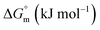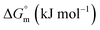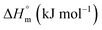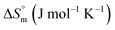Open Access Article
Open Access ArticleAssessment of the interaction between sodium dodecyl sulfate and trypsin enzyme through micellization and thermodynamic analysis
Md Al Amin Hossaina,
Ripa Aktera,
Md. Nasir Uddina,
Md. Tuhinur R. Joya,
Bulbul Ahmed b,
Dileep Kumar
b,
Dileep Kumar *cd,
Md Abdul Gonie,
K. M. Anis-Ul-Haque
*cd,
Md Abdul Gonie,
K. M. Anis-Ul-Haque a,
Md. Anamul Hoque
a,
Md. Anamul Hoque b and
Ajamaluddin Malik
b and
Ajamaluddin Malik f
f
aDepartment of Chemistry, Jashore University of Science and Technology, Jashore 7408, Bangladesh
bDepartment of Chemistry, Jahangirnagar University, Savar, Dhaka 1342, Bangladesh
cLaboratory for Chemical Computation and Modeling, Institute for Computational Science and Artificial Intelligence, Van Lang University, Ho Chi Minh City, Vietnam. E-mail: kumar.dileep@vlu.edu.vn
dFaculty of Applied Technology, Van Lang School of Technology, Van Lang University, Ho Chi Minh City, Vietnam
eDepartment of Biological and Physical Sciences, South Carolina State University, Orangeburg, SC 29117, USA
fDepartment of Biochemistry, College of Science, King Saud University, P.O. Box 2460, Riyadh 11451, Saudi Arabia
First published on 18th March 2026
Abstract
This study investigates the micellization behavior of a system comprising sodium dodecyl sulfate (SDS) and a biologically relevant protein, trypsin as well as the effects of different sodium salts and temperature on that system. The results reveal complex interactions among the surfactants, enzymes, and electrolytes, providing insights into the physicochemical mechanisms governing protein–surfactant systems. By systematically varying the concentrations of sodium salts (NaBr, CH3COONa(NaOAc), Na2SO4, and Na3PO4) and modulating the temperature from 290.55 K to 320.55 K, this research explores the self-association of SDS in the presence of trypsin. Conductometric analysis shows that sodium electrolytes reduce the critical micelle concentration (CMC) values of SDS, thereby enhancing micellization. Temperature influences this effect by generally favoring the micelle formation at lower temperatures while destabilizing the micelles at higher temperatures. The Gibbs free energy change 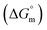 was found to be consistently negative (from −31.40 to −34.88 kJ mol−1), indicating the spontaneity of the micellization. Temperature-dependent variations in standard enthalpy
was found to be consistently negative (from −31.40 to −34.88 kJ mol−1), indicating the spontaneity of the micellization. Temperature-dependent variations in standard enthalpy 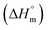 and entropy
and entropy 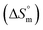 reveal that the entropy changes control the micelle formation at lower temperatures, while at higher temperatures, both entropy and enthalpy jointly control the process. Further, a molecular docking study was carried out using the CB-Dock2 online docking server with AutoDock Vina, which confirmed the binding interactions between the SDS surfactant and the trypsin protein. These findings have implications for biochemical processes, pharmaceutical formulation, and enzyme-based applications, where protein–surfactant interactions are critical.
reveal that the entropy changes control the micelle formation at lower temperatures, while at higher temperatures, both entropy and enthalpy jointly control the process. Further, a molecular docking study was carried out using the CB-Dock2 online docking server with AutoDock Vina, which confirmed the binding interactions between the SDS surfactant and the trypsin protein. These findings have implications for biochemical processes, pharmaceutical formulation, and enzyme-based applications, where protein–surfactant interactions are critical.
1. Introduction
Protein–surfactant interaction study is a vital research area for various applications, such as biotherapy,1,2 food processing,3–7 cosmetics,8 enzyme stabilization,9 and pharmaceutical formulation.10 Amphiphilic molecules such as surfactants can significantly impact protein metabolism by affecting aggregation, denaturation, and activity modulation.11–14 While a specific group of surfactants can refold denatured proteins or stabilize enzymes in harsh environments, which is critical for industrial applications,15,16 others can destabilize proteins by disrupting hydrophobic interactions, potentially impairing biological functions. These processes are governed not only by hydrophobic interactions but also by supramolecular forces, such as electrostatic (ion–ion), hydrogen bonding,17 ion-dipole,13 and dipole–dipole interactions.1,18,19 These forces collectively determine the binding affinity, structural conformation, and functional properties of protein–surfactant complexes.20,21 Hence, good understanding of the types of interactions between proteins and surfactants is crucial for investigating a wide range of physico-chemical and biological processes.20,22 More specifically, surfactants can alter the structure, stability, and functionality of essential proteins such as enzymes, resulting in enhanced stability in specific formulations or destabilization of the enzymes by disrupting the geometry of the active site.16,23 Such interference may impair the enzyme's metabolic action, leading to reduced catalytic efficiency or complete inactivation, which can have critical implications in pharmaceutical formulations and biotechnological processes.1,24 Surfactants can affect enzyme activity in two primary ways: first, by directly binding to the enzyme, which influences its structure (secondary and tertiary) and flexibility, thus altering its ability to catalyze reactions, and second, by modifying the environment in which the enzyme functions.22,25 In the presence of surfactants, they tend to adsorb on surfaces or incorporate into membranes, significantly affecting interfacial and membrane-bound enzymes. Surfactants can also protect enzymes from denaturation induced by surfaces by altering the characteristics of the surface, offering a protective effect.Trypsin is a serine protease enzyme that catalyzes the breakdown of proteins into smaller peptides in the digestive system.26,27 The pancreas secretes trypsinogen as an inactive precursor of trypsin, which is activated by enterokinase in the small intestine.28 Trypsin is essential for the digestion of dietary proteins and for the activation of other digestive enzymes. In addition to its digestive role, trypsin is used in various biotechnological applications, such as cell culture29 to dissociate adherent cells from culture vessels and protein purification to remove fusion tags in recombinant protein production. Trypsin's activity is highly dependent on its structural integrity and the surrounding ionic environment, making it sensitive to pH, temperature, and the presence of cofactors, inhibitors or surfactants.8 SDS, an anionic surfactant, significantly influences enzyme structure, solubility, stability, and function.30–33 Drug formulation is another important sector where SDS is actively involved to upgrade the activity of the drug molecule.34,35 Studies have shown that SDS binds to trypsin, where SDS can induce structural alterations in trypsin, affecting its stability and function.32,36 Additionally, microcalorimetric studies have measured the binding of SDS to trypsin at different pH levels, revealing cooperative binding behavior and enthalpy changes associated with protein unfolding.37 The partial unfolding of trypsin exposes buried active sites or functional domains, which might be beneficial for specific biochemical or industrial applications. The degree of these effects depends on different features such as SDS concentration, pH, temperature and ionic environment.
Salts can alter electrostatic interactions between trypsin and surfactants.38 The electrolyte effects on the micellization of surfactants can be interpreted by studying their positions in the Hofmeister series (lyotropic series).39–42 Different ions in the series have different abilities to influence various properties of water, proteins, and other characteristics of surfactant solutions, such as the critical micelle concentration (CMC), aggregation number, and stability of micelles.40 Depending on their position in the lyotropic series, ions can enhance or disrupt the hydrogen-bonding network of water.42 Ions that are higher in the Hofmeister series, known as kosmotropes, can strengthen the hydrogen-bonding network of water and enhance its structure. Conversely, chaotropic ions, which rank lower in the series, can weaken the water network by disrupting water's hydrogen bonds, thereby affecting the behavior of surfactants.41 Temperature also plays a vital role by influencing the flexibility of enzymes and surfactants, enhancing their interactions at higher temperatures, while excessive heat may cause enzyme denaturation and reduce surfactant efficacy. Numerous studies have previously investigated the interactions between the SDS surfactant and the trypsin protein.32,36 However, none of them attempted to assess the association nature of SDS with trypsin protein and the thermodynamic relations of the respective system in detail in the presence of sodium electrolytes. Trypsin is not used as a drug carrier, but surfactants are utilized as drug carriers. Additionally, the study of the interactions between surfactants and trypsin to understand the additive effects (such as electrolytes) on micelle formation is essential, as real-world systems, including pharmaceutical formulations, personal care products, food items, and surfactant solutions in industrial applications (such as petrochemistry), are not merely pure aqueous solutions but rather solutions containing a specific quantity of additives. Our primary aim was to demonstrate the interactions between these two components (surfactant and trypsin) in both aqueous and aqueous electrolyte solutions.
This study aimed to examine the potential interactions between SDS and trypsin protein in the presence of several sodium salts (NaBr, Na2SO4, Na3PO4, and NaOAc). Conductometric experiment investigates the impacts of temperature and the ionic environment, concentrating on notable alterations that affect trypsin + SDS micellization behavior by changing the ionic profile (sodium electrolytes) and concentration (0.05–8.00 mmol kg−1). The measurements were conducted within the temperature range of 290.55–320.55 K, with intervals of 5 K. Molecular docking simulations were also performed to investigate how SDS might bind to trypsin. This analysis aimed to identify potential binding sites. To characterize the aggregation of the SDS + trypsin mixture, various physicochemical parameters and various thermodynamic quantities were determined. The enthalpy–entropy compensation of micellization was determined and discussed in detail to investigate the trypsin + SDS interaction in the presence or absence of sodium-based salts at a certain concentration and temperature. Understanding these dynamics is essential for optimizing enzyme–surfactant systems in various industrial and research applications.
2. Materials and methods
2.1. Materials
Several chemicals (Table 1) were utilized in this study to carry out the experimental procedures and analyses. Double-distilled water (specific conductivity <1.50 µS cm−1) was used to prepare the solutions of trypsin and SDS with and without electrolytes.| Chemical | Origin | Mass fraction purity | CAS no. | Molecular weight (g mol−1) |
|---|---|---|---|---|
| SDS | Sigma Aldrich, USA | 0.99 | 151-21-3 | 288.38 |
| Trypsin | Sigma Aldrich, Switzerland | 0.99 | 9002-07-7 | 23300.0 |
| NaBr | Duksan Pure Chemicals Co., Ltd., South Korea | 0.98 | 7647-15-6 | 102.91 |
| Na2SO4 | Scharlau Chemicals, Spain | 0.99 | 7757-82-6 | 142.04 |
| Na3PO4 | Merck, Mumbai, India | 0.98 | 7601-54-9 | 163.94 |
| NaOAc | Research-Lab Fine Chem Industries, Mumbai, India | 0.99 | 127-09-3 | 82.03 |
2.2. Methods
3. Results and discussion
3.1. Conductometric investigation
 | ||
| Fig. 1 Representative plot of the specific conductivity vs. SDS concentration for a mixture of SDS and 0.5% (w/w) trypsin in 1.0 mmol kg−1 NaBr at 310.55 K. | ||
The interaction between counterions and the micelle surface (Helmholtz layer) reduces the overall micelle charge density.53 Therefore, the steep slope of the conductivity curve before the CMC reflects SDS dissociation, while the flattening after the CMC indicates micelle formation, where the conductivity is slightly influenced by the free surfactant ion concentration. Physicochemical parameters such as the degree of ionization (α) and counter-ion binding (β), linked to the CMC, provide insight into the micellization process and its stability. These parameters can be determined graphically from the conductivity vs. concentration curve (Fig. 1). The different evaluated physicochemical parameters are depicted in Table 2.
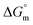 values for the mixture of SDS and trypsin in aqueous and numerous aq. sodium salt systems at 310.55 K
values for the mixture of SDS and trypsin in aqueous and numerous aq. sodium salt systems at 310.55 K
| Medium | Ctrypsin (% (w/w)) | Csalt (mmol kg−1) | Isalt (mmol kg−1) | CMC (mmol kg−1) | XCMC (×104) | S1 | S2 | β | |
|---|---|---|---|---|---|---|---|---|---|
| H2O | 0.005 | 0 | 0 | 8.28 | 1.49 | 64.61 | 32.30 | 0.50 | −34.12 |
| 0.1 | 8.31 | 1.50 | 64.15 | 32.08 | 0.50 | −34.11 | |||
| 0.5 | 8.45 | 1.52 | 64.83 | 33.08 | 0.49 | −33.82 | |||
| 1 | 8.75 | 1.58 | 119.8 | 63.51 | 0.47 | −33.23 | |||
| 1.5 | 9.95 | 1.79 | 110.3 | 61.75 | 0.44 | −32.07 | |||
| 2.5 | 11.28 | 2.03 | 108.5 | 61.84 | 0.43 | −31.39 | |||
| H2O + NaBr | 0.5 | 0.05 | 0.05 | 7.32 | 1.32 | 66.04 | 33.50 | 0.49 | −34.37 |
| 0.1 | 0.1 | 7.26 | 1.31 | 127.5 | 66.31 | 0.48 | −34.17 | ||
| 1 | 1 | 6.98 | 1.26 | 55.59 | 29.79 | 0.46 | −33.86 | ||
| 3 | 3 | 6.65 | 1.20 | 53.14 | 29.23 | 0.45 | −33.80 | ||
| 5 | 5 | 6.42 | 1.16 | 49.99 | 26.50 | 0.47 | −34.40 | ||
| 8 | 8 | 6.11 | 1.10 | 60.07 | 31.30 | 0.48 | −34.83 | ||
| H2O + NaOAc | 0.5 | 0.05 | 0.05 | 7.36 | 1.33 | 64.38 | 32.83 | 0.49 | −34.35 |
| 0.1 | 0.1 | 7.28 | 1.31 | 66.34 | 34.66 | 0.48 | −34.16 | ||
| 1 | 1 | 7.16 | 1.29 | 67.14 | 33.57 | 0.50 | −34.68 | ||
| 3 | 3 | 6.86 | 1.24 | 119.3 | 62.05 | 0.48 | −34.39 | ||
| 5 | 5 | 6.62 | 1.19 | 60.84 | 32.85 | 0.46 | −34.06 | ||
| 8 | 8 | 6.35 | 1.14 | 51.83 | 28.50 | 0.45 | −33.98 | ||
| H2O + Na2SO4 | 0.5 | 0.017 | 0.05 | 7.28 | 1.31 | 131.4 | 67.16 | 0.49 | −34.39 |
| 0.033 | 0.1 | 7.22 | 1.30 | 126.3 | 66.99 | 0.47 | −33.96 | ||
| 0.333 | 1 | 6.9 | 1.24 | 120.4 | 64.60 | 0.46 | −33.90 | ||
| 1.000 | 3 | 6.56 | 1.18 | 118.9 | 65.73 | 0.45 | −33.86 | ||
| 1.667 | 5 | 6.32 | 1.14 | 115.9 | 63.76 | 0.45 | −34.00 | ||
| 2.667 | 8 | 5.98 | 1.08 | 118.7 | 67.67 | 0.43 | −33.73 | ||
| H2O + Na3PO4 | 0.5 | 0.008 | 0.05 | 7.26 | 1.31 | 65.01 | 33.16 | 0.49 | −34.40 |
| 0.017 | 0.1 | 7.18 | 1.29 | 65.01 | 33.81 | 0.48 | −34.21 | ||
| 0.167 | 1 | 6.81 | 1.23 | 69.13 | 34.54 | 0.50 | −34.88 | ||
| 0.500 | 3 | 6.48 | 1.17 | 69.27 | 36.33 | 0.48 | −34.60 | ||
| 0.833 | 5 | 6.22 | 1.12 | 68.11 | 36.78 | 0.46 | −34.29 | ||
| 1.333 | 8 | 5.87 | 1.06 | 126.4 | 69.33 | 0.45 | −34.27 |
In the pre-micellar region, where surfactants exist primarily as monomers or small aggregates, conductivity increases linearly with concentration with a steeper slope, S1. However, beyond CMC, micelle formation causes a significant decrease in conductivity, which is represented by the less steep slope, S2. The parameter α can be calculated using these slopes from the following relation: α = S2/S1.48 Here, α offers a quantitative measure of the change in the system's behavior as surfactant molecules transition from pre-micellar to post-micellar conditions.54 The value of α indicates the magnitude of this transition; larger α values suggest a more abrupt change in the system's properties, while smaller values indicate a more gradual shift.55 Furthermore, α can be used to determine the value of β using the formula β = 1 − α,56,57 which is a good indicator for micellar stability.58 In this study, the obtained β values correlate directly with the micellar characteristics of the SDS and trypsin mixed systems; the values decreased with the increase in trypsin concentration (Table 2). This interference often results in increased CMC and reduced micellization efficiency.
The CMC initially increased slowly with increasing concentrations of trypsin protein (0.005–1.00%) (Table 2 and Fig. 2). However, a rapid rise in CMC was observed at higher concentrations (1.00–2.50%). This phenomenon can be attributed to trypsin's interference with surfactant aggregation, which hinders micelle formation.49,59 Similar results of the enhancement of CMC values for SDS and TTAB assembly by the introduction of pepsin have been reported in our previous studies in an aqueous environment.48,49 Trypsin can hinder the micellization of surfactants, like SDS, through various mechanisms.27,36 It can bind to negatively charged surfactant monomers via electrostatic interactions, reducing the availability of free monomers needed for micelle formation.27 This interaction effectively inhibits aggregation, leading to an increase in the CMC. Trypsin can also interfere with these hydrophobic interactions,36 hindering micellization. The hydrophobic regions of trypsin may interact with the hydrophobic tails of surfactant molecules, competing for the available space and disrupting the natural organization of surfactants into micelles. Additionally, trypsin can form stable protein–surfactant complexes,32,36 which compete with micelle formation and alter the typical structure and size of micelles. One of the previously published works36 describes the interactions between SDS and trypsin by applying different methods along with conformational changes and stability of trypsin, but they did not analyze the association behavior of SDS with trypsin and the thermodynamics of the system in the presence of electrolytes to illustrate the micellization and the interactions that we elaborately did in the current study.
In this study, we selected electrolytes (NaBr, NaOAc, Na2SO4, and Na3PO4) containing sodium cations and varying anions. Various concentrations of electrolytes were selected (NaBr from 0.05 to 8.00 mmol kg−1, NaOAc from 0.05 to 8.00 mmol kg−1, Na2SO4 from 0.02 to 2.50 mmol kg−1, and Na3PO4 from 0.01 to 1.50 mmol kg−1) to examine the effect of these salts on the association process of SDS + trypsin systems at a specific temperature. To enable a clear comparison between electrolytes, a constant ionic strength was maintained across the range of electrolytes used (Table 2). Our results demonstrate that introducing electrolytes into the SDS + trypsin system significantly affects the aggregation process. The reduction in CMC values upon electrolyte addition facilitated the association of SDS with trypsin (Fig. 3). At an identical ionic strength of all salts used, the CMC values for the studied system satisfy the following order: CMC (NaOAc) > CMC (NaBr) > CMC (Na2SO4) > CMC (Na3PO4). Electrolytes with anions of higher charge (SO42− and PO43−) more readily facilitate micellization than those with singly charged anions (OAc− and Br−) although OAc− is an organic anion with a mechanistically different behavior in surfactant solutions. Therefore, this study suggests that the higher the charge of anions for the three anions used (Br−, SO42−, and PO43−), the higher the effect in decreasing the CMC. This finding is supported by the literature, which indicates that these electrolytes promote surfactant aggregation.49,54,65 In our previous studies, we observed a decrease in the CMC values of SDS and TTAB surfactants when the pepsin protein was present in aqueous sodium salt media.48,49
 | ||
| Fig. 3 Plots for the CMC variation of SDS with 0.5% (w/w) trypsin in aqueous solutions of sodium salt media at 310.55 K. | ||
Electrolytes significantly alter surfactant properties by reducing the electrical double layer potential of the micelle and the viscosity of the solution.66 They also compress the electrical double layer surrounding the charged head groups of surfactant molecules, diminishing electrostatic repulsion between surfactants.67,68 Consequently, the surfactant molecules form micelles at a lower surfactant concentration. Because electrostatic interactions do not influence the hydrophobic effect due to surfactant molecules, both effects are independent. Additionally, electrolytes decrease solution viscosity by disrupting the structured hydration shell around the ionic head groups, leading to easier aggregation of surfactant monomers, which supports our study. For example, Na2SO4 and Na3PO4 are kosmotropic salts,40,42 which facilitate micelle formation by reducing the hydration of surfactant headgroups and lowering CMC. Na3PO4, with a stronger kosmotropic effect compared to Na2SO4, exerts an even stronger micelle-promoting effect. In contrast, organic salt NaOAc has a weaker kosmotropic effect; therefore, it offers moderate micellization. NaBr is a chaotropic salt that destabilizes micelle formation by increasing hydration and the CMC. This reflects the varying abilities of salts to influence water structure and ionic interactions, with kosmotropes promoting aggregation and chaotropes hindering it.40
Kosmotropic multivalent ions, like PO43− and SO42−, can decrease electrostatic repulsions between surfactant molecules more effetely compared to monovalent ions, like Br− and OAc−, which have a weaker screening effect. These observations also support our CMC trend. Organic electrolytes, like NaOAc, exhibit complex behavior in changing CMC. The CMC of surfactants decreases with an increase in the NaOAc concentration at a lower concentration range, following the kosmotropic nature.56,69 At lower concentrations, OAc− promotes the aggregation of surfactant molecules into micelles by a reduction in hydration. Additionally, the hydration of Na+ ions decreases the available free water, promoting surfactant aggregation into micelles at lower concentrations. However, a decrease in CMC values at lesser OAc− concentrations and a gradual increase in CMC at elevated OAc− concentrations have been described in the literature.56,70 The results observed in the present study reinforce the significance of the introduction of electrolytes in micellar systems for use in pharmaceutical and biological applications. The overall results of the current study have only a physiological relevance model, but the findings cannot provide any direct evidence of physiological relevance.
 | ||
| Fig. 4 Plots of the specific conductivity vs. concentration of SDS for the mixture of SDS and 0.5% (w/w) trypsin in 1 mmol kg−1 Na2SO4 solution at various temperatures. | ||
Unlike conductance patterns, CMC exhibits distinct behavior. As shown in Table 3 and Fig. 5, increasing the temperature initially decreases the CMC, which reaches a minimum before increasing again. In salt solutions with an identical ionic strength of 1 mmol kg−1, the CMC of SDS is lower than that in aqueous solutions, indicating that the salt promotes micellization at the temperature range investigated, which follows the order CMC (H2O) > CMC (NaOAc) > CMC (NaBr) > CMC (Na2SO4) > CMC (Na3PO4). This trend is consistent with observations in surfactant chemistry, where the CMC decreases with increasing temperature to a certain point, after which it increases.55,73 This behavior can be attributed to the interplay between enthalpy and entropy.74 The enthalpic interactions dominate at lower temperatures, leading to a decrease in CMC. However, as the temperature continues to increase, the entropic penalty associated with ordering water molecules around the hydrophobic tails becomes significant, causing the CMC to increase.
| Medium | Csalt (mmol kg−1) | Isalt (mmol kg−1) | T (K) | CMC (mmol kg−1) | XCMC (×104) | S1 | S2 | β |
|---|---|---|---|---|---|---|---|---|
| H2O | 0 | 0 | 290.55 | 7.54 | 1.36 | 55.75 | 30.11 | 0.46 |
| 295.55 | 7.32 | 1.32 | 58.55 | 31.03 | 0.47 | |||
| 300.55 | 6.90 | 1.24 | 117.6 | 61.12 | 0.48 | |||
| 305.55 | 7.35 | 1.32 | 122.3 | 66.04 | 0.46 | |||
| 310.55 | 8.45 | 1.52 | 119.8 | 64.71 | 0.46 | |||
| 315.55 | 8.75 | 1.58 | 123.5 | 67.55 | 0.45 | |||
| 320.55 | 8.92 | 1.61 | 132.2 | 74.02 | 0.44 | |||
| H2O + NaBr | 1 | 1 | 290.55 | 7.36 | 1.33 | 99.28 | 47.65 | 0.52 |
| 295.55 | 7.05 | 1.27 | 101.3 | 52.70 | 0.48 | |||
| 300.55 | 6.55 | 1.18 | 122.2 | 64.76 | 0.47 | |||
| 305.55 | 6.65 | 1.20 | 56.75 | 30.08 | 0.47 | |||
| 310.55 | 6.98 | 1.26 | 55.59 | 29.79 | 0.46 | |||
| 315.55 | 7.20 | 1.30 | 60.95 | 34.13 | 0.44 | |||
| 320.55 | 7.37 | 1.33 | 65.15 | 38.44 | 0.41 | |||
| H2O + NaOAc | 1 | 1 | 290.55 | 7.45 | 1.34 | 103.1 | 55.69 | 0.46 |
| 295.55 | 7.20 | 1.30 | 110.9 | 59.87 | 0.46 | |||
| 300.55 | 6.68 | 1.20 | 56.84 | 30.13 | 0.47 | |||
| 305.55 | 6.80 | 1.23 | 124.1 | 67.02 | 0.46 | |||
| 310.55 | 7.16 | 1.29 | 67.14 | 34.92 | 0.48 | |||
| 315.55 | 7.40 | 1.33 | 64.40 | 34.78 | 0.46 | |||
| 320.55 | 7.51 | 1.35 | 70.01 | 38.50 | 0.45 | |||
| H2O + Na2SO4 | 0.333 | 1 | 290.55 | 7.24 | 1.30 | 107.0 | 53.50 | 0.50 |
| 295.55 | 6.95 | 1.25 | 115.6 | 56.77 | 0.51 | |||
| 300.55 | 6.45 | 1.16 | 122.3 | 58.70 | 0.52 | |||
| 305.55 | 6.55 | 1.18 | 119.3 | 63.23 | 0.47 | |||
| 310.55 | 6.90 | 1.24 | 120.4 | 64.60 | 0.46 | |||
| 315.55 | 7.10 | 1.28 | 123.7 | 70.16 | 0.43 | |||
| 320.55 | 7.25 | 1.31 | 151.0 | 90.62 | 0.40 | |||
| H2O + Na3PO4 | 0.167 | 1 | 290.55 | 7.15 | 1.29 | 103.2 | 52.65 | 0.49 |
| 295.55 | 6.85 | 1.23 | 108.8 | 54.68 | 0.50 | |||
| 300.55 | 6.35 | 1.14 | 63.46 | 31.10 | 0.51 | |||
| 305.55 | 6.45 | 1.16 | 61.82 | 30.91 | 0.50 | |||
| 310.55 | 6.81 | 1.23 | 69.13 | 34.54 | 0.50 | |||
| 315.55 | 6.99 | 1.26 | 133.3 | 69.60 | 0.48 | |||
| 320.55 | 7.10 | 1.28 | 135.6 | 73.16 | 0.46 |
The dependence of CMC on temperature can be elucidated by considering the balance between hydrophilic and hydrophobic hydration around the surfactant monomers.75 As temperature increases, the kinetic energy of water molecules increases, leading to a disruption of the structured hydration shell around the hydrophilic head groups (reducing hydrophilic hydration). This desolvation of the polar head groups favors micellization. However, this effect can be counterbalanced by a simultaneous decrease in hydrophobic hydration. The structured water molecules surrounding the hydrophobic tails are also disrupted at higher temperatures (reducing hydrophobic hydration), making the environment less conducive to micellization. Thus, although reduced hydrophilic hydration promotes micelle formation, a corresponding decrease in hydrophobic hydration hinders it. In the SDS + trypsin mixture, the combined effects of reduced hydrophilic and hydrophobic hydration with rising temperature determine the overall micellization behavior. The decrease in hydrophilic hydration has a more pronounced effect as the temperature rises in the higher temperature range, leading to a decrease in CMC and vice versa.
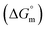 , standard enthalpy
, standard enthalpy 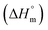 , and standard entropy
, and standard entropy 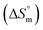 to understand the enzyme–surfactant interactions. The feasibility and spontaneity of the micellization process are primarily determined by evaluating
to understand the enzyme–surfactant interactions. The feasibility and spontaneity of the micellization process are primarily determined by evaluating 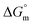 , where a negative value indicates that micelle formation occurs spontaneously.78
, where a negative value indicates that micelle formation occurs spontaneously.78Eqn (1) is employed to evaluate 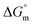 from the phase separation model:55,78
from the phase separation model:55,78
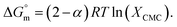 | (1) |
Eqn (2) may be used to solve the enthalpy of micellization 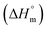 of the encapsulated system of trypsin and SDS:55,78
of the encapsulated system of trypsin and SDS:55,78
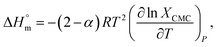 | (2) |
Given the above relationship, we can express:
| α = 1 − β. | (3) |
Therefore, eqn (1) and (2) can be written as follows:
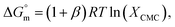 | (4) |
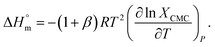 | (5) |
The empirical formula of temperature dependency ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) XCMC values is epitomized using the following equation:
XCMC values is epitomized using the following equation:
ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) XCMC = A + BT + CT2. XCMC = A + BT + CT2.
| (6) |
Differentiating ∂ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) XCMC with respect to T,
XCMC with respect to T,
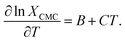 | (7) |
The values of A, B and C constants in eqn (6) were calculated using the method of least squares. The  of association was assessed by means of polynomial fitting curves (second order) of ln
of association was assessed by means of polynomial fitting curves (second order) of ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) XCMC vs. T of the employed system (Fig. 6). The values of the regression constant (A, B and C) are also presented in Table 4. After that, the enthalpy of micellization
XCMC vs. T of the employed system (Fig. 6). The values of the regression constant (A, B and C) are also presented in Table 4. After that, the enthalpy of micellization 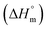 can be evaluated by applying eqn (8):
can be evaluated by applying eqn (8):
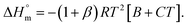 | (8) |
 | ||
Fig. 6 ln![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) XCMC vs. T plot (second order polynomial fitting) for the mixed system of SDS and 0.5% (w/w) trypsin in a 1 mmol per kg NaBr solution. XCMC vs. T plot (second order polynomial fitting) for the mixed system of SDS and 0.5% (w/w) trypsin in a 1 mmol per kg NaBr solution. | ||
| Medium | Isalt (mmol kg−1) | A | B (K−1) | C (K−2) |
|---|---|---|---|---|
| H2O | 0.0 | 27.3015 | −0.24458 | 0.00041 |
| H2O + NaBr | 1.0 | 31.6977 | −0.26730 | 0.00044 |
| H2O + NaOAc | 1.0 | 28.5394 | −0.24675 | 0.00041 |
| H2O + Na2SO4 | 1.0 | 30.6550 | −0.26060 | 0.00043 |
| H2O + Na3PO4 | 1.0 | 29.8383 | −0.25517 | 0.00042 |
Finally, it was likely to evaluate the 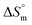 value for SDS + trypsin mixture association using the following equation:55,78
value for SDS + trypsin mixture association using the following equation:55,78
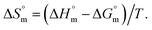 | (9) |
The 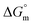 values were negative for both the aqueous and aqueous electrolyte media (Table 5), indicating that micellization occurring in these environments is thermodynamically favorable.48,57 The
values were negative for both the aqueous and aqueous electrolyte media (Table 5), indicating that micellization occurring in these environments is thermodynamically favorable.48,57 The 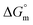 values for the four electrolytes were observed to fall within a nearly identical range, which indicates the uniformity in the thermodynamic behavior of the electrolytes in this system. This implies that the electrolytes and solvents have similar solvation properties and interaction patterns. The observed behavior of
values for the four electrolytes were observed to fall within a nearly identical range, which indicates the uniformity in the thermodynamic behavior of the electrolytes in this system. This implies that the electrolytes and solvents have similar solvation properties and interaction patterns. The observed behavior of 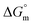 in the aqueous and aqueous electrolyte media reflects distinct temperature-dependent trends. Specially, in the case of aqueous medium, the increasing negative
in the aqueous and aqueous electrolyte media reflects distinct temperature-dependent trends. Specially, in the case of aqueous medium, the increasing negative 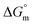 values demonstrate that micellization becomes more favoured thermodynamically as the temperature increases.48,49 A similar pattern is also obtained in the cases of aq. solutions of salts, with a minor exception.36,48,49
values demonstrate that micellization becomes more favoured thermodynamically as the temperature increases.48,49 A similar pattern is also obtained in the cases of aq. solutions of salts, with a minor exception.36,48,49
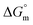 ,
,  , and
, and 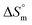 values for the mixture of SDS and 0.5% (w/w) trypsin in aq. sodium salt media at various temperatures
values for the mixture of SDS and 0.5% (w/w) trypsin in aq. sodium salt media at various temperatures
| Medium | T (K) | |||
|---|---|---|---|---|
| H2O | 290.55 | −31.40 | 6.485 | 130.4 |
| 295.55 | −32.27 | 2.380 | 117.2 | |
| 300.55 | −33.26 | −2.080 | 103.7 | |
| 305.55 | −33.12 | −6.767 | 86.24 | |
| 310.55 | −33.13 | −11.79 | 68.73 | |
| 315.55 | −33.30 | −17.01 | 51.64 | |
| 320.55 | −33.53 | −22.48 | 34.47 | |
| H2O + NaBr | 290.55 | −32.78 | 12.39 | 155.5 |
| 295.55 | −32.63 | 7.756 | 136.6 | |
| 300.55 | −33.22 | 3.109 | 120.9 | |
| 305.55 | −33.72 | −1.807 | 104.4 | |
| 310.55 | −33.86 | −7.005 | 86.46 | |
| 315.55 | −33.81 | −12.38 | 67.92 | |
| 320.55 | −33.54 | −17.81 | 49.09 | |
| H2O + NaOAc | 290.55 | −31.45 | 8.709 | 138.2 |
| 295.55 | −32.11 | 4.664 | 124.4 | |
| 300.55 | −33.15 | 0.330 | 111.4 | |
| 305.55 | −33.41 | −4.308 | 95.24 | |
| 310.55 | −34.22 | −9.376 | 80.01 | |
| 315.55 | −34.18 | −14.50 | 62.34 | |
| 320.55 | −34.42 | −19.94 | 45.17 | |
| H2O + Na2SO4 | 290.55 | −32.41 | 11.29 | 150.4 |
| 295.55 | −33.34 | 7.048 | 136.7 | |
| 300.55 | −34.41 | 2.428 | 122.6 | |
| 305.55 | −33.78 | −2.479 | 102.4 | |
| 310.55 | −33.90 | −7.578 | 84.76 | |
| 315.55 | −33.63 | −12.75 | 66.16 | |
| 320.55 | −33.37 | −18.03 | 47.86 | |
| H2O + Na3PO4 | 290.55 | −32.24 | 11.62 | 150.9 |
| 295.55 | −33.17 | 7.525 | 137.7 | |
| 300.55 | −34.24 | 3.071 | 124.2 | |
| 305.55 | −34.52 | −1.737 | 107.3 | |
| 310.55 | −34.88 | −6.846 | 90.27 | |
| 315.55 | −34.87 | −12.12 | 72.08 | |
| 320.55 | −34.88 | −17.58 | 53.98 |
The temperature-dependent behavior of  during the surfactant micellization differs notably between aqueous and aqueous electrolyte solutions (Table 5). In aqueous media, micellization exhibited a positive
during the surfactant micellization differs notably between aqueous and aqueous electrolyte solutions (Table 5). In aqueous media, micellization exhibited a positive  at lower temperatures (290.55 K and 295.55 K), indicating an endothermic process. However, in the higher temperature range (300.55–320.55 K),
at lower temperatures (290.55 K and 295.55 K), indicating an endothermic process. However, in the higher temperature range (300.55–320.55 K),  values become negative after an increasing trend. In contrast, the aqueous electrolyte medium displayed a similar pattern although the positive values of the
values become negative after an increasing trend. In contrast, the aqueous electrolyte medium displayed a similar pattern although the positive values of the  for micellization were observed up to 300.55 K instead of 295.55 K and then became increasingly negative, indicating a shift to an exothermic process at a higher temperature range (305.55–320.55 K). Again, micelle formation at this higher temperature requires higher concentrations of surfactant molecules, which is a consequence of the interplay between enthalpy and entropy. The temperature increase facilitates favorable enthalpy
for micellization were observed up to 300.55 K instead of 295.55 K and then became increasingly negative, indicating a shift to an exothermic process at a higher temperature range (305.55–320.55 K). Again, micelle formation at this higher temperature requires higher concentrations of surfactant molecules, which is a consequence of the interplay between enthalpy and entropy. The temperature increase facilitates favorable enthalpy 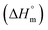 , but the entropy-driven disorder of surfactant molecules hinders aggregation, thus increasing the CMC. The temperature-dependent behavior of the
, but the entropy-driven disorder of surfactant molecules hinders aggregation, thus increasing the CMC. The temperature-dependent behavior of the 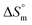 during the micellization of surfactants in aqueous and aqueous electrolyte media highlights consistent thermodynamic trends (Table 5). Across the entire temperature range, the
during the micellization of surfactants in aqueous and aqueous electrolyte media highlights consistent thermodynamic trends (Table 5). Across the entire temperature range, the 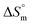 for the micellization of surfactants was positive in both aqueous and aqueous electrolyte media, suggesting that micelle formation increases in disorder or randomness in the system. Increasing temperature affects the change in
for the micellization of surfactants was positive in both aqueous and aqueous electrolyte media, suggesting that micelle formation increases in disorder or randomness in the system. Increasing temperature affects the change in  values, showing a decreasing trend regardless of the media. This consistent decrease in positive
values, showing a decreasing trend regardless of the media. This consistent decrease in positive 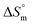 change suggests a general trend toward decreasing disorder with temperature increase although the micellization process is entropy-driven at all temperatures. Comparing the values of
change suggests a general trend toward decreasing disorder with temperature increase although the micellization process is entropy-driven at all temperatures. Comparing the values of 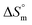 in different electrolyte media at a specific temperature (310.55 K) yields the following order:
in different electrolyte media at a specific temperature (310.55 K) yields the following order: 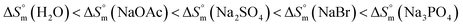 .
.
The positive  values observed for the trypsin + SDS mixture can be attributed to two factors: first, the transfer of hydrophobic components from their hydrated forms in the aqueous system to the micellar core, which disrupts the ordered “iceberg” structure of water, and second, the increased rotational freedom of the hydrophobic parts within the micelle than their more constrained state in the surrounding water.66,67 When combined, these positive
values observed for the trypsin + SDS mixture can be attributed to two factors: first, the transfer of hydrophobic components from their hydrated forms in the aqueous system to the micellar core, which disrupts the ordered “iceberg” structure of water, and second, the increased rotational freedom of the hydrophobic parts within the micelle than their more constrained state in the surrounding water.66,67 When combined, these positive  and
and 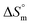 values strongly suggest evidence of nonpolar interaction forces between the surfactant and the drug or enzyme molecule.49 Temperature-dependent changes in enthalpy and entropy during micellization reveal whether the process is enthalpy-driven, entropy-driven, or governed by both contributions. In aqueous solutions, micellization exhibits a transition from an entropy-driven to an enthalpy-driven process. However, aqueous electrolyte solutions maintain an entropy-driven process at lower temperatures, transitioning to a process controlled by both entropy and enthalpy across the remaining temperature range. Binding studies reveal that trypsin + SDS interactions are spontaneous and driven by hydrogen bonding and van der Waals forces, which are observable through techniques like UV spectroscopy, circular dichroism, and fluorescence spectroscopy.36 Based on the observed entropy and enthalpy trends for SDS micellization, the interactions between trypsin and SDS likely involve a combination of electrostatic attraction and hydrophobic effects. At lower temperatures, the electrostatic interactions and hydrophobic effects likely dominate, leading to stronger binding between SDS and trypsin. As the temperature increases, the increase in entropy and enhanced solvation around the surfactant molecules could weaken the interaction, potentially leading to dissociation or reduced binding strength between SDS and trypsin at higher temperatures.
values strongly suggest evidence of nonpolar interaction forces between the surfactant and the drug or enzyme molecule.49 Temperature-dependent changes in enthalpy and entropy during micellization reveal whether the process is enthalpy-driven, entropy-driven, or governed by both contributions. In aqueous solutions, micellization exhibits a transition from an entropy-driven to an enthalpy-driven process. However, aqueous electrolyte solutions maintain an entropy-driven process at lower temperatures, transitioning to a process controlled by both entropy and enthalpy across the remaining temperature range. Binding studies reveal that trypsin + SDS interactions are spontaneous and driven by hydrogen bonding and van der Waals forces, which are observable through techniques like UV spectroscopy, circular dichroism, and fluorescence spectroscopy.36 Based on the observed entropy and enthalpy trends for SDS micellization, the interactions between trypsin and SDS likely involve a combination of electrostatic attraction and hydrophobic effects. At lower temperatures, the electrostatic interactions and hydrophobic effects likely dominate, leading to stronger binding between SDS and trypsin. As the temperature increases, the increase in entropy and enhanced solvation around the surfactant molecules could weaken the interaction, potentially leading to dissociation or reduced binding strength between SDS and trypsin at higher temperatures.
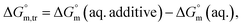 | (10) |
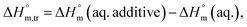 | (11) |
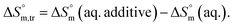 | (12) |
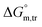 were found to be negative. Specially, abnormal positive values were observed in aqueous NaBr medium (3.00 mmol kg−1 at 310.55 K and 1 mmol kg−1 at 300.55 K), Na2SO4 medium (1 mmol kg−1 at 320.55 K) and NaOAc medium (1 mmol kg−1 at 295.55 K and 300.55 K), which need further advance investigation to find out the root cause behind this (Table S1 (SI)). The negative transfer values of
were found to be negative. Specially, abnormal positive values were observed in aqueous NaBr medium (3.00 mmol kg−1 at 310.55 K and 1 mmol kg−1 at 300.55 K), Na2SO4 medium (1 mmol kg−1 at 320.55 K) and NaOAc medium (1 mmol kg−1 at 295.55 K and 300.55 K), which need further advance investigation to find out the root cause behind this (Table S1 (SI)). The negative transfer values of 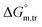 for both temperature and concentration ranges indicates that the micellization process is thermodynamically favoured by the presence of electrolytes. The positive transfer values for
for both temperature and concentration ranges indicates that the micellization process is thermodynamically favoured by the presence of electrolytes. The positive transfer values for 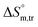 indicate that disorder increased associated with the aggregation of surfactant molecules into micelles in the electrolyte solution (Table S1 (SI)). The positive
indicate that disorder increased associated with the aggregation of surfactant molecules into micelles in the electrolyte solution (Table S1 (SI)). The positive 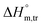 transfer suggests that the micellization process is endothermic in only electrolyte medium and may be solely an entropy-driven process (Table S1 (SI)). Despite these positive changes, the negative Gibbs free energy indicates that the micellization process remains energetically favorable, highlighting the stabilizing role of electrolytes in promoting micelle formation.
transfer suggests that the micellization process is endothermic in only electrolyte medium and may be solely an entropy-driven process (Table S1 (SI)). Despite these positive changes, the negative Gibbs free energy indicates that the micellization process remains energetically favorable, highlighting the stabilizing role of electrolytes in promoting micelle formation.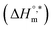 and compensation temperature (Tc) are derived from the plot of
and compensation temperature (Tc) are derived from the plot of  vs.
vs. 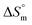 (Fig. 7) of micellization of SDS + pepsin with and without electrolytes according to eqn (13):74,80–83
(Fig. 7) of micellization of SDS + pepsin with and without electrolytes according to eqn (13):74,80–83
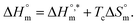 | (13) |
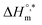 represents the intrinsic enthalpy contribution to micellization, independent of entropy and calculated from the y-intercept of the
represents the intrinsic enthalpy contribution to micellization, independent of entropy and calculated from the y-intercept of the  vs.
vs. 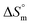 plot. Meanwhile, the compensation temperature (Tc) is the slope of the straight line of the plot, which reflects the point at which enthalpic and entropic effects are balanced in the overall free energy of micellization. Together, these parameters provide deeper mechanistic insight into the driving forces governing micelle formation in complex biochemical environments.
plot. Meanwhile, the compensation temperature (Tc) is the slope of the straight line of the plot, which reflects the point at which enthalpic and entropic effects are balanced in the overall free energy of micellization. Together, these parameters provide deeper mechanistic insight into the driving forces governing micelle formation in complex biochemical environments.
 | ||
Fig. 7 Plots of  vs. vs. 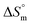 for the mixture of SDS and 0.5% (w/w) trypsin in (a) aqueous and (b) aq. NaOAc media at various temperatures. for the mixture of SDS and 0.5% (w/w) trypsin in (a) aqueous and (b) aq. NaOAc media at various temperatures. | ||
The enthalpy–entropy compensation variables of SDS micellization in the presence of trypsin and various sodium electrolytes are presented in Table 6. In pure water, micellization is moderately exothermic 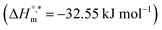 and occurs at a compensation temperature of 298.01 K. The addition of sodium electrolytes significantly alters both
and occurs at a compensation temperature of 298.01 K. The addition of sodium electrolytes significantly alters both 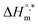 and Tc. For instance, systems containing Na3PO4 and NaOAc exhibited more negative
and Tc. For instance, systems containing Na3PO4 and NaOAc exhibited more negative 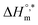 values, −33.83 and −33.84 kJ mol−1, respectively, and higher Tc values (299.70 K and 308.25 K, respectively), indicating enhanced enthalpic contributions and thermodynamic favorability of micellization. In contrast, NaBr and Na2SO4 lowered the compensation temperature (to 287.11 K and 282.38 K, respectively) and reduced the magnitude of
values, −33.83 and −33.84 kJ mol−1, respectively, and higher Tc values (299.70 K and 308.25 K, respectively), indicating enhanced enthalpic contributions and thermodynamic favorability of micellization. In contrast, NaBr and Na2SO4 lowered the compensation temperature (to 287.11 K and 282.38 K, respectively) and reduced the magnitude of 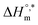 , suggesting a shift toward entropy-driven micellization. The Tc values of the 270–350 K range are reported for the aq. solutions of protein and small solutes,80 and the Tc values have been accepted as evidence of the presence of hydrophobic interactions.74
, suggesting a shift toward entropy-driven micellization. The Tc values of the 270–350 K range are reported for the aq. solutions of protein and small solutes,80 and the Tc values have been accepted as evidence of the presence of hydrophobic interactions.74
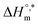 and Tc values for the mixture of SDS and 0.5% (w/w) trypsin in aq. sodium salt media at different temperatures
and Tc values for the mixture of SDS and 0.5% (w/w) trypsin in aq. sodium salt media at different temperatures
The exceptionally high R2 values (all above 0.999) indicate the consistent linear enthalpy–entropy compensation relationship across the entire temperature range studied (300.55–320.55 K) for each electrolyte condition illustrated in Table 6. This statistical robustness validates the reliability of the  and
and 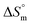 values derived from temperature-dependent conductometric measurements. More importantly, it confirms that trypsin does not introduce irregular thermodynamic behavior or significant conformational variability under these micellization conditions. Such tight compensation behavior typically indicates that micellization across the temperature range is governed by a single dominant mechanism with predictable energetic trade-offs. Therefore, in protein–surfactant systems, this suggests that trypsin remains involved in the micellization process, without significant aggregation, degradation, or secondary transitions under the studied temperature and salt conditions.
values derived from temperature-dependent conductometric measurements. More importantly, it confirms that trypsin does not introduce irregular thermodynamic behavior or significant conformational variability under these micellization conditions. Such tight compensation behavior typically indicates that micellization across the temperature range is governed by a single dominant mechanism with predictable energetic trade-offs. Therefore, in protein–surfactant systems, this suggests that trypsin remains involved in the micellization process, without significant aggregation, degradation, or secondary transitions under the studied temperature and salt conditions.
3.2. Molecular docking and analysis of the trypsin–SDS complex
The online docking server CB-Dock2, which is based on Auto-Dock-Vina, was used in this study for blind docking between trypsin and SDS. The 3D structure of trypsin was uploaded and docked with the 3D structure of SDS. The CB-Dock (Fig. 8) results revealed a high-affinity pose between trypsin and SDS, as determined by the Vina score, cavity size, and grid map. Interaction models were created based on the projected binding affinity between trypsin and SDS using a curvature-dependent surface-area model for the trypsin and SDS.84 The binding energy values (Vina score) for the top 5 binding poses were determined, with the interaction having binding energy, cavity volume, and the interacting residues, as presented in Table S2 (SI). The lowermost projected Vina score was −5.0 kcal mol−1. The agreeing binding pose is displayed in Fig. 8. Further binding poses with lower scores are shown in Fig. S1 (SI). | ||
| Fig. 8 Interaction between trypsin and SDS was analyzed via molecular docking using CB-Dock2 (molecular interaction shown in inset). | ||
The interaction between trypsin and the anionic surfactant SDS is governed by a combination of electrostatic and hydrophobic forces. Generally, the negatively charged sulfate headgroup of SDS electrostatically associates with positively charged residues on the protein surface, and the hydrophobic dodecyl alkyl chain of the surfactant interacts with non-polar residues.31,85 This cooperative binding disrupts the intricate non-covalent interactions, including H-bonding and van der Waals forces, which are essential for maintaining the protein's stable conformation. The formation of a trypsin–SDS complex may induce significant conformational changes, leading to unfolding and denaturation. A critical consequence of this structural perturbation may be the distortion of the catalytic triad and the deformation of the substrate-binding pocket. The loss of the native structure, driven by the amphiphilic nature of SDS, effectively abolishes trypsin's proteolytic function by preventing proper substrate recognition and catalysis.
4. Conclusions
This study elucidates the impacts of sodium electrolytes and temperature on the micellization behavior of the SDS and trypsin enzyme mixture. The SDS assembly process has been found to be significantly dependent on the presence of trypsin, the introduction of Na-salts and temperature alteration. Trypsin significantly causes a delay in SDS micellization by increasing the critical micelle concentration (CMC) and reducing the degree of micelle ionization, as a consequence of protein–surfactant interactions that hinder surfactant aggregation. The addition of sodium electrolytes lowers the CMC in the order NaOAc > NaBr > Na2SO4 > Na3PO4, reflecting the greater ability of higher-valent kosmotropic anions to screen electrostatic repulsions and reduce hydration of surfactant headgroups, thereby promoting tighter micelle packing. The micellization of the current system showed dependency on the temperature changes over the range studied in all the media employed. Thermodynamic analyses confirm that micellization is spontaneous across all conditions, driven by a complex interplay of enthalpic and entropic factors. The micellization of the SDS + trypsin mixture was found to be endothermic at lower temperatures, while the process became exothermic at higher temperatures studied. A positive entropy change is the dominating factor driving micellization at lower temperatures, whereas enthalpy becomes a more dominant factor at elevated temperatures. According to the thermodynamic analysis, the recommended interaction forces are hydrophobic, electrostatic and ion-dipole types. Negative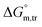 values further confirm the thermodynamic favorability of micellization despite a few anomalous cases requiring further investigation. Docking simulations show that SDS's extended nonpolar chain inserts into trypsin's hydrophobic surface regions, facilitating binding via hydrophobic interactions. Importantly, these variations occur without compromising protein stability or inducing significant conformational changes. Together, these findings provide comprehensive insights into how sodium electrolytes and temperature synergistically influence surfactant–enzyme self-assembly, offering valuable guidance for the design of stable biochemical formulations and efficient drug delivery systems.
values further confirm the thermodynamic favorability of micellization despite a few anomalous cases requiring further investigation. Docking simulations show that SDS's extended nonpolar chain inserts into trypsin's hydrophobic surface regions, facilitating binding via hydrophobic interactions. Importantly, these variations occur without compromising protein stability or inducing significant conformational changes. Together, these findings provide comprehensive insights into how sodium electrolytes and temperature synergistically influence surfactant–enzyme self-assembly, offering valuable guidance for the design of stable biochemical formulations and efficient drug delivery systems.
Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra07779f.Acknowledgements
Authors are grateful to the Ongoing Research Funding program (ORF-2026-727), King Saud University, Riyadh, Saudi Arabia.References
- T. A. Khan, H.-C. Mahler and R. S. K. Kishore, Eur. J. Pharm. Biopharm., 2015, 97, 60–67 CrossRef CAS PubMed.
- M. N. Jones, Chem. Soc. Rev., 1992, 21, 127–136 RSC.
- J.-S. Gong, W. Li, D.-D. Zhang, M.-F. Xie, B. Yang, R.-X. Zhang, H. Li, Z.-M. Lu, Z.-H. Xu and J.-S. Shi, Int. J. Mol. Sci., 2015, 16, 30061–30074 CrossRef CAS PubMed.
- P. Busti, S. Scarpeci, C. A. Gatti and N. J. Delorenzi, J. Agric. Food Chem., 1999, 47, 3628–3631 CrossRef CAS PubMed.
- M. G. Semenova, L. E. Belyakova, Y. N. Polikarpov, M. M. Il'in, T. A. Istarova, M. S. Anokhina and E. N. Tsapkina, Biomacromolecules, 2006, 7, 101–113 CrossRef CAS PubMed.
- J. G. Hansted, P. L. Wejse, H. Bertelsen and D. E. Otzen, Biochim. Biophys. Acta, 2011, 1814, 713–723 CrossRef CAS PubMed.
- S. Rouimi, C. Schorsch, C. Valentini and S. Vaslin, Food Hydrocoll., 2005, 19, 467–478 CrossRef CAS.
- S. Kumar, D. Saha, D. Ray and V. K. Aswal, Soft Matter, 2025, 21, 4979–4998 RSC.
- M. D. Womack, D. A. Kendall and R. C. MacDonald, Biochim. Biophys. Acta, 1983, 733, 210–215 CrossRef CAS PubMed.
- A. M. Morales, A. Sreedhara, J. Buecheler, S. Brosig, D. Chou, T. Christian, T. Das, I. de Jong, J. Fast, B. Jagannathan, E. M. Moussa, M. R. Nejadnik, I. Prajapati, A. Radwick, Y. Rahman and S. Singh, AAPS J., 2022, 25, 6 CrossRef PubMed.
- S. Tandon and P. M. Horowitz, J. Biol. Chem., 1987, 262, 4486–4491 CrossRef CAS PubMed.
- I. Fatma, V. Sharma, R. C. Thakur and A. Kumar, J. Mol. Liq., 2021, 341, 117344 CrossRef CAS.
- Interactions of Surfactants with Polymers and Proteins, ed. E. D. Goddard and K. P. Ananthapadmanabhan, CRC Press, Boca Raton, 1993 Search PubMed.
- Y. Li and J. S. Lee, Anal. Chim. Acta, 2019, 1063, 18–39 CrossRef CAS PubMed.
- N. B. Bam, J. L. Cleland and T. W. Randolph, Biotechnol. Prog., 1996, 12, 801–809 CrossRef CAS PubMed.
- V. V. Mozhaev, I. V. Berezin, K. Martinek and Y. Nosoh, Crit. Rev. Biochem., 1988, 23, 235–281 CrossRef CAS PubMed.
- P. Lundahl, E. Greijer, M. Sandberg, S. Cardell and K.-O. Eriksson, Biochim. Biophys. Acta, 1986, 873, 20–26 CrossRef CAS.
- C. La Mesa, J. Colloid Interface Sci., 2005, 286, 148–157 CrossRef CAS PubMed.
- D. Otzen, Biochim. Biophys. Acta, 2011, 1814, 562–591 CrossRef CAS PubMed.
- A. Mackie and P. Wilde, Adv. Colloid Interface Sci., 2005, 117, 3–13 CrossRef CAS PubMed.
- J. Maldonado-Valderrama and J. M. R. Patino, Curr. Opin. Colloid Interface Sci., 2010, 15, 271–282 CrossRef CAS.
- K. Holmberg, Colloids Surf., B, 2018, 168, 169–177 CrossRef CAS PubMed.
- M. Aguirre-Ramírez, H. Silva-Jiménez, I. M. Banat and M. A. Díaz De Rienzo, Biotechnol. Lett., 2021, 43, 523–535 CrossRef PubMed.
- V. Delorme, R. Dhouib, S. Canaan, F. Fotiadu, F. Carrière and J.-F. Cavalier, Pharm. Res., 2011, 28, 1831–1842 CrossRef CAS PubMed.
- D. N. Rubingh, Curr. Opin. Colloid Interface Sci., 1996, 1, 598–603 CrossRef CAS.
- E. Vandermarliere, M. Mueller and L. Martens, Mass Spectrom. Rev., 2013, 32, 453–465 CrossRef CAS PubMed.
- S. Ghosh and A. Banerjee, Biomacromolecules, 2002, 3, 9–16 CrossRef CAS PubMed.
- W. Kong, K. McConalogue, L. M. Khitin, M. D. Hollenberg, D. G. Payan, S. K. Böhm and N. W. Bunnett, Proc. Natl. Acad. Sci. U. S. A., 1997, 94, 8884–8889 CrossRef CAS PubMed.
- H.-L. Huang, H.-W. Hsing, T.-C. Lai, Y.-W. Chen, T.-R. Lee, H.-T. Chan, P.-C. Lyu, C.-L. Wu, Y.-C. Lu and S.-T. Lin, J. Biomed. Sci., 2010, 17, 1–10 CrossRef PubMed.
- M. L. Guzman, M. R. Marques, M. E. Olivera Me and E. S. Stippler, Results Pharma Sci., 2016, 6, 15–19 CrossRef PubMed.
- J. M. Khan, A. Malik, A. Ahmed, O. H. A. Alghamdi and M. Ahmed, J. Mol. Liq., 2020, 319, 114154 CrossRef CAS.
- S. Ghosh, Colloids Surf., B, 2008, 66, 178–186 CrossRef CAS PubMed.
- P. N. Moore, S. Puvvada and D. Blankschtein, Langmuir, 2003, 19, 1009–1016 CrossRef CAS.
- L. Maggi, M. L. Torre, P. Giunchedi and U. Conte, Int. J. Pharm., 1996, 135, 73–79 CrossRef CAS.
- S. Geng, Y. Wang, L. Wang, T. Kouyama, T. Gotoh, S. Wada and J.-Y. Wang, Sci. Rep., 2017, 7, 39202 CrossRef CAS PubMed.
- H. Ma, T. Zou, H. Li and H. Cheng, Int. J. Biol. Macromol., 2020, 162, 1546–1554 CrossRef CAS PubMed.
- D. E. Otzen, J. N. Pedersen, H. O. Rasmussen and J. S. Pedersen, Adv. Colloid Interface Sci., 2022, 308, 102754 CrossRef CAS PubMed.
- M. H. Najar, O. A. Chat, P. A. Bhat, M. A. Mir, G. M. Rather and A. A. Dar, Int. J. Biol. Macromol., 2021, 180, 121–128 CrossRef CAS PubMed.
- M. Cacace, E. Landau and J. Ramsden, Q. Rev. Biophys., 1997, 30, 241–277 CrossRef CAS PubMed.
- B. Kang, H. Tang, Z. Zhao and S. Song, ACS Omega, 2020, 5, 6229–6239 CrossRef CAS PubMed.
- X. He and A. G. Ewing, ChemBioChem, 2023, 24, e202200694 CrossRef CAS PubMed.
- K. P. Gregory, G. R. Elliott, H. Robertson, A. Kumar, E. J. Wanless, G. B. Webber, V. S. Craig, G. G. Andersson and A. J. Page, Phys. Chem. Chem. Phys., 2022, 24, 12682–12718 RSC.
- Y. Liu, X. Yang, J. Gan, S. Chen, Z.-X. Xiao and Y. Cao, Nucleic Acids Res., 2022, 50, W159–W164 CrossRef CAS PubMed.
- R. Srivastava and M. S. Alam, Int. J. Biol. Macromol., 2020, 145, 252–261 CrossRef CAS PubMed.
- S. Deep and J. C. Ahluwalia, Phys. Chem. Chem. Phys., 2001, 3, 4583–4591 RSC.
- Y. Liu, Y. Liu and R. Guo, J. Colloid Interface Sci., 2010, 351, 180–189 CrossRef CAS PubMed.
- R.-C. Lu, A.-N. Cao, L.-H. Lai, B.-Y. Zhu, G.-X. Zhao and J.-X. Xiao, Colloids Surf., B, 2005, 41, 139–143 CrossRef CAS PubMed.
- S. Kabir, M. A. A. Hossain, I. Jahan, B. Ahmed, A. Malik, M. A. Goni, M. A. Hoque and K. M. Anis-Ul-Haque, Int. J. Biol. Macromol., 2025, 291, 138990 CrossRef PubMed.
- M. A. A. Hossain, T. Islam, M. T. R. Joy, Z. Kowser, M. Z. Ahmed, M. T. Rehman, M. F. AlAjmi, S. Mahbub, M. A. Goni and M. A. Hoque, Int. J. Biol. Macromol., 2023, 253, 127478 CrossRef CAS PubMed.
- S. Mondal, M. L. Raposo, A. Ghosh, G. Prieto and S. Ghosh, Colloids Surf., A, 2019, 577, 167–174 CrossRef CAS.
- E. Fuguet, C. Ràfols, M. Rosés and E. Bosch, Anal. Chim. Acta, 2005, 548, 95–100 CrossRef CAS.
- S. Naderi Miqan, F. Farshchi Tabrizi and H. Abedini, J. Mol. Liq., 2015, 201, 59–67 CrossRef CAS.
- O. S. Zueva, V. S. Rukhlov and Y. F. Zuev, ACS Omega, 2022, 7, 6174–6183 CrossRef CAS PubMed.
- M. Al Amin Hossain, M. N. Sultana, J. M. Khan, M. T. R. Joy, S. C. Mohanta, M. R. Amin, M. F. Hossain, M. Irfan, M. Z. Ahmed, D. Kumar, M. A. Hoque and S. E. Kabir, Chem. Pap., 2022, 76, 1501–1511 CrossRef CAS.
- M. J. Rosen and J. T. Kunjappu, Surfactants and Interfacial Phenomena, John Wiley & Sons, 2012 Search PubMed.
- K. M. Anis-Ul-Haque, M. A. A. Hossain, M. J. Ferdous, M. A. Taher, J. M. Khan, A. Ahmad, M. T. R. Joy and M. A. Hoque, Mol. Phys., 2023, 122, e2253931 CrossRef.
- K. M. Anis-Ul-Haque, M. A. A. Hossain, N. Uddin, M. Jonayed, M. K. Gatasheh, D. Kumar, M. A. Goni, M. T. R. Joy and M. A. Hoque, Colloids Surf., A, 2024, 692, 133972 CrossRef CAS.
- B. Naskar, A. Dey and S. P. Moulik, J. Surfact. Deterg., 2013, 16, 785–794 CrossRef CAS.
- B. S. Rauniyar and A. Bhattarai, J. Mol. Liq., 2021, 323, 114604 CrossRef CAS.
- M. H. Weinberger, in Handbook of Research Methods in Cardiovascular Behavioral Medicine, ed. N. Schneiderman, S. M. Weiss and P. G. Kaufmann, Springer US, Boston, MA, 1989, pp. 133–143, DOI:10.1007/978-1-4899-0906-0_9.
- M. L. Fanani, N. E. Nocelli and Y. d. l. M. Zulueta Díaz, Biochim. Biophys. Acta, Biomembr., 2022, 1864, 183781 CrossRef CAS PubMed.
- M. Karar, S. Paul, A. Mallick and T. Majumdar, Chem. Phys. Lipids, 2018, 210, 122–128 CrossRef CAS PubMed.
- F. A. Wani, K. Behera and R. Patel, ChemistrySelect, 2022, 7, e202201928 CrossRef CAS.
- F. J. He and G. A. MacGregor, Kidney Int., 2004, 66, 2454–2466 CrossRef PubMed.
- M. R. Amin, S. A. Alissa, M. R. Molla, M. A. Rub, S. M. Wabaidur, M. A. Hoque and S. E. Kabir, J. Phys. Org. Chem., 2020, 33, e4057 CrossRef CAS.
- G. Para, E. Jarek and P. Warszynski, Adv. Colloid Interface Sci., 2006, 122, 39–55 CrossRef CAS PubMed.
- H. Hooshyar and R. Sadeghi, J. Chem. Eng. Data, 2015, 60, 983–992 CrossRef CAS.
- K. Maiti, D. Mitra, S. Guha and S. P. Moulik, J. Mol. Liq., 2009, 146, 44–51 CrossRef CAS.
- A. Amin, A. Ahmad Dar, M. Ahmad Bhat, M. Jan, N. Rehman, M. Amin Mir and G. Mohammad Rather, J. Dispers. Sci. Technol., 2008, 29, 406–414 CrossRef CAS.
- B. Paul, S. Islam and K. Ismail, J. Phys. Chem. B, 1998, 102, 7807–7812 CrossRef CAS.
- C. Vautier-Giongo and B. L. Bales, J. Phys. Chem. B, 2003, 107, 5398–5403 CrossRef CAS.
- E. Summerton, G. Zimbitas, M. Britton and S. Bakalis, J. Cryst. Growth, 2016, 455, 111–116 CrossRef CAS.
- E. Mohajeri and G. D. Noudeh, J. Chem., 2012, 9, 2268–2274 CrossRef CAS.
- C. Jolicoeur and P. R. Philip, Can. J. Biochem., 1974, 52, 1834–1839 CrossRef CAS.
- H. Naorem and S. D. Devi, J. Surf. Sci. Technol., 2006, 22, 89 CAS.
- H. B. Jones, S. A. Wells, E. J. Prentice, A. Kwok, L. L. Liang, V. L. Arcus and C. R. Pudney, FEBS J., 2017, 284, 2829–2842 CrossRef CAS PubMed.
- E. Noor, A. Flamholz, W. Liebermeister, A. Bar-Even and R. Milo, FEBS Lett., 2013, 587, 2772–2777 CrossRef CAS PubMed.
- M. A. Hoque, M. I. Ali, M. A. Rub, M. Rahman, S. Rana, M. M. Rahman, D. Kumar, N. Azum, A. M. Asiri and M. A. Khan, Int. J. Biol. Macromol., 2023, 228, 445–452 CrossRef CAS PubMed.
- A. Pan, A. K. Rakshit and S. P. Moulik, Colloids Surf., A, 2016, 495, 248 CrossRef CAS.
- R. Lumry and S. Rajender, Biopolym, 1970, 9, 1125–1227 CrossRef CAS PubMed.
- S. M. A. Ahsan, M. D. Hossain, M. A. Hoque and M. A. Khan, Indian J. Chem., Sect. A, 2016, 55, 160–169 Search PubMed.
- H. A. Bhuiyan, M. A. Rub, S. Mahbub, M. T. R. Joy, A. Mamun, D. Kumar, A. M. Asiri, M. A. Hoque and S. E. Kabir, J. Saudi Chem. Soc., 2022, 26, 101451 CrossRef CAS.
- L.-J. Chen, S.-Y. Lin and C.-C. Huang, J. Phys. Chem. B, 1998, 1024, 350–4356 Search PubMed.
- Y. Cao and L. Li, Bioinformatics, 2014, 30, 1674 CrossRef CAS PubMed.
- S. AlResaini, A. Malik, M. Alonazi, A. Alhomida and J. M. Khan, Int. J. Biol. Macromol., 2023, 231, 123237 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |