 Open Access Article
Open Access ArticleSelective detection of Cu2+ in aqueous medium using an acid hydrazide-based chemosensor: experimental and DFT/TDDFT studies
Joardar Gima,
Zannatul Kowser *a,
Dipa Debnatha,
Redika Sarmin Prieetya,
Miss. Tasnim Jahana,
Paul G. Waddell
*a,
Dipa Debnatha,
Redika Sarmin Prieetya,
Miss. Tasnim Jahana,
Paul G. Waddell b,
Most Tahera Khatuna,
Sahara Khatun Munnia,
Md. Rashed Khana and
Rashedul Islama
b,
Most Tahera Khatuna,
Sahara Khatun Munnia,
Md. Rashed Khana and
Rashedul Islama
aDepartment of Chemistry, Jashore University of Science and Technology, Jashore-7408, Bangladesh. E-mail: joardargim@gmail.com; zannatulkowser_che@just.edu.bd; debnathdipa018@gmail.com; redikaprity@gmail.com; tasnim.b920@gmail.com; msttaherakhatun92@gmail.com; saharakhatunmunni@gmail.com; rashedkhan039@gmail.com; riju3295@gmail.com
bFaculty of Science, Agriculture & Engineering, Newcastle University, Newcastle Upon Tyne, NE1 7RU, UK. E-mail: paul.waddell@ncl.ac.uk
First published on 20th January 2026
Abstract
An acid hydrazide-based Schiff base, N'-(4-hydroxybenzylidene)picolinohydrazide (HP), was designed and synthesized in a single-step process for the selective detection of Cu2+ ions. The structure of HP was thoroughly characterized by FTIR, 1H NMR, 13C NMR, HRMS, SCXRD and elemental analysis. The cation-sensing performance of HP was investigated using UV-Vis spectroscopy, revealing a rapid response with high selectivity and sensitivity toward Cu2+ in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, v/v) with a 10 mM HEPES buffer at pH 7.4, with a detection limit of 8.94 µM. Job's plot analysis confirmed a 1
4, v/v) with a 10 mM HEPES buffer at pH 7.4, with a detection limit of 8.94 µM. Job's plot analysis confirmed a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry, with an association constant of 4.26 × 104 M−1. FESEM analysis showed distinct morphological changes upon Cu2+ coordination. The SCXRD result exhibits good agreement with DFT and TD-DFT of the probe HP. The hole–electron analysis clearly reveals locally excited (LE) characteristics for the free ligand and intramolecular charge transfer (ICT) behavior for the HP–Cu2+ complex. Furthermore, the HP probe demonstrated practical applicability for Cu2+ detection in real water samples.
1 binding stoichiometry, with an association constant of 4.26 × 104 M−1. FESEM analysis showed distinct morphological changes upon Cu2+ coordination. The SCXRD result exhibits good agreement with DFT and TD-DFT of the probe HP. The hole–electron analysis clearly reveals locally excited (LE) characteristics for the free ligand and intramolecular charge transfer (ICT) behavior for the HP–Cu2+ complex. Furthermore, the HP probe demonstrated practical applicability for Cu2+ detection in real water samples.
1. Introduction
Ions of transition metals are crucial for human health and provide significant functions in diverse industries, such as medications, diagnostics, and catalysis.1 It has been proven that abnormalities in the amounts of specific metal ions may perturb normal biological activities.2 Copper is an essential micronutrient necessary for the growth and development of humans, animals, and plants. It is the third most prevalent trace element in the Earth's crust, behind iron and zinc.3 A component of almost every human tissue, copper has a role in many physiological functions, such as cellular metabolism, oxidative stress tolerance, and immunological response.4–6 Copper(II) ions (Cu2+) function as vital cofactors for numerous metalloenzymes, including cytochrome c oxidase, tyrosinase, and superoxide dismutase.7–9 Dysregulation of copper homeostasis has been associated with various neurodegenerative illnesses, including Alzheimer's dementia, Wilson's disease, Menkes syndrome, and amyotrophic lateral sclerosis (ALS).10–17 Moreover, copper has significant toxicity to microorganisms including algae, bacteria, viruses, and aquatic creatures.18 To safeguard human health, the World Health Organization (WHO) has set a maximum permissible level of 31.5 µM for Cu2+ in drinking water.19 The precise detection of copper ions, due to its biological and environmental importance, has garnered significant interest in life and environmental sciences.20,21 Despite the utilization of various analytical techniques—such as atomic absorption spectroscopy (AAS), anodic stripping voltammetry (ASV), and inductively coupled plasma optical emission spectrometry/mass spectrometry (ICP-OES/MS)—have been employed for copper ion detection,22–24 their extensive application is constrained by the necessity for costly equipment, skilled operators, and protracted procedures. In contrast, Chemosensors have developed as useful instruments for ion detection, owing to their simplicity, cost-efficiency, and capacity to deliver both qualitative and quantitative data. This allows us to quickly and effectively identify the target ions in a matter of seconds.In recent years, numerous chemosensors utilizing Schiff bases have been created, as the –C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N framework, when modified with suitable substituents, provides an advantageous platform for the identification of metal ions in aqueous or semi-aqueous environments.25 The electronic characteristics of Schiff bases can be precisely altered by structural alterations, resulting in unique spectroscopic reactions upon metal binding.
N framework, when modified with suitable substituents, provides an advantageous platform for the identification of metal ions in aqueous or semi-aqueous environments.25 The electronic characteristics of Schiff bases can be precisely altered by structural alterations, resulting in unique spectroscopic reactions upon metal binding.
In this study we have synthesized a novel Schiff base receptor, HP—an acid hydrazide derivative—via the condensation of 2-pyridinecarboxylic acid hydrazide and 4-hydroxybenzaldehyde. This receptor has significant sensitivity and selectivity for Cu2+ ions in an aqueous environment at pH 7.4. Interaction with Cu2+ ions lead HP to exhibit notable alterations in its UV-Vis absorbance spectrum, confirming complex formation. The suggested sensing mechanism is also validated by FTIR and DFT analysis. The structure of HP has been comprehensively characterized via FTIR, 1H NMR, 13C NMR, HRMS, SCXRD and elemental analysis. The probe has also been successfully employed in real sample analysis, underscoring its practical applicability.
2. Experimental
2.1 Materials and methods
All solvents and reagents used throughout the synthesis and spectroscopic analyses were of spectroscopic grade. The primary reagents, 2-pyridinecarboxylic acid hydrazide and 4-hydroxybenzaldehyde, were obtained from TCI (Japan) and utilized with no additional purification upon receipt. Fresh solutions of metal ions were made from their corresponding chloride salts, sourced from Sigma-Aldrich and TCI. FTIR (ATR) spectra were acquired utilizing a PerkinElmer Spectrum 2 spectrometer. The melting points were obtained using a Stuart SMP-30 instrument. 1H and 13C NMR spectra were obtained using Bruker Ascend 600 MHz NMR spectrometer, with chemical shifts (δ) expressed in parts per million (ppm) in relation to internal standards. Electrospray ionization mass spectrometry (ESI-MS) spectra were recorded in positive ion mode on a Thermo Scientific Orbitrap Exploris 120 mass spectrometer. UV-Visible absorption spectra were obtained using HITACHI UH-5300 double-beam spectrophotometer with a 1 cm path length quartz cuvette.2.2 X-ray data collection and structure determination
Single crystal diffraction data for HP were collected at 150 K on a XtaLAB Synergy HyPix-Arc 100 diffractometer using copper radiation (λCuKα = 1.54184 Å) equipped with an Oxford Cryosystems CryostreamPlus open-flow N2 cooling device. Intensities were corrected for absorption using a multifaceted crystal model created by indexing the faces of the crystal for which data were collected.26 Cell refinement, data collection and data reduction were undertaken via the software CrysAlisPro.27The structure was solved using XT28 and refined by XL29 using the Olex2 interface.30 All non-hydrogen atoms were refined anisotropically and hydrogen atoms were positioned with idealised geometry, with the exception of those bound to heteroatoms, the positions of which were located using peaks in the Fourier difference map. The displacement parameters of the hydrogen atoms were constrained using a riding model with U(H) set to be an appropriate multiple of the Ueq value of the parent atom. The crystallographic data of HP are listed in Table 1.
| CCDC 2515488 | |
| Empirical formula | C13H11N3O2 |
| Formula weight | 241.25 |
| Temperature/K | 150.0(2) |
| Crystal system | Orthorhombic |
| Space group | Pbca |
| a/Å | 11.9663(2) |
| b Å | 7.50140(10) |
| c/Å | 25.5657(4) |
| α/° | 90 |
| β/° | 90 |
| γ/° | 90 |
| Volume/Å3 | 2294.88(6) |
| Z | 8 |
| ρcalcg cm−3 | 1.397 |
| µ/mm−1 | 0.803 |
| F(000) | 1008.0 |
| Crystal size/mm3 | 0.18 × 0.06 × 0.05 |
| Radiation | CuKα (λ = 1.54184) |
| 2Θ range for data collection/° | 6.916 to 153.774 |
| Index ranges | −14 ≤ h ≤ 14, −6 ≤ k ≤ 9, −29 ≤ l ≤ 30 |
| Reflections collected | 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 496 496 |
| Independent reflections | 2268 [Rint = 0.0226, Rsigma = 0.0182] |
| Data/restraints/parameters | 2268/0/169 |
| Goodness-of-fit on F2 | 1.060 |
| Final R indexes [I> = 2σ (I)] | R1 = 0.0313, wR2 = 0.0832 |
| Final R indexes [all data] | R1 = 0.0344, wR2 = 0.0858 |
| Largest diff. Peak/hole/e Å−3 | 0.16/−0.21 |
2.3 Synthesis of probe HP
Scheme 1 illustrates the synthesis of HP. A mixture of 2-pyridine carboxylic acid hydrazide (0.567 g, 4.14 mmol) and 4-hydroxybenzaldehyde (0.506 g, 4.14 mmol) in ethanol (20 mL) was stirred for 4 h. A catalytic amount (2–3 drops) of acetic acid was added into the reaction. After the reaction completed as confirmed by TLC, the precipitate was filtered and washed with cold ethanol to give N'-(4-hydroxy benzylidene)picolinohydrazide. Crystals suitable for X-ray diffraction analysis were grown from the slow evaporation of a methanolic solution at room temperature. Yield: 76% mp: 223 °C. FT-IR (ATR, cm−1): 3237 (OH), 3198 (N–H), 1633 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1603 (C
O), 1603 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N). 1H NMR (600 MHz, DMSO-d6, ppm): δ 11.942 (1H, s, NH), 9.976 (1H, s, OH), 8.689–8.682 (1H, d, J = 4.2, H–Ar), 8.539 (1H, s, C
N). 1H NMR (600 MHz, DMSO-d6, ppm): δ 11.942 (1H, s, NH), 9.976 (1H, s, OH), 8.689–8.682 (1H, d, J = 4.2, H–Ar), 8.539 (1H, s, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) NH), 8.123–8.110 (1H, d, J = 7.8, H–Ar), 8.046–8.021 (1H, m, H–Ar), 7.652–7.629 (1H, m, H–Ar), 7.567–7.553(1H, d, J = 8.4, H–Ar), 6.856–6.842 (1H, d, J = 8.4, H–Ar). 13C NMR (150 MHz, CD3OD, ppm): δ 116.67, 123.77, 126.80, 128.11, 130.81, 138.92, 149.87, 150.56, 151.97, 161.47, 163.0. Elem. anal. cald. for HP (%): C, 64.72; H, 4.60; N, 17.42. Found: C, 61.94; H, 4.43; N, 16.16. HRMS-ESI(m/z) calcd for C13H11N3O2 [M + H]+242.0927, found 242.0922.
NH), 8.123–8.110 (1H, d, J = 7.8, H–Ar), 8.046–8.021 (1H, m, H–Ar), 7.652–7.629 (1H, m, H–Ar), 7.567–7.553(1H, d, J = 8.4, H–Ar), 6.856–6.842 (1H, d, J = 8.4, H–Ar). 13C NMR (150 MHz, CD3OD, ppm): δ 116.67, 123.77, 126.80, 128.11, 130.81, 138.92, 149.87, 150.56, 151.97, 161.47, 163.0. Elem. anal. cald. for HP (%): C, 64.72; H, 4.60; N, 17.42. Found: C, 61.94; H, 4.43; N, 16.16. HRMS-ESI(m/z) calcd for C13H11N3O2 [M + H]+242.0927, found 242.0922.
2.4 Procedure of measurement
The addition of methanol as a co-solvent was needed as probe HP is not wholly soluble in 100% aqueous medium. By dissolving the required amount, a stock solution of HP at a concentration of 1 mM was prepared. In tests, the solution of HP was further diluted to a suitable concentration (10 µM) with a mixed solution of MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4), and other metal ion solutions (1 mM) (e.g., Na+, Ni2+, NH4+, K+, Cu2+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Pb2+, Ag+) were made in the same solvent system. Then, in aqueous solvent medium MeOH/H2O (6
4 v/v, HEPES = 10 mM, pH 7.4), and other metal ion solutions (1 mM) (e.g., Na+, Ni2+, NH4+, K+, Cu2+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Pb2+, Ag+) were made in the same solvent system. Then, in aqueous solvent medium MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4), the UV-Vis spectra were recorded. Throughout the whole experiment, Millipore Milli-Q water was used. All UV-Vis studies were conducted at ambient temperature.
4 v/v, HEPES = 10 mM, pH 7.4), the UV-Vis spectra were recorded. Throughout the whole experiment, Millipore Milli-Q water was used. All UV-Vis studies were conducted at ambient temperature.
3. Results and discussion
3.1 Synthesis and characterization
The chemosensor HP was synthesized via a condensation reaction between 2-pyridine carboxylic acid hydrazide and 4-hydroxybenzaldehyde in good yield, as shown in Scheme 1. Sensor HP was fully characterized by various techniques such as FTIR, 1H NMR, 13C NMR, HRMS, SCXRD and elemental analysis. The results are well consistent with the expected chemical structure. The FTIR, 1H NMR, 13C NMR and HRMS spectra were shown in Fig. S1–S4 of the SI.3.2 UV-Vis spectral studies
Probe HP (10 µM) in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4) exhibited a clear absorption band at 323 nm with a shoulder peak at 223 nm in its UV-Vis absorption spectra (Fig. 1), the band at 323 nm was ascribed to n → π* transitions, whereas the band at 223 nm was designated for π → π* transitions. To examine the selective sensing capabilities of probe HP (10 µM) towards various metal ions (1 mM) of environmental and biological relevance, including Na+, Ni2+, NH4+, K+, Cu2+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Hg2+, Pb2+ and Ag+ these ions were introduced into a solution of HP in MeOH/H2O (6
4 v/v, HEPES = 10 mM, pH 7.4) exhibited a clear absorption band at 323 nm with a shoulder peak at 223 nm in its UV-Vis absorption spectra (Fig. 1), the band at 323 nm was ascribed to n → π* transitions, whereas the band at 223 nm was designated for π → π* transitions. To examine the selective sensing capabilities of probe HP (10 µM) towards various metal ions (1 mM) of environmental and biological relevance, including Na+, Ni2+, NH4+, K+, Cu2+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Hg2+, Pb2+ and Ag+ these ions were introduced into a solution of HP in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4). No notable spectrum alterations of probe HP were observed, with the exception of Cu2+. After the addition of Cu2+ to the solution of probe the absorption band of HP at 323 nm totally diminished, while a new band emerged at 280 nm. The results indicate that HP exhibits high sensitivity and selectivity for Cu2+ in a MeOH/H2O (6
4 v/v, HEPES = 10 mM, pH 7.4). No notable spectrum alterations of probe HP were observed, with the exception of Cu2+. After the addition of Cu2+ to the solution of probe the absorption band of HP at 323 nm totally diminished, while a new band emerged at 280 nm. The results indicate that HP exhibits high sensitivity and selectivity for Cu2+ in a MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4).
4 v/v, HEPES = 10 mM, pH 7.4).
 | ||
Fig. 1 UV-Vis absorption changes of probe HP (10 µM) in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4) in the presence of various metal ions. 4 v/v, HEPES = 10 mM, pH 7.4) in the presence of various metal ions. | ||
3.3 Interference of other metal ions
The interference test was conducted to assess the practical application of probe HP, as illustrated in Fig. 2. The competition experiments were performed by monitoring the absorbance variations before and after the addition of Cu2+ (1 × 10−3 M) to the HP solution containing various metal ion interferents (MeOH/H2O = 6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4) at 280 nm. The ligand, HP treated with Cu2+ in the presence of various mono- and divalent ions (Na+, Ni2+, NH4+, K+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Hg2+, Pb2+, and Ag+, each at 1 × 10−3 M) showed no significant spectral changes. These results indicate that the presence of other metal ions did not interfere with the detection of Cu2+ by the probe HP, as shown in Fig. 2. Furthermore, the interference study HP (10 µM) + Cu2+ (1 × 10−3 M) was extended to a wider concentration range of competing metal ions (up to 1 × 10−1 M), and even at this elevated concentration, no noticeable interference with the HP–Cu2+ signal was observed (Fig. S6).
4 v/v, HEPES = 10 mM, pH 7.4) at 280 nm. The ligand, HP treated with Cu2+ in the presence of various mono- and divalent ions (Na+, Ni2+, NH4+, K+, Fe3+, Cr3+, Al3+, Sn2+, Cd2+, Zn2+, Hg2+, Pb2+, and Ag+, each at 1 × 10−3 M) showed no significant spectral changes. These results indicate that the presence of other metal ions did not interfere with the detection of Cu2+ by the probe HP, as shown in Fig. 2. Furthermore, the interference study HP (10 µM) + Cu2+ (1 × 10−3 M) was extended to a wider concentration range of competing metal ions (up to 1 × 10−1 M), and even at this elevated concentration, no noticeable interference with the HP–Cu2+ signal was observed (Fig. S6).
 | ||
Fig. 2 UV-Vis absorption responses of probe HP (10 µM) in the presence of dual metal ions (Cu2+ and other specified metal ions, each at 1 × 10−3 M) in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4). 4 v/v, HEPES = 10 mM, pH 7.4). | ||
3.4 UV-Vis spectral titration and stoichiometry studies
To better understand the sensing ability of probe HP, we performed UV-Vis titrations of probe HP through the incremental addition of varying quantities of Cu2+ ions (0.1 to 12 equivalents) in a MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v) solution containing 10 mM HEPES at pH 7.4. In a 10 µM solution of HP, the titration graph shows a peak at 223 nm after the addition of 0.1 to 0.5 equivalents of Cu2+, which shifts to 230 nm for 1 to 12 equivalents of Cu2+. To gain a better understanding of the sensing ability of probe HP, UV-Vis titrations were performed by the incremental addition of varying concentrations of Cu2+ ions (0.1–12 equivalents) to a MeOH/H2O (6
4 v/v) solution containing 10 mM HEPES at pH 7.4. In a 10 µM solution of HP, the titration graph shows a peak at 223 nm after the addition of 0.1 to 0.5 equivalents of Cu2+, which shifts to 230 nm for 1 to 12 equivalents of Cu2+. To gain a better understanding of the sensing ability of probe HP, UV-Vis titrations were performed by the incremental addition of varying concentrations of Cu2+ ions (0.1–12 equivalents) to a MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, v/v) solution containing 10 mM HEPES at pH 7.4. In a 10 µM solution of HP, the titration spectrum exhibited a peak at 223 nm upon the addition of 0.1–0.5 equivalents of Cu2+, which shifted to 230 nm with the addition of 1–12 equivalents of Cu2+. Simultaneously, the absorption band of HP at 323 nm was diminished after the addition of 0.1–0.5 equivalents of Cu2+ and new absorption bands were appeared at 280 nm for 0.1–12 equivalents of Cu2+, indicating the formation of the HP–Cu2+ complex.
4, v/v) solution containing 10 mM HEPES at pH 7.4. In a 10 µM solution of HP, the titration spectrum exhibited a peak at 223 nm upon the addition of 0.1–0.5 equivalents of Cu2+, which shifted to 230 nm with the addition of 1–12 equivalents of Cu2+. Simultaneously, the absorption band of HP at 323 nm was diminished after the addition of 0.1–0.5 equivalents of Cu2+ and new absorption bands were appeared at 280 nm for 0.1–12 equivalents of Cu2+, indicating the formation of the HP–Cu2+ complex.
(Fig. 3). The stoichiometry of binding between HP and Cu2+ was found using the Job's plot approach, revealing a maximum absorbance at 0.5, as illustrated in Fig. 4. This particular point suggests that the stoichiometry of the binding between probe HP and Cu2+ is 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.
1.
 | ||
Fig. 3 UV-Vis absorption spectrum of probe HP (10 µM) in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4) with different concentrations (0.1–12 equiv.) of Cu2+. 4 v/v, HEPES = 10 mM, pH 7.4) with different concentrations (0.1–12 equiv.) of Cu2+. | ||
 | ||
Fig. 4 Job's Plot showing 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry between HP and Cu2+ in MeOH/H2O (6 1 binding stoichiometry between HP and Cu2+ in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4). 4 v/v, HEPES = 10 mM, pH 7.4). | ||
The limit of detection of probe HP was determined using the equation: detection limit = 3σ/m, where σ is the standard deviation of the blank solution, and m is the slope of the absorbance versus [Cu2+] calibration curve.31,32 The detection limits of HP toward Cu2+ ion is 8.94 µM (Fig. S5), this is much below the maximum allowable level of 31.5 µM for Cu2+ in drinking water as per World Health Organization standards (1993).19,33 Additionally, the Benesi–Hildebrand method was employed to ascertain the binding constant for HP and Cu2+ and was found to be 3.04 × 104 M−1.34–36 The 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry of the HP–Cu2+ complex was further confirmed by the linear relationship between 1/A − A0 and 1/[Cu2+] (Fig. 5).
1 binding stoichiometry of the HP–Cu2+ complex was further confirmed by the linear relationship between 1/A − A0 and 1/[Cu2+] (Fig. 5).
 | ||
Fig. 5 Benesi–Hildebrand plot (absorbance at 270 nm) of probe HP considering 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry with Cu2+. 1 binding stoichiometry with Cu2+. | ||
3.5 Microscopic studies
Furthermore, for a better understanding of the variations in surface topography, field emission scanning electron microscopy (FESEM) of HP and HP–Cu2+ were analyzed, as illustrated in Fig. 6a and b. FESEM photos reveal variations before and after complexation, displaying a highly crystalline morphology characterized by unique rod- or needle-like structures in probe HP. These elongated blocks exhibit smooth surfaces and sharp edges, indicating well-ordered molecular packing in the solid state. The morphology undergoes a significant transformation upon complexation with Cu2+. The formerly organized rod-shaped crystals are no longer discernible. | ||
| Fig. 6 Alterations in surface topography in FESEM images of (a) probe HP, (b) probe HP–Cu2+, and EDAX analysis of (c) probe HP and (d) probe HP–Cu2+. | ||
The chemical compositions of HP and HP–Cu2+ were determined using Energy Dispersive X-ray analysis (EDAX) (Fig. 6c and d), which directly confirms the presence of carbon (C), oxygen (O), nitrogen (N), and copper (Cu) components in the produced HP and HP–Cu2+ complex.
3.6 FT-IR analysis
FT-IR spectroscopy was utilized to elucidate the coordination mechanism and structural alterations throughout complexation. Characteristic vibrational bands of the free probe HP were identified at 3238 cm−1 (O–H stretching), 3201 cm−1 (N–H stretching), 1633 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching), and 1603 cm−1 (C
O stretching), and 1603 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching). Substantial spectrum alterations were found upon coordination with Cu2+ ions. The O–H and N–H stretching bands converged into a single broad absorption, indicating strong participation of these groups in coordination. The C
N stretching). Substantial spectrum alterations were found upon coordination with Cu2+ ions. The O–H and N–H stretching bands converged into a single broad absorption, indicating strong participation of these groups in coordination. The C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N stretching band changed marginally from 1603 to 1598 cm−1, indicating a contact between the imine nitrogen and the Cu2+. Conversely, the C
N stretching band changed marginally from 1603 to 1598 cm−1, indicating a contact between the imine nitrogen and the Cu2+. Conversely, the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching band remained fixed at 1633 cm−1, revealing no participation of the carbonyl group in metal coordination. These observations clearly corroborate the coordination of Cu2+ with the hydroxyl, amino, and imine nitrogen of probe HP (Fig. 7). The suggested binding mechanism is shown in Scheme 2, based on the aforementioned data.
O stretching band remained fixed at 1633 cm−1, revealing no participation of the carbonyl group in metal coordination. These observations clearly corroborate the coordination of Cu2+ with the hydroxyl, amino, and imine nitrogen of probe HP (Fig. 7). The suggested binding mechanism is shown in Scheme 2, based on the aforementioned data.
 | ||
| Fig. 7 FT-IR spectra of (a) probe HP (red) (b) probe HP–Cu2+(black). (Lower wavenumbers peak are at the inset). | ||
3.7 X-ray crystal structure and DFT-optimized structures
The main crystal parameters are reported in Table 1. The ORTEP and DFT-optimized diagrams of HP structures and their corresponding parameters are illustrated in Fig. 8. C13H11N3O2 is orthorhombic in the crystal system. The XRD-parameters of the HP ligand structure, like bond distances and the values of their angles, were matched with those derived from the computed DFT/B3LYP/6-311++G(d,p) calculation. An excellent match between calculated and measured results was illustrated, as can be seen in Fig. 9. The correlation between the experimental and calculated bond distances is 0.9718 (Fig. 9a and b). Similarly, the correlation between the calculated and experimental angles is 0.9608 (Fig. 9c and d). The DFT-optimized and ORTEP bond length and angles values have been reported in Tables S1 and S2. | ||
| Fig. 8 Ligand structure: (a) ORTEP diagram (50% probability) and (b) optimized ground state geometries at B3LYP/6-31G++(d,p). | ||
 | ||
| Fig. 9 (a) XRD/DFT-bond lengths histogram, (b) XRD/DFT-bond lengths graphical correlation, (c) XRD/DFT-angles histogram, and (d) DFT/Exp. angles correlation diagram. | ||
3.8 Frontier molecular orbital analysis
To elucidate the electronic structure, electron density distribution, kinetic stability, and reactivity of the sensor and its complex, Frontier Molecular Orbital Analysis was performed on isolated molecules. The HOMO is generally linked to electron-donating ability, whereas the LUMO is regarded as possessing electron-accepting aptitude. The HOMO and LUMO band gaps are crucial characteristics that influence the electronic behavior of molecules. The presence of positive frequencies for all the structures ensures minimum in the potential energy surface. The optimized geometries of HP and HP–Cu2+ complexes at the B3LYP/6-311++G(d,p) and LANL2DZ level of theory are shown in Fig. 10.Fig. 11 depicts the frontier molecular orbital (FMO) energies of the probe HP and it's complex. The computed HOMO and LUMO values for the free receptor HP are −6.37 eV and −1.70 eV, respectively, resulting in an energy gap of 4.67 eV. The coordination of Cu2+ with HP results in a reduction of the energy gap to 3.26 eV. Using the B3LYP/6-311++G(d,p) level of theory, the calculated FT-IR spectra of HP and its Cu2+ complex are displayed in Fig. 12. For receptor HP, four distinct absorption bands are observed at 1647, 1757, 3488, and 3831 cm−1, corresponding to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N, C
N, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, N–H, and O–H vibrations, respectively. Upon complexation with Cu2+, significant shifts in these vibrational bands are noted at 1654, 1577, 3412, and 3647 cm−1, respectively. These changes indicate strong interactions between the metal ion and the donor sites of the ligand.
O, N–H, and O–H vibrations, respectively. Upon complexation with Cu2+, significant shifts in these vibrational bands are noted at 1654, 1577, 3412, and 3647 cm−1, respectively. These changes indicate strong interactions between the metal ion and the donor sites of the ligand.
 | ||
| Fig. 12 FT-IR spectra of HP (left) and HP–Cu2+(right) complex calculated at the B3LYP/6-311++G(d,p) level of theory. | ||
Fig. 13 illustrates the graphical depiction of Mulliken atomic charges for the structures. The Mulliken data indicate that all hydrogen atoms possess positive charges, while the majority of carbon atoms exhibit negative charges.
 | ||
| Fig. 13 Mulliken atomic charges of HP and HP–Cu2+ obtained at the B3LYP/6 311++G(d,p) level of theory. | ||
| Compound | λ (nm) | E (eV) | fos | MO contributions (major contribution, <30%) |
|---|---|---|---|---|
| HP | 292.21 | 4.24 | 0.33 | HOMO > LUMO (78%) |
| 281.90 | 4.40 | 0.162 | H − 1 > LUMO (44%) | |
| 275.40 | 4.50 | 0.0248 | HOMO > L + 1(38%), HOMO > L + 2 (42%) | |
| HP–Cu2+ | 2389.34 | 0.518 | 0.0048 | HOMO(B) > LUMO(B) (98%) |
| 1262.17 | 0.982 | 0.0196 | H − 2(B) > LUMO(B) (36%), H − 1(B) > LUMO(B) (60%) | |
| 943.62 | 1.31 | 0.1052 | H − 2(B) > LUMO(B) (58%), H − 1(B) > LUMO(B) (36%) |
Table 3 summarizes the quantitative descriptors derived from Multiwfn analysis, which include Sr (spatial overlap integral), D (centroid distance between hole and electron), H (combined spatial extent of hole and electron), t (separation threshold), Δσ (difference in spatial spread), HDI (hole delocalization index), and EDI (electron delocalization index).
| Descriptor | HP | HP–Cu2+ complex |
|---|---|---|
| Excitation | S0 → S1 | S0 → S1 |
| E (eV) | 4.241 | 0.519 |
| Sr index (a.u.) | 0.64146 | 0.32662 |
| D index (Å) | 1.039 | 3.892 |
| H Index (Å) | 3.371 | 2.434 |
| t index (Å) | −1.717 | 2.188 |
| HDI | 8.61 | 20.20 |
| EDI | 6.68 | 10.52 |
| ECoul (eV) | 4.152 | 3.686 |
These metrics jointly facilitate the classification of electronic excitation characteristics. Characteristic intramolecular charge transfer (ICT) typically correlates with low Sr values (<0.6), substantial D values (>4.0 Å), and prominent H (>6.5 Å) or t (>0.3 Å) values, indicating considerable spatial separation. Conversely, local excitation (LE) transitions are generally characterized by elevated Sr (>0.6) and diminished D and H values, signifying localized excitation. Our work reveals that the S0 → S1 transition of the free ligand (HP) exhibits characteristic LE behavior, with Sr = 0.64, D = 1.039 Å, H = 3.371 Å, and t = −1.717 Å. The Cu2+ combination (HP–Cu2+) exhibits distinct ICT properties, indicated by Sr = 0.32, D = 3.892 Å, H = 2.434 Å, and t = 2.188 Å. These data validate a shift from a locally excited state in the unbound ligand to a charge transfer state following coordination with Cu2+.
The stability of the complex was assessed by NBO analysis by analyzing electron transfer from bonding and lone pair orbitals to antibonding orbitals (Table 4). The most significant intramolecular charge transfer (ICT) was found for the π(C19–C23) → LP*(σ) C23 interaction, with a stabilization energy of 26.51 kcal mol−1. Notable π → π* interactions comprise π(C16–C17) → π*(N14–C15) at 20.63 kcal mol−1, and π(C2–C3) → π*(C10–O12) with a stabilization energy of 17.00 kcal mol−1. The lone pair orbital LP(σ) on C19 significantly donates to the π*(C16–C17) acceptor, exhibiting a substantial stabilization energy of 75.68 kcal mol−1, which signifies considerable electron delocalization.
| No. | Donor NBO(I) | Acceptor NBO (j) | E(2) kcal mol−1a | E(j) − E(i) a.ub | F(i,j) a.uc |
|---|---|---|---|---|---|
| a E(2) means energy of hyperconjugative interactions (stabilization energy).b Energy difference between donor and acceptor i and j NBO orbitals.c F(i, j) is the Fock matrix element between i and j NBO orbitals. | |||||
| 1 | BD (π) C2–C3 | LP* (σ) C1 | 20.96 | 0.18 | 0.090 |
| 2 | BD (π) C2–C3 | BD* (π) C10–O12 | 17.00 | 0.20 | 0.074 |
| 3 | BD (π) C4–N13 | LP (σ) C5 | 15.50 | 0.18 | 0.082 |
| 4 | BD (π) C4–N13 | BD* (π) C2–C3 | 12.21 | 0.29 | 0.078 |
| 5 | BD (σ) C15–H30 | BD* (σ) N11–N14 | 5.59 | 0.86 | 0.088 |
| 6 | BD (π) C16–C17 | BD* (π) N14–C15 | 20.63 | 0.24 | 0.090 |
| 7 | BD (π) C16–C17 | BD* (π) C18–C21 | 9.68 | 0.32 | 0.073 |
| 8 | BD (π) C18–C21 | LP* (σ) C23 | 26.51 | 0.13 | 0.090 |
| 9 | BD (π) C18–C21 | BD* (π) C16–C17 | 7.17 | 0.25 | 0.057 |
| 10 | LP* (σ) C1 | BD* (π) C2–C3 | 45.09 | 0.10 | 0.108 |
| 11 | LP (σ) C5 | BD* (π) C4–N13 | 40.29 | 0.12 | 0.111 |
| 12 | LP (σ) N11 | BD* (π) C10–O12 | 21.70 | 0.30 | 0.104 |
| 13 | LP (σ) N11 | BD* (π) N14–C15 | 7.34 | 0.29 | 0.059 |
| 14 | LP (π) O12 | BD* (σ) C3–C10 | 7.28 | 0.82 | 0.099 |
| 15 | LP (π) O12 | BD* (σ) C10–N11 | 7.35 | 0.67 | 0.089 |
| 16 | LP (σ) N13 | BD* (σ) C2–C3 | 5.80 | 0.82 | 0.088 |
| 17 | LP (σ) N13 | BD* (σ) C4–C5 | 5.09 | 0.84 | 0.083 |
| 18 | LP (σ) C19 | BD* (π) C16–C17 | 75.68 | 0.10 | 0.124 |
| 19 | LP* (σ) C23 | BD* (π) C18–C21 | 20.87 | 0.16 | 0.099 |
| 20 | LP (π) O26 | LP* (σ) C23 | 36.79 | 0.18 | 0.127 |
| 21 | BD* (π) C10–O12 | BD* (π) C2–C3 | 16.06 | 0.08 | 0.074 |
| 22 | BD* (π) N14–C15 | BD* (π) C16–C17 | 33.83 | 0.04 | 0.069 |
| 23 | BD* (π) C16–C17 | BD* (π) C18–C21 | 23.31 | 0.05 | 0.070 |
| 24 | LP (σ) O12 | LP* (d) Cu28 | 10.01 | 0.76 | 0.111 |
| 25 | LP (π) O12 | LP* (d) Cu28 | 8.71 | 0.48 | 0.083 |
Additional significant interactions comprise LP(σ) C5 → π*(C4–N13) at 15.50 kcal mol−1 and LP(σ) N11 → π*(C10–O12) at 21.70 kcal mol−1, hence affirming the participation of heteroatoms in electron donation. The σ(C15–H30) → σ*(N11–N14) interaction provides a moderate stabilization energy of 5.59 kcal mol−1, indicative of hyperconjugation effects.
Metal coordination was validated by the donation of LP(O12) to LP*(Cu28), with energies of 10.01 and 8.71 kcal mol−1 for the two lone pair orbitals on oxygen engaging with the copper antibonding orbitals, indicating coordination bonding. Additional LP(π) interactions, such as LP(O26) → LP*(C23) (36.79 kcal mol−1), substantially enhance the overall stability.
The donor–acceptor interactions collectively indicate that electron delocalization via π and σ bonding frameworks, facilitated by lone pair electron donation and metal coordination, plays an important role for stabilizing the complex.
3.9 Application of chemosensor HP in real samples
Laboratory tap water samples were used to assess the practical usability of the HP probe for Cu2+ detection. The innate Cu2+ concentration in tap water was below the method's assessing limit; therefore, artificial Cu2+-contaminated tap water samples were made by spiking with known amounts of 10 µM and 20 µM of standard Cu2+ solutions. The spiked samples were subsequently evaluated using the same exact UV-Vis spectroscopic technique. The concentrations of Cu2+ were determined using the calibration curve (absorbance versus concentration) depicted in Fig. S5. The recovery values achieved varied from 94.10% to 96.43% (Table 5), demonstrating the method's high accuracy. The results indicate that HP demonstrates significant potential for the quantitative detection of Cu2+ ions in actual water samples, confirming its relevance in environmental monitoring.| Metal ion | Spiked amount (µM) | Recovered amount (µM) | Recovery % |
|---|---|---|---|
| Cu2+ | 10 | 9.41 | 94.10 |
| 20 | 19.28 | 96.43 |
3.10 Comparison of HP with other Schiff base chemosensors
The efficacy of the Schiff base probe HP in detecting Cu2+ was evaluated in comparison to several other stated Schiff base chemosensors, as detailed in Table 6. Table 6 illustrates that the present system exhibits several advantageous analytical features compared to other systems, including high sensitivity, high selectivity, a lower detection limit, straightforward operational technology, good solubility, and practical applicability. The synthesis of the proposed chemosensor HP involves a single step, utilizes less hazardous reagents, and does not produce any hazardous by-products.| Probe | Solvent | LOD | Binding constant | M![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) L ratio L ratio |
Ref. |
|---|---|---|---|---|---|
 |
CH3CN-Tris (20 mM, v/v = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, pH = 7.2) 1, pH = 7.2) |
11.4 µM | 1.47 × 105 M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
40 |
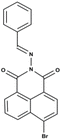 |
CH3CN | 12.5 µM | Not reported | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
31 |
 |
THF/H2O | 15.14 µM | 1.26 × 105 M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 2 |
41 |
 |
(DMSO/TrisHCl),1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1,v/v, buffer pH 7 1,v/v, buffer pH 7 |
19.7 µM | Not reported | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
42 |
 |
20% H2O/DMF | 15 µM | 2.42 × 108 M−2 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
43 |
 |
75%CH3CN 25% 0.01 M Tris–HCl buffer pH 7 | 69 µM | 1.65 × 103 M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
44 |
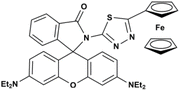 |
EtOH/HEPES (8![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, v/v, pH 7.2) 2, v/v, pH 7.2) |
650 µM | 1.73 × 104 M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
45 |
 |
MeOH/H2O (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) 1, v/v) |
10.67 µM | 2.69 × 104 M−1 | 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
46 |
 |
MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 v/v, HEPES = 10 mM, pH 7.4) 4 v/v, HEPES = 10 mM, pH 7.4) |
8.94 µM | 3.04 × 104 M−1 | 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
This work |
4. Conclusion
In summary, the compound HP was synthesized via a simple and efficient one-step methodology and characterized using FTIR, 1H NMR, 13C NMR, HRMS, SCXRD, and elemental analysis. HP functions as a chemodosimeter for Cu2+ ions, exhibiting distinct absorption changes in MeOH/H2O (6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4, v/v, HEPES = 10 mM, pH 7.4). Job's plot analysis indicated a 1
4, v/v, HEPES = 10 mM, pH 7.4). Job's plot analysis indicated a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry, while FTIR studies confirmed the complexation process with Cu2+ ions. FESEM imaging of the free ligand and its Cu2+ complex revealed pronounced morphological changes, further supporting successful coordination. The experimental findings are in excellent agreement with DFT and TD-DFT calculations. NBO analysis validated the stability of the HP–Cu2+ complex. The hole–electron analysis clearly reveals locally excited (LE) characteristics for the free ligand and intramolecular charge transfer (ICT) behavior for the HP–Cu2+ complex. The combination of cost-effective and readily available precursors, a high binding constant, and a lower limit of detection (LOD) compared to many reported Cu2+ chemosensors positions HP as a promising candidate for Cu2+ ion detection. Its practical applicability was demonstrated by the high sensitivity achieved in detecting trace concentrations of Cu2+ in real water samples.
1 binding stoichiometry, while FTIR studies confirmed the complexation process with Cu2+ ions. FESEM imaging of the free ligand and its Cu2+ complex revealed pronounced morphological changes, further supporting successful coordination. The experimental findings are in excellent agreement with DFT and TD-DFT calculations. NBO analysis validated the stability of the HP–Cu2+ complex. The hole–electron analysis clearly reveals locally excited (LE) characteristics for the free ligand and intramolecular charge transfer (ICT) behavior for the HP–Cu2+ complex. The combination of cost-effective and readily available precursors, a high binding constant, and a lower limit of detection (LOD) compared to many reported Cu2+ chemosensors positions HP as a promising candidate for Cu2+ ion detection. Its practical applicability was demonstrated by the high sensitivity achieved in detecting trace concentrations of Cu2+ in real water samples.
Author contributions statement
Joardar Gim: Performed the experiment and all computational studies, analyzed and interpreted the data, wrote the paper. Zannatul Kowser: conceptualized the whole experiment, supervised students, contributed materials, reagents, analysis tools, or data, analyzed and interpreted the data, wrote the paper, and revised the manuscript. Dipa Debnath: performed the experimental part. Redika Sarmin Prieety: performed the experimental part. Miss. Tasnim Jahan: performed the experimental part. Paul G. Waddell: single-crystal X-ray diffraction data collection, structure solution and refinement, crystallographic interpretation, manuscript writing, and critical revision. Most Tahera Khatun: performed the experimental part. Sahara Khatun Munni: performed the experimental part. Md. Rashed Khan: wrote the paper. Rashedul Islam: revised the manuscript.Conflicts of interest
The authors declare that they have no conflict of interest.Data availability
CCDC 2515488 contains the supplementary crystallographic data for this paper.47Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ra06131h.
Acknowledgements
Zannatul Kowser is thankful to Jashore University of Science and Technology Research Cell (JUSTRC), Jashore 7408, Bangladesh, under grant number 24-FoS-08 for the fiscal year 2024 to 2025 for providing financial support to carry out a part of the study. The authors gratefully acknowledge the Department of Chemistry, of Jashore University of Science and Technology (JUST) for providing data analysis support.References
- A. Taha, N. Farooq, N. Singh and A. A. Hashmi, J. Mol. Liq., 2024, 401, 124678 Search PubMed
.
- S.-H. Park, N. Kwon, J.-H. Lee, J. Yoon and I. Shin, Chem. Soc. Rev., 2020, 49, 143–179 RSC
.
- M. Prajapati, N. Pandey, S. Kalla, S. Bandaru and A. Sivaiah, Sens. Diagn., 2024, 3, 412–420 Search PubMed
.
- M. Valko, K. Jomova, C. J. Rhodes, K. Kuča and K. Musílek, Arch. Toxicol., 2016, 90, 1–37 Search PubMed
.
- L. Banci, I. Bertini, S. Ciofi-Baffoni, T. Kozyreva, K. Zovo and P. Palumaa, Nature, 2010, 465, 645–648 CrossRef CAS PubMed
.
- P. Matak, S. Zumerle, M. Mastrogiannaki, S. El Balkhi, S. Delga, J. R. R. Mathieu, F. Canonne-Hergaux, J. Poupon, P. A. Sharp, S. Vaulont and C. Peyssonnaux, PLoS One, 2013, 8, e59538 CrossRef CAS PubMed
.
- I. A. Koval, P. Gamez, C. Belle, K. Selmeczi and J. Reedijk, Chem. Soc. Rev., 2006, 35, 814 RSC
.
- E. L. Que, D. W. Domaille and C. J. Chang, Chem. Rev., 2008, 108, 1517–1549 Search PubMed
.
- S. Poormoradkhan Melal, B. Khalili, N. O. Mahmoodi and M. Pasandideh Nadamani, J Photochem Photobiol A Chem, 2025, 460, 116140 Search PubMed
.
- Z. K. Mathys and A. R. White, Adv. Neurobiol., 2017, 18, 199–216 Search PubMed
.
- A. Lucena-Valera, P. Ruz-Zafra and J. Ampuero, Med. Clin., 2023, 160, 261–267 Search PubMed
.
- J. Chen, Y. Jiang, H. Shi, Y. Peng, X. Fan and C. Li, Pflugers Arch, 2020, 472, 1415–1429 Search PubMed
.
- Q. Y. Chen, P. Wu, T. Wen, X. Qin, R. Zhang, R. Jia, J. Jin, F. Hu, X. Xie and J. Dang, Front. Aging Neurosci., 2022, 17(14), 970711 CrossRef PubMed
.
- L. McAlary, V. K. Shephard, G. S. A. Wright and J. J. Yerbury, J. Biol. Chem., 2022, 298, 101612 Search PubMed
.
- G. Mohammadi Ziarani, N. Shamkhali, Z. Panahande, M. Feizi-Dehnayebi and A. Badiei, Results Chem, 2025, 16, 102381 Search PubMed
.
- B. Thangaraj, M. Ponram, S. Ranganathan, B. Sambath, R. Cingaram, S. K. Iyer and K. Natesan Sundaramurthy, RSC Adv., 2023, 13, 26023–26030 RSC
.
- P. Deng, Y. Pei, M. Liu, W. Song, M. Wang, F. Wang, C. Wu and L. Xu, RSC Adv., 2021, 11, 7610–7620 RSC
.
- C. Lawrence, G. E. Sanders and C. Wilson, in Management of Animal Care and Use Programs in Research, Education, and Testing, CRC Press, 2nd edn, Boca Raton, Taylor & Francis, 2018, pp. 559–578 Search PubMed
.
- National Research Council (US) Committee, Copper in Drinking Water, National Academies Press (US), Washington (DC), 2000 Search PubMed.
- R. Kumar, B. Singh, P. Gahlyan, A. Verma, M. Bhandari, R. Kakkar and B. Pani, RSC Adv., 2024, 14, 23083–23094 RSC
.
- W.-Y. Zhu, K. Liu and X. Zhang, Sens. Diagn., 2023, 2, 665–675 RSC
.
- J. Pizarro, E. Flores, V. Jimenez, T. Maldonado, C. Saitz, A. Vega, F. Godoy and R. Segura, Sens Actuators B Chem, 2019, 281, 115–122 CrossRef CAS
.
- N. Li, D. Zhang, Q. Zhang, Y. Lu, J. Jiang, G. L. Liu and Q. Liu, Sens Actuators B Chem, 2016, 231, 349–356 CrossRef CAS
.
- L. Lago, O. R. B. Thomas and B. R. Roberts, J Chromatogr A, 2020, 1616, 460806 CrossRef CAS PubMed
.
- J. Wang, Y. Gao, Y. Zhang, D. Xu, Z. Li, Y. Su, F. Wang, N. Zhang, D. Zhang, K. Wang and Y. Chen, New J. Chem., 2025, 49, 9133–9144 RSC
.
- R. C. Clark and J. S. Reid, Acta Crystallogr A, 1995, 51, 887–897 CrossRef
.
- CrysAlisPro, Rigaku Oxford Diffraction, Tokyo, Japan Search PubMed
.
- G. M. Sheldrick, Acta Crystallogr., Sect. A:Found. Adv., 2015, 71, 3–8 CrossRef PubMed
.
- G. M. Sheldrick, Acta Crystallogr A, 2008, 64, 112–122 CrossRef CAS PubMed
.
- O. V. Dolomanov, L. J. Bourhis, R. J. Gildea, J. A. K. Howard and H. Puschmann, J. Appl. Crystallogr., 2009, 42, 339–341 CrossRef CAS
.
- G. Kaur, G. Kumar and I. Singh, J. Mol. Struct., 2025, 1319, 139252 CrossRef CAS
.
- P. Seenu and S. K. Iyer, Sens. Diagn., 2025, 4, 973–983 RSC
.
- R. Gajendhiran, A. K. R. Ahmed, S. Mithra, S. A. Majeed, A. S. S. Hameed, S. Jegadeeshwari, K. Muthu, M. NizamMohideen and A. K. Rahiman, Anal. Methods, 2025, 17, 7920–7935 RSC
.
- S. Sarkar, A. Chatterjee, K. Ghosh, N. N. Ghosh, A. Dutta, A. Kumar, A. K. Mandal and K. Biswas, New J. Chem., 2025, 49, 10991–11000 RSC
.
- S. Schoeman, N. Mama and L. Myburgh, New J. Chem., 2025, 49, 1745–1754 RSC
.
- B. Zhang, D. Zeng, Y.-X. Zhang, P. Pan, J. Wang, A. Shen, J.-X. Lu, Y.-J. Zhu, A.-P. Xing and J. Yuan, Spectrochim Acta A Mol Biomol Spectrosc, 2025, 340, 126310 Search PubMed
.
- T. Saleem, S. Khan, M. Yaqub, M. Khalid, M. Islam, M. Yousaf ur Rehman, M. Rashid, I. Shafiq, A. A. C. Braga, A. Syed, A. H. Bahkali, J. F. Trant and Z. Shafiq, New J. Chem., 2022, 46, 18233–18243 Search PubMed
.
- V. Pophristic, L. Goodman and N. Guchhait, J. Phys. Chem. A, 1997, 101, 4290–4297 Search PubMed
.
- F. Weinhold, Nature, 2001, 411, 539–541 CrossRef CAS PubMed
.
- X. Liu, P. Xu, X. Zhao, J. Ge, C. Huang, W. Zhu, C. Li, L. Du and M. Fang, Inorganica Chim Acta, 2019, 495, 118975 CrossRef CAS
.
- N. Saini, N. Prigyai, C. Wannasiri, V. Ervithayasuporn and S. Kiatkamjornwong, J Photochem Photobiol A Chem, 2018, 358, 215–225 CrossRef CAS
.
- E. Normaya, N. A. Baharu and M. N. Ahmad, J. Mol. Struct., 2020, 1212, 128094 CrossRef CAS
.
- A. Dhawale and D. R. Trivedi, ACS Omega, 2025, 10, 9527–9536 CrossRef CAS PubMed
.
- A. J. Weerasinghe, F. A. Abebe and E. Sinn, Tetrahedron Lett., 2011, 52, 5648–5651 CrossRef CAS
.
- H. Ye, F. Ge, X.-C. Chen, Y. Li, H. Zhang, B.-X. Zhao and J.-Y. Miao, Sens Actuators B Chem, 2013, 182, 273–279 CrossRef CAS
.
- A. K. Manna, J. Mondal, R. Chandra, K. Rout and G. K. Patra, J Photochem Photobiol A Chem, 2018, 356, 477–488 CrossRef CAS
.
- CCDC 2515488: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qfksj
.
| This journal is © The Royal Society of Chemistry 2026 |







