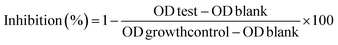Open Access Article
Open Access ArticleDual-targeting sulfamethoxazole derivatives overcome multidrug resistance in ESKAPE pathogens by inhibiting quorum sensing and efflux pumps
Benedicta Yayra Adzaha,
Prince Danan Biniyama,
Victoria Ohene-Adua,
Michael Oseia,
Kwabena Adu-Adjei a,
Victoria Elmesb,
Iain Goodallb,
Vuvor Faustinaa,
Yvonne Dogbeda Agboyibora,
Patrick Gyan
a,
Victoria Elmesb,
Iain Goodallb,
Vuvor Faustinaa,
Yvonne Dogbeda Agboyibora,
Patrick Gyan a,
Paul Quansaha,
Cyril Makafui Amengorc,
Joshua Boateng
a,
Paul Quansaha,
Cyril Makafui Amengorc,
Joshua Boateng b,
En Zhang
b,
En Zhang d,
Maxwell Ampomah-Wirekod,
Ernest Oyehe and
Cedric Dzidzor Kodjo Amengor
d,
Maxwell Ampomah-Wirekod,
Ernest Oyehe and
Cedric Dzidzor Kodjo Amengor *a
*a
aDepartment of Pharmaceutical Chemistry, Drug Discovery Unit, School of Pharmacy, University of Health and Allied Sciences, Ho, Ghana. E-mail: camengor@uhas.edu.gh; Tel: +233246456764
bSchool of Science, University of Greenwich, UK
cDepartment of Mathematics, University of Ghana, Legon, Ghana
dSchool of Pharmaceutical Sciences, Key Laboratory of Advanced Pharmaceutical Technology, Ministry of Education of China, Zhengzhou University, Zhengzhou 450001, PR China
eDepartment of Nursing, Faculty of Science, Methodist University Ghana, Ghana
First published on 13th February 2026
Abstract
The rapid emergence of pan-drug-resistant pathogens necessitates innovative antimicrobial strategies that overcome conventional resistance. This study reports the structure-guided design of sulfamethoxazole derivatives as dual inhibitors of quorum sensing (LasR) and efflux pumps (AcrB), alongside the classic dihydropteroate synthase (DHPS) target. High-throughput virtual screening of 54 derivatives, followed by MM-GBSA analysis, prioritized five novel compounds for synthesis. Their structures were confirmed by 1H/13C NMR, FT-IR, and mass spectrometry. All compounds demonstrated potent growth inhibition (MICs 15.625–125 µg mL−1) against Proteus mirabilis, Salmonella typhi, and Escherichia coli, though activity against the more resistant Pseudomonas aeruginosa was reduced. Critically, lead compounds SMX033 and SMX015 achieved >99% biofilm inhibition against P. mirabilis, and SMX033 showed significant efflux pump inhibition, directly validating our in silico predictions of a multi-target mechanism. In particular, compound SMX033 exhibited the best antimicrobial activity and lowest cytotoxicity of all the sulfamethoxazole derivatives with a CCC50 value of 286.20 µM against Vero cells. Despite their predicted in silico genotoxicity, these derivatives provide promising chemical scaffolds for combating multidrug-resistant infections by concurrently attenuating virulence and restoring antibiotic susceptibility.
1 Introduction
Bacterial-related infectious diseases pose a significant threat to global health due to their clinical diversity, high transmissibility, and the rapid emergence of antibiotic-resistant strains.1 These infections are the leading causes of morbidity and mortality worldwide.2 Antimicrobial resistance is a pressing global concern, as microorganisms develop resistance to existing antibiotics, rendering many treatments ineffective.3,4 This highlights the urgent need for novel antimicrobial agents with broad-spectrum efficacy, particularly against multidrug-resistant strains.Heterocyclic compounds have become a cornerstone in medicinal chemistry because of their potential versatile pharmacological activities.3 Therefore, synthetic medicinal chemists usually introduce a heterocyclic moiety, representing 85%, in the design of most drugs in order to enhance pharmacological activity while addressing metabolism and pharmacokinetic issues.3,4 Despite their ease of synthesis, the heterocyclic core imparts the clinical drug candidate with suitable pharmacokinetic properties.4 Most drugs fail to move from the bench to the patient because of poor pharmacokinetic profiles; therefore, the introduction of the heterocyclic groups can contribute to reducing the attrition rate of promising clinical candidates while improving the efficacy and toxicity profiles.5
The isoxazole class of heterocyclics continues to make an immense contribution to the drug discovery pipeline with the introduction of drugs possessing anti-inflammatory, antimicrobial, antioxidant, and other anti-infective properties.6,7 Drugs that possess the isoxazole group include sulfamethoxazole, sulfisoxazole, flucloxacillin, valdecoxib, leflunomide, and isoxicam8,9 (Fig. 1). One of the most common and clinically useful sulphonamides, sulfamethoxazole, is an antimicrobial drug used in combination with trimethoprim for the treatment of bacteria-related infections.10,11 It has established activity against Gram-positive and Gram-negative organisms, rendering it a broad-spectrum antibiotic, but it is gradually losing potency due to antimicrobial resistance.12 Current studies have demonstrated that sulfamethoxazole derivatives possess antimicrobial properties, and they represent promising clinical candidates. These include sulfamethoxazole ligand metal complexes, which showed potency against some ESKAPE pathogens (Staphylococcus aureus, Escherichia coli, and Klebsiella pneumoniae) and fungal strains of Aspergillus, and these complexes have the ability to inhibit bacterial biofilm formation.13,14 They have also been shown to inhibit the growth of virulent forms of Mycobacteria by blocking biofilm formation and efflux pumps of the bacteria.
Sulfonamides inhibit the para-amino benzoic acid (PABA) pathway, blocking folate synthesis required for DNA replication. However, the approach of drug discovery through targeting this bacteria mechanism is no longer effective due to antibiotic resistance.15,16 Moreover, the development of multi-drug resistance to sulfamethoxazole has also been attributed to mutation and generation of new resistance genes, quorum sensing, biofilm formation, presence of efflux pumps, and redundancy of the known inhibitory pathways.17–19 This has rendered the drug sulfamethoxazole/trimethoprim (co-trimoxazole) ineffective as the first choice of drug for the treatment of most infections.
The application of computational methods such as molecular docking and molecular dynamics simulations has greatly enhanced our understanding of how sulfamethoxazole derivatives interact with biological targets such as DHPS.20 These approaches provide high-resolution information about binding affinities, conformational dynamics, and molecular recognition.21 This combination of computational and experimental approaches has become an important part of current drug development, allowing for the rapid identification and optimization of new sulfamethoxazole compounds with enhanced pharmacological characteristics.22–24
In this current study, we employed a computationally guided approach to design a library of sulfamethoxazole derivatives targeting DHPS, LasR, and AcrB, and to explore their resulting potential new phenotypic antibacterial, efflux pump inhibitory, and biofilm inhibitory activities.
2 Experimental design
The study involved 54 sulfamethoxazole derivatives, with sulfamethoxazole as a benchmark drug.25 Ligands were designed using positional isomerization of 19 diverse substituents, electron donating (dimethylamino, hydroxy, methoxy, ethoxy, methylthio, methyl, acetamido), electron withdrawing (nitro, cyano, trifluoromethyl, carboxylic acid, methyl ester, sulfonic acid, formyl), and halogens (bromo, chloro, fluoro), on the aromatic ring of the sulfamethoxazole (SMX) core. Molecular docking and MMGBSA calculations were performed on DHPS, LasR, and AcrB proteins to estimate binding free energy. Five sulfamethoxazole derivatives were prioritized for synthesis from the in silico screening and antimicrobial evaluation, focusing on synthetic accessibility, structural diversity, and resource efficiency.3 Chemistry
The chemicals used for the synthesis were purchased from Merck® (Gillingham, UK) and were of analytical grade, except for the HPLC-grade solvents (methanol and acetonitrile) sourced from Fischer Scientific (Loughborough, UK). The progress of the reaction and the purity of the synthesized compounds were monitored by thin-layer chromatography (TLC) using aluminum plates pre-coated with silica gel, observed at a fluorescence wavelength of 254 nm. A calibrated pH meter (Eutech Instruments ECPH70042GS, Singapore) was employed to monitor the pH, which in turn offered information about the progress of the reaction. The purity of the compounds was further assessed by determining their melting points using a B-540 analyzer for melting points (Büchi Corporation, New Castle, DE, USA), and the reported melting points are uncorrected. The presence of key functional groups in the synthesized compounds was recorded using a PerkinElmer Spectrum 100 FT-IR spectrometer (PerkinElmer Inc., Waltham, MA, USA) equipped with attenuated total reflectance mode. NMR spectra were acquired on a Bruker Ascend III (500 MHz) spectrometer (Bruker BioSpin AG, Fällanden, Switzerland). Chemical shifts were reported in parts per million (δ) relative to tetramethylsilane (TMS), with the residual solvent serving as the lock solvent ([D6] DMSO, δ = 2.50 ppm for 1H and δ = 39.52, 77.5 ppm for 13C). High-resolution mass spectra were obtained using an HPLC (Shimadzu LC-2010A) in conjunction with a time-of-flight (quadrupole mass analyzer) mass spectrometer operating in positive ESI ionization mode.3.1 General procedure for the synthesis of the sulfamethoxazole hybrids25
A mixture of 3-amino-5-methylisoxazole (2.5 mmol, 1 equiv.) and substituted sulfonyl chlorides (2.5 mmol, 1.0 equiv.) was suspended in 30 mL of water. The pH of the suspension was adjusted and maintained at 8.0 by adding a 1 M Na2CO3 aqueous solution at room temperature and monitored using a pH meter probe. The progress of the reaction was tracked using TLC with a developing solvent of (CH2Cl2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) (Et2O) [60
(Et2O) [60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40]. Upon completion of the reaction, concentrated HCl was added slowly to lower the pH to 2.0. The resulting precipitate was collected by suction filtration, washed with water, and dried to yield the crude compound. The crude product was purified by silica column chromatography using a dichloromethane (CH2Cl2)
40]. Upon completion of the reaction, concentrated HCl was added slowly to lower the pH to 2.0. The resulting precipitate was collected by suction filtration, washed with water, and dried to yield the crude compound. The crude product was purified by silica column chromatography using a dichloromethane (CH2Cl2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) diethyl ether (Et2O) [60
diethyl ether (Et2O) [60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40] mobile phase through gradient elution to isolate the target compounds as solid products.
40] mobile phase through gradient elution to isolate the target compounds as solid products.
3.2 Spectral data
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1566 (–C
O), 1566 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm), 8.12 (2H, m, –ArH), 7.86 (1H, m, –ArH), 7.75 (1H, m, –ArH), 6.17 (1H, q, –ArNHisox), 2.95 (br, 1H, s, –NHSO2–), 2.36 (3H, d, J = 4.00 Hz –ArCH3); 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.04, 156.97, 134.44, 132.79, 132.60, 131.45, 125.38, 96.02, 12.67; HRMS (ESI): m/z calculated for C10H9N3O5S: 283.0263, found, [M + H]+: 284.0327.
CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm), 8.12 (2H, m, –ArH), 7.86 (1H, m, –ArH), 7.75 (1H, m, –ArH), 6.17 (1H, q, –ArNHisox), 2.95 (br, 1H, s, –NHSO2–), 2.36 (3H, d, J = 4.00 Hz –ArCH3); 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.04, 156.97, 134.44, 132.79, 132.60, 131.45, 125.38, 96.02, 12.67; HRMS (ESI): m/z calculated for C10H9N3O5S: 283.0263, found, [M + H]+: 284.0327.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1566 (–C
O), 1566 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm), 8.34 (2H, d. J = 8.00, –ArH), 8.08 (2H, d, J = 8.00, –ArH), 7.13 (1H, m, –ArH), 6.18–6.17 (1H, q, –ArNHisox), 2.56 (br, 1H, s, –NHSO2–), 2.36–2.35 (3H, d, J = 4.00 Hz –ArCH3); 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.17, 157.20, 145.12, 128.49, 124.47, 95.56, 50.20, 48.92, 49.78, 29.73, 12.67; HRMS (ESI): m/z calculated for C10H9N3O5S: 283.0263, found, [M + H]+: 284.0329.
CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm), 8.34 (2H, d. J = 8.00, –ArH), 8.08 (2H, d, J = 8.00, –ArH), 7.13 (1H, m, –ArH), 6.18–6.17 (1H, q, –ArNHisox), 2.56 (br, 1H, s, –NHSO2–), 2.36–2.35 (3H, d, J = 4.00 Hz –ArCH3); 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.17, 157.20, 145.12, 128.49, 124.47, 95.56, 50.20, 48.92, 49.78, 29.73, 12.67; HRMS (ESI): m/z calculated for C10H9N3O5S: 283.0263, found, [M + H]+: 284.0329.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1566 (–C
O), 1566 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.48 (br, 1H, s, –NHSO2–), 8.15 (2H, d, J = 8.00 Hz, –ArH), 7.91 (2H, d, J = 8.00 Hz, –ArH), 6.26–6.24 (1H, q, –ArNHisox), 3.95 (3H, s, –OCOCH3), 2.31 (3H, s, –ArCH3) 13C NMR (400 MHz, CDCl3 δ ppm) δ; δ; 171.38, 165.37, 142.67 134.64, 130.49, 127.13, 95.55, 52.74, 21.60, 12.73; HRMS (ESI): m/z calculated for C12H12N2O5S: 296.0567, found, [M + H]+: 297.0535.
CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.48 (br, 1H, s, –NHSO2–), 8.15 (2H, d, J = 8.00 Hz, –ArH), 7.91 (2H, d, J = 8.00 Hz, –ArH), 6.26–6.24 (1H, q, –ArNHisox), 3.95 (3H, s, –OCOCH3), 2.31 (3H, s, –ArCH3) 13C NMR (400 MHz, CDCl3 δ ppm) δ; δ; 171.38, 165.37, 142.67 134.64, 130.49, 127.13, 95.55, 52.74, 21.60, 12.73; HRMS (ESI): m/z calculated for C12H12N2O5S: 296.0567, found, [M + H]+: 297.0535.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1566 (–C
O), 1566 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.56 (br, 1H, s, –NHSO2–), 8.13 (1H, m, –ArH), 7.45 (1H, m, –ArH), 7.13–7.18 (2H, d, J = 8.00 Hz, –ArH), 6.11–6.10 (1H, q, –ArHisox), 2.31 (3H, s, –ArCH3) ; 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.01, 156.67, 138.03, 135.50, 134.56, 131.56, 127.79, 120.20, 95.27, 12.68; HRMS (ESI): m/z calculated for C10H935BrN2O3S: 315.9517, found, [M + H]+: 316.9591; calculated for C10H937BrN2O3S: 316.9517, found, [M + 2]+: 318.9565.
CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.56 (br, 1H, s, –NHSO2–), 8.13 (1H, m, –ArH), 7.45 (1H, m, –ArH), 7.13–7.18 (2H, d, J = 8.00 Hz, –ArH), 6.11–6.10 (1H, q, –ArHisox), 2.31 (3H, s, –ArCH3) ; 13C NMR (400 MHz, CDCl3 δ ppm) δ; 171.01, 156.67, 138.03, 135.50, 134.56, 131.56, 127.79, 120.20, 95.27, 12.68; HRMS (ESI): m/z calculated for C10H935BrN2O3S: 315.9517, found, [M + H]+: 316.9591; calculated for C10H937BrN2O3S: 316.9517, found, [M + 2]+: 318.9565.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1566 (–C
O), 1566 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.24 (br, 1H, s, –NHSO2–), 7.74–7.72 (2H, d, J = 8.00 Hz, –ArH), 7.70–7.68 (1H, d, J = 8.00 Hz, –ArH), 6.26–6.24 (1H, q, –ArNHisox), 2.40 (3H, s, –ArCH3), 2.37–2.36 (3H, d, J = 4.00 m –ArCH3) 13C NMR (400 MHz, CDCl3 δ ppm) δ 171.01, 157.53, 144.56, 136.02, 129.92, 127.10, 95.55, 21.60, 12.73; HRMS (ESI): m/z calculated for C11H12N2O3S: 252.0569, found, [M + H]+:253.0649.
CH–), 822, 798, 524 (Ar–C–H), 1H NMR (400 MHz, CDCl3 δ ppm) 8.24 (br, 1H, s, –NHSO2–), 7.74–7.72 (2H, d, J = 8.00 Hz, –ArH), 7.70–7.68 (1H, d, J = 8.00 Hz, –ArH), 6.26–6.24 (1H, q, –ArNHisox), 2.40 (3H, s, –ArCH3), 2.37–2.36 (3H, d, J = 4.00 m –ArCH3) 13C NMR (400 MHz, CDCl3 δ ppm) δ 171.01, 157.53, 144.56, 136.02, 129.92, 127.10, 95.55, 21.60, 12.73; HRMS (ESI): m/z calculated for C11H12N2O3S: 252.0569, found, [M + H]+:253.0649.4 Antimicrobial assay
4.1 Minimum inhibitory concentration assay
To determine the minimum inhibitory concentration (MIC), the micro-well dilution method outlined by Fetse et al. (2014) and Agyare et al. (2013) was used.26,27 This approach measures the development/growth of microorganisms at different antimicrobial concentrations. Aqueous solutions of the compounds were prepared at a concentration of 200 mg mL−1 by dissolving 0.6 g of each synthesized compound in 3 mL of sterile water. In 96-well plates, 100 µL of double-strength broth was added to each well, followed by the addition of the compounds at a series of doubling dilutions, with final concentrations ranging from 250 to 3.91 µg mL−1. The test organisms, standardized to an inoculum size of 1 × 106 CFU mL−1 using the 0.5 M MacFarland standard, were then added to each well at a volume of 20 µL. The plates were covered and incubated at 37 °C for 24 hours. After the incubation period, 25 µL of 1.25 mg mL−1 of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) solution was added to each well and further incubated at 37 °C for 30 minutes. The MIC was determined as the lowest concentration at which no visible change in colour from yellow to purple occurred following the addition of MTT, indicating no microbial growth.4.2 Crystal violet biofilm inhibition assay
The biofilm inhibitory activity against Proteus mirabilis, Salmonella typhi, Escherichia coli, and Pseudomonas aeruginosa was evaluated using a modified assay that retains crystal violet, adapted from Ofori et al. (2021).28 Bacterial cultures were grown on brain heart infusion (BHI) agar at 37 °C for 24 hours. Colonies were suspended in BHI broth and adjusted to 0.5 McFarland standard (∼1.5 × 108 CFU mL−1) using sterile saline. In sterile 96-well plates, 100 µL of standardized bacterial suspension was combined with 100 µL of BHI broth containing test compounds at final sub-MIC concentrations. Wells containing bacteria without compounds served as growth controls, while wells with BHI broth alone served as blanks. Plates were wrapped in aluminum foil and incubated statically at 37 °C for 24 hours. After incubation, planktonic cells were aspirated, and wells were washed three times with 200 µL phosphate-buffered saline (PBS, pH 7.4) to remove non-adherent cells. Adherent biofilms were fixed with 150 µL methanol for 15 minutes, stained with 0.1% (w/v) crystal violet for 15 minutes, and washed three times with distilled water. Bound dye was solubilized with 150 µL of 95% ethanol for 30 minutes. Absorbance was measured at 630 nm using a microplate reader (BioTek Synergy H1, USA). Percentage biofilm inhibition was calculated as:Each compound-pathogen combination was tested in triplicate wells across three independent experiments.
4.3 Ethidium bromide efflux pump inhibition assay
Efflux pump inhibition was assessed against P. mirabilis, S. typhi, E. coli, and P. aeruginosa using an ethidium bromide accumulation assay as reported by Danquah et al. (2018).29 Bacteria were cultured in nutrient broth with 0.4% glucose at 37 °C (150 rpm) until OD600 reached 0.8–1.0. Cells were centrifuged (3000×g, 10 min), washed twice with PBS, and resuspended in PBS to OD600 = 0.4. Aliquots (250 µL) of bacterial suspension were transferred into Eppendorf tubes containing sub-MIC concentrations (½ MIC) of test compounds. Controls included: positive control: 250 µL bacteria + 20 µg mL−1 verapamil. Negative control: 250 µL bacteria (no inhibitor). Blank: 250 µL PBS. Glucose (1.5 µL of 80% w/v) was added to energize efflux pumps. After pre-incubation (37 °C, 10 min), 2.5 µL EtBr (50 mg L−1) was added. Immediately, 250 µL aliquots were transferred to black 96-well plates. Fluorescence (λex = 530 nm, λem = 600 nm) was measured every minute for 60 min at 37 °C (BioTek Synergy H1). Relative fluorescence was plotted against time, and efflux pump inhibition was quantified by calculating the area under the curve (AUC) for fluorescence kinetics.30 All assays were performed in triplicate with three biological replicates.4.4. In vitro cytotoxicity
The in vitro cytotoxicity of the synthesized compounds were assessed using Vero cells (kidney cells from African green monkey: ATCC CCL-81) following the standard MTT method.30 The average absorbances of triplicate cells were recorded and used to determine the % growth inhibition was determined using the formula: 100 − [sample absorbance/control absorbance] × 100.5 Computational methods
5.1 Protein selection and preparation
Bacterial drug targets related to bacterial folic acid biosynthesis (3TZF), bacterial multidrug exporters (2W1B), and biofilm formation (6V7X) were considered in the molecular docking studies. Inhibiting folic acid biosynthesis eventually disrupts bacterial DNA replication, stability, and integrity, while targeting multidrug exporters and biofilm-related pathways reduces bacterial resistance and enhances treatment efficacy.31 The 3D structure of protein targets complexed with native ligands was retrieved from the Protein Data Bank (https://www.rcsb.org/). The protein structures were imported and pre-processed in Discovery Studio 2025 Client by deleting heteroatoms. Protein structures containing multiple identical chains were examined, and, for cases where the chains were structurally equivalent and each contained the active site, only one chain was retained to eliminate redundancy and minimize computational complexity. The protein structures were energy minimized using the AMBERf4SB force field, with Gasteiger charges assigned using Antechamber in Chimera to enhance the models for molecular docking and MMGBSA calculations.325.2 Ligand preparation and ADME-TOX
The 2D and 3D molecular structures of sulfamethoxazole and a curated library of 54 sulfamethoxazole derivatives were modeled and geometry-optimized using Avogadro molecular modeling software. Energy minimization was performed using the General Amber Force Field (GAFF) to refine the chemical structures and ensure accurate molecular geometries and electronic properties.33 Partial atomic charges were assigned via the Gasteiger method, and polar hydrogens were incorporated using UCSF Chimera. The final structures were exported in Sybyl Molecular Model (MOL2) and Structure Data File (SDF) formats for subsequent computational analyses, including molecular docking simulations and scoring assessments.Pharmacokinetic properties, such as absorption, distribution, metabolism, and excretion of the designed sulfamethoxazole derivative, were predicted from ADMETLAB 3.0 and SwissADME web servers. Toxicity parameters were assessed via Data Warrior software and ADMETLAB 3.0.
5.3 Molecular docking
Molecular docking was employed to explore the binding mechanisms of the sulfamethoxazole derivatives. A blind docking approach utilizing AutoDock Vina software was employed to generate the optimal binding poses and interactions between the modeled ligands and the prepared DHPS, AcrB, and LasR proteins.34 In this method the grid box was programmed to encompass the entire protein structure (bind docking) without predefining any residues as critical for binding. This approach ensures that the ligand has the opportunity to interact with any portion of the protein, allowing the identification of potential binding pockets and enabling the ligand to bind to the most favorable site. The molecular docking protocol was validated by retrieving the 3D structure of the native ligand co-crystallized with P. aeruginosa LasR protein from the Protein Data Bank. The ligand was redocked onto the protein structure using AutoDock Vina and a blind docking approach. Five exhaustiveness values (8, 64, 128, 512, and 1064) were tested to evaluate the influence of sampling thoroughness on the redocking accuracy.35 An improvement in accuracy was observed when increasing exhaustiveness from 8 to 64, while higher values provided no substantial additional benefit. An exhaustiveness value of 64 was selected for all subsequent docking runs. Root mean square deviation and interaction maps were assessed to ensure the accuracy of the docking protocol. The consistency of key interactions between the redocked ligand and the protein was also checked in relation to the experimentally determined binding site. This multi-faceted validation approach ensures the reliability, reproducibility, and accuracy of the docking protocol.5.4 Ligand efficiency
Ligand efficiency is a metric for evaluating the binding efficiencies of a ligand relative to its size. Ligand efficiency was calculated using the following formula:where ΔG is the binding free energy (in kcal mol−1), HAC represents heavy atom count, which is the number of non-hydrogen atoms in the ligand.
5.5 MM-PBSA calculations
The Molecular Mechanics Generalized Born Surface Area (MM-GBSA) method was used to estimate the binding free energy (ΔGbinding) between the protein and the optimized docked poses of each ligand. Binding free energies were calculated as the difference in free energy between the ligand–protein complex and the unbound states of the protein and ligand.36 The binding free energy was computed according to the following relationship:| ΔGbinding = ΔGcomplex − ΔGptotein − ΔGligand |
5.6 Molecular dynamics simulations
SMX015 and SMX033, selected based on docking performance and experimental validation, were subjected to molecular dynamics (MD) simulations in complex with DHPS and LasR, together with their respective apo forms. Simulations were performed using GROMACS 2023,37 with system preparation carried out using the CHARMM-GUI Solution Builder.38 Systems were solvated in TIP3P water boxes extending 10 Å from the protein surface and neutralized with 0.15 M NaCl.39 Protein parameters were assigned using the CHARMM36m force field, and ligand parameters were generated with CGenFF.40 Following energy minimization (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 steps, steepest descent), systems were equilibrated under NVT (2 ns, 303.15 K, v-rescale thermostat) and NPT (5 ns, 1 bar, C-rescale barostat) conditions. Production simulations were conducted for 100 ns using a leapfrog integrator (2 fs timestep). Hydrogen bonds were constrained with LINCS, long-range electrostatics were treated using PME (1.2 nm cutoff), and van der Waals interactions were smoothly switched between 1.0 and 1.2 nm.
000 steps, steepest descent), systems were equilibrated under NVT (2 ns, 303.15 K, v-rescale thermostat) and NPT (5 ns, 1 bar, C-rescale barostat) conditions. Production simulations were conducted for 100 ns using a leapfrog integrator (2 fs timestep). Hydrogen bonds were constrained with LINCS, long-range electrostatics were treated using PME (1.2 nm cutoff), and van der Waals interactions were smoothly switched between 1.0 and 1.2 nm.
Trajectory analyses were performed using GROMACS 2023.1. Structural stability and dynamics were evaluated using RMSD, RMSF, radius of gyration (Rg), and solvent-accessible surface area (SASA). Principal component analysis (PCA) was used to identify dominant conformational motions, and non-bonded interaction energies were computed to characterize ligand-binding energetics.
6 Results and discussion
6.1 Integrated computational profiling
| Compound ID | MW (Da) | TPSA (Å2) | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S S |
GI absorption | PPB | BBB permeant |
|---|---|---|---|---|---|---|
| SMX033 | 252.06 | 72.2 | −3.3378 | High | 92.00938 | No |
| SMX015 | 296.05 | 98.5 | −2.90194 | High | 92.75873 | No |
| SMX019 | 315.95 | 72.2 | −3.49824 | High | 94.69174 | No |
| SMX003 | 283.03 | 115.34 | −3.07732 | High | 85.5729 | No |
| SMX001 | 283.03 | 115.34 | −3.22374 | High | 95.448 | No |
| SMX | 253.05 | 98.22 | −2.64664 | High | 61.27968 | No |
| Compound ID | hERG | DILI | BSEP | Nephrotoxicity-DI | Neurotoxicity-DI |
|---|---|---|---|---|---|
| SMX033 | 0.075765 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 766 766 |
0.987![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 738 738 |
0.113![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 391 391 |
0.034758 |
| SMX015 | 0.043284 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 995 995 |
0.916![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 285 285 |
0.043675 | 0.012197 |
| SMX019 | 0.042108 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 762 762 |
0.998![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 342 342 |
0.068108 | 0.042637 |
| SMX003 | 0.061045 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 999 999 |
0.749![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 857 857 |
0.068014 | 0.001445 |
| SMX001 | 0.035825 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 983 983 |
0.809![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 684 684 |
0.069278 | 0.006155 |
| SMX | 0.036709 | 0.999![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 988 988 |
0.004074 | 0.071402 | 0.102![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 531 531 |
Predicted properties were classified using an empirical decision scale: values between 0.0–0.3 were considered excellent, 0.3–0.7 moderate, and 0.7–1.0 poor facilitating rapid prioritization of compounds.
The derivatives were found to have favourable oral absorption potential, but aqueous solubility was a major limitation, particularly for those featuring strong electron-withdrawing groups. This property could be improved by perhydrating the isoxazole ring of the potential drug-like derivatives.40 Positional isomerism also influenced distribution and metabolic stability. para-Substituted derivatives consistently demonstrated enhanced intestinal permeability relative to ortho or meta isomers.41 This geometric dependence extends to plasma protein binding, where ortho-substituted derivatives exhibit elevated affinity due to restricted conformational freedom promoting hydrophobic interactions.42 Metabolic susceptibility follows distinct electron gradients, with electron-donating groups reducing CYP450 inhibition liabilities and nitro and cyano substituents elevating risk for CYP3A4 and CYP2C19.43 Toxicity profiling revealed organ-specific risk stratification. Hepatotoxicity (DILI) correlated strongly with BSEP inhibition, which is linked to bile acid accumulation and mitochondrial dysfunction.44 Cardiotoxicity risks diverged significantly, with SMX052 showing a high predicted risk of severe hERG inhibition, which is potentially fatal. Neurotoxicity and nephrotoxicity probability depend on transport inhibition profiles, underscoring the necessity of integrated assessment of off-target effects at hepatic or renal transporters. Compound SMX012 (para-carboxylic acid) showed favourable solubility, negligible hERG inhibition, less BBB permeability, and low DILI risk, positioning it as a candidate for consideration. Trifluoromethyl derivatives were removed due to high PPB and BSEP inhibition. Furthermore, SMX016 (hepatotoxic) and SMX052 (cardiotoxic) were excluded due to intractable safety liabilities, while moderate-risk candidates such as SMX042 may require medicinal chemistry refinement to reduce CYP2C9 inhibition.
The molecular docking protocol was initially validated by redocking the bound ligand N-3-oxo-dodecanosyl-homoserine lactone (OHN) and sulphonamide drug (08D) with LasR and DHPS, respectively, using Autodock Vina software (Fig. 2 &3). As expected, redocked ligands tended to have lower calculated energy values. A root mean square deviation (RMSD) of 0.20 Å was obtained for the highest OHN-ranked pose, while a value of 1.12 Å was recorded for the lowest-ranked OHN pose. The RMSD values of the sulphonamide drug (O8D) were between 0.03 and 0.67 Å. These low RMSD values (RMSD < 2 Å) suggest that the docking protocol and scoring function employed can accurately reproduce the native binding pose.45 Furthermore, the redocked poses of the top-ranked eight complexes maintained all key interactions, implying that they were oriented in a biologically relevant manner within the binding sites, making the redocking parameters suitable for docking the designed compounds against the protein targets.46
The docking scores of the designed compounds ranged from −4.13 to −6.78 kcal mol−1, with the benchmark drug, sulfamethoxazole, exhibiting a score of −4.79 kcal mol−1. The results revealed a spectrum of binding affinities among the compounds, indicating potential for further investigation. The compounds were divided into four clusters based on docking score (D-score) and ligand efficiency, with more negative D-scores indicating stronger predicted binding affinities (Fig. 4). Electron-withdrawing (EWG) substituents consistently showed stronger binding affinities than electron-donating counterparts, as the sulfonamide group served as a hydrogen bond acceptor. The most potent derivative, SMX-018, possesses a para-sulfonic acid moiety, a strong EWG, achieving a docking score of −6.78 kcal mol−1. Analogues with other strong EWGs, such as carboxylic acids and trifluoromethyl groups, show moderate docking scores. Compounds functionalized with electron-donating groups exhibit moderate binding, with their docking scores clustering in a less negative range.47 The least potent compounds are those substituted with strong para-directed EDGs, such as p-dimethylamino, p-ethoxy, and p-nitro. The ligand efficiency (L.E.) metric generally trends with the D-score, confirming that the improved binding of EWG analogues is not an artifact of increased molecular size but a more efficient interaction.48
Sulfamethoxazole inhibits DHPS through conserved interactions, including the electrostatic anchoring of its sulfonyl group to Lys221 and π-stacking of its aniline ring with Phe190.49 To address the limitations related to resistance, our structure-guided optimization procedure yielded sulfamethoxazole derivatives that created a comprehensive interaction network with the DHPS active site, as shown in Fig. 5a–d. Protein-ligand analysis identified hydrogen bonds with residues Arg235, Ser222, Thr62, Pro64, Gly189, and Lys221, as well as extended hydrophobic interactions through π-stacking with Phe28 and Phe190, and pi-cation contacts with Lys221 and Arg255. The strategic recruitment of Arg255, a residue essential for pterin-site recognition, may synergize with hydrogen bond interactions involving Thr62, Arg63, and Pro64 to enhance stability.50 This binding strategy follows the substrate envelope principle by confining interactions within the native volume of p-aminobenzoic acid (PABA), thus avoiding mutation-prone regions (e.g., Phe33, Pro69, Thr67).51 By utilizing the phylogenetically conserved Arg255, this approach connects the PABA and pterin sites, leading to a hybrid inhibitory mechanism that is less prone to single-point mutations (Table 3).52
| Compound | Substituent (position) | Type | Relative strength | D-Score | L.E. |
|---|---|---|---|---|---|
| SMX | p-Amino | EDG | Very strong | −4.79 | −0.28 |
| SMX001 | o-Nitro | EWG | Strong | −4.17 | −0.22 |
| SMX002 | m-Nitro | EWG | Strong | −4.75 | −0.25 |
| SMX003 | p-Nitro | EWG | Strong | −4.34 | −0.23 |
| SMX004 | o-Cyano | EWG | Strong | −4.13 | −0.23 |
| SMX005 | m-Cyano | EWG | Strong | −5.01 | −0.28 |
| SMX006 | p-Cyano | EWG | Strong | −5.53 | −0.31 |
| SMX007 | o-Trifluoromethyl | EWG | Strong | −5 | −0.25 |
| SMX008 | m-Trifluoromethyl | EWG | Strong | −5.54 | −0.28 |
| SMX009 | p-Trifluoromethyl | EWG | Strong | −4.69 | −0.23 |
| SMX010 | o-Carboxylic acid | EWG | Moderate | −6.23 | −0.33 |
| SMX011 | m-Carboxylic acid | EWG | Moderate | −5.79 | −0.3 |
| SMX012 | p-Carboxylic acid | EWG | Moderate | −5.88 | −0.31 |
| SMX013 | o-Methyl ester | EWG | Moderate | −5.32 | −0.27 |
| SMX014 | m-Methyl ester | EWG | Moderate | −5.51 | −0.28 |
| SMX015 | p-Methyl ester | EWG | Moderate | −4.82 | −0.24 |
| SMX016 | o-Sulfonic acid | EWG | Strong | −5.48 | −0.27 |
| SMX017 | m-Sulfonic acid | EWG | Strong | −5.23 | −0.26 |
| SMX018 | p-Sulfonic acid | EWG | Strong | −6.78 | −0.34 |
| SMX019 | o-Bromo | EWG | Weak | −5.18 | −0.3 |
| SMX020 | m-Bromo | EWG | Weak | −5.08 | −0.3 |
| SMX021 | p-Bromo | EWG | Weak | −5.44 | −0.32 |
| SMX022 | o-Chloro | EWG | Weak | −4.79 | −0.28 |
| SMX023 | m-Chloro | EWG | Weak | −5 | −0.29 |
| SMX024 | p-Chloro | EWG | Weak | −5.33 | −0.31 |
| SMX025 | o-Fluoro | EWG | Weak | −4.57 | −0.27 |
| SMX026 | m-Fluoro | EWG | Weak | −5.43 | −0.32 |
| SMX027 | p-Fluoro | EWG | Weak | −4.54 | −0.27 |
| SMX028 | o-Formyl | EWG | Strong | −4.8 | −0.27 |
| SMX029 | m-Formyl | EWG | Strong | −5.29 | −0.29 |
| SMX030 | p-Formyl | EWG | Strong | −4.89 | −0.27 |
| SMX031 | o-Methyl | EDG | Weak | −4.79 | −0.28 |
| SMX032 | m-Methyl | EDG | Weak | −4.73 | −0.28 |
| SMX033 | p-Methyl | EDG | Weak | −4.77 | −0.28 |
| SMX034 | o-Methoxy | EDG | Strong | −5.03 | −0.28 |
| SMX035 | m-Methoxy | EDG | Weak | −5.12 | −0.28 |
| SMX036 | p-Methoxy | EDG | Strong | −4.63 | −0.26 |
| SMX037 | o-Hydroxy | EDG | Strong | −5.17 | −0.3 |
| SMX038 | m-Hydroxy | EDG | Weak | −5.1 | −0.3 |
| SMX039 | p-Hydroxy | EDG | Strong | −4.69 | −0.28 |
| SMX040 | o-Methylamino | EDG | Strong | −4.87 | −0.27 |
| SMX041 | m-Methylamino | EDG | Weak | −4.7 | −0.26 |
| SMX042 | p-Methylamino | EDG | Strong | −5.22 | −0.29 |
| SMX043 | o-Dimethylamino | EDG | Very strong | −4.91 | −0.26 |
| SMX044 | m-Dimethylamino | EDG | Weak | −4.84 | −0.25 |
| SMX045 | p-Dimethylamino | EDG | Very strong | −4.2 | −0.22 |
| SMX046 | o-Ethoxy | EDG | Strong | −4.17 | −0.22 |
| SMX047 | m-Ethoxy | EDG | Weak | −5.28 | −0.28 |
| SMX048 | p-Ethoxy | EDG | Strong | −4.28 | −0.23 |
| SMX049 | o-Acetamido | EDG | Moderate | −5.06 | −0.25 |
| SMX050 | m-Acetamido | EDG | Moderate | −6.22 | −0.31 |
| SMX051 | p-Acetamido | EDG | Moderate | −4.45 | −0.22 |
| SMX052 | o-Methylthio | EDG | Moderate | −4.93 | −0.27 |
| SMX053 | m-Methylthio | EDG | Moderate | −4.94 | −0.27 |
| SMX054 | p-Methylthio | EDG | Moderate | −4.48 | −0.25 |
MMGBSA is a computationally efficient method that combines molecular mechanics energies for both protein and ligand atoms with solvation terms derived from the Generalized Born model.53 This approach was utilized to refine the docking poses and served as an additional selection criterion to advance the designed compounds to the next stage. Following MMGBSA refinement in the PRIME module, twenty-four (24) derivatives progressed to the next phase of computational profiling (Table 4). These derivatives exhibited superior MM-GBSA binding energies against DHPS compared to sulfamethoxazole (SMX: −45.03 kcal mol−1), thereby confirming the effectiveness of substitution strategies across various chemical classes. High-affinity compounds included derivatives featuring electron donating groups (e.g., SMX051: −58.46 kcal mol−1), electron-withdrawing groups (e.g., SMX018: −49.62, SMX023: −49.35, SMX033: −47.86 kcal mol−1), and polar functionalities (e.g., SMX050: −49.68 kcal mol−1).
| Derivative | Bind | Coulomb | Covalent | Hbond | Lipo | Packing | Solv_GB | vdW |
|---|---|---|---|---|---|---|---|---|
| SMX | −45.03 | −21.71 | 3.26 | −2.95 | −9.82 | −3.98 | 24.36 | −34.18 |
| SMX005 | −46.93 | −19.81 | 2.28 | −1.09 | −9.72 | −1.23 | 19.48 | −36.82 |
| SMX010 | −47.24 | −35.96 | 3.31 | −2.07 | −9.15 | −0.94 | 29.28 | −31.71 |
| SMX012 | −47.66 | −28.35 | 2.82 | −2.74 | −9.79 | −2.33 | 26.67 | −33.94 |
| SMX015 | −45.86 | −15.39 | 2.53 | −2.16 | −10.76 | −2.44 | 21.86 | −39.49 |
| SMX017 | −46.96 | −26.88 | 3.85 | −1.54 | −8.76 | −1.11 | 24.52 | −37.03 |
| SMX018 | −49.62 | −25.01 | 2.57 | −2.62 | −8.73 | −2.75 | 21.79 | −34.86 |
| SMX020 | −46.24 | −21.66 | 2.97 | −2.15 | −10.82 | −1.99 | 21.78 | −34.38 |
| SMX023 | −49.35 | −27.66 | 3.01 | −2.17 | −12.92 | −3.92 | 30.58 | −36.28 |
| SMX024 | −46.12 | −21.79 | 2.98 | −2.47 | −13.07 | −4.14 | 25.75 | −33.39 |
| SMX026 | −45.54 | −27.88 | 3 | −2.18 | −11.57 | −3.95 | 30.83 | −33.77 |
| SMX029 | −49.17 | −31.5 | 3.99 | −2.89 | −10.15 | −3.88 | 33.52 | −38.25 |
| SMX030 | −47.51 | −32.69 | 3.33 | −2.68 | −10.25 | −4.12 | 33.39 | −34.48 |
| SMX032 | −47.01 | −20.31 | 3.03 | −2.19 | −13.09 | −3.93 | 25.26 | −35.78 |
| SMX033 | −47.86 | −9.19 | 0.12 | −1.43 | −12.61 | −1.41 | 11.55 | −34.89 |
| SMX035 | −45.45 | −21.53 | 2.75 | −2.24 | −12.87 | −3.97 | 29.83 | −37.41 |
| SMX037 | −45.46 | −24.98 | 0.31 | −1.34 | −8.85 | −0.89 | 19.21 | −28.92 |
| SMX038 | −46.21 | −25.92 | 3.25 | −2.32 | −11.62 | −4.02 | 28.28 | −33.86 |
| SMX041 | −48.43 | −22.47 | 3.46 | −2.28 | −12.99 | −4.01 | 27.05 | −37.18 |
| SMX042 | −46.89 | −21.07 | 3.21 | −2.94 | −10.85 | −3.99 | 25.82 | −37.07 |
| SMX044 | −45.53 | −10.35 | 0.95 | −1.46 | −12.85 | −0.97 | 14.37 | −35.21 |
| SMX047 | −49.28 | −18.47 | 3.73 | −2.16 | −12.75 | −2.06 | 22.54 | −40.11 |
| SMX050 | −49.68 | −32.27 | 7.06 | −3.11 | −11.44 | −3.97 | 35.22 | −41.16 |
| SMX051 | −58.46 | −43.82 | 5.1 | −3.58 | −11.15 | −3.97 | 40.32 | −41.36 |
| SMX053 | −50.59 | −26.91 | 5.73 | −2.12 | −13.11 | −3.89 | 30.82 | −41.1 |
The differences in the MMGBSA binding free energy between sulfamethoxazole and the filtered derivatives indicated that the derivatives fit better into the DHPS binding pockets or adjacent binding clefts. This improved fit creates strong polar and nonpolar contacts with the active site residues, effectively holding the derivatives within the binding site or in allosteric clefts.54 The successful candidates exhibited high positive solvation energy values, implying a desolvation penalty upon binding; however, this penalty was outweighed by the strong van der Waals interactions and coulombic energies. The performance of the top binders was attributed to their coulombic energy and favourable van der Waals forces, which are significantly more advantageous than those of sulfamethoxazole. This evidence suggests that the successful derivatives enhance the formation of stronger salt bridges, dipole–dipole interactions, or charge-assisted hydrogen bonds and van der Waals contact with the DHPS active site. It also implies that modifications improving electrostatic fit without compromising van der Waals contacts are likely to achieve better binding to the DHPS protein target.55 The free binding energies from the simulation further confirmed the strength of ligand binding to the target and their favorable interactions with DHPS, as evidenced by the docking studies.
Derivatives that exhibited superior MMGBSA scores compared to sulfamethoxazole were selected for molecular docking and MMGBSA calculations with the LasR and AcrB proteins. In the context of quorum-sensing inhibition, the top 24 DHPS-targeting derivatives were evaluated against Pseudomonas aeruginosa LasR. While sulfamethoxazole (SMX) demonstrated an MMGBSA score of −43.30 kcal mol−1, 17 derivatives displayed enhanced binding free energies (dBind), particularly SMX018 (−52.81 kcal mol−1) and SMX047 (−51.10 kcal mol−1). Docking analysis indicated competitive displacement of the autoinducer 3-oxo-C12-HSL through hydrogen bonding with residues including Thr115, Ser129, Thr75, Tyr56, Leu39, and Gly38. Additional interactions included π–π stacking with Tyr64, pi-sulfur interactions with Asp73, and hydrophobic interactions with Val76, Leu125, Ala127, Leu40, Leu36, and Ala50 as illustrated in Fig. 6a–d.
Moreover, these derivatives caused allosteric disruption by occupying the Ile52–Ala50 hydrophobic subpocket, which impaired dimerization and DNA binding, resembling the mechanism of phage protein Aqs1.56 The incorporation of halogen atoms in compounds such as SMX019 and SMX020 enhanced potency via halogen bonding (e.g., with Val76) and hydrophobic filling of subpockets. Structure–activity relationships suggested that optimal hydrophobic rings were essential for acyl pocket occupation, with increased rigidity leading to reduced entropic penalties and improved hydrogen bonding with the sulphonamide functional group.57 Biologically, these derivatives can disrupt the LasR-RhlR-PQS hierarchy, thereby inhibiting virulence factors such as elastase and pyocyanin.
For efflux pump inhibition, the derivatives were evaluated against E. coli AcrB. SMX exhibited a free binding energy of −34.17 kcal mol−1, while 7 derivatives showed improved affinity. The derivatives target the periplasmic vestibule, engaging Phe664 and Pro669 (Fig. 7a–d), which are critical for van der Waals interactions with native substrates such as deoxycholate, ciprofloxacin, and ethidium. Notably, deoxycholate binds in the same vestibule region and forms a hydrogen bond with Ser715, in addition to hydrophobic interactions with Phe664,58 supporting the biological relevance of this site. Novel interactions include hydrogen bonds with Arg717 (unused by ciprofloxacin/ethidium), creating a steric block at the vestibule entrance.1,4 Pi–sulfur bonds with Met575 and hydrogen bonds with Pro718, Gly720, Asn719, Ala665, Asn667, Met649, Thr648, Thr678, and Gln830 further enhance entropy to counter vestibule flexibility.58
Mechanistically, vestibule confinement arrests AcrB in the “loose” (L) state, as was observed in the deoxycholate-bound AcrB structure, preventing rotational transition to export-competent conformations (“tight”/“open” states).
Following the computational modeling, five derivatives (SMX001, SMX003, SMX015, SMX019, and SMX033) were prioritized for synthesis and biological evaluation. This selection was designed to rigorously validate our computational predictions and probe critical structure–activity relationships. Specifically, SMX001 and SMX003 tested the hypothesis that electron-withdrawing groups enhance binding affinity to DHPS. SMX015 was chosen to evaluate whether incorporating polar functionality enabled high-affinity dual engagement with both DHPS and LasR targets. To probe the predicted role of halogen bonding in LasR inhibition, SMX019 was selected. Finally, SMX033 served as a critical benchmark for ADME/toxicity, boasting a favorable predicted solubility and safety profile, in addition to its strong van der Waals contacts within AcrB's hydrophobic vestibule.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Et2O (diethyl ether) [60
Et2O (diethyl ether) [60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40], all corresponding sulfamethoxazole derivatives were obtained in 65–78% yields (Fig. 8 and Table 5).
40], all corresponding sulfamethoxazole derivatives were obtained in 65–78% yields (Fig. 8 and Table 5).
| Compound | –R | Molecular formula | Calculated molecular weight | Melting point (oC) | Yield (%) |
|---|---|---|---|---|---|
| SMX001 | o-NO2 | C10H9N3O5S | 283.03 | 116–118 | 78 |
| SMX003 | p-NO2 | C10H9N3O5S | 283.03 | 118–120 | 71 |
| SMX015 | p-COOCH3 | C11H12N2O3S | 296.06 | 126–129 | 75 |
| SMX019 | o-Br | C10H9BrN2O3S | 315.95 | 164–167 | 75 |
| SMX033 | p-CH3 | C11H12N2O3S | 252.06 | 125–127 | 65 |
The compounds were confirmed by their structures using 1H NMR, 13C NMR spectroscopy and HRMS. The 1H NMR spectrum of compound SMX033 showed a broad peak between 6.24 and 6.26 ppm, indicating the formation of the sulfonamide group. The doublet peaks observed downfield at 7.68 to 7.70 ppm correspond to the p-disubstituted sulfonyl aromatic portion. The HRMS analysis revealed the molecular ion peak calculated for [M + H]+ as 252.0569, while the observed peak was 253.0649. Based on the structural evaluation, compound SMX033 was designated as 4-methyl-N-(5-methylisoxazol-3-yl)benzenesulfonamide. The primary amino group of sulfamethoxazole was converted through synthesis to generate compounds with electron-withdrawing and donating groups on the benzene sulfonyl portion, which were synthesized in good to excellent yields (60–78%).
6.1.4.1 Minimum inhibitory concentration (MIC) and cytotoxicity. The MICs of the synthesized compounds against test organisms (Proteus mirabilis, Salmonella typhi, Escherichia coli, and Pseudomonas aeruginosa) were determined using the broth microdilution technique. Fig. 9 shows the MIC values of the compounds within the range of 15.625 to 125 µg mL−1. All the synthesized derivatives exhibited MIC values <100 µg mL−1 against the test organisms, except Pseudomonas aeruginosa, which showed MIC values >100 µg mL−1 for SMX001, SMX003, and SMX019. The cytotoxicity data on the compounds is shown in Table 6.
| Compound | –R | Molecular formula | aMIC (µg mL−1) ± SEM | bCC50 (µM) ± SEM | SIc |
|---|---|---|---|---|---|
| a MIC: minimum inhibitory concentration of compounds SMX015 and SMX033 demonstrating highest inhibitory activity against P. mirabilis, S. typhi, and E. coli (MIC: 15.625 µg mL−1).b CC50 = 50% cytotoxic concentration on Vero cells.c SI = selective index (CC50/MIC); ciprofloxacin positive control. aIC50 and bCC50 values represent triplicate determinations (three determinations from three different experiments). | |||||
| SMX001 | o-NO2 | C10H9N3O5S | 441.00 ± 0.050 | 112.22 ± 0.060 | 0.25 |
| SMX003 | p-NO2 | C10H9N3O5S | 441.00 ± 0.002 | 125 ± 0.060 | 0.28 |
| SMX015 | p-COOCH3 | C11H12N2O3S | 61.98 ± 0.060 | 186.00 ± 0.005 | 3.00 |
| SMX019 | o-Br | C10H9BrN2O3S | 395.63 ± 0.025 | 121.10 ± 0.060 | 0.30 |
| SMX033 | p-CH3 | C11H12N2O3S | 61.98 ± 0.020 | 286.20 ± 0.001 | 4.62 |
| Ciprofloxacin | — | C17H18FN3O3 | 3.91 ± 0.0001 | Not deterermined | — |
The in vitro antibiotic potential of the five (5) sulfamethoxazole derivatives was assessed individually using the high-throughput spotting assay method. The MIC of the five (5) synthesized compounds was determined alongside ciprofloxacin and DMSO as positive and negative controls, respectively. All compounds showed significant antibacterial activity against P. mirabilis, S. typhi, and E. coli (MICs 15.625–62.5 µg mL−1), although they were less potent than ciprofloxacin (3.91 µg mL−1). Compounds SMX015 and SMX033 demonstrated high inhibitory activity against P. mirabilis, S. typhi, and E. coli (MIC: 15.625 µg mL−1). Compounds SMX001, SMX003, and SMX019 recorded inhibitory activity against S. typhi (MIC: 15.625 µg mL−1). Additionally, compound SMX001 also recorded inhibitory activity (MIC: 15.625) against E. coli. Activity against the more resistant Pseudomonas aeruginosa was markedly reduced (MICs 62.5–125 µg mL−1) compared to ciprofloxacin (3.91 µg mL−1), indicating substantial intrinsic resistance. Compound SMX033 features a methyl group at the para position, whilst compounds SMX015 and SMX001 feature p-methyl ester and o-nitro groups, respectively. SMX033 demonstrated comparable or improved activity relative to sulfamethoxazole against most tested Gram-negative pathogens. Compound SMX033 consistently demonstrated significant antibacterial activity against all tested pathogens, suggesting its potential as a broad-spectrum agent. Pseudomonas aeruginosa, the most resistant pathogen, was susceptible to SMX033 and SMX019 at 62.5 µg mL−1 and SMX001, SMX003, and SMX015 MIC at 125 µg mL−1, indicating moderate inhibition. The observed MIC values (15.625–125 µg mL−1) against Gram-negative pathogens align with the computational ranking based on MMGBSA binding free energies. Notably, SMX033 (p-CH3) and SMX015 (p-COOCH3) consistently exhibited the lowest MICs (most potent activity), particularly against P. mirabilis, S. typhi, and E. coli (15.625 µg mL−1). Their strong computational scores directly support this strong inhibitory activity. SMX033 showed favourable MM-GBSA scores relative to the parent compound for each target individually. Its high affinity, predicted to stem from optimal van der Waals contacts in the hydrophobic vestibules of AcrB and LasR, translates to effective bacterial growth inhibition by simultaneously disrupting folate synthesis, quorum sensing, and efflux. The potent activity of SMX015 aligns with its in silico profile, where it ranked among the top compounds for affinity against DHPS, LasR and AcrB in separate MM-GBSA calculations. Its polar ester group was predicted to facilitate dual engagement, which is consistent with its potent and broad-spectrum MIC results.59 The reduced activity against P. aeruginosa (MICs: 62.5–125 µg mL−1) for all compounds was anticipated. This pathogen possesses a notoriously impermeable outer membrane and highly efficient efflux systems, which our in silico models, focused on target binding, would not fully account for. P. aeruginosa is highly resistant and could require different derivatives or scaffolds.60 However, the fact that SMX033 and SMX015 still showed the best activity (MIC = 62.5 µg mL−1) against this resilient pathogen underscores the strength of the multi-target approach, as overcoming its defence likely requires potent inhibition of multiple resistance pathways simultaneously.61 Compound SMX033 is arguably the most potent when compared with ciprofloxacin because the substitution of the methyl in the phenyl ring present at the para position of the isoxazole nucleus enhanced the antimicrobial activity and decreased the cytotoxic activity of compound SMX033. Compound SMX033 exhibited reduced cytotoxicity against Vero cell lines, with a CCC50 value of 286.20 µM suggesting its high selectivity towards the pathogens.
6.1.4.2 Biofilm formation inhibition. Biofilms are structured communities of microorganisms embedded in an extracellular matrix, offering protection against antimicrobials.62 The ability to inhibit biofilm formation is critical for treating chronic infections, as biofilms are notoriously resistant to conventional antibiotics. By preventing biofilm development, the compounds could reduce microbial persistence and improve treatment outcomes. Biofilm inhibition might result from interference with adhesion, quorum sensing, or EPS synthesis.62–64 Biofilm formation inhibition effects of the compounds SMX033, SMX019, SMX003, SMX015, and SMX001 at sub-MIC concentrations against Proteus mirabilis, Salmonella typhi, Escherichia coli, and Pseudomonas aeruginosa were analysed and represented in Fig. 10a and b. The antibiotic's subminimal inhibitory concentration (sub-MIC) can influence the biofilm formation capacity of bacterial pathogens, thereby impacting the pathogenesis and infection outcomes.65 The compounds exhibited excellent biofilm inhibition. All derivatives, particularly SMX033, SMX015, and SMX001, achieved exceptional inhibition (>98.5%) against the tested organisms, significantly outperforming the parent drug ciprofloxacin (94.35) (Fig. 10a and b). This result directly correlates with the molecular docking predictions. Our models indicated that these derivatives competitively displace the native autoinducer (OHN) in the LasR binding pocket by forming key hydrogen bonds with residues Thr75, Tyr56, and Ser129. Furthermore, they were predicted to cause allosteric disruption by occupying the critical Ile52–Ala50 hydrophobic subpocket, impairing LasR dimerization and DNA binding, a mechanism known to inhibit biofilm formation profoundly.66 The superior experimental biofilm inhibition of SMX019 (o-Br) against E. coli aligns with its in silico prediction of enhanced potency through halogen bonding (e.g., with Val76 in LasR). The strong experimental biofilm inhibition across the board confirms the computational prediction that targeting LasR is a viable and effective strategy for these sulfamethoxazole derivatives.
Absorbance reflects remaining biofilm biomass, where lower values indicate greater inhibition.67 Each value is the average of replicate readings for three determinations with their standard deviation (SD). The negative control, which contained no antimicrobial treatment, served as a baseline for maximum biofilm formation in the organism.
6.1.4.3 Ethidium bromide efflux pump inhibition assay. Efflux pumps are membrane proteins found in bacteria that actively expel toxic substances including antibiotics out of the pathogen cell.68 Some of these efflux pumps are non-specific, removing a wide range of drug molecules, while others are selective. Efflux pump inhibitors are compounds that block the activity of these pumps, thereby increasing the intracellular concentration of antibiotics and restoring their effectiveness.69 The relevance of efflux pump inhibition still remains intact as it aids in combating antibiotic resistance. Overexpression of efflux pumps is a common mechanism of multidrug resistance, especially in pathogens like Pseudomonas aeruginosa, Escherichia coli, and Acinetobacter baumannii.70 Efflux pump inhibitors (EPIs) enhance drug potency by retaining antibiotics inside bacteria cells, thereby lowering the required dose of antibiotic and reducing side effects. Combining an antibiotic (even older and less tolerated drugs) with EPIs can offer a synergistic effect.71,72 Ethidium bromide was utilized in the evaluation of the efflux pump inhibition potential of the compounds. Because it may intercalate DNA, which breaks down DNA structure and causes cell death, the dye ethidium bromide (EB) possesses antibiotic activity.73 The efflux pump is the sole defense mechanism that bacteria use to fend off the effects of ethidium bromide.74 This method has been used in many studies to investigate the presence or absence of efflux pumps in combination with traditional efflux pump inhibitors.75,76 To evaluate the compound's effectiveness, the efflux pump inhibition experiment used ethidium bromide (EtBr) as a substrate. The premise of this experiment is that EtBr has strong intracellular fluorescence. As it keeps building up, it binds to DNA and creates a growing fluorescence. When they build up inside the bacterial cells, the fluorescence intensity rises, indicating that the efflux pump mechanism is being inhibited. A low fluorescence measurement occurs when the cell pumps out EtBr in the absence of efflux pump inhibitory action. The fluorescence intensity was recorded at an interval of everyone (1) minute for a total period of sixty (60)-minutes (Fig. 11a–d). Compound SMX033 (CH3-para) demonstrated a remarkable inhibition of the efflux pump and was the strongest EPI overall.
SMX033 was predicted to bind strongly in the periplasmic vestibule of AcrB, interacting with key residues such as Phe664 and Pro669 through significant van der Waals and hydrophobic interactions. The methyl group at the para position was computationally identified as essential for occupying the hydrophobic pocket formed by Phe664, a residue crucial for substrate binding and expulsion. By obstructing this vestibule, SMX033 is expected to stabilize AcrB in the “loose” (L) state, thereby preventing the conformational changes necessary for antibiotic export. The experimental observation of high ethidium bromide (EtBr) accumulation serves as direct functional evidence of this mechanism.77 Additionally, the moderate to satisfactory efflux pump inhibitor (EPI) activity of SMX015, SMX019, SMX003, and SMX001 correlates with their in silico profiles. Their diverse substituents (ester, bromo, and nitro) were anticipated to interact with the vestibule through different mechanisms, such as forming hydrogen bonds with Arg717 or pi–sulfur interactions with Met575, which explains their measurable, albeit lower, efflux inhibition compared to SMX033.78,79
7 Molecular dynamics simulation studies
To complement the molecular docking analyses and provide a dynamic validation of the binding stability and conformational effects induced by the prioritized sulfamethoxazole derivatives, molecular dynamics (MD) simulations of 100 ns duration were performed. The top two candidates, SMX033 and SMX015, were simulated in complex with the primary enzyme target dihydropteroate synthase (DHPS) and the quorum-sensing regulator LasR. Their trajectories were compared to those of the corresponding apo proteins. Analyses included root-mean-square deviation (RMSD) for complex stability, root-mean-square fluctuation (RMSF) for residue-specific flexibility, radius of gyration (Rg) and solvent-accessible surface area (SASA) for global compactness, and principal component analysis (PCA) for essential collective motions. These detailed simulations provide a rigorous, time-evolved perspective on ligand-induced stabilization, directly supporting the static binding poses obtained from docking.7.1 Stability and dynamics of DHPS complexes
The stability of the DHPS-ligand complexes was quantitatively assessed by monitoring the backbone RMSD relative to the initial minimized structure over the entire simulation trajectory (Fig. 12A and B). The RMSD for the SMX033-DHPS complex reached equilibrium rapidly, maintaining a low and stable profile with a mean value of 0.129 ± 0.020 nm. Similarly, the SMX015-DHPS complex exhibited a mean RMSD of 0.128 ± 0.030 nm. In contrast, the apo DHPS simulation displayed significantly higher and more variable instability, with a mean RMSD of 0.316 ± 0.070 nm. The substantially lower mean values and reduced standard deviations of the ligand-bound systems demonstrate that the binding of either SMX033 or SMX015 effectively constrains the protein backbone, inducing a more rigid and conformationally stable state compared to the apo protein. To elucidate localized effects, the per-residue flexibility was calculated via RMSF (Fig. 12C and D). The global backbone flexibility, expressed as the mean RMSF across all residues, was reduced in the presence of SMX033 (0.105 ± 0.069 nm) and SMX015 (0.111 ± 0.086 nm) relative to the apo protein (0.132 ± 0.107 nm). A critical observation was the pronounced dampening of fluctuations in key active site residues responsible for substrate binding and inhibitor interactions. For example, in the apo simulation, residues Phe28, Arg63, and Pro64 exhibited high fluctuations of 0.434 nm, 0.427 nm, and 0.556 nm, respectively. Upon binding SMX033, these fluctuations were reduced to 0.219 nm, 0.263 nm, and 0.366 nm. A comparable stabilizing trend was observed for the SMX015 complex. Furthermore, residues involved in critical hydrogen-bonding networks with the sulfonamide core and substituents, such as Thr62, Lys221, and Ser222, consistently demonstrated lower mobility in the bound states. This ligand-induced reduction in flexibility across the active site pocket indicates a stable occupation that restricts inherent dynamical motions, likely contributing to sustained enzyme inhibition.The global structural analysis of ligand binding were evaluated through the radius of gyration (Rg) and solvent-accessible surface area (SASA) (Fig. 13). The mean Rg for DHPS decreased from 1.827 ± 0.019 nm in the apo state to 1.811 ± 0.016 nm with SMX015 and 1.824 ± 0.017 nm with SMX033. In addition, the mean SASA decreased from 138.2 ± 4.1 nm2 (apo) to 134.2 ± 3.8 nm2 (SMX015) and 135.4 ± 3.5 nm2 (SMX033). The paired reduction in both Rg and SASA signifies a ligand-mediated contraction of the protein structure. This compaction is consistent with the stabilization of binding site loops and the burial of hydrophobic surfaces upon optimal ligand engagement, leading to a more globular and less solvent-exposed conformation.
To capture the large-scale, functionally relevant motions of DHPS, principal component analysis (PCA) was performed on the Cα atom trajectories (Fig. 14). The total variance captured by the first two principal components (PC1 and PC2) was substantially lower for the ligand-bound states: 7.36 ± 2.11 for SMX015 and 5.18 ± 1.64 for SMX033, compared to 12.44 ± 3.87 for the apo protein. This marked reduction in collective motion variance indicates that binding of either derivative confines the protein's conformational sampling to a significantly more restricted subspace. The conformational space area, calculated from the projection of trajectories onto PC1 and PC2, was reduced by approximately 33% and 44% for the SMX015 and SMX033 complexes, respectively.
 | ||
| Fig. 14 SMX015-and SMX033-induced conformational landscapes of DHPS revealed by principal component analysis. (A) apo protein; (B) SMX015; (C) SMX033. | ||
This analysis confirms that both ligands not only stabilize local binding site residues but also effectively dampen the global, low-frequency motions of the entire DHPS enzyme, effectively locking it into a stable conformation that is less prone to unproductive dynamics.
7.2 Stability and dynamics of LasR complexes
MD simulations of the LasR complexes revealed distinct dynamic profiles for the two ligands, correlating with their roles as quorum-sensing inhibitors. The backbone RMSD for the LasR-SMX033 complex averaged 0.306 ± 0.062 nm, which was lower than both the apo protein (0.331 ± 0.095 nm) and the SMX015 complex (0.338 ± 0.081 nm) (Fig. 15A and B). This suggests SMX033 forms a marginally more stable complex with LasR. RMSF analysis demonstrated that both ligands reduced the overall flexibility of LasR (mean RMSF: apo = 0.202 ± 0.149 nm; SMX015 = 0.164 ± 0.067 nm; SMX033 = 0.157 ± 0.062 nm). Notably, SMX033 binding specifically reduced fluctuations in key hydrophobic cavity residues vital for autoinducer binding and protein dimerization (Fig. 15C and D). For instance, fluctuations of Leu36, Ile52, and Tyr64 were reduced to 0.089 nm, 0.111 nm, and 0.153 nm, respectively, from their higher apo values. While SMX015 also exhibited a stabilizing effect, the magnitude of fluctuation reduction in these specific residues was less pronounced. This residue-specific stabilization supports the docking prediction that SMX033 occupies and rigidifies the native ligand-binding site, potentially disrupting the conformational changes required for functional dimerization.Analysis of the global parameters Rg and SASA indicated distinct ligand-dependent effects on the tertiary structure of LasR. The SMX015 complex showed a slight decrease in mean Rg (1.912 ± 0.032 nm) and SASA (131.4 ± 3.0 nm2) relative to apo-LasR (1.914 ± 0.036 nm; 133.0 ± 3.6 nm2), indicating a mild compacting effect (Fig. 16). In contrast, the LasR-SMX033 complex displayed a significant increase in both mean Rg (2.032 ± 0.025 nm) and SASA (137.4 ± 2.7 nm2). This ligand-induced expansion suggests that SMX033 binding triggers a distinct allosteric response, potentially causing structural rearrangements that propagate beyond the binding pocket. This expansion may be linked to the disruption of the dimerization interface, providing a dynamic rationale for its potent anti-biofilm activity via quorum-sensing inhibition.
PCA of the LasR trajectories further differentiated the ligands' effects. The total variance for the SMX033-bound system (8.38 ± 2.24) was dramatically lower than for the apo protein (24.07 ± 5.87) and the SMX015-bound system (11.39 ± 3.12) (Fig. 17). The profound reduction in collective motion for the SMX033 complex indicates it imposes a more restriction on the intrinsic flexibility and essential dynamics of LasR. Although SMX015 also reduced the conformational space area, its effect was less extensive. The capability of SMX033 to significantly confine LasR's large-scale motions likely highlights its efficacy in biofilm inhibition, as it may more effectively prevent the conformational transitions necessary for LasR activation, DNA binding, and subsequent virulence gene expression.
The MD simulation analysis provide a robust, atomistic rationale for the experimentally observed biological efficacy of SMX033 and SMX015. For DHPS, the pronounced stabilization of the active site architecture, suppression of key residue flexibility, and damping of global motions by both ligands correlate directly with their potent growth inhibitory activity (MIC = 15.625 µg mL−1). This dynamic stabilization affirms their role as direct, high-affinity enzyme inhibitors. The superior biofilm inhibition (>99%) exhibited by both compounds, and particularly by SMX033, is dynamically supported by their stable binding and modulation of LasR. The unique dynamic signature of SMX033 characterized by strong binding pocket stabilization, induction of an expanded protein state, and marked restriction of collective motions offers a compelling mechanism for its function as a quorum-sensing inhibitor. This aligns with the experimental observation of near-complete biofilm eradication. Consistent favorable dynamic behavior of SMX033 across both targets supports its prioritization as the lead compound.
8 Conclusion
This study has successfully demonstrated a rational, integrated computational and experimental approach to design novel sulfamethoxazole derivatives that have potential to simultaneously target bacterial viability, virulence, and resistance mechanisms. Our structure-guided design yielded a focused library of derivatives, from which five candidates were prioritized based on superior predicted binding affinities towards DHPS, the quorum-sensing regulator LasR, and the efflux pump transporter AcrB. The synthesis of these compounds was achieved efficiently, and their structures were unequivocally confirmed via NMR and HRMS. Biological evaluation revealed potent, broad-spectrum antimicrobial activity (MICs 15.625–125 µg mL−1) against key ESKAPE pathogens, with lead compounds SMX033 and SMX015 exhibiting exceptional potency. Crucially, the experimental data strongly correlated with our in silico predictions. The derivatives achieved near-complete biofilm inhibition (>99%) by disrupting LasR-mediated quorum sensing, and SMX033 emerged as a potent efflux pump inhibitor, directly corroborating our AcrB binding models. Furthermore, the markedly low cytotoxicity of SMX033 (CC50 = 286.20 µM) underscores its promising selective toxicity and potential as a therapeutic scaffold. This work provides compelling proof-of-concept for a multi-targeting strategy to overcome multidrug resistance. By inhibiting DHPS, LasR, and AcrB concurrently, these sulfamethoxazole derivatives can attack the pathogen on multiple fronts, thereby mitigating the evolutionary pressure that leads to resistance. The excellent correlation between our computational predictions and experimental results establishes a validated blueprint for the potential rational design of next-generation antimicrobial agents. Future work will focus on in vivo efficacy studies, further medicinal chemistry optimization to improve potency and aqueous solubility and expanding this strategy to a broader chemical space to combat the ever-growing threat of pan-drug-resistant infections.Author contributions
Cedric D. K. Amengor: supervised the project, developed the overall conceptual framework, oversaw the synthesis, wrote the original draft, and proofread the manuscript. Benedicta Yayra Adzah carried out the synthesis of the compounds. Prince Danan Biniyam: designed the computational conceptual framework and methodology, authored the molecular docking and MMGBSA, and proofread the manuscript. Ernest Oyeh, Vuvor Faustina and Yvonne Dogbeda Agboyibor assisted in the characterisation of the compounds. Victoria O. Adu: provided computational technical support for ADME analysis, assisted in data curation, and proofread the manuscript. Michael Osei: provided computational technical support for molecular docking, assisted in data curation, and proofread the manuscript. Kwabena Adu-Adjei: provided computational technical support for ADME analysis, assisted in data curation, and proofread the manuscript. Patrick Gyan, En Zhang, Ampomah-Wireko: contributed to the writing of the antitubercular section and proofread the manuscript. Paul Quansah and Cyril Makafui Amengor: provided mathematical and statistical support for antimicrobial data analysis. Joshua Boateng: interpreted the HRMS data and edited the first draft of the manuscript. Victoria Elmes: carried out the NMR analysis and provided technical expertise on the NMR instrumentation Iain Goodall: carried out the HRMS analysis and provided technical expertise on the mass spectrometer.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The original contributions and data obtained in the study are presented in the article.Supplementary information (SI): description of the spectroscopic data of the compounds. See DOI: https://doi.org/10.1039/d5ra08024j.
Acknowledgements
The authors would like to express their sincere gratitude to the technicians of the Synthetic Medicinal Chemistry and Computational Division at the School of Pharmacy, University of Health and Allied Sciences, for their invaluable support.References
- C. Xu, L. Kong, H. Gao, X. Cheng and X. Wang, Front. Microbiol., 2022, 13, 822689 CrossRef PubMed.
- K. P. High, Ageing Res. Rev., 2004, 3, 1–14 CrossRef PubMed.
- M. A. Salam, M. Y. Al-Amin, M. T. Salam, J. S. Pawar, N. Akhter, A. A. Rabaan and M. A. Alqumber, Healthcare, 2023, 11, 1946 CrossRef PubMed.
- A. P. Taylor, R. P. Robinson, Y. M. Fobian, D. C. Blakemore, L. H. Jones and O. Fadeyi, Org. Biomol. Chem., 2016, 14, 6611–6637 RSC.
- J. Jampilek, Molecules, 2019, 24, 3839 CrossRef CAS PubMed.
- G. Shabir, A. Saeed, W. Zahid, F. Naseer, Z. Riaz, N. Khalil and F. Albericio, Pharmaceuticals, 2023, 16, 1162 CrossRef CAS PubMed.
- S. Kumari, A. V. Carmona, A. K. Tiwari and P. C. Trippier, J. Med. Chem., 2020, 63, 12290–12358 CrossRef CAS PubMed.
- G. J. Martis and S. L. Gaonkar, RSC Adv., 2025, 15, 8213–8243 RSC.
- S. Li, Y. Mei, L. Jiang, X. Yang, W. Zeng and Y. Du, RSC Med. Chem., 2025 10.1039/D4MD00777H.
- S. Ramlakhan, V. Singh, J. Stone and A. Ramtahal, Clin. Med. Insights Pediatr., 2014, 8, CMPed–S8100 CrossRef PubMed.
- H. Sørum, in Antimicrobial Resistance in Bacteria of Animal Origin, ed. F. M. Aarestrup, ASM Press, 2005, pp. 213–238 Search PubMed.
- B. Jubeh, Z. Breijyeh and R. Karaman, Molecules, 2020, 25, 2888 CrossRef CAS PubMed.
- M. Pervaiz, A. Riaz, A. Munir, Z. Saeed, S. Hussain, A. Rashid and A. Adnan, J. Mol. Struct., 2020, 1202, 127284 CrossRef CAS.
- M. Claudel, J. V. Schwarte and K. M. Fromm, Chemistry, 2020, 2, 849–899 Search PubMed.
- S. Wang, H. Y. Zhang, L. Wang, Z. J. Duan and I. Kennedy, Food Addit. Contam., 2006, 23, 362–384 Search PubMed.
- S. Ballav, D. Jana, K. C. Guchhait, B. Das, M. Manna, S. Hazra and C. Ghosh, in Biofilm Associated Antimicrobial Resistance and its Recovery, CRC Press, 2023, pp. 66–81 Search PubMed.
- R. Alenazy, J. King Saud Univ. Sci., 2022, 34, 102275 CrossRef.
- V. C. Scoffone, G. Trespidi, G. Barbieri, S. Irudal, E. Perrin and S. Buroni, Antibiotics, 2021, 10, 863 CrossRef CAS PubMed.
- L. M. A. Ghany, N. Ryad, M. S. Abdel-Aziz, H. M. Abd El-Lateef, I. Zaki and B. Y. Beshay, J. Mol. Struct., 2024, 1309, 138170 CrossRef CAS.
- F. Giordanetto, P. W. Fowler, M. Saqi and P. V. Coveney, Philos. Trans. R. Soc. London, A, 2005, 363, 2055–2073 CrossRef CAS PubMed.
- C. A. Salubi, Molecular Docking Investigation and Pharmacokinetic Properties Prediction of Some Benzimidazole Analogues as Dihydropteroate Synthase (DHPS) Inhibitors, 2023 Search PubMed.
- Z. Breijyeh and R. Karaman, Antibiotics, 2023, 12, 628 CrossRef CAS PubMed.
- T. Hanai, Front. Med. Chem., 2009, 4, 86–129 Search PubMed.
- H. Azevedo-Barbosa, B. P. do Vale, G. Guidolin Rossi, F. dos Santos Siqueira, K. Bordignon Guterres, M. M. A. de Campos and D. Teixeira Carvalho, Chem. Biodiversity, 2021, 18, e2100066 CrossRef CAS PubMed.
- D. Das, N. Sahu, S. Roy, P. Dutta, S. Mondal, E. L. Torres and C. Sinha, Spectrochim. Acta, Part A, 2015, 137, 560–568 Search PubMed.
- J. P. Fetse, J. O. Kyekyeku, E. Dueve and K. B. Mensah, Br. J. Pharm. Res., 2014, 4, 2642 Search PubMed.
- C. Agyare, A. S. Dwobeng, N. Agyepong, Y. D. Boakye, K. B. Mensah, P. G. Ayande and M. Adarkwa-Yiadom, Adv. Pharmacol. Pharm. Sci., 2013, 2013, 692613 Search PubMed.
- M. Ofori, C. A. Danquah, S. Ativui, P. Doe and W. A. Asamoah, Biomed. Pharmacol. J., 2021, 14(4), 1905–1915 CAS.
- C. A. Danquah, P. A. B. Minkah, T. A. Agana, P. Moyo, M. Tetteh, I. O. D. Junior and V. J. Maharaj, et al., Natural Products as Antibiofilm Agents, Focus on bacterial biofilms, 2022, p. 203 Search PubMed.
- D. Anokwah, E. Asante-Kwatia, J. Asante, D. Obeng-Mensah, C. A. Danquah, I. K. Amponsah and A. Y. Mensah, Microorganisms, 2023, 12, 7 Search PubMed.
- R. Mowla, Y. Wang, S. Ma and H. Venter, Biochim. Biophys. Acta, Biomembr., 2018, 1860, 878–886 CrossRef CAS PubMed.
- K. Azeem, S. Fatima, A. Ali, A. Ubaid, F. M. Husain and M. Abid, Life, 2025, 15, 49 Search PubMed.
- X. He, S. Liu, T. S. Lee, B. Ji, V. H. Man, D. M. York and J. Wang, ACS Omega, 2020, 5, 4611–4619 Search PubMed.
- P. Bhadra and S. W. Siu, Langmuir, 2019, 35, 9622–9633 Search PubMed.
- H. Grabski, L. Hunanyan, S. Tiratsuyan and H. Vardapetyan, F1000Research, 2019, 8, 324 Search PubMed.
- N. Singh and A. Warshel, Proteins: Struct., Funct., Bioinf., 2010, 78, 1705–1723 Search PubMed.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 Search PubMed.
- S. Jo, X. Cheng, J. Lee, S. Kim, S. J. Park, D. S. Patel and W. Im, J. Comput. Chem., 2017, 38, 1114–1124 Search PubMed.
- R. Shukla and T. Tripathi, in Computer-aided Drug Design, Springer, Singapore, 2020, pp. 133–161 Search PubMed.
- K. Vanommeslaeghe, E. Hatcher, C. Acharya, S. Kundu, S. Zhong, J. Shim and A. D. Mackerell Jr, J. Comput. Chem., 2010, 31, 671–690 CrossRef CAS PubMed.
- G. J. Martis and S. L. Gaonkar, RSC Adv., 2025, 15, 8213–8243 Search PubMed.
- M. A. Subbaiah and N. A. Meanwell, J. Med. Chem., 2021, 64, 14046–14128 Search PubMed.
- E. Marsault and M. L. Peterson, J. Med. Chem., 2011, 54, 1961–2004 CrossRef CAS PubMed.
- K. J. Coe, PhD Thesis, University of Washington, 2008 Search PubMed.
- C. Saran and K. L. Brouwer, Toxicol. Pathol., 2023, 51, 405–413 CrossRef CAS PubMed.
- Z. Wang, H. Sun, X. Yao, D. Li, L. Xu, Y. Li and T. Hou, Phys. Chem. Chem. Phys., 2016, 18, 12964–12975 Search PubMed.
- R. Jakhar, M. Dangi, A. Khichi and A. K. Chhillar, Curr. Bioinf., 2020, 15, 270–278 Search PubMed.
- A. H. Bakheit, M. W. Attwa, A. A. Kadi, H. A. Ghabbour and H. M. Alkahtani, Crystals, 2023, 13, 1020 Search PubMed.
- J. W. M. Nissink and S. Degorce, Future Med. Chem., 2013, 5, 753–767 Search PubMed.
- C. A. Salubi, Molecular Docking Investigation and Pharmacokinetic Properties Prediction of Some Benzimidazole Analogues as Dihydropteroate Synthase (DHPS) Inhibitors, 2023 Search PubMed.
- C. R. Bourne, Antibiotics, 2014, 3, 1–28 CrossRef PubMed.
- M. N. Nalam, A. Ali, M. D. Altman, G. S. Reddy, S. Chellappan, V. Kairys, A. Ozen, H. Cao, M. K. Gilson, B. Tidor, T. M. Rana and C. A. Schiffer, J. Virol., 2010, 84, 5368–5378 Search PubMed.
- K. A. Ayers, PhD Thesis, The University of Tennessee Health Science Center, 2009 Search PubMed.
- S. Genheden and U. Ryde, Expert Opin. Drug Discovery, 2015, 10, 449–461 Search PubMed.
- M. Zürcher and F. Diederich, J. Org. Chem., 2008, 73, 4345–4361 Search PubMed.
- H. X. Zhou and X. Pang, Chem. Rev., 2018, 118, 1691–1741 CrossRef CAS PubMed.
- M. Shah, V. L. Taylor, D. Bona, Y. Tsao, S. Y. Stanley, S. M. Pimentel-Elardo and K. L. Maxwell, Mol. Cell, 2021, 81, 571–583 CrossRef CAS PubMed.
- P. Di Fruscia, A. Carbone, G. Bottegoni, F. Berti, F. Giacomina, S. Ponzano and F. Bertozzi, J. Med. Chem., 2021, 64, 13327–13355 Search PubMed.
- D. Drew, M. M. Klepsch, S. Newstead, R. Flaig, J. W. De Gier, S. Iwata and K. Beis, Mol. Membr. Biol., 2008, 25, 677–682 Search PubMed.
- L. Bibens, J. P. Becker, A. Dassonville-Klimpt and P. Sonnet, Pharmaceuticals, 2023, 16, 425 Search PubMed.
- J. Mehla, G. Malloci, R. Mansbach, C. A. López, R. Tsivkovski, K. Haynes and H. I. Zgurskaya, mBio, 2021, 12, 10–1128 CrossRef PubMed.
- M. Baym, L. K. Stone and R. Kishony, Science, 2016, 351, aad3292 CrossRef PubMed.
- H. C. Flemming and J. Wingender, Nat. Rev. Microbiol., 2010, 8, 623–633 CrossRef CAS PubMed.
- N. Rabin, Y. Zheng, C. Opoku-Temeng, Y. Du, E. Bonsu and H. O. Sintim, Future Med. Chem., 2015, 7, 493–512 Search PubMed.
- S. Singh, S. Datta, K. B. Narayanan and K. N. Rajnish, J. Genet. Eng. Biotechnol., 2021, 19, 140 CrossRef PubMed.
- R. Elawady, A. G. Aboulela, A. Gaballah, A. A. Ghazal and A. N. Amer, BMC Infect. Dis., 2024, 24, 1065 Search PubMed.
- X. Wang, M. Liu, C. Yu, J. Li and X. Zhou, Mol. Biomed., 2023, 4, 49 Search PubMed.
- M. E. Skogman, P. M. Vuorela and A. Fallarero, J. Antibiot., 2012, 65, 453–459 Search PubMed.
- A. Gaurav, P. Bakht, M. Saini, S. Pandey and R. Pathania, Microbiology, 2023, 169, 001333 Search PubMed.
- M. AlMatar, O. Albarri, E. A. Makky and F. Köksal, Pharmacol. Rep., 2021, 73, 1–16 Search PubMed.
- L. Huang, C. Wu, H. Gao, C. Xu, M. Dai, L. Huang and G. Cheng, Antibiotics, 2022, 11, 520 CrossRef CAS PubMed.
- A. Lamut, L. Peterlin Mašič, D. Kikelj and T. Tomašič, Med. Res. Rev., 2019, 39, 2460–2504 CrossRef CAS PubMed.
- L. Zhang, X. Tian, L. Sun, K. Mi, R. Wang, F. Gong and L. Huang, Pharmaceutics, 2024, 16, 170 CrossRef CAS PubMed.
- S. Nafisi, A. A. Saboury, N. Keramat, J. F. Neault and H. A. Tajmir-Riahi, J. Mol. Struct., 2007, 827, 35–43 Search PubMed.
- S. Pal, A. Misra, S. Banerjee and B. Dam, J. King Saud Univ. Sci., 2020, 32, 939–945 CrossRef.
- P. W. Limaverde, F. F. Campina, F. A. da Cunha, F. D. Crispim, F. G. Figueredo, L. F. Lima and S. R. Tintino, Food Chem. Toxicol., 2017, 109, 957–961 Search PubMed.
- S. R. Tintino, C. D. Oliveira-Tintino, F. F. Campina, P. Weslley Limaverde, P. S. Pereira, J. P. Siqueira-Junior and V. Q. Balbino, Med. Chem. Res., 2018, 27, 261–267 Search PubMed.
- J. Olmsted III and D. R. Kearns, Biochemistry, 1977, 16, 3647–3654 Search PubMed.
- D. Olender, J. Żwawiak and L. Zaprutko, Pharmaceuticals, 2018, 11, 54 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |