 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Lonidamine-1,3,4-oxadiazole derivatives with antiproliferative effects on HCT116 colon cancer cell lines: biological evaluation, ADMET, and computational studies
Raveendra Madhukar Bhatab,
A. N. Priyadarshinic,
M. S. Sudhanvac,
Gangadhar V. Muddapurd,
Kawthar Alhussienie,
Raman Kumar Kb and
Rangappa S. Keri *a
*a
aCentre for Nano and Material Sciences, Jain (Deemed-to-be University), Jain Global Campus, Jakkasandra Post, Kanakapura Road, Ramanagara District, Bangalore, Karnataka 562112, India. E-mail: sk.rangappa@jainuniversity.ac.in; keriphd@gmail.com; Tel: +918027577199 Tel: +919620667075
bAurigene Pharmaceutical Services, Bangalore, Karnataka, India
cAdichunchanagiri Institute for Molecular Medicine, Adichunchanagiri Institute of Medical Sciences, Adichunchanagiri University, BG Nagara-571448, Karnataka, India
dDepartment of Physics, KLE Technological University, Hubballi-580031, Karnataka, India
eDepartment of Pharmaceutical Chemistry, Faculty of Pharmacy, Universiti Malaya, Kuala Lumpur 50603, Malaysia
First published on 11th March 2026
Abstract
In recent years, cancer has emerged as a significant challenge for healthcare systems, posing challenges to researchers to develop new treatments. Among various cancers, colorectal cancer is a major cause of death worldwide. To develop novel compounds that strongly inhibit colon cancer cells, lonidamine-1,3,4-oxadiazole derivatives 7(a–h) were designed and synthesized. The prepared compounds were characterized by various spectral techniques, including NMR (1H and 13C), mass spectra, and HPLC. Cytotoxicity tests conducted on a colorectal cancer cell line indicated that compound (7d) demonstrated significant antiproliferative effects, achieving the lowest IC50 value of 12.91 ± 1.58 µM, thereby making it the most effective among the compounds tested. This compound induces apoptosis, as evidenced by Hoechst/PI double staining, and mitigates cell migration, demonstrating its antiproliferative and antimigratory capabilities. Molecular docking, dynamics simulations, and DFT studies helped clarify the structure–activity relationship (SAR) and mechanisms of action. ADME and toxicity predictions also supported its drug-like properties. SAR analysis identified key substituents influencing activity, guiding further optimization. Overall, compound (7d) appears to be a promising candidate for colon cancer therapy, though additional studies are necessary to assess its clinical potential. These findings could be used for designing novel cancer therapeutic or preventive LND-derived agents.
1 Introduction
Colon cancer, also known as colorectal cancer (CRC), is one of the most common types of cancer that originates in the colon of the digestive tract. It contributes to significant morbidity and mortality.1 CRC is the third most often diagnosed malignancy and the second largest cause of cancer-related mortality.2,3 In 2020, the global figures indicated 1.9 million new cases and fatalities, an increase from 1.8 million new cases and 861![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths in 2018.4,5 Additionally, it is projected that 107
000 deaths in 2018.4,5 Additionally, it is projected that 107![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 320 more cases will be identified by 2025, and approximately 53
320 more cases will be identified by 2025, and approximately 53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 individuals are expected to die from colon cancer in 2024.6 The worldwide burden of CRC is projected to rise by 60% by 2030, leading to an estimated 2.2 million new cases and 1.1 million deaths.7
000 individuals are expected to die from colon cancer in 2024.6 The worldwide burden of CRC is projected to rise by 60% by 2030, leading to an estimated 2.2 million new cases and 1.1 million deaths.7
Colon cancer frequently begins with the formation of tiny, benign clusters of cells known as polyps in the colon, which can eventually lead to cancer.8 Age, smoking, inflammatory bowel disease, inheritance, and dietary habits all have a substantial impact on the development of colon cancer.9 Colon cancer displays no visible early signs, and it is typically identified in its middle or late stages. Surgery, chemotherapy, immunotherapy, radiation, and targeted therapies are the most common treatment methods for colon cancer today.10 Among these, chemotherapy plays a crucial role throughout colon cancer treatment, especially for advanced cases.11 FDA-approved frontline chemotherapeutic drugs such as folinic acid, methotrexate, 5-fluorouracil, bevacizumab (avastin), oxaliplatin, and celecoxib (celebrex) are used to treat CRC.12,13 However, these medications have limitations such as adverse side effects, short half-lives, limited tumor selectivity, absorption in the upper GI tract, and drug resistance, which can lead to treatment failure and disease progression.14,15 As a result, there is a pressing need to find innovative treatment options with considerable activity and efficacy against CRC.
Oxadiazoles are novel heterocyclic compounds with substantial applications in synthetic and medicinal chemistry.16 Oxadiazoles have four regioisomers: 1,2,3-, 1,2,4-, 1,2,5-, and 1,3,4.16 Among these region isomers, 1,3,4-oxadiazoles play an important role in medicinal chemistry and display a wide range of biological actions,17 such as anti-tuberculosis,18 anticancer,19 antimalarial,20 antibacterial,21 anticonvulsant, antidiabetic,22 and anti-inflammatory.23 In addition, various medications having a 1,3,4-oxadiazole core are available in the market, including raltegravir (antiretroviral drug), furamizole (antibiotic drug), nesapidil (antiarrhythmic drug), zibotentan (anticancer agent), fenadiazole (hypnotic drug), and tiodazosin (adrenergic receptor antagonist)24 (Fig. 1). In contrast, lonidamine (LND) is an indazole derivative (1-[(2,4-dichlorophenyl) methyl]-1H-indazole-3-carboxylic acid) that was formerly employed as an anti-spermatogenic drug but is now known for its anti-tumor and pro-apoptotic properties.25,26 This drug has undergone pre-clinical testing and has been examined in Phase II and Phase III trials targeting lung cancer, and it has been demonstrated to be safe but with limited efficacy.27 When combined with other pharmacophores or chemotherapeutics, LND demonstrated increased anticancer effects in the treatment of breast, prostate, and ovarian tumors.28,29
With this in mind, and to minimize LND's side effects while improving its pharmacological profile and therapeutic efficacy, we've continued our efforts to develop new anticancer agents.30,31 We developed a rational strategy to replace LND's carboxyl functional group with a 1,3,4-oxadiazole bioisostere. This modification is anticipated to enhance the compound's pharmacokinetic and pharmacodynamic properties while preserving its mechanism of action. We also evaluated the in vitro anticancer activity against HCT116 colon cancer cell lines and conducted in silico analyses, including molecular docking simulations, density functional theory (DFT) assessments, ADME (absorption, distribution, metabolism, and excretion) evaluations, and toxicity studies. This analysis provided insights into how the compounds interact with the active site. We discussed the findings regarding structure–activity relationships and identified the most promising compounds as potential treatment options.
2 Design strategy
LND is a well-known anti-cancer agent that disrupts glycolysis by inhibiting mitochondrial hexokinase, leading to energy depletion in tumor cells. Despite its promising activity, LND has been investigated for the treatment of various cancers such as colon cancer, breast cancer, non-small cell lung cancer (NSCLC), astrocytoma, and squamous cell carcinoma.32 Additionally, 1,3,4-oxadiazoles have demonstrated various mechanisms of action and have been involved in the discovery and development of anticancer drugs.33,34 Despite its promising activity, LND has poor water solubility and limited bioavailability, mainly because of the carboxylic acid (–COOH) group. To overcome these limitations, bioisosteric replacement strategies have been explored. The design strategy involves replacing the amide linker in LND with a 1,3,4-oxadiazole group to enhance metabolic stability and biological activity. This scaffold modification preserves the primary pharmacophoric activity of LND while allowing the insertion of a 1,3,4-oxadiazole at the terminal position. The redesigned scaffold allows for additional customization by selectively modifying substituents (R1, R2, R3) on aryl or heteroaryl rings to change electrical, lipophilic, and steric properties. The rational combination of LND and important 1,3,4-oxadiazole is predicted to improve target selectivity, binding affinity, and anti-proliferative effects via dual or increased interactions with biological targets (Fig. 2).3 Results and discussion
3.1 Synthesis of novel bifunctional compounds for the study
A new series of lonidamine-1,3,4-oxadiazole compounds 7(a–h) were strategically designed and synthesized to enhance the therapeutic potential of the parent compound. The targeted compounds 7(a–h) were synthesized following the strategy outlined in Scheme 1. The synthesis commenced with the preparation of LND (5) from methyl 1H-indazole-3-carboxylate and 2,4-dichloro-1-(chloromethyl) benzene in the presence of potassium carbonate in DMF, using previously published approaches with minor modifications.31 In the next step, a series of structurally diverse hydrazides 6(a–h), including aliphatic, aromatic, and heterocyclic derivatives, were selected for condensation with LND (5). These hydrazides were chosen to introduce electronic and steric diversity into the final molecules, allowing for a comprehensive evaluation of structure–activity relationships (SAR). The reaction was carried out in the presence of phosphoryl chloride under a nitrogen atmosphere, facilitating cyclodehydration to form the desired LND-1,3,4-oxadiazole molecules 7(a–h). This design approach integrates the pharmacophoric properties of LND with the versatility and drug-like nature of the oxadiazole ring, offering a promising platform for the development of new anticancer agents. | ||
| Scheme 1 Synthetic scheme outlining the preparation of target compounds 7(a–h): (i) K2CO3, DMF, rt, 16 h, (ii) NaOH, THF: H2O, 70 °C, 2 h, (iii) POCl3 (neat), reflux. | ||
3.2 Biological studies
Uncontrolled cell proliferation is a hallmark of cancer, leading to the formation of tumor masses through evasion of the programmed cell death mechanism. One promising approach in cancer therapy involves the use of small molecules to inhibit proliferation and impair the migratory and invasive potential of cancer cells, ultimately triggering cell death. In this study, we screened a library of synthesized small molecules for their anticancer activity against HCT116 colorectal cancer cells. We employed the MTT assay, a colorimetric technique that assesses mitochondrial activity to gauge cell viability and metabolic health. Table 1 displays the IC50 values of LND, as a reference, and synthesized compounds 7(a–h). The IC50 value, which indicates the concentration needed to inhibit 50% of cell viability, is a key measure of a compound's potency. Among the tested compounds, compound (7d) demonstrated the strongest anti-proliferative effect, showing an IC50 of 12.91 ± 1.58 µM, which indicates a significant inhibition of cell growth. The compounds (7c) and (7g) had higher potency than other compounds, with IC50 values of 15.43 ± 0.93 and 19.41 ± 0.68 µM, respectively (Table 1). Selectivity index of the screened drug was measured by comparing CC50 of small molecule against normal cells and IC50 against cancer cells (Table 1).| IC50 (µM) | |||
|---|---|---|---|
| Compounds | HCT116 | HEK | Selectivity index |
| 7a | 25.29 ± 1.12 | 58.34 ± 1.78 | 2.31 |
| 7b | 25.95 ± 1.32 | 52.75 ± 1.11 | 2.03 |
| 7c | 15.43 ± 0.93 | 56.21 ± 1.39 | 3.64 |
| 7d | 12.91 ± 1.58 | 72.72 ± 1.69 | 5.63 |
| 7e | 22.23 ± 0.89 | 78.97 ± 1.53 | 3.55 |
| 7f | 29.69 ± 1.95 | 89.81 ± 1.98 | 3.02 |
| 7g | 19.41 ± 0.68 | 64.88 ± 1.27 | 3.34 |
| 7h | 27.93 ± 1.88 | 62.38 ± 1.65 | 2.23 |
| Lonidamine | 88.45 ± 0.55 | 258.11 ± 3.21 | 2.92 |
The trypan blue dye exclusion assay further supported these observations. This assay differentiates the viable and nonviable cells by penetrating into the cytosol of the cells based on the membrane integrity. Compound (7d) induced a concentration-dependent reduction in viable cell number, confirming its cytotoxic effect (Fig. 3B). To further confirm the induction of cell death by compound (7d), Hoechst/PI dual staining was employed. Hoechst stains the nuclei of all cells, while PI selectively enters cells with compromised membranes, typically late apoptotic or necrotic cells. Treatment with compound (7d) led to visible nuclear deformations, membrane rupture, and chromatin condensation, indicating apoptotic cell death (Fig. 3C).
Similarly, acridine orange/PI (AO/PI) dual staining was conducted to distinguish between early apoptotic, late apoptotic, and necrotic cells (Fig. 3D). Acridine orange stains live and early apoptotic cells green, while PI stains late apoptotic stains due to compromised membrane integrity, further supporting the potent anti-proliferative effect of compound (7d) against colorectal cancer cells in a concentration-dependent manner (Fig. 3E and F). Cell migration and cell invasion play crucial roles in the metastasis of cancer cells to form secondary tumors. Interestingly, Compound (7d) mitigates the migration of colorectal cancer cells in a concentration-dependent manner, which was demonstrated by an in vitro migration assay (Fig. 3G). Compound (7d) not only restrains the proliferation of colorectal cancer cells but also hinders the migratory potential of colorectal cancer cells, exhibiting its potent anti-migratory potential and anti-proliferative potential.
3.3 Theoretical studies
| Compounds | Binding affinity (kcal mol−1) | H-bonds (residues) | π–π interactions | Hydrophobic (alkyl/π-alkyl) | Halogen | Other |
|---|---|---|---|---|---|---|
| 7a | −11.2 | TYR143 | HIS121, LEU83 | LEU117, LEU138, VAL118 | Cl–LEU83 | — |
| 7b | −10.4 | THR144 | HIS121, TYR143 | LEU138, ALA140, ILE142, PRO141 | Cl–PRO135 | — |
| 7c | −11.8 | PRO141 | HIS121, TYR143 | LEU138, ALA140, ILE142 | Cl–PRO135 | — |
| 7d | −10.8 | TYR143 | HIS121, LEU83 | LEU117, LEU138, VAL118 | Cl–LEU83 | — |
| 7e | −10.5 | THR144, PRO135 | HIS121, TYR143 | LEU138, ALA140, ILE142, PRO141 | — | — |
| 7f | −11.2 | THR144 | HIS121, TYR143 | LEU138, ALA140, ILE142, PRO141 | Cl–PRO135 | — |
| 7g | −11.8 | THR144, PRO135 | HIS121, TYR143 | LEU138, ALA140, ILE142, PRO141 | Cl–PRO135 | π–sulfur (HIS121) |
| 7h | −10.9 | THR144, PRO135 | HIS121, TYR143 | LEU117, LEU138, ALA140, ILE142 | Cl–PRO135 | Unfavorable bump (THR144) |
| Lonidamine (positive control) | −8.1 | TYR143 (H-bond) | HIS121 (π–π stacked) | LEU83, LEU117, VAL118 (alkyl, π-alkyl) | — | Reference interactions |
3.3.2.1 Root-mean-square deviation. Molecular dynamics (MD) simulations were carried out to evaluate the structural stability and binding behavior of the MMP2–7XJO complex with lonidamine (LND) as the positive control and the screened ligands proposed as colon cancer inhibitors. The stability of each protein–ligand system was assessed using root-mean-square deviation (RMSD) across the simulation trajectory. The MMP2–7XJO–LND complex displayed noticeable fluctuations before reaching a relatively stable regime, with RMSD values stabilizing after ∼50 ns at approximately 0.19–0.28 nm (mean ≈ 0.25 ± 0.03 nm) (Fig. 5). In contrast, the complexes formed with compounds (7c) and (7d) exhibited consistently lower RMSD profiles over the same period, remaining within 0.16–0.26 nm (7c; mean ≈ 0.22 ± 0.03 nm) and 0.16–0.27 nm (7d; mean ≈ 0.20 ± 0.03 nm), indicating reduced conformational drift and improved dynamic stability compared with LND. Conversely, compound (7g) showed higher deviations (approximately 0.22–0.36 nm, mean ≈ 0.29 ± 0.04 nm), suggesting a less stable complex under the simulated conditions, while compound (7a) exhibited larger variability overall despite intermittent low RMSD values (mean ≈ 0.27 ± 0.10 nm). Collectively, these RMSD trends support tighter and more persistent binding for 7d (best), followed by 7c, relative to the reference inhibitor LND, highlighting compound 7d as the most promising candidate for further validation as an MMP2 inhibitor with potential relevance to colon cancer therapy.
3.3.2.2 Root-mean-square fluctuation (RMSF). The RMSF profile indicates that the protein remains largely stable across the trajectory, with flexibility concentrated in two main regions. The dominant dynamic hotspot is the loop spanning GLU69–ASP80 (GLU69, HIS70, GLY71, ASP72, GLY73, TYR74, PRO75, PHE76, ASP77, GLY78, LYS79, ASP80), which shows the highest residue mobility and represents the primary “breathing” segment of the structure. Across the simulated systems, the maximum fluctuations occur within this loop for most conditions: 7g peaks at PHE76 (6.1517 Å), 7c peaks at ASP72 (4.4247 Å), 7a peaks at GLY73 (6.7580 Å), and lonidamine peaks at ASP72 (4.0913 Å), confirming that residues ASP72/GLY73/TYR74/PRO75/PHE76 are repeatedly the most flexible core of this region. In contrast, 7d shows its strongest peak at the C-terminus (ASP168 = 5.5965 Å), with elevated motion also at PRO167, consistent with the expected intrinsic flexibility of terminal residues (SER166–PRO167–ASP168) (Fig. 6). A secondary moderate-mobility site is observed around PRO135–MET139, appearing notably in the lonidamine system (e.g., PRO135), suggesting an additional loop/turn region that may respond to ligand-dependent conformational changes. Overall, the RMSF pattern supports a stable structural core with localized flexibility dominated by the GLU69–ASP80 loop, while terminal residues (166–168) contribute condition-dependent fluctuations, particularly in 7d.
3.3.2.3 Radius of gyration (Rg). The radius of gyration (Rg) was evaluated over 500 ns to assess the global compactness and overall structural stability of the protein in the simulated systems (7a, 7c, 7d, 7g, and lonidamine). Overall, most systems maintained a relatively stable Rg around ∼14.9–15.0 Å, indicating preservation of the global fold with only minor fluctuations. In particular, 7c (14.93 ± 0.10 Å) and 7d (14.92 ± 0.12 Å) exhibited the lowest variability, suggesting that these complexes remain consistently compact throughout the simulation. The lonidamine-bound system also retained a stable compact profile (14.95 ± 0.12 Å), supporting minimal ligand-induced global expansion.
By contrast, the 7a trajectory displayed the most pronounced time-dependent change in compactness (15.15 ± 0.32 Å overall). During the first 0–300 ns, 7a remained comparable to the other systems (14.93 ± 0.09 Å), but a clear transition occurred after 300 ns, where Rg increased to a higher state (15.48 ± 0.25 Å) and remained elevated through the late-stage window (350–500 ns: 15.59 ± 0.16 Å), with occasional excursions reaching 16.16 Å (Fig. 7). This pattern suggests a distinct global expansion event in 7a, consistent with ligand-associated or conformational rearrangements that increase the overall molecular dimensions. The 7g system showed intermediate variability (14.98 ± 0.22 Å) with transient expansions early to mid-trajectory (maximum 15.84 Å), followed by a more compact late-stage behavior (350–500 ns: 14.85 ± 0.16 Å). Taken together, the Rg analysis indicates that the protein remains globally stable and compact in most systems (7c, 7d, and lonidamine), whereas 7a undergoes a marked compactness shift toward a more expanded conformational state, potentially reflecting ligand-dependent opening or broader structural reorganization.
3.3.2.4 Molecular mechanics/generalized Born surface area (MM/GBSA) binding free energy analysis. The results from the MM/GBSA calculations indicate that the protein's binding affinity varies significantly across the different systems. System 7a exhibited the most favorable binding with a total free energy (ΔGtotal) of −33.17 ± 2.81 kcal mol−1, reflecting a stronger binding interaction. This system showed substantial contributions from both the van der Waals (ΔEvdW = −44.12 ± 3.25 kcal mol−1) and electrostatic (ΔEele = −6.89 ± 2.67 kcal mol−1) components, which were balanced by a solvation free energy (ΔGsolv = 17.84 ± 2.10 kcal mol−1) that did not significantly reduce its overall stability. In comparison, 7c demonstrated weaker binding (ΔGtotal = −27.57 ± 4.88 kcal mol−1) with more significant contributions from solvation (ΔGsolv = 15.96 ± 5.97 kcal mol−1), suggesting that while the van der Waals and electrostatic contributions (ΔEvdW = −35.29 ± 5.60 kcal mol−1, ΔEele = −8.24 ± 5.06 kcal mol−1) were favorable, the solvation free energy did not contribute sufficiently to stabilize the protein–ligand complex. Similarly, 7d showed a balanced energy distribution (ΔGtotal = −30.33 ± 2.43 kcal mol−1), with moderate contributions from van der Waals and electrostatic energies, although the solvation energy (ΔGsolv = 18.12 ± 2.71 kcal mol−1) was slightly higher compared to 7a. The 7g system (ΔGtotal = −32.95 ± 3.02 kcal mol−1) exhibited a similar trend with substantial van der Waals and electrostatic energy contributions, but with a less favorable solvation energy (ΔGsolv = 17.99 ± 2.70 kcal mol−1) (Table 3). This suggests that the system may exhibit slightly weaker binding than 7A, but still maintains a strong affinity. Finally, the lonidamine-bound system showed the highest solvation energy (ΔGsolv = 26.46 ± 3.15 kcal mol−1), but its overall binding affinity was still relatively strong (ΔGtotal = −33.22 ± 2.90 kcal mol−1), which indicates that the ligand's influence on the protein–ligand binding dynamics is substantial despite the high solvation contributions.
| Systems | ΔEvdW | ΔEele | ΔEGB | ΔEsurf | ΔGgas | ΔGsolv | ΔGtotal |
|---|---|---|---|---|---|---|---|
| a ΔEvdW: van der Waals energy; ΔEele: electrostatic energy; ΔGGB: solvation-free energy change part based on the generalized Born model; ΔEsurf: free energy change contributed by the surface area; ΔGgas: change in free energy in the gas phase; ΔGsolv: solvation free energy change; ΔGbinding: total free energy change. Relationship: ΔGtotal = ΔGgas + ΔGsolv = ΔEvdW + ΔEele + ΔEGB + ΔEsurf. | |||||||
| 7a | −44.12 ± 3.25 | −6.89 ± 2.67 | 20.97 ± 2.13 | −3.13 ± 0.22 | −51.01 ± 3.37 | 17.84 ± 2.10 | −33.17 ± 2.81 |
| 7c | −35.29 ± 5.60 | −8.24 ± 5.06 | 18.93 ± 6.28 | −2.97 ± 0.41 | −43.53 ± 9.66 | 15.96 ± 5.97 | −27.57 ± 4.88 |
| 7d | −44.72 ± 2.86 | −3.73 ± 3.41 | 21.53 ± 2.72 | −3.41 ± 0.16 | −48.45 ± 3.67 | 18.12 ± 2.71 | −30.33 ± 2.43 |
| 7g | −43.68 ± 3.11 | −7.26 ± 2.89 | 21.08 ± 2.82 | −3.10 ± 0.27 | −50.93 ± 4.11 | 17.99 ± 2.70 | −32.95 ± 3.02 |
| Lonidamine | −42.01 ± 2.63 | −17.68 ± 3.65 | 29.37 ± 3.24 | −2.90 ± 0.17 | −59.68 ± 4.85 | 26.46 ± 3.15 | −33.22 ± 2.90 |
3.3.3.1 2D projection of trajectories. Principal component analysis (PCA) of the MD trajectories (50
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames) demonstrated ligand-dependent reshaping of the sampled conformational landscape. The apo system exhibited the most compact distribution in the PC1–PC2 subspace (total variance, Var(PC1) + Var(PC2) = 94.120), whereas ligand binding increased dispersion to varying extents. Among the complexes, 7a showed the broadest sampling (total variance = 448.276; PC1 range −28.041 to 44.659; PC2 range −25.760 to 33.218), followed by 7g (total variance = 316.281), indicating substantially enhanced conformational heterogeneity relative to apo. Compounds 7c and 7d occupied intermediate conformational space (157.984 and 131.239, respectively), while lonidamine displayed a comparatively confined distribution (116.614). A two-center separation check further supported distinct population structure for 7a and 7g (center distances 37.77 and 31.74), consistent with sampling of multiple dominant conformational regions, whereas 7d showed the weakest separation (13.14), indicating more limited partitioning within the reduced space (Fig. 8).
000 frames) demonstrated ligand-dependent reshaping of the sampled conformational landscape. The apo system exhibited the most compact distribution in the PC1–PC2 subspace (total variance, Var(PC1) + Var(PC2) = 94.120), whereas ligand binding increased dispersion to varying extents. Among the complexes, 7a showed the broadest sampling (total variance = 448.276; PC1 range −28.041 to 44.659; PC2 range −25.760 to 33.218), followed by 7g (total variance = 316.281), indicating substantially enhanced conformational heterogeneity relative to apo. Compounds 7c and 7d occupied intermediate conformational space (157.984 and 131.239, respectively), while lonidamine displayed a comparatively confined distribution (116.614). A two-center separation check further supported distinct population structure for 7a and 7g (center distances 37.77 and 31.74), consistent with sampling of multiple dominant conformational regions, whereas 7d showed the weakest separation (13.14), indicating more limited partitioning within the reduced space (Fig. 8).
3.3.3.2 Free energy landscape (FEL). The FEL projected onto PC1–PC2 revealed ligand-dependent reshaping of the conformational basins. The apo system displayed multiple low-energy minima, indicating the presence of more than one metastable conformation, although these states were confined within a comparatively bounded region of PC space. In contrast, 7a produced the most expanded and distinctly partitioned landscape, with well-separated low-energy basins across negative and positive PC1, supporting pronounced multi-state behavior. Compound 7g similarly exhibited several low-energy minima distributed across PC space, consistent with dynamic sampling of multiple substates and potential interconversion along connecting corridors. Compounds 7c and 7d showed intermediate landscapes with two to several dominant basins, suggesting moderate conformational heterogeneity under ligand binding. lonidamine displayed a dominant low-energy basin accompanied by secondary minima, indicating stabilization of a preferred conformational state while retaining access to alternate substrates (Fig. 9).
3.4 In silico evaluation of physicochemical and ADME properties
A series of compounds 7(a–h) were evaluated based on their physicochemical properties, pharmacokinetic profiles, drug-likeness, and synthetic accessibility to assess their potential as orally active drug candidates using SwissADME.35 The key parameters analyzed included hydrogen bond acceptors (HBA), hydrogen bond donors (HDA), topological polar surface area (TPSA), lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), aqueous solubility (log
P), aqueous solubility (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S), gastrointestinal absorption (GIA), skin permeability (log
S), gastrointestinal absorption (GIA), skin permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp), Lipinski's rule compliance, and synthetic accessibility (SA) (Table 4). All of these compounds fulfilled Lipinski's rule of five, suggesting excellent drug-likeness. Notably, compound (7h) emerged as the most promising option, with no violations of Lipinski's rule,36 strong gastrointestinal absorption, and a modest log
Kp), Lipinski's rule compliance, and synthetic accessibility (SA) (Table 4). All of these compounds fulfilled Lipinski's rule of five, suggesting excellent drug-likeness. Notably, compound (7h) emerged as the most promising option, with no violations of Lipinski's rule,36 strong gastrointestinal absorption, and a modest log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P (5.09), indicating a desirable balance of hydrophilicity and lipophilicity. In contrast, compound (7f) had the highest lipophilicity (log
P (5.09), indicating a desirable balance of hydrophilicity and lipophilicity. In contrast, compound (7f) had the highest lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 6.11) and the lowest water solubility (log
P = 6.11) and the lowest water solubility (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S = −7.48), which corresponded to its low anticipated oral absorption. Despite complying with Lipinski's requirements, this makes it a less favorable candidate. All compounds have TPSA values ≤ 140 Å2, indicating acceptable membrane permeability. Compound (7g) exhibited the largest TPSA (84.98 Å2), which may affect its permeability, although it maintained high GIA.37 The synthetic accessibility (SA) scores varied from 3.17 to 3.37 across the series, indicating that all compounds are reasonably easy to synthesis, with compounds (7a) and (7e) having slightly lower SA values (3.17), indicating a more favorable synthetic profile.38 In terms of skin permeability, all compounds had relatively low log
S = −7.48), which corresponded to its low anticipated oral absorption. Despite complying with Lipinski's requirements, this makes it a less favorable candidate. All compounds have TPSA values ≤ 140 Å2, indicating acceptable membrane permeability. Compound (7g) exhibited the largest TPSA (84.98 Å2), which may affect its permeability, although it maintained high GIA.37 The synthetic accessibility (SA) scores varied from 3.17 to 3.37 across the series, indicating that all compounds are reasonably easy to synthesis, with compounds (7a) and (7e) having slightly lower SA values (3.17), indicating a more favorable synthetic profile.38 In terms of skin permeability, all compounds had relatively low log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp values (−4.65 to −5.72), which is typical for drug-like molecules and not a limiting factor for drug development. Overall, compound (7h) has the most balanced and positive profile across all parameters tested, making it a promising candidate for additional preclinical testing.
Kp values (−4.65 to −5.72), which is typical for drug-like molecules and not a limiting factor for drug development. Overall, compound (7h) has the most balanced and positive profile across all parameters tested, making it a promising candidate for additional preclinical testing.
| Comp. | Physiochemical properties | Pharmacokinetics | Drug likeness | Medicinal chemistry | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| HBA | HDA | TPSA | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Po/w Po/w |
Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) S S |
GIA | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Kp Kp |
Lipinski | Vio | SA | |
| 7a | 4 | 0 | 56.74 | 5.11 | −6.56 | High | −4.89 | Yes | 1 | 3.17 |
| 7b | 4 | 0 | 56.74 | 5.51 | −6.94 | High | −5.72 | Yes | 1 | 3.31 |
| 7c | 5 | 0 | 65.97 | 5.14 | −6.73 | High | −5.09 | Yes | 1 | 3.29 |
| 7d | 4 | 0 | 56.74 | 5.66 | −7.22 | High | −4.65 | Yes | 1 | 3.18 |
| 7e | 5 | 0 | 56.74 | 5.40 | −6.67 | High | −4.93 | Yes | 1 | 3.18 |
| 7f | 7 | 0 | 56.74 | 6.11 | −7.48 | Low | −4.68 | Yes | 1 | 3.32 |
| 7g | 4 | 0 | 84.98 | 5.10 | −6.87 | High | −5.12 | Yes | 1 | 3.24 |
| 7h | 4 | 0 | 56.74 | 5.09 | −6.56 | High | −4.84 | Yes | 0 | 3.37 |
Bioavailability radar plots displaying six essential physicochemical parameters related to oral bioavailability: lipophilicity (LIPO), size, polarity (POLAR), solubility (INSOLU), saturation (INSATU), and molecular flexibility (FLEX). The pink region shows the ideal drug-likeness range. The radar plot analysis offers a clear visual comparison of the biological and physicochemical profiles of LND and its analogues. LND has a balanced but moderate profile across most evaluated parameters, which reflects its established but limited activity. The analogues show clear differences, highlighting how structural changes impact their properties. Among the analogues, compound (7d) stands out with the best profile. It has higher scores across several descriptors, suggesting it may be more potent and drug-like. Compound (7c) shows moderate improvements over LND but is still less effective than (7d). Compound (7g) presents a different pattern, demonstrating strong activity in some areas while performing weaker in others, indicating potential selectivity. The other analogues show varying trends, suggesting that adjusting substituents can lead to different activity and toxicity outcomes. In summary, the radar analysis indicates that substitution patterns significantly affect the physicochemical and biological behavior of the oxadiazole derivatives. Compound (7d) is the most promising analogue when compared to LND (Fig. 10).
The BOILED-Egg diagram provides a visual prediction of the gastrointestinal absorption (HIA) and blood–brain barrier (BBB) penetration for compounds 7(a–h) and LND.39 In this plot, the white region (egg white) represents compounds likely to be absorbed in the gastrointestinal tract, while the yellow region (yolk) indicates the potential to cross the blood–brain barrier. Compounds 7(a–h) are expected to have high gastrointestinal absorption, indicated by the white ellipse. However, they stay outside the BBB penetration zone, marked by the yellow region, which means there is limited exposure to the central nervous system (CNS). Notably, compound (7d), the most active in the series, is well within the optimal HIA region and outside the BBB zone. This suggests it has good oral absorption and a lower chance of causing side effects in the CNS (Fig. 11).
 | ||
| Fig. 11 The BOILED-Egg diagram illustrates the predicted gastrointestinal absorption (HIA) and blood–brain barrier (BBB) penetration of lonidamine and 7(a–h). | ||
3.5 Toxicity
A theoretical analysis of the toxicity of the LND derivatives 7(a–h), alongside the LND, was conducted using Protox-II,40 with the results presented in Table 5. The LD50 values show that all tested compounds fall within toxicity class 4, which means they are harmful if swallowed, according to the GHS classification. Most compounds show an inactive profile for hepatotoxicity, carcinogenicity, mutagenicity, and cytotoxicity. Only a few predictions of immunotoxicity or hepatotoxicity are noted, such as for (7c) and (7e). Notably, compound (7d), the most active of the sequence, has a completely inert toxicity profile across all endpoints. This provides a sufficient safety margin for future development.| Comp. code | Predicted LD50 (mg kg−1) | Predicted toxicity class | Hepatotoxicity | Carcinogenicity | Immunotoxicity | Mutagenicity | Cytotoxicity |
|---|---|---|---|---|---|---|---|
| 5 | 900 | 4 | Active (0.53) | Inactive (0.84) | Inactive (0.98) | Inactive (0.86) | Inactive (0.67) |
| 7a | 1140 | 4 | Inactive (0.58) | Inactive (0.65) | Inactive (0.96) | Inactive (0.68) | Inactive (0.78) |
| 7b | 1140 | 4 | Inactive (0.57) | Inactive (0.66) | Inactive (0.97) | Inactive (0.68) | Inactive (0.76) |
| 7c | 1140 | 4 | Active (0.54) | Inactive (0.66) | Active (0.90) | Inactive (0.63) | Inactive (0.70) |
| 7d | 1140 | 4 | Inactive (0.58) | Inactive (0.65) | Inactive (0.91) | Inactive (0.68) | Inactive (0.78) |
| 7e | 1140 | 4 | Active (0.57) | Inactive (0.65) | Active (0.63) | Inactive (0.66) | Inactive (0.78) |
| 7f | 1140 | 4 | Inactive (0.54) | Inactive (0.63) | Inactive (0.85) | Inactive (0.64) | Inactive (0.76) |
| 7g | 1140 | 4 | Inactive (0.54) | Inactive (0.65) | Inactive (0.98) | Inactive (0.62) | Inactive (0.74) |
| 7h | 1140 | 4 | Inactive (0.53) | Inactive (0.64) | Inactive (0.94) | Inactive (0.62) | Inactive (0.76) |
The toxicity prediction models for compounds (7c), (7d), and (7g) revealed significant variances in toxicity profiles. Compound (7c) had probable hepatotoxicity (probability 0.54) and immunotoxicity (0.90), but it was neither carcinogenic, mutagenic, or cytotoxic. It also confirmed blood–brain barrier (BBB) permeability with high confidence (0.90). In contrast, compound (7d) demonstrated a superior profile, being inactive for all examined endpoints except BBB permeability (0.95), indicating that it is safer than (7c) (Table 6 and Fig. 12). Also, the compounds (7c), (7d), and (7g) shows clear safety trends within the group. Compound (7c) is non-carcinogenic, non-mutagenic, and non-cytotoxic. However, it has potential liver toxicity and significant immune-related risks, which may limit how it can be used therapeutically. On the other hand, compound (7d) has a better profile. It shows inactivity in all major toxicity areas and has strong permeability through the blood–brain barrier, making it suitable for targeting the central nervous system. Among the three, compound (7g) has the most appealing profile, with all toxicity end points predicted to be inactive, including immune toxicity, and a high chance of penetrating the blood–brain barrier. Overall, these findings suggest that (7g) is the safest and most promising candidate. Compound (7d) also has positive qualities, while (7c) raises concerns due to its immune and liver risks.
| Classification | Target | Shorthand | Prediction | Probability | ||||
|---|---|---|---|---|---|---|---|---|
| 7c | 7d | 7g | 7c | 7d | 7g | |||
| Organ toxicity | Hepatotoxicity | Dili | Active | Inactive | Inactive | 0.54 | 0.58 | 0.54 |
| Toxicity end points | Carcinogenicity | Carcino | Inactive | Inactive | Inactive | 0.66 | 0.65 | 0.65 |
| Toxicity end points | Immunotoxicity | Immuno | Active | Inactive | Inactive | 0.90 | 0.91 | 0.98 |
| Toxicity end points | Mutagenicity | Mutagen | Inactive | Inactive | Inactive | 0.63 | 0.68 | 0.62 |
| Toxicity end points | Cytotoxicity | Cyto | Inactive | Inactive | Inactive | 0.70 | 0.78 | 0.74 |
| Toxicity end points | BBB-barrier | BBB | Active | Active | Active | 0.90 | 0.95 | 0.94 |
3.6 Plausible structural–activity relationship (SAR) study
The SAR is based on the synthesized compounds indicated in Scheme 1 and their correlation with biological activity (Table 1). The cytotoxic effect study provided vital insights into the SAR that governs their anticancer efficacy. Compound (7d) is the most active analogue of the LND-1,3,4-oxadiazole family, and its potency stems from three structural properties. The terminal 2,4-dichlorophenyl group enhances lipophilicity and hydrophobic fit inside the binding pocket. This corresponds to the peak activity recorded. The distal 4-chlorophenyl ring enhances hydrophobic contacts and alters electronic properties, promoting π-stacking and target engagement. The central oxadiazole linker is important because it keeps the two aryl rings aligned and offers hydrogen-bond acceptor capability. All of these features work together to provide an ideal shape and polarity for high binding affinity. This explains why (7d) exhibits more activity than its other counterparts. However, while the numerous-chloro substituents improve potency, they may have a deleterious impact on solubility and metabolic clearance. This emphasizes the importance of careful optimization in achieving a balance between efficacy and drug-like features (Fig. 13).4 Conclusion
This study focuses on the design and synthesis of a series of lonidamine and 1,3,4-oxadiazole derivatives by substituting the amide linker with a 1,3,4-oxadiazole scaffold. The compounds were evaluated for their ability to inhibit the growth of the HCT116 colorectal cancer cells. Among the screened compounds, compound (7d) exhibited significant anticancer activity, showing the lowest IC50 value of 12.91 ± 1.58 µM, in contrast to the parent compound LND, which had an IC50 of 88.45 ± 0.55 µM. The induction of apoptosis by compound (7d) against HCT116 confirms its role as a potent anticancer agent. Fluorescence microscopic studies showed that the compound induces cell death through apoptosis, not via necrosis. Additional computational studies, including molecular docking, molecular dynamics simulations, and DFT analysis, provided insights into how these compounds interact with the target protein, supporting their biological activity. All designed ligands 7(a–h) showed stronger binding affinities (−10.4 to −11.8 kcal mol−1) than the positive control LND (−8.1 kcal mol−1). Compounds (7c) and (7g) proved to be the most potent, exhibiting various stabilizing interactions. DFT analysis provided insights into the chemical reactivity and binding potential of the compounds, while molecular electrostatic potential maps highlighted favorable interaction sites. Furthermore, ADME profiling and toxicity predictions indicated excellent pharmacokinetic properties and acceptable safety margins, suggesting these molecules as promising drug candidates. Structure–activity relationship analysis identified key structural features necessary for activity, providing useful guidance for designing improved analogs. Overall, this research identifies compound (7d) as a promising lead for further optimization and preclinical testing in the treatment of colorectal cancer. Future studies (in vivo) aimed at enhancing the cytotoxicity of these compounds will be published later.5 Experimental
5.1 Chemistry
5.1.3.1 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-phenyl-1,3,4-oxadiazole (7a).
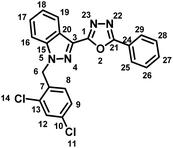 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (lonidamine) (5) and benzohydrazide (6a) afforded the pure title product as white solid. Yield; 32%; m.p 191–193 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.33 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.17–8.09 (m, 2H), 7.91 (d, J = 8.7 Hz, 1H), 7.73–7.59 (m, 5H), 7.48 (t, J = 7.2 Hz, 1H), 7.40 (dd, J = 2.2, 8.4 Hz, 1H), 7.03 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6). 13C NMR (101 MHz, DMSO-d6) δ = 163.46, 159.60, 140.91, 133.45, 133.24, 133.19, 132.14, 130.93, 129.62, 129.52, 129.13, 127.94, 127.83, 126.77, 123.51, 123.15, 121.51, 121.12, 110.91, 49.90. MS-ESI (m/z): 420.90 (M + 1)+; HPLC: 92.19%; RT: 5.942 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (lonidamine) (5) and benzohydrazide (6a) afforded the pure title product as white solid. Yield; 32%; m.p 191–193 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.33 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.17–8.09 (m, 2H), 7.91 (d, J = 8.7 Hz, 1H), 7.73–7.59 (m, 5H), 7.48 (t, J = 7.2 Hz, 1H), 7.40 (dd, J = 2.2, 8.4 Hz, 1H), 7.03 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6). 13C NMR (101 MHz, DMSO-d6) δ = 163.46, 159.60, 140.91, 133.45, 133.24, 133.19, 132.14, 130.93, 129.62, 129.52, 129.13, 127.94, 127.83, 126.77, 123.51, 123.15, 121.51, 121.12, 110.91, 49.90. MS-ESI (m/z): 420.90 (M + 1)+; HPLC: 92.19%; RT: 5.942 min.
5.1.3.2 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-(p-tolyl)-1,3,4-oxadiazole (7b).
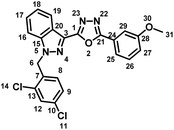 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 4-methylbenzohydrazide (6b) afforded the pure title product as an off-white solid. Yield: 43%; m.p.186–187 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.32 (d, J = 8.1 Hz, 1H, indazole-H-16, 19), 8.01 (d, J = 8.2 Hz, 2H), 7.90 (d, J = 8.7 Hz, 1H), 7.72 (d, J = 2.2 Hz, 1H), 7.60 (t, J = 7.7 Hz, 1H), 7.49–7.43 (m, 3H), 7.40 (dd, J = 2.2, 8.4 Hz, 1H), 7.02 (d, J = 8.4 Hz, 1H, H-8), 5.94 (s, 2H, –CH2–, H-6), 2.45–2.40 (m, 3H, –CH3, H-30). 13C NMR (101 MHz, DMSO-d6) δ = 164.04, 159.85, 142.83, 141.40, 133.94, 133.70, 131.38, 130.55, 130.17, 129.62, 128.41, 128.33, 127.22, 123.96, 121.98, 121.62, 120.88, 111.37, 50.38, 21.67. MS-ESI (m/z): 435.0 (M + 1)+; HPLC: 98.28%; RT: 6.076 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 4-methylbenzohydrazide (6b) afforded the pure title product as an off-white solid. Yield: 43%; m.p.186–187 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.32 (d, J = 8.1 Hz, 1H, indazole-H-16, 19), 8.01 (d, J = 8.2 Hz, 2H), 7.90 (d, J = 8.7 Hz, 1H), 7.72 (d, J = 2.2 Hz, 1H), 7.60 (t, J = 7.7 Hz, 1H), 7.49–7.43 (m, 3H), 7.40 (dd, J = 2.2, 8.4 Hz, 1H), 7.02 (d, J = 8.4 Hz, 1H, H-8), 5.94 (s, 2H, –CH2–, H-6), 2.45–2.40 (m, 3H, –CH3, H-30). 13C NMR (101 MHz, DMSO-d6) δ = 164.04, 159.85, 142.83, 141.40, 133.94, 133.70, 131.38, 130.55, 130.17, 129.62, 128.41, 128.33, 127.22, 123.96, 121.98, 121.62, 120.88, 111.37, 50.38, 21.67. MS-ESI (m/z): 435.0 (M + 1)+; HPLC: 98.28%; RT: 6.076 min.
5.1.3.3 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-(3-methoxyphenyl)-1,3,4-oxadiazole (7c).
 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6c) afforded the pure title product as an off white solid. Yield: 63%; m.p.176–178 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.18 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 7.98–7.86 (m, 3H), 7.72 (d, J = 2.1 Hz, 1H), 7.70–7.63 (m, 2H), 7.58 (t, J = 7.6 Hz, 1H), 7.50–7.36 (m, 2H), 7.08 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6), 3.88 (s, 3H, –OCH3, H-31). 13C NMR (101 MHz, DMSO-d6) δ = 165.42, 160.03, 150.21, 141.02, 133.81, 133.57, 133.37, 132.91, 131.22, 131.15, 130.23, 129.14, 127.82, 127.50, 127.04, 126.82, 123.95, 123.16, 122.65, 121.33, 111.08, 52.42, 50.11. MS-ESI (m/z): 452.0 (M + 2)+; HPLC: 98.33%; RT: 6.694 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6c) afforded the pure title product as an off white solid. Yield: 63%; m.p.176–178 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.18 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 7.98–7.86 (m, 3H), 7.72 (d, J = 2.1 Hz, 1H), 7.70–7.63 (m, 2H), 7.58 (t, J = 7.6 Hz, 1H), 7.50–7.36 (m, 2H), 7.08 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6), 3.88 (s, 3H, –OCH3, H-31). 13C NMR (101 MHz, DMSO-d6) δ = 165.42, 160.03, 150.21, 141.02, 133.81, 133.57, 133.37, 132.91, 131.22, 131.15, 130.23, 129.14, 127.82, 127.50, 127.04, 126.82, 123.95, 123.16, 122.65, 121.33, 111.08, 52.42, 50.11. MS-ESI (m/z): 452.0 (M + 2)+; HPLC: 98.33%; RT: 6.694 min.
5.1.3.4 2-(4-Chlorophenyl)-5-(1-(2,4-dichlorobenzyl)-1H-indazol-3-yl)-1,3,4-oxadiazole (7d).
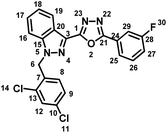 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6d) afforded the pure title product as an off white solid. Yield: 51%; m.p.167–168 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.32 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.13 (d, J = 8.7 Hz, 2H), 7.91 (d, J = 8.7 Hz, 1H), 7.76–7.67 (m, 3H), 7.61 (t, J = 7.5 Hz, 1H), 7.50–7.43 (m, 1H), 7.40 (dd, J = 2.1, 8.4 Hz, 1H), 7.04 (d, J = 8.4 Hz, 1H, H-8), 5.94 (s, 2H, –CH2–, H-6). 13C NMR (101 MHz, DMSO-d6) δ = 162.73, 159.73, 140.91, 136.83, 133.48, 133.27, 133.15, 130.99, 129.67, 129.51, 129.14, 128.59, 127.96, 127.83, 123.55, 122.06, 121.50, 121.09, 110.94, 49.92. MS-ESI (m/z): 454.85 (M + 1)+; HPLC: 94.89%; RT: 8.585 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6d) afforded the pure title product as an off white solid. Yield: 51%; m.p.167–168 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.32 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.13 (d, J = 8.7 Hz, 2H), 7.91 (d, J = 8.7 Hz, 1H), 7.76–7.67 (m, 3H), 7.61 (t, J = 7.5 Hz, 1H), 7.50–7.43 (m, 1H), 7.40 (dd, J = 2.1, 8.4 Hz, 1H), 7.04 (d, J = 8.4 Hz, 1H, H-8), 5.94 (s, 2H, –CH2–, H-6). 13C NMR (101 MHz, DMSO-d6) δ = 162.73, 159.73, 140.91, 136.83, 133.48, 133.27, 133.15, 130.99, 129.67, 129.51, 129.14, 128.59, 127.96, 127.83, 123.55, 122.06, 121.50, 121.09, 110.94, 49.92. MS-ESI (m/z): 454.85 (M + 1)+; HPLC: 94.89%; RT: 8.585 min.
5.1.3.5 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-(3-fluorophenyl)-1,3,4-oxadiazole (7e).
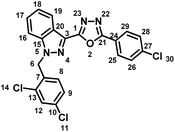 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6e) afforded the pure title product as an off-white solid. Yield: 44%; m.p.162–164 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.34 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.01–7.85 (m, 2H), 7.76–7.68 (m, 2H), 7.62 (br t, J = 7.8 Hz, 1H), 7.57–7.32 (m, 4H), 7.03 (d, J = 8.5 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6); 19F NMR (376 MHz, DMSO-d6) δ = −111.23 (s, 1F); 13C NMR (101 MHz, DMSO-d6) δ = 183.61, 161.22 (d, J = 273.2 Hz, 1C), 140.92, 133.48, 133.25, 133.15, 131.94 (br d, J = 8.0 Hz, 1C), 130.94, 129.47, 129.14, 127.91 (br d, J = 13.1 Hz, 1C), 127.19, 125.69 (br d, J = 90.8 Hz, 1C), 123.59, 123.12 (d, J = 2.9 Hz, 1C), 121.53, 121.13, 119.24, 119.03, 113.51 (br d, J = 23.3 Hz, 1C), 110.94, 49.94. MS-ESI (m/z): 438.7 (M + 1)+; HPLC: RT: 96.09%; RT: 5.881 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and 3-methoxybenzohydrazide (6e) afforded the pure title product as an off-white solid. Yield: 44%; m.p.162–164 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.34 (d, J = 8.2 Hz, 1H, indazole-H-16, 19), 8.01–7.85 (m, 2H), 7.76–7.68 (m, 2H), 7.62 (br t, J = 7.8 Hz, 1H), 7.57–7.32 (m, 4H), 7.03 (d, J = 8.5 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6); 19F NMR (376 MHz, DMSO-d6) δ = −111.23 (s, 1F); 13C NMR (101 MHz, DMSO-d6) δ = 183.61, 161.22 (d, J = 273.2 Hz, 1C), 140.92, 133.48, 133.25, 133.15, 131.94 (br d, J = 8.0 Hz, 1C), 130.94, 129.47, 129.14, 127.91 (br d, J = 13.1 Hz, 1C), 127.19, 125.69 (br d, J = 90.8 Hz, 1C), 123.59, 123.12 (d, J = 2.9 Hz, 1C), 121.53, 121.13, 119.24, 119.03, 113.51 (br d, J = 23.3 Hz, 1C), 110.94, 49.94. MS-ESI (m/z): 438.7 (M + 1)+; HPLC: RT: 96.09%; RT: 5.881 min.
5.1.3.6 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-(4-(trifluoromethyl)phenyl)-1,3,4-oxadiazole (7f).
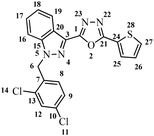 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid and 4-trifluoromethyl benzohydrazide (6f) afforded the pure title product a white solid. Yield: 28%; m.p.196–198 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.33 (d, J = 8.2 Hz, 3H), 8.01 (d, J = 8.4 Hz, 2H), 7.93 (d, J = 8.7 Hz, 1H), 7.73 (d, J = 2.2 Hz, 1H), 7.62 (t, J = 7.8 Hz, 1H), 7.48 (t, J = 7.5 Hz, 1H), 7.41 (dd, J = 2.2, 8.4 Hz, 1H), 7.05 (d, J = 8.4 Hz, 1H, H-8), 5.96 (s, 2H, –CH2–, H-6), 19F NMR (376 MHz, DMSO-d6) δ = −61.59 (s, 1F)·13C NMR (101 MHz, DMSO-d6) δ = 162.46, 160.09, 140.94, 133.49, 133.28, 133.14, 131.74, 131.43, 130.99, 129.43, 129.15, 127.91 (br d, J = 16.0 Hz, 1C), 127.66, 126.99, 126.45 (br d, J = 3.6 Hz, 1C), 123.62, 121.55, 121.08, 110.98, 49.96. MS-ESI (m/z): 488.9 (M + 1)+; HPLC: 88%; RT: 6.177 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid and 4-trifluoromethyl benzohydrazide (6f) afforded the pure title product a white solid. Yield: 28%; m.p.196–198 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.33 (d, J = 8.2 Hz, 3H), 8.01 (d, J = 8.4 Hz, 2H), 7.93 (d, J = 8.7 Hz, 1H), 7.73 (d, J = 2.2 Hz, 1H), 7.62 (t, J = 7.8 Hz, 1H), 7.48 (t, J = 7.5 Hz, 1H), 7.41 (dd, J = 2.2, 8.4 Hz, 1H), 7.05 (d, J = 8.4 Hz, 1H, H-8), 5.96 (s, 2H, –CH2–, H-6), 19F NMR (376 MHz, DMSO-d6) δ = −61.59 (s, 1F)·13C NMR (101 MHz, DMSO-d6) δ = 162.46, 160.09, 140.94, 133.49, 133.28, 133.14, 131.74, 131.43, 130.99, 129.43, 129.15, 127.91 (br d, J = 16.0 Hz, 1C), 127.66, 126.99, 126.45 (br d, J = 3.6 Hz, 1C), 123.62, 121.55, 121.08, 110.98, 49.96. MS-ESI (m/z): 488.9 (M + 1)+; HPLC: 88%; RT: 6.177 min.
5.1.3.7 2-(1-(2,4-Dichlorobenzyl)-1H-indazol-3-yl)-5-(thiophen-2-yl)-1,3,4-oxadiazole (7g).
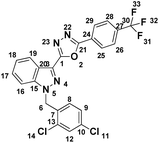 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and thiophene 2-carbohydrazide (6g) afforded the pure title product a white solid. Yield: 35%; m.p.173–174 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.30 (d, J = 8.1 Hz, 1H), 8.00 (dd, J = 1.2, 5.0 Hz, 1H), 7.96 (dd, J = 1.2, 3.7 Hz, 1H), 7.90 (d, J = 8.5 Hz, 1H), 7.72 (d, J = 2.1 Hz, 1H), 7.61 (ddd, J = 1.0, 7.1, 8.4 Hz, 1H), 7.46 (t, J = 7.3 Hz, 1H), 7.39 (dd, J = 2.2, 8.4 Hz, 1H), 7.33 (dd, J = 3.7, 4.9 Hz, 1H), 7.01 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6); 13C NMR (101 MHz, DMSO-d6) δ = 159.84, 159.00, 140.93, 133.48, 133.25, 133.20, 130.91, 130.73, 129.45, 129.16, 128.94, 128.00, 127.85, 124.04, 123.57, 121.52, 121.13, 110.94, 104.42, 49.94. MS-ESI (m/z): 426.85 (M + 1)+; HPLC: 94.9%; RT: 6.728 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and thiophene 2-carbohydrazide (6g) afforded the pure title product a white solid. Yield: 35%; m.p.173–174 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.30 (d, J = 8.1 Hz, 1H), 8.00 (dd, J = 1.2, 5.0 Hz, 1H), 7.96 (dd, J = 1.2, 3.7 Hz, 1H), 7.90 (d, J = 8.5 Hz, 1H), 7.72 (d, J = 2.1 Hz, 1H), 7.61 (ddd, J = 1.0, 7.1, 8.4 Hz, 1H), 7.46 (t, J = 7.3 Hz, 1H), 7.39 (dd, J = 2.2, 8.4 Hz, 1H), 7.33 (dd, J = 3.7, 4.9 Hz, 1H), 7.01 (d, J = 8.4 Hz, 1H, H-8), 5.95 (s, 2H, –CH2–, H-6); 13C NMR (101 MHz, DMSO-d6) δ = 159.84, 159.00, 140.93, 133.48, 133.25, 133.20, 130.91, 130.73, 129.45, 129.16, 128.94, 128.00, 127.85, 124.04, 123.57, 121.52, 121.13, 110.94, 104.42, 49.94. MS-ESI (m/z): 426.85 (M + 1)+; HPLC: 94.9%; RT: 6.728 min.
5.1.3.8 2-Cyclopentyl-5-(1-(2,4-dichlorobenzyl)-1H-indazol-3-yl)-1,3,4-oxadiazole (7h).
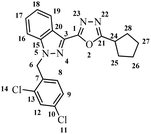 (2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and cyclopentane carbohydrazide (6h) afforded the pure title product a white solid. Yield: 37%; m.p.180–182 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.22 (d, J = 8.2 Hz, 1H), 7.87 (d, J = 8.7 Hz, 1H), 7.75–7.67 (m, 1H), 7.63–7.53 (m, 1H), 7.48–7.36 (m, 2H), 6.98 (d, J = 8.4 Hz, 1H, H-8), 5.90 (s, 2H, –CH2–, H-6), 3.56–3.39 (m, 1H, –CH–, H-24), 2.16–2.07 (m, 2H, –CH2–, H-25), 2.02–1.85 (m, 2H, –CH2–, H-28), 1.80–1.63 (m, 4H, –CH2–, H-26, 27). 13C NMR (101 MHz, DMSO-d6) δ = 169.26, 159.42, 140.85, 133.42, 133.22, 130.90, 129.84, 129.11, 127.84, 127.84, 123.32, 121.31, 121.05, 110.82, 49.80, 35.12, 30.55, 25.07. MS-ESI (m/z): 412.8 (M + 1)+; HPLC: 92.73%; RT: 6.964 min.
(2,4-Dichlorobenzyl)-1H-indazole-3-carboxylic acid (5) and cyclopentane carbohydrazide (6h) afforded the pure title product a white solid. Yield: 37%; m.p.180–182 °C; 1H NMR (400 MHz, DMSO-d6) δ = 8.22 (d, J = 8.2 Hz, 1H), 7.87 (d, J = 8.7 Hz, 1H), 7.75–7.67 (m, 1H), 7.63–7.53 (m, 1H), 7.48–7.36 (m, 2H), 6.98 (d, J = 8.4 Hz, 1H, H-8), 5.90 (s, 2H, –CH2–, H-6), 3.56–3.39 (m, 1H, –CH–, H-24), 2.16–2.07 (m, 2H, –CH2–, H-25), 2.02–1.85 (m, 2H, –CH2–, H-28), 1.80–1.63 (m, 4H, –CH2–, H-26, 27). 13C NMR (101 MHz, DMSO-d6) δ = 169.26, 159.42, 140.85, 133.42, 133.22, 130.90, 129.84, 129.11, 127.84, 127.84, 123.32, 121.31, 121.05, 110.82, 49.80, 35.12, 30.55, 25.07. MS-ESI (m/z): 412.8 (M + 1)+; HPLC: 92.73%; RT: 6.964 min.
5.2 Biological activity
5.3 Computational studies
Author contributions
Rangappa Keri: resources, writing – original draft, project administration, funding acquisition, data curation, conceptualization methodology, funding acquisition. Raveendra Madhukar Bhat: data curation, conceptualization methodology, data curation. Priyadarshini A. N.: methodology, visualization, validation. Dr Sudhanva M. S.: visualization, resources, formal analysis, data curation. Gangadhar V Muddapur: software, methodology, formal analysis. Kawthar Alhussinei: software, methodology, formal analysis. Raman Kumar K: visualization.Conflicts of interest
The authors declared that they have no conflicts of interest.Abbreviations
| ADMET | Absorption, distribution, metabolism, excretion, and toxicity |
| AO | Acridine orange |
| ANOVA | Analysis of variance |
| ADP | Adenosine diphosphate |
| BBB | Blood brain barrier |
| CNS | Central nervous system |
| CRC | Colorectal cancer |
| DFT | Density functional theory |
| DNA | Deoxyribonucleic acid |
| FBS | Fetal bovine serum |
| FDA | Food and drug administration |
| FEL | Free energy landscape |
| GCO | Global cancer observatory |
| h | Hour |
| HK-2 | Hexokinase 2 |
| HOMO | Highest occupied molecular orbital |
| HPLC | High-performance liquid chromatography |
| IARC | International agency for research on cancer |
| LND | Lonidamine |
| LUMO | Lowest unoccupied molecular orbital |
| IC50 | Half-maximal inhibitory concentration |
| MEP | Molecular electrostatic potentials |
| µg | Micro gram |
| mL | Milli litre |
| m.p. | Melting points |
| MTT | 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide |
| M.W. | Molecular weight |
| NMR | Nuclear magnetic resonance |
| NSCLC | Non-small cell lung cancer |
| OLED | Organic light emitting diode |
| PCA | Principal component analysis |
| PDB | Protein data bank |
| PI | Propidium iodide |
| PROTAC | Proteolysis targeting chimeras |
| Rg | Radius of gyration |
| RMSD | Root mean square deviation |
| RMSF | Root mean square fluctuation |
| ROS | Reactive oxygen species |
| SAR | Structure activity relationship |
| SASA | Solvent accessible surface area |
| SEM | standard error of the mean |
| TLC | Thin layer chromatography |
| TMS | Tetramethylsilane |
| TPSA | Topological polar surface area |
| WHO | World health organization |
Data availability
All the data is contained in the manuscript. More data can be obtained from the corresponding author through request email.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra07852k.
Acknowledgements
We acknowledge Jain University, Bangalore for the financial support (Minor research project – JU/MRP/CNMS/119/2025). RB thanks to Aurigene Pharmaceutical Services, Bangalore, for providing the necessary facilities.References
- R. B. Liszbinski, G. G. Romagnoli, C. M. Gorgulho, C. R. Basso, V. A. Pedrosa and R. Kaneno, anti-EGFR-coated gold nanoparticles in vitro carry 5-fluorouracil to colorectal cancer cells, Materials, 2020, 13, 375, DOI:10.3390/ma13020375.
- H. Brenner, M. Kloor and C. P. Pox, Colorectal cancer, Lancet, 2014, 383, 1490–1502, DOI:10.1016/S0140-6736(13)61649-9.
- J. H. Pan, B. Abernathy, Y. J. Kim, J. H. Lee, J. H. Kim, E. C. Shin and J. K. Kim, Cruciferous vegetables, and colorectal cancer prevention through microRNA regulation: A review, Crit. Rev. Food Sci. Nutr., 2018, 58, 2026–2038, DOI:10.1080/10408398.2017.1300134.
- R. L. Siegel, K. D. Miller, S. A. Fedewa, D. J. Ahnen, R. G. S. Meester, A. Barzi and A. Jemal, Colorectal cancer statistics, 2017, Ca-Cancer J. Clin., 2017, 67, 177–193, DOI:10.3322/caac.21395.
- R. L. Siegel, K. D. Miller, H. E. Fuchs and A. Jemal, Cancer Statistics, Ca-Cancer J. Clin., 2021, 71, 7–33, DOI:10.3322/caac.21654.
- A. Haque, D. Brazeau and A. R. Amin, Perspectives on natural compounds in chemoprevention and treatment of cancer: An update with new promising compounds, Eur. J. Cancer, 2021, 149, 165–183, DOI:10.1016/j.ejca.2021.03.009.
- G. Mohan Shankar, M. Swetha, C. K. Keerthana, T. P. Rayginia and R. J. Anto, Cancer chemoprevention: A strategic approach using phytochemicals, Front. Pharmacol, 2022, 13(12), 809308, DOI:10.3389/fphar.2021.809308.
- C. Wu, Systemic therapy for colon cancer, Surg. Oncol. Clin. N. Am., 2018, 27, 235–242, DOI:10.1016/j.soc.2017.11.001.
- V. Barresi, Colorectal cancer: from pathophysiology to novel therapeutic approaches, Biomedicines, 2021, 9, 1858, DOI:10.3390/biomedicines9121858.
- J. C. Fabregas, B. Ramnaraign and T. J. George, Clinical updates for colon cancer care in 2022, Clin. Colorectal Cancer, 2022, 21, 198–203, DOI:10.1016/j.clcc.2022.05.006.
- A. M. Gonzalez-Angulo and J. Fuloria, The role of chemotherapy in colon cancer, Ochsner J., 2002, 4, 163–167 Search PubMed.
- N. Petrelli, H. O. Douglass, L. Herrera, D. Russell, D. M. Stablein, H. W. Bruckner, R. J. Mayer, R. Schinella, M. D. Green and F. M. Muggia, The modulation of fluorouracil with leucovorin in metastatic colorectal carcinoma: A prospective randomized phase III trial. gastrointestinal tumor study group, J. Clin. Oncol., 1989, 7, 1419–1426, DOI:10.1200/JCO.1989.7.10.1419.
- C. Erlichman, S. Fine, A. Wong and T. Elhakim, A randomized trial of fluorouracil and folinic acid in patients with metastatic colorectal carcinoma, J. Clin. Oncol., 1988, 6, 469–475, DOI:10.1200/JCO.1988.6.3.469.
- F. G. A. Jansman, D. T. Sleijfer, J. C. de Graaf, J. L. L. M. Coenen and J. R. B. J. Brouwers, Management of chemotherapy- induced adverse effects in the treatment of colorectal cancer, Drug Saf., 2001, 24, 353–367, DOI:10.2165/00002018-200124050-00002.
- V. T. DeVita, T. S. Lawrence and S. A. Rosenberg, Devita, Hellman, and Rosenberg's Cancer: Principles & Practice of Oncology, Wolters Kluwer, 2015 Search PubMed.
- J. Boström, A. Hogner, A. Llinàs, E. Wellner and A. T. Plowright, Oxadiazoles in medicinal chemistry, J. Med. Chem., 2012, 55, 1817–1830, DOI:10.1021/jm2013248.
- R. M. Bhat, V. Hegde, S. Budagumpi, V. Adimule and R. S. Keri, Benzimidazole-oxadiazole hybrids-development in medicinal chemistry: An overview, Chem. Biol. Drug Des., 2024, 104, e14609, DOI:10.1111/cbdd.14609.
- S. K. Verma, R. Verma, S. Verma, Y. Vaishnav, S. P. Tiwari and K. P. Rakesh, Anti-tuberculosis activity and its structure-activity relationship (SAR) studies of oxadiazole derivatives: A key review, Eur. J. Med. Chem., 2021, 209, 112886, DOI:10.1016/j.ejmech.2020.112886.
- T. Glomb, K. Szymankiewicz and P. Świątek, Anti-cancer activity of derivatives of 1,3,4-oxadiazole, Molecules, 2018, 23, 3361, DOI:10.3390/molecules23123361.
- G. Verma, G. Chashoo, A. Ali, M. F. Khan, W. Akhtar, I. Ali, M. Akhtar, M. M. Alam and M. Shaquiquzzaman, Synthesis of pyrazole acrylic acid based oxadiazole and amide derivatives as antimalarial and anticancer agents, Bioorg. Chem., 2018, 77, 106–124, DOI:10.1016/j.bioorg.2018.01.007.
- M. Xu, P. Wu, F. Shen, J. Ji and K. P. Rakesh, Chalcone derivatives and their antibacterial activities: Current development, Bioorg. Chem., 2019, 91, 103133, DOI:10.1016/j.bioorg.2019.103133.
- R. V. Shingalapur, K. M. Hosamani, R. S. Keri and M. H. Hugar, Derivatives of benzimidazole pharmacophore: Synthesis, anticonvulsant, antidiabetic and DNA cleavage studies, Eur. J. Med. Chem., 2010, 45, 1753–1759, DOI:10.1016/j.ejmech.2010.01.007.
- H. S. Abd-Ellah, M. Abdel-Aziz, M. E. Shoman, E. A. M. Beshr, T. S. Kaoud and A.-S. F. F. Ahmed, New 1,3,4-oxadiazole/oxime hybrids: Design, synthesis, anti-inflammatory, COX inhibitory activities and ulcerogenic liability, Bioorg. Chem., 2017, 74, 15–29, DOI:10.1016/j.bioorg.2017.06.003.
- K. Rana, Salahuddin and J. K. Sahu, Significance of 1,3,4-oxadiazole containing compounds in new drug development, Curr. Drug Abuse Rev., 2021, 13, 90–100, DOI:10.2174/2589977512666201221162627.
- Q. Peng, Q. Zhou, D. Zhong, F. Pan, H. Liang and J. Zhou, Stable RNA interference of hexokinase II gene inhibits human colon cancer LoVo cell growth in vitro and in vivo, Cancer Biol. Ther., 2008, 7, 1128–1135, DOI:10.4161/cbt.7.7.6199.
- K. Nath, L. Guo, B. Nancolas, D. S. Nelson, A. A. Shestov, S.-C. Lee, J. Roman, R. Zhou, D. B. Leeper, A. P. Halestrap, I. A. Blair and J. D. Glickson, Mechanism of antineoplastic activity of lonidamine, Biochim. Biophys. Acta Rev. Canc, 2016, 1866, 151–162, DOI:10.1016/j.bbcan.2016.08.001.
- P. Pronzato, D. Amoroso, G. Bertelli, P. Conte, M. Cusimano, G. Ciottoli, M. Gulisano, R. Lionetto and R. Rosso, Phase II study of lonidamine in metastatic breast cancer, Br. J. Cancer, 1989, 59, 251–253, DOI:10.1038/bjc.1989.51.
- H. Chen, F. Chen, W. Hu and S. Gou, Effective platinum(IV) prodrugs conjugated with lonidamine as a functional group working on the mitochondria, J. Inorg. Biochem., 2018, 180, 119–128, DOI:10.1016/j.jinorgbio.2017.11.017.
- D. Cervantes-Madrid, Y. Romero and A. Dueñas-González, Reviving lonidamine and 6-diazo-5-oxo-l-norleucine to be used in combination for metabolic cancer therapy, BioMed Res. Int., 2015, 2015, 1–13, DOI:10.1155/2015/690492.
- R. M. Bhat, V. Hegde, A. K. Shettar, M. R. Alam, M. F. AlAjmi, J. H. Hoskeri and R. S. Keri, Mimics of benserazide-an oxazole derivatives: Synthesis, molecular docking/simulation study, and cytotoxicity assessment on lung and cervical cancer cell lines, J. Mol. Struct., 2025, 1341, 142585, DOI:10.1016/j.molstruc.2025.142585.
- R. M. Bhat, V. Potluri, B. Kattula, V. Hegde, M. F. AlAjmi, M. R. Alam, P. K. Prasad and R. S. Keri, Hexokinase 2 inhibition study of lonidamine-tacrine/quinazoline hybrids: Design, synthesis, molecular docking, and dynamics simulations, J. Mol. Struct., 2025, 1344, 142996, DOI:10.1016/j.molstruc.2025.142996.
- Y. Huang, G. Sun, X. Sun, F. Li, L. Zhao, R. Zhong and Y. Peng, The potential of lonidamine in combination with chemotherapy and physical therapy in cancer treatment, Cancers, 2020, 12, 3332, DOI:10.3390/cancers12113332.
- A. Vaidya, D. Pathak and K. Shah, 1,3,4-oxadiazole and its derivatives: A review on recent progress in anticancer activities, Chem. Biol. Drug Des., 2021, 97, 572–591, DOI:10.1111/cbdd.13795.
- G. Kumar, R. Kumar, A. Mazumder, Salahuddin and U. Kumar, 1,3,4-Oxadiazoles as anticancer agents: A review, Recent Pat. Anticancer Drug Discov., 2024, 19, 257–267, DOI:10.2174/1574892818666230727102928.
- https://www.swissadme.ch/.
- P. Ertl, B. Rohde and P. Selzer, Fast calculation of molecular polar surface area as a sum of fragment-based contributions and its application to the prediction of drug transport properties, J. Med. Chem., 2000, 43, 3714–3717, DOI:10.1021/jm000942e.
- J. Ali, P. Camilleri, M. B. Brown, A. J. Hutt and S. B. Kirton, Revisiting the general solubility equation: in silico prediction of aqueous solubility incorporating the effect of topographical polar surface area, J. Chem. Inf. Model., 2012, 52, 420–428, DOI:10.1021/ci200387c.
- C. A. Lipinski, Lead- and drug-like compounds: the rule-of-five revolution, Drug Discov. Today Technol., 2004, 1, 337–341, DOI:10.1016/j.ddtec.2004.11.007.
- A. Daina and V. Zoete, A BOILED-egg to predict gastrointestinal absorption and brain penetration of small molecules, ChemMedChem, 2016, 11, 1117–1121, DOI:10.1002/cmdc.201600182.
- https://tox.charite.de/protox3.
- J. G. Swathantraiah, S. M. Srinivasa, A. K. Belagal Motatis, A. Uttarkar, S. Bettaswamygowda, S. B. Thimmaiah, V. Niranjan, S. Rangappa, R. K. Subbegowda and T. N. Ramegowda, Novel 1,2,5-trisubstituted benzimidazoles potentiate apoptosis by mitochondrial dysfunction in panel of cancer cells, ACS Omega, 2022, 7, 46955–46971, DOI:10.1021/acsomega.2c06057.
- Y. R. Girish, B. A. Kumar, K. S. S. Kumar, V. K. Hamse, K. Prashantha, M. S. Sudhanva and R. Shobith, Identification of novel benzimidazole-based small molecule targeting dual targets Tankyrase and Bcl2 to induce apoptosis in Colon cancer, J. Mol. Struct., 2022, 1269, 133813, DOI:10.1016/j.molstruc.2022.133813.
- J. Eberhardt, D. Santos-Martins, A. F. Tillack and S. Forli, AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings, J. Chem. Inf. Model., 2021, 61, 3891–3898, DOI:10.1021/acs.jcim.1c00203.
- O. Trott and A. J. Olson, AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading, J. Comput. Chem., 2010, 31, 455–461, DOI:10.1002/jcc.21334.
- C. HMADA, K. Belfon, I. Ben-Shalom, J. Berryman, S. Brozell, D. Cerutti, T. Cheatham, G. Cisneros, V. Cruzeiro and T. Darden, Amber2022, University of California, San Francisco, 2022 Search PubMed.
- C. Tian, K. Kasavajhala, K. A. Belfon, L. Raguette, H. Huang, A. N. Migues, J. Bickel, Y. Wang, J. Pincay and Q. Wu, FF19SB: Amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution, J. Chem. Theory Comput., 2019, 16(1), 528–552, DOI:10.1021/acs.jctc.9b00591.
- X. He, V. H. Man, W. Yang, T.-S. Lee and J. Wang, A fast and high-quality charge model for the next generation general AMBER force field, J. Chem. Phys., 2020, 15 3, 114502, DOI:10.1063/5.0019056.
- D. R. Roe and T. E. Cheatham III, PTRAJ and CPPTRAJ: Software for processing and analysis of molecular dynamics trajectory data, J. Chem. Theory Comput., 2013, 9, 3084–3095, DOI:10.1021/ct400341p.
- C. Wei, C. H. Heh and S. P. Chin, Selective stabilization of LRRK2 kinase conformations reveals distinct modes of action for type I and II inhibitors, Cell Biochem. Biophys., 2025, 1–13, DOI:10.1007/s12013-025-01877-1.
- C. Wei, C. H. Heh and S. P. Chin, Exploring the impact of LRRK2 WD40 G2294R mutation on conformation and dimerisation dynamics: insights from molecular dynamics simulation, J. Cell. Biochem., 2025, 126(2), e70011, DOI:10.1002/jcb.70011.
- B. R. Miller III, T. D. McGee Jr, J. M. Swails, N. Homeyer, H. Gohlke and A. E. Roitberg, MMPBSA. py: An efficient program for end-state free energy calculations, J. Chem. Theory Comput., 2012, 8(9), 3314–3321, DOI:10.1021/ct300418h.
| This journal is © The Royal Society of Chemistry 2026 |












