Open Access Article
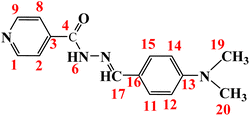
Open Access ArticleSpectroscopic, computational, and biological activity studies of bivalent metal complexes of (E)-N′-(4-(dimethylamino)benzylidene) isonicotinohydrazide
Osama'a A. Y. Al-Samrai a,
Mohamed E. Eissab,
Khalid Tuama Abdullahc,
Tarek A. Yousef*b,
Mona H. Alhalafid and
Ahmed S. M. Al-Janabi
a,
Mohamed E. Eissab,
Khalid Tuama Abdullahc,
Tarek A. Yousef*b,
Mona H. Alhalafid and
Ahmed S. M. Al-Janabi *e
*e
aDepartment of Chemistry, College of Education for Pure Sciences, University of Samarra, Samarra, Iraq
bCollege of Science, Chemistry Department, Imam Mohammad Ibn Saud Islamic University (IMSIU), Riyadh 11623, Saudi Arabia
cDepartment of Chemistry, College of Education for Pure Science, University of Diyala, Diyala, Iraq
dDepartment of Chemistry, College of Science Al-Zulfi, Majmaah University, Al-Majmaah 11952, Saudi Arabia
eDepartment of Chemistry, College of Science, Tikrit University, Tikrit, Iraq
First published on 22nd January 2026
Abstract
(E)-N′-(4-(Dimethylamino)benzylidene) isonicotinohydrazide (HL) ligand with bidentate O, N-donor configuration and its coordination complexes with Zn(II), Cd(II), and Pt(II) metal centers were synthesized. All substances were characterized using elemental analysis, molar conductivity, thermal analysis, NMR (1H and 13C), and FT-IR spectral data. The HL ligand was coordinated as a bidentate chelating ligand via the oxygen of the carbonyl group and the nitrogen of the azomethine group. The thermal properties of the compounds were assessed by simultaneous thermogravimetric-differential thermal analysis measurements, and the findings related to the hypothesized structures. The structures were theoretically examined using density functional theory computations with the B3LYP functional and with the 6-31+G(d,p) basis set for C, H, N, and O atoms, and the LANL2DZ effective core potential for the metal centers (Zn, Cd, and Pt) for the coordination compounds. Molecular electrostatic potential was computed to examine the reactive characteristics of the compounds. The compounds were evaluated in vitro for their antibacterial effects on Escherichia coli and Staphylococcus aureus. The Pt(II) complex displayed the highest antibacterial effect, whereas the Zn(II) complex exhibited moderate inhibitory activity, and the Cd(II) complexes were less active.
1 Introduction
Hydrazones are a class of organic compounds characterized by the presence of the (–NH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–) functional group, typically synthesized via the acid-catalyzed condensation of hydrazides with aldehydes or ketones.1 The general formula is R1R2C
C–) functional group, typically synthesized via the acid-catalyzed condensation of hydrazides with aldehydes or ketones.1 The general formula is R1R2C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–NR3R4, where the R groups can be alkyl, aryl, or most importantly, acyl (giving rise to acyl hydrazones, R′–C(O)–NH–N
N–NR3R4, where the R groups can be alkyl, aryl, or most importantly, acyl (giving rise to acyl hydrazones, R′–C(O)–NH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CR2).1–3 The significance of hydrazones in coordination chemistry stems from their ability to act as multidentate ligands with a multitude of potential donor atoms: the imine nitrogen (N-imine), the carbonyl oxygen (O-carbonyl), the amide oxygen (O-amide), and, in tailored derivatives, additional groups like pyridyl nitrogen or phenolic oxygen.4–10 This donor versatility allows them to form stable complexes with almost every metal in the periodic table. A key feature of hydrazone ligands is their tautomerism. They can exist in an amide form (R–C(O)–NH–N
CR2).1–3 The significance of hydrazones in coordination chemistry stems from their ability to act as multidentate ligands with a multitude of potential donor atoms: the imine nitrogen (N-imine), the carbonyl oxygen (O-carbonyl), the amide oxygen (O-amide), and, in tailored derivatives, additional groups like pyridyl nitrogen or phenolic oxygen.4–10 This donor versatility allows them to form stable complexes with almost every metal in the periodic table. A key feature of hydrazone ligands is their tautomerism. They can exist in an amide form (R–C(O)–NH–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CR2) or an iminol form (R–C(OH)
CR2) or an iminol form (R–C(OH)![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–N
N–N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CR2) (Fig. 1).11
CR2) (Fig. 1).11
Metal coordination often selects for and stabilizes one tautomer over the other, influencing the electronic properties of the resulting complex. Furthermore, the amide proton (–NH–) is acidic. It can be deprotonated upon coordination or by reaction with base, converting the ligand from neutral to anionic and dramatically enhancing its chelating ability and the stability of the resulting complex.5,6,12–15 The bonding modes of hydrazones are diverse, but the most common involve neutral bidentate chelation via the imine nitrogen and the carbonyl oxygen, forming a stable five- or six-membered chelate ring, or deprotonated bidentate chelation after deprotonation and becoming monoanionic, often creating a highly stable six-membered chelate ring with an [O, N] donor set. This mode is ubiquitous with bivalent transition metals. Also, coordination can be mono- or polydentate (this is a common motif for stabilizing the high oxidation states in metal ions).12–15
Hydrazone ligands and their complexes have numerous practical applications in various fields, including biological and medicinal applications (such as anticancer agents, for their antimicrobial properties, and for their antioxidant activity).6,7 They are also excellent chromogenic and fluorogenic agents, enabling the development of sensitive optical sensors for metal ions.9–13 They also serve as suitable scaffolds for catalysis and materials science, acting as effective catalysts for oxidation reactions and organic transformations.3,16–18 The stable metal-chelate environment provided by deprotonated hydrazones makes them suitable for various applications. As part of our interest in oxygen- and nitrogen-containing ligands,19–24 we herein report the synthesis of hydrazone ligand complexes with bivalent metal centers and characterization by different spectroscopic techniques. Furthermore, density functional theory (DFT) calculations were performed on all the compounds, and molecular electrostatic potential (MEP) surfaces were calculated to determine the reactivity centers. Additionally, we evaluated the antibacterial activity of the prepared complexes against Escherichia coli and Staphylococcus aureus species.
2 Experimental section
2.1 General methods
All chemicals and solvents used in the preparation were used without purification. Melting point apparatus type (Stuarts SMP50). Infrared spectra of the ligand and its complexes were recorded using an FTIR-IRXross by Shimadzu in the region between 400 and 4000 cm−1 using Quest ATR Crystal Diamond. The molar conductivity of the prepared complexes was measured using a 10−3 mol L−1 solution in dimethyl sulfoxide (DMSO) at room temperature (25 °C) using a digital conductivity meter (CD-2005). 1H-NMR and 13C-NMR spectra were measured with a Bruker Avance III spectrometer (300 MHz) using DMSO-d6 solvent. Thermogravimetric analysis (TG) and differential thermal analysis (DTA) were conducted using a DTG-60 Shimadzu instrument. The samples were heated at a rate of 10 °C min−1 under an inert nitrogen atmosphere using platinum pans.2.2 Synthesis of Schiff base ligand (E)-N′-(4-(dimethylamino)benzylidene) isonicotinohydrazide (HL)
The ligand (E)-N′-(4-(dimethylamino)benzylidene) isonicotinohydrazide (HL) was prepared by a condensation reaction between 4-(dimethylamino)benzaldehyde (DMAB) and isoniazid (INH). INH (3.64 mmol, 0.50 g) was added to a 50-mL round-bottom flask containing 25 mL of ethanol, followed by heating and stirring for 15 minutes. After that, a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio of DMAB (3.64 mmol, 0.54 g) was dissolved in 10 mL of ethanol containing a few drops of glacial acetic acid, resulting in a clear yellow solution. The mixture was refluxed at 70 °C for 3 hours, after which yellow crystals were formed. The mixture was then left to evaporate at room temperature, yielding a solid product, which was then washed with ether and dried in a vacuum oven. The purity of the compound was confirmed using thin-layer chromatography (TLC).
1 molar ratio of DMAB (3.64 mmol, 0.54 g) was dissolved in 10 mL of ethanol containing a few drops of glacial acetic acid, resulting in a clear yellow solution. The mixture was refluxed at 70 °C for 3 hours, after which yellow crystals were formed. The mixture was then left to evaporate at room temperature, yielding a solid product, which was then washed with ether and dried in a vacuum oven. The purity of the compound was confirmed using thin-layer chromatography (TLC).
HL: yellow crystalline solid. Product yield 98% (0.96 g), m.p. 206 °C, chemical formula C15H16N4O. FT-IR (ATR, cm−1): 3388.9 ν(N–H), 3192.2 ν(C–H) aromatic, 2939.5, 2906.7 ν(C–H) aliphatic, CH3, 2906.7 ν(C–H) aliphatic, 1662.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) amide I, 1602.9 ν(C
O) amide I, 1602.9 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine, 1589.3 ν(C
N) imine, 1589.3 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) aromatic, 1546.9, 1523.8 ν(C–N) Car–N, ν(N–H) amide II, 1325.1 ν(C–N) Car–N, 1230.6 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.82 (s, 1H, H6, NH), 8.78 (d, JHH = 8.0 Hz, 2H, H1,9), 8.35 (s, 1H, H17, N
C) aromatic, 1546.9, 1523.8 ν(C–N) Car–N, ν(N–H) amide II, 1325.1 ν(C–N) Car–N, 1230.6 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.82 (s, 1H, H6, NH), 8.78 (d, JHH = 8.0 Hz, 2H, H1,9), 8.35 (s, 1H, H17, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.83 (d, JHH = 8.0 Hz, 2H, H2,8), 7.58 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 8.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.55 (C4), 152.16 (C13), 150.73 (C1, C9), 150.35 (C17), 141.30 (C3), 129.16 (C11, C15), 121.96 (C2, C8), 121.65 (C16), 112.23 (C12, C14), 40.20 (C19, C20).
CH), 7.83 (d, JHH = 8.0 Hz, 2H, H2,8), 7.58 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 8.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.55 (C4), 152.16 (C13), 150.73 (C1, C9), 150.35 (C17), 141.30 (C3), 129.16 (C11, C15), 121.96 (C2, C8), 121.65 (C16), 112.23 (C12, C14), 40.20 (C19, C20).
2.3 Synthesis of [Zn(HL)(Cl)2]·H2O (1)
A colorless solution of zinc chloride hydrate (ZnCl2·4H2O) (0.51 mmol, 0.106 g) in ethanol (10 mL) was added to a hot yellow solution of ligand HL (0.51 mmol, 0.13 g) in ethanol (15 mL). The resulting mixture was refluxed in a water bath at 50 °C for 20 hours, affording a light-orange solution. The reaction mixture was subsequently allowed to evaporate slowly at ambient temperature. Upon complete removal of the solvent, a solid product was obtained, washed thoroughly with diethyl ether, and dried under vacuum. The obtained complex was isolated as an orange precipitate (m.p. 244 °C, yield 0.18 g, 83%).(1) Orange. Product yield 84% (0.18 g), m.p. 244 °C, chemical formula C15H16Cl2N4OZn·H2O, molar conductivity in DMSO solvent: 5.65 (Ω−1 cm−1 mol−1). FT-IR (ATR, cm−1): 3375.4 ν(N–H), 3197.9, 3041.7 ν(C–H) aromatic, 2910.5 ν(C–H) aliphatic, 2868.1, 2816 ν (C–H) aliphatic CH3, 1656.8 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), amide I, 1611 ν(C
O), amide I, 1611 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine, 1595.1, 1525.6 ν(C
N) imine, 1595.1, 1525.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) aromatic, 1525.6 ν(N–H) amide II, 1546.9 ν(C–N) Car–N, 1232.5 ν(C–N) Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.82 (s, 1H, H6, NH), 8.78 (d, JHH = 12.0 Hz, 2H, H1,9), 8.34 (s, 1H, H17, N
C) aromatic, 1525.6 ν(N–H) amide II, 1546.9 ν(C–N) Car–N, 1232.5 ν(C–N) Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.82 (s, 1H, H6, NH), 8.78 (d, JHH = 12.0 Hz, 2H, H1,9), 8.34 (s, 1H, H17, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.57 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.42 (C4), 152.18 (C17), 150.57 (C1, C9), 150.44 (C13), 141.71 (C3), 129.18 (C11, C15), 122.21 (C2, C8), 121.60 (C16), 112.24 (C12, C14), 40.22 (C19, C20).
CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.57 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.42 (C4), 152.18 (C17), 150.57 (C1, C9), 150.44 (C13), 141.71 (C3), 129.18 (C11, C15), 122.21 (C2, C8), 121.60 (C16), 112.24 (C12, C14), 40.22 (C19, C20).
Complexes [Cd(HL)(Cl)2] (2), [Cd(HL)(NO3)2]·2H2O (3), and [Pt(HL)Cl2]·0.5H2O (4) were prepared and isolated using a similar method.
(2) Dark brown. Product yield 80% (0.09 g), m.p. 296.1 °C, chemical formula C15H16CdCl2N4O, molar conductivity in DMSO solvent: 7.35 (Ω−1 cm−1 mol−1). FT-IR (ATR, cm−1): 3331.1 ν(N–H), 3182.6 ν(C–H) aromatic, 3034.0 ν(C–H) aliphatic, 2906.7 ν(C–H) aliphatic CH3, 1633.7 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) amide I, 1610.6 ν(C
O) amide I, 1610.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine, 1595.1, 1579.7 ν(C
N) imine, 1595.1, 1579.7 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) aromatic, 1525.7 ν(N–H) amide II, 1498.7 ν(C–N), Car–N, 1228.7 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.81 (s, 1H, H6, NH), 8.79 (d, JHH = 12.0 Hz, 2H, H1,9), 8.33 (s, 1H, H17, N
C) aromatic, 1525.7 ν(N–H) amide II, 1498.7 ν(C–N), Car–N, 1228.7 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.81 (s, 1H, H6, NH), 8.79 (d, JHH = 12.0 Hz, 2H, H1,9), 8.33 (s, 1H, H17, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.57 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.39 (C4), 152.19 (C17), 150.68 (C1, C9), 150.45 (C13), 141.77 (C3), 129.18 (C11, C15), 122.20 (C2, C8), 121.58 (C16), 112.25 (C12, C14), 40.24 (C19, C20).
CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.57 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.39 (C4), 152.19 (C17), 150.68 (C1, C9), 150.45 (C13), 141.77 (C3), 129.18 (C11, C15), 122.20 (C2, C8), 121.58 (C16), 112.25 (C12, C14), 40.24 (C19, C20).
(3) Orange. Product yield 91% (0.08 g), m.p. 213.6 °C, chemical formula C15H16CdN6O7·2H2O, molar conductivity in DMSO solvent: 6.42 (Ω−1 cm2 mol−1). FT-IR (ATR, cm−1): 3396.6 ν(N–H), 3230.8 ν(C–H) aromatic, 3097.7 ν(C–H) aliphatic, 2918.3, 2862.4 ν(C–H) aliphatic CH3, 1627.9 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) amide I, 1610.6 ν(C
O) amide I, 1610.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine, 1581.6 ν(C
N) imine, 1581.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) aromatic, 1525.7 ν(N–H) amide II; ν(N
C) aromatic, 1525.7 ν(N–H) amide II; ν(N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) NO3, 1546.9 ν(C–N) Car–N, 1296.2, 1224.7 ν(N–O) NO3, 1226.7 ν(C–N) Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.86 (s, 1H, H6, NH), 8.76 (s, 2H, H1,9), 8.33 (s, 1H, H17, N
O) NO3, 1546.9 ν(C–N) Car–N, 1296.2, 1224.7 ν(N–O) NO3, 1226.7 ν(C–N) Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 11.86 (s, 1H, H6, NH), 8.76 (s, 2H, H1,9), 8.33 (s, 1H, H17, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.56 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.24 (C4), 152.19 (C17), 150.70 (C1, C9), 150.47 (C13), 141.70 (C3), 129.18 (C11, C15), 122.21 (C2, C8), 121.57 (C16), 112.24 (C12, C14), 40.20 (C19, C20).
CH), 7.86 (d, JHH = 12.0 Hz, 2H, H2,8), 7.56 (d, JHH = 12.0 Hz, 2H, H11,15), 6.76 (d, JHH = 12.0 Hz, 2H, H12,14), 2.98 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 161.24 (C4), 152.19 (C17), 150.70 (C1, C9), 150.47 (C13), 141.70 (C3), 129.18 (C11, C15), 122.21 (C2, C8), 121.57 (C16), 112.24 (C12, C14), 40.20 (C19, C20).
(4) Reddish orange. Product yield 92% (0.12 g), m.p 322.8 °C, chemical formula C15H16Cl2N4OPt, molar conductivity in DMSO solvent: 10.05 (Ω−1 cm2 mol−1). FTIR (ATR, cm−1): 3516.2 ν(N–H), 3099.6 ν(C–H) aromatic, 3024.4 ν(C–H) aliphatic, 2902.8, 2812.2 ν(C–H) aliphatic CH3, 1674.2 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) amide I, 1608.6 ν(C
O) amide I, 1608.6 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine, 1591.2 ν(C
N) imine, 1591.2 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) aromatic, 1525.6 ν(N–H) amide II, 1363.6 ν(C–N), Car–N, 1232.5 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 12.16 (s, 1H, H6, NH), 9.14 (d, JHH = 12.0 Hz, 2H, H1,9), 8.38 (s, 1H, H17, N
C) aromatic, 1525.6 ν(N–H) amide II, 1363.6 ν(C–N), Car–N, 1232.5 ν(C–N), Calp–N. 1H-NMR (300 MHz, DMSO-d6, δ ppm): 12.16 (s, 1H, H6, NH), 9.14 (d, JHH = 12.0 Hz, 2H, H1,9), 8.38 (s, 1H, H17, N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CH), 7.98 (d, JHH = 12.0 Hz, 2H, H2,8), 7.58 (d, JHH = 12.0 Hz, 2H, H11,15), 6.78 (d, JHH = 12.0 Hz, 2H, H12,14), 2.99 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 159.85 (C4), 152.88 (C17), 152.28 (C1, C9), 150.03 (C13), 143.87 (C3), 129.28 (C11, C15), 124.52 (C2, C8), 121.44 (C16), 112.27 (C12, C14), 40.27 (C19, C20).
CH), 7.98 (d, JHH = 12.0 Hz, 2H, H2,8), 7.58 (d, JHH = 12.0 Hz, 2H, H11,15), 6.78 (d, JHH = 12.0 Hz, 2H, H12,14), 2.99 (s, 6H, H19,20, 2CH3). 13C-NMR (75 MHz, DMSO-d6, δ ppm): 159.85 (C4), 152.88 (C17), 152.28 (C1, C9), 150.03 (C13), 143.87 (C3), 129.28 (C11, C15), 124.52 (C2, C8), 121.44 (C16), 112.27 (C12, C14), 40.27 (C19, C20).
2.4 DFT study
DFT is a computational method used to calculate molecular system properties such as energy, structure, vibrational frequencies, and chemical reactions. It uses electron density, focusing on the spatial distribution of electrons.25,26 All calculations were performed using the Gaussian 16 program.27 The ligand and its complexes were optimized using DFT with the B3LYP functional and the 6-31+G(d,p) basis set for ligand atoms C, H, N, and O,28 and Lanl2dZ (Los Alamos National Laboratory Double Zeta) effective core potential (ECP) basis set for the central metal ion (Zn, Cd, and Pt). This function effectively represents the electronic structure of atoms and molecules in quantum chemistry calculations. Lanl2dZ is particularly useful for studying molecules containing heavy transition metals. The B3LYP functional (Becke, 3-parameter, Lee–Yang–Parr) has aided researchers in achieving satisfactory geometries for transition metal complexes with low computational costs.29–332.5 Antibacterial activity
The antibacterial activities of the synthesized compounds were screened against two pathogenic bacterial species (Staphylococcus aureus and Escherichia coli) using the disc diffusion method at 10−2, 10−3, 10−4, 10−5, and 10−6 M of DMSO solutions of the tested compounds. The zone of inhibition values (in mm) of the free ligand and its complexes against the bacterial strains were measured, and the results were compared with those for the standard streptomycin. For more details, see the SI (Exp. SI 1).3 Results and discussion
Ligand HL was prepared by a condensation reaction between DMAB and INH to afford a yellow crystalline solid in high yield (98%) (Scheme 1). The treatment of the prepared ligand (HL) with metal salts, ZnCl2·4H2O, CdCl2·H2O, Cd(NO3)2·4H2O, and K2PtCl4, afforded a colored ppt in high yields (80–93%) (Scheme 1). TLC verified the purity of the ligand and its complexes.The molar conductivity data are indicative of the non-electrolytic character of all four coordination complexes,34 in accordance with the coordination of the ligand in its natural form. The results of the physical and spectroscopic characterizations are due to the prepared ligand (HL) being coordinated in a bidentate manner through the oxygen of the carbonyl group and the nitrogen of the azomethine group to give a tetrahedral geometry around Zn(II) and Cd(II) in complexes (1) and (2), whereas it gave an octahedral geometry around Cd(II) in complex (3) and a square planar geometry around Pt(II) ion in complex (4).
3.1 Thermal analysis
Thermal analysis of samples was performed under nitrogen to prevent oxidation, ensuring observed events reflected material properties. Measurements were made using a platinum crucible for its excellent thermal conductivity and stability. A controlled heating rate of 10 °C min−1 was applied from room temperature to 1000 °C, enabling precise tracking of mass loss and thermal transitions. This method comprehensively characterizes thermal stability, decomposition, and composition, providing valuable data on material thermal properties (Table 1). The ligand (HL) initially lost water, forming ZnO and CdO residues in complexes. [Zn(HL)Cl2]·H2O showed multi-step decomposition, releasing chlorine and organic fragments. [Cd(HL)(NO3)2]·2H2O lost water hydrate35 and nitrates, yielding stable cadmium oxide. [Pt(HL)Cl2]·0.5H2O demonstrated platinum oxide stability after ligand decomposition. Overall, thermal behavior is influenced by chemical structure and metal coordination, offering insights for applications.36,37| Compound | TGA | DTA max | Mass loss (%) | Assignment | Residues |
|---|---|---|---|---|---|
| Onset–endset (°C) | (°C) | Exp. (calc.) | |||
| HL | 146.50–173.66 | 133.86 | 6.76 (6.71) | H2O | |
| 301.63–345.21 | 411.12 | 72.16 (78.36) | C13H12N3 | ||
| 667.73–846.83 | 668.43 | 16.31 (16.42) | C2H6N | ||
| [Zn(HL)Cl2]·H2O (1) | 114.42–145.58 | 93.23 | 4.23 (4.26) | H2O | |
| 256.90–300.32 | 284.17 | 18.38 (18.47) | C5H4N | ||
| 332.02–312.55 | 406.57 | 16.18 (16.77) | Cl2 | ||
| 601.00–761.53 | 629.40 | 42.16 (41.22) | C10H12N3 | ||
| 19.05 (19.25) | ZnO | ||||
| [Cd(HL)Cl2] (2) | 301.76–319.28 | 317.41 | 26.36 (26.37) | C6H5N3 | |
| 362.43–372.00 | 378.6 | 9.68 (9.75) | C2H6N | ||
| 539.52–587.53 | 440.9 | 15.88 (15.69) | Cl2 | ||
| 639.53–703.79 | 786.41 | 19.79 (19.73) | C7H5 | ||
| 28.29 (28.44) | CdO | ||||
| [Cd(HL)(NO3)2]·2H2O (3) | 37.73–112.37 | 152.04 | 6.66 (6.66) | 2H2O | |
| 251.56–290.18 | 284.77 | 22.82 (22.93) | 2NO3 | ||
| 356.88–371.72 | 362.68 | 16.56 (16.48) | C7H5 | ||
| 380.72–438.27 | 399.07 | 22.19 (22.21) | C6H6N3 | ||
| 711.56–748.54 | 615.64 | 8.15 (8.15) | C2H6N | ||
| 23.62 (23.74) | CdO | ||||
| [Pt(HL)Cl2]·0.5H2O (4) | 28.83–76.12 | 88.51 | 1.68 (1.65) | 0.5H2O | |
| 233.54–273.06 | 171.45 | 8.20 (8.10) | C2H6N | ||
| 275.02–331.26 | 380.16 | 13.25 (13.04) | Cl2 | ||
| 433.60–668.77 | 424.62 | 36.56 (38.70) | C13H12N3 | ||
| 40.31 (38.85) | PtO |
The ligand exhibits three main thermal events (Fig. 2A). The first event (6.76%) corresponds to the loss of water (H2O) between 56.72 and 179.16 °C. The second event shows a significant mass loss (72.16%) related to the decomposition of the structure (C13H12N3), occurring at a peak temperature of 411.12 °C. The final thermal event leads to a smaller mass loss of 16.31% attributed to further decomposition of (C2H6N) at 668.43 °C. The [Zn(HL)Cl2]·H2O (1) complex (Fig. 2B) also exhibits water loss (4.23%) in the initial temperature range (31.43–156.73 °C). The subsequent mass losses (18.38% and 16.18%) correspond to the decomposition of C5H4N and chlorine gas (Cl2), respectively. The major decomposition event (42.16%) at 629.40 °C reflects the breakdown of the ligand complex (C10H12N3). Residual material (19.05%) is identified as ZnO.
For [Cd(HL)Cl2] (2) complex (Fig. 2C), the initial step shows a significant mass loss (26.36%) attributed to C6H5N3, and the complex shows excellent thermal stability up to approximately 265 °C, after which smaller losses (9.68% for C2H6N and 15.88% for Cl2) occur. The final decomposition event (19.79%) indicates the presence of C7H5. A residue of 28.29% is attributed to CdO, suggesting stability of cadmium oxide after decomposition.
In the thermal analysis of [Cd(HL)(NO3)2]·2H2O (3) (Fig. 2D), the first mass loss (6.66%) corresponds to water loss at lower temperatures. This is followed by significant mass losses due to the decomposition of nitrates (22.82% for 2NO3) and additional organic components (16.56% for C7H5 and 22.19% for C6H6N3). Finally, the compound loses 8.15% of C2H6N with a resultant residue of 23.62% as CdO. The platinum complex (4) (Fig. 2E) displays mass loss below 100 °C, indicating loss of water of crystallization. The second thermal event starts only above 97 °C and is characterized by a relatively strong endothermic DTA peak accompanied by about 8.20% mass loss, which corresponds to the evolution of a small organic fragment, C2H6N, followed by Cl2 loss (13.25%). The major decomposition (36.56%) occurs at higher temperatures, indicating a breakdown of C13H12N3, with a final residue of 40.31% corresponding to PtO, which signifies the stability of platinum oxide after thermal degradation.
3.2 DFT study of prepared compounds
Optimized geometries of the studied complexes were obtained via DFT, showing coordination between metal centers and ligand donor atoms, with bond parameters aligning with experimental values (Fig. 3). The HOMO–LUMO gap analysis supported the stability of these complexes (Fig. 4), which provided insights into their electronic and reactive behavior. Molecular orbital analysis revealed strong overlaps between metal d-orbitals and ligand orbitals, confirming effective electron delocalization and metal–ligand interactions (Table 2).3.3 Infrared spectra (theoretical and experimental)
Theoretical IR spectra, generated through DFT calculations, were compared with experimental data to verify ligand coordination. Shifts in vibrational frequencies, particularly in C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, C
O, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, and M–O/M–N stretching modes, confirmed complex formation.28 The agreement between theoretical and experimental spectra supports the computational models used and further validates the observed vibrational characteristics of the complexes.
N, and M–O/M–N stretching modes, confirmed complex formation.28 The agreement between theoretical and experimental spectra supports the computational models used and further validates the observed vibrational characteristics of the complexes.
Table 3 presents the experimental ATR-IR vibrational frequencies of the free ligand and its metal complexes alongside the corresponding DFT-calculated values, both unscaled and scaled. A close agreement is observed between the experimental and scaled computational data, confirming the validity of the theoretical approach. The ATR-FT-IR spectra of the free ligand (HL) and its metal complexes [Zn(HL)Cl2]·H2O (1), [Cd(HL)Cl2] (2), [Cd(HL)(NO3)2]·2H2O (3) and [Pt(HL)Cl2]·0.5H2O (4) were recorded in the range of 400–4000 cm−1 (Fig. SI1–SI4) and compared with the corresponding theoretical vibrational frequencies obtained by DFT calculations. The free ligand was calculated at the B3LYP/6-31+G(d,p) level, while the metal complexes were optimized using a GEN basis set, applying 6-31+G(d,p) for C, H, N, O atoms and the LanL2DZ effective core potential for Zn, Cd, and Pt.29 Since DFT methods typically overestimate harmonic vibrational frequencies, the calculated values were scaled by the recommended factor of 0.964, from the Computational Chemistry Comparison and Benchmark Database (CCCBDB, NIST), which resulted in excellent agreement with the experimental data.30–33
| Assignment | HL | [Zn(HL)Cl2]·H2O | [Cd(HL)Cl2] | [Cd(HL)(NO3)2]·2H2O | [Pt(HL)Cl2]·0.5H2O | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ATR | Calculated | ATR | Calculated | ATR | Calculated | ATR | Calculated | ATR | Calculated | ||||||
| Exp. | (Un) | (Sc) | Exp. | (Un) | (Sc) | Exp. | (Un) | (Sc) | Exp. | (Un) | (Sc) | Exp. | (Un) | (Sc) | |
| a (un) 6-31+G(d,p) without scaling factor; (sc) 6-31+G(d,p) with scaling factor of 0.964; ar. = aromatic; alp. = aliphatic | |||||||||||||||
| ν(NH) | 3388.9 | 3507.8 | 3381.6 | 3375.4 | 3560.3 | 3432.2 | 3331.1 | 3550.4 | 3422.5 | 3396.6 | 3560.8 | 3432.6 | 3510.4 | 3585.5 | 3456.5 |
| ν(C–H) ar. | 3192.2 | 3229.7 | 3113.5 | 3197.9 | 3240.2 | 3123.6 | 3182.6 | 3239.2 | 3122.5 | 3230.7 | 3244.0 | 3127.1 | 3099.6 | 3246.7 | 3129.9 |
| 3177.2 | 3062.9 | 3041.7 | 3162.0 | 3048.2 | |||||||||||
| ν(C–H) aliphatic | 2906.7 | 3010.4 | 2902.1 | 2910.5 | 3062.5 | 2952.3 | 3034.0 | 3055.4 | 2945.3 | 3097.6 | 3057.5 | 2947.4 | 3024.4 | 3077.3 | 2966.6 |
| ν(C–H) aliphatic CH3 | 2939.5 | 3064.9 | 2954.6 | 2868.1 | 3014.8 | 2906.3 | 2906.7 | 3074.1 | 2963.4 | 2918.3 | 3077.6 | 2966.8 | 2902.8 | 3065.5 | 2955.2 |
| 2906.7 | 3000.1 | 2892.1 | 2816 | 2959.3 | 2852.8 | 3062.9 | 2952.6 | 2862.3 | 3066.8 | 2956.3 | 2812.2 | 3025.0 | 2916.1 | ||
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) amide I O) amide I |
1662.6 | 1759.0 | 1695.7 | 1656.8 | 1691.4 | 1630.6 | 1633.7 | 1696.7 | 1635.6 | 1627.9 | 1691.3 | 1630.4 | 1674.2 | 1659.2 | 1599.5 |
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine N) imine |
1602.9 | 1681.5 | 1621.0 | 1611 | 1673.1 | 1612.9 | 1610.6 | 1672.5 | 1612.2 | 1610.5 | 1671.6 | 1611.4 | 1608.6 | 1617.8 | 1559.5 |
ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) ar. C) ar. |
1589.3 | 1656.1 | 1596.5 | 1595.1 | 1640.8 | 1581.8 | 1595.1 | 1641.2 | 1582.1 | 1581.6 | 1640.4 | 1581.3 | 1591.2 | 1637.7 | 1578.8 |
| 1523.8 | 1591.2 | 1534.0 | 1525.6 | 1584.0 | 1527.0 | 1579.7 | |||||||||
| ν(N–H) amide II | 1523.8 | 1552.1 | 1496.3 | 1525.6 | 1584.0 | 1527.0 | 1525.7 | 1583.2 | 1526.2 | 1525.6 | 1584.7 | 1527.6 | 1525.6 | 1581.0 | 1524.1 |
| ν(C–N) C)ar.)–N | 1546.9 | 1603.4 | 1545.7 | 1546.9 | 1602.6 | 1545.0 | 1498.7 | 1562.5 | 1506.2 | 1546.9 | 1563.5 | 1507.2 | 1363.6 | 1406.4 | 1355.8 |
| 1325.1 | 1389.8 | 1339.8 | |||||||||||||
ν(N![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) NO3 O) NO3 |
— | — | — | — | — | — | — | — | — | 1525.6 | 1592.8 | 1535.4 | — | — | — |
| ν(N–O) NO3 | — | — | — | — | — | — | — | — | — | 1296.1 | 1296.2 | 1249.5 | — | — | — |
| — | — | — | — | — | — | — | — | — | 1224.7 | 1280.3 | 1234.1 | — | — | — | |
| ν(C–N) C(alp.)–N | 1230.6 | 1271.5 | 1225.8 | 1232.5 | 1273.3 | 1227.5 | 1228.7 | 1273.3 | 1227.4 | 1226.7 | 1272.6 | 1226.7 | 1232.5 | 1272.5 | 1226.7 |
| ν (M–O) | — | — | — | 520.7 | — | — | 516.9 | — | — | 520.8 | — | — | 528.1 | — | — |
| ν (M–N) | — | — | — | 476.4 | — | — | 457.1 | — | — | 422.4 | — | — | 460.9 | — | — |
The broad absorption bands observed in the 3380–3330 cm−1 region correspond to ν(N–H) stretching modes, which remain almost unchanged upon complexation, confirming that the amide N–H group does not participate in bonding.34 Aromatic ν(C–H) stretching vibrations were detected near 3190–3170 cm−1 and aliphatic ν(C–H) in the 2900–3050 cm−1 range, showing good correlation with the scaled theoretical predictions.36 The characteristic amide I ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) band at 1660–1670 cm−1 for the free ligand shifted to 1657–1628 cm−1 for the complexes, clearly indicating coordination through the carbonyl oxygen atom.37 Similarly, the ν(C
O) band at 1660–1670 cm−1 for the free ligand shifted to 1657–1628 cm−1 for the complexes, clearly indicating coordination through the carbonyl oxygen atom.37 Similarly, the ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) imine stretching vibration at 1600–1620 cm−1 shifted downwards by ∼10–20 cm−1 upon complexation, confirming the involvement of the azomethine nitrogen atom.38 The aromatic skeletal ν(C
N) imine stretching vibration at 1600–1620 cm−1 shifted downwards by ∼10–20 cm−1 upon complexation, confirming the involvement of the azomethine nitrogen atom.38 The aromatic skeletal ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C) bands at 1590–1520 cm−1 remained essentially unchanged, suggesting no direct interaction of the aromatic ring with the metal center, although intensity variations reflected electronic redistribution upon coordination.
C) bands at 1590–1520 cm−1 remained essentially unchanged, suggesting no direct interaction of the aromatic ring with the metal center, although intensity variations reflected electronic redistribution upon coordination.
In the low-frequency region, new absorptions appeared for all complexes at 520.7, 516.9, 520.8, and 528.1 cm−1 (ν(M–O)) and at 476.4, 457.1, 422.4 and 460.9 cm−1 (ν(M–N)) for [Zn(HL)Cl2]·H2O (1), [Cd(HL)Cl2] (2), [Cd(HL)(NO3)2]·2H2O (3) and [Pt(HL)Cl2]·0.5H2O (4), respectively. These bands are absent in the spectrum of free ligand, confirming the formation of new M–O and M–N bonds.39–41
In the case of the nitrate complex [Cd(HL)(NO3)2]·2H2O (3), additional strong absorption bands at 1525, 1296, and 1280 cm−1 were observed, corresponding to ν(NO2)as and ν(NO2)s modes of the nitrate groups. The difference between the asymmetric and symmetric stretching frequencies (Δν) was calculated to be 229 cm−1 (1525–1296 cm−1) and 245 cm−1 (1525–1280 cm−1).42 These values are significantly increased compared to free ionic nitrate (≈170–200 cm−1),42 thereby confirming a chelating bidentate coordination mode. In this arrangement, each nitrate group binds through two oxygen atoms to cadmium, forming five-membered chelate rings. Together with the donor atoms of the ligand (carbonyl oxygen and imine nitrogen), this provides the Cd(II) ion with a six-coordinate environment, resulting in a distorted octahedral geometry. The observed splitting and shifts of the nitrate bands, combined with the large Δν values, provide strong spectroscopic evidence for this structural assignment.43
Overall, the combined experimental and theoretical IR analysis, supported by CCCBDB scaling,31 demonstrates that the ligand coordinates via C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and C
O and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N groups, while in [Cd(HL)(NO3)2]·2H2O (3), the nitrates act as bidentate chelating ligands, leading to an octahedral arrangement around the Cd center. Overall, the combined experimental and theoretical analysis clearly demonstrates that metal coordination induces significant modifications in the electronic environment of the donor groups, while peripheral moieties remain largely unaffected, thereby supporting the proposed coordination modes of the complexes.
N groups, while in [Cd(HL)(NO3)2]·2H2O (3), the nitrates act as bidentate chelating ligands, leading to an octahedral arrangement around the Cd center. Overall, the combined experimental and theoretical analysis clearly demonstrates that metal coordination induces significant modifications in the electronic environment of the donor groups, while peripheral moieties remain largely unaffected, thereby supporting the proposed coordination modes of the complexes.
3.4 NMR spectra (theoretical and experimental)
Simulated NMR spectra of the ligand and its complexes using DFT showed chemical shifts in close agreement with experimental results.44 Upon coordination with metal centers, noticeable upfield and downfield shifts were observed, reflecting changes in the electronic environments of protons and carbons. The consistency between computational predictions and experimental results confirmed the structural assignments and highlighted the electronic effects induced by metal–ligand interactions. The results are listed in Tables 4 and 5.| Atoms | HL | (1) | (2) | (3) | (4) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | ||||||
| δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | |
| a (iso) isotropic shielding | |||||||||||||||
| H19, H20 | 2.98 | 28.61 | 2.98 | 2.98 | 28.54 | 3.06 | 2.98 | 28.55 | 3.05 | 2.98 | 28.55 | 3.05 | 2.99 | 28.53 | 3.06 |
| H12, H14 | 6.76 | 24.72 | 6.87 | 6.76 | 24.68 | 6.91 | 6.76 | 24.70 | 6.89 | 6.76 | 24.68 | 6.92 | 6.78 | 24.73 | 6.86 |
| H11, H15 | 7.58 | 24.04 | 7.55 | 7.57 | 23.90 | 7.70 | 7.57 | 23.92 | 7.68 | 7.56 | 23.87 | 7.73 | 7.58 | 23.98 | 7.61 |
| H2, H8 | 7.83 | 23.73 | 7.86 | 7.86 | 23.67 | 7.92 | 7.86 | 23.70 | 7.90 | 7.86 | 23.60 | 8.00 | 7.98 | 23.71 | 7.88 |
| H17 | 8.35 | 23.51 | 8.08 | 8.34 | 23.41 | 8.18 | 8.33 | 23.38 | 8.21 | 8.33 | 23.21 | 8.38 | 8.38 | 23.30 | 8.29 |
| H1, H9 | 8.78 | 22.61 | 8.98 | 8.78 | 22.51 | 9.08 | 8.79 | 22.52 | 9.07 | 8.76 | 22.49 | 9.10 | 9.14 | 22.48 | 9.11 |
| H6 | 11.82 | 22.29 | 9.30 | 11.82 | 22.04 | 9.56 | 11.81 | 22.11 | 9.48 | 11.86 | 21.87 | 9.72 | 12.16 | 22.27 | 9.33 |
| Atoms | HL | (1) | (2) | (3) | (4) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | Exp. | Calc. | ||||||
| δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | δ ppm | (iso) | δ ppm | |
| a (iso) isotropic shielding | |||||||||||||||
| C19, C20 | 40.20 | 152.52 | 41.58 | 40.22 | 152.29 | 41.82 | 40.24 | 152.35 | 41.75 | 40.20 | 152.24 | 41.86 | 40.27 | 152.19 | 41.92 |
| C12, C14 | 112.23 | 83.17 | 110.94 | 112.24 | 82.63 | 111.48 | 112.25 | 82.81 | 111.29 | 112.24 | 82.69 | 111.41 | 112.27 | 84.21 | 109.90 |
| C16 | 121.65 | 73.43 | 120.68 | 121.60 | 78.34 | 115.77 | 121.58 | 77.54 | 116.57 | 121.57 | 78.45 | 115.66 | 121.44 | 80.49 | 113.61 |
| C2, C8 | 121.96 | 72.18 | 121.92 | 122.21 | 72.15 | 121.96 | 122.20 | 72.06 | 122.04 | 122.21 | 72.33 | 121.77 | 124.52 | 73.13 | 120.97 |
| C11, C15 | 129.16 | 62.82 | 131.28 | 129.81 | 66.54 | 127.57 | 129.18 | 67.21 | 126.90 | 129.18 | 67.78 | 126.33 | 129.28 | 61.24 | 132.87 |
| C3 | 141.30 | 51.27 | 142.84 | 141.71 | 56.51 | 137.60 | 141.77 | 55.04 | 139.06 | 141.70 | 56.07 | 138.04 | 143.87 | 60.35 | 133.75 |
| C17 | 150.35 | 48.66 | 145.44 | 152.18 | 40.35 | 153.75 | 152.19 | 40.66 | 153.45 | 152.19 | 39.65 | 154.46 | 152.88 | 37.06 | 157.04 |
| C1, C9 | 150.73 | 44.04 | 150.07 | 150.57 | 43.56 | 150.54 | 150.68 | 43.69 | 150.41 | 150.70 | 43.62 | 150.48 | 152.28 | 43.36 | 150.74 |
| C13 | 152.16 | 44.81 | 149.30 | 150.44 | 43.25 | 150.86 | 150.45 | 43.78 | 150.32 | 150.47 | 43.19 | 150.91 | 151.03 | 42.30 | 151.80 |
| C4 | 161.55 | 35.14 | 158.97 | 161.42 | 32.18 | 161.93 | 161.39 | 32.22 | 161.88 | 161.24 | 32.35 | 161.75 | 159.85 | 29.62 | 164.49 |
Table 4 presents a comparative analysis between the experimental NMR chemical shifts (δ, ppm) and the calculated isotropic shielding values (σiso) obtained from DFT studies for the free ligand (HL) and its metal complexes with Zn(II), Cd(II), and Pt(II) ions. The data reveal a strong correlation between theoretical and experimental results, confirming the reliability of the computational model. For the aliphatic protons (H19, H20), the shifts remain nearly constant around 3.0 ppm across all systems, indicating that metal coordination has little influence on these sites. In contrast, the aromatic protons (H12, H14; H11, H15; H2, H8; H17) exhibit noticeable downfield shifts upon complexation, reflecting the electron-withdrawing effects of the coordinated metal centers and the resulting deshielding. The most pronounced changes are observed for H1 and H9, which are located near the donor sites; their signals shift significantly downfield (8.7–9.1 ppm), providing strong evidence for their direct involvement in metal coordination. Interestingly, the H6 proton, which resonates at 11.82 ppm in the spectrum of the free ligand, shifts upfield (9.3–9.7 ppm) upon complexation, consistent with coordination-induced changes in its electronic environment. Overall, complexation strongly affects protons near donor atoms, while peripheral protons remain largely unaffected. The agreement between the experimental and calculated data validates the proposed structures and underscores the influence of metal–ligand interactions on the electronic environments of individual protons.
The experimental 1H and 13C NMR spectra of the free ligand (HL) and its metal complexes (1–4) were recorded in DMSO-d6 (Fig. SI5–SI9). In the 1H NMR spectrum of the free ligand, the aromatic protons (H1, H2, H8, H9, H11, H12, H14, H15) appeared in the region δ 6.76–8.78 ppm. The most downfield resonance at δ 11.82 ppm was assigned to the hydrazinic –NH proton (H6), which is strongly deshielded due to its acidic nature and involvement in hydrogen bonding. For the 13C NMR spectra, the aromatic and imine carbons resonated in the range δ 112.23–161.55 ppm, with the most deshielded signals corresponding to the imine carbon (C17) and the carbonyl carbon (C4), reflecting the influence of electronegative nitrogen atoms and the electronic structure of the conjugated system.37
Theoretical chemical shifts were calculated using the gauge-including atomic orbital (GIAO) method.44,45 The geometries were optimized at the B3LYP/6-31+G(d,p) level for the free ligand for C, H, N, and O atoms, and LANL2DZ for Zn, Cd, and Pt in the complexes. The isotropic shielding values were used to calculate the isotropic chemical shifts (δ) with respect to tetramethylsilane (TMS) according to the equation:
| (TMS)δisox = σTMSiso − σisox |
The values of σisoTMS used in this study are 194.10 and 31.59 ppm for the 13C and 1H NMR spectra, respectively. Upon complexation, a general downfield shift was observed for most proton and carbon signals, which is indicative of the ligand's coordination to the metal center. This coordination reduces the electron density on the ligand, causing a deshielding effect. Analysis of the 13C NMR data reveals the coordination mode, with the most significant shifts observed for the carbonyl carbon (C4) and the imine carbon (C17) (Fig. SI10–SI14). The carbonyl carbon (C4) signal shows a notable shift across the series, moving from δ 161.55 ppm for the free ligand (HL) to the range of δ 159.85–161.42 ppm for the metal complexes. Concurrently, the imine carbon (C17) signal shifts significantly downfield from δ 150.35 ppm for the free ligand to the range of δ 152.18–154.46 ppm for the complexes (Table 4).46,47
The correlation plots of experimental versus calculated chemical shifts showed high linearity, with R2 values greater than 0.88 for 1H NMR and 0.98 for 13C NMR across all compounds (Fig. 5). The performance of the B3LYP method with respect to the prediction of the relative shielding within the molecules is quite good. Notably, the 13C NMR chemical shift calculations gave a significantly better correlation coefficient (R2 > 0.98) than the 1H NMR calculations (R2 > 0.88). Based on the 1H and 13C chemical shift data collected in Tables 4 and 5, one can deduce qualitatively that the NMR chemical shifts of the ligand (HL) and its complexes are described very well by the selected DFT method combined with the basis set. Based on the definitive evidence from the large shifts of the C4 and C17 signals, this validates the proposed coordination48 mode of the ligand to the Zn(II), Cd(II), and Pt(II) centers in a bidentate fashion through the carbonyl oxygen and the imine nitrogen.49,50
3.5 Molecular electrostatic potential (ESP) analysis
ESP maps illustrate charge distributions across the complexes, highlighting electron-rich regions near electronegative atoms and electron-deficient zones around the metal centers.51 Variations in ligand substitution were shown to significantly influence these charge distributions, thereby modifying the complexes' reactivity and interaction tendencies with biomolecules or substrates. Such results demonstrate the importance of ESP in predicting molecular reactivity and interaction patterns. ESP is a valuable quantum chemical property that describes the distribution of electrical charges within a molecule. It serves as a powerful tool for understanding various aspects of molecular behavior and intermolecular interactions. ESP analysis helps predict the strength, type, and duration of interactions between molecules. It also assists in determining whether the interaction between two molecules involves hydrogen bonding or van der Waals forces.52,53ESP analysis identifies the active sites within a molecule. Regions with negative ESP are susceptible to nucleophilic attack, while areas with positive ESP are prone to electrophilic attack.54 The ESP of the HL ligand and its complexes was visualized using a color scheme of red, green, and blue, as presented in Fig. 6. Red represents the electrophilic region, characterized by high electron density and susceptibility to attack. Green corresponds to areas with neutral electrostatic potential, indicating a balanced distribution of electrons. Finally, blue indicates the nucleophilic region, characterized by low electron density and suitability for attack.25–27,52
3.6 Biological activity
The pharmacological importance of various Schiff bases and hydrazone ligands has paved the way for active research in synthetic and medicinal chemistry. As a result, we prepared a new hydrazone ligand and its complexes and screened them for their antimicrobial activities against two different microbial strains. All the synthesized compounds were screened using the disc diffusion method for their antibacterial activities against Staphylococcus aureus and Escherichia coli.The zone of inhibition activity of the compounds was compared with that of the standard streptomycin for antibacterial activity, which was investigated under similar conditions. The inhibition zone values are measured in mm and represented as 10−2, 10−3, 10−4, 10−5, and 10−6 M depending on the diameter and clarity. The measured zone of inhibition values of the free ligand and its complexes against the bacterial strains are shown in Table 6 and Fig. 7.
| Compound | Diameter of inhibition zone (mm) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Staphylococcus aureus | Escherichia coli | ||||||||||
| 10−2 | 10−3 | 10−4 | 10−5 | 10−6 | 10−2 | 10−3 | 10−4 | 10−5 | 10−6 | ||
| HL | 14 | 11 | N.S. | N.S. | N.S. | 11 | 9 | N.S. | N.S. | N.S. | |
| (1) | 21 | 14 | 8 | N.S. | N.S. | 15 | 14 | 8 | N.S. | N.S. | |
| (2) | 18 | 12 | 6 | N.S. | N.S. | 13 | 11 | N.S. | N.S. | N.S. | |
| (3) | 18 | 11 | 6 | N.S. | N.S. | 13 | 10 | N.S. | N.S. | N.S. | |
| (4) | 23 | 16 | 11 | 9 | 6 | 19 | 16 | 11 | N.S. | N.S. | |
| Streptomycin | 29 | 25 | — | — | — | 25 | 23 | — | — | — | |
 | ||
| Fig. 7 Biological activity histogram of the prepared compounds against Staphylococcus aureus and Escherichia coli at 10−2 and 10−3 M. | ||
The results showed that all compounds exhibit a clear dose-dependent response at higher concentrations (10−2 M and 10−3 M), resulting in larger zones of inhibition. In contrast, activity diminishes or becomes negligible (N.S. = not sensitive) at lower concentrations (10−4 M to 10−6 M).
Also, the metal complexes generally exhibit greater antibacterial activity than the free ligand (HL), indicating that coordination with metals enhances the biological efficacy. In addition, the results indicate that none of the tested compounds is more potent than streptomycin at these concentrations.
The [Pt(HL)Cl2]·0.5H2O (4) complex shows the highest activity among all synthesized compounds against Staphylococcus aureus with inhibition zones up to 23 mm at 10−2 M and still measurable activity (6 mm) even at 10−6 M. The [ZnCl2(HL)]·H2O complex and both cadmium complexes [Cd(HL)Cl2] (2) and [Cd(HL)(NO3)2]·2H2O (3) show moderate activity, with zones of around 18–21 mm at 10−2 M. Again, [Pt(HL)Cl2]·0.5H2O (4) is the most active complex against Escherichia coli, with zones of 19 mm at 10−2 M and 16 mm at 10−3 M. [Zn(HL)Cl2]·H2O (1) shows better activity than the cadmium complexes and the free ligand. HL is the least effective, with a zone of only 11 mm at 10−2 M.
The platinum(II) complex demonstrates superior and broad-spectrum antibacterial activity, maintaining efficacy even at very low concentrations. Zinc and cadmium complexes show moderate improvement over the free ligand, but their activity drops sharply with dilution. The results suggest that metal coordination improves antibacterial properties, possibly due to increased membrane permeability, stability, or synergistic effects between the metal and ligand.36 These findings indicate that metal-based complexes—especially platinum derivatives—could serve as promising candidates for developing new antibacterial agents, particularly against Gram-positive (S. aureus) and Gram-negative (E. coli) bacteria.
4 Conclusions
The Schiff base ligand (E)-N′-(4-(dimethylamino)benzylidene) isonicotinohydrazide (HL) and its bivalent metal complexes with Zn(II), Cd(II), and Pt(II) were successfully synthesized and thoroughly characterized using elemental analysis, molar conductivity, thermal analysis, and spectroscopic techniques (FT-IR, NMR). The ligand was found to act as a bidentate chelating agent, coordinating through the carbonyl oxygen and azomethine nitrogen atoms. Theoretical studies using DFT calculations supported the experimental findings, providing optimized geometries, molecular ESP maps, and HOMO–LUMO energy gaps that corroborated the structural and electronic properties of the compounds. The thermal stability and decomposition pathways of the complexes were elucidated through TG-DTA measurements, confirming the formation of stable metal oxides as end products. Antibacterial evaluation revealed that all the metal complexes exhibited enhanced activity compared to the free ligand, with the Pt(II) complex demonstrating the most potent and broad-spectrum efficacy against both Staphylococcus aureus and Escherichia coli, even at low concentrations. The Zn(II) and Cd(II) complexes also showed significant activity, though less than that of the Pt(II) complex.Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are available in the supplementary information (SI) of this article. Supplementary information is available. See DOI: https://doi.org/10.1039/d5ra07625k.Acknowledgements
This work was supported and funded by the Deanship of Scientific Research at Imam Mohammad Ibn Saud Islamic University (IMSIU) (grant number IMSIU-DDRSP2601).References
- A. Furst, R. C. Berlo and S. Hooton, Chem. Rev., 1965, 65, 51–68 CrossRef CAS.
- J. de Oliveira Carneiro Brum, T. C. C. Franca, S. R. LaPlante and J. D. F. Villar, Mini-Rev. Med. Chem., 2020, 20, 342–368 CrossRef CAS PubMed.
- L. A. Tatum, X. Su and I. Aprahamian, Acc. Chem. Res., 2014, 47, 2141–2149 CrossRef CAS PubMed.
- N. Ribeiro and I. Correia, Front. Chem. Biol., 2024, 3, 1398873 Search PubMed.
- M. M. Shakdofa, M. H. Shtaiwi, N. Morsy and T. M. Abdel-rassel, Curr. Med. Chem., 2014, 13, 187–218 CAS.
- B. Kumar, J. Devi, A. Dubey and M. Kumar, Chem. Biodiversity, 2024, 21, e202401116 CrossRef CAS PubMed.
- D. A. Tafere, M. Gebrezgiabher, F. Elemo, T. Sani, T. B. Atisme, T. G. Ashebr and I. N. Ahmed, RSC Adv., 2025, 15, 6191–6207 RSC.
- R. Narang, B. Narasimhan and S. Sharma, Curr. Med. Chem., 2012, 19, 569–612 CrossRef CAS PubMed.
- I. Hamzi, J. Fluoresc., 2025, 35, 2569–2621 CrossRef CAS PubMed.
- G. More, D. Raut and K. Aruna, Appl. Organomet. Chem., 2018, 32, e4124 CrossRef.
- P. Ball and C. Nicholls, Dyes Pigm., 1982, 3, 5–26 CrossRef CAS.
- J. Dilworth, Coord. Chem. Rev., 1976, 21, 29–62 CrossRef CAS.
- C. Radunsky, J. Kösters and J. Müller, Inorg. Chim. Acta, 2015, 428, 14–20 CrossRef CAS.
- E. Miminoshvili, J. Struct. Chem., 2009, 50, 168–175 CrossRef.
- D. Kumar, S. Chadda, J. Sharma and P. Surain, Bioinorg. Chem. Appl., 2013, 2013, 981764 Search PubMed.
- N. P. Belskaya, A. I. Eliseeva and V. A. Bakulev, Russ. Chem. Rev., 2015, 84, 1226 CrossRef CAS.
- D. K. Kölmel and E. T. Kool, Chem. Rev., 2017, 117, 10358–10376 CrossRef PubMed.
- L. Stegbauer, K. Schwinghammer and B. V. Lotsch, Chem. Sci., 2014, 5, 2789–2793 Search PubMed.
- S. Shaaban, T. A. Yousef, A. S. Al-Janabi, T. Alammar, M. Alaasar, K. Shalabi, A. A. Al-Karmalawy, H. Ferjani, A. Al-Dakhil and A. M. Abu-Dief, Polyhedron, 2025, 279, 117652 CrossRef CAS.
- R. O. Khalid, A. S. Al-Janabi, A. A. Al-Bayati and M. Alheety, J. Mol. Struct., 2025, 1323, 140721 CrossRef CAS.
- K. T. Abdullah, A. M. Etheb, W. H. Majeed, A. S. Al-Janabi and N. J. Hussien, Edelweiss Appl. Sci. Technol., 2025, 9, 2363–2373 CrossRef.
- T. A. Yousef and A. S. Al-Janabi, Heliyon, 2024, 10, e37310 CrossRef CAS PubMed.
- T. A. Yousef, L. H. Al-Jibori, A. S. Fiahan, A. O. Elzupir, M. M. Abou-Krisha and A. S. Al-Janabi, J. Mol. Struct., 2024, 1308, 138073 CrossRef CAS.
- A. S. Al-Janabi, K. H. Oudah, S. A. Aldossari, M. A. Khalaf, A. M. Saleh, M. R. Hatshan, N. B. Altheeb and S. F. Adil, J. Saudi Chem. Soc., 2023, 27, 101619 CrossRef CAS.
- M. Gouda, H. Ferjani, H. M. Abd El-Lateef, M. M. Khalaf, S. Shaaban and T. A. Yousef, Int. J. Mol. Sci., 2022, 23, 2716 CrossRef CAS PubMed.
- S. Shaaban, H. Ferjani, T. Yousef and M. Abdel-Motaal, J. Inorg. Organomet. Polym. Mater., 2022, 32, 1878–1890 CrossRef CAS.
- T. Yousef, O. Alduaij, S. F. Ahmed, G. A. El-Reash and O. El-Gammal, J. Mol. Struct., 2016, 1125, 788–799 CrossRef CAS.
- R. M. Silverstein and G. C. Bassler, J. Chem. Educ., 1962, 39, 546 CrossRef CAS.
- Y. Yang, M. N. Weaver and K. M. Merz Jr., J. Phys. Chem. A, 2009, 113, 9843–9851 CrossRef CAS PubMed.
- R. D. Johnson III, NIST Computational Chemistry Comparison and Benchmark Database, Report NIST Standard Reference Database Number 101, NIST, Gaithersburg, 2022, DOI:10.18434/T47C7Z.
- G. F. Liu, X. J. Ma, S. H. Ma, H. W. Zhao, M. W. Ma, M. Ge and W. F. Wang, Chin. J. Chem., 2008, 26, 1257–1261 CrossRef CAS.
- R. Sghyar, M. Lahyaoui, N. Aflak, O. Moussaoui, A. Chda, R. Bencheikh, E. M. El Hadrami, N. K. Sebbar, A. S. Alanazi and M. Hefnawy, Molecules, 2024, 29, 3855 CrossRef CAS PubMed.
- A. F. Rodrigues-Oliveira, F. W. M. Ribeiro, G. Cervi and T. C. Correra, ACS Omega, 2018, 3, 9075–9085 CrossRef CAS PubMed.
- W. J. Geary, Coord. Chem. Rev., 1971, 7, 81–122 CrossRef CAS.
- H. A. Al-Abdulkarim, H. A. Qasem, M. R. Aouad, M. S. Khushaim, E. S. Al-Farraj, A. Abdou and A. M. Abu-Dief, Appl. Organomet. Chem., 2024, 38, e7486 Search PubMed.
- H. A. K. Kyhoiesh and K. J. Al-Adilee, Chem. Pap., 2022, 76, 2777–2810 Search PubMed.
- A. A. Al-Shamry, M. M. Khalaf, H. M. A. El-Lateef, T. A. Yousef, G. G. Mohamed, K. M. K. El-Deen, M. Gouda and A. M. Abu-Dief, Materials, 2022, 16, 83 Search PubMed.
- R. N. Patel, Y. Singh, Y. P. Singh and R. J. Butcher, J. Coord. Chem., 2016, 69, 2377–2390 Search PubMed.
- L. A. Alnuaimy, Rafidain J. Sci., 2022, 31, 62–76 CrossRef.
- M. Y. Nassar, I. S. Ahmed, H. A. Dessouki and S. S. Ali, J. Basic Environ. Sci., 2018, 5, 60–71 CrossRef CAS.
- W. Mahmoud, N. Mahmoud and G. G. Mohamed, Appl. Organomet. Chem., 2017, 31, e3858 CrossRef.
- R. D. Marchenko, T. S. Sukhikh, A. A. Ryadun and A. S. Potapov, Molecules, 2021, 26, 5400 Search PubMed.
- Y. Xin-Tong, C. Xiang-Yu, L. Yang, C. Man-Sheng and Z. Kristallogr, New Cryst. Struct., 2024, 239, 665–667 Search PubMed.
- K. Wolinski, J. F. Hinton and P. Pulay, J. Am. Chem. Soc., 2002, 112, 8251–8260 Search PubMed.
- D. H. Kim, H. M. Eun and H.-S. Choi, Bull. Korean Chem. Soc., 2000, 21, 148–150 CAS.
- R. M. Ahmed, E. I. Yousif and M. J. Al-Jeboori, Sci. World J., 2013, 2013, 754868 Search PubMed.
- S. İrişli, S. Günnaz, Ö. Özcan, A. Arı, M. Maral, A. Erdem, D. Özel and F. Yurt, Appl. Organomet. Chem., 2024, 38, e7540 CrossRef.
- P. Barraud, M. Schubert and F. H. Allain, J. Biomol. NMR, 2012, 53, 93–101 CrossRef CAS PubMed.
- A. Hussain, K. Mariappan, D. C. Cork, L. D. Lewandowski, P. K. Shrestha, S. Giri, X. Wang and A. G. Sykes, RSC Adv., 2021, 11, 34181–34192 Search PubMed.
- R. Khalaf, A. Faheim, S. Ibrahim, Z. El-Shafiey and H. Saleh, Green Energy Environ. Technol., 2025, 4(1), 1–24 Search PubMed.
- E. Scrocco and J. Tomasi, in Advances in Quantum Chemistry, Elsevier, 1978, vol. 11, pp. 115–193 Search PubMed.
- O. A. Al-Samrai, O. R. A. Samarrai, T. A. Yousef, M. M. Abou-Krisha, M. H. Alhalafi and A. S. Al-Janabi, Chem. Phys. Impact, 2024, 8, 100581 CrossRef.
- K. A. Ford, Mol. Pharmaceutics, 2013, 10, 1171–1182 Search PubMed.
- L.-Q. Chai, X.-F. Zhang and L.-J. Tang, J. Mol. Struct., 2021, 1245, 131028 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |