 Open Access Article
Open Access ArticleFirst-principles study of solvation effects on propylene carbonate reduction on defective hard carbon, for advanced sodium-ion battery anodes
Jin-Song Kima,
Myong-Jin Wona,
Won Pakb,
Kum-Hui Pakb,
Chol Ryua and
Chol-Jun Yu *a
*a
aComputational Materials Design, Faculty of Materials Science, Kim Il Sung University, Taesong District, Pyongyang, Democratic People’s Republic of Korea. E-mail: cj.yu@ryongnamsan.edu.kp
bPyongyang Teacher-Training College, Mangyongtae District, Pyongyang, Democratic People’s Republic of Korea
First published on 26th January 2026
Abstract
Hard carbon (HC) is an attractive anode material for grid-scale sodium-ion batteries (SIBs), but its structural design must be improved through understanding the reaction mechanism. In this work, we investigate the adsorption characteristics of Na atoms with a propylene carbonate (PC) molecule, and the decomposition reaction mechanism of the Na–PC complex on a HC sheet, under PC solvent conditions using first-principles calculations. Using perfect and defective graphene cluster models, which include point defects such as mono-vacancy (MV), di-vacancy (DV) and Stone–Wales (SW), we calculate the binding energies of one or two Na atoms, revealing that the binding strength of the Na atom is in the order of SW < perfect < DV < MV and adding another Na atom lowers the binding energy due to the Na–Na interaction. For adsorption of the Na–PC complex, our calculations demonstrate that binding energies of Na2PC are lower than those of NaPC, where the PC solvent molecules enhance the binding strength of the adsorbate complex on the HC surface. Furthermore, we investigate the decomposition reaction mechanism of the PC molecule by calculating reaction heats and activation barriers, finding that one-electron reduction reactions are endothermic while two-electron ones are exothermic without reaction barriers at room temperature. These findings contribute to our atomistic understanding of the initial charge process of the HC anode and thus aid improvement of the first cycle efficiency of SIBs.
1 Introduction
With increasing concerns about global warming and climate change, there is a growing need to switch over to renewable clean energy sources and electric vehicles (EVs). For this, the development of advanced rechargeable batteries with high performance and low cost has been attracting significant attention over the past decade.1–3 At present, lithium-ion batteries (LIBs) dominate the portable electronics market and show great potential for electrochemical energy storage (EES) and EVs, owing to their high energy density and long lifetime, but they face challenges such as the increasing cost and possible safety concerns such as bursting.4 Due to the high abundance and low cost of Na resources and their superior safety, sodium ion batteries (SIBs) have attracted renewed interest as the most promising alternative to LIBs.5,6 Over the past few years, SIB technologies have rapidly evolved with advanced cathode and anode materials, towards industrial production and commercial application for large-scale EES and EVs.7–9Among the developed anode materials, hard carbon (HC) has been most widely used for commercialized SIBs due to its advantages of relatively high specific capacity (200–500 mAh g−1), low potential plateau below 0.1 V, long cycling life, low cost and environmental friendliness.10–12 In fact, HC has been mostly produced by high-temperature (600–1900 °C) pyrolysis of biomass-derived precursors, such as wood waste, straw, pine cone, fruit peel and so on,13–17 under Ar or N2 flow. In spite of such merits, the initial coulombic efficiency (ICE) of HC anodes in SIBs (30–80%)14,18 is much lower than that of the graphite anode in LIBs, which is a major hurdle to industrial application. The low ICE of the HC anode is generally ascribed to large irreversible Na loss caused by electrolyte decomposition during the formation of a solid electrolyte interphase (SEI),19–22 which originates from a defective structure and large specific area of HC.23–25 The SEI layer comprises small domains of inorganic sodium salts, organic solvents and additives, which are formed on the anode surface upon organic electrolyte decomposition during the first cycle, allowing the transport of Na+ ions and protection of the anode surface.12,20 Together with control of the HC structure, proper selection of sodium salt (NaClO4 or NaPF6) and organic solvent – among ethylene carbonate (EC), propylene carbonate (PC, C4H6O3), diethyl carbonate (DEC), etc. – is crucial in the improvement of ICE and the rate capability of SIBs.26–28 This can be achieved by performing a deep analysis of the HC structure and SEI formation reaction, thereby revealing the underlying mechanism.29
For the HC anode, the sodium storage and SEI formation mechanisms are still in debate, severely hindering further improvement of rate capability and specific capacity.30–33 As Stevens and Dahn proposed, the HC structure is explained as a “house of cards”, which is composed of disordered graphitic domains with randomly oriented defective graphene nanosheets and interstitial nanoporous regions.34,35 In this model, the sodium storage is explained by three processes: Na adsorption on the surface, intercalation into graphitic domains and pore filling.30 In these processes, defects on the graphene sheets, such as mono-vacancy (MV), di-vacancy (DV) and Stone–Wales (SW), play a decisive part in Na adsorption, intercalation and diffusion.36–38 Therefore, somewhat in-depth understanding of the defect influence on the performance improvement has been obtained by a number of theoretical studies.38–44 For instance, Youn et al.39 demonstrated the stable formation of Na clusters with nanometer-size and 3–6 layers between two graphene sheets and suitable micropores with defective graphitic domains in HC, using density functional theory (DFT) calculations. Soto et al.45 performed ab initio molecular dynamics (AIMD) simulations of the sodiated HC to show the SEI formation through decomposition of EC and PF−6 anions on the edge of graphite layers. The migration pathways of Na+ cations through the SEI were identified by performing DFT calculations.45,46
To the best of our knowledge, however, theoretical works on the influence of organic solvent on the SEI formation on HC are rare. In our previous works, we investigated the initial process of SEI formation on HC considering the solvation effect41 and sodiation mechanism of HC considering point defects and PC solvent effects.40 In this work, we continue exploring this effect by identifying the reaction of PC decomposition on the graphene sheet that contains a point defect such as MV, DV and SW and discussing the reaction barriers at finite temperature.
2 Computational method
All first-principle calculations were performed using the all-electron atomic orbital method within the DFT framework as implemented in the NWChem (Version 7.2) package.47 The Gaussian-type basis functions of 6-31G** were used as a basis set and the B3LYP functional48,49 was used for the exchange-correlation (XC) interaction. To account for the PC solvent effect, we used the solvation model based on density (SMD),50 which is a universal continuum model with applicability to any charged and uncharged solute in any solvent as a dielectric medium. For the SMD input parameters of PC solvent, the dielectric constant (dielec) was set to 64.92,51,52 the macroscopic surface tension (solg) was set to 41.28 cal (mol−1·Å−2) and the index of refraction (soln) was 1.419,53 and the hydrogen bond acidity (sola) and hydrogen bond basicity (solb) were set to 0.0 and 0.4, respectively.54 The aromaticity (solc) and electronegative halogenicity (solh) were considered to be 0 from the structural characteristics of PC. The quasi-Newton optimization method was applied for geometry optimizations of the molecular complexes using the DRIVER module with the default convergence criteria; the maximum and root mean square gradient in the coordinates are 0.00045 and 0.00030, and the maximum and root mean square of the Cartesian step are 0.0018 and 0.0012 (see input files in SI). We conducted a search for a transition state (or saddle point) using the Driver module, which allows to calculate the eigenvalues of the Hessian matrix to search for a saddle point structure with only one negative eigenvalue, i.e., imaginary frequency. For connectivity via the intrinsic reaction coordinate (IRC), we used the MEPGS module in the NWChem package, which performs a search for the two critical points on the potential energy surface connected to a saddle point of the molecule (Gonzalez & Schlegel IRC optimization). The frequency/thermal corrections and solvent were applied consistently to IRC optimizations.Since HC is known to have a hybrid structure of stacked and randomly arranged graphene sheets, the HC model can be a graphene sheet with a few layers in the atomic scale.40,41,55 Fig. 1 shows the mechanism of Na storage within the porous structure (A) and Na adsorption with the PC solvent molecule at the surface (B), solvated in the PC electrolyte, which can be ascribed to the adsorption model on the graphene sheet. As the number of stacking layers was known to barely affect the Na absorption in previous work,39 the interaction model of electrolyte with HC is considered with a mono-layer graphene sheet. Considering that there are two modelling methods of the supercell model for graphene – with periodic boundary condition38,43,44 and cluster model56,57 – we chose the cluster model with edges passivated with hydrogen, that is, C48H18, as shown in Fig. 1. The C48H18 cluster model can be considered as having the minimum planar size that can incorporate the PC solvent molecule. Visualization of molecular structures was performed using the VESTA.58
3 Results and discussion
3.1 Na binding characteristics
First, we investigated the binding characteristics of Na atoms on the HC surface modelled by the C48H18 cluster. The total charge of the whole molecular complex, that is, Na atom on C48H18 cluster, is zero under vacuum conditions, due to one Na+ cation adsorption on the graphene sheet within the micropore region or on the HC outer surface, which are negatively charged. Among numerous Na adsorption sites on the graphene sheet without and with a point defect such as MV, DV and SW, the lowest energy site was selected for each graphene sheet after geometry optimizations (see Fig. S1). For the geometry optimizations, we adopted two methods: (1) the constrained optimization method with freezing the terminating hydrogen atoms and (2) the full geometry optimization. During the full geometry optimization, unlike the supercell model using the periodic boundary conditions, the cluster model exhibited in general undesirable graphene sheet distortion due to the finite size of graphene cluster when considering adsorption on the graphene sheet, which can be addressed by immobilizing the terminating H atoms.59Fig. 2 shows the optimized structures of Na-adsorbed C48H18 clusters without and with a vacancy defect such as MV, DV and SW on the graphene sheet. In the cases of perfect and SW-defective models, the planar structures of graphene sheet were observed for both H-fixed and full optimization methods, whereas the swollen and hollow structures at the centre of the graphene cluster, where the Na atom was absorbed, were observed by the full geometry optimization in the cases of MV- and DV-defective graphene sheets, respectively. Similar structural features were observed in the two Na atoms-absorbed HC models, as shown in Fig. 3. During the geometry optimizations, the PC solvation effects were also considered through the SMD solvation model, resulting in a slight increase of Na–graphene and Na–Na distances as shown in Fig. 2 and 3, with the blue-coloured values in parentheses.
To evaluate the binding strength of Na atoms with HC, we calculated the binding energy per Na atom as follows,
 | (1) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 44 and 0.75 eV
44 and 0.75 eV![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 60 obtained using the 4 × 4 and 6 × 6 supercell periodic models, respectively. Under the PC solvent conditions, the Eb value of 1.33 eV is also in reasonable agreement with our previous DFT calculation of 1.23 eV obtained using the 6 × 6 supercell periodic model.40 For justification of the cluster size, we considered different cluster sizes by systematically increasing the number of carbon atoms from 24 to 54 with an interval of 6, confirming that the present size of cluster model is sufficiently reliable for absorption properties and electronic structure (see Table S1 and Fig. S2 and S3, SI). For the SW HC model, meanwhile, the Eb values obtained by the H-fixed optimization method (0.82 and 1.30 eV) were slightly larger than those (0.74 and 1.23 eV) by the full geometry optimization method (under the vacuum and solvent conditions) due to the slight difference between the optimized structures, although they looked similar each other. For the MV and DV HC models, which have quite different structures according to the optimization method, the binding energies of the Na atom (2.30 and 1.10 eV) on the bended graphene sheet resulting from the full optimization under the vacuum conditions, were found to be lower than those (2.39 and 1.35 eV) on the planar graphene sheet resulting from the H-fixed optimization.
60 obtained using the 4 × 4 and 6 × 6 supercell periodic models, respectively. Under the PC solvent conditions, the Eb value of 1.33 eV is also in reasonable agreement with our previous DFT calculation of 1.23 eV obtained using the 6 × 6 supercell periodic model.40 For justification of the cluster size, we considered different cluster sizes by systematically increasing the number of carbon atoms from 24 to 54 with an interval of 6, confirming that the present size of cluster model is sufficiently reliable for absorption properties and electronic structure (see Table S1 and Fig. S2 and S3, SI). For the SW HC model, meanwhile, the Eb values obtained by the H-fixed optimization method (0.82 and 1.30 eV) were slightly larger than those (0.74 and 1.23 eV) by the full geometry optimization method (under the vacuum and solvent conditions) due to the slight difference between the optimized structures, although they looked similar each other. For the MV and DV HC models, which have quite different structures according to the optimization method, the binding energies of the Na atom (2.30 and 1.10 eV) on the bended graphene sheet resulting from the full optimization under the vacuum conditions, were found to be lower than those (2.39 and 1.35 eV) on the planar graphene sheet resulting from the H-fixed optimization.
| Model | Structure | Eb for Na atom (eV) | Eb for Na2 atoms (eV) | ||||
|---|---|---|---|---|---|---|---|
| Vac. | Sol. | Prev. | Vac. | Sol. | Prev. | ||
| a Ref. 44.b Ref. 60.c Ref. 42. | |||||||
| Perfect | Fully optimized | 0.76 | 1.33 | 0.62a, 0.75 (1.14)b | 0.51 | 1.05 | 0.60 (0.93)c |
| H-fixed optimized | 0.76 | 1.33 | 0.54 | 1.16 | |||
| MV | Fully optimized | 2.30 | 2.62 | 2.00a, 1.96 (2.15)b | 1.85 | 2.24 | |
| H-fixed optimized | 2.39 | 2.92 | 1.56 | 2.03 | |||
| DV | Fully optimized | 1.10 | 1.73 | 1.73 (2.00)b | 1.07 | 1.41 | 1.22 (1.60)c |
| H-fixed optimized | 1.35 | 1.88 | 0.87 | 1.27 | |||
| SW | Fully optimized | 0.74 | 1.23 | 0.73 | 1.11 | ||
| H-fixed optimized | 0.82 | 1.30 | 0.78 | 1.20 | |||
We tested different XC functionals for the choosing the B3LYP functional, which is known to sometimes underestimate dispersion interactions. Malyi et al.60 found that the inclusion of dispersion correction (DFT-D) increased the binding energy as shown with the values in parentheses in Table 1. We calculated the binding energies of the Na atom to four different HC sheets using B3LYP+D3/6-31G** and PBE0+D3/def2-TZVP methods, revealing that the inclusion of Grimme’s dispersion (D3) correction slightly increased Eb by 0.07–0.17 eV except for the DV model (−0.1 eV), while the later method clearly increased Eb by 0.19–0.49 eV (Table S2, SI).
The same tendencies were found for the cases under solvent conditions. These indicate that the binding strength of the Na atom with defective HC is weakened by bending of the graphene sheet. By considering the PC solvent effect, the binding energy of Na atoms with HC was found to be enhanced for the perfect and defective graphene sheets compared with those under vacuum conditions. From the view point of the continuum solvation model (also called implicit solvation model), this indicates that the binding strength of the Na atom with HC is enhanced by the electrostatic interaction between the charge distribution of solute and the electric polarization field of the solvent. It is worth noting that the binding strength of the Na atom is in the order of SW < perfect < DV < MV, among which the MV system showed a unique swollen structure by the H-fixed optimization.
For the cases of two Na atoms adsorbed on the graphene sheet, on the other hand, the binding energies per Na atom were found to be lowered compared to those of one Na atom adsorption, as listed in Table 1. This can be attributed to the reduction of Na–graphene binding due to the additional Na–Na interaction, as has been already mentioned in previous works.39,42 To estimate the decreasing effect in binding energy due to the Na–Na interaction, we evaluated the binding energy change as follows,
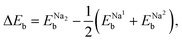 | (2) |
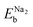 is the binding energy per Na atom in the two Na atoms adsorbed graphene sheet, corresponding to eqn (1) with n = 2, and
is the binding energy per Na atom in the two Na atoms adsorbed graphene sheet, corresponding to eqn (1) with n = 2, and  and
and 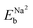 are the binding energies of Na1 and Na2 in the one Na atom adsorbed graphene sheets, respectively. For the perfect and SW-defective graphene sheets, two Na atoms were found to be adsorbed on the centre of two hexagonal and two heptagonal rings with Na–Na distances of 3.75 and 3.51 Å as shown in Fig. 3(a) and (d), respectively. For the fully optimized perfect model under vacuum conditions, the
are the binding energies of Na1 and Na2 in the one Na atom adsorbed graphene sheets, respectively. For the perfect and SW-defective graphene sheets, two Na atoms were found to be adsorbed on the centre of two hexagonal and two heptagonal rings with Na–Na distances of 3.75 and 3.51 Å as shown in Fig. 3(a) and (d), respectively. For the fully optimized perfect model under vacuum conditions, the 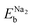 value was calculated to be 0.51 eV in agreement with the previous calculation (0.60 eV),42 with the ΔEb value of 0.14 eV. The
value was calculated to be 0.51 eV in agreement with the previous calculation (0.60 eV),42 with the ΔEb value of 0.14 eV. The 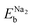 and ΔEb values were calculated to be 0.73 and 0.10 eV for the fully optimized structure under vacuum conditions. On the other hand, the adsorption of two Na atoms on the MV and DV defective graphene sheets occurred on the hexagonal ring and defect-containing large ring as shown in Fig. 3(b) and (c), respectively. For the fully optimized MV and DV cases under vacuum conditions, the
and ΔEb values were calculated to be 0.73 and 0.10 eV for the fully optimized structure under vacuum conditions. On the other hand, the adsorption of two Na atoms on the MV and DV defective graphene sheets occurred on the hexagonal ring and defect-containing large ring as shown in Fig. 3(b) and (c), respectively. For the fully optimized MV and DV cases under vacuum conditions, the  values were found to be 1.85 and 1.07 eV with the ΔEb values of 0.09 and 0.23 eV, respectively. The decreasing tendency of Na2 binding with graphene due to the Na–Na interaction compared to Na binding is in the order of MV < SW < perfect < DV, indicating that the Na–graphene binding near the DV defect is most affected by the Na–Na interaction. In contrast with one Na atom adsorption, the binding strength of Na atoms by the full optimization was found to be stronger than that by the H-fixed optimization for MV and DV systems, implying that the strong Na–Na interaction seriously affects the Na–graphene binding.
values were found to be 1.85 and 1.07 eV with the ΔEb values of 0.09 and 0.23 eV, respectively. The decreasing tendency of Na2 binding with graphene due to the Na–Na interaction compared to Na binding is in the order of MV < SW < perfect < DV, indicating that the Na–graphene binding near the DV defect is most affected by the Na–Na interaction. In contrast with one Na atom adsorption, the binding strength of Na atoms by the full optimization was found to be stronger than that by the H-fixed optimization for MV and DV systems, implying that the strong Na–Na interaction seriously affects the Na–graphene binding.
3.2 Adsorption of Na atoms and PC molecule
To investigate the SEI formation on HC, we then considered the decomposition reaction of the PC solvent molecule on the Na-adsorbed graphene cluster models and thus the initial reaction structures of Na–PC (NaPC) and Na2–PC (Na2PC) complexes on the C48H18 models. Note that the NaPC complex model corresponds to the one electron reaction of Na+ + e− + PC → NaPC, while the Na2PC complex model is responsible for the two electrons reaction of 2Na+ + 2e− + PC → Na2PC. The H-fixed constrained geometry optimizations were performed to get the optimized structures for NaPC and Na2PC complexes.Fig. 4 shows the optimized structures of the NaPC complex on the C48H18 clusters with perfect and defective graphene sheets. To discuss the bonding characteristics between the Na atom and PC molecule as well as the graphene sheet, we measured the Na–PC bond length (dNa–PC or dNa–O) and the Na–graphene distance (dNa–G). For the cases of perfect, MV and SW graphene sheets, the distances of dNa–PC and dNa–G were found to be somewhat similar with small Δd = dNa–PC − dNa–G below 0.16 Å, indicating that the Na atom is well balanced in binding distances between PC solvent and graphene sheet. For the DV-defective graphene sheet, however, the Na atom is more attracted by the defective graphene sheet than by the PC solvent molecule with the clearly larger Δd value of 0.26 Å. The adsorption height of the NaPC complex above the HC sheet dNa–G decreases in the order perfect > SW > MV > DV, a similar trend to that of the isolated Na atom.
Fig. 5 shows the optimized structures of the Na2PC complex on the HC models. For the adsorption of the Na2PC complex, the spatial arrangement of two Na atoms and a PC molecule mainly depends on the adsorption sites of the two Na atoms on the graphene sheet. As mentioned above, the two Na atoms were adsorbed above the centres of two hexagonal and heptagonal rings for the perfect and SW-defective graphene sheets, and therefore, the Na–Na axis of the Na2PC complex is parallel to the graphene sheet with almost the same dNa–G value. For the cases of MV- and DV-defective graphene sheets, however, the Na–Na axis of the Na2PC complex is inclined with different dNa–G values between Na above the defect site (MV or DV) and Na above the hexagonal ring. In fact, the adsorption heights of the Na atom were found to be 2.27 and 2.29 Å above the hexagonal ring, while 2.41 and 2.07 Å above the defective ring for the MV and DV defects, respectively. For the case of the MV HC model, a few C atoms around the defect on the graphene sheet were found to apparently rise upward, resulting in the higher adsorption height of the Na atom than that above the hexagonal ring, but the nearest C–Na distance was found to be the shortest at 2.38 Å, among the four different HC models. On the contrary, the planar feature of the DV-defective graphene sheet was found to be retained, resulting in the lower adsorption height of the Na atom on the DV defect site. When the adsorption height of the Na2PC complex is estimated as the average of Na1– and Na2–graphene distances by 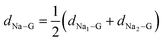 , the adsorption height dNa–G was decreased in the order of MV (2.347 Å) > perfect (2.271 Å) > SW (2.236 Å) > DV (2.173 Å).
, the adsorption height dNa–G was decreased in the order of MV (2.347 Å) > perfect (2.271 Å) > SW (2.236 Å) > DV (2.173 Å).
The binding energy of the NaPC complex with the HC model was calculated as follows,
| Eb = −(ENaPC–G − EG − ENaPC), | (3) |
 | ||
| Fig. 6 Binding energy of NaPC and Na2PC adsorbate complex on the perfect and defective graphene sheets with a point defect such as MV, DV and SW. | ||
To get a meaningful understanding of Na binding, we calculated the charge density differences with a Mulliken charge population analysis. Upon adsorption of a NaPC or Na2PC complex on different graphene sheets, the charge density difference is calculated as follows,
| Δρ(r) = ρNaPC–G(r) − ρNaPC(r) − ρG(r), | (4) |
For the case of NaPC adsorption, the Mulliken charge analysis shows that the charge populations of different graphene models are in the order of perfect (0.77e) > SW (0.72e) > MV (0.68e) > DV (0.62e) (Fig. S4, SI). Note that the Hirshfeld charge analysis gives the same order (Fig. S5, SI). Therefore, it can be said that the presence of defects on the graphene sheet decreases the electron-accepting ability (reduction ability) of the HC surface and thus the electron-donating ability (oxidation ability) of the adsorbate complex NaPC. From the viewpoint of reduction ability, it is expected that at the perfect HC surface more electron accumulation is achieved by adding the PC solvent molecule, resulting in enhanced ionic binding between the neutral NaPC and neutral HC surface. However, considering the initial charging process, which is the Na+ cation adsorption on the negatively charged HC anode resulting in the reaction Na+ + e− → Na, the lower reduction ability of HC is desirable. In fact, the negatively charged HC anode should donate an electron to the Na+ cation of the NaPC cluster during the initial charging process, and thus the HC should be oxidized. Therefore, it is concluded that the DV-defective graphene is the most desirable anode among the four different models due to its lowest reduction ability with the shortest absorption distance of NaPC, which is favourable for donating electrons. This agrees well with the experimental observation when Na atoms are stored in defective or disordered carbon materials.61
Upon adsorption of Na2PC, a larger amount of electrons were transferred from the Na2PC complex to the graphene sheet. From the Mulliken charge analysis, it was found that the charge populations of the graphene sheet are in the order of perfect (1.26e) > DV (1.15e) > SW (1.12e) > MV (1.09e). Therefore, the MV-defective graphene has the lowest reduction ability, as the distortion of a few carbon atoms near the MV defect leads to the shortest C–Na distance. The order of charge populations along the different HC models is different from that upon NaPC adsorption, which is attributed to the overall charge transferring effect depending on the adsorption sites of two Na atoms. For the perfect and SW-defective graphene sheets, the two Na atoms were adsorbed on the two hexagonal rings and two heptagonal rings, for which the Mulliken charge populations were found to be 0.63e and 0.56e, respectively. For the cases of MV- and DV-defective graphene sheets, where one Na atom was adsorbed on the vacancy defect and another Na atom was adsorbed on the hexagonal ring, the overall reduction ability was affected by the Na–Na bond, resulting in the reverse order of Mulliken charge populations compared with those upon NaPC adsorption. The oxidation ability of the PC solvent molecule was found to also be enhanced upon Na2PC adsorption, as the Mulliken charge of the PC molecule became larger: e.g., −0.15e for NaPC versus −0.18e for Na2PC on the MV-defective graphene sheet. The frontier molecular orbitals including the highest occupied molecular orbital (HOMO) and the lowest unoccupied molecular orbital (LUMO) are shown in Fig. S5 and S6 (SI).
3.3 Decomposition reaction mechanism of PC solvent
Based on the initial reaction structures, we investigated the decomposition reaction of the PC solvent molecule on HC under the PC solvation conditions to get an insight into SEI formation. The decomposition reaction of the PC molecule is considered as the reduction reaction of PC by receiving an electron (one-electron reaction) or two electrons (two-electron reaction) from the HC anode, producing a propylene (C3H6) and NaCO3 or sodium carbonate (Na2CO3),62 as depicted in Fig. 8. Through the one-electron reaction pathway, the ring-cleavage reaction is expected to occur, followed by formation of the linear NaPC complex, which is then decomposed into a NaCO3 radical and propylene (CH3–CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH3). The NaCO3 radical subsequently binds with an additional Na+ cation to form sodium carbonate (Na2CO3), which is one of the major SEI components. The one-electron reaction can be described as follows,
CH3). The NaCO3 radical subsequently binds with an additional Na+ cation to form sodium carbonate (Na2CO3), which is one of the major SEI components. The one-electron reaction can be described as follows,| PC + Na+ + e−→NaPC (ring-cleavage) → NaCO3 + C3H6, NaCO3 + Na+ + e− → Na2CO3 | (5) |
In the two-electron reaction pathway, meanwhile, the two C–O bonds of the PC molecule are simultaneously broken, and then Na2CO3 and C3H6 are produced as follows,
| PC + 2Na+ + 2e− → Na2CO3 + C3H6 | (6) |
Although there exist several mechanisms for reactions of producing different SEI components, we considered only the production of Na2CO3 in the present work.
 | ||
| Fig. 8 Reaction pathways for PC decomposition on HC by receiving (a) one electron or (b) two electrons from the graphene sheet. | ||
The geometries of the final products of Na2CO3 and C3H6 were optimized on the different graphene models under the PC solvation conditions. Then we conducted searches for transition states (or saddle points) with only one imaginary frequency for the one-electron reaction [eqn (5)], which correspond to the ring-opening NaPC complexes, using the three starting structures as shown in Fig. 9. By identifying the optimized molecular structures for initial, transition and final states of the reactions, we could determine the activation barriers with the reaction heats, which were calculated by the Gibbs free energy difference between reactants and products by considering the zero-point correction (ZPC) and thermal correction (TC) to the total energy. The solvation effects were also considered through the SMD solvation model.
 | ||
| Fig. 9 Starting structures of the NaPC complex for conducting a search for a transition state with only one imaginary frequency. | ||
Table 2 summarizes the calculation results (see Fig. 10). In the case of the one-electron reaction, the transition states were found in the PC reduction decomposition reaction on the perfect and SW-defective graphene sheet, corresponding to the TS3 state with two C–O bonds breaking in Fig. 9 (see Fig. S7, SI). In the TS structure on the perfect graphene sheet, the nearest C–O bond length of the PC molecule was shifted from 1.45 Å to 3.94 Å. Meanwhile, in the TS structure on the SW graphene sheet, the molecular orientation of C3H6 was directed towards the oxygen atom of NaCO3, which was bound with the hydrogen atom of C3H6 through hydrogen bonding with a bond distance of dO–H = 2.44 Å. With these states, the activation barriers were determined to be 28.25 and 13.47 kcal mol−1 at 0 K while 6.25 and 12.32 kcal mol−1 at 298.15 K for the perfect and SW-defective graphene, respectively. It was found that the adsorption site of the NaCO3 product at final state was different from that of the NaPC reactant at the initial state. In fact, the adsorption site of NaCO3 was found to move to the adjacent hexagonal carbon ring in the case of perfect graphene, while to the middle point between the two neighboring hexagonal rings from the two heptagonal rings in the SW graphene.
| Model | PC + Na+ + e− → NaCO3 + C3H6 (kcal mol−1) | PC + 2Na+ + 2e− → Na2CO3 + C3H6 (kcal mol−1) | ||||||
|---|---|---|---|---|---|---|---|---|
| ΔE (0 K) | ΔG (298.15 K) | ΔE (0 K) | ΔG (298.15 K) | |||||
| TS | FS | TS | FS | IM | FS | IM | FS | |
| Perfect | 28.25 | 22.64 | 6.25 | 4.05 | −34.93 | −35.48 | −42.21 | −40.31 |
| SW | 13.47 | 8.03 | 12.32 | 9.95 | −39.80 | −42.00 | −47.33 | −44.29 |
| MV | 0.0 | 34.08 | 0.0 | 23.90 | −18.25 | −37.18 | −20.99 | −39.58 |
| DV | 0.0 | 13.63 | 0.0 | 5.82 | 2.78 | 0.48 | −1.36 | −0.36 |
 | ||
| Fig. 10 Schematic view for summarizing the proposed PC decomposition on different defect types. Solid (dotted) lines show the one-(two-)electron reactions. | ||
For the cases of MV- and DV-defective graphene sheets, meanwhile, no transition states were found and no moving in the absorption sites of Na atom occurred. This indicates that the PC decomposition on the MV- and DV-defective graphene through one-electron reaction, which is accompanied by simple C–O bond breakage, occurs more easily than on the perfect and SW-defective graphene, where diffusion as well as bond breaking occurs. Therefore, the MV and DV defects are favourable for the formation of a thin and steady SEI layer which could endure the sodiation/desodiation process. The experimental findings well agree with this.37 The facile reactions on the MV- and DV-defective graphene can be associated with the stronger adhesion of the NaPC complex to the graphene with the higher binding energies near the MV and DV defects. Moreover, the second lowest endothermic reaction heat of 5.82 kcal mol−1 at room temperature was found for the DV-defective model, while the lowest endothermic reaction heat of 4.05 kcal mol−1 with the relatively low activation barrier was found for the perfect graphene. For the case of MV-defective graphene, however, the endothermic reaction heat was found to be highest, 23.90 kcal mol−1, at room temperature. Therefore, the one-electron reaction pathway can be said to be favourable on the perfect and DV-defective HC surfaces.
In the two-electron cases, the reactions on all the different HC models were found to be exothermic without transition states, i.e., barrierless reactions, except on the DV-defective graphene at 0 K with an activation barrier. As shown in Table 2 and Fig. 10, the reaction heats at room temperature were calculated to be negative for the perfect, MV- and SW-defective graphene models, but positive for the DV-defective graphene model. On the perfect, MV and SW HC models, the two C–O bonds of the Na2PC complex were firstly broken, resulting in the formation of Na2CO3 and C3H6 (intermediate states: IM), and then Na2CO3 was transferred to the energetically stable adsorption site (see Fig. S8, SI). On the perfect and SW models in particular, the exothermic reaction heat from the breaking of the two C–O bonds of the Na2PC complex was found to account for most of the total reaction heat, and then the endothermic reaction heat for transport of Na2CO3 at room temperature was found to be only 1.90 and 3.04 kcal mol−1, respectively. For the case of the MV model, both the two C–O bonds breaking and the Na2CO3 transfer were found to be exothermic with reaction heats of −20.99 and −18.59 kcal mol−1 at room temperature. In the molecular structure of Na2CO3 on the MV-defective graphene sheet, we observed a bond between the O atom (intervening between two Na atoms) and the C atom near the MV defect with a bond length of 1.37 Å, indicating that robuster SEI can be formed on the MV-defective HC surface.
On the DV-defective graphene sheet, meanwhile, only the two C–O bonds breaking occurred without transfer of Na2CO3. The binding distance between Na2CO3 and DV-defective graphene was found to be very short (1.46 Å), resulting in the formation of robust SEI similarly with MV-defective graphene. For the energetics, the reaction was found to be endothermic at 0 K with a reaction heat of 0.48 kcal mol−1 and activation barrier of 2.78 kcal mol−1, while exothermic at room temperature with reaction heat of −0.36 kcal mol−1. As such, the reaction heat is considerably lower compared with other graphene models, which might be attributed to the deformation of DV defect involved in the PC decomposition reaction. On the DV-defective HC surface, the energy released during the PC reduction decomposition reaction is almost compensated by the energy required for the deformation of carbon atoms near the DV defect, resulting in considerably low reaction heat. On the contrary, the surface deformation (relaxation of C atom positions of graphene) occurs with requirement of little energy on the perfect and SW-defective graphene sheets.
It was known that the strong binding energy between the Na ions and defects lead to low irreversible capacity, because certain Na ions are trapped by the defects and then become irreversible.18 Hence, the HC materials with high concentration of defects are generally associated with insufficient reversible capacity and ICE. With this regard, our work demonstrated that the MV or DV defects induce robust SEI formation with no or low activation barriers, indicating that the concentration tuning of such defects is the key factor for increasing the reversible capacity and ICE. It is worth noting that the low concentration of defects and higher ICE in HC materials can be achieved by slow heating rate during hard carbon synthesis.
4 Conclusions
In this work, we have investigated the binding characteristics of Na atoms in combination with PC molecule and decomposition reactions of the Na–PC complex on different types of HC sheet using perfect and defective graphene cluster models under PC solvent conditions, by performing first-principles calculations. We firstly calculated the binding energies of one or two Na atoms with the perfect and defective graphene sheets, which include a point defect such as MV, DV and SW, revealing that the binding strength of the Na atom was increased in the order SW < perfect < DV < MV and lowered by adding a Na atom due to the Na–Na interaction. Our calculations for adsorption of NaPC and Na2PC complexes demonstrated that the increasing tendency of binding strength was the same as Na atom adsorption, binding energies of Na2PC were lower than those of NaPC, and PC solvent molecules enhanced the binding strength of the adsorbate complex on the HC surface. Moreover, we studied the decomposition reaction mechanism of the PC molecule by calculating the reaction heats and activation barriers for the one-electron reduction reactions, which were found to be endothermic with activation barriers only for the perfect and SW graphene sheets, and the two-electron reduction reactions, which were exothermic without reaction barriers at room temperature. This work highlights the importance of tailoring graphene structure for improving the initial coulombic efficiency and forming the solid-electrolyte interphase of SIBs.Author contributions
Jin-Song Kim performed the calculations and drafted the first manuscript. Myong-Jin Won, Won Pak, Kum-Hui Pak and Chol Ryu assisted with the post-processing of calculation results, and contributed to useful discussions. Chol-Jun Yu developed the original project and supervised the work. All authors reviewed the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information (SI): input files, Cartesian coordinates, Figures for Na adsorption sites, charge population, frontier molecular orbitals, and molecular structures of reactants, intermediates and products during PC decomposition reactions. See DOI: https://doi.org/10.1039/d5ra07677c.Acknowledgements
This work is supported as part of the fundamental research project “Design of New Energy Materials” (No. 2021-12) funded by the State Commission of Science and Technology, DPR Korea. Computations in this work have been done on the HP Blade System C7000 (HP BL460c) that is owned and managed by the Faculty of Materials Science, Kim Il Sung University.References
- D. Wu, H. Li, B. Kang, L. Lu, X. Sun and F. Wu, Appl. Phys. Lett., 2023, 123, 020401 Search PubMed.
- K. M. Tan, T. S. Babu, V. K. Ramachandaramurthy, P. Kasinathan, S. G. Solanki and S. K. Raveendran, J. Energy Storage, 2021, 39, 102591 Search PubMed.
- J. Janek and W. G. Zeier, Nat. Energy, 2016, 1, 16141 CrossRef.
- Y. Yang, E. G. Okonkwo, G. Huang, S. Xu, W. Sun and Y. He, Energy Storage Mater., 2021, 36, 186–212 CrossRef.
- N. Tapia-Ruiz, A. R. Armstrong, H. Alptekin, M. A. Amores, H. Au, J. Barker, R. Boston, W. R. Brant, J. M. Brittain and Y. Chen, et al., J. Phys. Energy, 2021, 3, 031503 CrossRef CAS.
- C. Vaalma, D. Buchholz, M. Weil and S. Passerini, Nat. Rev. Mater., 2018, 3, 18013 Search PubMed.
- R. Usiskin, Y. Lu, J. Popovic, M. Law, P. Balaya, Y. S. Hu and J. Maier, Nat. Rev. Mater., 2021, 6, 1020–1035 CrossRef CAS.
- A. Rudola, A. J. R. Rennie, R. Heap, S. S. Meysami, A. Lowbridge, F. Mazzali, R. Sayers, C. J. Wright and J. Barker, J. Mater. Chem. A, 2021, 9, 8279–8302 RSC.
- J. Deng, W.-B. Luo, S.-L. Chou, H.-K. Liu and S.-X. Dou, Adv. Energy Mater., 2018, 8, 1701428 Search PubMed.
- H. Liu, M. Baumann, X. Dou, J. Klemens, L. Schneider, A.-K. Wurba, M. Haringer, P. Scharfer, H. Ehrenberg, W. Schabel, J. Fleischer, N. von der Aβen and M. Weil, J. Energy Storage, 2022, 56, 105964–105975 Search PubMed.
- Z. Lu, H. Yang, Y. Guo, H. Lin, P. Shan, S. Wu, P. He, Y. Yang, Q.-H. Yang and H. Zhou, Nat. Commun., 2024, 15, 3497 Search PubMed.
- S. Aina, B. Tratnik, A. Vizintin, E. Tchernychova, M. P. Lobera, R. Dominko and M. Bernechea, J. Power Sources, 2024, 610, 234730 CrossRef CAS.
- Z. Tang, R. Zhang, H. Wang, S. Zhou, Z. Pan, Y. Huang, D. Sun, Y. Tang, X. Ji, K. Amine and M. Shao, Nat. Commun., 2023, 14, 6024 CrossRef CAS PubMed.
- Y. Luo, Y. Xu, X. Li, K. Zhang, Q. Pang and A. Qin, Nanomaterials, 2023, 13, 881–895 CrossRef CAS PubMed.
- T. Zhang, J. Mao, X. Liu, M. Xuan, K. Bi, X. L. Zhang, J. Hu, J. Fan, S. Chen and G. Shao, RSC Adv., 2017, 7, 41504–41511 Search PubMed.
- N. Zhang, Q. Liu, W. Chen, M. Wan, X. Li, L. Wang, L. Xue and W. Zhang, J. Power Sources, 2018, 378, 331–337 Search PubMed.
- I. Izanzar, M. Dahbi, M. Kiso, S. Doubaji, S. Komaba and I. Saadoune, Carbon, 2018, 137, 165–173 Search PubMed.
- M. Zhang, Y. Li, F. Wu, Y. Bai and C. Wu, Nano Energy, 2021, 82, 105738 Search PubMed.
- M. Carboni, J. Manzi, A. R. Armstrong, J. Billaud, S. Brutti and R. Younesi, ChemElectroChem, 2019, 6, 1745–1753 CrossRef CAS.
- J. Fondard, E. Irisarri, C. Courrèges, M. R. Palacin, A. Ponrouch and R. Dedryvère, J. Electrochem. Soc., 2020, 167, 070526 Search PubMed.
- H. S. Hirsh, B. Sayahpour, A. Shen, W. Li, B. Lu, E. Zhao, M. Zhang and Y. S. Meng, Energy Storage Mater., 2021, 42, 78–87 CrossRef.
- A. Bouibes, N. Takenaka, K. Kubota, S. Komaba and M. Nagaoka, RSC Adv., 2022, 12, 971–984 RSC.
- L. Xiao, H. Lu, Y. Fang, M. L. Sushko, Y. Cao, X. Ai, H. Yang and J. Liu, Adv. Energy Mater., 2018, 8, 1703238 Search PubMed.
- J.-L. Xia, D. Yan, L.-P. Guo, X.-L. Dong, W.-C. Li and A.-H. Lu, Adv. Mater., 2020, 2000447 CrossRef CAS.
- Z. Li, Y. Chen, Z. Jian, H. Jiang, J. J. Razink, W. F. Stickle, J. C. Neuefeind and X. Ji, Chem. Mater., 2018, 30, 4536–4542 CrossRef CAS.
- G. G. Eshetu, T. Diemant, M. Hekmatfar, S. Grugeon, R. J. Behm, S. Laruelle, M. Armand and S. Passerini, Nano Energy, 2019, 55, 327–340 CrossRef CAS.
- K. Pan, H. Lu, F. Zhong, X. Ai, H. Yang and Y. Cao, Appl. Mater. Int., 2018, 10, 39651–39660 CrossRef CAS PubMed.
- Q. Liu, R. Xu, D. Mu, G. Tan, H. Gao, N. Li, R. Chen and F. Wu, Carbon Energy, 2021, 4, 458–479 Search PubMed.
- K. Schutjajew, T. Tichter, J. Schneider, M. Antonietti, C. Roth and M. Oschatz, Phys. Chem. Chem. Phys., 2021, 23, 11488–11500 Search PubMed.
- M. A. Reddy, M. Helen, A. Groβ, M. Fichtner and H. Euchner, ACS Energy Lett., 2018, 3, 2851–2857 Search PubMed.
- O. Lenchuk, P. Adelhelm and D. Mollenhauer, Phys. Chem. Chem. Phys., 2019, 21, 19378–19390 Search PubMed.
- H. Moriwake, A. Kuwabara, C. A. J. Fisher and Y. Ikuhara, RSC Adv., 2017, 7, 36550–36554 Search PubMed.
- R. Morita, K. Gotoh, M. Fukunishi, K. Kubota, S. Komaba, N. Nishimura, T. Yumura, K. Deguchi, S. Ohki, T. Shimizue and H. Ishida, J. Mater. Chem. A, 2016, 4, 13183–13193 Search PubMed.
- D. A. Stevens and J. R. Dahn, J. Electrochem. Soc., 2000, 147, 1271–1273 Search PubMed.
- D. A. Stevens and J. R. Dahn, J. Electrochem. Soc., 2001, 148, A803–A811 Search PubMed.
- D. Datta, J. Li and V. B. Shenoy, ACS Appl. Mater. Interfaces, 2014, 6, 1788–1795 Search PubMed.
- C. M. Ghimbeu, J. Górka, V. Simone, L. Simonin, S. Martinet and C. Vix-Guterl, Nano Energy, 2018, 44, 327–335 Search PubMed.
- S. Yang, S. Li, S. Tang, D. Shen, W. Dong and W. Sun, Surf. Sci., 2017, 658, 31–37 Search PubMed.
- Y. Youn, B. Gao, A. Kamiyama, K. Kubota, S. Komaba and Y. Tateyama, npj Comput. Mater., 2021, 7, 48 Search PubMed.
- C. Ryu, S.-B. Rim, Y. Kang and C.-J. Yu, RSC Adv., 2023, 13, 5627–5633 RSC.
- C. Ryu, J.-S. Kim, S.-B. Rim, S.-H. Choe and C.-J. Yu, Appl. Surf. Sci., 2022, 573, 151495 Search PubMed.
- Z. Liang, X. Fan, W. Zheng and D. J. Singh, ACS Appl. Mater. Interfaces, 2017, 9, 17076–17084 CrossRef CAS.
- C. Bommier, X. Ji and P. A. Greaney, Chem. Mater., 2019, 31, 658–677 Search PubMed.
- E. Olsson, G. Chai, M. Doveb and Q. Cai, Nanoscale, 2019, 11, 5274–5284 RSC.
- F. A. Soto, A. Marzouk, F. El-Mellouhi and P. B. Balbuena, Chem. Mater., 2018, 30, 3315–3322 Search PubMed.
- H. Yildirim, A. Kinaci, M. K. Y. Chan and J. P. Greeley, ACS Appl. Mater. Interfaces, 2015, 7, 18985–18996 CrossRef CAS PubMed.
- E. Aprà, E. J. Bylaska, W. A. de Jong, N. Govind, K. Kowalski, T. P. Straatsma, M. Valiev, H. J. J. van Dam, Y. Alexeev and J. Anchell, et al., J. Chem. Phys., 2020, 152, 184102 CrossRef PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- C. Li, W. Yang and R. G. Parr, Phys. Rev. B:Condens. Matter Mater. Phys., 1988, 37, 785–789 CrossRef PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 CrossRef CAS PubMed.
- A. Ponrouch, E. Marchante, M. Courty, J.-M. Tarascon and M. R. Palacín, Energy Environ. Sci., 2012, 5, 8572–8583 Search PubMed.
- H. Jia, H. Onishi, N. von Aspern, U. Rodehorst, K. Rudolf, B. Billmann, R. Wagner, M. Winter and I. Cekic-Laskovic, J. Power Sources, 2018, 397, 343–351 CrossRef CAS.
- D. R. Lide, CRC Handbook of Chemistry and Physics, 89th Edition, CRC Press, Boca Raton, FL, Internet Version 2009 Search PubMed.
- M. J. Kamlet, J.-L. M. Abboud, M. H. Abraham and R. W. Taft, J. Org. Chem., 1983, 48, 2877–2887 Search PubMed.
- P.-C. Tsai, S.-C. Chung, S.-K. Linb and A. Yamada, J. Mater. Chem. A, 2015, 3, 9763–9768 Search PubMed.
- R. Morita, K. Gotoh, M. Fukunishi, K. Kubota, S. Komaba, N. Nishimura, T. Yumura, K. Deguchi, S. Ohki, T. Shimizue and H. Ishida, J. Mater. Chem. A, 2016, 4, 13183–13193 Search PubMed.
- A. Mohajeri and A. Omidvar, Synth. Metals, 2018, 241, 39–46 Search PubMed.
- K. Momma and F. Izumi, J. Appl. Crystallogr., 2011, 44, 1272–1276 Search PubMed.
- J. Van der Mynsbrugge, S. L. C. Moors, K. D. Wispelaere and V. V. Speybroeck, ChemCatChem, 2014, 6, 1906–1918 Search PubMed.
- O. I. Malyi, K. Sopiha, V. V. Kulish, T. L. Tan, S. Manzhos and C. Persson, Appl. Surf. Sci., 2015, 333, 235–243 CrossRef CAS.
- A. Ponrouch, A. R. Goni and M. R. Palacín, Electrochem. Commun., 2013, 27, 85–88 CrossRef CAS.
- A. N. Dey and B. P. Sullivan, J. Electrochem. Soc., 1970, 117, 222–224 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |






