Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Novel cinnamic acid-based N-benzyl pyridinium analogs: potent dual cholinesterase inhibitors with neuroprotective properties for Alzheimer's disease
Maryam Esmkhania,
Mohammad Mahdavib,
Shahrzad Javanshir *a and
Aida Iraji
*a and
Aida Iraji *cd
*cd
aPharmaceutical and Heterocyclic Compounds Research Laboratory, Department of Chemistry, Iran University of Science and Technology, Tehran, 16846-13114, Iran
bEndocrinology and Metabolism Research Center, Endocrinology and Metabolism Clinical Sciences Institute, Tehran University of Medical Sciences, Tehran, Iran
cStem Cells Technology Research Center, Shiraz University of Medical Sciences, Shiraz, Iran
dPharmaceutical Sciences Research Center, Shiraz University of Medical Sciences, Shiraz, Iran
First published on 16th February 2026
Abstract
This study reports the design and synthesis of a novel series of cinnamic acid-based analogs bearing an N-benzyl pyridinium moiety against Alzheimer's disease (AD), aiming at dual inhibition of acetylcholinesterase (AChE) and butyrylcholinesterase (BChE), alongside neuroprotective effects. A total of 15 derivatives were synthesized, among which compound 7b exhibited the most potent dual inhibition (AChE IC50 = 0.89 µM; BChE IC50 = 0.11 µM), and significant neuroprotection against H2O2-induced oxidative stress in SH-SY5Y cells, with no cytotoxicity under the tested concentration. Structure–activity relationship (SAR) analysis revealed that small electron-withdrawing substituents (e.g. ortho-fluoro, methyl) enhanced inhibitory activity, whereas meta and para substitutions generally reduced potency. Enzyme kinetics also determined compound 7b to be a competitive inhibitor of AChE (Ki = 0.49 µM). Furthermore, molecular docking and molecular dynamics simulations identified stable binding interactions in the active sites of AChE and BChE. All these findings support the potential of these compounds as effective multi-target-directed ligands (MTDLs) for AD, displaying coordinated inhibition of cholinesterase, neuroprotection, and low toxicity.
1. Introduction
Alzheimer's disease (AD) is a chronic, progressive brain disorder and the most prevalent cause of cognitive decline, responsible for about 60–80% of all cases of dementia. It is mostly found in people over 65 years of age, with the exception of the early presentation in some cases.1 It is associated with loss of memory, problems thinking, confusion, and changed behavior. Gradually, it leads to complete dependency and eventually death. At a biological level, AD has been linked to the generation of amyloid-beta (Aβ) plaques in synapses, tau protein tangles in neurons, inflammation, oxidative stress, and overall loss of nerve cells.2–4Despite a history of research during these years, no definitive treatment for AD has been found.5 Current medications are intended to slow the rate of cognitive decline, emphasizing the importance of early diagnosis and timely treatment. It is approved that the cholinergic system is excessively damaged in AD.6 Acetylcholine (ACh), a neurotransmitter essential for memory formation and learning, is rapidly degraded by two enzymes acetylcholinesterase (AChE) and butyrylcholinesterase (BChE).7 Whereas AChE is more active in the early phases of AD, BChE becomes significant with disease progression. Thus, dual inhibition of AChE and BChE has been a promising approach to enhance cholinergic transmission and overcome cognitive manifestations. Some FDA-approved medications, such as donepezil, rivastigmine, and galantamine, are cholinesterase inhibitors that offer symptomatic relief with modest cognitive and functional improvement.8
In addition to cholinesterase inhibition, the neuroprotective medications aim to hinder or decelerate the loss of neurons by addressing oxidative stress, mitochondrial dysfunction, neuroinflammation, and excitotoxicity processes presently recognized as playing a major role in AD pathogenesis.9,10 Antioxidant and neuroprotective drugs are studied for their potential to halt or reverse neurodegenerative activity. The combination of cholinesterase inhibition with neuroprotection may provide a synergistic effect and not only symptomatic control but also modification of the disease.11 Hence, highly efficacious AChE and BChE inhibitors with minimal cytotoxicity and neuroprotection effects as categorized as the ideal therapeutic agents. In this context, various structural scaffolds have been explored, including iminochromene-2H-carboxamide [11], triazine [12], carbazole-benzylpiperidine [13], and N-cyclohexylimidazo[1,2-a]pyridine [14]. As a result, a novel class of cinnamic acid-derived analogues attached to benzyl pyridinium derivatives was designed to combine cognitive improvement, neuroprotection, and low toxicity in a single entity. Fifteen derivatives were synthesized and screened against AChE and BChE. Molecular docking, molecular dynamics simulations, kinetic analysis, cytotoxicity tests, and neuroprotection assays of the active analogues were performed and validate their therapeutic value towards AD.
2. Results and discussion
2.1 Designing
Donepezil (A, Fig. 1) is known as an FDA-approved drug for the management of AD. Considering this drug, different derivatives were designed and synthesized. A series of donepezil-based hybrids incorporating N-benzyl pyridinium groups (B, Fig. 1) was developed to enhance binding to both the catalytic active site (CAS) and peripheral anionic site (PAS) of AChE. These compounds demonstrated significant AChE inhibition, with structure–activity relationship (SAR) analysis highlighting that electron-donating groups, such as methoxy at the para-position of the benzyl ring, enhanced activity, likely due to strengthened π–π interactions with the PAS. A three-carbon spacer between the core and pyridinium ring was optimal for dual-site binding.12 Additionally, 2-benzofuran carboxamide derivatives (C, Fig. 1) with N-benzyl pyridinium moieties showed potent BChE inhibition (IC50 = 0.054–2.7 µM), with compound C being the most potent (IC50 = 0.054 µM), over 100 times stronger than donepezil, with excelled in AChE inhibition (IC50 = 2.1 µM). SAR indicated that 3-pyridinium derivatives outperformed 4-pyridinium ones, and ortho- or meta-substituted benzyl groups were more effective than para-substituted ones.13Notably, cinnamic acid-based analogs with N-benzyl pyridinium groups, particularly compound D, exhibited exceptional AChE potency (IC50 = 12.1 nM for electric eel AChE; 8.6 nM for human AChE) with high selectivity over BChE (selectivity index ∼215). Structure activity relationships (SAR) showed that meta and para substituents fluorine or methoxy on the benzyl group improved potency, while bulky or electron-withdrawing groups reduced activity. Kinetic and docking studies confirmed D as a mixed-type inhibitor, engaging both CAS and PAS via π–π stacking, π–cation interactions, and hydrogen bonds.14 Thiazole-pyridinium hybrids (E, Fig. 1) demonstrated dual action against AChE and Aβ aggregation, with E showing strong AChE inhibition (IC50 = 0.40 µM) and superior Aβ inhibition compared to donepezil (20.4% and 42.7% vs. 14.7% at 10 µM). According to SAR, the ortho and meta substituted compounds demonstrated stronger inhibition against AChE than the para substituted derivatives. Compound E exhibited significant neuroprotection against H2O2-induced oxidative stress in PC12 cells, comparable to donepezil, and favorable blood–brain barrier permeability.15 For 4-aryl-4-oxo-N-phenyl-2-aminylbutyramides (F), SAR revealed that alkyl substituents near a hydrogen-bond acceptor, like a carbonyl enhanced AChE potency with heterocyclic amines (like piperidinyl) modulating activity and selectivity.16
In conclusion, combining cinnamic acid scaffolds with N-benzyl pyridinium groups emerged as an ideal pharmacophore for potent and selective AChE and BChE inhibition to target AD.
2.2 Synthesis
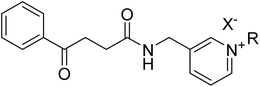
The synthesis of 1-benzyl-3-[(4-oxo-4-phenylbutanamido)methyl]pyridine derivatives was carried out in three main steps shown in Scheme 1. Initially, 4-oxo-4-phenylbutanoic acid (3) was synthesized by reacting succinic anhydride (2) with anhydrous aluminum chloride in dry benzene (1) under reflux conditions for 24 hours. In the second step, the synthesized acid was activated using TBTU in the presence of triethylamine in DMF and then reacted with 3-picolylamine (4) at room temperature to form the corresponding amide, 4-oxo-4-phenyl-N-(pyridin-3-ylmethyl)butanamide (5). In the final step, this amide was dissolved in acetonitrile and reacted with different benzyl halide derivatives (6) under reflux. In each step the reaction completion was monitored via thin layer chromatography (TLC). These steps led to the successful synthesis of a series of substituted pyridine-based derivatives with potential biological significance.2.3 Structure–activity relationship analysis of AChE inhibitors
The parent compound 7a (benzyl, AChE IC50 = 3.77 ± 0.55 µM) acts as our reference, showing a moderate level of AChE potency. Among fluoro substitutions analog, 7b (2-fluorobenzyl, AChE IC50 = 0.89 ± 0.12 µM) shows a notable improvement in potency compared to 7a, suggesting that the small, electronegative fluorine enhances binding through favorable electronic interactions with the residues in the AChE active site (Table 1).| Compound | R | X | IC50 (µM) against AChE | IC50 (µM) against BChE | Selectivity profile |
|---|---|---|---|---|---|
| a Mean ± SE.b Positive control. | |||||
| 7a | Benzyl | Br | 3.77 ± 0.55 | 1.02 ± 0.29 | 3.69 |
| 7b | 2-Fluorobenzyl | Br | 0.89 ± 0.12 | 0.11 ± 0.08 | 8.09 |
| 7c | 3-Fluorobenzyl | Cl | 19.73 ± 0.96 | 0.49 ± 0.16 | 40.26 |
| 7d | 4-Fluorobenzyl | Br | 4.89 ± 1.41 | 11.09 ± 0.18 | 0.44 |
| 7e | 2-Chlorobenzyl | Cl | 4.04 ± 0.05 | 1.02 ± 0.00 | 3.69 |
| 7f | 3-Chlorobenzyl | Cl | 16.79 ± 0.27 | 0.98 ± 0.13 | 17.32 |
| 7g | 4-Chlorobenzyl | Cl | 24.55 ± 0.79 | 0.68 ± 0.26 | 36.10 |
| 7h | 4-Bromobenzyl | Br | 17.09 ± 2.77 | 2.66 ± 0.22 | 6.42 |
| 7i | 4-Triflurobenzyl | Br | 7.83 ± 2.75 | 6.61 ± 0.21 | 1.18 |
| 7j | 4-Nitrobenzyl | Br | >50 | 25.22 ± 3.28 | >1.98 |
| 7k | 4-Methoxybenzyl | Br | 20.43 ± 1.33 | 1.09 ± 0.03 | 18.73 |
| 7l | 4-Methylbenzyl | Cl | 13.21 ± 2.23 | 15.31 ± 0.24 | 0.86 |
| 7m | 2-Methylbenzyl | Cl | 15.17 ± 1.48 | 0.50 ± 0.16 | 30.34 |
| 7n | Methyl | I | 0.90 ± 0.11 | 1.73 ± 0.31 | 0.51 |
| Donepezilb | — | 0.079 ± 0.05 | 10.6 ± 2.1 | 0.007 | |
Compound 7c (3-fluorobenzyl, AChE IC50 = 19.73 ± 0.96 µM) reduces potency considerably, which implies meta-fluoro disrupts AChE binding. Compound 7d (4-fluorobenzyl, AChE IC50 = 4.89 ± 1.41 µM) reduces potency modestly compared to 7a, which implies that para-fluoro is more tolerated than meta-fluoro but less favored than ortho-fluoro. Among chloro substitutions, 7e (2-chloro benzyl, AChE IC50 = 4.04 ± 0.05 µM) is marginally less potent than 7a, which suggests minimal steric hindrance by the chlorine atom. Compound 7f (3-chlorobenzyl, AChE IC50 = 16.79 ± 0.27 µM) is significantly less potent, suggesting unfavorable position at the meta position. Compound 7g (4-chlorobenzyl, AChE IC50 = 24.55 ± 0.79 µM) reduces the potency further, which suggests that para-chloro is not well tolerated for AChE binding, which may be due to steric or electronic mismatch. Compound 7h (4-bromobenzyl, AChE IC50 = 17.09 ± 2.77 µM) loses potency significantly, likely due to steric hindrance from the larger bromine atom. Compound 7i (4-trifluoromethylbenzyl, AChE IC50 = 7.83 ± 2.75 µM) is of moderate potency, with the electron-withdrawing hydrophobic trifluoromethyl group partially compensating for steric effects. Compound 7j (4-nitrobenzyl, AChE IC50 > 50 µM) is inactive due to the nitro group's strong electron-withdrawing nature and steric bulk. Compound 7k (4-methoxybenzyl, AChE IC50 = 20.43 ± 1.33 µM) reduces potency significantly, possibly because of the electron-donating nature or steric bulk of the methoxy group. Compound 7l (4-methylbenzyl, AChE IC50 = 13.21 ± 2.23 µM) reduces potency, indicating that para-methyl is less sterically disruptive than other para substitutions. Compound 7m (2-methylbenzyl, AChE IC50 = 15.17 ± 1.48 µM) is less active than 7a, revealing that ortho-methyl is responsible for steric hindrance.
On the other hand, compound 7n (methyl, AChE IC50 = 0.90 ± 0.11 µM) stands out with the highest AChE potency, hinting that the smaller, aliphatic methyl group boosts binding affinity, likely because it reduces steric hindrance or fits better in the AChE active site.
SAR analysis indicates that small substituents, such as the methyl group in 7n or ortho-fluoro in 7b are optimal for AChE inhibition, with 7n being the most potent. Meta and para substitution, particularly with larger or strongly electron-modifying groups such as chloro (7f, 7g), bromo (7h), nitro (7j), or methoxy (7k), reduce potency significantly. Ortho substitution is generally better tolerated, with fluoro being optimal. The trifluoromethyl group in 7i is moderately active but less than the unsubstituted benzyl. Optimization of AChE inhibition in the future should be aimed at small, electron-withdrawing ortho substituents or minimal alkyl groups for optimal potency.
The selectivity profile of the series for BChE over AChE is expressed as the selectivity index (SI = IC50 AChE/IC50 BChE). Most derivatives exhibited preferential inhibition of BChE, with SI values ranging from 3.69 to 40.26. Compound 7c (SI = 40.26), 7g (SI = 36.10), and 7m (SI = 30.34) displayed the highest BChE selectivity, while 7b showed excellent dual potency (IC50 AChE = 0.89 µM, IC50 BChE = 0.11 µM) with good BChE selectivity (SI = 8.09). In contrast, compounds 7d, 7l, and 7n showed slight preference for AChE (SI < 1).
2.4 Structure–activity relationship analysis of BChE inhibitors
Parent compound 7a (benzyl, BChE IC50 = 1.02 ± 0.29 µM, AChE IC50 = 3.77 ± 0.55 µM) is the baseline with good BChE activity and moderate AChE inhibition with a selectivity ratio (BChE IC50/AChE IC50) of 3.7, showing selectivity for BChE.For comparison, 7n (methyl, BChE IC50 = 1.73 ± 0.31 µM, AChE IC50 = 0.90 ± 0.11 µM) has less BChE activity but more AChE activity, with the selectivity ratio of approximately 0.52 in favor of AChE. This suggests the benzyl skeleton is more favorable towards BChE inhibition, due to hydrophobic or π–π interactions in the BChE active site. Halogen substitutions provide hints about electronic and steric effects. Among the fluoro replacements, 7b (2-fluorobenzyl, BChE IC50 = 0.11 ± 0.08 µM, AChE IC50 = 0.89 ± 0.12 µM) is most potent at BChE with a selectivity ratio of approximately 8.09, meaning the small electronegative fluorine at the ortho position enhances binding via favorable interactions. Compound 7c (3-fluorobenzyl, BChE IC50 = 0.49 ± 0.16 µM, AChE IC50 = 19.73 ± 0.96 µM) increases the potency of BChE over 7a with a selectivity ratio of approximately 40.3, presenting very high selectivity towards BChE, but is less potent compared to 7b.
The compound 7d, characterized by 4-fluorobenzyl, has a BChE IC50 = 11.09 ± 0.18 µM, AChE IC50 = 4.89 ± 1.41 µM) significantly reducing BChE potency, with a selectivity ratio of approximately 0.44, favoring AChE, suggesting that para-fluoro disrupts BChE binding, possibly due to unfavorable electronic or steric effects. Regarding chloro substitutions, the compound identified as 7e (2-chlorobenzyl, BChE IC50 = 1.02 ± 0.00 µM, AChE IC50 = 4.04 ± 0.05 µM) maintains BChE potency equivalent to 7a, with a selectivity ratio of approximately 4.0.
Compound 7f (3-chlorobenzyl, BChE IC50 = 0.98 ± 0.13 µM, AChE IC50 = 16.79 ± 0.27 µM) slightly improves BChE potency, with a selectivity ratio of approximately 17.1, suggesting favorable electronic or steric interactions at the meta position. The compound 7g (4-chlorobenzyl, BChE IC50 = 0.68 ± 0.26 µM; AChE IC50 = 24.55 ± 0.79 µM), greatly increases the potency towards BChE. The selectivity ratio is about 36.1, which indicates that chlorine is well tolerated at the para position, possibly because of favorable electronic or hydrophobic interactions. Compound 7h (4-bromobenzyl, BChE IC50 = 2.66 ± 0.22 µM, AChE IC50 = 17.09 ± 2.77 µM) decreases BChE potency compared to 7a, with a selectivity ratio of approximately 6.4, likely due to the larger bromine causing steric hindrance. Compound 7i (4-trifluoromethylbenzyl, BChE IC50 = 6.61 ± 0.21 µM, AChE IC50 = 7.83 ± 2.75 µM) reduces BChE potency compared to 7a, with a selectivity ratio of approximately 1.2, but is better than other para substitutions like nitro or methyl, suggesting that the electron-withdrawing and hydrophobic trifluoromethyl group partially supports binding, though steric effects limit efficacy. Compound 7j shows a negligible enzyme inhibitory effect (selectivity greater than 2.0), likely due to the strong electron withdrawing group and steric bulk of the nitro group, which disrupted some important interactions (4-nitrobenzyl, BChE IC50 25.22 ± 3.28 µM, AChE IC50 > 50 µM). On the other hand, compound 7k produced an inhibition pattern in some degree similar to that of 7a or nearly equal, which showed good potency for the inhibition of BChE, with an IC50 of 1.09 ± 0.03 µM, and poor inhibition for AChE, with an IC50 of 20.43 ± 1.33 µM, providing a selectivity ratio of approximately 18.7, thereby suggesting that the para-position of the electron-donating methoxy group is beneficial. Compound 7l (4-methylbenzyl, BChE IC50 = 15.31 ± 0.24 µM, AChE IC50 = 13.21 ± 2.23 µM) significantly reduces BChE potency, with a selectivity ratio of approximately 0.86, indicating that the para-methyl group is poorly tolerated, possibly due to steric effects. Compound 7m (2-methylbenzyl, BChE IC50 = 0.50 ± 0.16 µM, AChE IC50 = 15.17 ± 1.48 µM) improves BChE potency compared to 7a, with a selectivity ratio of approximately 30.3, suggesting that the ortho-methyl group enhances hydrophobic or van der Waals interactions without significant steric hindrance.
SAR pointed at small electron-withdrawing substituents mostly, on the ortho position, like fluoro in 7b, and small hydrophobic groups, like methyl in 7m were best for BChE inhibition, often with high selectivity against AChE. Fluoro in 7c and chloro in 7f at the meta position increase BChE potency and selectivity as well. Para substitution is mostly disfavored, except chloro in 7g and methoxy in 7k, which still show good potency and selectivity. On the other hand, bulky or strong electron-modifying groups at the para position, such as nitro in 7j and bromo in 7h, reduce the BChE activity. The benzyl scaffold was generally found to be preferred for BChE, whereas methyl in 7n prefers AchE, which could be due to the bulkier active site of BChE.
2.5 Kinetic study
To determine the mechanism of inhibition, a kinetic study of 7b as a potent inhibitor was done against AChE. In the reciprocal Lineweaver–Burk plot (Fig. 2a), the Km gradually increased and Vmax remained unchanged with increasing inhibitor concentration, indicating a competitive inhibition. Furthermore, the plot of the Km versus different concentrations of the inhibitor gave an estimate of the inhibition constant, Ki of 0.49 µM (Fig. 2b). | ||
| Fig. 2 (a) The Lineweaver–Burk plot of the 7b against AChE, (b) the secondary plot between Km and various concentrations of 7b. | ||
2.6 Cell cytotoxicity and neuroprotection
Cell viability of all derivatives was assessed on the SH-SY5Y neuroblastoma cell line 72 h after treatment, and the results are summarized in Table 2 and Fig. 3.| Compound | CC50 (µM) | SI vs. AChE (CC50/IC50 AChE) | SI vs. BChE (CC50/IC50 BChE) |
|---|---|---|---|
| 7a | 121.48 ± 6.52 | 29.6 | 118.6 |
| 7b | 111.46 ± 11.22 | 97.1 | 1013 |
| 7c | 86.42 ± 2.91 | 4.3 | 176.4 |
| 7d | 94.42 ± 3.31 | 19.3 | 8.5 |
| 7e | 72.23 ± 1.68 | 17.8 | 70.8 |
| 7f | 78.11 ± 7.96 | 4.6 | 79.7 |
| 7g | 70.35 ± 8.18 | 2.86 | 103.4 |
| 7h | 75.08 ± 9.59 | 4.4 | 28.2 |
| 7i | 60.66 ± 3.57 | 7.74 | 9.2 |
| 7j | 54.02 ± 3.51 | — | 2.1 |
| 7k | 73.26 ± 3.11 | 3.6 | 67.2 |
| 7l | 75.98 ± 2.14 | 5.7 | 4.96 |
| 7m | 74.94 ± 1.32 | 4.94 | 149.9 |
| 7n | 118.35 ± 11.81 | 131 | 86.4 |
From the data, most compounds show low cytotoxicity at both concentrations. At 50 µM, compounds 7j (54.0%) and 7i (60.7%) exhibited the strongest reduction in cell viability, while 7g, 7e, 7f, 7k, 7m, 7h, and 7l reduced viability to approximately 70–76%, indicating moderate effects. Compounds 7c, 7d, and others remained above 86% viability, and remarkably, 7n (118.3%) and 7a (121.5%) slightly increased viability compared to control, which may reflect mild proliferative stimulation. At the lower 10 µM concentration, nearly all compounds displayed viability above 77–138%, with several (7n, 7f, 7m, 7a, 7e, 7k, 7c) exceeding 100%, again suggesting good tolerability and possible metabolic activation at lower doses.
To assess the therapeutic window, selectivity indices (SI) and CC50 were calculated (Table 2). The lead compound 7b demonstrated exceptionally high selectivity, particularly against BChE (SI = 1013), along with a strong SI of 97.1 against AChE, underscoring its favorable profile as a potent dual inhibitor with minimal neurotoxicity. Among the series of several derivatives, including 7a, 7c, and 7m achieve SI > 100 for at least one enzyme, considering these analogs as ideal lead candidates.
The ability of compound 7b to protect against neurotoxicity was tested in SH-SY5Y cells under oxidative stress induced by hydrogen peroxide (H2O2). H2O2 exposure caused cell viability to drop to about 59% compared to controls. Treatment with compound 7b at 3 µM produced a pronounced neuroprotective effect, resulting in a significant increase in viability to 72% (Fig. 4).
2.7 Molecular docking
Docking studies were conducted to evaluate the interaction of compounds with AChE. The suitability of the docking protocol was assessed by redocking the native ligand into the AChE active site, yielding root-mean-square deviation (RMSD) values below 2 Å, thereby validating the docking process. The superimposed structure of donepezil and compound 7b is exhibited in Fig. 5. It is shown that 7b has a binding pose similar to that of donepezil within the active site. Compound 7b recorded a binding energy of −9.43 kcal mol−1, indicating high binding affinity. The cinnamic acid moiety of 7b exhibited π–π stacking and π–cation interactions with Hip447 (protonated form of His residue), a key residue of the catalytic triad that plays a role in inhibiting AChE. Furthermore, the carbonyl group formed hydrogen bonds with Ser203, a catalytic triad residue, and with Glh122 (protonated Glu). The carbonyl of the amide linker also donated a hydrogen bond with Tyr124 in the PAS. Furthermore, the terminal 2-fluorobenzyl pyridinium was involved in π–π stacking interactions with Tyr341 and π–cation interactions with Trp286, both of which are located in the PAS region and are important in ligand recognition and inhibition. | ||
| Fig. 5 Superimposed binding poses of compound 7b (blue) and donepezil (green) within the active site of AChE. | ||
To elucidate the influence of substituent position on binding behavior, additional docking studies were performed for analogs 7c–g. The 7c (3-fluorobenzyl) exhibited a lower binding energy (−8.608 kcal mol−1) and showed π–π stacking with Tyr337 and Trp86, as well as hydrogen bonding to Phe295, suggesting weaker stabilization within the active site than 7b. In contrast, the para substituted analog 7d displayed a slightly improved binding energy (−9.559 kcal mol−1), forming π–π stacking with Trp86 and Tyr341, as well as hydrogen bonds with Phe295.
Among the chloro-substituted derivatives, 7e bearing ortho-chloro substitution showed the strongest binding energy (−10.559 kcal mol−1), likely due to enhanced hydrophobic interactions and ideal pose within the binding pocket. In comparison, meta (7f) and para chloro (7g) analogs displayed slightly reduced binding energies, indicating that ortho substitution better stabilization within the AChE active site (Table 3).
| Compound | AChE | BCHE | ||
|---|---|---|---|---|
| Binding energy (kcal mol−1) | Type of interaction and residues | Binding energy (kcal mol−1) | Type of interaction and residues | |
| Donepezil | −9.342 | π–cation with Trp86 | −7.590 | H-bond with Ser198 |
| π–cation with Tyr337 | π–π stacking with Trp82 | |||
| π–cation with Phe338 | π–cation with Tyr332 | |||
| π–cation with Tyr341 | ||||
| Salt bridge with Asp74 | ||||
| H-bound with Phe295 | ||||
| π–π stacking Trp286 | ||||
| 7b | −9.43 | π–π stacking Hip447 | −9.04 | Two π–π stacking with Trp231 |
| π–cation with Hip447 | H-bond with Ser198 H-bond with Gly117 | |||
| H-bond with Ser203 | π–cation with Asp70 | |||
| H-bond with Glh122 | π–π stacking with Tyr332 | |||
| H-bond with Tyr124 | ||||
| π–cation with Trp286 | ||||
| 7c | −8.608 | π–π stacking with Tyr337 | −8.641 | π–π stacking with Trp332 |
| π–π stacking with Trp86 | π–cation with Phe329 | |||
| π–cation with Trp86 | π–π stacking with Trp430 | |||
| H-bond with Phe295 | H-bonding with His438 | |||
| π–π stacking with Trp286 | ||||
| 7d | −9.559 | π–π stacking with Trp86 | −7.919 | π–π stacking with Trp82 |
| π–π stacking with Tyr341 a π–cation with Asp74 | π–cation with Asp70 | |||
| Two H-bonds with Phe295 | H-bond with Ser198 | |||
Molecular docking was performed against BChE, and the binding poses of compound 7b and donepezil are shown in Fig. 6. The aromatic ring of the cinnamic moiety in compound 7b formed two π–π stacking interactions with Trp231. Additionally, the carbonyl group of the cinnamic acid formed two hydrogen bonds, one with Ser198 (a key residue of the catalytic triad) and the other with Gly117 (located in the oxyanion hole); both are crucial for catalytic function. The pyridinium moiety underwent a π–cation interaction with Asp70, while the terminal 2-fluorobenzyl group performed a π–π stacking interaction with Tyr332 instead, which relates to PAS of BChE.These findings highlight the dual binding potential of compound 7b, showing effective interactions with both the CAS and the PAS in both cholinesterase enzymes.
 | ||
| Fig. 6 Superimposed binding poses of compound 7b (blue) and donepezil (green) within the active site of BChE. | ||
Compound 7c (3-fluorobenzyl) exhibited π–π stacking with Trp332, a pyridinium π–cation interaction with Phe329, additional π–π stacking with Trp430, and hydrogen bonding with His438. In 7d (4-fluorobenzyl), π–π stacking occurred with Trp82, the pyridinium π–cation interacted with Asp70, and a hydrogen bond was formed with Ser198. The chloro-substituted compounds 7e–g exhibited approximately similar binding energy, suggesting that the substituent position does not significantly affect overall binding affinity against BChE.
2.8 Molecular dynamics simulations
Molecular dynamics simulations were performed to evaluate the stability of the AChE–7b complex, compared with apo AChE form. The RMSD fluctuations of AChE were natural, with an average of 1.2 Å. On the contrary, the RMSD of the AChE–7b complex was much less compared to the unbound state, averaging only 0.6 Å. This highlights the stability of AChE–7b complex, thus making compound 7b a prospective strong AChE inhibitor (Fig. 7). | ||
| Fig. 7 RMSD plot of AChE backbone (Apo, velvet color) and compound 7b (orange) throughout the 100 ns of the simulation time. | ||
The interactions of compound 7b within AChE active site were further explored by MD simulation; the 4-oxo-4-phenylbutanamido moiety formed three hydrogen bonds with Gly122, Tyr124, and Ser203 (Fig. 8a). A persistent hydrogen bond was evidenced to Tyr124 and was present for about 96% of the simulation time. The pyridinium linker stabilized the complex through π–cation interaction with Trp286 and through π–π stacking with Tyr341, both residues being located at the PAS. RMSF analysis of AChE in complex with 7b showed most residues below 2 Å RMSF, confirming protein structural stability over simulation. These interactions were preserved across MD snapshots, providing further evidence for the stability of the AChE–7b complex (Fig. 8b).
 | ||
| Fig. 8 (a) 2D interaction diagram and (b) timeline interaction of compound 7b-AChE complex, which is responsible for over 30% of MD simulation time. | ||
The RMSF analysis of compound 7b is carried on inside the AChE active site. The RMSD less than 2 Å indicates proper fitting in the active site. All atoms of 7b recorded RMSF values of less than 1 Å, showing that the ligand is properly anchored in the active site. Such stable binding ensured that, throughout the simulation, both CAS and PAS sites on the enzyme were occupied, making it a potential candidate for an efficient inhibitor (Fig. 9).
2.9 In silico drug-likeness, ADME, and toxicity studies
In silico ADMET profiling of the most potent compound 7b (Fig. 10 and Table 4) revealed a highly favorable profile relative to the reference drug, donepezil (Fig. 11 and Table 5). Both compounds fulfil Lipinski's rule with no violations, but 7b exhibits lower lipophilicity (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P 2.43 vs. 4.19), higher TPSA (50.05 vs. 38.77 Å2), and full compliance with the Pfizer 3/75 rule compared with donepezil. Compound 7b shows good intestinal absorption, comparable Caco-2 permeability, markedly lower plasma protein binding (34.51% vs. 87.74%), similar volume of distribution, reduced CYP substrate liability, slightly lower clearance, longer half-life, and no predicted mutagenicity, carcinogenicity, or skin sensitization overall indicating superior drug-like properties, lower drug–drug interaction risk, and potentially improved brain availability compared to donepezil, thereby supporting 7b as a promising and safe anti-AD.
P 2.43 vs. 4.19), higher TPSA (50.05 vs. 38.77 Å2), and full compliance with the Pfizer 3/75 rule compared with donepezil. Compound 7b shows good intestinal absorption, comparable Caco-2 permeability, markedly lower plasma protein binding (34.51% vs. 87.74%), similar volume of distribution, reduced CYP substrate liability, slightly lower clearance, longer half-life, and no predicted mutagenicity, carcinogenicity, or skin sensitization overall indicating superior drug-like properties, lower drug–drug interaction risk, and potentially improved brain availability compared to donepezil, thereby supporting 7b as a promising and safe anti-AD.
| Parameters | Value | Parameters | Value | ||
|---|---|---|---|---|---|
| Medicinal chemistry | Lipinski rule | Accepted | Distribution | PPB | 34.51 |
| Pfizer rule | Accepted | VD (volume distribution) | 1.685 | ||
| SAscore | Accepted | Metabolism | CYP1A2 inhibitor | — | |
| Absorption | Caco-2 permeability | −5.112 | CYP1A2 substrate | — | |
| HIA (human intestinal absorption) | + | CYP2C9 inhibitor | — | ||
| Toxicity | Carcinogenicity | — | CYP2C9 substrate | — | |
| AMES toxicity | — | CYP2D6 inhibitor | + | ||
| Skin sensitization | — | CYP2D6 substrate | — | ||
| LC50FM | 3.323 | Excretion | CL (clearance rate) | 9.206 | |
| LC50DM | 4.724 | T1/2 | 0.827 |
| Parameters | Value | Parameters | Value | ||
|---|---|---|---|---|---|
| Medicinal chemistry | Lipinski rule | Accepted | Distribution | PPB | 87.74 |
| Pfizer rule | Rejected | VD (volume distribution) | 1.589 | ||
| SAscore | Accepted | Metabolism | CYP1A2 inhibitor | — | |
| Absorption | Caco-2 permeability | −4.793 | CYP1A2 substrate | + | |
| HIA (human intestinal absorption) | — | CYP2C9 inhibitor | — | ||
| Toxicity | Carcinogenicity | — | CYP2C9 substrate | — | |
| AMES toxicity | — | CYP2D6 inhibitor | + | ||
| Skin sensitization | — | CYP2D6 substrate | + | ||
| LC50FM | 5.338 | Excretion | CL (clearance rate) | 10.635 | |
| LC50DM | 6.367 | T1/2 | 0.164 |
3. Conclusion
The findings of the current research demonstrate effective design and synthesis of a novel series of cinnamic acid analogues conjugated with N-benzyl pyridinium, which are potential dual inhibitors of AChE and BChE and exhibit excellent neuroprotective activity. Compound 7b demonstrated promising inhibitory potential with IC50 = 0.89 µM against AChE and IC50 = 0.11 µM against BChE. SAR research demonstrated that electron-withdrawing small substituents, like ortho-fluoro groups, substantially elevate potency and selectivity, while bulk or electron-donating meta- and para-substitutions reduce potency. Kinetic assays confirmed 7b as a competitive AChE inhibitor (Ki = 0.49 µM), and molecular docking and molecular dynamics simulations supported its stable binding to both CAS and PAS of AChE and BChE via π–π stacking, π–cation interactions, and hydrogen bonding. Furthermore, this study highlights 7b low cytotoxicity to SH-SY5Y neuroblastoma cells at 10 µM and a potent neuroprotective effect against oxidative stress induced by H2O2. Overall, the current study showed cinnamic acid-derived N-benzyl pyridinium analogues as promising MTDLs of AD with optimal cognitive enhancement, neuroprotection, and safety.4. Methods and materials
All the solvents and reagents were purchased from Merck Chemical Company and used without further purification. The melting points were determined using an Electrothermal 9100 instrument. The 1HNMR and 13CNMR spectra were recorded with a Bruker 500 using TMS as an internal standard. Chemical shifts were reported at room temperature in ppm scale with DMSO-d6 as the solvents due to the poor solubility of the salts in non-polar solvents. SH-SY5Y cells were obtained from the Pasteur Institute of Iran (https://en.pasteur.ac.ir/).4.1 Chemistry
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to give the pure compound 5 as an off-white solid (yield: 0.233 g, 87%).
1) to give the pure compound 5 as an off-white solid (yield: 0.233 g, 87%).4.1.3.1 1-Benzyl-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7a). Yield: 85%. Cream powder. 1H NMR (400 MHz, DMSO) δ 9.20 (d, J = 6.4 Hz, 2H, (H-2,6 pyridinium)), 8.85 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.55–8.48 (m, 1H), 8.18 (dd, J = 8.1, 6.0 Hz, 1H), 8.02–7.95 (m, 2H), 7.65 (td, J = 7.2, 1.3 Hz, 1H), 7.61–7.49 (m, 4H), 7.42 (dd, J = 5.1, 1.9 Hz, 3H), 5.92 (s, 2H, CH2–N+ (benzyl)), 4.50 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.33 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.62 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm. 13C NMR (101 MHz, DMSO) δ: 29.56, 33.84, 39.36, 39.56, 39.77, 39.81, 39.98, 40.19, 40.40, 40.61, 63.76, 128.35, 128.52, 129.21, 129.66, 129.83, 133.73, 134.79, 136.90, 141.71, 143.37, 143.65, 144.66, 172.76 (CONH), 199.68 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.2 1-(2-Fluorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7b). Yield: 70%, dark yellow solid. 1H NMR (500 MHz, DMSO) δ 9.08 (s, 1H, (H-2 pyridinium)), 9.03 (d, J = 6.1 Hz, 1H, (H-6 pyridinium)), 8.93 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.56 (d, J = 8.0 Hz, 1H), 8.17 (dd, J = 8.1, 6.0 Hz, 1H), 7.97–7.92 (m, 2H), 7.65–7.60 (m, 1H), 7.59–7.44 (m, 6H), 7.41 (td, J = 7.3, 1.4 Hz, 1H), 6.04 (s, 2H, CH2–N+ (benzyl)), 4.49 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.29 (t, J = 6.5 Hz, 2H, (CH2–CO–Ph)), 2.59 (t, J = 6.5 Hz, 2H, (CH2–CH2–CO)) ppm, δ13CNMR (126 MHz, DMSO): 29.16, 33.39, 39.11, 39.27, 39.36, 39.44, 39.57, 39.61, 39.77, 39.94, 40.11, 61.23, 125.75, 127.80, 127.89, 128.07, 128.65, 130.01, 131.34, 131.37, 133.13, 133.21, 136.47, 141.11, 143.41, 143.52, 144.69, 172.20 (CONH), 199.04 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm, HRMS (ESI) (m/z): found for C23H22FN2O2+ [M + H]+ 377.0838.
O) ppm, HRMS (ESI) (m/z): found for C23H22FN2O2+ [M + H]+ 377.0838.
4.1.3.3 1-(3-Fluorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7c). Cream solid; yield: 73%, 1H NMR (500 MHz, DMSO) δ 9.13 (d, J = 6.8 Hz, 2H, (H-2,6 pyridinium)), 8.73 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.50 (d, J = 8.1 Hz, 1H), 8.17 (dd, J = 8.1, 5.8 Hz, 1H), 7.99–7.93 (m, 2H), 7.63 (t, J = 7.3 Hz, 1H), 7.55–7.41 (m, 4H), 7.38 (d, J = 7.6 Hz, 1H), 7.25 (td, J = 8.6, 2.7 Hz, 1H), 5.89 (s, 2H, CH2–N+ (benzyl)), 4.49 (d, J = 5.9 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.5 Hz, 2H, (CH2–CO–Ph)), 2.59 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm, δ 13C NMR (101 MHz, DMSO): 29.52, 33.80, 39.34, 39.54, 39.65, 39.75, 39.96, 40.17, 40.38, 63.05, 116.88, 125.51, 128.34, 128.57, 129.20, 131.77, 131.77, 133.75, 139.47, 141.78, 143.49, 144.78, 172.77 (CONH), 190.12 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.4 1-(4-Fluorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7d). Cream solid; yield: 76%, 1H NMR (400 MHz, DMSO) δ 9.24–9.17 (m, 2H, (H-2,6 pyridinium)), 8.80 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.56–8.49 (m, 2H), 8.27 (dd, J = 8.1, 2.1 Hz, 1H), 8.20 (dd, J = 8.1, 6.0 Hz, 1H), 8.02 (dd, J = 7.8, 1.3 Hz, 1H), 7.98–7.94 (m, 2H), 7.73 (t, J = 8.0 Hz, 1H), 7.67–7.62 (m, 1H), 7.53 (t, J = 7.7 Hz, 2H), 6.04 (s, 2H CH2–N+ (benzyl)), 4.49 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.32 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.60 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm, δ13CNMR(126 MHz, DMSO): 28.95, 29.21, 33.40, 39.09, 39.26, 39.34, 39.43, 39.59, 39.76, 39.93, 40.09, 61.83, 115.90, 116.07, 127.79, 127.95, 128.64, 130.42, 130.44, 131.39, 131.46, 133.12, 136.47, 141.11, 142.95, 143.01, 144.23, 161.53, 163.49, 172.22 (CONH), 199.12 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O).
O).
4.1.3.5 1-(2-Chlorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7e). Yield: 91%. Cream-pink powder. 1H NMR (400 MHz, DMSO) δ 9.09 (s, 1H, (H-2 pyridinium)), 9.05 (d, J = 6.1 Hz, 1H, (H-6 pyridinium)), 8.98 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.58 (d, J = 8.0 Hz, 1H), 8.19 (dd, J = 8.0, 6.0 Hz, 1H), 8.00–7.93 (m, 2H), 7.64 (dd, J = 8.3, 6.5 Hz, 1H), 7.59–7.42 (m, 6H), 6.05 (s, 2H, CH2–N+ (benzyl)), 4.51 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.60 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm. 13C NMR (101 MHz, DMSO) δ: 29.56, 33.85, 61.72, 128.33, 128.42, 128.60, 129.19, 130.52, 131.74, 131.86, 131.95, 133.65, 133.72, 136.87, 141.61, 143.86, 144.09, 145.15, 172.75 (CONH), 199.58 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.6 1-(3-Chlorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7f). Yield: 78%. Cream powder. 1H NMR (500 MHz, DMSO) δ 9.27–9.21 (m, 2H, (H-2,6 pyridinium)), 8.82 (t, J = 19.1, 5.9 Hz, 1H, (N–H amide)), 8.50 (dd, J = 14.9, 8.1 Hz, 1H), 8.20–8.16 (m, 1H), 8.16–8.09 (m, 1H), 7.98–7.92 (m, 2H), 7.86 (q, J = 2.0 Hz, 1H), 7.62–7.56 (m, 3H), 7.49 (t, J = 7.7 Hz, 2H), 7.35 (t, J = 7.9 Hz, 1H), 5.94 (s, 2H, CH2–N+ (benzyl)), 4.48 (d, J = 5.7 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.5 Hz, 2H, (CH2–CO–Ph)), 2.60 (t, J = 6.5 Hz, 2H, (CH2–CH2–CO)) ppm. 13C NMR (126 MHz, DMSO): 29.18, 33.44, 39.08, 39.24, 39.37, 39.41, 39.58, 39.75, 39.91, 40.08, 62.19, 122.21, 127.86, 127.97, 128.05, 128.08, 128.70, 131.29, 131.65, 131.68, 132.26, 133.21, 136.41, 136.59, 141.18, 143.00, 143.05, 143.07, 143.18, 144.28, 144.35, 172.27 (CONH), 199.16 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.7 1-(4-Chlorobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7g). Yield: 89%. Pink powder. 1H NMR (400 MHz, DMSO) δ 9.21 (t, J = 3.5 Hz, 2H, (H-2,6 pyridinium)), 8.95 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.52 (d, J = 8.3 Hz, 1H), 8.18 (dd, J = 8.0, 6.1 Hz, 1H), 8.01–7.93 (m, 2H), 7.67–7.60 (m, 3H), 7.55–7.46 (m, 4H), 5.92 (s, 2H, CH2–N+ (benzyl)), 4.49 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.32 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.62 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm. 13C NMR (101 MHz, DMSO) δ: 29.56, 33.83, 39.36, 39.57, 39.78, 39.83, 39.98, 40.19, 40.40, 40.61, 62.83, 128.34, 128.53, 129.19, 129.63, 131.30, 133.66, 133.72, 134.65, 136.90, 141.71, 143.47, 143.68, 144.74, 172.75 (CONH), 199.66 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm, HRMS (ESI) (m/z): found for C23H22ClN2O2+ [M + H]+ 393.0533.
O) ppm, HRMS (ESI) (m/z): found for C23H22ClN2O2+ [M + H]+ 393.0533.
4.1.3.8 1-(4-Bromoobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7h). Beige solid; yield: 75%, 1H NMR (500 MHz, DMSO) δ 9.24–9.18 (m, 2H, (H-2,6 pyridinium)), 8.94 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.54–8.49 (m, 1H), 8.16 (dd, J = 8.1, 6.0 Hz, 1H), 7.98–7.92 (m, 2H), 7.65–7.58 (m, 3H), 7.53–7.41 (m, 4H), 5.93 (s, 2H, CH2–N+ (benzyl)), 4.48 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.30 (t, J = 6.5 Hz, 2H, (CH2–CO–Ph)), 2.61 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm, δ13CNMR(101 MHz, DMSO): 29.50, 33.77, 39.34, 39.54, 39.75, 39.96, 40.17, 40.38, 40.59, 44.80, 62.70, 124.47, 124.50, 124.60, 128.31, 128.67, 129.18, 129.51, 130.37, 130.54, 130.59, 133.73, 136.87, 140.19, 141.68, 141.83, 143.76, 144.05, 145.02, 148.32, 172.79 (CONH), 199.66 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.9 3-((4-Oxo-4-phenylbutanamido)methyl)-1-(4-(trifluoromethyl)benzyl)pyridin-1-ium bromide (7i). Beige solid; yield: 76%, 1H NMR (500 MHz, DMSO) δ 9.23–9.18 (m, 2H, (H-2,6 pyridinium)), 8.85 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.51 (d, J = 8.1 Hz, 1H), 8.17 (dd, J = 8.1, 6.0 Hz, 1H), 8.03 (s, 1H), 7.98–7.93 (m, 2H), 7.85 (d, J = 7.8 Hz, 1H), 7.77 (d, J = 7.8 Hz, 1H), 7.64 (dt, J = 12.6, 7.6 Hz, 2H), 7.51 (t, J = 7.6 Hz, 2H), 5.99 (s, 2H, CH2–N+ (benzyl)), 4.48 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.5 Hz, 2H, (CH2–CO–Ph)), 2.59 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)) ppm. 13C NMR (101 MHz, DMSO) δ 29.5, 33.77, 39.34, 39.44, 39.54, 39.75, 39.96, 40.17, 40.38, 40.59, 45.9, 62.7, 114.58, 124.47, 124.5, 124.6, 128.31, 128.67, 129.18, 129.51, 130.37, 130.54, 130.59, 133.73, 136.87, 140.19, 141.68, 141.83, 143.76, 144.05, 145.02, 148.32, 172.79 (CONH), 199.66 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.10 1-(4-Nitrobenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7j). Dark pink solid; yield: 87%, 1H NMR (400 MHz, DMSO) δ 9.19–9.15 (m, 2H, (H-2,6 pyridinium)), 8.81 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.53 (d, J = 8.0 Hz, 1H), 8.20 (dd, J = 8.0, 6.1 Hz, 1H), 8.01–7.94 (m, 2H), 7.80 (d, J = 8.2 Hz, 2H), 7.73 (d, J = 8.1 Hz, 2H), 7.68–7.62 (m, 1H), 7.53 (t, J = 7.7 Hz, 2H), 6.01 (s, 2H, CH2–N+ (benzyl)), 4.50 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.32 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.61 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)). 13C NMR (101 MHz, DMSO) δ (101 MHz, DMSO): 29.50, 33.78, 39.34, 39.55, 39.76, 39.86, 39.97, 40.17, 40.38, 40.59, 63.00, 123.03, 125.74, 126.44, 126.48, 126.52, 126.56, 128.32, 128.63, 129.18, 129.93, 130.23, 133.73, 136.88, 139.19, 141.79, 143.65, 143.94, 144.90, 172.77 (CONH), 199.67 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.11 1-(4-Methoxybenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium bromide (7k). Purple solid; yield: 75%, 1H NMR (500 MHz, DMSO) δ 9.10 (d, J = 8.2 Hz, 2H, (H-2,6 pyridinium)), 8.73 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.48 (d, J = 8.1 Hz, 1H), 8.14 (dd, J = 8.1, 5.9 Hz, 1H), 7.97 (d, J = 7.7 Hz, 2H), 7.64 (t, J = 7.3 Hz, 1H), 7.52 (t, J = 7.6 Hz, 2H), 7.32 (t, J = 7.9 Hz, 1H), 7.14 (t, J = 2.1 Hz, 1H), 7.07 (d, J = 7.6 Hz, 1H), 6.97 (dd, J = 8.3, 2.6 Hz, 1H), 5.80 (s, 2H, CH2–N+ (benzyl)), 4.48 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.73 (s, 3H, (–OCH3)), 3.31 (t, 2H, (CH2–CO–Ph)), 2.58 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)). δ13CNMR(126 MHz, DMSO): 13.48, 19.21, 23.07, 29.06, 33.32, 39.04, 39.20, 39.37, 39.54, 39.70, 39.87, 40.04, 55.22, 57.52, 57.55, 63.42, 114.52, 114.75, 120.65, 127.85, 128.02, 128.72, 130.42, 133.26, 135.53, 136.43, 141.23, 142.81, 143.17, 144.20, 159.66, 172.28 (CONH), 199.21 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.12 1-(4-Methylbenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7l). Cream solid; yield: 80%, 1H NMR (500 MHz, DMSO) δ 9.14 (d, J = 6.9 Hz, 2H, (H-2,6 pyridinium)), 8.80 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.49 (d, J = 8.0 Hz, 1H), 8.18–8.12 (m, 1H), 7.96 (d, J = 7.7 Hz, 2H), 7.62 (t, J = 7.4 Hz, 1H), 7.51 (t, J = 7.6 Hz, 2H), 7.45 (d, J = 7.8 Hz, 2H), 7.20 (d, J = 7.7 Hz, 2H), 5.84 (s, 2H, CH2–N+ (benzyl)), 4.48 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.60 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)), 2.25 (s, 3H) ppm, δ13CNMR (126 MHz, DMSO): 20.70, 29.10, 33.35, 39.02, 39.19, 39.35, 39.52, 39.69, 39.78, 39.85, 39.95, 40.02, 63.13, 91.62, 125.77, 127.49, 127.81, 127.92, 128.16, 128.67, 128.76, 128.86, 129.65, 131.27, 133.17, 136.46, 138.91, 139.68, 141.15, 142.99, 144.10, 144.50, 172.20, 174.73 (CONH), 199.13 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm.
O) ppm.
4.1.3.13 1-(2-Methylbenzyl)-3-((4-oxo-4-phenylbutanamido)methyl)pyridin-1-ium chloride (7m). Yield: 74%. Dark cream powder. 1H NMR (400 MHz, DMSO) δ 9.00 (s, 1H, (H-6 pyridinium)), 8.97 (d, J = 6.1 Hz, 1H), 8.89 (t, J = 5.9 Hz, 1H, (N–H amide)), 8.55 (d, J = 8.0 Hz, 1H), 8.21–8.15 (m, 1H), 7.99–7.94 (m, 2H), 7.68–7.62 (m, 1H), 7.53 (t, J = 7.6 Hz, 2H), 7.35–7.30 (m, 2H), 7.26–7.22 (m, 1H), 7.11 (d, J = 7.6 Hz, 1H), 5.95 (s, 2H, CH2–N+ (benzyl)), 4.50 (d, J = 5.8 Hz, 2H, (–CH2–CONH)), 3.31 (t, J = 6.4 Hz, 2H, (CH2–CO–Ph)), 2.59 (t, J = 6.4 Hz, 2H, (CH2–CH2–CO)), 2.30 (s, 3H) ppm. 13C NMR (101 MHz, DMSO) δ 19.31, 29.53, 33.81, 39.85, 62.07, 127.16, 128.33, 128.48, 129.2, 129.33, 129.81, 131.37, 132.77, 133.73, 136.88, 137.34, 141.62, 143.61, 143.88, 144.85, 172.75 (CONH), 199.58 (C
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm, HRMS (ESI) (m/z): found for C24H25N2O2+ [M + H]+ 373.1218.
O) ppm, HRMS (ESI) (m/z): found for C24H25N2O2+ [M + H]+ 373.1218.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) ppm, HRMS (ESI) (m/z): found for C17H19N2O2+ [M + H]+ 283.0973.
O) ppm, HRMS (ESI) (m/z): found for C17H19N2O2+ [M + H]+ 283.0973.4.2 Screening of AChE and BChE inhibitory activity
Cholinesterase inhibitory activities of all derivatives were assessed using the modified Ellman method using butyrylcholinesterase (BChE, E.C. 3.1.1.8, from horse serum), and acetylcholinesterase (AChE, E.C. 3.1.1.7, Type V-S, lyophilized powder)174.3 Enzyme kinetic studies
The mode of inhibition for the most active compound, 7b, against AChE was investigated using acetylthiocholine as a substrate at concentrations ranging from 0.1 to 1 mM. A Lineweaver–Burk plot was generated, plotting the reciprocal of the substrate concentration (1/[S]) against the reciprocal of the enzyme rate (1/V) across various inhibitor concentrations. This plot was used to identify the type of inhibition and determine the Michaelis–Menten constant (Km) value.18,194.4 Cell viability
SH-SY5Y cells were cultured in DMEM/F12 medium supplemented with 15% FBS, penicillin, and streptomycin, and maintained at 37 °C with 5% CO2 cell viability was evaluated using the MTT assay. Cells were seeded in 96-well plates, treated with various concentrations of test compounds for 72 hours, and incubated with MTT reagent (0.5 mg mL−1) for 2 hours. The resulting formazan crystals were dissolved in DMSO, and absorbance was measured at 540 nm. Untreated cells served as the 100% viability control.20,214.5 Neuroprotectivity assay on SH-SY5Y
SH-SY5Y cells were cultured and incubated for 24 hours. Subsequently, 100 µM H2O2 was added to each well of a 96-well plate, except for the healthy control group. After 4 hours of H2O2 exposure, various concentrations of the test compound were added. Following a 72 hour incubation period, cell viability was assessed using the MTT assay.224.6 Molecular docking
The molecular docking of compound 7b was performed using the Schrödinger Suites Maestro molecular modeling platform. The X-ray crystallographic structures of AChE and BChE used for this purpose were obtained from the RCSB Protein Data Bank with PDB IDs 4EY7 and 4BDS.234.7 MD simulations
The starting model for the molecular dynamics simulations was obtained by imposing induced fit docking to AChE. The MD simulations themselves were conducted using Desmond v5.3 of Schrodinger's Suite, Maestro, following previously reported procedures.244.8 Prediction of pharmacokinetic properties of 7c and donepezil
The physicochemical and biological absorption, distribution, metabolism, excretion, and toxicity (ADMET) properties of the selected compounds were studied. These predictions were generated using reliable tools, including PKCSM (https://biosig.lab.uq.edu.au/pkcsm/) and ADMETmesh https://admetlab3.scbdd.com/and https://admetmesh.scbdd.com/.Author contributions
M. E. synthesized compounds and contributed to the characterization of compounds. A. I. performed in silico and biological studies. S. J. and M. M. supervised the synthetic part of the study. All authors read and approved the final version of the article.Conflicts of interest
There are no conflicts to declare.Data availability
The datasets generated and/or analyzed during the current study are available in the Worldwide ProteinData Bank with the PDB IDs of the 4EY7 and 4BDS repository. Supplementary information: 1H NMR and 13C NMR spectra, is available. See DOI: https://doi.org/10.1039/d5ra06941f.Acknowledgements
The authors wish to thank the support of the Vice-Chancellor for Research of Shiraz University of Medical Sciences (grant number = IR.SUMS.REC.1404.267).References
- R. Tenchov, J. M. Sasso and Q. A. Zhou, ACS Chem. Neurosci., 2024, 15, 3800–3827 CrossRef CAS PubMed.
- Z. Breijyeh and R. Karaman, Molecules, 2020, 25, 5789 CrossRef CAS.
- M. Yazdani, N. Edraki, R. Badri, M. Khoshneviszadeh, A. Iraji and O. Firuzi, Mol. Diversity, 2020, 24, 641–654 CrossRef CAS.
- Z. Haghighijoo, S. Akrami, M. Saeedi, A. Zonouzi, A. Iraji, B. Larijani, H. Fakherzadeh, F. Sharifi, S. M. Arzaghi, M. Mahdavi and N. Edraki, Bioorg. Chem., 2020, 103, 104146 CrossRef CAS PubMed.
- M. A. Better, Alzheimers Dement, 2023, 19, 1598–1695 CrossRef PubMed.
- N. Oliyaei, M. Moosavi-Nasab, N. Tanideh and A. Iraji, Brain Res. Bull., 2023, 193, 11–21 CrossRef CAS.
- H. Pourtaher, A. Hasaninejad and A. Iraji, Sci. Rep., 2022, 12, 15236 CrossRef CAS PubMed.
- H. Pourtaher, A. Hasaninejad, S. Zare, N. Tanideh and A. Iraji, Sci. Rep., 2023, 13, 11952 CrossRef CAS.
- J. Zhang, Y. Zhang, J. Wang, Y. Xia, J. Zhang and L. Chen, Signal Transduction Targeted Ther., 2024, 9, 211 CrossRef CAS.
- H. Xing, S. Yue, R. Qin and X. Du, Int. J. Mol. Sci., 2025, 26, 3905 CrossRef CAS PubMed.
- A. P. Kamath, P. G. Nayak, J. John, S. Mutalik, A. K. Balaraman and N. Krishnadas, Neuropharmacology, 2024, 259, 110096 Search PubMed.
- J.-S. Lan, T. Zhang, Y. Liu, J. Yang, S.-S. Xie, J. Liu, Z.-Y. Miao and Y. Ding, Eur. J. Med. Chem., 2017, 133, 184–196 Search PubMed.
- F. Abedinifar, S. M. F. Farnia, M. Mahdavi, H. Nadri, A. Moradi, J. B. Ghasemi, T. T. Küçükkılınç, L. Firoozpour and A. Foroumadi, Bioorg. Chem., 2018, 80, 180–188 CrossRef CAS.
- J.-S. Lan, J.-W. Hou, Y. Liu, Y. Ding, Y. Zhang, L. Li and T. Zhang, J. Enzyme Inhib. Med. Chem., 2017, 32, 776–788 CrossRef CAS.
- G. Ghotbi, M. Mahdavi, Z. Najafi, F. H. Moghadam, M. Hamzeh-Mivehroud, S. Davaran and S. Dastmalchi, Bioorg. Chem., 2020, 103, 104186 CrossRef CAS PubMed.
- M. D. Vitorović-Todorović, I. O. Juranić, L. M. Mandić and B. J. Drakulić, Bioorg. Med. Chem., 2010, 18, 1181–1193 CrossRef.
- H. Pourtaher, A. Hasaninejad and A. Iraji, Sci. Rep., 2022, 12, 15236 CrossRef CAS PubMed.
- A. Iraji, P. Nikfar, M. Nazari Montazer, M. Karimi, N. Edraki, M. Saeedi and S. S. Mirfazli, Sci. Rep., 2024, 14, 21115 CrossRef CAS.
- B. Sadeghian, A. Sakhteman, Z. Faghih, H. Nadri, N. Edraki, A. Iraji, I. Sadeghian and Z. Rezaei, J. Mol. Struct., 2020, 1221, 128793 CrossRef CAS.
- M. Noori, N. Dastyafteh, S. Safapoor, M. Khalili Ghomi, R. Tanideh, K. Zomorodian, H. Hamedifar, M. Dara, S. Zare, C. Irajie, S. Javanshir, H. Rastegar, N. Panahi, B. Larijani, M. Mahdavi, M. H. Hajimiri and A. Iraji, Int. J. Biol. Macromol., 2023, 253, 127392 CrossRef CAS.
- N. Edraki, A. Iraji, O. Firuzi, Y. Fattahi, M. Mahdavi, A. Foroumadi, M. Khoshneviszadeh, A. Shafiee and R. Miri, J. Iran. Chem. Soc., 2016, 13, 2163–2171 CrossRef CAS.
- A. Iraji, D. Sharghei-Boroujeni, A. Aliabadi, M. Dara, M. H. Hashempur, R. Hariri, D. A. M. Khassaki, M. Saeedi and T. Akbarzadeh, J. Mol. Struct., 2025, 143337 CrossRef CAS.
- H. Pourtaher, Y. Mohammadi, A. Hasaninejad and A. Iraji, RSC Med. Chem., 2024, 15, 207–222 RSC.
- A. Iraji, R. Hariri, M. H. Hashempur, M. Ghasemi, H. Pourtaher, M. Saeedi and T. Akbarzadeh, BMC Chem., 2025, 19, 97 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |