Open Access Article
Open Access ArticleDesign, synthesis, in vitro, and in silico studies on promising α-glucosidase inhibitors based on a quinazolinone–thiophene scaffold
Parsa Baghershahia,
Navid Dastyaftehb,
Mohammad Halimic,
Maryam Mohammadi-Khanaposhtani d,
Seyedeh Niloufar Ghafourie,
Milad Nooria,
Fahimeh Ghasemif,
Somayeh Mojtabavi
d,
Seyedeh Niloufar Ghafourie,
Milad Nooria,
Fahimeh Ghasemif,
Somayeh Mojtabavi g,
Mohammad Ali Faramarzi
g,
Mohammad Ali Faramarzi g,
Mohammadreza Torabif,
Majid Alikhanih,
Bagher Larijanii,
Shahrzad Javanshir
g,
Mohammadreza Torabif,
Majid Alikhanih,
Bagher Larijanii,
Shahrzad Javanshir *a and
Mohammad Mahdavi*i
*a and
Mohammad Mahdavi*i
aPharmaceutical and Heterocyclic Compounds Research Laboratory, Department of Chemistry, Iran University of Science and Technology, Tehran 16846-13114, Iran. E-mail: shjavan@iust.ac.ir
bCenter for Research of Endemic Parasites of Iran, Tehran University of Medical Sciences, Tehran, Iran
cDepartment of Biology, Babol Branch, Islamic Azad University, Babol, Iran
dCellular and Molecular Biology Research Center, Health Research Institute, Babol University of Medical Sciences, Babol, Iran
eSchool of Chemistry, College of Science, University of Tehran, Tehran, Iran
fDepartment of Medical Biotechnology, School of Advanced Technologies in Medicine, Tehran University of Medical Sciences, Tehran, Iran
gDepartment of Pharmaceutical Biotechnology, Faculty of Pharmacy and Biotechnology Research Center, Tehran University of Medical Sciences, Tehran, Iran
hDepartment of Internal Medicine, School of Medicine, Rheumatology Research Center Shariati Hospital, Tehran University of Medical Sciences, Iran
iEndocrinology and Metabolism Research Center, Endocrinology and Metabolism Clinical Sciences Institute, Tehran University of Medical Sciences, Tehran, Iran. E-mail: momahdavi@tums.ac.ir
First published on 29th January 2026
Abstract
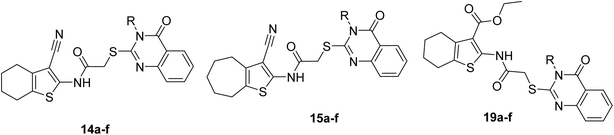
To discover novel inhibitors for α-glucosidase, a new scaffold featuring a quinazolinone–thiophene skeleton was designed through molecular hybridization. Eighteen derivatives, 14a–f, 15a–f, and 19a–f, were synthesized from this scaffold. These compounds were tested against yeast α-glucosidase. The in vitro enzymatic assays showed that all but one of the new compounds were active against α-glucosidase. The most potent compound was 19d, which inhibited the activity 11.7 times more effectively than acarbose, a standard inhibitor. In vitro kinetic studies demonstrated that this compound is a competitive inhibitor. Furthermore, in silico docking studies showed that compound 19d interacts with key residues in the active sites of both homology-modeled yeast and human α-glucosidase with favorable binding energies. Additionally, molecular dynamics simulations indicated that 19d formed a stable complex with this enzyme. Density functional theory (DFT) calculations provided further insight into the potency of the lead compounds. Based on the in vitro results, the most potent compounds were further examined in silico for druglikeness, pharmacokinetics, and toxicity. These evaluations revealed that, in terms of druglikeness and pharmacokinetics, the new compounds were similar to acarbose, while they showed better toxicity profiles. Furthermore, the most potent compounds exhibited low cytotoxicity against normal NIH-3T3 cells.
Introduction
Today, diabetes mellitus is one of the most serious medical problems in all societies.1 In the main types of this disease (type 1 and type 2), unusually high blood sugar levels or hyperglycemia are typical symptoms.2 Hyperglycemia is a dangerous metabolic condition that causes irreparable complications in the body.3 The main type of diabetes mellitus is type 2, which affects 90–95% of diabetic patients.4 In this type, there is resistance to insulin action, and oral hypoglycemic medications are the main drugs to control this type of diabetes.5 One of the most important categories of these oral drugs is α-glucosidase inhibitors such as acarbose, miglitol, and voglibose.α-Glucosidase is a carbohydrate hydrolase that is found in the small intestine (brush border). This enzyme, in the process of carbohydrate digestion, hydrolyzes 1,4-glycosidic linkages of oligosaccharides and disaccharides to monosaccharides.6 These process leads to postprandial hyperglycemia (PPH) and therefore, α-glucosidase inhibitors are useful in the control of PPH.7 Although α-glucosidase inhibitors retards glucose absorption, undesirable gastrointestinal side effects such as diarrhea, bloating, and flatulence impede their application.8,9 According to the mentioned facts, the development of strong α-glucosidase inhibitors with low side effects is an attractive goal for medicinal chemists.10–12
One approach in medicinal chemistry for developing new drugs is molecular hybridization theory.13 In this method, active pharmacophores are chosen from effective bioactive compounds and connected through conventional chemical reactions. Recently, the application of this theory in designing α-glucosidase inhibitors has gained significant popularity.14–16
Quinazolinone is a bicyclic heterocycle containing two nitrogen atoms and an oxygen atom. This heterocycle is found in several derivatives with high inhibitory activity against α-glucosidase.17–19 Compounds A are simple derivatives of quinazolinone that showed significant anti-α-glucosidase activity (Fig. 1).20 On the other hand, quinazolinone derivatives B and C, with more complex structures compared to the compounds A, also showed high inhibitory activity against α-glucosidase (Fig. 1).21,22 As can be seen, a thio group in the 2-position of the quinazolinone ring was observed.22 On the other hand, thiophene derivatives D–F demonstrated significant anti-α-glucosidase activity (Fig. 1).23–25 In this study, derivatives C, E, and F were selected as templates for further development and discussion of structure–activity relationships.
In our continued interest in the development of active inhibitors against α-glucosidase by molecular hybridization, we linked quinazolinone and thiophene heterocycle together by thioacetamide as a linker and designed a new series of quinazolinone–thiophene hybrids. It should be noted that the thioacetamide moiety is found in several series of potent α-glucosidase inhibitors.26 These derivatives were evaluated against α-glucosidase by in vitro enzymatic inhibition assay, in vitro kinetic study, in silico molecular docking, and in silico molecular dynamics. We also predicted druglikeness, pharmacokinetics, and toxicity of the target new compounds using an online server.
Materials and methods
General procedure for synthesis of 2-aminobenzamide derivatives 3a–f
A mixture of isatoic anhydride (1, 10 mmol) and amine derivatives 2a–f (10 mmol) in acetic acid (10 mL) and ethanol (50 mL) was stirred at reflux conditions for 8 h. After that, cold water was added to the mixture, and the pure 2-aminobenzamide derivatives 3a–f were separated by filtration.General procedure for the synthesis of quinazolinone derivatives 5a–f
2-Aminobenzamide derivatives 3a–f (10 mmol) and CS2 (4, 30 mmol) in KOH (10 mmol) and ethanol (50 mL) were stirred at reflux conditions for 24 h. The obtained products were quinazolinone derivatives 5a–f that were precipitated by the addition of cold water and were collected by filtration.Synthesis of 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carbonitrile (9) and 2-amino-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophene-3-carbonitrile (10)
In this step, cyclohexanone (6, 10 mmol) or cycloheptanone (7, 10 mmol) reacted with malononitrile (8, 10 mmol), S8 (20 mmol), and NEt3 (10.1 mmol) in ethanol (50 mL) at 100 °C for 3 h. To obtain the mixture, water was added, and 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carbonitrile (9) or 2-amino-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophene-3-carbonitrile (10) precipitates were collected by filtration.Synthesis of 2-chloro-N-(3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)acetamide (12) and 2-chloro-N-(3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)acetamide (13)
A mixture of compound 9 (10 mmol) or 10 (10 mmol) and chloroacetyl chloride (11) in 1,4-dioxane was stirred at RT for 2 h. After that, 2-chloro-N-(3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)acetamide (12) and 2-chloro-N-(3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)acetamide (13) were formed, and by adding cold water, were precipitated. Then, these compounds were separated by filtration.General procedure for synthesis of 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophens 14a–f and 3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophens 15a–f
In this step, a mixture of quinazolinone derivatives 5a–f (1 mmol) and compound 12 or 13 (1 mmol) in DMF (10 mL) was stirred at 50 °C for 8 h. After this time, cold water was added to the reaction mixture, and the precipitated 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophens 14a–f and 3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophens 15a–f were collected by filtration.Synthesis of ethyl 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (17)
A mixture of cyclohexanone (6, 10 mmol), ethyl 2-cyanoacetate (16, 10 mmol), S8 (20 mmol), and NEt3 (10.1 mmol) in ethanol (50 mL) was heated at 100 °C for 3 h. After that, water was added to the obtained mixture, and the precipitated ethyl 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (17) was collected by filtration.Synthesis of ethyl 2-(2-chloroacetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (18)
A mixture of ethyl 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (17, 10 mmol) and compound 11 (10 mmol) in 1,4-dioxane was stirred at RT for 2 h. Then, ethyl 2-(2-chloroacetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (18) was precipitated by adding cold water and was separated by filtration.General procedure for synthesis of 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylates 19a–f
In the final step, a mixture of compound 18 (1 mmol) and quinazolinone derivatives 5a–f (1 mmol) in DMF (10 mL) was stirred at 50 °C for 8 h. After that, cold water was added to the reaction mixture, and the obtained products 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylates 19a–f were separated by filtration.N-(3-Cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)-2-((3-ethyl-4-oxo-3,4-dihydroquinazolin-2-yl) thio) acetamide (14a)
Cream solid; yield: 75%; MP = 145–147 °C; IR (KBr, vmax) 3242 (NH), 3035 (CH aromatic), 2980 (CH aliphatic), 1660 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 12.06 (s, 1H), 8.06 (d, J = 7.0 Hz, 1H), 7.76 (t, J = 7.6 Hz, 1H), 7.45–7.41 (m, 2H), 4.38 (s, 2H), 4.12 (q, J = 7.0 Hz, 2H), 2.59–2.53 (m, 2H), 1.80–1.70 (m, 4H), 1.31 (t, J = 7.0 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.55, 160.65, 156.13, 147.10, 147.02, 135.18, 131.32, 127.88, 126.88, 126.51, 126.11, 119.26, 114.75, 93.20, 35.75, 23.99, 23.78, 23.02, 22.15, 13.49; anal. calcd: C21H20N4O2S2; C, 59.41; H, 4.75; N, 13.20; found; C, 59.59; H, 4.90; N, 13.38.
O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 12.06 (s, 1H), 8.06 (d, J = 7.0 Hz, 1H), 7.76 (t, J = 7.6 Hz, 1H), 7.45–7.41 (m, 2H), 4.38 (s, 2H), 4.12 (q, J = 7.0 Hz, 2H), 2.59–2.53 (m, 2H), 1.80–1.70 (m, 4H), 1.31 (t, J = 7.0 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.55, 160.65, 156.13, 147.10, 147.02, 135.18, 131.32, 127.88, 126.88, 126.51, 126.11, 119.26, 114.75, 93.20, 35.75, 23.99, 23.78, 23.02, 22.15, 13.49; anal. calcd: C21H20N4O2S2; C, 59.41; H, 4.75; N, 13.20; found; C, 59.59; H, 4.90; N, 13.38.
N-(3-Cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)-2-((4-oxo-3-propyl-3,4-dihydroquinazolin-2-yl)thio)acetamide (14b)
Cream solid; yield: 78%; MP = 150–152 °C; IR (KBr, vmax) 3354 (NH), 3030 (CH aromatic), 2885 (CH aliphatic), 1661 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 12.05 (s, 1H), 8.06 (dd, J = 8.2, 1.6 Hz, 1H), 7.79–7.73 (m, 1H), 7.47–7.39 (m, 2H), 4.38 (s, 2H), 4.07–3.99 (m, 2H), 2.55 (d, J = 4.5 Hz, 2H), 1.75 (d, J = 6.9 Hz, 6H), 0.96 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.52, 160.85, 156.33, 147.06, 146.98, 135.19, 131.32, 127.88, 126.94, 126.52, 126.11, 119.21, 114.74, 93.21, 46.18, 35.80, 23.98, 23.77, 23.02, 22.14, 21.46, 11.59; anal. calcd: C22H22N4O2S2; C, 60.25; H, 5.06; N, 12.78; found; C, 60.33; H, 5.21; N, 12.95.
O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 12.05 (s, 1H), 8.06 (dd, J = 8.2, 1.6 Hz, 1H), 7.79–7.73 (m, 1H), 7.47–7.39 (m, 2H), 4.38 (s, 2H), 4.07–3.99 (m, 2H), 2.55 (d, J = 4.5 Hz, 2H), 1.75 (d, J = 6.9 Hz, 6H), 0.96 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.52, 160.85, 156.33, 147.06, 146.98, 135.19, 131.32, 127.88, 126.94, 126.52, 126.11, 119.21, 114.74, 93.21, 46.18, 35.80, 23.98, 23.77, 23.02, 22.14, 21.46, 11.59; anal. calcd: C22H22N4O2S2; C, 60.25; H, 5.06; N, 12.78; found; C, 60.33; H, 5.21; N, 12.95.
2-((3-Butyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)-N-(3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)acetamide (14c)
Cream solid; yield: 69%; MP = 153–155 °C; IR (KBr, vmax) 3358 (NH), 3020 (CH aromatic), 2865 (CH aliphatic), 1663 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1.1H NMR (400 MHz, DMSO-d6) δ 12.05 (s, 1H), 8.06 (dd, J = 7.8, 1.6 Hz, 1H), 7.79–7.74 (m, 1H), 7.49–7.39 (m, 2H), 4.38 (s, 2H), 4.07 (t, J = 7.8 Hz, 2H), 2.60–2.54 (m, 2H), 1.78–1.72 (m, 4H), 1.73–1.65 (m, 2H), 1.47–1.33 (m, 2H), 0.96 (t, J = 7.3 Hz, 3H);13C NMR (101 MHz, DMSO-d6) δ 166.24, 160.58, 156.19, 147.02, 146.94, 134.85, 131.07, 127.63, 126.82, 126.40, 126.00, 119.08, 114.62, 92.78, 45.73, 35.51, 23.88, 23.62, 22.93, 22.06, 21.33, 17.41, 11.50; anal. calcd: C23H24N4O2S2; C, 61.04; H, 5.35; N, 12.38; found; C, 61.21; H, 5.49; N, 12.52.
O) cm−1.1H NMR (400 MHz, DMSO-d6) δ 12.05 (s, 1H), 8.06 (dd, J = 7.8, 1.6 Hz, 1H), 7.79–7.74 (m, 1H), 7.49–7.39 (m, 2H), 4.38 (s, 2H), 4.07 (t, J = 7.8 Hz, 2H), 2.60–2.54 (m, 2H), 1.78–1.72 (m, 4H), 1.73–1.65 (m, 2H), 1.47–1.33 (m, 2H), 0.96 (t, J = 7.3 Hz, 3H);13C NMR (101 MHz, DMSO-d6) δ 166.24, 160.58, 156.19, 147.02, 146.94, 134.85, 131.07, 127.63, 126.82, 126.40, 126.00, 119.08, 114.62, 92.78, 45.73, 35.51, 23.88, 23.62, 22.93, 22.06, 21.33, 17.41, 11.50; anal. calcd: C23H24N4O2S2; C, 61.04; H, 5.35; N, 12.38; found; C, 61.21; H, 5.49; N, 12.52.
N-(3-Cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)-2-((4-oxo-3-phenyl-3,4-dihydroquinazolin-2-yl)thio)acetamide (14d)
Brown solid; yield: 88%; MP = 145–147 °C; IR (KBr, vmax) 3375 (NH), 3040 (CH aromatic), 2985 (CH aliphatic), 1664 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 11.98 (s, 1H), 8.08 (dd, J = 8.0, 1.5 Hz, 1H), 7.87–7.78 (m, 1H), 7.61 (d, J = 2.0 Hz, 1H), 7.60 (d, J = 1.8 Hz, 2H), 7.54–7.44 (m, 3H), 4.23 (s, 2H), 2.58–2.54 (m, 2H), 1.74 (t, J = 3.7 Hz, 4H);13C NMR (101 MHz, DMSO) δ 166.52, 161.06, 157.15, 147.52, 146.94, 136.21, 135.44, 131.30, 130.54, 130.06, 129.90, 127.86, 127.09, 126.59, 126.33, 120.01, 114.74, 93.17, 36.20, 23.98, 23.78, 23.04, 22.15; anal. calcd: C25H20N4O2S2; C, 63.54; H, 4.27; N, 11.86; found; C, 63.68; H, 4.48; N, 12.02.
O) cm−1; 1H NMR (400 MHz, DMSO-d6) δ 11.98 (s, 1H), 8.08 (dd, J = 8.0, 1.5 Hz, 1H), 7.87–7.78 (m, 1H), 7.61 (d, J = 2.0 Hz, 1H), 7.60 (d, J = 1.8 Hz, 2H), 7.54–7.44 (m, 3H), 4.23 (s, 2H), 2.58–2.54 (m, 2H), 1.74 (t, J = 3.7 Hz, 4H);13C NMR (101 MHz, DMSO) δ 166.52, 161.06, 157.15, 147.52, 146.94, 136.21, 135.44, 131.30, 130.54, 130.06, 129.90, 127.86, 127.09, 126.59, 126.33, 120.01, 114.74, 93.17, 36.20, 23.98, 23.78, 23.04, 22.15; anal. calcd: C25H20N4O2S2; C, 63.54; H, 4.27; N, 11.86; found; C, 63.68; H, 4.48; N, 12.02.
2-((3-Benzyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)-N-(3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)acetamide (14e)
Cream solid; yield: 71%; MP = 149–151 °C; IR (KBr, vmax) 3258 (NH), 3020 (CH aromatic), 2965 (CH aliphatic), 1661 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 12.04 (s, 1H), 8.11 (d, J = 7.1 Hz, 1H), 7.83–7.78 (m, 1H), 7.50–7.45 (m, 2H), 7.38–7.30 (m, 5H), 5.35 (s, 2H), 4.34 (s, 2H), 2.58–2.53 (m, 2H), 1.89–1.52 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.43, 161.27, 156.79, 147.13, 146.98, 136.01, 135.49, 131.33, 129.12, 128.01, 127.90, 127.36, 127.14, 126.73, 126.23, 119.16, 114.75, 93.22, 47.55, 36.01, 23.98, 23.78, 23.02, 22.15; anal. calcd: C26H22N4O2S2; C, 64.18; H, 4.56; N, 11.51; found; C, 64.34; H, 4.73; N, 11.74.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 12.04 (s, 1H), 8.11 (d, J = 7.1 Hz, 1H), 7.83–7.78 (m, 1H), 7.50–7.45 (m, 2H), 7.38–7.30 (m, 5H), 5.35 (s, 2H), 4.34 (s, 2H), 2.58–2.53 (m, 2H), 1.89–1.52 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.43, 161.27, 156.79, 147.13, 146.98, 136.01, 135.49, 131.33, 129.12, 128.01, 127.90, 127.36, 127.14, 126.73, 126.23, 119.16, 114.75, 93.22, 47.55, 36.01, 23.98, 23.78, 23.02, 22.15; anal. calcd: C26H22N4O2S2; C, 64.18; H, 4.56; N, 11.51; found; C, 64.34; H, 4.73; N, 11.74.
Ethyl 2-(2-((4-oxo-3-phenethyl-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (14f)
Browne solid; yield: 68%; MP = 154–156 °C; IR (KBr, vmax) 3231 (NH), 3010 (CH aromatic), 2955 (CH aliphatic), 1664 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 12.06 (s, 1H), 8.07 (d, J = 6.6 Hz, 1H), 7.80–7.74 (m, 1H), 7.47–7.42 (m, 2H), 7.37–7.26 (m, 5H), 4.40 (s, 2H), 4.25 (t, J = 8.0 Hz, 2H), 3.02 (t, J = 8.0 Hz, 2H), 2.57–2.53 (m, 2H), 1.76–1.72 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.48, 160.75, 156.15, 147.05, 146.99, 138.15, 135.27, 131.32, 129.15, 129.12, 127.90, 127.22, 126.93, 126.60, 126.16, 119.23, 114.74, 93.22, 46.12, 35.84, 33.89, 23.99, 23.78, 23.03, 22.15; anal. calcd: C27H24N4O2S2 C, 64.78; H, 4.83; N, 11.19; found; C, 64.97; H, 5.06; N, 11.32.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 12.06 (s, 1H), 8.07 (d, J = 6.6 Hz, 1H), 7.80–7.74 (m, 1H), 7.47–7.42 (m, 2H), 7.37–7.26 (m, 5H), 4.40 (s, 2H), 4.25 (t, J = 8.0 Hz, 2H), 3.02 (t, J = 8.0 Hz, 2H), 2.57–2.53 (m, 2H), 1.76–1.72 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.48, 160.75, 156.15, 147.05, 146.99, 138.15, 135.27, 131.32, 129.15, 129.12, 127.90, 127.22, 126.93, 126.60, 126.16, 119.23, 114.74, 93.22, 46.12, 35.84, 33.89, 23.99, 23.78, 23.03, 22.15; anal. calcd: C27H24N4O2S2 C, 64.78; H, 4.83; N, 11.19; found; C, 64.97; H, 5.06; N, 11.32.
N-(3-Cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)-2-((3-ethyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)acetamide (15a)
Cream solid; yield: 78%; MP = 160–162 °C; IR (KBr, vmax) 3264 (NH), 3015 (CH aromatic), 2950 (CH aliphatic), 1662 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1;1H NMR (400 MHz, DMSO) δ 11.96 (s, 1H), 8.06 (dd, J = 8.2, 1.2 Hz, 2H), 7.80–7.72 (m, 1H), 7.48–7.39 (m, 2H), 4.38 (s, 2H), 4.12 (q, J = 7.0 Hz, 2H), 2.71–2.60 (m, 4H), 1.83–1.75 (m, 2H), 1.65–1.52 (m, 4H), 1.31 (t, J = 7.0 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.56, 160.65, 156.11, 147.10, 144.99, 136.07, 135.15, 131.53, 126.88, 126.51, 126.14, 119.27, 115.23, 95.75, 35.65, 31.82, 28.81, 28.71, 28.05, 27.36, 13.49; anal. calcd: C22H22N4O2S2; C, 60.25; H, 5.06; N, 12.78; found; C, 60.45; H, 5.27; N, 12.95.
O) cm−1;1H NMR (400 MHz, DMSO) δ 11.96 (s, 1H), 8.06 (dd, J = 8.2, 1.2 Hz, 2H), 7.80–7.72 (m, 1H), 7.48–7.39 (m, 2H), 4.38 (s, 2H), 4.12 (q, J = 7.0 Hz, 2H), 2.71–2.60 (m, 4H), 1.83–1.75 (m, 2H), 1.65–1.52 (m, 4H), 1.31 (t, J = 7.0 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.56, 160.65, 156.11, 147.10, 144.99, 136.07, 135.15, 131.53, 126.88, 126.51, 126.14, 119.27, 115.23, 95.75, 35.65, 31.82, 28.81, 28.71, 28.05, 27.36, 13.49; anal. calcd: C22H22N4O2S2; C, 60.25; H, 5.06; N, 12.78; found; C, 60.45; H, 5.27; N, 12.95.
N-(3-Cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)-2-((4-oxo-3-propyl-3,4-dihydroquinazolin-2-yl)thio)acetamide (15b)
Cream solid; yield: 65%; MP = 164–166 °C; IR (KBr, vmax) 3252 (NH), 3020 (CH aromatic), 2960 (CH aliphatic), 1666 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.95 (s, 1H), 8.06 (d, J = 7.0 Hz, 1H), 7.80–7.70 (m, 1H), 7.47–7.39 (m, 2H), 4.38 (s, 2H), 4.03 (t, J = 6.0 Hz, 2H), 2.75–2.58 (m, 4H), 1.82–1.76 (m, 2H), 1.76–1.69 (m, 2H), 1.63–1.51 (m, 4H), 0.96 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.53, 160.85, 156.31, 147.06, 144.98, 136.06, 135.15, 131.51, 126.93, 126.51, 126.14, 119.22, 115.22, 95.75, 46.16, 35.71, 31.82, 28.80, 28.71, 28.05, 27.36, 21.46, 11.59; anal. calcd: C23H24N4O2S2; C, 61.04; H, 5.35; N, 12.38; found; C, 61.26; H, 5.59; N, 12.52.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.95 (s, 1H), 8.06 (d, J = 7.0 Hz, 1H), 7.80–7.70 (m, 1H), 7.47–7.39 (m, 2H), 4.38 (s, 2H), 4.03 (t, J = 6.0 Hz, 2H), 2.75–2.58 (m, 4H), 1.82–1.76 (m, 2H), 1.76–1.69 (m, 2H), 1.63–1.51 (m, 4H), 0.96 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.53, 160.85, 156.31, 147.06, 144.98, 136.06, 135.15, 131.51, 126.93, 126.51, 126.14, 119.22, 115.22, 95.75, 46.16, 35.71, 31.82, 28.80, 28.71, 28.05, 27.36, 21.46, 11.59; anal. calcd: C23H24N4O2S2; C, 61.04; H, 5.35; N, 12.38; found; C, 61.26; H, 5.59; N, 12.52.
2-((3-Butyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)-N-(3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)acetamide (15c)
Cream solid; yield: 69%; MP = 168–170 °C; IR (KBr, vmax) 3275 (NH), 3035 (CH aromatic), 2945 (CH aliphatic), 1669 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.96 (s, 1H), 8.06 (dd, J = 8.2, 1.5 Hz, 1H), 7.81–7.72 (m, 1H), 7.49–7.39 (m, 2H), 4.37 (s, 2H), 4.06 (t, J = 8.0 Hz, 2H), 2.74–2.60 (m, 4H), 1.84–1.76 (m, 2H), 1.74–1.67 (m, 2H), 1.63–1.53 (m, 4H), 1.40 (h, J = 7.4 Hz, 2H), 0.95 (t, J = 7.3 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.55, 160.83, 156.30, 147.06, 145.00, 136.07, 135.16, 131.52, 126.92, 126.53, 126.14, 119.21, 115.23, 95.75, 44.48, 35.71, 31.82, 30.07, 28.81, 28.71, 28.06, 27.36, 20.11, 14.04; anal. calcd: C24H26N4O0S2; C, 61.78; H, 5.62; N, 12.01; found; C, 61.96; H, 5.80; N, 12.23.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.96 (s, 1H), 8.06 (dd, J = 8.2, 1.5 Hz, 1H), 7.81–7.72 (m, 1H), 7.49–7.39 (m, 2H), 4.37 (s, 2H), 4.06 (t, J = 8.0 Hz, 2H), 2.74–2.60 (m, 4H), 1.84–1.76 (m, 2H), 1.74–1.67 (m, 2H), 1.63–1.53 (m, 4H), 1.40 (h, J = 7.4 Hz, 2H), 0.95 (t, J = 7.3 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 166.55, 160.83, 156.30, 147.06, 145.00, 136.07, 135.16, 131.52, 126.92, 126.53, 126.14, 119.21, 115.23, 95.75, 44.48, 35.71, 31.82, 30.07, 28.81, 28.71, 28.06, 27.36, 20.11, 14.04; anal. calcd: C24H26N4O0S2; C, 61.78; H, 5.62; N, 12.01; found; C, 61.96; H, 5.80; N, 12.23.
N-(3-Cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)-2-((4-oxo-3-phenyl-3,4-dihydroquinazolin-2-yl)thio)acetamide (15d)
Cream solid; yield: 71%; MP = 168–170 °C; IR (KBr, vmax) 3295 (NH), 3030 (CH aromatic), 2975 (CH aliphatic), 1671 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.89 (s, 1H), 8.08 (d, J = 7.7 Hz, 1H), 7.86–7.77 (m, 1H), 7.63–7.57 (m, 2H), 7.58–7.43 (m, 5H), 4.23 (s, 2H), 2.70–2.62 (m, 4H), 1.82–1.76 (m, 2H), 1.60–1.54 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.52, 161.07, 157.12, 147.53, 144.93, 136.21, 136.04, 135.41, 131.50, 130.54, 130.06, 129.90, 127.09, 126.59, 126.36, 120.01, 115.23, 95.71, 36.11, 31.83, 28.80, 28.71, 28.07, 27.36; anal. calcd: C26H22N4O2S2 C, 64.18; H, 4.56; N, 11.51; found; C, 64.35; H, 4.72; N, 11.69.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.89 (s, 1H), 8.08 (d, J = 7.7 Hz, 1H), 7.86–7.77 (m, 1H), 7.63–7.57 (m, 2H), 7.58–7.43 (m, 5H), 4.23 (s, 2H), 2.70–2.62 (m, 4H), 1.82–1.76 (m, 2H), 1.60–1.54 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.52, 161.07, 157.12, 147.53, 144.93, 136.21, 136.04, 135.41, 131.50, 130.54, 130.06, 129.90, 127.09, 126.59, 126.36, 120.01, 115.23, 95.71, 36.11, 31.83, 28.80, 28.71, 28.07, 27.36; anal. calcd: C26H22N4O2S2 C, 64.18; H, 4.56; N, 11.51; found; C, 64.35; H, 4.72; N, 11.69.
2-((3-Benzyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)-N-(3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)acetamide (15e)
Cream solid; yield: 61%; MP = 172–174 °C; IR (KBr, vmax) 3290 (NH), 3050 (CH aromatic), 2980 (CH aliphatic), 1670 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.94 (s, 1H), 8.11 (d, J = 8.3 Hz, 1H), 7.85–7.76 (m, 1H), 7.52–7.45 (m, 2H), 7.40–7.27 (m, 5H), 5.35 (s, 2H), 4.34 (s, 2H), 2.72–2.61 (m, 4H), 1.83–1.77 (m, 2H), 1.64–1.53 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.44, 161.26, 156.78, 147.13, 144.95, 144.95, 136.08, 136.01, 135.46, 131.55, 129.12, 128.01, 127.35, 127.14, 126.74, 126.25, 119.17, 115.22, 95.76, 47.54, 35.92, 33.16, 31.82, 28.81, 28.05, 27.36; anal. calcd: C27H24N4O2S2; C, 64.78; H, 4.83; N, 11.19; found; C, 64.96; H, 4.98; N, 11.41.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.94 (s, 1H), 8.11 (d, J = 8.3 Hz, 1H), 7.85–7.76 (m, 1H), 7.52–7.45 (m, 2H), 7.40–7.27 (m, 5H), 5.35 (s, 2H), 4.34 (s, 2H), 2.72–2.61 (m, 4H), 1.83–1.77 (m, 2H), 1.64–1.53 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.44, 161.26, 156.78, 147.13, 144.95, 144.95, 136.08, 136.01, 135.46, 131.55, 129.12, 128.01, 127.35, 127.14, 126.74, 126.25, 119.17, 115.22, 95.76, 47.54, 35.92, 33.16, 31.82, 28.81, 28.05, 27.36; anal. calcd: C27H24N4O2S2; C, 64.78; H, 4.83; N, 11.19; found; C, 64.96; H, 4.98; N, 11.41.
N-(3-Cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)-2-((4-oxo-3-phenethyl-3,4-dihydroquinazolin-2-yl)thio)acetamide (15f)
Cream solid; yield: 78%; MP = 205–207 °C; IR (KBr, vmax) 3210 (NH), 3020 (CH aromatic), 2920 (CH aliphatic), 1666 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.97 (s, 1H), 8.07 (d, J = 7.1 Hz, 1H), 7.80–7.74 (m, 1H), 7.48–7.42 (m, 2H), 7.39–7.25 (m, 5H), 4.40 (s, 2H), 4.25 (t, J = 8.0 Hz, 2H), 3.02 (t, J = 8.0 Hz, 2H), 2.72–2.62 (m, 4H), 1.85–1.74 (m, 2H), 1.66–1.50 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.49, 160.75, 156.14, 147.05, 144.99, 138.15, 136.07, 135.24, 131.53, 129.15, 129.12, 127.22, 126.93, 126.60, 126.19, 119.25, 115.23, 95.77, 46.11, 35.76, 33.89, 31.82, 28.81, 28.71, 28.06, 27.36; HRMS: (M = 514.2886); anal. calcd: C28H29N4O2S2; C, 65.35; H, 5.09; N, 10.89; found; C, 65.51; H, 5.23; N, 10.97.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.97 (s, 1H), 8.07 (d, J = 7.1 Hz, 1H), 7.80–7.74 (m, 1H), 7.48–7.42 (m, 2H), 7.39–7.25 (m, 5H), 4.40 (s, 2H), 4.25 (t, J = 8.0 Hz, 2H), 3.02 (t, J = 8.0 Hz, 2H), 2.72–2.62 (m, 4H), 1.85–1.74 (m, 2H), 1.66–1.50 (m, 4H); 13C NMR (101 MHz, DMSO) δ 166.49, 160.75, 156.14, 147.05, 144.99, 138.15, 136.07, 135.24, 131.53, 129.15, 129.12, 127.22, 126.93, 126.60, 126.19, 119.25, 115.23, 95.77, 46.11, 35.76, 33.89, 31.82, 28.81, 28.71, 28.06, 27.36; HRMS: (M = 514.2886); anal. calcd: C28H29N4O2S2; C, 65.35; H, 5.09; N, 10.89; found; C, 65.51; H, 5.23; N, 10.97.
Ethyl 2-(2-((3-ethyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19a)
Cream solid; yield: 65%; MP = 135–137 °C; IR (KBr, vmax) 3215 (NH), 3020 (CH aromatic), 2960 (CH aliphatic), 1667 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.71 (s, 1H), 8.06 (dd, J = 8.1, 1.2 Hz, 3H) 7.79–7.71 (m, 1H), 7.48–7.40 (m, 2H), 4.31 (s, 2H), 4.21 (q, J = 7.1 Hz, 2H), 4.15 (q, J = 7.1 Hz, 2H), 2.69–2.63 (m, 2H), 2.61–2.54 (m, 2H), 1.74–1.61 (m, 4H), 1.35 (t, J = 7.1 Hz, 3H), 1.25 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.72, 165.53, 160.79, 155.46, 146.95, 146.41, 135.02, 130.64, 126.83, 126.75, 126.67, 126.28, 118.90, 111.28, 60.32, 45.14, 35.66, 26.09, 24.05, 22.77, 22.64, 14.34, 11.41; anal. calcd: C23H25N3O4S2; C, 58.58; H, 5.34; N, 8.91; found; C, 58.72; H, 5.56; N, 9.07.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.71 (s, 1H), 8.06 (dd, J = 8.1, 1.2 Hz, 3H) 7.79–7.71 (m, 1H), 7.48–7.40 (m, 2H), 4.31 (s, 2H), 4.21 (q, J = 7.1 Hz, 2H), 4.15 (q, J = 7.1 Hz, 2H), 2.69–2.63 (m, 2H), 2.61–2.54 (m, 2H), 1.74–1.61 (m, 4H), 1.35 (t, J = 7.1 Hz, 3H), 1.25 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.72, 165.53, 160.79, 155.46, 146.95, 146.41, 135.02, 130.64, 126.83, 126.75, 126.67, 126.28, 118.90, 111.28, 60.32, 45.14, 35.66, 26.09, 24.05, 22.77, 22.64, 14.34, 11.41; anal. calcd: C23H25N3O4S2; C, 58.58; H, 5.34; N, 8.91; found; C, 58.72; H, 5.56; N, 9.07.
Ethyl 2-(2-((4-oxo-3-propyl-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19b)
Cream solid; yield: 65%; MP = 142–144 °C; IR (KBr, vmax) 3230 (NH), 3030 (CH aromatic), 2930 (CH aliphatic), 1666 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1;1H NMR (400 MHz, DMSO) δ 11.70 (s, 1H), 8.06 (dd, J = 8.2, 1.5 Hz, 1H), 7.79–7.71 (m, 1H), 7.49–7.40 (m, 2H), 4.31 (s, 2H), 4.21 (q, J = 7.1 Hz, 2H), 4.06 (t, J = 8.0 Hz, 2H), 2.70–2.63 (m, 2H), 2.61–2.53 (m, 2H), 1.80 (h, J = 7.5 Hz, 2H), 1.73–1.63 (m, 4H), 1.25 (t, J = 7.1 Hz, 3H), 0.97 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.76, 165.65, 155.64, 146.99, 146.46, 135.17, 130.87, 126.88, 126.79, 126.71, 126.32, 119.96, 119.27, 111.93, 60.85, 46.19, 35.75, 26.24, 24.16, 22.88, 22.69, 21.40, 14.51, 11.61; anal. calcd: C24H27N3O4S2; C, 59.36; H, 5.60; N, 8.65; found C, 59.55; H, 5.73; N, 8.80.
O) cm−1;1H NMR (400 MHz, DMSO) δ 11.70 (s, 1H), 8.06 (dd, J = 8.2, 1.5 Hz, 1H), 7.79–7.71 (m, 1H), 7.49–7.40 (m, 2H), 4.31 (s, 2H), 4.21 (q, J = 7.1 Hz, 2H), 4.06 (t, J = 8.0 Hz, 2H), 2.70–2.63 (m, 2H), 2.61–2.53 (m, 2H), 1.80 (h, J = 7.5 Hz, 2H), 1.73–1.63 (m, 4H), 1.25 (t, J = 7.1 Hz, 3H), 0.97 (t, J = 7.4 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.76, 165.65, 155.64, 146.99, 146.46, 135.17, 130.87, 126.88, 126.79, 126.71, 126.32, 119.96, 119.27, 111.93, 60.85, 46.19, 35.75, 26.24, 24.16, 22.88, 22.69, 21.40, 14.51, 11.61; anal. calcd: C24H27N3O4S2; C, 59.36; H, 5.60; N, 8.65; found C, 59.55; H, 5.73; N, 8.80.
Ethyl 2-(2-((3-butyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19c)
Cream solid; yield: 65%; MP = 139–141 °C; IR (KBr, vmax) 3230 (NH), 3035 (CH aromatic), 2950 (CH aliphatic), 1669 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.71 (s, 1H), 8.06 (dd, J = 8.3, 1.6 Hz, 1H), 7.79–7.72 (m, 1H), 7.49–7.40 (m, 2H), 4.30 (s, 2H), 4.22 (q, J = 7.1 Hz, 2H), 4.10 (t, J = 7.8 Hz, 2H), 2.71–2.62 (m, 2H), 2.59–2.54 (m, 2H), 1.78–1.71 (m, 2H), 1.71–1.66 (m, 4H), 1.41 (h, J = 7.3 Hz, 2H), 1.25 (t, J = 7.1 Hz, 3H), 0.97 (t, J = 7.3 Hz, 3H); 13C NMR (101 MHz, DMSO-d6) δ 165.75, 165.67, 155.61, 147.36, 147.01, 135.17, 130.87, 126.87, 126.79, 126.73, 126.32, 120.76, 120.02, 111.92, 60.83, 44.54, 35.74, 30.02, 26.24, 26.14, 22.88, 22.69, 20.14, 14.53, 14.06.; anal. calcd: C25H29N3O4S2; C, 60.10; H, 5.85; N, 8.41; found; C, 60.57; H, 5.96; N, 8.58.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.71 (s, 1H), 8.06 (dd, J = 8.3, 1.6 Hz, 1H), 7.79–7.72 (m, 1H), 7.49–7.40 (m, 2H), 4.30 (s, 2H), 4.22 (q, J = 7.1 Hz, 2H), 4.10 (t, J = 7.8 Hz, 2H), 2.71–2.62 (m, 2H), 2.59–2.54 (m, 2H), 1.78–1.71 (m, 2H), 1.71–1.66 (m, 4H), 1.41 (h, J = 7.3 Hz, 2H), 1.25 (t, J = 7.1 Hz, 3H), 0.97 (t, J = 7.3 Hz, 3H); 13C NMR (101 MHz, DMSO-d6) δ 165.75, 165.67, 155.61, 147.36, 147.01, 135.17, 130.87, 126.87, 126.79, 126.73, 126.32, 120.76, 120.02, 111.92, 60.83, 44.54, 35.74, 30.02, 26.24, 26.14, 22.88, 22.69, 20.14, 14.53, 14.06.; anal. calcd: C25H29N3O4S2; C, 60.10; H, 5.85; N, 8.41; found; C, 60.57; H, 5.96; N, 8.58.
Ethyl 2-(2-((4-oxo-3-phenyl-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19d)
Cream solid; yield: 72%; MP = 147–149 °C; IR (KBr, vmax) 3290 (NH), 3045 (CH aromatic), 2970 (CH aliphatic), 1668 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.70 (s, 1H), 8.07 (dd, J = 7.9, 1.5 Hz, 1H), 7.85–7.78 (m, 1H), 7.65–7.60 (m, 2H), 7.59–7.53 (m, 4H), 7.48 (t, J = 7.1 Hz, 1H), 7.59–7.45 (m, 5H), 4.28 (q, J = 7.1 Hz, 2H), 4.14 (s, 2H), 2.73–2.63 (m, 2H), 2.63–2.54 (m, 2H), 1.73–1.66 (m, 4H), 1.27 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.74, 165.68, 161.09, 161.09, 156.37, 147.44, 146.47, 136.07, 130.88, 130.09, 130.02, 127.02, 126.79, 126.55, 120.08, 120.06, 112.01, 89.82, 60.93, 35.98, 26.24, 24.17, 22.89, 22.70, 14.53; HRMS: (M = 520.2258); anal. calcd: C29H29BrFN3O5S2; C, 62.41; H, 4.85; N, 8.09; found; C, 62.58; H, 5.02; N, 8.29.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.70 (s, 1H), 8.07 (dd, J = 7.9, 1.5 Hz, 1H), 7.85–7.78 (m, 1H), 7.65–7.60 (m, 2H), 7.59–7.53 (m, 4H), 7.48 (t, J = 7.1 Hz, 1H), 7.59–7.45 (m, 5H), 4.28 (q, J = 7.1 Hz, 2H), 4.14 (s, 2H), 2.73–2.63 (m, 2H), 2.63–2.54 (m, 2H), 1.73–1.66 (m, 4H), 1.27 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.74, 165.68, 161.09, 161.09, 156.37, 147.44, 146.47, 136.07, 130.88, 130.09, 130.02, 127.02, 126.79, 126.55, 120.08, 120.06, 112.01, 89.82, 60.93, 35.98, 26.24, 24.17, 22.89, 22.70, 14.53; HRMS: (M = 520.2258); anal. calcd: C29H29BrFN3O5S2; C, 62.41; H, 4.85; N, 8.09; found; C, 62.58; H, 5.02; N, 8.29.
Ethyl 2-(2-((3-benzyl-4-oxo-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19e)
Cream solid; yield: 69%; MP = 156–158 °C; IR (KBr, vmax) 3310 (NH), 3040 (CH aromatic), 2980 (CH aliphatic), 1672 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.61 (s, 1H), 8.10 (dd, J = 8.0, 1.5 Hz, 1H), 7.84–7.73 (m, 1H), 7.55–7.44 (m, 2H), 7.40–7.23 (m, 5H), 5.38 (s, 2H), 4.28 (s, 2H), 4.22 (q, J = 7.1 Hz, 2H), 2.72–2.62 (m, 2H), 2.63–2.52 (m, 2H), 1.75–1.61 (m, 4H), 1.25 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.72, 165.55, 161.27, 156.26, 147.09, 146.38, 135.90, 135.41, 130.90, 129.11, 127.99, 127.29, 127.05, 126.88, 126.82, 126.43, 119.23, 111.99, 60.85, 47.62, 36.05, 26.25, 24.15, 22.88, 22.70, 14.54; anal. calcd: C28H27N3O4S2; C, 63.02; H, 5.10; N, 7.87; found: C, 63.21; H, 5.28; N, 8.02.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.61 (s, 1H), 8.10 (dd, J = 8.0, 1.5 Hz, 1H), 7.84–7.73 (m, 1H), 7.55–7.44 (m, 2H), 7.40–7.23 (m, 5H), 5.38 (s, 2H), 4.28 (s, 2H), 4.22 (q, J = 7.1 Hz, 2H), 2.72–2.62 (m, 2H), 2.63–2.52 (m, 2H), 1.75–1.61 (m, 4H), 1.25 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.72, 165.55, 161.27, 156.26, 147.09, 146.38, 135.90, 135.41, 130.90, 129.11, 127.99, 127.29, 127.05, 126.88, 126.82, 126.43, 119.23, 111.99, 60.85, 47.62, 36.05, 26.25, 24.15, 22.88, 22.70, 14.54; anal. calcd: C28H27N3O4S2; C, 63.02; H, 5.10; N, 7.87; found: C, 63.21; H, 5.28; N, 8.02.
Ethyl 2-(2-((4-oxo-3-phenethyl-3,4-dihydroquinazolin-2-yl)thio)acetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (19f)
Cream solid; yield: 78%; MP = 205–207 °C; IR (KBr, vmax) 3260 (NH), 3040 (CH aromatic), 2980 (CH aliphatic), 1669 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.78 (s, 1H), 8.07 (d, J = 6.9 Hz, 1H), 7.81–7.72 (m, 1H), 7.50–7.42 (m, 2H), 7.40–7.19 (m, 5H), 4.33 (s, 2H), 4.33–4.24 (m, 2H), 4.18 (q, J = 7.1 Hz, 2H), 3.17–3.01 (m, 2H), 2.61–2.55 (m, 2H), 2.58 (s, 2H), 1.69 (d, J = 5.0 Hz, 4H), 1.21 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.74, 165.68, 160.76, 155.39, 154.55, 146.98, 146.52, 142.69, 138.23, 135.25, 130.87, 129.14, 129.12, 127.22, 126.80, 126.40, 119.30, 111.96, 60.85, 46.14, 35.75, 35.11, 33.80, 26.24, 24.16, 22.69, 14.49; anal. calcd: C29H29N3O4S2; C, 63.60; H, 5.34; N, 7.67; found; C, 63.75; H, 5.49; N, 7.88.
O) cm−1; 1H NMR (400 MHz, DMSO) δ 11.78 (s, 1H), 8.07 (d, J = 6.9 Hz, 1H), 7.81–7.72 (m, 1H), 7.50–7.42 (m, 2H), 7.40–7.19 (m, 5H), 4.33 (s, 2H), 4.33–4.24 (m, 2H), 4.18 (q, J = 7.1 Hz, 2H), 3.17–3.01 (m, 2H), 2.61–2.55 (m, 2H), 2.58 (s, 2H), 1.69 (d, J = 5.0 Hz, 4H), 1.21 (t, J = 7.1 Hz, 3H); 13C NMR (101 MHz, DMSO) δ 165.74, 165.68, 160.76, 155.39, 154.55, 146.98, 146.52, 142.69, 138.23, 135.25, 130.87, 129.14, 129.12, 127.22, 126.80, 126.40, 119.30, 111.96, 60.85, 46.14, 35.75, 35.11, 33.80, 26.24, 24.16, 22.69, 14.49; anal. calcd: C29H29N3O4S2; C, 63.60; H, 5.34; N, 7.67; found; C, 63.75; H, 5.49; N, 7.88.
In vitro α-glucosidase inhibition and kinetics
The in vitro anti-α-glucosidase activity and kinetic study on quinazolinone–thiophene derivatives 14a–f, 15a–f, and 19a–f were evaluated according to the previously reported methods on Saccharomyces cerevisiae α-glucosidase.26Molecular docking
For the docking study, we prepared a homology model of α-glucosidase using the described method by Imran et al. because in the in vitro study, we used Saccharomyces cerevisiae α-glucosidase, which does not have a crystallographic structure in the Protein Data Bank.27,28 Firstly, an appropriate protein with high similarity to Saccharomyces cerevisiae α-glucosidase was found by SWISS-MODEL. This protein was Saccharomyces cerevisiae isomaltase with PDB ID of 3 A4A (https://www.rcsb.org/structure/3A4A). After that, this protein was subjected to sequence alignment and homology model using the automated homology modeling pipeline SWISS-MODEL (managed by the Swiss Institute of Bioinformatics), and the quality of the obtained homology model was verified using PROCHECK.29 According to these steps, a pdb file for S. cerevisiae α-glucosidase was constructed and converted to pdbq coordinate using AutodockTools (version 1.5.6). By the latter software, the 3D structures of the acarbose and selected compounds (that were obtained by MarvineSketch 5.8.3, 2012, ChemAxon, http://www.chemaxon.com) were converted to pdbqt coordinates. pdbqt coordinates of the selected ligands (acarbose and the most potent compounds) were used as input files for the AUTOGRID program. In this program, for each atom type in the ligand, maps were calculated with 0.375 Å spacing between the following grid points: the center was placed at x = 12.5825, y = 7.8955, and z = 12.519, and the dimensions of the active site box were set at 40 × 40 × 40 Å. Flexible ligand dockings were applied for the selected ligands. Each docked system was carried out by 75 runs of the AUTODOCK search by the Lamarckian genetic algorithm. The best poses in terms of binding energy for each compound were selected for further analysis using BIOVIA Discovery Studio v.3.5.To evaluate the binding potential within a human-specific context, a molecular docking study of the most potent compound was conducted against human α-glucosidase.30 The crystal structure of the enzyme in complex with acarbose (PDB ID: 2QMJ) was retrieved from the RCSB Protein Data Bank (https://www.rcsb.org). The active site coordinates were defined based on the crystallographic position of the co-crystallized acarbose ligand using BIOVIA Discovery Studio Visualizer 2019. Docking was performed with a grid box centered at coordinates (x = −20.174, y = −6.206, z = −5.281 Å) and dimensions of 50 × 50 × 50 Å, encompassing the catalytic site. A flexible ligand docking protocol with 100 independent runs was employed. The most favorable binding pose for each ligand, as determined by the docking algorithm's scoring function, was selected for detailed interaction analysis. All molecular visualizations and interaction analyses were performed using BIOVIA Discovery Studio Visualizer 2019.
Molecular dynamics
The molecular dynamics (MD) behavior of α-glucosidase in complex with acarbose and the most potent inhibitor was investigated using the GROMACS 2024.1 simulation package, which was executed on Ubuntu 18.04.5 LTS.31 The protein topology was generated using the CHARMM36 force field and ligand topology was generated using the CHARMM General Force Field (CGenFF). Ligand and protein structures were converted to GROMACS-compatible.gro format and manually merged using Notepad++. The resulting protein–ligand complex was placed at the center of a cubic simulation box with a 1.0 nm buffer from the box edges. Solvation was performed using the SPC216 water model, and 15 water molecules were substituted with Na+ ions to neutralize the system's net negative charge. Energy minimization was conducted using the steepest descent algorithm for up to 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps, with convergence defined as a maximum force below 10.0 kJ mol−1. Equilibration was carried out in two phases: first, under constant volume (NVT ensemble) using the velocity-rescale (v-rescale) thermostat at 300 K, with a coupling constant of 0.1 ps for 500 ps; second, under constant pressure (NPT ensemble) using the Parrinello–Rahman barostat with a coupling constant of 5.0 ps for a duration of 1000 ps. Long-range electrostatics were computed using the Particle Mesh Ewald (PME) method, and van der Waals interactions were handled with a cut-off scheme. The cut-off distances were set at 1.0 nm for electrostatics and 1.2 nm for van der Waals interactions. Following system stabilization, each compound was subjected to three independent 100 nanosecond molecular dynamics simulations.
000 steps, with convergence defined as a maximum force below 10.0 kJ mol−1. Equilibration was carried out in two phases: first, under constant volume (NVT ensemble) using the velocity-rescale (v-rescale) thermostat at 300 K, with a coupling constant of 0.1 ps for 500 ps; second, under constant pressure (NPT ensemble) using the Parrinello–Rahman barostat with a coupling constant of 5.0 ps for a duration of 1000 ps. Long-range electrostatics were computed using the Particle Mesh Ewald (PME) method, and van der Waals interactions were handled with a cut-off scheme. The cut-off distances were set at 1.0 nm for electrostatics and 1.2 nm for van der Waals interactions. Following system stabilization, each compound was subjected to three independent 100 nanosecond molecular dynamics simulations.
Free binding energy calculations
The binding free energy of the protein–ligand complex was calculated using the molecular mechanics Poisson–Boltzmann surface area (MM/PBSA) method, with entropic contributions neglected. The binding free energy (ΔGbind) was defined as the sum of the molecular mechanics interaction energy (MMIE) and the solvation energy (SE):| ΔGbind = MMIE + SE |
| MMIE = EvdW + Eelec |
SE accounts for the solvation effects and is divided into polar and nonpolar contributions:
| SE = PSE + SASA energy | (1) |
Polar solvation energy (PSE) is derived from the Poisson–Boltzmann equation:
| PSE = PSEcomplex − (PSEprotein + PSEligand) | (2) |
Nonpolar solvation energy (SASA energy) is estimated based on solvent-accessible surface area (SASA):
| SASAenergy = SASAcomplex − (SASAprotein + SASAligand) | (3) |
Binding free energy calculations were carried out using 200 snapshots taken at 100 ps intervals from the last 20 ns of the molecular dynamics trajectory, ensuring representative sampling of protein–ligand interactions during the equilibrated phase of the simulation.
Druglikeness and ADMET prediction
Druglikeness and ADMET prediction of acarbose and the most potent compounds 19d, 15f, 15b, and 15e were determined from the preADMET online server (https://preadmet.bmdrc.org/).32Density functional theory Calculations
The frontier molecular orbital (FMO) energies and FMO-based chemical reactivity descriptors of the synthesized quinazolinone–thiophene derivatives 14a–f, 15a–f, and 19a–f were investigated using density functional theory (DFT).33 Geometry optimizations of all compounds were initially performed at the B3LYP/def2-SVP level of theory, followed by single-point energy evaluations on the geometry-optimized structures at the same theoretical level.34 All DFT calculations were executed using the ORCA software package (version 6.1.0).35 Chemical reactivity descriptors, including HOMO and LUMO energies, energy gap, chemical potential, global hardness, and global softness, were calculated employing equations derived from Koopmans' theory.36 Input and output files were generated and visualized using the ORCA-enhanced Avogadro graphical interface and ChemCraft visualization program.37Antioxidant activity
The antioxidant activity of the compound at different concentrations was evaluated using the DPPH radical scavenging assay. A series of sample dilutions were prepared and mixed with the DPPH solution, and the decrease in absorbance (517 nm) was measured after incubation. The radical scavenging activity was calculated and expressed as inhibition percentage (% inhibition). Trolox was used as the reference antioxidant standard for comparison.Result and discussion
Chemistry
Various derivatives 14a–f, 15a–f, and 19a–f with a quinazolinone–thiophene scaffold were synthesized through the reactions depicted in Fig. 2. First, quinazolinone derivatives 5a–f were synthesized in two steps: (1) reaction between isatoic anhydride (1) and amine derivatives 2a–f in acetic acid and ethanol to produce 2-aminobenzamide derivatives 3a–f, and (2) reaction between the latter derivatives with carbon disulfide (CS2, 4) in alcoholic potassium hydroxide. On the other hand, for the synthesis of target compounds, three thiophene derivatives were required: 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carbonitrile (9), 2-amino-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophene-3-carbonitrile (10), and ethyl 2-amino-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (17). For the synthesis of compounds 9 and 10, cyclohexanone (6) or cycloheptanone (7) reacted with malononitrile (8) and S8 in the presence of NEt3 in ethanol, and for the synthesis of compound 17, cyclohexanone (6) reacted with ethyl 2-cyanoacetate (16) and S8 in the presence of NEt3 in ethanol. After that, compounds 9, 10, and 17, reacted with chloroacetyl chloride (11) in 1,4-dioxan to give 2-chloro-N-(3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophen-2-yl)acetamide (12), 2-chloro-N-(3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen-2-yl)acetamide (13), and ethyl 2-(2-chloroacetamido)-4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate (18), respectively. In the final step, compounds 12, 13, and 18 reacted with quinazolinone derivatives 5a–f in DMF to give target compounds 14a–f, 15a–f, and 19a–f.Anti-α-glucosidase activity of the new compounds
Our synthesized new compounds were tested against yeast α-glucosidase (Table 1). In this assignment, like most research in the field of α-glucosidase inhibitors, acarbose was used as the positive control. In comparison to acarbose, all the newly synthesized quinazolinone–thiophene derivatives, except one compound, were active against α-glucosidase. The most active compound was 11.7 times more potent than acarbose, and the least active compound was negligibly more potent than acarbose.| Compound | R | IC50 (µM) |
|---|---|---|
| 14a | –CH2–CH3 | 328.7 ± 0.9 |
| 14b | –CH2–CH2–CH3 | 648.7 ± 0.7 |
| 14c | –CH2–CH2–CH2–CH3 | 601.7 ± 1.4 |
| 14d | –Ph | 410.4 ± 1.1 |
| 14e | –CH2–Ph | 469.6 ± 0.3 |
| 14f | –CH2–CH2–Ph | 721.5 ± 0.3 |
| 15a | –CH2–CH3 | 622.5 ± 0.7 |
| 15b | –CH2–CH2–CH3 | 184.7 ± 1.0 |
| 15c | –CH2–CH2–CH2–CH3 | 433.9 ± 1.2 |
| 15d | –Ph | 454.8 ± 0.6 |
| 15e | –CH2–Ph | 210.8 ± 0.4 |
| 15f | –CH2–CH2–Ph | 168.4 ± 0.6 |
| 19a | –CH2–CH3 | > 750 |
| 19b | –CH2–CH2–CH3 | 467.5 ± 0.2 |
| 19c | –CH2–CH2–CH2–CH3 | 324.8 ± 1.0 |
| 19d | –Ph | 64.0 ± 0.7 |
| 19e | –CH2–Ph | 377.5 ± 1.4 |
| 19f | –CH2–CH2–Ph | 445.1 ± 1.5 |
| Acarbose | — | 750.1 ± 1.3 |
Structure–activity relationships (SARs)
Structurally, new synthesized quinazolinone–thiophene derivatives were divided into three groups: (1) 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophens 14a–f, (2) 3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophens 15a–f, and (3) ethyl 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylates 19a–f. In each group, substituents on N-3 of the quinazolinone ring were altered for optimization of anti-α-glucosidase activity. As observed in Table 1, the most active compound and the only inactive compound among the synthesized compounds belonged to the third series. SARs of these derivatives demonstrated that compound 19d with a phenyl substituent on N-3 of the quinazolinone ring was the most potent compound in this series and among all synthesized compounds. Insertion of one or two methylene groups between the phenyl ring and the N-3 unit of the quinazolinone ring, as in the cases of compounds 19e and 19f, anti-α-glucosidase activity diminished to 5.9 and 6.9-fold, respectively. On the other hand, replacing the phenyl group of compound 19d with an ethyl group resulted in the loss of inhibitory activity, as observed in compound 19a. Introduction of n-propyl and n-butyl groups on N-3 unit of the quinazolinone ring instead of the phenyl group of compound 19d, as in the cases of compounds 19b and 19c, created a significant decrease in anti-α-glucosidase activity.Among the 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophene series 14a–f, no compound showed a significant effect against α-glucosidase. In contrast, the second, third, and fourth potent compounds among all the synthesized compounds belonged to the 3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophene series 15a–f. The SAR study demonstrated that in this series, after the most potent compound 19d of ethyl 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate series, phenethyl, n-propyl, and benzyl derivatives 15f, 15b, and 15e of 3-cyano-5,6,7,8-tetrahydro-4H-cyclohepta[b]thiophen series showed significant inhibition effects against α-glucosidase. The remaining derivatives in the latter series did not show substantial anti-α-glucosidase activity.
Comparison of anti-α-glucosidase activity of new title compounds with the used templates C, E, and F
The comparison of IC50 values of the 3-phenethyl-quinazolin-2-thio derivatives of the template compound C with their corresponding analogs of new quinazolinone–thiophene derivatives is shown in Fig. 3.22 As can be seen in this figure, the most potent compound among the 3-phenethyl-quinazolin-2-thio derivatives of the template compound C was less potent than the new compound 15f and more potent than the new compounds 14f and 19f. | ||
| Fig. 3 The comparison of the anti-α-glucosidase activity of the template compounds A with their corresponding analogs of new quinazolinone–thiophene derivatives. | ||
In the design of compounds 14a–f and 15a–f, the 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophene scaffold of compound E was used.24 As can be seen in Fig. 4, a comparison of compound E with the most potent compounds among the compounds 14a–f (compound 14a) and 15a–f (compound 15f) demonstrated that modifications in this project did not improve anti-α-glucosidase activity because compound E was 6.9-fold more potent than acarbose, while compounds 14a and 15f were 2.3 and 4.5 folds more potent than acarbose, respectively.
 | ||
| Fig. 4 The comparison of anti-α-glucosidase activity of the reported 3-cyano-4,5,6,7-tetrahydrobenzo[b]thiophene derivative E with new thiophen derivatives 14a and 15f. | ||
As was mentioned in the introduction, the ethyl 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate moiety was selected for compound F.25 Comparison of the IC50 value of this compound with the most potent compound among the newly synthesized ethyl 4,5,6,7-tetrahydrobenzo[b]thiophene-3-carboxylate derivatives 19a–f (compound 19d), demonstrated that our modification improved inhibitory activity against α-glucosidase because compound 19d was 11.7-fold more potent than acarbose, while compound F was 9.7 times more potent than acarbose (Fig. 5).
Kinetic study
The inhibition mode of the title compounds against α-glucosidase was determined by a kinetic study. Compound 19d, based on its IC50 value, was selected as a representative α-glucosidase inhibitor for kinetic study using Lineweaver–Burk plot analysis. As shown in Fig. 6a, the graphs of 1/V and 1/[S] produce lines with different slopes by having the same Vmax but increasing Michaelis constant (Km) as the concentration of compound 19d increased. The above results indicated that compound 19d was a competitive α-glucosidase inhibitor. The calculation of Ki was performed by secondary plot between 1/Vmax and various concentrations of compound 19d (Fig. 6b). The Ki value for compound 19d was 69 µM.Docking study
In order to gain insight into the interaction modes of new synthesized compounds in the α-glucosidase's active site, a molecular docking study was conducted using AutodockTools (version 1.5.6) on modeled α-glucosidase.27–29 The reliability of the molecular docking protocol was validated by re-docking the co-crystallized ligand (maltose) into the active site, yielding a satisfactory root-mean-square deviation (RMSD) value of less than 2.0 Å. To start this process, acarbose, as a standard inhibitor and compound 19d, as the most potent new compound, were placed in the active site, and the obtained superimposed structure is shown in Fig. 7. | ||
| Fig. 7 Superimposition of the structure of acarbose (pink) and compound 19d (orange) in the α-glucosidase's active site. | ||
According to our previous reported works, acarbose, with a binding energy (BE) of −4.4 kcal mol−1 interacted with residues His279, Thr301, Asn241, Thr307, Pro309, Ser308, Arg312, Glu304, and Gln322.24 The binding mode of the most potent compound 19d showed that this compound established two hydrogen bonds with residues Glu304 and Gln322 and two non-classical hydrogen bonds with Arg312 and Gly306 (Fig. 8). Moreover, compound 19d via the sulfur atom of the thiophene ring interacted with amino acid Phe311. Furthermore, several hydrophobic interactions between this compound and active site residues His279, His239, Thr301, Val305, Pro309, Arg312, and Phe157 were also observed. The BE of this compound was −9.04 kcal mol−1. The second potent compound 15f established two hydrogen bonds with His279 and Ser308 and the following π-interactions with active site residues: a π-sulfur with His239 via thioacetamide moiety, a π-anion with Glu304, a π–π with His279, and two weak π-alkyl interactions with Arg312 and Pro309 (Fig. 8). The BE of compound 15f was −8.99 kcal mol−1.
The interaction modes of the third and fourth potent compounds, 15b and 15e, are shown in Fig. 9. Compound 15b formed two hydrogen bonds with His279 and Asn241 via cyno group. The sulfur atom of the thiophene ring interacted with residues Arg312 and His239. This compound also established hydrophobic interactions with Val305, His279, Thr301, His239, and Pro309. The BE of compound 15b was −8.99 kcal mol−1. The fourth potent compound 15e, with a BE value of −8.74 kcal mol−1, formed two hydrogen bonds with Glu304 and His279, two π–π interactions with His279 and His239, and three hydrophobic interactions with Pro309 and Arg312.
The binding energy values of the studied compounds in the docking study, compounds 15b, 15e, 15f, and 19d, are shown in Table 2. For better comparison, IC50 values of these compounds were obtained from nonlinear regression of triplicate measurements (n = 3) and compared by one-way ANOVA with Tukey post hoc tests; significant differences relative to acarbose are indicated (p < 0.05 A) and tabulated in Table 2. A review of these results shows that the docking studies largely corroborate the findings obtained from the in vitro experiments.
| Compound | Binding energy (kcal mol−1) | IC50 against α-glucosidase (µM) |
|---|---|---|
| 15b | −8.99 | 184.7 ± 1.0 |
| 15e | −8.74 | 210.8 ± 0.4 |
| 15f | −8.99 | 168.4 ± 0.6 |
| 19d | −9.04 | 64.0 ± 0.7 |
| Acarbose | −4.4 | 750.1 ± 1.3 |
To further validate the inhibitory potential of the most potent analog (compound 19d) and provide a direct comparison with the clinical inhibitor acarbose, we performed a molecular docking study within the active site of human α-glucosidase (PDB ID: 2QMJ). Superimposed structure of acaerbose and compound 19d in the active site of human α-glucosidase was shown in Fig. 10. The applied docking procedure was validated by extracting and re-docking the co-crystallized inhibitor acarbose into the α-glucosidase active site, RMSD < 2 Å.
 | ||
| Fig. 10 Superimposed structure of acarbose (pink) and compound 19d (orange) in the human α-glucosidase's active site. | ||
Detailed interaction modes of acarbose and compound 19d was shown in Fig. 11. As can be seen in this figure, acarbose established eleven hydrogen bonds with residues Gln603 (two interactions), Asp443 (two interactions), Asp54 (two interactions), Asp203 (two interactions), Arg526 (two interactions), and Tyr605 (one interaction). This drug also created a non-classical hydrogen bond with Asp327, a π-lone pair interaction with Tyr299, a sulfur interaction with Met444, and a hydrophobic interaction with Phe575 (Fig. 11). Fig. 11 illustrates the 2D and 3D interaction modes of compound 19d within the active site of human α-glucosidase. Compound 19d forms three hydrogen bonds with residues Thr205, Arg526, and Arg202, as well as two sulfur-containing interactions with Met444. Additionally, two π-anion interactions with Asp203 and Asp542, along with a π-cation interaction with Arg526, are observed. The interaction profile of compound 19d also includes several hydrophobic interactions with residues Leu473, Thr204, Phe450, Trp406, Tyr299, and Lys480. The binding energy (BE) calculations revealed values of −5.99 kcal mol−1 for acarbose and −8.02 kcal mol−1 for compound 19d in the active site of human α-glucosidase. These computational results indicate that the novel compound 19d exhibits a stronger predicted binding affinity and, consequently, a potentially superior inhibitory potency against human α-glucosidase compared to the clinical drug acarbose.
Molecular dynamics
The binding of a ligand to the binding site of a receptor, similar to other molecular interactions, is a dynamic process. Thus, simulating and studying the behavior of a receptor–ligand complex in a realistic environment, including water and ions, can provide valuable insights into the stability and flexibility of the receptor–ligand complex. In this study, molecular dynamics (MD) simulations were performed on the docking files of acarbose, a standard inhibitor, and compound 19d, the most potent inhibitor against α-glucosidase based on in vitro studies. The simulations were conducted in an explicit hydration environment to assess the stability, flexibility, and intermolecular interactions between the protein and the compounds throughout the simulation period.A two-phase simulation approach was employed. First, a 10 ns simulation was performed for all complexes, where both acarbose and compound 19d exhibited stability within the active site of α-glucosidase. Following this, the simulation was extended for 90 ns to further investigate the behavior of the compounds in the enzyme's active site. Remarkably, the compounds remained stable during the extended simulation. In the next stage, the simulation trajectories of the complexes were subjected to a more detailed analysis.
Root-mean-square deviation (RMSD) and radius of gyration (Rg) were calculated for all saved structures during the MD simulation. The variations in these parameters over time were used to evaluate the stability of the complexes. Additionally, the root mean square fluctuation (RMSF) of the backbone atoms was calculated to assess the residual flexibility throughout the simulation period.
The RMSD results are shown in Fig. 12. As indicated, the RMSD of the backbone atoms of α-glucosidase in free enzyme and in complex with compound 19d and acarbose displayed minimal fluctuations over time, consistently remaining below 0.3 nm, suggesting a stable protein structure. The average RMSD values for α-glucosidase in free enzyme and in complexes with acarbose and 19d were 0.253 nm, 0.251 nm, and 0.249 nm, respectively. The RMSD values for acarbose and 19d in complex with α-glucosidase remained under 0.2 nm throughout the simulation. The average RMSD values for acarbose and 19d in complex with α-glucosidase were 0.120 nm and 0.075 nm, respectively. These findings confirm the stability of both the enzyme and the ligands during the entire simulation period.
Protein stability can be evaluated by analyzing its compactness during MD simulation. The radius of gyration (Rg) of α-glucosidase was calculated to determine the protein's compactness over the course of the simulation (Fig. 13). The average Rg of α-glucosidase was 2.496 nm for free enzyme, 2.504 nm for the acarbose complex, and 2.510 nm for the 19d complex. Throughout the simulation, the Rg values fluctuated between 2.450 nm and 2.550 nm for all complexes. These consistent values indicate minimal changes in the protein's compactness, suggesting that the overall structure of α-glucosidase remained stable during the simulation period.
 | ||
| Fig. 13 Time dependence of the radius of gyration graph of α-glucosidase in complex with acarbose (green), compound 19d (red) and free enzyme (black). | ||
α-Glucosidase is a large protein composed of 579 residues, divided into multiple domains with distinct structures and functions. As shown in Fig. 10, the fluctuations across different regions of this large protein are not uniform. However, the RMSF values of the residues in free enzyme and in complex with acarbose and/or compound 19d demonstrate a remarkable similarity. As expected, residues engaged in non-bonded interactions with the ligands display lower fluctuations. The active site of α-glucosidase is located within a cleft formed by the A and B domains. As shown in Fig. 11, the residues of “A domain” have a noticeably higher fluctuation in the free enzyme versus the ligand-bound enzyme, which can be an indicator of non-bonded interactions of the residues of this domain with the ligands in the ligand-bound enzyme. As is typical for many proteins, the loop regions of α-glucosidase exhibit the highest level of fluctuations. In particular, residues in the B domain loop and the active site lid show the highest RMSF values in both free and ligand-bound enzymes.
Fig. 14 presents the fluctuations of heavy atoms in acarbose and 19d. The root mean square fluctuation (RMSF) of these atoms remains below 0.15 nm. Such limited fluctuation indicates strong and stable interactions between the ligands and α-glucosidase, suggesting that their movements are significantly restricted by intermolecular forces, thus maintaining the integrity of the ligand–enzyme complex.
Analysis of the MD trajectories indicated that the number of hydrogen bonds formed between the ligands and α-glucosidase varied over time (Fig. 15). In the α-glucosidase–acarbose complex, the number of hydrogen bonds predominantly fluctuated between 4 and 8, suggesting the formation of a strong and stable complex. Docking results predicted that compound 19d forms two hydrogen bonds within the α-glucosidase binding site (Fig. 8). In contrast, MD simulations revealed that the number of hydrogen bonds in the α-glucosidase–19d complex mainly ranged between two and three. Such discrepancies between docking and MD simulation outcomes are expected, as both ligand and receptor undergo conformational fluctuations during MD simulations, giving rise to a dynamic and diverse set of interactions. Nevertheless, subsequent binding free energy analysis demonstrated that the cumulative effect of these interactions favored the stable binding of compound 19d to α-glucosidase.
 | ||
| Fig. 15 The numbers of hydrogen bonds between compound 19d (A), and acarbose (B) with α-glucosidase binding site residues during MD simulation. | ||
Free binding energy calculations
The binding affinity of a ligand toward a protein receptor can be evaluated using the molecular mechanics/Poisson–Boltzmann surface area (MM/PBSA) approach, which also provides insight into the dominant interactions governing ligand–receptor complex formation. Binding energies estimated solely from molecular docking are generally less reliable because they are based on a single structural snapshot. In contrast, MD simulations generate multiple snapshots over time, enabling a more accurate calculation of binding free energy. The results of the MM/PBSA binding free energy analysis are summarized in Table 3. Both acarbose and compound 19d exhibited negative binding free energies in this study. The average MM/PBSA binding free energy of the reference inhibitor acarbose with α-glucosidase was −122.4 kJ mol−1, whereas compound 19d showed a binding free energy of −91.9 kJ mol−1. Fig. 16 illustrates the binding energy fluctuations over the final 20 ns of the MD simulation. For both complexes, the binding energies remained within a narrow negative range, indicating stable ligand–protein interactions. Although compound 19d displayed a weaker binding energy compared to acarbose, it remained stably accommodated within the α-glucosidase binding pocket. Notably, a binding free energy of −91.9 kJ mol−1 is sufficient to support stable complex formation between a small molecule such as 19d and α-glucosidase. Decomposition of the binding free energy revealed that molecular mechanics interaction energies (van der Waals and electrostatic contributions) were favorable, whereas solvation energies (including polar solvation and SASA terms) were unfavorable for both α-glucosidase–acarbose and α-glucosidase–19d complexes. Specifically, van der Waals and electrostatic energies contributed negatively, while solvation energies contributed positively to the overall binding free energy in both systems. | ||
| Fig. 16 Diagram of binding energy changes during the last 20 ns of simulation time. α-glucosidase in complex with acarbose (green) and 19d (red). | ||
Density functional theory calculations
DFT analyses were performed to explore the electronic structure and reactivity of the studied compounds. The key parameters included frontier molecular orbital energies (HOMO and LUMO), energy gap (Egap), dipole Moment (Debye), electronegativity (χ), electrophilicity index (ω), chemical potential (µ), global softness (σ), and dipole moment, are summarized in Table 4.| Compound | HOMO (eV) | LUMO (eV) | Egap | Debye | χ | µ | σ | ω |
|---|---|---|---|---|---|---|---|---|
| 14a | −6.1295 | −1.4998 | 4.6297 | 6.3311 | 3.8147 | −3.8147 | 0.4320 | 3.1431 |
| 14b | −6.0133 | −1.6578 | 4.3555 | 2.0063 | 3.8356 | −3.8356 | 0.4592 | 3.3777 |
| 14c | −6.0824 | −1.5746 | 4.5078 | 4.9187 | 3.8285 | −3.8285 | 0.4436 | 3.2579 |
| 14d | −6.2017 | −1.4329 | 4.7688 | 3.2745 | 3.8173 | −3.8173 | 0.4193 | 3.0586 |
| 14e | −6.0542 | −1.6084 | 4.4458 | 5.1029 | 3.8313 | −3.8313 | 0.4500 | 3.3034 |
| 14f | −6.1669 | −1.4861 | 4.6808 | 1.8426 | 3.8265 | −3.8265 | 0.4273 | 3.1338 |
| 15a | −6.2478 | −1.3654 | 4.8824 | 2.9181 | 3.8066 | −3.8066 | 0.4096 | 2.9728 |
| 15b | −6.0986 | −1.5523 | 4.5463 | 4.2075 | 3.8255 | −3.8255 | 0.4400 | 3.2254 |
| 15c | −6.1742 | −1.4728 | 4.7014 | 3.6892 | 3.8235 | −3.8235 | 0.4254 | 3.1159 |
| 15d | −6.2215 | −1.4219 | 4.7996 | 2.1478 | 3.8217 | −3.8217 | 0.4167 | 3.0421 |
| 15e | −6.0879 | −1.5896 | 4.4983 | 5.4362 | 3.8388 | −3.8388 | 0.4446 | 3.3509 |
| 15f | −6.1628 | −1.4907 | 4.6721 | 2.5634 | 3.8268 | −3.8268 | 0.4279 | 3.1389 |
| 19a | −6.0314 | −1.6429 | 4.3885 | 7.2846 | 3.8372 | −3.8372 | 0.4557 | 3.3586 |
| 19b | −6.1187 | −1.5234 | 4.5953 | 4.9618 | 3.8211 | −3.8211 | 0.4353 | 3.1742 |
| 19c | −6.1765 | −1.4712 | 4.7053 | 3.1189 | 3.8239 | −3.8239 | 0.4251 | 3.1141 |
| 19d | −6.2098 | −1.4396 | 4.7702 | 2.3867 | 3.8247 | −3.8247 | 0.4191 | 3.0599 |
| 19e | −6.0923 | −1.5868 | 4.5055 | 5.8124 | 3.8396 | −3.8396 | 0.4440 | 3.3632 |
| 19f | −6.1681 | −1.4849 | 4.6832 | 3.0046 | 3.8265 | −3.8265 | 0.4272 | 3.1346 |
The calculated HOMO energies ranged from −6.25 to −6.01 eV, indicating similar electron donation.38 The LUMO energies are clustered between −1.66 and −1.36 eV, suggesting comparable electron acceptance.38 The HOMO–LUMO Egap, an important indicator of chemical stability and reactivity, varies between 4.35 and 4.88 ev.39 Compounds with relatively smaller energy gaps (e.g., 14b, 19a, and 19b) are expected to exhibit higher chemical reactivity and lower kinetic stability, whereas compounds with larger gaps (e.g., 15a and 14d) are predicted to be more chemically stable and less reactive. Overall, the moderate Egap values suggest a balance between stability and reactivity, which is favorable for potential biological applications.40 The calculated dipole moments (Debyes) span a wide range, from ∼1.8 to ∼7.3 Debye, reflecting significant differences in molecular polarity and charge distribution. The χ values are narrowly distributed around 3.80–3.84 eV. Softness values (0.41–0.46 eV−1) were higher in reactive compounds like 14b and 19a, indicating greater polarizability.40 Narrower bandgaps, higher softness, and elevated electrophilicity (ω) correlated with better α-glucosidase inhibition, enhancing charge transfer and binding. For instance, 15b, 15e, 15f, and 19d exhibited balanced properties, likely improving interactions with enzyme residues. These points was also further confirmed by frontier molecular orbitals of the most potent compounds 19d and 15f (Fig. 17).
The electrostatic potential (ESP) maps of compounds 15f and 19d (Fig. 18) exhibit comparable global charge distribution patterns, characterized by distinct electron-rich and electron-deficient regions over the molecular surfaces. In both compounds, pronounced negative electrostatic potential is mainly localized around the carbonyl oxygen atoms and heterocyclic nitrogen atoms of the quinazolinone core, indicating strong hydrogen-bond acceptor capability. Conversely, regions of positive electrostatic potential are primarily associated with hydrogen-bearing atoms and peripheral alkyl or aromatic moieties.
 | ||
| Fig. 18 Electrostatic potential (ESP) maps of 19d and 15f (B3LYP). Red (negative) to blue (positive) gradient highlights interaction sites. | ||
The electrophilicity index (ω) was key predictor of bioactivity, with higher values aiding interactions with electron-rich amino acid residues (e.g., Asp, Glu, and His) within the enzyme active site.40 Notably, 19d, the most potent compound in this series, exhibits a balanced combination of a moderate energy gap, suitable softness, and an electrophilicity index comparable to many less active analogues.
Overall, the combined DFT and biological results demonstrate that electronic reactivity descriptors provide meaningful insight into the observed inhibitory trends. Compounds that achieve an optimal balance between electrophilicity, softness, and molecular polarity exhibit superior anti-α-glucosidase activity.
In silico drug-likeness, pharmacokinetics, and toxicity studies
Drug-likeness study and prediction of pharmacokinetics and toxicity (ADMET) of the positive control acarbose and most potent compounds 15b, 15e, 15f, and 19d were conducted using by PreADMET as an online software and the obtained data were listed in Table 5. This table showed that, except compound 15b, acarbose and all the studied new compounds 15e, 15f, and 19d did not follow of Lipinski ‘Rule of five’. Compound 15b and acarbose had poor permeability to Caco2 cell, while compounds 15e, 15f, and 19d had moderate permeability to these cells. Moreover, human intestinal absorption (HIA) of new compounds 15b, 15e, 15f, and 19d was high, while acarbose did not have HIA. Blood–brain barrier (BBB) permeability of acarbose and compounds 15b, 15e, 15f, and 19d was in the acceptable range. All these compounds were mutagenic. Moreover, compounds 15b, 15e, 15f, and 19d did not have carcinogenic effects on mice and rats, while acarbose had carcinogenic effects on mice. Cardiotoxicity (hERG inhibition) risk of new compounds 15b, 15e, 15f, and 19d was low, while this risk in acarbose was ambiguous.| Entry | Compound | ||||
|---|---|---|---|---|---|
| 15b | 15e | 15f | 19d | Acarbose | |
| a The recommended ranges for Caco2: <25 poor, >500 great, HIA: >80% is high <25% is poor, and BBB = −3.0–1.2. | |||||
| Rule of five | Suitable | Violated | Violated | Violated | Violated |
| Caco2a | 17.8621 | 28.5028 | 28.9142 | 26.6316 | 9.44448 |
| HIAa | 98.225697 | 98.471431 | 98.332416 | 98.404245 | 0.000000 |
| BBBa | 0.107165 | 0.15091 | 0.165436 | 0.342645 | 0.0271005 |
| Ames_test | Mutagen | Mutagen | Mutagen | Mutagen | Mutagen |
| Carcino_Mouse | Negative | Negative | Negative | Negative | Positive |
| Carcino_Rat | Negative | Negative | Negative | Negative | Negative |
| hERG_inhibition | Low risk | Low risk | Low risk | Low risk | Ambiguous |
Anti-proliferative assay
The potential cytotoxicity of the most potent compounds was evaluated against normal NIH-3T3 fibroblast cells using the MTT assay.41 NIH-3T3 cells were purchased from Iranian Biological Resource Center (IBRC), Tehran, Iran. Obtained data showed that the tested compounds 15b, 15f, and 19d and the reference chemotherapeutic agent cisplatin exhibited the IC50 values 67.4 ± 10.7 40.6 ± 1.2 µM, 20.1 ± 3.6, and 6.9 ± 0.8 µM, respectively. Therefore cisplatin demonstrated significantly greater cytotoxicity in comparison to all test compounds.Antioxidant activity
The antioxidant potential of our most potent compound 19d was evaluated in comparison to the standard antioxidant trolox. The results, presented in Fig. 19, clearly indicate that compound 19d is nearly inactive relative to trolox. As shown in this figure, our synthesized molecule demonstrates negligible radical scavenging activity and therefore cannot be classified as an effective antioxidant under the assay conditions used. | ||
| Fig. 19 Comparison of antioxidant activity between compound 19d and the reference standard trolox, measured using the DPPH radical scavenging assay. | ||
Conclusion
A new series of quinazolinone–thiophene derivatives (compounds 14a–f, 15a–f, and 19a–f) was synthesized and evaluated for yeast α-glucosidase inhibitory potential. All analogs, except compound 19a, exhibited inhibitory potentials in the IC50 values range of 64.0 to 721.5 µM as compared to the standard drug acarbose with an IC50 value of 750.1 µM. The most potent compound among the synthesized compounds was compound 19d that is a competitive inhibitor against yeast α-glucosidase. Therefore, compound 19d and other most potent compounds 15f, 15b, and 15e were docked in the active site of yeast α-glucosidase that constructed via homology modeling. The obtained results of the docking study were confirmed by in vitro anti-α-glucosidase activity results and an additional docking evaluation specifically on human α-glucosidase was performed for compound 19d, revealing a favorable binding energy. Furthermore, a dynamic study on the complex of this compound and α-glucosidase demonstrated that compound 19d formed a stable complex with α-glucosidase. DFT calculations providing a rational explanation for the superior inhibitory potency of the most potent compounds. The most potent compounds also showed low cytotoxicity against normal NIH-3T3 cells. Moreover, in silico pharmacokinetic studies predicted that the most potent compounds had druglikeness and pharmacokinetic profiles comparable to acarbose, but with a better toxicity profile.Author contributions
MM, BL, and SJ conceived, designed, and supervised this study. PB, ND, MN, and SNG synthesized and interpreted analytical data of compounds. In vitro experiments were performed by FG, MT, SM and MAF. In silico studies were performed and written by MH and MM-K. The manuscript was drafted by MM-K and MA. MM reviewed and edited the drafts.Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra08206d.References
- G. Roglic, Int. J. Noncommun. Dis., 2016, 1, 3–8 Search PubMed.
- D. Dabelea, E. J. Mayer-Davis, S. Saydah, G. Imperatore, B. Linder, J. Divers, R. Bell, A. Badaru, J. W. Talton, T. Crume and A. D. Liese, J. Am. Med. Assoc., 2014, 311, 1778–1786 Search PubMed.
- A. Kulkarni, A. R. Thool and S. Daigavane, Cureus, 2024, 16 Search PubMed.
- A. B. Olokoba, O. A. Obateru and L. B. Olokoba, Oman Med. J., 2012, 27, 269 CrossRef CAS PubMed.
- V. Thakare, S. S. Shende, P. A. Shirure and O. C. Swami, Int. J. Med. Sci., 2017, 5, 749 Search PubMed.
- C. W. Choi, Y. H. Choi, M. R. Cha, D. S. Yoo, Y. S. Kim, G. H. Yon, K. S. Hong, Y. H. Kim and S. Y. Ryu, J. Agric. Food Chem., 2010, 58, 9988–9993 CrossRef CAS PubMed.
- A. D. Baron, Diabetes Res. Clin. Pract., 1998, 40, S51–S55 CrossRef CAS PubMed.
- A. M. Dirir, M. Daou, A. F. Yousef and L. F. Yousef, Phytochem. Rev., 2022, 21, 1049–1079 Search PubMed.
- M. Taha, F. Rahim, S. Imran, N. H. Ismail, H. Ullah, M. Selvaraj, M. T. Javid, U. Salar, M. Ali and K. M. Khan, Bioorg. Chem., 2017, 74, 30–40 Search PubMed.
- J. Wang, S. Lu, R. Sheng, J. Fan, W. Wu and R. Guo, Mini-Rev. Med. Chem., 2020, 20, 1791–1818 Search PubMed.
- S. M. Khirallah, H. M. Ramadan, H. A. A. Aladl, N. O. Ayaz, L. A. Kurdi, M. Jaremko, S. Z. Alshawwa and E. M. Saied, Pharmaceuticals, 2022, 15, 1576 CrossRef CAS PubMed.
- H. Kashtoh and K. H. Baek, Plants, 2022, 11, 2722 CrossRef CAS PubMed.
- C. Viegas-Junior, A. Danuello, V. da Silva Bolzani, E. J. Barreiro and C. A. M. Fraga, Curr. Med. Chem., 2022, 14, 1829–1852 CrossRef PubMed.
- E. S. Moghadam, A. M. Al-Sadi, M. S. Moghadam, B. Bayati, S. Mojtabavi, M. A. Faramarzi, M. Amini and R. Abdel-Jalil, Future Med. Chem., 2024, 16, 2395–2410 Search PubMed.
- G. Wang, M. Chen, J. Wang, Y. Peng, L. Li, Z. Xie, B. Deng, S. Chen and W. Li, Bioorg. Med. Chem. Lett., 2017, 27, 2957–2961 CrossRef CAS PubMed.
- M. M. Khalifa, H. M. Sakr, A. Ibrahim, A. M. Mansour and R. R. Ayyad, J. Mol. Struct., 2022, 1250, 131768 CrossRef CAS.
- M. Wei, W. M. Chai, R. Wang, Q. Yang, Z. Deng and Y. Peng, Bioorg. Med. Chem., 2017, 25, 1303–1308 Search PubMed.
- F. Azimi, H. Azizian, M. Najafi, F. Hassanzadeh, H. Sadeghi-Aliabadi, J. B. Ghasemi, M. A. Faramarzi, S. Mojtabavi, B. Larijani, L. Saghaei and M. Mahdavi, Bioorg. Chem., 2021, 114, 105127 Search PubMed.
- M. M. Khalifa, H. M. Sakr, A. Ibrahim, A. M. Mansour and R. R. Ayyad, J. Mol. Struct., 2022, 1250, 131768 Search PubMed.
- M. Wei, W. M. Chai, R. Wang, Q. Yang, Z. Deng and Y. Peng, Bioorg. Med. Chem., 2017, 25, 1303–1308 Search PubMed.
- A. Yavari, M. Mohammadi-Khanaposhtani, S. Moradi, S. Bahadorikhalili, R. Pourbagher, N. Jafari, M. A. Faramarzi, E. Zabihi, M. Mahdavi, M. Biglar and B. Larijani, Med. Chem. Res., 2021, 30, 702–711 CrossRef CAS.
- M. Saeedi, M. Mohammadi-Khanaposhtani, P. Pourrabia, N. Razzaghi, R. Ghadimi, S. Imanparast, M. A. Faramarzi, F. Bandarian, E. N. Esfahani, M. Safavi and H. Rastegar, Bioorg. Chem., 2019, 83, 161–169 CrossRef CAS PubMed.
- Y. N. Mabkhot, A. Barakat, A. M. Al-Majid and M. I. Choudhary, Int. J. Mol. Sci., 2013, 14, 5712–5722 Search PubMed.
- J. H. Zhang, H. X. Xie, Y. Li, K. M. Wang, Z. Song, K. K. Zhu, L. Fang, J. Zhang and C. S. Jiang, Bioorg. Med. Chem. Lett., 2021, 52, 128413 Search PubMed.
- H. X. Xie, J. Zhang, Y. Li, J. H. Zhang, S. K. Liu, J. Zhang, H. Zheng, G. Z. Hao, K. K. Zhu and C. S. Jiang, Bioorg. Chem., 2021, 115, 105236 CrossRef CAS PubMed.
- M. Mohammadi-Khanaposhtani, S. Rezaei, R. Khalifeh, S. Imanparast, M. A. Faramarzi, S. Bahadorikhalili, M. Safavi, F. Bandarian, E. N. Esfahani, M. Mahdavi and B. Larijani, Bioorg. Chem., 2018, 80, 288–295 Search PubMed.
- S. Imran, M. Taha, N. H. Ismail, S. M. Kashif, F. Rahim, W. Jamil, M. Hariono, M. Yusuf and H. Wahab, Eur. J. Med. Chem., 2015, 105, 156–170 Search PubMed.
- S. Imran, M. Taha, N. H. Ismail, S. M. Kashif, F. Rahim, W. Jamil, H. Wahab and K. M. Khan, Chem. Biol. Drug Des., 2016, 87, 361–373 CrossRef CAS PubMed.
- F. Kiefer, K. Arnold, M. Künzli, L. Bordoli and T. Schwede, Nucleic Acids Res., 2008, 37, D387–D392 CrossRef PubMed.
- K. Zheng, Y. Wu, Q. Dai, X. Yan, Y. Liu, D. Sun, Z. Yu, S. Jiang, Q. Ma and W. Jiang, Int. J. Biol. Macromol., 2024, 266, 131126 CrossRef CAS PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1, 19–25 CrossRef.
- J. A. Pradeepkiran, S. B. Sainath and K. V. L. Shrikanya, in Brucella Melitensis, Academic Press, 2021, pp. 133–176 Search PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648–5652 Search PubMed.
- M. Bursch, J. M. Mewes, A. Hansen and S. Grimme, Angew. Chem., 2022, 134, e202205735 Search PubMed.
- F. Neese, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2012, 2, 73–78 Search PubMed.
- T. Koopmans, Physica, 1934, 1, 104–113 Search PubMed.
- M. D. Hanwell, D. E. Curtis, D. C. Lonie, T. Vandermeersch, E. Zurek and G. R. Hutchison, J. Cheminf., 2012, 4, 17 Search PubMed.
- R. G. Parr, in Horizons of Quantum Chemistry, Springer, 1979, pp. 5–15 Search PubMed.
- P. Geerlings, F. De Proft and W. Langenaeker, Chem. Rev., 2003, 103, 1793–1874 Search PubMed.
- R. G. Parr, L. V. Szentpály and S. Liu, J. Am. Chem. Soc., 1999, 121, 1922–1924 Search PubMed.
- M. Bashiri, A. Jarrahpour, S. M. Nabavizadeh, S. Karimian, B. Rastegari, E. Haddadi and E. Turos, Med. Chem. Res., 2021, 30, 258–284 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |