast extract concentration to assess the effect of FPH as a major nitrogen source and to replace YE completely, as a part of utilising an economical nitrogen source. This change in concentration of FPH to replace YE didn't show any marginal effect on DLA yield. The regression coefficients for the model were R2 = 0.97, Rpred2 = 0.83, and Radj2 = 0.95. The ANOVA results for the DLA response showed a significant model (P < 0.05) and a lack of fit (P > 0.05). The same is demonstrated in Table S5.
Table 4 Central composite design for optimization of parameters for DLA production
| Run order |
FWH (g L−1) |
pH |
KH2PO4 (g L−1) |
DLA (g g−1) (experimental) |
DLA (g g−1) CCD predicted |
DLA (g g−1) (ANN predicted) |
| 1 |
20 |
7 |
2 |
0.61 ± 0.03 |
0.605 |
0.615 |
| 2 |
60 |
7 |
2 |
0.57 ± 0.02 |
0.607 |
0.588 |
| 3 |
20 |
10 |
2 |
0.36 ± 0.01 |
0.372 |
0.366 |
| 4 |
60 |
10 |
2 |
0.34 ± 0.01 |
0.392 |
0.343 |
| 5 |
20 |
7 |
6 |
0.55 ± 0.02 |
0.560 |
0.558 |
| 6 |
60 |
7 |
6 |
0.60 ± 0.02 |
0.649 |
0.562 |
| 7 |
20 |
10 |
6 |
0.30 ± 0.01 |
0.324 |
0.305 |
| 8 |
60 |
10 |
6 |
0.36 ± 0.01 |
0.431 |
0.336 |
| 9 |
6.36 |
8.5 |
4 |
0.39 ± 0.01 |
0.408 |
0.576 |
| 10 |
73.63 |
8.5 |
4 |
0.58 ± 0.02 |
0.499 |
0.517 |
| 11 |
40 |
5.97 |
4 |
0.51 ± 0.01 |
0.500 |
0.177 |
| 12 |
40 |
11.02 |
4 |
0.17 ± 0.00 |
0.121 |
0.802 |
| 13 |
40 |
8.5 |
0.63 |
0.80 ± 0.03 |
0.786 |
0.827 |
| 14 |
40 |
8.5 |
7.36 |
0.83 ± 0.03 |
0.781 |
0.903 |
| 15 |
40 |
8.5 |
4 |
0.88 ± 0.04 |
0.901 |
0.903 |
| 16 |
40 |
8.5 |
4 |
0.90 ± 0.04 |
0.901 |
0.903 |
| 17 |
40 |
8.5 |
4 |
0.88 ± 0.04 |
0.901 |
0.903 |
| 18 |
40 |
8.5 |
4 |
0.90 ± 0.04 |
0.901 |
0.903 |
| 19 |
40 |
8.5 |
4 |
0.88 ± 0.04 |
0.901 |
0.903 |
| 20 |
40 |
8.5 |
4 |
0.87 ± 0.03 |
0.901 |
0.903 |
3D surface plots were created for the response DLA yield across all the interacting variables. Fig. 2A shows the 3D surface plot of FWH versus pH, highlighting the domination of FWH in DLA production, but with the increase in the concentration of glucose to 40 g L−1, an inhibition in the DLA yield was observed. Even the pH played a determinant role in DLA yield, but after (pH 8.5), there was a significant drop in the DLA yield. This could be due to the alkaline pH affecting cellular homeostasis by disrupting membrane potential.54,55 The second surface plot of FWH versus KH2PO4 is shown in Fig. 2B, which signifies that an increase in FWH concentration improved the lactic acid production up to 40 (g L−1) only.56 At the same time, KH2PO4 didn't show a direct effect on DLA yield but played a supporting role of stabilising pH, supporting cell growth, enhancing metabolic efficiency and providing essential ions and cofactors up to 4 (g L−1). Later, it inhibited the DLA yield. The final 3D plot of KH2PO4 versus pH showed that though pH played a determinant role in the DLA production, after (pH 8.5) there was a steep decrease in the DLA yield, and KH2PO4 was inherently contributing to the DLA yield for a limited extent only53 (Fig. 2C).
 |
| | Fig. 2 3D surface plots for CCD studies for DLA optimisation studies (A) FWH versus pH (B) FWH versus KH2PO4 (C) pH versus KH2PO4; food waste hydrolysate (FWH). | |
3.6. Artificial neural network and genetic algorithm-based optimisation
The output of the CCD was used as input for an ANN-based optimisation study. ANN has three stages: first, training the input variables to predict the target response; second, validating the model; and third, testing the model. A total of 1000 epochs were considered for training, with 70% for training, 15% for validation and 15% for testing. The ANN output depends on the number of hidden layers formed from the given three input layers, resulting in a final 1-output layer with the lowest minimum mean square error (MSE). In the present study, the ANN at epoch 4 with 8 hidden layers and 1 output predicted response for 20 CCD runs, with run number 17 showing the highest lactic acid yield of 0.912 (g g−1), Fig. 3A and B. Further, the ANN-trained output was used as input for genetic algorithm-based studies with the following parameters: 100 generations, population size of 20, crossover probability of 0.8, and mutation probability of 0.01. The genetic algorithm generated an output response at 50th iteration with 0.944 (g g−1) of DLA yield and 12.6% increment compared to initial Placket–Burman studies (Fig. 3C). Following the output generated the optimised global input variables are as follows FWH with glucose concentration 49.98 (g L−1), pH (8.52) and KH2PO4 concentration of 3.93 (g L−1) (Fig. 3D). With the GA based optimization after CCD studies overall there was 12.6% increase in the DLA yield.
 |
| | Fig. 3 ANN and GA optimisation studies for D-lactic acid production (A) regression results of training, validation, text and overall regression (B) ANN normalization chart (C) GA iterations (D) final optimised GA values for the parameters. | |
3.7. Validation of the optimized parameters
The output of the ANN-GA-optimised variables was used in the validation study, as described in the methodology section. The verification studies, including shake flask study experiments, results supported the GA prediction with a DLA yield of 0.92 (g g−1). Additionally, substrate utilisation increased from 40 (g L−1) to 50 (g L−1) in the CCD studies, as predicted by ANN-GA optimisation. Biomass accumulation was also marginally improved compared with the CCD and PB studies. Further scale-up studies were performed to understand the effect of optimised parameters in the bioreactor (Table 5). The scale up considerations for bioreactor studies are given in Tables S7 and S8.
Table 5 Comparative yields of strains producing lactic acid
| Strain |
Type of organism |
Substrate |
Batch or fed batch |
Yield (g g−1) |
References |
| C. sonorensis |
Mutant |
Glucose |
Batch |
0.94 |
57 |
| K. pneumoniae |
Mutant |
Glucose |
Fed batch |
0.91 |
58 |
| S. cerevisiae |
Mutant |
Glucose |
— |
— |
59 |
| S. cerevisiae |
Mutant |
Glucose |
Fed batch |
0.80 |
61 |
| K. pneumoniae |
Wild type |
Glycerol |
Fed batch |
0.82 |
62 |
| S. cerevisiae |
Mutant |
Glucose |
Fed batch |
0.83 |
60 |
| Komagataella phaffii |
Mutant |
Methanol |
— |
— |
63 |
| S. cerevisiae |
Mutant |
Xylose |
Batch |
0.84 |
64 |
| Lactobacillus delbrueckii |
Wild type |
Food waste |
Batch |
0.91 |
65 |
| Bacillus coagulans DSM1 |
Wild type |
Bread waste hydrolysate |
Fed batch |
0.85 |
34 |
| Yarrowia lipolytica (YDl-Acs) |
Mutant |
Food waste hydrolysate |
Batch |
0.94 |
This study |
3.8. Kinetic modelling
3.8.1. Growth kinetics. Batch kinetics studies of Y. lipolytica YDl-Acs producing DLA performed in a bioreactor showed regular standard growth pattern with FWH as the sole carbon source. A significant increase in the biomass was observed between 12 and 36 hours; thereafter, there was a slow decline in the biomass accumulation. The logistic eqn (5) was used to fit the growth kinetics data, which revealed that the maximum specific growth rate (µm) was 0.124 h−1 and the initial biomass concentration (X0) was −0.384 (g L−1). The slope and intercept of the logistic equation will represent (µm) & (X0), and also experimental values matched the model-predicted values. The maximum growth (Xm) of 9.7 (g L−1) was achieved by 48 hours, but later the biomass accumulation was stationary because of depletion of nutrients. The logistic equation with maximum growth data sufficiently explained the sigmoidal growth pattern of Y. lipolytica.
3.8.2. Product kinetics. Though biomass is also an important parameter, in non-growth-associated fermentation studies, the pattern of product kinetics plays a crucial role in determining overall yield, titer, and productivity. The Luedeking–Piret model was used to evaluate the DLA product kinetics through eqn (9). The product data points at the stationary phase were considered to calculate the growth-associated constant (m) and the non-growth-associated constant (n). In the present study, the (mexp) value was 0.701 & (nexp) was 0.21, whereas (mpre) was 0.285, and (npre) was 0.11. With a slight difference between m and n, the lactic acid production showed a mixed fermentation pattern. The possible reason for the mixed fermentation could be the high accumulation of lactic acid during the stationary-phase growth and under microaerobic conditions.26 Several yeast and bacterial studies support the above theory of a high lactic acid yield at the stationary phase.66
3.8.3. Substrate kinetics. An initial glucose substrate concentration of 50 (g L−1) of glucose was maintained, further glucose concentration gradually declined with an increase in the biomass and DLA concentration and maximum glucose depletion was observed between 12–60 hours. The logistic eqn (16), plotted against substrate kinetics data and its intercept, yielded the maintenance coefficient (ms). The maintenance coefficient (ms) is a parameter that describes the total substrate uptake rate for cellular maintenance. The maintenance coefficient for the present study was analysed using the intercept of the logistic eqn (16), which was 0.11 (gsubstrate gbiomass−1 h−1). Studies reported by C. Li et al. (2022) on producing succinic acid with glycerol using Y. lipolytica reported ms values of 0.118 g (g−1 h−1), which is within the reasonably reported range.49
3.8.4. Matlab model testing. Matlab 2016b was used to analyse and estimate the kinetic parameters (µm, X0, Xm, m, n, p, q, Yx/s, and Yp/s). The following parameters were used to test the model. The predicted and experimental parameters of the kinetic model were tabulated in Table 6. Fig. 4 showed the relationship between the predicted and experimental values of X, P, and S, thereby reiterating the accuracy of batch studies of DLA production. A Simulink model was used to simulate studies on DLA production by Y. lipolytica (YDl-Acs). The R2 values for X, P, and S were 0.92, 0.97, and 0.96, respectively, confirming the model's accuracy with experimental results (Fig. 5).
Table 6 Experimental and predicted parameters for the kinetic model for DLA production
| Model parameters |
Experimental value |
Predicted value |
| X0 (g L−1) |
0.38 |
0.38 |
| Xm (g L−1) |
9.70 |
9.68 |
| µm (h−1) |
0.12 |
0.09 |
| Alpha (g gbiomass−1) |
0.72 |
0.28 |
| Beta (U g−1 h−1) |
0.18 |
0.10 |
| p (gbiomass gglucose−1) |
1.78 |
1.64 |
| Q (gglucose gbiomass−1 h−1) |
0.14 |
0.11 |
 |
| | Fig. 4 Comparative values of experimental and model predicted kinetic parameters for batch production of D-lactic acid. | |
 |
| | Fig. 5 Matlab predicted kinetic model for batch production of D-lactic acid. | |
3.9. Purification and characterisation of DLA
The fermentation broth obtained had a DLA concentration of 43.5 (g L−1). Butanol forms butyl lactate upon association with LA, which then moves to the upper organic layer, leaving other organic impurities in the layer below. Protein impurities were concentrated at the junction of two layers. Initially, fermentation was performed with glucose as the sole carbon source, yielding a high DLA yield; the product was purified with less complexity than when FWH was used as the carbon source. The DLA produced from glucose was purified with 92.7% recovery, and DLA purity increased 22.5-fold. Compared with FWH, when FWH was used as a carbon source, DLA was extracted with 83.51% recovery and purity was 16.72-fold higher. Studies reported that varying the load of crude lactic acid significantly impacts the effect of impurities in food waste on phase separation. In a study on utilising fruit waste to produce lipase, it was reported that varying the crude load by 25% reduced the effect of impurities on the separation of molecules. Chakraborty and Ghalsasi (2025) reported that FTIR characterisation of the produced DLA depicted a broad peak at 3312 and 1612. Similar results were observed in the present study and are shown in (Fig. S3).67 1H NMR studies with CDCl3 as solvent were analysed for (A) DLA standard, (B) Fermented broth and (C) Purified DLA. Bourafai-Aziez et al. (2022) reported that the presence of a doublet at 1.3–1.45 ppm and a quartet at 4.3 ppm emphasised the presence of LA; similar results were observed in the present study with the DLA standard and purified DLA (Fig. 6).68 Similarly, studies by Chemama et al. (2020)69 reported that 1H NMR characterisation of DLA showed chemical shifts reported in parts per million (ppm) at 1.3 (CH3) and 4.3 (CH). Similar results were reported in the present study and also by Pulido et al. (2021).70
 |
| | Fig. 6 1H NMR characterization of D-lactic acid (A) DLA standard (B) unpurified fermentation broth (C) purified DLA. | |
4. Conclusion
The successful use of FWH and FPH as the sole carbon and nitrogen sources for producing DLA from engineered Y. lipolytica highlights a promising approach for sustainable bioproduction. FPH proved to be an effective nitrogen source, while optimisation studies identified the ideal conditions for maximising DLA yields. Substrate inhibition models, based on an extended Monod model, explained the impact of threshold substrate concentrations on specific growth rates and DLA production. Kinetic modelling using MATLAB confirmed the model's accuracy by comparing experimental and predicted values for biomass, product formation, and substrate consumption. Downstream processing of the fermentation broth revealed that FWH exhibited inhibitory effects, as evidenced by its lower recovery efficiency than glucose. Despite these challenges, the use of waste biomass, such as FWH, offers an environmentally friendly and economically viable carbon source, contributing to waste valorisation and reducing reliance on food-based feedstocks, thereby addressing environmental sustainability and safety concerns.
Author contributions
Chandukishore T., Ashish A. Prabhu are involved in designing the idea for the current study. Chandukishore T. carried out the experiment. Ashish A. Prabhu and K. Narsimhulu supervised the findings of this work. Chandukishore T. wrote the manuscript and interpreted the results. All authors provided critical feedback and helped shape the research, analysis, and manuscript.
Conflicts of interest
The authors declare that they have no competing interests.
Data availability
All data generated or analyzed during this study are included in the manuscript.
Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra08276e.
Acknowledgements
We acknowledge NIT Warangal for providing the necessary facilities to accomplish this project. The authors also thank Dr Rodrigo Ledesma-Amaro from Imperial College London, UK, for providing strains and plasmids.
References
- T. A. Swetha, V. Ananthi, A. Bora, N. Sengottuvelan, K. Ponnuchamy, G. Muthusamy and A. Arun, Int. J. Biol. Macromol., 2025, 234, 123703 CrossRef PubMed.
- G. G. N. Thushari and J. D. M. Senevirathna, Heliyon, 2020, 6, e04709 CrossRef.
- M. M. Islam, S. Chaudry, A. W. Thornton, N. Haque, D. Lau, M. Bhuiyan and B. K. Pramanik, J. Cleaner Prod., 2025, 496, 145132 CrossRef CAS.
- H. Rahaman, S. Hosen, A. Gafur and R. Habib, Results Mater., 2020, 8, 100125 CrossRef.
- T. Chandukishore, A. P. Ashish and K. Narasimhulu, Sep. Purif. Rev., 2024, 1–17 Search PubMed.
- N. M. Salatein, A. F. Omara, A. R. Mansour and I. S. Fahim, Results Eng., 2025, 25, 104184 CrossRef CAS.
- G. Policastro, F. Carraturo, M. Compagnone, M. Giugliano, M. Guida, V. Luongo, R. Napolitano and M. Fabbricino, Bioresour. Technol., 2021, 340, 125595 CrossRef CAS PubMed.
- S. K. Maity, D. Agrawal, S. Gadkari, K. R. Vanapalli, Y. C. Yong, D. Zhu, C. Chen and V. Kumar, ACS Sustain. Chem. Eng., 2025, 18, 6538–6553 CrossRef.
- P. Sivagurunathan, T. Raj, P. S. Chauhan, P. Kumari, A. Satlewal, R. P. Gupta and R. Kumar, Biochem. Eng. J., 2022, 187, 108668 CrossRef CAS.
- L. Song, S. Liu, R. Liu, D. Yang and X. Dai, Sci. Total Environ., 2022, 840, 156479 CrossRef CAS.
- W. C. Lam, D. Pleissner and C. S. K. Lin, Biomolecules, 2013, 3, 651–661 CrossRef.
- S. Ranganathan, S. Dutta, J. A. Moses and C. Anandharamakrishnan, Heliyon, 2020, 6, e04891 CrossRef CAS PubMed.
- D. Pleissner and C. S. K. Lin, Sustainable Chem. Processes, 2013, 1, 1–6 Search PubMed.
- M. O. Ramadhan and M. N. Handayani, IOP Conf. Ser.: Mater. Sci. Eng., 2020, 980, 012082 CAS.
- L. Panzella, F. Moccia, R. Nasti, S. Marzorati, L. Verotta and A. Napolitano, Front. Nutr., 2020, 7, 1–27 Search PubMed.
- L. Bhatia, D. S. V. G. K. Kaladhar, T. Sarkar, H. Jha and B. Kumar, Energy, Ecol. Environ., 2024, 9, 455–485 CrossRef CAS.
- V. Novy, B. Brunner and B. Nidetzky, Microb. Cell Fact., 2018, 17, 59 CrossRef PubMed.
- A. K. Michalczyk, S. Garbaczewska, B. Morytz, A. Białek and J. Zakrzewski, Fermentation, 2021, 7, 1–11 CrossRef.
- M. Papizadeh, M. Rohani, S. N. Hosseini, S. A. Shojaosadati, H. Nahrevanian, M. Talebi and M. R. Pourshafie, AMB Express, 2020, 10, 53 CrossRef CAS.
- T. Lian, W. Zhang, Q. Cao, S. Wang, F. Yin, T. Zhou, F. Zhang and H. Dong, Chem. Eng. J., 2023, 471, 144689 CrossRef CAS.
- S. Ngasotter, Int. J. Bio-Resour. Stress Manage., 2020, 11, 370–380 Search PubMed.
- A. P. Tom, J. S. Jayakumar, M. Biju, J. Somarajan and M. A. Ibrahim, Energy Nexus, 2021, 4, 100022 Search PubMed.
- T. B. Ribeiro, M. R. G. Maia, A. J. M. Fonseca, B. Marques, C. Caleja, A. Rosa, R. Martins, A. Almeida, M. J. Mota, T. Aires, C. M. R. Rocha, J. A. Teixeira, A. R. J. Cabrita, L. Barros and M. Pintado, Food Rev. Int., 2024, 1–39 CAS.
- M. T. Gao, M. Hirata, E. Toorisaka and T. Hano, Bioresour. Technol., 2006, 97, 2414–2420 Search PubMed.
- T. Chandukishore, S. Das, K. Narasimhulu and A. A. Prabhu, Sustainable Chem. Pharm., 2023, 35, 101230 Search PubMed.
- T. Chandukishore, K. Narasimhulu, R. Ledesma-Amaro and A. A. Prabhu, Chem. Eng. J., 2025, 165519 CrossRef CAS.
- P. Wongsirichot, M. Costa, B. Dolman, M. Freer, A. Welfle and J. Winterburn, Resour. Conserv. Recycl., 2022, 185, 106499 CrossRef CAS.
- V. Narisetty, A. A. Prabhu, R. R. Bommareddy, R. Cox, D. Agrawal, A. Misra, M. A. Haider, A. Bhatnagar, A. Pandey and V. Kumar, ACS Sustain. Chem. Eng., 2022, 10, 10858–10869 Search PubMed.
- K. Dutta, V. V. Dasu, B. Mahanty and A. A. Prabhu, Chem. Biochem. Eng. Q., 2015, 29, 437–445 CrossRef CAS.
- G. Broli, H. Nygaard, H. Sletta, K. Sandnes and I. M. Aasen, Process Biochem., 2021, 102, 157–164 CrossRef CAS.
- S. Kumar, N. Yadav, L. Nain and S. K. Khare, Bioresour. Technol., 2020, 318, 124260 CrossRef CAS PubMed.
- A. Komesu, J. A. R. d. Oliveira, L. H. d. S. Martins, M. R. Wolf Maciel and R. Maciel Filho, BioResources, 2017, 12, 4364–4383 CAS.
- C. G. Liu, X. M. Hao, Y. H. Lin and F. W. Bai, Sci. Rep., 2016, 6, 1–7 CrossRef.
- R. Cox, V. Narisetty, S. Nagarajan, D. Agrawal, V. V. Ranade, K. Salonitis, J. Venus and V. Kumar, Fuel, 2022, 313, 122976 CrossRef CAS.
- S. Lane, H. Xu, E. J. Oh, H. Kim, A. Lesmana, D. Jeong, G. Zhang, C. S. Tsai, Y. S. Jin and S. R. Kim, Sci. Rep., 2018, 8, 1–12 Search PubMed.
- C. Li, S. Gao, X. Li, X. Yang and C. S. K. Lin, Biotechnol. Biofuels, 2018, 11, 1–12 CrossRef.
- P. Hapeta, E. J. Kerkhoven and Z. Lazar, Biotechnol. Rep., 2020, 27, e00521 CrossRef PubMed.
- A. Schön, J. Switulla, L. Luksch, J. Pesl, R. Kölling and D. Einfalt, Beverages, 2024, 10, 93 CrossRef.
- J. Zhang, N. Zhao, L. Guo, P. Li, S. Gu, J. Yuan and M. Fan, LWT, 2023, 187, 115376 CrossRef CAS.
- O. Konzock, S. Zaghen, J. Fu and E. J. Kerkhoven, iScience, 2022, 25, 105703 Search PubMed.
- S. I. Aspmo, S. J. Horn and V. G. H. Eijsink, FEMS Microbiol. Lett., 2005, 248, 65–68 CrossRef CAS PubMed.
- S. Thitiprasert, P. Jaiaue, N. Amornbunchai, J. Thammakes, J. Piluk, P. Srimongkol, S. Tanasupawat and N. Thongchul, Sci. Rep., 2024, 14, 1–12 Search PubMed.
- H. Zhang, Y. Zhang, S. K. C. Chang, Y. Luo, H. Hong and Y. Tan, J. Cleaner Prod., 2024, 447, 141076 Search PubMed.
- S. Krull, S. Brock, U. Prüße and A. Kuenz, Fermentation, 2020, 6, 97 Search PubMed.
- S. Shi, J. Li, W. Guan and D. Blersch, RSC Adv., 2018, 8, 31267–31274 Search PubMed.
- T. F. Ferreira, F. F. Martins, C. A. Cayres, P. F. F. Amaral, D. d. A. Azevedo and M. A. Z. Coelho, Fermentation, 2023, 9, 21 CrossRef CAS.
- M. Muloiwa, S. Nyende-Byakika and M. Dinka, S. Afr. J. Chem. Eng., 2020, 33, 141–150 Search PubMed.
- A. A. Prabhu and V. Venkata Dasu, Prep. Biochem. Biotechnol., 2017, 47, 953–962 Search PubMed.
- C. Li, Y. Xiao, Z. Sang, Z. Yang, T. Xu, X. Yang, J. Yan and C. S. K. Lin, Chem. Eng. J., 2022, 442, 136273 Search PubMed.
- C. Pateraki, H. Almqvist, D. Ladakis, G. Lidén, A. A. Koutinas and A. Vlysidis, Biochem. Eng. J., 2016, 114, 26–41 CrossRef CAS.
- N. Ribeiro-Filho, R. Linforth, N. Bora, C. D. Powell and I. D. Fisk, Food Res. Int., 2022, 162, 112044 Search PubMed.
- A. A. Prabhu, B. Mandal and V. V. Dasu, Korean J. Chem. Eng., 2017, 34, 1109–1121 Search PubMed.
- S. A. van der Hoek, M. Rusnák, I. H. Jacobsen, J. L. Martínez, D. B. Kell and I. Borodina, FEBS Lett., 2022, 596, 1356–1364 Search PubMed.
- E. Eskes, M. A. Deprez, T. Wilms and J. Winderickx, Curr. Genet., 2018, 64, 155–161 Search PubMed.
- A. Serra-Cardona, D. Canadell and J. Ariño, Microb. Cell, 2015, 2, 182–196 Search PubMed.
- T. Liu, L. Sun, C. Zhang, Y. Liu, J. Li, G. Du, X. Lv and L. Liu, Bioresour. Technol., 2023, 379, 129023 CrossRef CAS.
- M. Ilmén, K. Koivuranta, L. Ruohonen, V. Rajgarhia, P. Suominen and M. Penttilä, Microb. Cell Fact., 2013, 12, 1–15 CrossRef.
- X. Feng, L. Jiang, X. Han, X. Liu, Z. Zhao, H. Liu, M. Xian and G. Zhao, Microb. Cell Fact., 2017, 16, 1–11 Search PubMed.
- B. K. Jang, Y. Ju, D. Jeong, S. K. Jung, C. K. Kim, Y. S. Chung and S. R. Kim, J. Fungi, 2021, 7, 938 Search PubMed.
- S. Baek, E. Y. Kwon, S. Bae, B. Cho, S. Kim and J. Hahn, Biotechnol. J., 2017, 12, 1700015, DOI:10.1002/biot.201700015.
- S. H. Baek, E. Y. Kwon, Y. H. Kim and J. S. Hahn, Appl. Microbiol. Biotechnol., 2016, 100, 2737–2748 CrossRef CAS.
- X. Feng, Y. Ding, M. Xian, X. Xu, R. Zhang and G. Zhao, Bioresour. Technol., 2014, 172, 269–275 Search PubMed.
- J. Ding, G. Holzwarth, M. H. Penner, J. Patton-Vogt and A. T. Bakalinsky, FEMS Microbiol. Lett., 2015, 362, 1–7 CrossRef CAS PubMed.
- B. Choi, A. Tafur Rangel, E. J. Kerkhoven and Y. Nygård, Metab. Eng., 2024, 84, 23–33 CrossRef CAS PubMed.
- K. Il Kim, W. K. Kim, D. K. Seo, I. S. Yoo, E. K. Kim and H. H. Yoon, Appl. Biochem. Biotechnol., Part A, 2003, 107, 637–647 Search PubMed.
- E. Abedi and S. M. B. Hashemi, Heliyon, 2020, 6, e04974 CrossRef CAS PubMed.
- A. Chakraborty and P. Ghalsasi, Biomass Convers. Biorefin., 2025, 6871–6888 Search PubMed.
- A. Bourafai-Aziez, D. Jacob, G. Charpentier, E. Cassin, G. Rousselot, A. Moing and C. Deborde, Molecules, 2022, 27, 5614 CrossRef CAS PubMed.
- T. Chemama, C. Pumas, Y. Tragoolpua and N. Thongwai, Chiang Mai J. Sci., 2020, 47, 403–417 CAS.
- N. Pulido, J. M. Guevara-Morales, A. Rodriguez-López, Á. Pulido, J. Díaz, R. A. Edrada-Ebel and O. Y. Echeverri-Peña, Metabolites, 2021 DOI:10.3390/metabo11120891.
|
| This journal is © The Royal Society of Chemistry 2026 |
Click here to see how this site uses Cookies. View our privacy policy here.  Open Access Article
Open Access Article * and
K. Narasimhulu
* and
K. Narasimhulu
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 tons of lactic acid (LA) are produced, and demand for LA is increasing by 10% annually.6 LA can be produced by both chemical and biological methods, with 90% of LA production occurring via fermentation.7 The cost of lactic acid depends on its applications, which range from 3.0 to 4.0 $ per kg, which was 1.5–3.0 $ per kg as of 2011.8 Recent market trends reported by the Tridge group indicate that in 2024, the global lactic price ranged from $2.74 per kg to $13.68 per kg. Also, there was a significant decline in the price from $ 1.06 per kg to $6.58 per kg. The reasons are sudden shifts in demand and supply. Annually, 3.3 × 105 tons of lactic acid are produced through the biochemical route using glucose derived from food-based materials as the substrate, contributing to the ongoing food-versus-fuel debate.9 Producing LA primarily from first-generation feedstocks may increase production costs. Further exploiting second- and third-generation renewable feedstocks derived from agro-industrial waste may substantially reduce lactic acid production costs.10 According to the Food Wastage Index Report 2021, globally, 1.3 billion tonnes of food waste are generated annually.11 Of the overall food waste generated, a minor portion is redistributed, composted, or treated by other methods; the majority ends up in landfills.12 Food waste is complex, making it difficult for microbes to hydrolyse it due to their limited metabolic capabilities.13 Food waste contains valuable nutrients and can be effectively used to produce fungal enzymes, such as glucoamylase and proteases, which efficiently break it down.14 Several studies are reporting the utilisation of food waste, either directly or after treatment, to produce bioethanol, biodiesel, organic acids, pigments, antibiotics, enzymes, etc.13,15,16
000 tons of lactic acid (LA) are produced, and demand for LA is increasing by 10% annually.6 LA can be produced by both chemical and biological methods, with 90% of LA production occurring via fermentation.7 The cost of lactic acid depends on its applications, which range from 3.0 to 4.0 $ per kg, which was 1.5–3.0 $ per kg as of 2011.8 Recent market trends reported by the Tridge group indicate that in 2024, the global lactic price ranged from $2.74 per kg to $13.68 per kg. Also, there was a significant decline in the price from $ 1.06 per kg to $6.58 per kg. The reasons are sudden shifts in demand and supply. Annually, 3.3 × 105 tons of lactic acid are produced through the biochemical route using glucose derived from food-based materials as the substrate, contributing to the ongoing food-versus-fuel debate.9 Producing LA primarily from first-generation feedstocks may increase production costs. Further exploiting second- and third-generation renewable feedstocks derived from agro-industrial waste may substantially reduce lactic acid production costs.10 According to the Food Wastage Index Report 2021, globally, 1.3 billion tonnes of food waste are generated annually.11 Of the overall food waste generated, a minor portion is redistributed, composted, or treated by other methods; the majority ends up in landfills.12 Food waste is complex, making it difficult for microbes to hydrolyse it due to their limited metabolic capabilities.13 Food waste contains valuable nutrients and can be effectively used to produce fungal enzymes, such as glucoamylase and proteases, which efficiently break it down.14 Several studies are reporting the utilisation of food waste, either directly or after treatment, to produce bioethanol, biodiesel, organic acids, pigments, antibiotics, enzymes, etc.13,15,16
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4) in a submerged hydrolysis at 55 °C, 150 rpm. Further samples were taken at regular intervals until the FWH was recorded, the optimum glucose concentration, which was analysed using the GOD-POD assay. The obtained FWH was concentrated at 60 °C using a rotary evaporator by vacuum evaporation.27
4) in a submerged hydrolysis at 55 °C, 150 rpm. Further samples were taken at regular intervals until the FWH was recorded, the optimum glucose concentration, which was analysed using the GOD-POD assay. The obtained FWH was concentrated at 60 °C using a rotary evaporator by vacuum evaporation.27
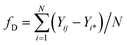
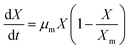
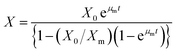
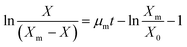
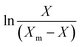 and time (h).
and time (h).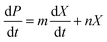
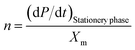
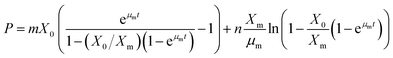
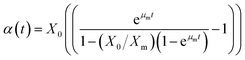
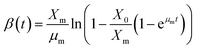
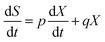
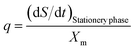
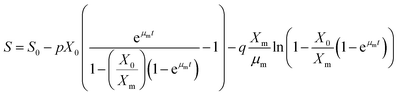
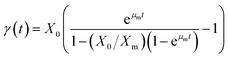
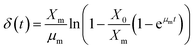
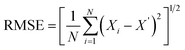
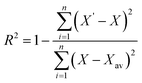
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm and 25 °C. The latter supernatant was filtered using a 0.22 µm syringe filter and used for organic acid and glucose analysis. A cell pellet was used to estimate biomass concentration according to the protocol described by Chandukishore et al. (2025).26 Lactic acid & acetic acid were analysed through HPLC using Rezex ROA-Organic Acid H+ (8%) ion exclusion column (7.8 × 150 mm, Phenomenex) with a flow rate of 0.4 (mL minute−1) and oven temperature of 40 °C with 5 mM H2SO4 as the mobile phase equipped with a photodiode detector at 210 nm wavelength. Glucose was estimated using the GOD-POD kit (Excel Diagnostics) at a wavelength of 505 nm.
000 rpm and 25 °C. The latter supernatant was filtered using a 0.22 µm syringe filter and used for organic acid and glucose analysis. A cell pellet was used to estimate biomass concentration according to the protocol described by Chandukishore et al. (2025).26 Lactic acid & acetic acid were analysed through HPLC using Rezex ROA-Organic Acid H+ (8%) ion exclusion column (7.8 × 150 mm, Phenomenex) with a flow rate of 0.4 (mL minute−1) and oven temperature of 40 °C with 5 mM H2SO4 as the mobile phase equipped with a photodiode detector at 210 nm wavelength. Glucose was estimated using the GOD-POD kit (Excel Diagnostics) at a wavelength of 505 nm.
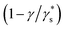 , which explains the lower, upper, and threshold limits of the optimal substrate concentration. The Haldane model is the preferred model to explain substrate inhibition at high substrate concentrations, and inhibition is explained by a toxic substrate with an inhibition constant (KI). The Yano model with two positive constants explained the substrate inhibition with both toxic and non-toxic substrates.29 The other inhibition models did not satisfactorily explain the inhibition kinetics of the lactic acid from FWH. Optimization of industrial production of biochemicals with fermentation poses several challenges in bioprocess optimization. Substrate inhibition models address many such problems by aiding bioreactor design for high-density fermentation and fermentation at high substrate concentrations.49 They also help in deciding the substrate concentration for feeding strategies in fed-batch fermentation. In continuous culture systems, they help determine reactor sizing and dilution rate. Even large-scale fermentation is directly affected by substrate concentration both economically and technically.48 In the Luong model, the critical substrate concentration (Sm) also has practical importance, helping to understand the threshold concentration beyond which microbial growth ceases completely.50 The model's compatibility was calculated using the Akaike information criterion (AIC) as defined in eqn (7). The Haldane and Luong models were chosen for AIC analysis because they achieved the Highest R2. The AIC values for the two models are tabulated in Table 2. The ΔAIC values were analysed using 3- and 4-parameter models; positive ΔAIC values indicate that the simpler Haldane model can comfortably explain the substrate inhibition kinetics of glucose (Fig. S1).
, which explains the lower, upper, and threshold limits of the optimal substrate concentration. The Haldane model is the preferred model to explain substrate inhibition at high substrate concentrations, and inhibition is explained by a toxic substrate with an inhibition constant (KI). The Yano model with two positive constants explained the substrate inhibition with both toxic and non-toxic substrates.29 The other inhibition models did not satisfactorily explain the inhibition kinetics of the lactic acid from FWH. Optimization of industrial production of biochemicals with fermentation poses several challenges in bioprocess optimization. Substrate inhibition models address many such problems by aiding bioreactor design for high-density fermentation and fermentation at high substrate concentrations.49 They also help in deciding the substrate concentration for feeding strategies in fed-batch fermentation. In continuous culture systems, they help determine reactor sizing and dilution rate. Even large-scale fermentation is directly affected by substrate concentration both economically and technically.48 In the Luong model, the critical substrate concentration (Sm) also has practical importance, helping to understand the threshold concentration beyond which microbial growth ceases completely.50 The model's compatibility was calculated using the Akaike information criterion (AIC) as defined in eqn (7). The Haldane and Luong models were chosen for AIC analysis because they achieved the Highest R2. The AIC values for the two models are tabulated in Table 2. The ΔAIC values were analysed using 3- and 4-parameter models; positive ΔAIC values indicate that the simpler Haldane model can comfortably explain the substrate inhibition kinetics of glucose (Fig. S1).