 Open Access Article
Open Access ArticleBiocatalytic treatment of cresols in aqueous solution with soybean peroxidase
Mohammadreza Haghighatnama a,
Samira Narimannejad
a,
Samira Narimannejad a,
Nihar Biswas
a,
Nihar Biswas a and
Keith E. Taylor
a and
Keith E. Taylor *b
*b
aDepartment of Civil and Environmental Engineering, University of Windsor, Windsor, ON N9B 3P4, Canada
bDepartment of Chemistry and Biochemistry, University of Windsor, Windsor, ON N9B 3P4, Canada. E-mail: taylor@uwindsor.ca
First published on 22nd January 2026
Abstract
Soybean peroxidase (SBP) from the seed coat is instrumental in catalyzing the oxidation of various aqueous aryl-amino and -hydroxyl compounds in the presence of hydrogen peroxide. This capability positions SBP as a promising agent for both industrial and domestic wastewater applications. To assess SBP's effectiveness in removing o-, m-, and p-cresol from water, experiments were conducted using a stirred batch reactor. The goal was to determine the optimal conditions for achieving at least 95% removal of these compounds. Key parameters examined included pH, SBP activity, and hydrogen peroxide concentration for 1.0 mM cresol. The experiments confirmed SBP's efficiency in eliminating the cresols from water. High-performance liquid chromatography identified the optimal conditions for removal of the isomers: for o-cresol, removal was achieved with an SBP activity of 0.7 U mL−1, pH 9.0, and hydrogen peroxide concentration of 1.2 mM; for m-cresol, SBP 0.8 U mL−1, pH 8.0, and hydrogen peroxide 1.1 mM; for p-cresol SBP 0.3 U mL−1, pH 7.0, and hydrogen peroxide 1.0 mM. Half-lives under these conditions were 2–9 min. Mass spectrometry confirmed the formation of product oligomers up to the decamer stage, and residual UV analysis was consistent with the oligomerization–precipitation pathway. These findings highlight SBP's potential to significantly reduce o-, m-, and p-cresol levels in water, demonstrating its utility in environmental remediation efforts.
Introduction
Cresols, a group of three methylphenol isomers, are priority pollutants with a host of toxicological effects, including classification as possible human carcinogens.1 They are introduced into the environment through industrial processes, especially as by-products of coal gasification and production of fragrances, resins, antioxidants, dyes, and insecticides.2,3 Representative regulatory thresholds include 0.1 µg kg−1 per day (US CDC) and 0.05 µg kg−1 per day (US EPA) for human exposure,1 4 µg L−1 (Canadian freshwater standard as mono- and dihyrdric-phenols) for aquatic life,3 200 mg L−1 (US Federal limit) for solid waste leachate, 50 µg L−1 (New Jersey water quality standard) for groundwater.4 Occurrence in Canada and the US of cresols in surface water was generally below 30 µg L−1 with hotspots into the 100 s of µg L−1, whereas in ground water, means were below 150 µg L−1 with hotspots in the 1000 s of µg L−1; leachates had a median of 112 µg L−1 with a maximum around 700 µg L−1; sewage treatment plant influents ranged into the 100 s of µg L−1, while effluents were below 30 µg L−1.2 The potential for human exposure to cresols is well summarized by the CDC from the US Toxics Release Inventory data showing onsite generation of cresol wastes around a million pounds.1 Conventional biological and physicochemical treatment methods exist but drawbacks have been noted, particularly sludge generation, energy consumption, and slowness.5–7 Enzymatic treatment, an advanced oxidation process, has proven to be particularly effective for removing phenolic compounds under mild conditions, with enzymes such as tyrosinases, laccases, and peroxidases demonstrating high efficiency.8–10 Biocatalytic remediation using peroxidases for phenolics, anilines, dyes, and other aromatic compounds in wastewater has been a particular focus11,12 and in this regard, cresols have been frequently studied,12,13 including with soybean peroxidase (SBP).14–16 Our preliminary studies showed SBP to be highly effective in treating o-, m-, and p-cresols in water.14,17 Its exceptional thermal and kinetic stability,18 ease of extraction, and availability from a byproduct (the seed coat, or hull) in the processing of a huge agricultural commodity make it an ideal choice for scale-up to industrial application19 compared to other commonly studied peroxidases such as horseradish peroxidase (HRP) and lignin peroxidase (LiP). At present, economic advantage falls to SBP since it may be extracted from the hulls without compromising their residual value as a feed additive.20 In contrast, HRP derives from an agricultural niche crop and its extraction would leave a toxic pulp to be disposed of, while LiP production by fermentation would leave the biomass to be disposed of. This relative ranking could change in future if heterologous expression systems, reported recently for HRP21 and LiP,22 substantially reduce the cost of enzyme. For the present, SBP is the peroxidase of choice for water treatment. A pro-forma cost analysis for SBP is given in SI to indicate its potential for low-cost production. Notably, crude enzyme preparations often outperform purified forms, and the main cost would lie in concentrating the extract, if necessary, rather than in purification,20,23,24 recently reviewed by Feng et al.25,26The peroxidase mechanism consists of two-electron oxidation of the enzyme heme group to a reactive intermediate, Compound I, followed by two successive one-electron oxidations of phenolic substrate molecules (via reactive enzyme intermediate Compound II), generating a phenoxyl free radical at each step and returning the enzyme to its resting state.27 These radicals couple to form oxidative dimers, which then oligomerize through repeated peroxidase cycles until they reach their solubility limit and precipitate.15,27 From the perspective of fit into the peroxidase active site, Uyama argues for the involvement of radical transfer in elaboration of oligomers beyond the dimer–trimer stage, i.e. – for example a monomer radical abstracts a hydrogen atom from a dimer or trimer hydroxyl group, and then that newly formed dimer or trimer radical couples with a phenoxyl radical to form a trimer or tetramer, respectively.15 The mechanism of oxidative phenolic radical coupling, including resonance-delocalization of the unpaired electron density and consequent regiochemistry, with chemical agents has been recently reviewed,28,29 as has its use for natural product synthesis.30 The enzymatic oligomerization – precipitation pathway of phenoxyl radical coupling has been extensively reviewed and these reviews briefly cover comparison of SBP and HRP with n-alkylphenols, including cresols, with SBP being better with bulkier substituents.15,19,31,32 Calorimetric measurements were used to estimate the relative stabilities of cresyloxy radicals,33 from which it is inferred that the radical reactivities are m-cresyloxy < p-cresyloxy < o-cresyloxy (i.e. – o-cresyloxy radical is the least stable, most reactive); measured values of second-order rate constants for radical coupling in water (pH 8, 12 °C) for H-, o-, m-, p-of (5.6, 3.2, 1.5, 2.2) × 108 M−1 s−1 are in accord with the calorimetric ranking. In the absence of SBP data, rate constants (second-order, M−1 s−1 pH 7, 22 °C) are given for the reaction of HRP Compounds I and II with phenol and the cresols (H-, o-, m-, p-in the following) as determined by pre-steady-state methods: for compound I (H-, –, m-, p- (2.9, –, 7.8, 42; o-not given) × 106);34 for Compound II (H-, o-, m-, p-, (0.32, 0.081, 0.40, 1.06) × 106);35 the latter were closely matched by others.36 Thus, the Compound II decay with phenolic substrate is rate-limiting in the peroxidase cycle by 1–2 orders of magnitude. The relative ranking of these Compound II rate constants does not parallel the ranking of cresoloxy radical self-coupling rate constants, which suggests that the enzymic process is rate-limiting, not the coupling reaction.
This study aims to optimize conditions for achieving at least 95% removal (an arbitrary threshold to allow comparison with previous work in the lab; any given application may require optimization for a different target removal) of cresols from water using SBP in the presence of hydrogen peroxide at room temperature, thence to determine the time course and initial conversion kinetics. Our preliminary studies,14,17 will be enhanced by these kinetics and confirmation of the oligomerization – precipitation pathway by ultraviolet (UV) absorption spectrometry, high performance liquid chromatography (HPLC), and high-resolution mass spectrometry (HRMS). By improving the efficiency of cresol conversion, this research contributes to the development of sustainable and cost-effective biocatalytic wastewater treatment methods.
Experimental
Chemicals and reagents
All chemicals used in this study, including o-cresol (C85700-5G), m-cresol (65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 996-1ML-F), and p-cresol (C85751), bovine liver catalase, hydrochloric acid (HCl), 4-aminoantipyrine (4-AAP), formic acid, and phenol (99%) were obtained from Millipore Sigma Canada (Oakville, ON). The crude solid SBP (Industrial Grade, lot #18541NX was provided by Organic Technologies (Coshocton, OH). A 30% aqueous hydrogen peroxide solution, stored at 4 °C, monobasic and dibasic sodium phosphate were sourced from ACP Chemicals Inc. (Montreal, QC), while tris(hydroxymethyl)aminomethane was acquired from EM Science (Germany). Acetonitrile (ACN, lot #23F2662004) was obtained from VWR International Inc. (Mississauga, ON). Buffers (40 mM) were prepared, covering a pH range from 1.0 to 10.0:37 Citrate buffer was used for pH values of 4.0, 4.4, 5.0, and 5.4; phosphate buffer for pH values of 6.0, 6.5, 7.0, and 7.5; and tris(hydroxymethyl)aminomethane (tris) buffer for pH values of 8.0 and 9.0; pHs were measured at room temperature, 22 ± 2 °C. Syringe filters with a 0.22 µm pore size were supplied by Biosharp Life Science (China), and norm-jet syringes were from Henke Sass Wolf (Germany). Deionized water was from Culligan (Windsor, ON). The SBP stock solution was prepared by mixing 1.4 g of crude solid enzyme with 100 mL of distilled water at low speed (approximately 400 rpm) for 24 h. The suspension was centrifuged at 4000 rpm for 25 min, and the supernatant was taken as the stock solution and stored at 4 °C. For the catalase stock solution, solid bovine liver catalase was removed from the freezer and allowed to equilibrate to room temperature for at least 30 minutes. One gram of catalase was then stirred at 600 rpm with 100 mL of water for about 4 hours. The catalase solution was stored at 4 °C when not in use.
996-1ML-F), and p-cresol (C85751), bovine liver catalase, hydrochloric acid (HCl), 4-aminoantipyrine (4-AAP), formic acid, and phenol (99%) were obtained from Millipore Sigma Canada (Oakville, ON). The crude solid SBP (Industrial Grade, lot #18541NX was provided by Organic Technologies (Coshocton, OH). A 30% aqueous hydrogen peroxide solution, stored at 4 °C, monobasic and dibasic sodium phosphate were sourced from ACP Chemicals Inc. (Montreal, QC), while tris(hydroxymethyl)aminomethane was acquired from EM Science (Germany). Acetonitrile (ACN, lot #23F2662004) was obtained from VWR International Inc. (Mississauga, ON). Buffers (40 mM) were prepared, covering a pH range from 1.0 to 10.0:37 Citrate buffer was used for pH values of 4.0, 4.4, 5.0, and 5.4; phosphate buffer for pH values of 6.0, 6.5, 7.0, and 7.5; and tris(hydroxymethyl)aminomethane (tris) buffer for pH values of 8.0 and 9.0; pHs were measured at room temperature, 22 ± 2 °C. Syringe filters with a 0.22 µm pore size were supplied by Biosharp Life Science (China), and norm-jet syringes were from Henke Sass Wolf (Germany). Deionized water was from Culligan (Windsor, ON). The SBP stock solution was prepared by mixing 1.4 g of crude solid enzyme with 100 mL of distilled water at low speed (approximately 400 rpm) for 24 h. The suspension was centrifuged at 4000 rpm for 25 min, and the supernatant was taken as the stock solution and stored at 4 °C. For the catalase stock solution, solid bovine liver catalase was removed from the freezer and allowed to equilibrate to room temperature for at least 30 minutes. One gram of catalase was then stirred at 600 rpm with 100 mL of water for about 4 hours. The catalase solution was stored at 4 °C when not in use.
Enzyme activity assay and pH
The standard catalytic unit (U) for measuring SBP activity is defined as the amount of enzyme required to consume 1.0 µmol of H2O2 per minute in the assay. Enzyme activity was determined by monitoring the rate of formation of a pink chromophore at 510 nm. This chromophore results from the oxidative coupling of phenol (10 mM) and 4-AAP (2.4 mM)) in the presence of H2O2 (0.2 mM), in 50 mM phosphate buffer, pH 7.4, with SBP serving as the catalyst. To facilitate reagent preparation, a 10X concentrate of 100 mM phenol and 0.5 M phosphate buffer (pH 7.4) and 100 mM H2O2 in water (510 µL 30% H2O2 made up to 50 mL) were used: 5.0 mL of 10X concentrate plus 0.10 mL 100 mM H2O2 plus 25 mg of AAP were made up to 47.5 mL with water. For the assay, SBP stock solution was diluted 50-fold in buffer, and 50 µL of that dilution was mixed with 950 µL of reagent solution and monitored at 510 nm for 30 s at 5 s intervals. Spectrometer software calculated the rate and activity. For the blank, 950 µL of reagent and 50 µL of distilled water were mixed.Experimental protocols
The tests in this study were conducted in batch reactors at room temperature (approximately 22 °C). Batch reactors consisted of glass vials (30 mL) containing 20 mL of a solution made up of deionized water, 40 mM buffer, 1 mM cresol, SBP, and H2O2. After adding the SBP and H2O2 to the reactor, the mixture was stirred with a Teflon-coated stir bar placed on a magnetic stirrer. The reaction was initiated by the addition of H2O2, and after three hours, 100 µL of catalase stock solution (1.0 g/100 mL) was added to rapidly quench the reaction. The residual substrate concentration was determined via HPLC analysis after the sample was microfiltered using pre-conditioned 0.22 µm polyethersulfone (PES) syringe filters.Analytical equipment
A Waters (Mississauga, ON) HPLC system with binary pumps (model 1525), a dual-wavelength absorbance detector (model 2487), and an autosampler (model 717 plus) was utilized to precisely measure the remaining amount of the molecule. Breeze software was used to operate this system, which was connected to a computer. Table 1 provides the mobile phase ratios, flow rate, column temperature, and column employed. Acetonitrile was solvent A and 0.1% formic acid in water was solvent B.| Substrate | Mobile phase ratio | Flow (mL min−1) | λmax (nm) | Column temperature | Column | |
|---|---|---|---|---|---|---|
| Pump A (75%) | Pump B (25%) | |||||
| o-cresol | Acetonitrile | 30% formic acid (0.1%) | 1.0 | 270 | 40 °C | Symmetry C18 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm (4.6 mm × 100 mm) µm (4.6 mm × 100 mm) |
| m-cresol | 271 | |||||
| p-cresol | 277 | |||||
UV-vis measurements were conducted with an Agilent (Mississauga, ON) diode array UV-vis spectrophotometer (model 8453 controlled by a Hewlett Packard Vectra ES/12 computer) with a range of 190–1100 nm and 1 nm resolution.
An Agilent 6546 instrument was used for LC-HRMS analysis at the Brock University Mass Spectrometry Facility (St. Catharines, ON). LC analysis was performed with a 5 µL injection volume at a flow rate of 0.4 mL min−1, using a column temperature of 30 °C. The mobile phases consisted of water with 0.1% formic acid (A) and methanol (B). The gradient started at 5% B at 0 min, increased to 100% B by 5 min, and was held at 100% B until 14 min, followed by a 2 min post-run equilibration; MS analysis used the Jetstream SI negative mode for the ion source and a 5 µL injection volume. The probe's mass-to-charge ratio (m/z) acquisition range was 100–1500. After enzymatic treatment, all substrates were filtered using a 0.22 µm cut-off filter. For measurement, two solutions were prepared: one containing filtered reaction solution and another consisting of a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of the sample suspension and ACN, filtered after mixing for 2 h. Reaction mixtures under specified conditions were analyzed to identify potential oligomerization products resulting from the enzymatic treatment. Since buffers, particularly phosphate, interfere with product peaks in MS, the buffer content for MS samples in batch reactors was reduced to 10 mM.
1 mixture of the sample suspension and ACN, filtered after mixing for 2 h. Reaction mixtures under specified conditions were analyzed to identify potential oligomerization products resulting from the enzymatic treatment. Since buffers, particularly phosphate, interfere with product peaks in MS, the buffer content for MS samples in batch reactors was reduced to 10 mM.
Results and discussion
pH optimization
The activity and stability of SBP are influenced by the pH of the reaction medium, a crucial biocatalytic parameter that impacts its industrial use. Biocatalysts such as SBP are characterized by specific 3-dimensional structures for which the state of ionizable amino acid residue side chains plays a critical role in the catalytic activity and conformational stability of the enzyme. Moreover, the pH of the reaction medium influences both the ionization state of the reducing substrate and the efficiency of electron transfer.38 Because of its solvent-exposed heme edge, SBP's catalytic activity is controlled by pH, although it exhibits considerable activity over a wide pH range, pH 3.0 to pH 10.0.39 For example, when guaiacol is used as the reducing substrate, SBP demonstrates peak activity between pH 5.5 and 6.0.40 SBP derived from seed hulls may efficiently oxidize a variety of substrates over a broad pH range.41The effect of pH on substrate conversion was examined under stringent conditions, defined as a pH range over which there was insufficient enzyme available so that any pH effects would be clearly discerned (Fig. 1). For these experiments at 1.0 mM cresol, 1.5 mM peroxide was chosen as being non-limiting based on previous experience in the lab. Initial range-finding experiments were conducted by UV-vis monitoring (data not shown), which expedited confirmation by HPLC analysis. The trends for the two methods were the same, with HPLC consistently showing better removal by 2–10% (example given in Supplemental information, Fig. S1). From Fig. 1, pHs of 7.0, 8.0, and 9.0 were chosen for further experiments with p-, m-, and o-cresols, respectively. For o-cresol, pH 9.0 was the compromise choice for the most effective pH for two reasons: more basic pHs are inconvenient for wastewater treatment, and concern about the progressive ionization of the substrate due to its pKa (10.29).42,43 Analogously, pHs 8.0 and 7.0, were chosen for m- and p-cresol, respectively.
These studies have been conducted in buffers; however, in any real-wastewater, treatment samples would not be buffered but adjusted to the optimal pHs determined here (with continuous pH control, if necessary), as a starting point for re-optimization.
SBP optimization
Since the cost of the enzyme might be a major obstacle to applying enzymatic treatment to actual wastewater, it is essential to use the minimum effective enzyme activity to achieve 95% removal. Only then can its cost be calculated. Preliminary range-finding was conducted by UV-vis analysis, subsequently confirmed by HPLC analysis (Fig. 2). The studies were conducted at the pH optima determined above, and the H2O2 concentration was kept at the non-limiting values shown. For o-cresol at pH 9.0 with 1.5 mM H2O2, increasing SBP activities from 0.5 to 0.9 U mL−1 significantly improved removal efficiency, achieving 5% remaining with 0.7 U mL−1 SBP. Similarly, for m-cresol at pH 8.0 with 1.1 mM H2O2, 0.8 U mL−1, and for p-cresol at pH 7.0 with 1.5 mM H2O2, the remaining percentage dropped to ≤5% at 0.8 and 0.3 U mL−1 SBP, respectively. Thus, 0.7, 0.8, and 0.3 U mL−1 SBP were selected for further studies with o-, m-, and p-cresols, respectively.Hydrogen peroxide optimization
As a co-substrate, hydrogen peroxide is essential to the enzymatic treatment process because it is the stoichiometric oxidant. H2O2 concentrations too high can cause peroxidase inactivation, while too low can cause insufficient substrate conversion. One mole of H2O2 produces two moles of free radicals from two moles of substrate in the oxidation-reduction mechanism of the peroxidase cycle (i.e. – peroxide:phenol stoichiometry from one turn of the cycle is 0.5). This stoichiometry, however, may change depending on the endogenous catalase activity of the peroxidase and on further enzymatic cycles (there is an upper limit of peroxide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) phenol ratio of 1.0 for an infinite polymer, thus any excess peroxide demand beyond this limit is attributed to catalase activity), which may produce higher oligomers from the first-generated dimers.41 To achieve >95% removal within a 3 hours reaction time, experiments were conducted to optimize the hydrogen peroxide concentration utilizing the respective pH optima and ideal activity of SBP for each substrate, determined above, Fig. 3. The data show that 1.2, 1.1, and 1.0 mM peroxide are optimal for 1.0 mM o-, m-, and p-cresols, respectively. The peroxide demand of these substrates is also in accord with that found for many other substrates with SBP.44–46 Excess peroxide is to be avoided in these reactions as it can lead to reversible (via Compound III) and/or irreversible (via compound P-670) enzyme inactivation.31
phenol ratio of 1.0 for an infinite polymer, thus any excess peroxide demand beyond this limit is attributed to catalase activity), which may produce higher oligomers from the first-generated dimers.41 To achieve >95% removal within a 3 hours reaction time, experiments were conducted to optimize the hydrogen peroxide concentration utilizing the respective pH optima and ideal activity of SBP for each substrate, determined above, Fig. 3. The data show that 1.2, 1.1, and 1.0 mM peroxide are optimal for 1.0 mM o-, m-, and p-cresols, respectively. The peroxide demand of these substrates is also in accord with that found for many other substrates with SBP.44–46 Excess peroxide is to be avoided in these reactions as it can lead to reversible (via Compound III) and/or irreversible (via compound P-670) enzyme inactivation.31
Summary of optimum conditions for cresols treatment
The ideal circumstances for removing at least 95% of cresols from water using SBP at room temperature are displayed in Table 2. Removal was characterized by two techniques, HPLC, the more reliable one, and UV, the less so, being based on small residual UV absorbances and the possible ‘colour correction’ thereto, as noted in the footnote (also demonstrated in Fig. S1). The phenomenon of SBP sorption to polyphenolic resins formed in such reactions has been quantitatively characterized.47 The two techniques are in good agreement as to their respective residuals, regardless of the ambiguity of colour correction. More importantly, the residual UV may be taken as a proxy for total organic carbon measurements, confirming that virtually all the aromatic starting material indicated as converted by HPLC has indeed been removed in the precipitate, according to the residual UV. This assertion assumes that the reaction products are aromatic oxidative oligomers as discussed in the Introduction and confirmed by mass spectrometry, below. Differences in efficiency are unlikely to be due to the cresol pKa s, since at their respective optimal pHs, all are predominantly (≥90%) in the –OH form. Thus, the pH-dependence of the enzymatic reaction is influenced by the ionization behavior of the enzyme, favouring the p-isomer. Relative to p-, the o- and m-isomers are disfavoured sterically or electronically, respectively.| Optimum condition | o-cresol (1 mM) | m-cresol (1 mM) | p-cresol (1 mM) |
|---|---|---|---|
| a pKa Values in Water Compilation;48 phenol pKa in the same source 9.95.b Residual UV measurements were ‘colour-corrected’ for contributions due to SBP and catalase heme groups at the respective analytical wavelengths; the first values shown are for full correction, as if none of the added enzymes were removed with the reaction precipitate; the values in parentheses are for no correction, as if all the added enzymes were removed with the reaction precipitate. | |||
| pH (pKaa) | 9.0 (10.3) | 8.0 (10.1) | 7.0 (10.2) |
| H2O2 (mM) | 1.2 | 1.1 | 1.0 |
| SBP (U mL−1) | 0.7 | 0.8 | 0.3 |
| Removal % (HPLC) | 95 | 95 | 98 |
| Removal % (UV)b | 97 (90) | 100 (96) | 98 (92) |
Optimization was conducted one factor at a time, for the sake of expedience and to assemble baseline information on SBP with this class of compounds, but if this enzymatic method were to be considered for implementation on a real wastewater, it would be re-optimized by response-surface methodology to take account of possible parameter interactions,49 starting with the parameter ranges developed here; it would also deal with the probable mixtures of compounds and any matrix effects due to non-substrates, inhibitors, etc. in the sample.
Comparison of treatment efficiencies
The optimum SBP activities for treatment of the cresols are listed with those of several other compounds in SI Table S1. For quantitative comparison, the activities are normalized with respect to the respective substrate concentrations. The cresols fall in the moderate SBP requirement range of 0.1 to 1.0 U mL−1 mM−1. Alternatively, efficiency could be characterized in terms of ‘enzyme turnovers’, the number of substrate molecules converted per molecule of enzyme in its catalytic lifetime.50 In that case, the cresols show turnovers in the range of 2000–5000. In a treatment scenario, the only sludge generated would be the cresol oligomers, which could be used as or formulated into resins, if in sufficient quantities; barring that, the sludge could be sent to a restricted landfill or incinerated for energy content.Time course of reaction
Designing a treatment facility requires careful consideration of reaction time since it affects reactor volume and ultimate cost. Three hours was chosen arbitrarily for the reaction time from previous studies in the lab. Finding the shortest time to achieve over 95% pollutant conversion is crucial for efficiency. Substrate conversion was monitored for three hours under ideal conditions to determine the residual substrate concentrations. Eqn (1) and (2) were used to calculate the initial first-order half-lives of the cresols from the initial pseudo-first-order rate constants, k. It is anticipated that the gradual loss of the unmonitored substrate, hydrogen peroxide, and loss of enzyme activity will cause these processes to lose first-order behaviour very quickly. However, this initial rate constant will help compare the reactivity of the enzyme with different substrates in the absence of a thorough kinetic analysis.| C = C0e−kt | (1) |
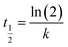 | (2) |
Fig. 4 illustrates the time course of reactions for the cresols under their respective optimized conditions, with first-order exponential fits applied to the initial reaction stages, as shown in the legend, and the rate constants are used to calculate the half-lives given in Comparison of SBP Kinetic Efficiencies, below. After 60 minutes, all cresols had reached 89 ± 2% conversion.
Comparison of SBP kinetic efficiencies
Half-lives derived from initial first-order rate constants of several substrates with the same SBP preparation are given in SI Table S2. For quantitative comparison, the half-lives are normalized with respect to enzyme activity and compared with several other compounds tabulated previously.51 The cresols' normalized half-lives fall within the moderate range, 1–6 min. U mL−1, of the tabulated compounds. Kinetic reactivity is likely governed in part by radical stability; the more stable the radical, the lower the cresol's reactivity with the enzyme's activated heme group. However, all three isomers will have significant unpaired electron density at their o- and p-positions, not meta-. The location of the ring methyl group in those radicals will have little influence on the unpaired electron density, but it will influence steric accessibility to the enzyme heme group. The normalized half-lives in Table S2, shortest for p-cresol, somewhat longer for m-cresol, and several-fold longer for o-cresol, parallel their order of increasing steric hindrance, and also the ranking of HRP Compound II with the cresol isomers given in the Introduction. The initial reactivities notwithstanding, Fig. 4 illustrates that with the appropriate enzyme activity, the curves converge with the passage of time. Differences in cresol isomer reactivities are accommodated by reaction time and enzyme activity; the specific regiochemistry of product formation is irrelevant in this work, as long as the oligomers precipitate to enable removal.Mass spectrometry (MS)
The peroxidase catalytic cycle produces free radicals that couple non-enzymatically. Because unpaired electron density is delocalized around the ring (see, for example, the computation for phenol in52), these radicals can connect C–O and C–C with o-, p-orientation (O–O coupling is unstable). The coupling reactions are oxidative; for example, an oxidative dimer would be symbolized, M2 – 2H, where M represents the monomer molecular weight. Thus, isomeric coupling products form depending on resonance-stabilized radical electron density delocalization.53 Until the oligomers precipitate due to reaching their solubility limit, the peroxidase cycle continues. Understanding the nature of the products is essential for determining their potential toxicity, and MS analysis helps with this process. MS analysis cannot differentiate between isomeric structures but it does provide insight into their specific constitutions. An electrospray ionization technique was used for MS analysis (in high-resolution negative-ion mode). Probable formulas have been derived for the molecular weight of the oligomers from mass-to-charge ratio, or m/z, in mass spectrometry data. The isotope abundance was considered during the analysis to verify 13C consistent with the given formulas. All the observed peaks were for formulas related to proton loss since negative-ion spectra were collected. Table 3 gives the data for the o-cresol reaction filtrate. Oligomers up to the nonamer stage were present in the filtered reaction mixture. For the molecular formulas, mass accuracy using high-resolution mass spectrometry of less than 10 ppm is considered unambiguous. The oligomerization observed here is in accord with many others, for example, triclosan,15,53 phenol,54 and acetaminophen.55| Symbols | Molecular formula | Mono-isotopic Mass (m/z) | Theoretical Mass (m/z) for H loss | Observed Mass (m/z) | Mass accuracy (ppm) | Detected |
|---|---|---|---|---|---|---|
| M | C7H8O | 108.0575 | 107.0502 | 107.0501 | −1.29 | * |
| M2 – 2H | C14H14O2 | 214.0994 | 213.0921 | 213.0922 | 0.45 | * |
| M3 – 4H | C21H20O3 | 320.1425 | 319.1340 | 319.1339 | −0.21 | * |
| M4 – 6H | C28H26O4 | 426.1831 | 425.1758 | 425.1755 | −0.78 | * |
| M5 – 8H | C35H32O5 | 532.2250 | 531.2177 | 531.2170 | −1.31 | * |
| M6 – 10H | C42H38O6 | 638.2668 | 637.2596 | 637.2587 | −1.35 | * |
| M7 – 12H | C49H44O7 | 744.3087 | 743.3014 | 743.3007 | −0.98 | * |
| M8 – 14H | C56H50O8 | 850.3506 | 849.3433 | 849.3420 | −1.52 | * |
| M9 – 16H | C63H56O9 | 956.3924 | 955.3852 | 955.3846 | −0.58 | * |
| M10 – 18H | C70H62O10 | 1062.4343 | 1061.4265 |
Because the MS approach was unable to handle the presence of reaction solids, a sample of the reaction suspension was diluted with an equal volume of acetonitrile, allowed to stand for three hours with occasional vortexing, and then filtered to obtain information on any precipitated oligomers brought into solution with the organic solvent. The corresponding Table for this o-cresol suspension, given as SI, Table S3, showed no qualitative difference with Table 3. Yang et al. reported an electrochemical approach to aqueous o-cresol oligomerization and precipitation, for which they detected oligomers up to the dodecamer stage.56 Analogous tables for m-cresol (Tables S4 and S5) and p-cresol (Tables S6 and S7) are also given in the SI. For m-cresol, only dimers were seen in the aqueous filtrate, whereas the aqueous-organic filtrate showed oligomers up to the octamer stage. For p-cresol, up to octamers were seen in the aqueous filtrate, whereas the aqueous-organic filtrate showed oligomers up to the nonamer stage.
The foregoing qualitative analysis only partially supported the hypothesis that higher oligomers were present in the precipitates than in the aqueous reaction mixtures. Semi-quantitative analysis of the integrated ion-current chromatograms provided further insights into the distribution of oligomers in different filtrates. These findings highlight the influence of molecular structure on oligomerization and solubility. The differences among the cresol isomers suggest that radical coupling efficiency and solubility constraints vary with positional isomerism. The limited solubility of m-cresol oligomers in aqueous conditions implies that steric or electronic factors restrict their further radical-mediated coupling, while o- and p-cresols undergo extended polymerization. ACN, as a solubilizing agent, enables the detection of higher oligomers that have precipitated. This suggests that solvent choice for reactions conducted in aqueous-organic mixtures would be important for studying oligomerization pathways and that modifying reaction conditions could help control product distribution.15,57 Future work could explore how solvent polarity, pH, or additional SBP influence the extent and nature of oligomer formation, particularly for industrial or environmental applications where cresol derivatives play a role in polymer synthesis or pollutant treatment, respectively.
Conclusions
This work was carried out to provide data from which a future, more complicated, study of real wastewater could be done. Real wastewater would likely contain several substrates and other constituents, which could influence the course of the enzymic reaction (the matrix effect). Soybean peroxidase demonstrated remarkable efficacy in the removal of cresols from aqueous solutions, achieving removal efficiencies of 95% or better under optimized reaction conditions. The enzymatic treatment exhibited rapid initial kinetics, with half-lives ranging from 2 to 9 minutes. Residual UV analysis and high-resolution mass spectrometry confirmed the formation of expected oligomeric transformation products, supporting the hypothesis that SBP catalyzes cresol removal primarily through an oxidative coupling and oligomerization pathway. Thus, SBP has large potential for sustainable, cost-effective treatment of industrial waste or process water. This would enable a circular economy in the soy value chain, as the crude seedcoat extract is suitable for the treatment, leaving the seedcoat residue available for its intended use in the formulation of animal feed.Author contributions
Mohammadreza Haghighatnama, Samira Narimannejad, Keith Taylor, and Nihar Biswas contributed to the study conception, design, and analysis. Material preparation and data collection were performed by Mohammadreza Haghighatnama and Samira Narimannejad. The first draft of the manuscript was written by Mohammadreza Haghighatnama. Mohammadreza Haghighatnama, Samira Narimannejad, Keith Taylor, and Nihar Biswas commented on and contributed to subsequent versions of the manuscript. Mohammadreza Haghighatnama, Samira Narimannejad, Keith Taylor, and Nihar Biswas read and approved the final manuscript.Conflicts of interest
The authors declare no relevant financial or non-financial competing interest.Data availability
All the data supporting the findings of this study are available within the paper or its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra08019c.Acknowledgements
Financial support from the Natural Sciences and Engineering Research Council of Canada and the University of Windsor is gratefully acknowledged.References
- ATSDR, Toxicological Profile for Cresols, 2015, https://wwwn.cdc.gov/TSP/ToxProfiles/ToxProfiles.aspx?id=946%26tid=196.
- Health Canada, Draft Assessment – Substituted Phenols Group, Government of Canada, 2015, https://www.canada.ca/en/environment-climate-change/services/evaluating-existing-substances/draft-assessment-substituted-phenols-group.html?utm_source=chatgpt.com Search PubMed.
- eCFR.gov, Code of Federal Regulations, 2006. https://www.ecfr.gov/current/title-40/chapter-I/subchapter-I/part-261/subpart-C/section-261.24 Search PubMed.
- NJDEP, Ground Water Quality Standard for o-, m-, and p-Cresol, 2015, https://dep.nj.gov/wp-content/uploads/bears/cresols_mixed_fact_sheet.pdf.
- O. B. Adesina, C. William and E. I. Oke, Evolution in water treatment: Exploring traditional self-purification methods and emerging technologies for drinking water and wastewater treatment: A review, World News Nat. Sci., 2024, 53, 169–185 CAS.
- L. Y. Jun, L. S. Yon and N. Mubarak, et al., An overview of immobilized enzyme technologies for dye and phenolic removal from wastewater, J. Environ. Chem. Eng., 2019, 7, 102961 CrossRef CAS.
- S. Salehi, K. Abdollahi and R. Panahi, et al., Applications of biocatalysts for sustainable oxidation of phenolic pollutants: A review, Sustainability, 2021, 13, 8620 CrossRef CAS.
- K. Sellami, A. Couvert and N. Nasrallah, et al., Peroxidase enzymes as green catalysts for bioremediation and biotechnological applications: A review, Sci. Total Environ., 2022, 806, 150500 Search PubMed.
- M. K. Al-Sakkaf, I. Basfer and M. Iddrisu, et al., An up-to-date review on the remediation of dyes and phenolic compounds from wastewaters using enzymes immobilized on emerging and nanostructured materials: Promises and challenges, Nanomaterials, 2023, 13, 2152 Search PubMed.
- A. Rostami, A. Abdelrasoul and Z. Shokri, et al., Applications and mechanisms of free and immobilized laccase in detoxification of phenolic compounds—a review, Korean J. Chem. Eng., 2022, 39, 821–832 CrossRef CAS.
- K. A. Almaqdi, R. Morsi and B. Alhayuti, et al., LC-MSMS based screening of emerging pollutant degradation by different peroxidases, BMC Biotechnol., 2019, 19, 83 CrossRef CAS PubMed.
- S. Saikia, R. D. Gogoi and M. Yadav, et al., Isolation, purification and characterization of peroxidase from Raphanus sativus and its applications in biotransformation of cresols, Biocatal. Agric. Biotechnol., 2022, 46, 102540 CrossRef CAS.
- K. Yamada, T. Shibuya and M. Noda, et al., Influence of position of substituent groups on removal of chlorophenols and cresols by horseradish peroxidase and determination of optimum conditions, Biosci., Biotechnol., Biochem., 2007, 71, 2503–2510 CrossRef CAS PubMed.
- N. Caza, J. Bewtra and N. Biswas, et al., Removal of phenolic compounds from synthetic wastewater using soybean peroxidase, Water Res., 1999, 33, 3012–3018 CrossRef CAS.
- H. Uyama, Synthesis of Poly (aromatic) s I: Oxidoreductase as Catalyst, Enzymatic Polymerization towards Green Polymer Chemistry, S. Kobayshi et al., Springer Nature, Singapore, 2019, pp. 267–305 Search PubMed.
- H. Wright and J. A. Nicell, Characterization of soybean peroxidase for the treatment of aqueous phenols, Bioresour. Technol., 1999, 70, 69–79 CrossRef CAS.
- M. Haghighatnama, S. Narimannejad, N. Biswas, et al., Enzymatic Treatment of Cresols in Water, in: Turbulence & Energy Laboratory Annual Conference, Springer, 2024, pp.53–67 Search PubMed.
- J. A. Kamal and D. V. Behere, Kinetic stabilities of soybean and horseradish peroxidases, Biochem. Eng. J., 2008, 38, 110–114 CrossRef CAS.
- M. Ayala and E. Torres, Peroxidases as potential industrial biocatalysts, Heme Peroxidases, E. Raven and B. Dunford, Royal Society of Chemistry, London, 2015 Search PubMed.
- A. Steevensz, L. G. Cordova Villegas and W. Feng, et al., Soybean peroxidase for industrial wastewater treatment: a mini review, J. Environ. Eng. Sci., 2014, 9, 181–186 CrossRef.
- M. Crnoglavac Popović, M. Stanišić and R. Prodanović, State of the art technologies for high yield heterologous expression and production of oxidoreductase enzymes: Glucose oxidase, cellobiose dehydrogenase, horseradish peroxidase, and laccases in yeasts P. pastoris and S. cerevisiae, Fermentation, 2024, 10, 93 CrossRef.
- M. I. Sánchez-Ruiz, I. Ayuso-Fernández, D. Linde, et al., Ligninolytic peroxidases: A guide for their heterologous expression in E. coli and protocols to evaluate their activity on lignin, Methods in Enzymology, Elsevier, 2025, pp. 361–379 Search PubMed.
- A. Bassi, Z. Geng and M. Gijzen, Enzymatic removal of phenol and chlorophenols using soybean seed hulls, Eng. Life Sci., 2004, 4, 125–130 CrossRef CAS.
- K. Wilberg, C. Assenhaimer and J. Rubio, Removal of aqueous phenol catalysed by a low purity soybean peroxidase, J. Chem. Technol. Biotechnol., 2002, 77, 851–857 CrossRef CAS.
- S. Feng, H. H. Ngo and W. Guo, et al., Roles and applications of enzymes for resistant pollutants removal in wastewater treatment, Bioresour. Technol., 2021, 335, 125278 CrossRef CAS.
- S. Feng, W. Guo and A. Ding, et al., Enzyme sources in wastewater treatment: Their influence on enzymatic bioremediation and large-scale applications, Chem. Eng. J., 2025, 161891 CrossRef CAS.
- H. B. Dunford, Heme peroxidase kinetics, Heme Peroxidases, E. Raven and B. Dunford, Royal Society of Chemistry, London, 2015 Search PubMed.
- J. Qian, X. Zhang and Y. Jia, et al., Oxidative polymerization in water treatment: chemical fundamentals and future perspectives, Environ. Sci. Technol., 2025, 59, 1060–1079 CrossRef CAS.
- H. Choi, S. E. Suh and H. Kang, Advances in Exploring Mechanisms of Oxidative Phenolic Coupling Reactions, Adv. Synth. Catal., 2024, 366, 4347–4384 CrossRef CAS.
- M. C. Carson and M. C. Kozlowski, Recent advances in oxidative phenol coupling for the total synthesis of natural products, Nat. Prod. Rep., 2024, 41, 208–227 RSC.
- M. Reihmann and H. Ritter, Synthesis of phenol polymers using peroxidases, Enzyme-Catalyzed Synthesis of Polymers, Springer, 2005, pp. 1–49 Search PubMed.
- I. Wazeer, H. F. Hizaddin and S. Mulyono, et al., Removal of cresol contaminants from aqueous media using hydrophobic deep eutectic solvents, J. Mol. Liq., 2024, 407, 125082 CrossRef CAS.
- L. S. Richard, C. E. Bernardes and H. P. Diogo, et al., Energetics of cresols and of methylphenoxyl radicals, J. Phys. Chem., 2007, 111, 8741–8748 Search PubMed.
- D. Job and H. B. Dunford, Substituent effect on the oxidation of phenols and aromatic amines by horseradish peroxidase compound I, Eur. J. Biochem., 1976, 66, 607–614 Search PubMed.
- J. Sakurada, R. Sekiguchi and K. Sato, et al., Kinetic and molecular orbital studies on the rate of oxidation of monosubstituted phenols and anilines by horseradish peroxidase compound II, Biochemistry, 1990, 29, 4093–4098 CrossRef CAS PubMed.
- P. K. Patel, M. S. Mondal and S. Modi, et al., Kinetic studies on the oxidation of phenols by the horseradish peroxidase compound II, Biochim. Biophys. Acta Protein Struct. Mol. Enzymol., 1997, 1339, 79–87 CrossRef CAS PubMed.
- G. Gomori, [16] Preparation of Buffers for Use in Enzyme Studies, 1955 Search PubMed.
- L. Parsiavash, A. Saboora and S. Z. Moosavi Nejad, Investigating on the Stability of Peroxidase Extracted from Soybean (Glycine max var. Williams) and Effects of Na+ and K+ Ions on its Activity, J. Cell Mol. Res., 2015, 7, 94–101 CAS.
- J. A. Kamal and D. V. Behere, Activity, stability and conformational flexibility of seed coat soybean peroxidase, J. Inorg. Biochem., 2003, 94, 236–242 CrossRef CAS.
- Z. Geng, K. J. Rao and A. S. Bassi, et al., Investigation of biocatalytic properties of soybean seed hull peroxidase, Catal. Today, 2001, 64, 233–238 CrossRef CAS.
- B. Valderrama, M. Ayala and R. Vazquez-Duhalt, Suicide inactivation of peroxidases and the challenge of engineering more robust enzymes, Chem. Biol., 2002, 9, 555–565 CrossRef CAS.
- M. Sobiesiak, Chemical structure of phenols and its consequence for sorption processes, Phenolic Compounds-Natural Sources, Importance and Applications, 2017, pp. 3–26 Search PubMed.
- W. Spirit, Health and Safety Guide (Health and Safety Guide No. 103), United Nations Environment Programme, International Labour Organisation, World Health Organization, Geneva, 1996 Search PubMed.
- D. Mukherjee, K. Taylor and N. Biswas, Soybean peroxidase-induced treatment of dye-derived arylamines in water, Water, Air, Soil Pollut., 2018, 229, 283 CrossRef.
- N. Mashhadi, K. E. Taylor and N. Jimenez, et al., Removal of selected pharmaceuticals and personal care products from wastewater using soybean peroxidase, Environ. Manag., 2019, 63, 408–415 CrossRef PubMed.
- M. Sharifzadeh, S. Narimannejad and K. E. Taylor, et al., Enzymatic removal of the sulfa drugs sulfamethoxazole and sulfamerazine from synthetic wastewater by soybean peroxidase, Environ. Sci. Pollut. Res., 2024, 31, 64760–64771 CrossRef CAS PubMed.
- K. W. Beemer, K. E. Taylor and N. Biswas, Sorption properties of peroxidase-catalysed polyphenolic resin enable aromatics' capture, J. Environ. Eng. Sci., 2019, 14, 90–96 CrossRef.
- R. Williams, pKa values in water compilation, ACS Division of Organic Chemistry, 2022, pp. 1–33, https://organicchemistrydata.org/hansreich/resources/pka/pka_data/pka-compilation-williams.pdf Search PubMed.
- L. G. Cordova-Villegas, A. Y. Cordova-Villegas and K. E. Taylor, et al., Response surface methodology for optimization of enzyme-catalyzed azo dye decolorization, J. Environ. Eng., 2019, 145, 04019013 CrossRef.
- A. Kaur, K. E. Taylor and N. Biswas, Soybean peroxidase-catalyzed degradation of a sulfonated dye and its azo-cleavage product, J. Chem. Technol. Biotechnol., 2021, 96, 423–430 Search PubMed.
- B. N. Ziayee, N. Mashhadi and K. E. Taylor, et al., Elimination of selected heterocyclic aromatic emerging contaminants from water using soybean peroxidase, Environ. Sci. Pollut. Res., 2021, 28, 37570–37579 CrossRef.
- N. Mashhadi, K. E. Taylor and N. Biswas, et al., Biocatalytic oligomerization of azoles; experimental and computational studies, Environ. Sci.:Water Res. Technol., 2021, 7, 1103–1113 RSC.
- J. Li, J. Peng and Y. Zhang, et al., Removal of triclosan via peroxidases-mediated reactions in water: Reaction kinetics, products and detoxification, J. Hazard. Mater., 2016, 310, 152–160 CrossRef CAS PubMed.
- Q. Huang, J. Tang and Jr W. J. Weber, Precipitation of enzyme-catalyzed phenol oxidative coupling products: Background ion and pH effects, Water Res., 2005, 39, 3021–3027 CrossRef CAS PubMed.
- J. Lu, Q. Huang and L. Mao, Removal of acetaminophen using enzyme-mediated oxidative coupling processes: I. Reaction rates and pathways, Environ. Sci. Technol., 2009, 43, 7062–7067 CrossRef CAS PubMed.
- S.-Q. Yang, Y.-H. Cui and W.-J. Xue, et al., Mechanism of o-cresol polymerization and its carbon recovery from aqueous solution by a cathode/Fe2+/H2O2 process, Chem. Eng. J., 2024, 499, 156385 CrossRef CAS.
- E. E. Hood and S. Narimannejad, Compositions and Methods for Degradation of Pollutants by Plant-Produced Manganese peroxidase, Patent US20250161732A1, 2025 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |




