Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Are we there yet with XNA aptamers?
Santiago Chaillou†
 ,
Saphyra Thonon†
,
Saphyra Thonon† ,
Sten Reynders
,
Sten Reynders ,
Eveline Lescrinier
,
Eveline Lescrinier and
Vitor B. Pinheiro
and
Vitor B. Pinheiro *
*
KU Leuven, Rega Institute for Medical Research, Department of Pharmaceutical and Pharmacological Sciences, Herestraat, 49, Leuven 3000, Belgium. E-mail: v.pinheiro@kuleuven.be
First published on 29th April 2026
Abstract
Since their original development, aptamers (or nucleic acid oligomers of defined sequence that are capable of high-affinity and high-specificity binding to a target) have been heralded as possible and promising new therapeutic modalities. Increasingly versatile and customizable aptamer selection (SELEX) processes, alongside incorporation of novel synthetic nucleic acid (XNA) chemistries, have played a key role in the development of therapeutically relevant molecules. Nonetheless, despite the significant progress in the last 40 years, clinical applications of aptamers remain a nascent field. This review summarises some of the key developments in the field in the last 40 years, highlighting the progress made in aptamer selection, XNA chemistries and computational analyses. We discuss the intricacies and limitations of the current gold standard in aptamer SELEX and underscore the added complexities of XNA incorporation. Reflecting on the distinction between the antibody and aptamer fields, we advocate how a more mature understanding of this class of molecules could be driving the next generation of applications.
Introduction
Aptamers are often defined as short single-stranded nucleic acids (e.g. DNA or RNA) that adopt stable three-dimensional structures capable of high-affinity and high-specificity binding to a target, which can include small organic compounds, peptides, proteins, other nucleic acids,1,2 and even whole cells.3 They are generally selected in vitro through an iterative process called systematic evolution of ligands by exponential enrichment (SELEX)4 or variations thereof.Like antibodies, aptamers are highly versatile molecules that enable a wide range of applications including diagnostics (e.g. facilitating rapid and sensitive detection of disease markers, pathogens, and toxins5), biosensors (e.g. monitor dynamic changes in analyte concentrations, providing real-time information in clinical and environmental settings6) and therapeutics (e.g. Macugen and Izervay targeting neovascular wet age-related macular degeneration and geographic atrophy secondary to age-related macular degeneration, respectively7,8).
Nonetheless, the narrow chemical and biological stability of natural DNA and RNA represents a significant development barrier for aptamer applications, and a major one for therapeutic aptamer development. It can also be argued that the small chemical diversity among natural nucleobases and the structural rigidity of the (deoxy)ribose and phosphate backbones may further limit the potential structures (and therefore functions) that can be isolated.
Chemical modification of nucleic acids is possible and widely used,9 as universally accepted in the field of therapeutic antisense oligonucleotides (ASOs). While some of the groundwork for nucleotide modifications dates back to the 1960's10–12 – including the discovery of 2′-fluoro-(2′-F), 2′-O-methyl-(2′-OMe) and phosphate-backbone modifications – the first generation oligonucleotide modifications appeared decades later with therapeutic ASOs to equip them with inherent nuclease resistance: in 1986, the antiviral effect of a methylphosphonate oligonucleotide was reported,13 swiftly followed by the more successfully adopted and widely studied phosphorothioate (PS) modification in 1987.14 The progress made in the antisense oligonucleotide field allowed for the rapid introduction of chemical modifications to aptamers after their emergence in the 1990s,4,5 with the earliest publications appearing in 1994, incorporating 5-(1-pentynyl)-2′-deoxyuridine modifications,15 and in 1995, applying 2′-amino-2′-deoxypyrimidine and 2′-OMe modifications.16,17 Historically, 2′-F and 2′-OMe modifications are amongst the most popular aptamer chemical modifications, exemplified by Macugen and Izervay (relying on those 2′-sugar modifications along with conjugation with biocompatible polymers – e.g. PEG or cholesterol and 3′-end capping).
Whereas those initial modifications are still widely used in therapeutic oligonucleotides, the available toolkit has grown rapidly to include chemical modifications of nucleobase, sugar, phosphate and combinations thereof. Modifications can increase nuclease resistance, such as phosphorothioates18 and HNA,19 or introduce novel chemical moieties that alter the chemical properties of the molecule, such as the neutral backbone of phNA20 and PNA,21 or the hydrophobic moieties used in SOMAmers.22 Almost invariably, they also introduce changes to the sequence-function landscape that make mapping between landscapes impossible – that is, a bona fide aptamer in one nucleic acid chemistry does not necessarily remain an aptamer once the chemistry in changed.
The term ‘XNA’ (xenobiotic nucleic acids, also known as synthetic nucleic acids) was coined in literature in 2009 referring to sugar modifications.23 However, since then, XNA has been used more broadly for chemically modified nucleic acids for any sugar and/or backbone modification.24 In 2019, a new classification system was proposed, using XNA, DZA and DNY according to the position of the modification.24 While this distinction is important to consider (because of their different chemical and structural impacts), for the purpose of this review, we discuss XNA in its broadest term, concentrating on backbone and nucleobase modifications most relevant for aptamer development. Chemical structures (and the position of modification compared to the natural nucleotides) of the XNA chemistries discussed in this review are depicted in Fig. 1.
Selection of aptamers directly in XNA is increasingly possible, but development lags compared to RNA and DNA. Here, we review critically the current state of the field of XNA aptamers and highlight some of the persistent bottlenecks that affect not only XNA aptamers but the aptamer field in general. The aptamer field is extensive, and we apologize to our colleagues whose work we cannot include in this brief review. Other excellent reviews on aptamers,3,25–29 chemical modification of nucleic acids and XNAs,9,30–32 and in silico tools for aptamer development33–36 are available elsewhere.
A brief history of aptamers
The 1990's – first examples
The development of SELEX is a watershed moment for nucleic acid aptamers, but it did not emerge in a vacuum. As highlighted by Joyce in 1989 (ref. 37), all tools required to carry out the directed evolution of nucleic acids in vitro were available. Less than a year later, two groups published groundbreaking results reporting the first aptamers – Tuerk and Gold isolating the first RNA aptamer against the T4 DNA polymerase,4 and Ellington and Szostak isolating ligands against small molecules.5The first decade in the aptamer field laid a solid foundation for subsequent decades of aptamer research (Fig. 2). Initially, the field quickly diversified its targets through the isolation of aptamers selected against various proteins, such as the thrombin-binding aptamer,38 and small molecules like ATP.39 Concurrently, significant technical advancements in SELEX methodologies further propelled this growth. By 1995, researchers had already isolated 49 aptamers, demonstrating a wide range of affinities spanning from 30 pM to 12 mM.40 Early research on aptamers also included enhancing aptamer properties through increasing their chemical diversity. Notably, some of these aptamers already incorporated chemically modified nucleosides like 2′-F, 2′-NH2,40 2′-OMe17 and PS,41 and mirror images of RNA molecules had been developed.42,43
Elucidating the structures of aptamers was of interest for the field from its very beginning. By 1997, several aptamer structures had been elucidated through NMR spectroscopy, such as the RNA aptamer complexes binding adenosine monophosphate, flavin mononucleotide, arginine/citrulline, and tobramycin.44
Reaching the end of the decade, the first applications of aptamer technology emerged with aptamers being integrated into molecular diagnostic arrays.45
The 2000's – focus on selection
The field of aptamer technology expanded significantly in the new millennium, driven by theoretical advancements and the proposal of aptamers as therapeutic and diagnostic agents.46 Methodologically, the early 2000s saw substantial progress with the integration of pivotal SELEX technologies such as magnetic beads47 and fluorescently labelled nucleic acid strands.48 Innovations like capillary electrophoresis-SELEX (CE-SELEX),49 FACS-SELEX using cell sorting,50 and Microfluidics SELEX (M-SELEX)51 integrated with micromagnetic principles expanded the capabilities of aptamer selection methods. The era also witnessed the advent of the first automated aptamer discovery and the introduction of non-SELEX methodologies.52Advancements in aptamer selection methodologies enabled researchers to develop aptamers for new classes of targets, including whole cells and RNA. Cell-SELEX was successfully demonstrated through the isolation of aptamers against pathogenic bacteria (e.g. Mycobacterium tuberculosis53), cancer cells (e.g. liver cancer-specific aptamers54) and viral RNA domains (e.g. aptamers against Hepatitis C virus55). Moreover, the integration of Cell-SELEX with other novel selection methodologies facilitated the development of in vivo SELEX.56
The growth in aptamer technology paved the way for novel practical applications. The development of ‘escort aptamers’ enabled their use in in vivo diagnostics and treatments, marking a first contribution of aptamers to precision medicine.57 Additionally, aptamer beacons allowed direct, fast and sensitive protein detection.58 Beyond diagnostics, natural aptamers were also identified and adopted for genetic regulation through riboswitches, modulating gene expression by binding metabolites.59 The introduction of aptamer biosensors marked the first step in a leading application in the field.60
Chemical diversity in aptamer design flourished during this decade with the development of novel chemistries such as boron-containing aptamers,61 hexitol nucleic acid-containing aptamers (HNA),62 4′-thioRNA,63 locked nucleic acids (LNA),64 2′-fluoro-D-arabino-RNA (FANA),65 cationic-charged modified DNA aptamers (e.g. THM),66 and the first combinations of modified chemistries, such as LNA with 2′-OMe.67 These advancements culminated in a pivotal moment with the FDA approval of Macugen in 2004 for the treatment of neovascular (wet) age-related macular degeneration,68 validating aptamers as viable therapeutic agents (while also establishing viable manufacturing and formulation strategies for aptamers) and highlighting their potential in biomedical research.
The 2000s also witnessed the emergence of computational tools essential for advancing aptamer research. Tools like Mfold69 and ViennaRNA70 provided the first predictions of RNA secondary structures critical for a better understanding of aptamer folding and stability. QGRS Mapper facilitated the identification of G-quadruplex structures within aptamers, enhancing their structural characterization.71 The era also saw the first pipeline for the prediction and visualization of 3D aptamer structures with 3D-DART.72 Moreover, during these years the first version of the Galaxy Platform was developed.73
Rigid docking methods employed by tools like ZDOCK74 and GRAMM-X75 facilitated exploration of aptamer–target binding surfaces, while flexible docking approaches emerged with tools like Rosetta, originally developed for proteins in 2004 (ref. 76) and later applied to nucleic acids in 2007.77 Molecular dynamics simulations with AMBER78 and GROMACS 4 (ref. 79) enabled researchers to simulate the dynamic behaviour of aptamer–target complexes, offering deeper insights into their stability and functional mechanisms. These computational tools laid the foundation for future in silico tools.
The 2010's – integrating new technologies
In the following decade, additional SELEX methodologies were developed, now focused on overcoming previously described limitations of the process. Techniques like graphene oxide-assisted SELEX (GO-SELEX, utilizing graphene oxide for non-specific adsorption of ssDNA)80 and Capture SELEX (employing nucleic acid library immobilization on magnetic beads),81 emerged to bypass target immobilization prior to selection. The integration of Next-Generation Sequencing (NGS) in SELEX methodologies represented a paradigm shift, offering an alternative to traditional Sanger sequencing.82 This innovation led to the development of analytical tools such as AptaPLEX,83 AptaMotif,84 AptaCluster,85 AptaMut,86 and AptaTRACE,87 culminating in their combination into the AptaSUITE platform.88 AptaSUITE enables researchers to demultiplex NGS data, identify sequence motifs, cluster sequences based on locality-sensitive hashing (LSH), predict mutant affinity, and analyse structural enrichment of k-mers, becoming a cornerstone in contemporary aptamer NGS sequencing analysis.Additional software tools for NGS data analysis emerged during this period, including FASTAptamer,89 facilitating sequence clustering based on Levenshtein edit distances, and APTANI,90 which leverages structural enrichment of k-mers and secondary structure information for aptamer identification. The update of Galaxy in 2016 provided an improved versatile web server platform for comprehensive nucleic-acid sequence analysis.91
Tools for predicting 2D structures, such as ViennaRNA package 2.0,92 NUPACK,93 and RNAstructure,94 were complemented by specialized software for G-quadruplex prediction like G4RNA95 and G4hunter.96 In 3D structure modelling, tools like RNAComposer,97 SimRNA,98 and 3D-Nus99 enhanced predictions despite their limitations in accuracy.
Advancements in predicting aptamer–target binding surfaces were achieved through tools like Hexserver100 for rigid docking and HADDOCK,101 AutoDock,102 and AutoDock Vina103 for flexible docking. Molecular dynamics simulations using CHARMM104 and AMOEBA105 were first used to provide insights into the dynamic behaviour of aptamer–target complexes. Machine learning applications also gained traction, exemplified by Apta-LoopEnc, which uses Support Vector Machines (SVM) to predict aptamer sequences based on their binding affinity using SELEX data from high and low affinity cycles.106
The 2010s also witnessed a significant growth in combining aptamers with diverse technologies. Innovations included bispecific aptamers,107 aptamer–drug conjugates108 for targeted drug delivery, integration of aptamers with DNA origami109 for enhanced structural interactions, and aptamer-functionalized lipopolymer delivery systems110 facilitating CRISPR-Cas9 plasmid transport for gene editing. Furthermore, advancements in aptamer display techniques were achieved through DNA nanostructure-decorated surfaces.111
The period witnessed new advancements in the chemical expansion of aptamers, such as the introduction of threose nucleic acids (TNA),112 and the incorporation of a genetically expanded alphabet using Ds,113 P and Z114 nucleobases. Additionally, there were innovations such as functionalized DNA structures like Clickmers115 and DNA-scaffolded peptides.116 Combinations of chemistries, such as TNA with 7-deaza-7-phenyl modified guanosines117 and 2′-OMe-DNA with HNA, were explored further.118
Significant milestones during this period include the development of the first aptamer with an uncharged backbone (phNA)20 and the creation of Slow Off-rate Modified Aptamers (SOMAmer),119 which later paved the way for the SOMAscan proteomics platform.22
Throughout the decade, clinical trials explored aptamer therapeutic potential across diverse medical conditions like macular degeneration, cancer, and diabetes. Some of the candidates did not progress past phase I or II, such as RB006, ARC1779, NU172, and ARC19499, aimed at addressing hemostasis; AS14, in the field of cancer treatment; and E10030 (Fovista™), for macular degeneration. Notably, other aptamers showed promise, such as NOX-A12 for cancer treatment, prompting TME Pharma to seek FDA Fast-Track Designation in February 2024. Similarly, NOX-E36 completed successful phase I and II trials for diabetes mellitus, indicating potential for further development as a treatment option.120
The 2020's – possible watershed moment
In the current decade, computational tools aimed at advancing aptamer research have proliferated. These tools represent a shift from single-criterion strategies to multi-dimensional scoring methods, exemplified by SMART-Aptamer,121 RaptRanker,122 and APTANI2,123 offering more nuanced perspectives on aptamer sequence analysis.Deep learning has propelled significant advancements in computational biology. Tools like AlphaFold 3 (ref. 124) and RoseTTAFold2NA125 showcase this progress, utilizing deep neural networks to predict protein and RNA structures with unprecedented accuracy. Similarly, innovations such as Alchemy_RNA2126 and Ares127 apply machine learning techniques to predict nucleic acid structures, leveraging evolutionary and biophysical data for improved insights.
This technological evolution parallels advancements in other areas of computational biology, such as AptaNet's aptamer–protein interactions prediction using machine learning,128 and generative aptamer design facilitated by Raptgen.129 Furthermore, the last few years also saw a milestone in aptamer discovery with the introduction of the first aptamer found using a machine learning-guided refinement and discovery protocol.130
Regarding novel aptamer chemistries, the field presented the integration of cubane moieties into aptamer structures131 – arguably combining the resulting improved stability with unique hydrogen bonding capabilities.
Clinical trials in the 2020s have underscored the diagnostic and therapeutic potential of aptamers across various applications. Diagnostic aptamers include the Saliva-based COVID-19 DNA Aptamer Test (NCT04974203) for rapid COVID-19 diagnosis, the Tenofovir (TFV) Aptamer-Based Biosensor (NCT04870671) for monitoring AIDS treatment adherence, molecular biosensors for the detection of bladder cancer (NCT02957370), and a labelled ssDNA aptamer for colorectal cancer diagnosis (NCT03385148).120 Therapeutic trials feature EYE001 (NCT00239928) for macular degeneration, NOX-H94 (NCT02079896) for anaemia of chronic disease, BT200 (NCT04677803) for hereditary bleeding disorders, and 68GA-Sgc8 (NCT03385148) against colorectal cancer. Notably, Izervay received FDA approval in 2023 for the treatment of geographic atrophy secondary to macular degeneration, marking a significant milestone in aptamer-based therapeutics.120
In 2022, the field achieved a pivotal standardization milestone with the publication of minimum aptamer publication standards,132 building upon initial suggestions from 2009.133 However, despite these advancements and the field's gain in popularity (Fig. 3), the literature still lacks comprehensive theoretical frameworks beyond traditional SELEX methodologies, highlighting an ongoing gap in aptamer theory despite the prolific practical and clinical advancements.
 | ||
| Fig. 3 Publication analysis by year of publications containing the words “aptamer” or “aptamers” until December 31st, 2025 using the Web of Science (WoS) (https://mjl.clarivate.com/). | ||
Limits despite success
Since its development, SELEX and its many variations have remained the gold standard in the field for the isolation of the highest affinity binders. Starting from a diverse library of random sequences, SELEX typically follows iterative cycles that can broadly be divided into four phases: (1) (re)generation of the library, (2) incubation with the target, (3) partitioning of binders from non-binders and (4) amplification of the retained sequences. The fundamentals of this selection process can be found in Fig. 4. Traditionally, stringency is increased between selection cycles – e.g. decreased target concentration, decreased incubation time or more elaborate washing steps – allowing the high-affinity binders within the library to outcompete the weaker ones. Once sufficient enrichment of high-affinity binders is observed, the remaining sequences are analyzed, and several aptamer-candidates are chosen for further characterization. In addition to affinity and specificity measurements, characterization can include stability tests (e.g. serum nuclease degradation134) and functionality descriptions (e.g. quantification of inhibitory effect135), with the goal of finding aptamer sequence(s) that satisfy the requirements of the selection. In a final stage, aptamers can also be fine-tuned, through truncations, chemical modifications, or conjugations.28 | ||
| Fig. 4 Overview of the aptamer selection process. SELEX consists of iterative rounds of library (re)generation, incubation with the target, partitioning and amplification of the retained sequences. However, depending on the chemistry used, the detailed SELEX process will look different. Library generation: the first step in the aptamer selection process is the library design and subsequent library generation. In most cases, a DNA library is first generated. For RNA or XNA chemistries, the DNA library is transcribed to either of them. For some XNA chemistries libraries are generated through direct chemical synthesis. Detailed reviews on considerations when designing your library can be found elsewhere.136–139 Main SELEX protocol: while the incubation with the target and partitioning steps are comparable between chemistries, subsequent amplification of binders and library regeneration require additional steps for RNA and XNA. To make amplifications possible, sequences need to be reverse transcribed into cDNA. After amplification these sequences are then transcribed back to the original chemistry. For RNA, established polymerases such as T7 RNA polymerase are used. For XNA, (reverse) transcriptases that can process these chemistries need to be available, incurring technological limitations. In some cases, i.e. with phosphorothioate, XNA can be directly amplified from the eluted sequences, resulting in shorter selection cycles. Various strategies to obtain ssXNA after PCR amplification exist but is here represented by the strategy utilizing magnetic beads and elution by denaturation. A common notable alternative to the depicted magnetic beads-based SELEX is Capture SELEX,140 in which the aptamer library is immobilized on magnetic beads instead of the target. Screening and deep sequencing: after the last selection round, the enriched sequences are determined and analysed using custom scripts, or through available platforms such as The Galaxy Project for sequence data handling, or AptaSUITE141 for further computational analyses. | ||
Research into aptamers has predominantly focused on addressing the inherent challenges associated with the selection process, i.e. SELEX modifications to optimize and streamline the SELEX protocol. Those SELEX modifications can encompass counter-selection steps (where similar targets are used in a negative selection round), alternative library designs (such as pre-structured libraries), or improved selection conditions, e.g. adaptations based on the intended application, promoting selection of the high-affinity binders or the addition of competitors. While those optimization strategies have increased the success rate in obtaining high-affinity aptamers against a diverse range of targets, SELEX still encounters both experimental challenges and theoretical limitations.142
On the practical front, SELEX remains a time- and labour-intensive procedure. Despite the emergence of innovative strategies aimed at enhancing or complementing SELEX, selection strategies continue (mostly) to function as black boxes, with few tools to monitor the progression of the selection campaign.143 In addition, the field lacks predictive tools to assess the likelihood of selection success before its execution.
When it comes to data analysis, the disclosure of analysis pipelines, software, or codes is infrequent. Moreover, there is a notable scarcity of detailed criteria for sequence selection and limited information on the number of sequences tested before arriving at a successful outcome. The field exhibits a significant survivor bias, and the absence of a repository for ‘difficult’ targets or unsuccessful selection attempts perpetuates a cycle of repeated mistakes among scientists across various laboratories. Nonetheless, in recent years, the field has become more aware of and concerned with the need for standardization, as exemplified by the publication of the minimum aptamer publication standards.132
Nonetheless, we believe that the field needs to go further, establishing a formal (and quantitatively robust) definition for aptamers. The current definition, which is qualitative, is ambiguous and leaves too much dependent on the context of the characterization, which often goes unreported. For instance, the reporting of a DNA sequence identical to a previously reported RNA aptamer against hen egg lysozyme:144 despite high affinities, compatible with aptamer definitions, the natural affinity of nucleic acids for a positively charged lysozyme protein, through electrostatic interactions, was not considered.139 In fact, much of the declined trust in the aptamer field stems, in our opinion, from the lack of guidelines and standards, mirroring the challenges faced by therapeutic antibodies due to overlooked specificity concerns.145–147
The problems caused by the lack of robust definition are exacerbated when chemical modifications are introduced or if different nucleic acid chemistries (i.e. XNAs) are used for aptamer development. In those cases, affinity alone is not enough to describe an aptamer, and specificity is not always clearly defined. Post-SELEX replacement of nucleic acid moieties with chemical analogues often compromises aptamer function63,148,149 but direct selection in XNA incurs greater technical challenges than those of SELEX alone.
XNA aptamer selection
The complexity of the selection process increases when chemistries other than the traditional ones are used. XNA modifications can be introduced to aptamers either post-SELEX, or through adjusted SELEX protocols (Table 1). In the first method, researchers systematically introduce modifications to a previously selected aptamer and a number of modified aptamers are analyzed to evaluate their affinity, selectivity, and stability. Although this method is widely used in the field,8,67,148,150–153 it is time-consuming and labour-intensive due to the unknown role and impact a given XNA substitution could have on aptamer folding and function. Furthermore, due to the usual trade-off between affinity and aptamer chemical modification it is rare to identify fully-substituted aptamers this way.3,63,148 (green) have been incorporated during selection,
(green) have been incorporated during selection,  (pink) only after selection and
(pink) only after selection and  (blue) both during and after selection. Where reported, targets screened for specificity are noted. No off-target binding was reported (unless otherwise stated in the table). The polymerases used in the SELEX process are listed with polymerases used directly for the XNA chemistry shown in green (if commercially available) or red (if not commercially available). N.R. – not reported. N.A. not applicable
(blue) both during and after selection. Where reported, targets screened for specificity are noted. No off-target binding was reported (unless otherwise stated in the table). The polymerases used in the SELEX process are listed with polymerases used directly for the XNA chemistry shown in green (if commercially available) or red (if not commercially available). N.R. – not reported. N.A. not applicable
The alternative solution is to adapt the selection protocol to directly integrate XNA chemistries. However, this requires a re-evaluation of every step in the SELEX process – from library generation to selection cycles – to ensure optimal efficiency and success.
Creating libraries for DNA is relatively straightforward, often involving the purchase of pre-synthesized libraries from specialized companies. For RNA libraries, a common practice involves transcribing DNA libraries, facilitated by commercially available kits. However, with XNA, building blocks are not often commercially available (thus requiring specialist chemistry for synthesis), chemical synthesis of libraries is often not possible (because of lower chemical couplings of the building blocks), and synthesis of the nucleotide triphosphate equivalent is not as efficient as for natural (deoxy)ribonucleotides.
Consequently, XNA SELEX adds two extra requirements: polymerases that can synthesize XNA libraries for selection and enzymes that can access the XNA information in the library post-selection. While some base and sugar modifications can be efficiently incorporated by natural or commercially available engineered polymerase variants, most backbone-modified XNAs rely on specialized enzymes that are not widely available (see next section).
Other steps in the SELEX process are also affected. While conventional DNA-based SELEX protocols can rely on enzymes like lambda exonuclease to degrade 5′-phosphorylated template strands in double-stranded DNA to obtain a library of single strands, XNA-based SELEX often requires alternatives, e.g. isolation through PAGE,154,155 degradation of the DNA templates through DNase I,156 magnetic beads155,157 or other innovative solutions158,159 – invariably increasing the duration and cost of SELEX.
Additionally, while the field has made great progress since the turn of the century in terms of computational analyses of aptamer selection and 2D/3D structure predictions, knowledge of XNA structures is very limited, curtailing possible structure prediction efforts.
Polymerases as a bottleneck for XNA aptamer discovery
The existence of a given XNA chemistry, and even the demonstration that it can be processed by an engineered polymerase, does not necessarily imply that its full sequence landscape (and therefore functional landscape) is experimentally accessible. Efficient SELEX requires polymerases capable of efficient synthesis (XNA from DNA templates and DNA from XNA templates) in reaction conditions that are often challenging, e.g. biased sequences during library synthesis, or low template concentrations after NA recovery.XNA synthesis is constrained by the stability of the synthesised duplex, and by the polymerase's sequence tolerance and error profile. The reported tPhoNA (phosphonomethyl-threosyl nucleic acid) showed significant stalling when two Ts were incorporated sequentially.160 Similarly, early TNA polymerases showed significant G:G mispairing.161 In both cases, the result would be the selective exclusion of certain sequences from the libraries under selection. Such constraints have long been recognised in nucleic acid evolution studies, where the success of selection experiments depends on how amenable to replication a sequence is, as much as on its binding capability.162,163
For DNA selections, polymerase performance has historically been a decisive enabling factor. Thermostable DNA polymerases, particularly Taq and its derivatives, allow detection and amplification from extremely small amounts of template, while maintaining sufficient fidelity to propagate functional sequences across selection cycles.164 The ability to amplify minute amounts of template directly, affects selection stringency: higher stringency conditions recover fewer nucleic acid molecules, and only robust amplification permits those stringent conditions to be applied without losing the evolving population.165 Moreover, DNA PCR amplification bias has been extensively studied, enabling researchers to mitigate its impact during SELEX and preserve a substantial fraction of library diversity, even for structured templates.166,167
A similar principle applies to RNA selections, which typically include transcription, reverse transcription, and PCR amplification steps. Reverse transcriptases such as M-MLV RT, AMV RT, and thermostable variants (e.g. SuperScript derivatives) are routinely used to convert selected RNA into cDNA prior to PCR.165,168 Their efficiency is sufficiently high that even structured RNAs can often be recovered and propagated.
For XNA chemistries that structurally resemble DNA/RNA (e.g. most modifications on the 2′ position or the nucleobase itself) or where modification of all four bases is not coveted, natural polymerases or commercially available engineered ones can still perform adequately (e.g. Therminator™ DNA polymerase incorporating TNA112). Nucleobase modifications can often be tolerated by natural polymerases and therefore can be directly amplified157,169–171 or amplified in their reverse transcription or transcription steps.172–174 Sugar or backbone modifications perform less well, e.g. PS, which can be directly amplified using Taq polymerase.175,176
For more divergent XNA chemistries, engineered XNA polymerases are essential and, despite advances in the field, remain a significant practical constraint. Consequently, their use for in vitro selection of XNA aptamers remains rare. In 2005, probably the first report of fully substituted XNA aptamer selections, Burmeister et al. used a T7 triple-mutant Y639F/H784A/K378R for the selection of a fully modified 2′-OMe aptamer binding VEGF.177 Significant advances in DNA polymerase engineering have enabled new families of XNA polymerases to be developed, and with that the expansion of the available repertoire of XNAs accessible to SELEX.19 The first report of in vitro selected HNA (1,5-anhydrohexitol nucleic acids) aptamers against HIV-1 TAR and hen egg lysozyme was quickly followed by multiple reports covering a wide range of nucleic acid chemistries, summarized in Table 1.
At present, most XNA polymerases derive from archaeal hyperthermophilic B-family DNA polymerases.178–181 Although bearing lower processivity than equivalent A-family polymerases, B-family enzymes generally exhibit greater substrate tolerance, making them suitable starting points for XNA polymerase engineering. A wide range of selected and engineered polymerases are available (both for XNA synthesis and XNA reverse transcription),19,182–184 but comparison of their biochemical activity highlights common engineering trade-offs.178 Mutations that enable XNA synthesis typically expand the substrate spectrum of the enzymes (rather than create polymerases specialist on the new chemistry) and do not remove their DNA polymerase activity. This broader substrate spectrum is often linked to decrease fidelity (beyond mutations that simply inactivate the error proofing mechanisms).3,27,185 Similar XNA chemistry variation is also observed among the engineered transcriptases.
In the context of SELEX, those features are not neutral. Lower amplification efficiencies and lower fidelity, limit the stringency that can be applied in a SELEX round, favour amplification of sequences that are easier to replicate, and selectively bias SELEX towards areas of the sequence space that are less rugged. As such, enrichment reflects not only function, but also the compatibility of the sequences with the XNA chemistry and polymerases being used. This process is exacerbated in XNA chemistries where exponential amplification is not possible, i.e. systems where XNA synthesis (or RT) relies on a transcription-like linear amplification or in direct synthesis of XNA from a DNA template – processes that limit sequence enrichment.178,186
There are also additional consequences to the SELEX process. Classical SELEX typically requires approximately 8–15 selection rounds before enrichment is achieved.5,165 Each selection round creates a new opportunity for polymerase-mediated effects to emerge, such as biases, parasites and amplification artefacts. Indeed, the accumulation of PCR byproducts, amplification parasites and artefacts is well documented even in SELEX for natural nucleic acids.166,187
Factors impacting XNA diversity and adoption
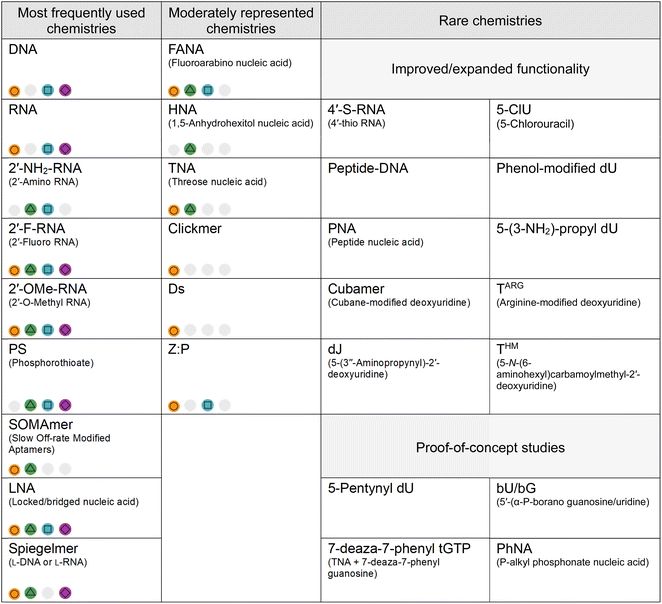
While we highlight representatives from a range of XNA chemistries in Table 1, that does not reflect the unequal frequency of XNA chemistries in the aptamer literature, which is shaped by feasibility rather than theoretical chemical potential. Multiple factors play into which chemistries dominate in the research field: (1) the time a chemistry has had to mature, (2) selection compatibility (both enzymatic and chemical synthesis), (3) a balance between benefit vs. cost-effectiveness and (4) the extent to which the XNA has been characterised (e.g. reached clinical trials). That is summarised in Table 2. – good compatibility with selection (efficient enzymatic/chemical synthesis),
– good compatibility with selection (efficient enzymatic/chemical synthesis),  – improved biological or chemical stability,
– improved biological or chemical stability,  – synthetic nucleotide triphosphates are commercially available,
– synthetic nucleotide triphosphates are commercially available,  – known pharmacokinetic properties,
– known pharmacokinetic properties,  – not known or not applicable. For the category of rare chemistries, XNAs are subdivided into those researched for their improved functionality (e.g. 4′-S-RNA increases nuclease resistance; cubamer adds cationic side chains) and those that use XNA aptamer selection as a proof of principle (often to demonstrate polymerase capabilities)
– not known or not applicable. For the category of rare chemistries, XNAs are subdivided into those researched for their improved functionality (e.g. 4′-S-RNA increases nuclease resistance; cubamer adds cationic side chains) and those that use XNA aptamer selection as a proof of principle (often to demonstrate polymerase capabilities)
Firstly, modifications appearing earlier in the literature tend to still be more heavily featured in XNA aptamers to date. Modifications such as 2′-F and 2′-OMe were developed decades ago, in 1961 (ref. 12) and 1965,11 respectively, and were incorporated into aptamers shortly after the emergence of the research field, primarily as strategies to enhance nuclease resistance. By the time one of the earliest mentions of FANA aptamers appeared in literature in 2007,65 2′-F/OMe modifications already had accumulated more than a decade of substantial methodological refinement and empirical validation. Similarly, frequently reported sugar modifications such as LNA67,148 already appear in literature as post-SELEX modified aptamers before the 2010's – preceding the use of specialised engineered polymerases. In comparison, chemistries such as HNA and TNA for which the tools have only been reported in 2012 and where optimization is still ongoing, lag behind the field.19,112,188,189
Secondly, as discussed above, polymerases play a crucial role in SELEX and it thus contributes substantially to the disparity among XNA chemistries. Unsurprisingly, the prevalence of reported XNA aptamers mirrors the availability of robust polymerase systems. The most frequently used chemistries (each reported in more than 100 aptamers) include 2′-F/OMe,8,17,177,190–194 SOMAmers,174,193,195 LNA,67,148,156,158,196,197 spiegelmers,198–203 and PS,18,135,176,204–207 have established SELEX pipelines and can be amplified using commercially available or robust engineered polymerases. In contrast, chemistries such as HNA remain less frequently reported despite favourable properties (including strong nuclease resistance, serum stability and duplex thermostability). HNA-based selections have yielded high-affinity ligands, even achieving low-nanomolar binding constants when combined with 2′-OMe-modified bases.118 Similarly, TNA aptamers have reached low-nanomolar affinities following the development of a robust KOD-RI polymerase.117,154,184,208,209 Nevertheless, because both platforms rely on specialized engineered polymerases and tailored synthetic methodologies, their broader implementation remains limited.
Accessibility to XNA precursors also affects which XNAs are used. Many heavily modified XNAs – such as some 5′-substituted dUTPs,210–212 Cubamer,157 dJ,213 phosphonate backbones,214 TARG,172 and THM66,173 – require bespoke synthetic routes, appear infrequently in the literature and are limited to a few specialist research groups.
Equally, solid-phase synthesis of selected sequences should be accessible. While polymerases are used during selection rounds, solid-phase synthesis (chemical synthesis of oligonucleotides using phosphoramidite chemistry) is usually necessary to create sequences for further characterisation or production of the selected aptamer. At that stage, the efficiency of the solid-phase coupling and the cost-effectiveness of the process become relevant bottlenecks. For example, while both 2′-F and 2′-NH2 modifications are relatively easily synthesised enzymatically, 2′-F quickly became preferred over 2′-NH2 because of higher coupling efficiency and additional deprotection steps required during purification.215
Last, is the potential for a given XNA to provide improved functionality, whether by providing greater chemical (or biological) stability, or by introducing greater chemical diversity to the oligomer (thus allowing a wider repertoire of intra- and inter-molecular interactions). For example, 2′-F/OMe modifications are routinely added to RNA aptamers as they improve endonuclease resistance due to the 2′ modification replacing the required 2′-hydroxyl group. SOMAmers and clickmers216–219 both expand functional diversity by incorporating hydrophobic or aromatic side chains – mimicking amino acid side chains – while simultaneously enhancing stability and, in many cases, increase binding affinity.119,174 Their translation into commercial platforms illustrates the importance of accessible standard protocols and sought-after novel interactions.
Adoption of certain chemistries in therapeutic contexts is limited by our understanding of their pharmacokinetic behaviour. Only a small number of aptamer chemistries have progressed to clinical trials (DNA, RNA, 2′-F/OMe, spiegelmers) and consequently their pharmacokinetic properties are comparatively well characterized.8,190,220 Many other modifications have only been tested in in vivo mouse or rat models. For instance, SOMAmers demonstrated slower plasma clearance (dependent on the number and nature of modifications) in rat models.221 For other XNA chemistries, pharmacokinetic insights remain more limited and are often derived indirectly. Therapeutic antisense oligonucleotides have given insight into the pharmacokinetics of chemistries such as LNA and PS, which are expected to mirror aptamer behaviour.222,223 However, quantitative pharmacokinetic parameters for many newer or more obscure XNAs remain largely undefined.
Antibodies vs. aptamers
Given the more than two decades of head start, it is not surprising that antibodies have established themselves as a dominant force in both the diagnostic and therapeutic markets. In the period from 2018 to 2023, regulatory agencies such as the FDA and EMA granted approvals for 109 monoclonal antibodies and only 16 nucleic acid-based therapeutics, with merely one belonging to the aptamer category. This underrepresentation of aptamers highlights not only the nascent state of the field but also the limited understanding of the therapeutic potential of aptamers.224,225Determination of nucleic acid structures has historically been more difficult than that of proteins. This is reflected in the number of available structures in public databases, where proteins in the PDB database226 surpass 1.7 million structures, providing a comprehensive understanding of protein configurations, compared to little under 60 thousand for nucleic acids (35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 426 for DNA and 19
426 for DNA and 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 635 for RNA) in the PDB and NDB databases. The lower number of available structures, the narrower chemical spectrum of nucleic acids, and our incomplete understanding of interactions beyond Watson–Crick base-pairing currently limit our progress on the de novo structure prediction and on the structure-based design of aptamers.
635 for RNA) in the PDB and NDB databases. The lower number of available structures, the narrower chemical spectrum of nucleic acids, and our incomplete understanding of interactions beyond Watson–Crick base-pairing currently limit our progress on the de novo structure prediction and on the structure-based design of aptamers.
X-ray crystallography and NMR spectroscopy, each bearing their own advantages and challenges, are the common approaches for aptamer structure determination. High-quality crystals are difficult to isolate for nucleic acids and as a result, many of the available structures have been obtained through NMR. However, NMR is only suitable for smaller complexes (typically aptamers of 30–50 nucleotides and smaller proteins), due to narrow chemistry resulting in spectral overlap that emerges quickly with increasing number of nucleotides. Newer strategies, such as cryo-electron microscopy (cryo-EM), are gaining traction but unbound aptamer structure will remain difficult to determine (because of their natural structural plasticity).227
The accuracy of aptamer structure prediction tools, typically trained by the relatively limited known nucleic acid structures, lags significantly compared to protein structure prediction methods. At CASP15, prediction strategies such as those used by the Yang-Server team – trRosettaX2 (an improved version of trRosettaX228) and AlphaFold2 (ref. 229) – reached TM-scores of 0.876 with backbone accuracies of 0.96 Å RMSD for proteins. The latest release from DeepMind, AlphaFold 3, promises even higher accuracy for protein monomers and enhanced predictions for protein–protein interactions.124
In contrast, CASP15 marked the first inclusion of RNA structure prediction.230 Across 12 RNA targets of varying complexity – ranging from those with identifiable templates to those without – predictions exhibited RMSD values spanning 2 Å to nearly 17 Å. The field did not anticipate achieving atomic-level precision and the current emphasis remains on accurately reproducing overall folding shapes.124 Nonetheless, in silico approaches for nucleic acid structure determination are gathering pace (see next section).
A gap between antibodies and aptamers is also present in the diagnostic landscape, where the antibody-based diagnostic market, valued at US $20,000 million in 2017 and projected to reach US $35,000 million by 2026, reflects the widespread adoption and established role of antibody-based diagnostics in disease detection and monitoring.231 Despite the potential advantages of aptamers (i.e. small size, cheaper production with low batch-to-batch variability, better thermal stability for transport and storage) that would make them highly relevant for point-of-care testing,232 only 10 are undergoing clinical trials for the development of diagnostics.
Still, aptamer-based diagnostic technologies are entering the market:233 companies like NeoVentures (https://neoaptamers.com/), who provide diagnostics, detection kits and aptamer-based affinity columns; and Aptamer Science Inc. (https://aptsci.com/en/), who commercialize several biomarker detection kits. Another notable example is the collaboration between SomaLogic and Illumina, announced in 2022, which combines the SomaScan® Proteomics Assay with high throughput NGS platforms to support multi-omic profiling for novel biomarker discovery.
Finally, the two technologies are not incompatible and can be combined, e.g. with aptamer–antibody conjugates, where aptamer pharmacokinetic properties can be improved by the conjugation to the larger antibodies (or other proteins), deep tissue penetration can be enhanced, or their conjugation can work synergistically for a more potent inhibitory effect.234
XNA aptamer structure determination through molecular modelling
As mentioned above, while progress is being made in the development of computational tools for aptamer structure prediction, the “AlphaFold” moment for the field has not happened yet and will require advancements in both computational algorithms and the availability of high-quality structural data for aptamers.235 Still, companies are already entering the market delivering custom DNA and RNA aptamers isolated in silico from AI-based platforms (e.g. Xelari, https://www.xelari.com).The scarcity of experimental XNA aptamer structures may complicate the use of AI-based methods for structure prediction, but it does not preclude the exploration of alternative computational strategies. In this regard, molecular modelling offers an interesting complementary route, in line with integrative structure-modelling approaches recently proposed for RNA and protein-RNA complexes.236
Molecular modelling of XNA aptamers faces two major challenges: (1) accurate parametrisation of the XNA chemistry and (2) predicting the aptamer folding from scratch. Parametrisation requires deriving force field parameters that accurately describe the structural and energetic behaviour of the biomolecule, including both bonded and non-bonded interactions.237,238 While well-established force fields exist for DNA and RNA239,240 they are generally insufficient to accurately describe the diverse chemical modifications present in XNAs, requiring customised parameters.
Obtaining accurate XNA parameters typically involves quantum mechanical (QM) calculations to model the linker and (or) the nucleoside. Significant progress has been made in recent years, particularly in conformational sampling of five- and six-membered (sugar) rings.241 For instance, the pucker.rs toolkit242 simplifies the generation of coordinates required for QM-based conformational sampling of sugar rings and linkers. Representative structures obtained from such sampling can then be used for force field parametrisation. Bonded and non-bonded parameters are typically derived using tools such as ParamFit, while partial charges are determined separately via ESP/RESP calculations.243–246
This methodology has been successfully applied in several studies to parametrise diverse XNA oligonucleotides, including HNA, dXyNA, TNA and tPhoNA. Using the resulting force fields, molecular dynamics (MD) simulations were performed, and the predicted structures obtained were found to be in good agreement with available experimental data.245,247 These examples demonstrate that such computational pipelines are feasible and can yield reliable structural models for chemically modified oligonucleotides. Crucially, this approach could, in principle, also be extended to XNA aptamers. Most recently, AmberTools 2025 introduced the modXNA module,248 providing a framework for XNA parameterisation within AMBER and simplifying access to parameters for modified nucleotides. While current validations focus on relatively small modifications (e.g., phosphorothioate, 2′F, 2′OMe, 5FU, methylated bases), it represents an encouraging step towards broader accessibility of XNA modelling tools.
The second challenge – predicting the folding of XNA aptamers – remains inherently more complex. Conventional MD simulations often struggle to explore the full folding landscape, as the system can become trapped in local energy minima. To overcome these limitations, enhanced sampling methods such as replica-exchange molecular dynamics (REMD) and simulated annealing, originally developed in the field of protein folding, have proven useful and can be applied to study aptamer folding.249,250 In REMD, several copies of the system are simulated simultaneously at different temperatures, occasionally swapping configurations to help it explore otherwise inaccessible states. Simulated annealing involves heating the system and then slowly cooling it, allowing it to sample a wider range of conformations. Simple restraints can guide the folding toward plausible structures, which are then refined in longer simulations to access stability. This strategy was successfully applied to predict the folded structure of an HNA aptamer, though its implementation remains challenging and often involves rounds of trial-and-error approaches.251
In parallel, for DNA and RNA aptamers, computational workflows have been developed to predict how an aptamer folds in complex with its target. These typically involve secondary structure prediction, 3D modelling, docking to the target, and subsequent MD simulations to evaluate stability and interactions.33,35 However, these workflows have so far been applied only to DNA and RNA, and extending them to XNA systems will require further development.
In the long term, as molecular modelling of XNA aptamers becomes increasingly accurate, the resulting structural ensembles could provide valuable training data for AI models. This would help alleviate the current scarcity of experimental structures and gradually bridge the gap between physics-based simulations and data-driven predictive approaches, though this remains a long-term goal.
The future of (XNA) aptamers
The aptamer field has undergone significant evolution across its various branches since its beginning in the 90 s. Throughout much of that time, the focus in the field has been to enhance the efficiency and reproducibility of the aptamer discovery process.252 Early successes, targeting proteins with high pI like lysozyme47,253–255 and thrombin,211,256–259 may have contributed to the complexities in aptamer development being underestimated, which has led to the historical paradigm that aptamer isolation is contingent upon luck to encounter a high-affinity sequence in the initial nucleic acid population.260–262 Recent insights, however, highlight the importance of the starting population composition,261,262 which has been combined with computational tools to construct improved libraries.263NGS has accelerated the identification of putative aptamers by enabling high-throughput analysis of nucleic acid populations, yet it introduces novel challenges, particularly in data interpretation.264 Traditional analysis methods predominantly focus on sequence-level information, often overlooking crucial structural attributes that govern aptamer functionality. Innovations in clustering algorithms designed to group aptamer sequences based on their structural profiles represent a fast-growing area of research,264 but their adoption in aptamer discovery pipelines is still emerging.
Lastly, the expanding repertoire of available XNA chemistries has direct consequences to the aptamer selection toolkit, offering unique properties that may enhance pharmacokinetic profiles or introduce novel chemical characteristics198,199 – but for which in silico tools are scarce.265
Nevertheless, increased effort without a change in paradigm will not be enough to bring aptamers into the limelight. From a more theoretical perspective, several critical aspects of the field demand attention.
First, there is a need for a more precise and objective definition of aptamers. The absence of a clear, universally accepted definition has led to ambiguity in distinguishing aptamers from single stranded nucleic acids with weak or nonspecific binding capabilities.144,266 In addition, there have been historical misconceptions in the field regarding the correlation between high affinity and high selectivity.267 It took nearly a decade for these assumptions to be challenged and revised.268 Therefore, there is a recognized need to establish a formal definition that comprehensively incorporates both affinity and selectivity in a paradigm that can be experimentally validated.
Second, although aptamers are often compared to antibodies, they are a unique class of biopolymers and require their own development paradigm. Aptamers are more complex than proteins, capable of reversible changes in response to environmental conditions,269,270 and exist within a very different sequence/functional landscape. In practical terms, it means that we have to rewrite our expectations on biophysical behaviour of aptamers and to develop tools that capture their true potential (or at least highlight their strength).
Third, the narrow chemical and biological stability of natural nucleic acids means that XNAs, irrespective of which nucleotide moiety is chemically modified, will become a central aspect of the future of therapeutic (and possibly also diagnostic) aptamers. The different XNA chemistries counteract the perceived chemical monotony of nucleic acids (i.e. the four natural nucleobases versus the 20 natural amino acids) and have been expanded through efforts to develop novel base pairs,271 or through the development XNA synthesis methods beyond polymerase-catalysed synthesis.272
Beyond the research itself, we must standardize publication criteria, by clearly describing selection protocols that are rigorous and reproducible, transparently detailing our NGS data analysis pipelines, and report our candidate selection methodologies.273
Solutions to these problems are possible and given the fast pace of research in the field, likely to emerge still within this decade. We look forward to see how the field will evolve and whether aptamers can finally live up to their potential.
Conflicts of interest
There are no conflicts to declare.Data availability
Our review makes reference to many published reports, but we have not included any original data or analyses in the current manuscript. As such, there are no data that can be made available.References
- F. Ducongé and J.-J. Toulmé, RNA, 1999, 5, S1355838299991318 CrossRef PubMed.
- S. Chaillou, P.-E. Stamou, L. L. Torres, A. B. Riesco, W. Hazelton and V. B. Pinheiro, Nucleic Acids Res., 2022, 50, 9568–9579 CrossRef CAS PubMed.
- J. Zhou and J. Rossi, Nat. Rev. Drug Discovery, 2017, 16, 181–202 CrossRef CAS PubMed.
- C. Tuerk and L. Gold, Science, 1990, 249, 505–510 CrossRef CAS.
- A. D. Ellington and J. W. Szostak, Nature, 1990, 346, 818–822 CrossRef CAS.
- U. Chadha, P. Bhardwaj, R. Agarwal, P. Rawat, R. Agarwal, I. Gupta, M. Panjwani, S. Singh, C. Ahuja, S. K. Selvaraj, M. Banavoth, P. Sonar, B. Badoni and A. Chakravorty, J. Ind. Eng. Chem., 2022, 109, 21–51 CrossRef CAS.
- A. M. Khanani, S. S. Patel, G. Staurenghi, R. Tadayoni, C. J. Danzig, D. A. Eichenbaum, J. Hsu, C. C. Wykoff, J. S. Heier, D. R. Lally, J. Monés, J. S. Nielsen, V. S. Sheth, P. K. Kaiser, J. Clark, L. Zhu, H. Patel, J. Tang, D. Desai, G. J. Jaffe, A. Alezzandrini, A. A. Francone, J. Bafalluy, S. Bainttein, J. Luna Pinto, M. Saravia, M. Vidosevich, C. Zeolite, F. Furno Sola, A. Chang, E. E. G. Cornish, T. Nguyen, O. Findl, A. Haas, M. Kralinger, S. Sacu, L. D. Postelmans, M. Farah, M. Maia, M. Nehemy, F. Ali, M. Brent, M. Dollin, J. Gonder, A. Kherani, A. Merkur, R. Tuli, M. M. Lopera, F. Rodriguez, M. Bradvica, J. Ernest, K. Kalijurand, K. Noor, Y. Cohen, C. Creuzot-Garcher, F. De Bats, F. Devin, C. Français-Maury, L. Kodjikian, J. F. Korobelnik, Y. Le Mer, M. Quaranta El Maftouhi, S. Razavi, E. Souied, R. Tadayoni, M. Weber, N. Eter, N. Feltgen, S. Grisanti, P. Walter, R. Liegl, K. Lorenz, G. Spital, S. Priglinger, M. Spitzer, M. Volker, T. Krohne, C. Jochmann, C. P. Lohmann, C. Framme, A. Kerenyi, A. Papp, A. Seres, E. Toth-Molnar, A. Tsorbatzoglou, A. Vajas, B. Varsanyi, G. Vogt, A. Bar, E. Eting, D. Hauser, J. Levy, N. Mathalone, H. Morori-Katz, I. Rosenblatt, S. Soudry-Zayit, O. Trivizky, F. Bandello, A. P. Ciardella, M. Figus, F. Giansanti, P. Lanzetta, C. Mariotti, L. Mastropasqua, E. Midena, F. Parmeggiani, F. Ricci, F. Simonelli, G. Staurenghi, F. Viola, M. Varano, G. Laganovska, S. Cisiecki, W. Jedrzejewski, J. Kaluzny, M. Misiuk-Hojło, S. Abengoechea, J. A. Iribarren, F. J. Ascaso, J. M. Cubero, R. Gallego-Pinazo, F. Gomez-Ulla De Irazazabal, I. J. Mestre, J. M. Mones I Carilla, J. Montero Moreno, J. María Ruiz Moreno, L. Sararols Ramsay, A. Garcia Layana, L. Downey, P. Abraham, D. V. Alfaro, N. Bagheri, I. Barbazetto, J. Benevento, P. Bernstein, G. Bertolucci, A. Bhavsar, W. Bridges, H. L. Brooks Jr, J. Brown, A. Brucker, C. M. Calvo, A. Capone, J. Carlson, C. Chan, E. Chang, B. Chan-Kai, N. Chaudhry, S. Chen, K. Csaky, H. Cummings, C. Danzig, A. Dessouki, D. Dyer, A. Eaton, D. Eichenbaum, D. Faber, R. Feldman, N. Finnen, W. Freeman, R. Frenkel, C. Gonzales, V. Gonzalez, J. Gross, S. Gupta, E. Hall, M.-K. Han, J. Heier, V. Hershberger, P. Higgins, J. Hsu, M. Ip, E. Jablon, B. Jewart, V. John, J. Jonisch, B. Joondeph, C. Kay, A. Khanani, G. T. Kokame, R. Kwun, M. Lai, D. Lally, K. Laud, A. Lavina, M. Lee, P. Lin, H. Lin, N. Manoharan, D. Marcus, A. Martidis, F. McCabe, J. Nielsen, J. Osher, J. Palmer, S. Patel, J. Pearlman, S. Perkins, A. Pirouz, J. Qureshi, J. Randolph, N. Piri, P. Rosenfeld, D. Saperstein, R. Scartozzi, S. Schwartz, A. Sharma, A. Sharma, V. Sheth, M. Singer, D. Spinak, E. Suan, H. Tabandeh, A. Tabassian, E. Uchiyama, M. Varenhorst, A. Wagner, D. Warrow, J. Wells III, R. Wong, K. Wong, C. Wykoff, S. Xavier and E. Ysasaga, Lancet, 2023, 402, 1449–1458 CrossRef CAS.
- E. W. M. Ng, D. T. Shima, P. Calias, E. T. Cunningham, D. R. Guyer and A. P. Adamis, Nat. Rev. Drug Discovery, 2006, 5, 123–132 CrossRef CAS PubMed.
- M. Egli and M. Manoharan, Nucleic Acids Res., 2023, 51, 2529–2573 CrossRef CAS PubMed.
- F. Eckstein, J. Am. Chem. Soc., 1966, 88, 4292–4294 CrossRef CAS.
- Y. Furukawa, K. Kobayashi, Y. Kanai and M. Honjo, Chem. Pharm. Bull., 1965, 13, 1273–1278 CrossRef CAS PubMed.
- J. F. Codington, I. Doerr, D. Van Praag, A. Bendich and J. J. Fox, J. Am. Chem. Soc., 1961, 83, 5030–5031 CrossRef CAS.
- C. C. Smith, L. Aurelian, M. P. Reddy, P. S. Miller and P. O. Ts’o, Proc. Natl. Acad. Sci. U. S. A., 1986, 83, 2787–2791 CrossRef CAS PubMed.
- M. Matsukura, K. Shinozuka, G. Zon, H. Mitsuya, M. Reitz, J. S. Cohen and S. Broder, Proc. Natl. Acad. Sci. U. S. A., 1987, 84, 7706–7710 CrossRef CAS.
- J. A. Latham, R. Johnson and J. J. Toole, Nucleic Acids Res., 1994, 22, 2817–2822 CrossRef CAS PubMed.
- D. Jellinek, L. S. Green, C. Bell, C. K. Lynott, N. Gill, C. Vargeese, G. Kirschenheuter, D. P. C. McGee and P. Abesinghe, Biochemistry, 1995, 34, 11363–11372 CrossRef CAS PubMed.
- L. S. Green, D. Jellinek, C. Bell, L. A. Beebe, B. D. Feistner, S. C. Gill, F. M. Jucker and N. Janjić, Chem. Biol., 1995, 2, 683–695 CrossRef CAS PubMed.
- D. J. King, D. A. Ventura, A. R. Brasier and D. G. Gorenstein, Biochemistry, 1998, 37, 16489–16493 CrossRef CAS PubMed.
- V. B. Pinheiro, A. I. Taylor, C. Cozens, M. Abramov, M. Renders, S. Zhang, J. C. Chaput, J. Wengel, S.-Y. Peak-Chew, S. H. McLaughlin, P. Herdewijn and P. Holliger, Science, 2012, 336, 341–344 CrossRef CAS PubMed.
- S. Arangundy-Franklin, A. I. Taylor, B. T. Porebski, V. Genna, S. Peak-Chew, A. Vaisman, R. Woodgate, M. Orozco and P. Holliger, Nat. Chem., 2019, 11, 533–542 CrossRef CAS PubMed.
- Y. Brudno, M. E. Birnbaum, R. E. Kleiner and D. R. Liu, Nat. Chem. Biol., 2010, 6, 148–155 CrossRef CAS PubMed.
- L. Gold, J. J. Walker, S. K. Wilcox and S. Williams, New Biotechnol., 2012, 29, 543–549 CrossRef CAS PubMed.
- P. Herdewijn and P. Marlière, Chem. Biodiversity, 2009, 6, 791–808 CrossRef CAS.
- J. C. Chaput and P. Herdewijn, Angew. Chem., Int. Ed., 2019, 58, 11570–11572 CrossRef CAS PubMed.
- S. M. Nimjee, R. R. White, R. C. Becker and B. A. Sullenger, Annu. Rev. Pharmacol. Toxicol., 2017, 57, 61–79 CrossRef CAS PubMed.
- G. Mayer, Angew. Chem., Int. Ed., 2009, 48, 2672–2689 CrossRef CAS PubMed.
- P. Röthlisberger and M. Hollenstein, Adv. Drug Delivery Rev., 2018, 134, 3–21 CrossRef PubMed.
- J. P. Elskens, J. M. Elskens and A. Madder, Int. J. Mol. Sci., 2020, 21, 4522 CrossRef CAS PubMed.
- S. Liu, Y. Xu, X. Jiang, H. Tan and B. Ying, Biosens. Bioelectron., 2022, 208, 114168 CrossRef CAS PubMed.
- E. Eremeeva and P. Herdewijn, Curr. Opin. Biotechnol., 2019, 57, 25–33 CrossRef CAS PubMed.
- A. I. Taylor, G. Houlihan and P. Holliger, Cold Spring Harbor Perspect. Biol., 2019, 11, a032490 CrossRef CAS PubMed.
- A. Khvorova and J. K. Watts, Nat. Biotechnol., 2017, 35, 238–248 CrossRef CAS PubMed.
- D. Sun, M. Sun, J. Zhang, X. Lin, Y. Zhang, F. Lin, P. Zhang, C. Yang and J. Song, TrAC, Trends Anal. Chem., 2022, 157, 116767 CrossRef CAS.
- N. A. Ahmad, R. Mohamed Zulkifli, H. Hussin and M. H. Nadri, J. Mol. Graphics Modell., 2021, 105, 107872 CrossRef CAS PubMed.
- T. N. Navien, R. Thevendran, H. Y. Hamdani, T.-H. Tang and M. Citartan, Biochimie, 2021, 180, 54–67 CrossRef CAS PubMed.
- N. Komarova, D. Barkova and A. Kuznetsov, Int. J. Mol. Sci., 2020, 21, 8774 CrossRef CAS PubMed.
- G. F. Joyce and T. Inoue, Nucleic Acids Res., 1989, 17, 711–722 CAS.
- L. C. Bock, L. C. Griffin, J. A. Latham, E. H. Vermaas and J. J. Toole, Nature, 1992, 355, 564–566 CrossRef CAS PubMed.
- M. Sassanfar and J. W. Szostak, Nature, 1993, 364, 550–553 CrossRef CAS.
- L. Gold, B. Polisky, O. Uhlenbeck and M. Yarus, Annu. Rev. Biochem., 1995, 64, 763–797 CrossRef CAS PubMed.
- S. Jhaveri, B. Olwin and A. D. Ellington, Bioorg. Med. Chem. Lett., 1998, 8, 2285–2290 CrossRef CAS PubMed.
- A. Nolte, S. Klußmann, R. Bald, V. A. Erdmann and J. P. Fürste, Nat. Biotechnol., 1996, 14, 1116–1119 CrossRef CAS PubMed.
- S. Klußmann, A. Nolte, R. Bald, V. A. Erdmann and J. P. Fürste, Nat. Biotechnol., 1996, 14, 1112–1115 CrossRef PubMed.
- D. J. Patel, A. K. Suri, F. Jiang, L. Jiang, P. Fan, R. A. Kumar and S. Nonin, J. Mol. Biol., 1997, 272, 645–664 CrossRef CAS.
- E. Brody, M. Willis, J. Smith, S. Jayasena, D. Zichi and L. Gold, Mol. Diagn., 1999, 4, 381–388 CrossRef CAS PubMed.
- B. Vant-Hull, L. Gold and D. A. Zichi, Curr. Protoc. Nucleic Acid Chem., 2000, 9.1.1–9.1.16 Search PubMed.
- J. C. Cox and A. D. Ellington, Bioorg. Med. Chem., 2001, 9, 2525–2531 CrossRef CAS PubMed.
- R. Stoltenburg, C. Reinemann and B. Strehlitz, Anal. Bioanal. Chem., 2005, 383, 83–91 CrossRef CAS PubMed.
- S. D. Mendonsa and M. T. Bowser, J. Am. Chem. Soc., 2004, 126, 20–21 CrossRef CAS.
- M. L. Raddatz, A. Dolf, E. Endl, P. Knolle, M. Famulok and G. Mayer, Angew. Chem., Int. Ed., 2008, 47, 5190–5193 CrossRef CAS PubMed.
- X. Lou, J. Qian, Y. Xiao, L. Viel, A. E. Gerdon, E. T. Lagally, P. Atzberger, T. M. Tarasow, A. J. Heeger and H. T. Soh, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 2989–2994 CrossRef CAS PubMed.
- M. Berezovski, M. Musheev, A. Drabovich and S. N. Krylov, J. Am. Chem. Soc., 2006, 128, 1410–1411 CrossRef CAS PubMed.
- F. Chen, J. Zhou, F. Luo, A.-B. Mohammed and X.-L. Zhang, Biochem. Biophys. Res. Commun., 2007, 357, 743–748 CrossRef CAS PubMed.
- D. Shangguan, L. Meng, Z. C. Cao, Z. Xiao, X. Fang, Y. Li, D. Cardona, R. P. Witek, C. Liu and W. Tan, Anal. Chem., 2008, 80, 721–728 CrossRef CAS.
- K. Kikuchi, J. Biochem., 2003, 133, 263–270 CrossRef CAS.
- J. Mi, Y. Liu, Z. N. Rabbani, Z. Yang, J. H. Urban, B. A. Sullenger and B. M. Clary, Nat. Chem. Biol., 2010, 6, 22–24 CrossRef CAS PubMed.
- B. J. Hicke and A. W. Stephens, J. Clin. Invest., 2000, 106, 923–928 CrossRef CAS PubMed.
- N. Hamaguchi, A. Ellington and M. Stanton, Anal. Biochem., 2001, 294, 126–131 CrossRef CAS PubMed.
- W. Winkler, A. Nahvi and R. R. Breaker, Nature, 2002, 419, 952–956 CrossRef CAS PubMed.
- F. Lisdat, D. Utepbergenov, R. F. Haseloff, I. E. Blasig, W. Stöcklein, F. W. Scheller and R. Brigelius-Flohé, Anal. Chem., 2001, 73, 957–962 CrossRef CAS PubMed.
- S. M. Lato, Nucleic Acids Res., 2002, 30, 1401–1407 CrossRef CAS.
- G. Kolb, S. Reigadas, C. Boiziau, A. van Aerschot, A. Arzumanov, M. J. Gait, P. Herdewijn and J.-J. Toulmé, Biochemistry, 2005, 44, 2926–2933 CrossRef CAS PubMed.
- S. Hoshika, Nucleic Acids Res., 2004, 32, 3815–3825 CrossRef CAS PubMed.
- R. N. Veedu, B. Vester and J. Wengel, ChemBioChem, 2007, 8, 490–492 CrossRef CAS PubMed.
- C. G. Peng and M. J. Damha, Nucleic Acids Res., 2007, 35, 4977–4988 CrossRef CAS PubMed.
- M. Mehedi Masud, M. Kuwahara, H. Ozaki and H. Sawai, Bioorg. Med. Chem., 2004, 12, 1111–1120 CrossRef CAS PubMed.
- C. Di Primo, I. Rudloff, S. Reigadas, A. A. Arzumanov, M. J. Gait and J.-J. Toulmé, FEBS Lett., 2007, 581, 771–774 CrossRef CAS PubMed.
- E. S. Gragoudas, A. P. Adamis, E. T. Cunningham, M. Feinsod and D. R. Guyer, N. Engl. J. Med., 2004, 351, 2805–2816 CrossRef CAS PubMed.
- M. Zuker, Nucleic Acids Res., 2003, 31, 3406–3415 CrossRef CAS PubMed.
- I. L. Hofacker, Nucleic Acids Res., 2003, 31, 3429–3431 CrossRef CAS PubMed.
- O. Kikin, L. D'Antonio and P. S. Bagga, Nucleic Acids Res., 2006, 34, W676–W682 CrossRef CAS.
- M. van Dijk and A. M. J. J. Bonvin, Nucleic Acids Res., 2009, 37, W235–W239 CrossRef CAS PubMed.
- B. Giardine, C. Riemer, R. C. Hardison, R. Burhans, L. Elnitski, P. Shah, Y. Zhang, D. Blankenberg, I. Albert, J. Taylor, W. Miller, W. J. Kent and A. Nekrutenko, Genome Res., 2005, 15, 1451–1455 CrossRef CAS.
- R. Chen, L. Li and Z. Weng, Proteins: Struct., Funct., Bioinf., 2003, 52, 80–87 CrossRef CAS PubMed.
- A. Tovchigrechko and I. A. Vakser, Nucleic Acids Res., 2006, 34, W310–W314 CrossRef CAS PubMed.
- C. A. Rohl, C. E. M. Strauss, K. M. S. Misura and D. Baker, 2004, pp. 66–93.
- R. Das and D. Baker, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 14664–14669 CrossRef CAS PubMed.
- D. A. Case, T. E. Cheatham, T. Darden, H. Gohlke, R. Luo, K. M. Merz, A. Onufriev, C. Simmerling, B. Wang and R. J. Woods, J. Comput. Chem., 2005, 26, 1668–1688 CrossRef CAS PubMed.
- B. Hess, C. Kutzner, D. van der Spoel and E. Lindahl, J. Chem. Theory Comput., 2008, 4, 435–447 CrossRef CAS PubMed.
- J.-W. Park, R. Tatavarty, D. W. Kim, H.-T. Jung and M. B. Gu, Chem. Commun., 2012, 48, 2071–2073 RSC.
- R. Stoltenburg, N. Nikolaus and B. Strehlitz, J. Anal. Methods Chem., 2012, 2012, 1–14 CrossRef.
- T. Schütze, B. Wilhelm, N. Greiner, H. Braun, F. Peter, M. Mörl, V. A. Erdmann, H. Lehrach, Z. Konthur, M. Menger, P. F. Arndt and J. Glökler, PLoS One, 2011, 6, e29604 CrossRef.
- J. Hoinka and T. Przytycka, Methods, 2016, 106, 82–85 CrossRef CAS PubMed.
- J. Hoinka, E. Zotenko, A. Friedman, Z. E. Sauna and T. M. Przytycka, Bioinformatics, 2012, 28, i215–i223 CrossRef PubMed.
- J. Hoinka, A. Berezhnoy, Z. E. Sauna, E. Gilboa and T. M. Przytycka, AptaCluster - A Method to Cluster HT-SELEX Aptamer Pools and Lessons from its Application, 2014, pp. 115–128 Search PubMed.
- J. Hoinka, A. Berezhnoy, P. Dao, Z. E. Sauna, E. Gilboa and T. M. Przytycka, Nucleic Acids Res., 2015, 43, 5699–5707 CrossRef CAS.
- P. Dao, J. Hoinka, M. Takahashi, J. Zhou, M. Ho, Y. Wang, F. Costa, J. J. Rossi, R. Backofen, J. Burnett and T. M. Przytycka, Cell Syst., 2016, 3, 62–70 CrossRef CAS.
- J. Hoinka, R. Backofen and T. M. Przytycka, Mol. Ther.–Nucleic Acids, 2018, 11, 515–517 CrossRef CAS PubMed.
- K. K. Alam, J. L. Chang and D. H. Burke, Mol. Ther.–Nucleic Acids, 2015, 4, e230 CrossRef CAS PubMed.
- J. Caroli, C. Taccioli, A. De La Fuente, P. Serafini and S. Bicciato, Bioinformatics, 2016, 32, 161–164 CrossRef CAS PubMed.
- E. Afgan, D. Baker, M. van den Beek, D. Blankenberg, D. Bouvier, M. Čech, J. Chilton, D. Clements, N. Coraor, C. Eberhard, B. Grüning, A. Guerler, J. Hillman-Jackson, G. Von Kuster, E. Rasche, N. Soranzo, N. Turaga, J. Taylor, A. Nekrutenko and J. Goecks, Nucleic Acids Res., 2016, 44, W3–W10 CrossRef CAS PubMed.
- R. Lorenz, S. H. Bernhart, C. Höner zu Siederdissen, H. Tafer, C. Flamm, P. F. Stadler and I. L. Hofacker, Algorithms Mol. Biol., 2011, 6, 26 CrossRef PubMed.
- J. N. Zadeh, C. D. Steenberg, J. S. Bois, B. R. Wolfe, M. B. Pierce, A. R. Khan, R. M. Dirks and N. A. Pierce, J. Comput. Chem., 2011, 32, 170–173 CrossRef CAS PubMed.
- S. Bellaousov, J. S. Reuter, M. G. Seetin and D. H. Mathews, Nucleic Acids Res., 2013, 41, W471–W474 CrossRef PubMed.
- J.-M. Garant, J.-P. Perreault and M. S. Scott, Biochimie, 2018, 151, 115–118 CrossRef CAS PubMed.
- V. Brázda, J. Kolomazník, J. Lýsek, M. Bartas, M. Fojta, J. Šťastný and J.-L. Mergny, Bioinformatics, 2019, 35, 3493–3495 CrossRef.
- M. Biesiada, K. Pachulska-Wieczorek, R. W. Adamiak and K. J. Purzycka, Methods, 2016, 103, 120–127 CrossRef CAS PubMed.
- M. J. Boniecki, G. Lach, W. K. Dawson, K. Tomala, P. Lukasz, T. Soltysinski, K. M. Rother and J. M. Bujnicki, Nucleic Acids Res., 2016, 44, e63 CrossRef PubMed.
- L. P. P. Patro, A. Kumar, N. Kolimi and T. Rathinavelan, J. Mol. Biol., 2017, 429, 2438–2448 CrossRef CAS PubMed.
- G. Macindoe, L. Mavridis, V. Venkatraman, M.-D. Devignes and D. W. Ritchie, Nucleic Acids Res., 2010, 38, W445–W449 CrossRef CAS PubMed.
- S. J. de Vries, M. van Dijk and A. M. J. J. Bonvin, Nat. Protoc., 2010, 5, 883–897 CrossRef CAS PubMed.
- S. Cosconati, S. Forli, A. L. Perryman, R. Harris, D. S. Goodsell and A. J. Olson, Expert Opin. Drug Discovery, 2010, 5, 597–607 CrossRef CAS PubMed.
- O. Trott and A. J. Olson, J. Comput. Chem., 2010, 31, 455–461 CrossRef CAS PubMed.
- E. J. Denning, U. D. Priyakumar, L. Nilsson and A. D. Mackerell, J. Comput. Chem., 2011, 32, 1929–1943 CrossRef CAS PubMed.
- J. C. Wu, G. Chattree and P. Ren, Theor. Chem. Acc., 2012, 131, 1138 Search PubMed.
- Q. Yang, S.-P. Wang, X.-L. Yu, X.-H. Yang, Q.-P. Guo, L.-J. Tang, J.-H. Jiang and R.-Q. Yu, Chemom. Intell. Lab. Syst., 2017, 170, 32–37 CrossRef CAS.
- X. Liu, H. Yan, Y. Liu and Y. Chang, Small, 2011, 7, 1673–1682 CrossRef CAS PubMed.
- G. Zhu, L. Meng, M. Ye, L. Yang, K. Sefah, M. B. O'Donoghue, Y. Chen, X. Xiong, J. Huang, E. Song and W. Tan, Chem.–Asian J., 2012, 7, 1630–1636 CrossRef CAS PubMed.
- Y. Sakai, Md. S. Islam, M. Adamiak, S. C.-C. Shiu, J. A. Tanner and J. G. Heddle, Genes, 2018, 9, 571 CrossRef.
- C. Liang, F. Li, L. Wang, Z.-K. Zhang, C. Wang, B. He, J. Li, Z. Chen, A. B. Shaikh, J. Liu, X. Wu, S. Peng, L. Dang, B. Guo, X. He, D. W. T. Au, C. Lu, H. Zhu, B.-T. Zhang, A. Lu and G. Zhang, Biomaterials, 2017, 147, 68–85 CrossRef CAS PubMed.
- Y. Wen, H. Pei, Y. Wan, Y. Su, Q. Huang, S. Song and C. Fan, Anal. Chem., 2011, 83, 7418–7423 CrossRef CAS.
- H. Yu, S. Zhang and J. C. Chaput, Nat. Chem., 2012, 4, 183–187 CrossRef CAS PubMed.
- M. Kimoto, R. Yamashige, K. Matsunaga, S. Yokoyama and I. Hirao, Nat. Biotechnol., 2013, 31, 453–457 CrossRef CAS PubMed.
- K. Sefah, Z. Yang, K. M. Bradley, S. Hoshika, E. Jiménez, L. Zhang, G. Zhu, S. Shanker, F. Yu, D. Turek, W. Tan and S. A. Benner, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 1449–1454 CrossRef CAS.
- F. Tolle, G. M. Brändle, D. Matzner and G. Mayer, Angew. Chem., Int. Ed., 2015, 54, 10971–10974 CrossRef CAS PubMed.
- C. Guo, C. P. Watkins and R. Hili, J. Am. Chem. Soc., 2015, 137, 11191–11196 CrossRef CAS PubMed.
- H. Mei, J.-Y. Liao, R. M. Jimenez, Y. Wang, S. Bala, C. McCloskey, C. Switzer and J. C. Chaput, J. Am. Chem. Soc., 2018, 140, 5706–5713 CrossRef CAS PubMed.
- E. Eremeeva, A. Fikatas, L. Margamuljana, M. Abramov, D. Schols, E. Groaz and P. Herdewijn, Nucleic Acids Res., 2019, 47, 4927–4939 CrossRef CAS PubMed.
- L. Gold, D. Ayers, J. Bertino, C. Bock, A. Bock, E. N. Brody, J. Carter, A. B. Dalby, B. E. Eaton, T. Fitzwater, D. Flather, A. Forbes, T. Foreman, C. Fowler, B. Gawande, M. Goss, M. Gunn, S. Gupta, D. Halladay, J. Heil, J. Heilig, B. Hicke, G. Husar, N. Janjic, T. Jarvis, S. Jennings, E. Katilius, T. R. Keeney, N. Kim, T. H. Koch, S. Kraemer, L. Kroiss, N. Le, D. Levine, W. Lindsey, B. Lollo, W. Mayfield, M. Mehan, R. Mehler, S. K. Nelson, M. Nelson, D. Nieuwlandt, M. Nikrad, U. Ochsner, R. M. Ostroff, M. Otis, T. Parker, S. Pietrasiewicz, D. I. Resnicow, J. Rohloff, G. Sanders, S. Sattin, D. Schneider, B. Singer, M. Stanton, A. Sterkel, A. Stewart, S. Stratford, J. D. Vaught, M. Vrkljan, J. J. Walker, M. Watrobka, S. Waugh, A. Weiss, S. K. Wilcox, A. Wolfson, S. K. Wolk, C. Zhang and D. Zichi, PLoS One, 2010, 5, e15004 CrossRef CAS PubMed.
- https://clinicaltrials.gov/.
- J. Song, Y. Zheng, M. Huang, L. Wu, W. Wang, Z. Zhu, Y. Song and C. Yang, Anal. Chem., 2020, 92, 3307–3314 CrossRef CAS PubMed.
- R. Ishida, T. Adachi, A. Yokota, H. Yoshihara, K. Aoki, Y. Nakamura and M. Hamada, Nucleic Acids Res., 2020, 48, e82 CrossRef CAS PubMed.
- J. Caroli, M. Forcato and S. Bicciato, Bioinformatics, 2020, 36, 2266–2268 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C.-C. Hung, M. O'Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- M. Baek, R. McHugh, I. Anishchenko, H. Jiang, D. Baker and F. DiMaio, Nat. Methods, 2024, 21, 117–121 CrossRef CAS PubMed.
- K. Chen, Y. Zhou, S. Wang and P. Xiong, Proteins., 2023, 91(12), 1771–1778 CrossRef CAS PubMed.
- R. J. L. Townshend, S. Eismann, A. M. Watkins, R. Rangan, M. Karelina, R. Das and R. O. Dror, Science, 2021, 373, 1047–1051 CrossRef CAS PubMed.
- N. Emami and R. Ferdousi, Sci. Rep., 2021, 11, 6074 CrossRef CAS PubMed.
- N. Iwano, T. Adachi, K. Aoki, Y. Nakamura and M. Hamada, Nat. Comput. Sci., 2022, 2, 378–386 CrossRef PubMed.
- A. Bashir, Q. Yang, J. Wang, S. Hoyer, W. Chou, C. McLean, G. Davis, Q. Gong, Z. Armstrong, J. Jang, H. Kang, A. Pawlosky, A. Scott, G. E. Dahl, M. Berndl, M. Dimon and B. S. Ferguson, Nat. Commun., 2021, 12, 2366 CrossRef CAS PubMed.
- Y.-W. Cheung, P. Röthlisberger, A. E. Mechaly, P. Weber, F. Levi-Acobas, Y. Lo, A. W. C. Wong, A. B. Kinghorn, A. Haouz, G. P. Savage, M. Hollenstein and J. A. Tanner, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 16790–16798 CrossRef CAS PubMed.
- M. McKeague, V. Calzada, L. Cerchia, M. DeRosa, J. M. Heemstra, N. Janjic, P. E. Johnson, L. Kraus, J. Limson, G. Mayer, M. Nilsen Hamilton, D. Porciani, T. Kumar Sharma, B. Suess, J. A. Tanner and S. Shigdar, Aptamers, 2022, 6, 10–18 Search PubMed.
- E. J. Cho, J.-W. Lee and A. D. Ellington, Annu. Rev. Anal. Chem., 2009, 2, 241–264 CrossRef CAS PubMed.
- L. Santana-Viera, J. P. Dassie, M. Rosàs-Lapeña, S. Garcia-Monclús, M. Chicón-Bosch, M. Pérez-Capó, L. del Pozo, S. Sanchez-Serra, O. Almacellas-Rabaiget, S. Maqueda-Marcos, R. López-Alemany, W. H. Thiel, P. H. Giangrande and O. M. Tirado, Mol. Ther.–Nucleic Acids, 2023, 32, 758–772 CrossRef CAS PubMed.
- Y. Higashimoto, T. Matsui, Y. Nishino, J. Taira, H. Inoue, M. Takeuchi and S. Yamagishi, Microvasc. Res., 2013, 90, 64–70 CrossRef CAS PubMed.
- M. Legiewicz, C. Lozupone, R. Knight and M. Yarus, RNA, 2005, 11, 1701–1709 CrossRef CAS PubMed.
- A. Brown, J. Brill, R. Amini, C. Nurmi and Y. Li, Angew. Chem., 2024, 136(16), e202318665 CrossRef.
- M. Vorobyeva, A. Davydova, P. Vorobjev, D. Pyshnyi and A. Venyaminova, Int. J. Mol. Sci., 2018, 19, 470 CrossRef PubMed.
- K. Pobanz and A. Lupták, Methods, 2016, 106, 14–20 CrossRef CAS PubMed.
- R. Stoltenburg, N. Nikolaus and B. Strehlitz, J. Anal. Methods Chem., 2012, 2012, 1–14 CrossRef PubMed.
- J. Hoinka, R. Backofen and T. M. Przytycka, Mol. Ther.–Nucleic Acids, 2018, 11, 515–517 CrossRef CAS PubMed.
- A. V. Lakhin, V. Z. Tarantul and L. V. Gening, Acta Naturae, 2013, 5(4), 34–43 CrossRef CAS PubMed.
- R. Stoltenburg, C. Reinemann and B. Strehlitz, Anal. Bioanal. Chem., 2005, 383, 83–91 CrossRef CAS PubMed.
- A. S. R. Potty, K. Kourentzi, H. Fang, P. Schuck and R. C. Willson, Int. J. Biol. Macromol., 2011, 48, 392–397 CrossRef CAS PubMed.
- H. Ausserwöger, M. M. Schneider, T. W. Herling, P. Arosio, G. Invernizzi, T. P. J. Knowles and N. Lorenzen, Nat. Rev. Chem., 2022, 6, 844–861 CrossRef PubMed.
- A. Schonbrunn, Mol. Endocrinol., 2014, 28, 1403–1407 CrossRef PubMed.
- C. B. Saper, J. Histochem. Cytochem., 2009, 57, 1–5 CrossRef CAS PubMed.
- F. Darfeuille, Nucleic Acids Res., 2004, 32, 3101–3107 CrossRef CAS PubMed.
- N. A. Kolganova, A. M. Varizhuk, R. A. Novikov, V. L. Florentiev, G. E. Pozmogova, O. F. Borisova, A. K. Shchyolkina, I. P. Smirnov, D. N. Kaluzhny and E. N. Timofeev, Artif. DNA PNA XNA, 2014, 5, e28422 CrossRef PubMed.
- M. Kimoto, M. Nakamura and I. Hirao, Nucleic Acids Res., 2016, 44(15), 7487–7494 CAS.
- K. Matsunaga, M. Kimoto, C. Hanson, M. Sanford, H. A. Young and I. Hirao, Sci. Rep., 2015, 5, 18478 CrossRef CAS PubMed.
- D. Shangguan, Z. Tang, P. Mallikaratchy, Z. Xiao and W. Tan, ChemBioChem, 2007, 8, 603–606 CrossRef CAS PubMed.
- A. Pasternak, F. J. Hernandez, L. M. Rasmussen, B. Vester and J. Wengel, Nucleic Acids Res., 2011, 39, 1155–1164 CrossRef CAS PubMed.
- L. Zhang and J. C. Chaput, Molecules, 2020, 25, 4194 CrossRef CAS PubMed.
- N. Minakawa, M. Sanji, Y. Kato and A. Matsuda, Bioorg. Med. Chem., 2008, 16, 9450–9456 CrossRef CAS PubMed.
- H. Hoshino, Y. Kasahara, M. Kuwahara and S. Obika, J. Am. Chem. Soc., 2020, 142, 21530–21537 CrossRef CAS PubMed.
- Y.-W. Cheung, P. Röthlisberger, A. E. Mechaly, P. Weber, F. Levi-Acobas, Y. Lo, A. W. C. Wong, A. B. Kinghorn, A. Haouz, G. P. Savage, M. Hollenstein and J. A. Tanner, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 16790–16798 CrossRef CAS PubMed.
- Y. Kasahara, Y. Irisawa, H. Ozaki, S. Obika and M. Kuwahara, Bioorg. Med. Chem. Lett., 2013, 23, 1288–1292 CrossRef CAS PubMed.
- A. E. Rangel, Z. Chen, T. M. Ayele and J. M. Heemstra, Nucleic Acids Res., 2018, 46, 8057–8068 CrossRef CAS PubMed.
- C. Liu, C. Cozens, F. Jaziri, J. Rozenski, A. Maréchal, S. Dumbre, V. Pezo, P. Marlière, V. B. Pinheiro, E. Groaz and P. Herdewijn, J. Am. Chem. Soc., 2018, 140, 6690–6699 CrossRef CAS PubMed.
- M. R. Dunn, A. C. Larsen, W. J. Zahurancik, N. E. Fahmi, M. Meyers, Z. Suo and J. C. Chaput, J. Am. Chem. Soc., 2015, 137, 4014–4017 CrossRef CAS PubMed.
- D. S. Wilson and J. W. Szostak, Annu. Rev. Biochem., 1999, 68, 611–647 CrossRef CAS PubMed.
- G. F. Joyce, Annu. Rev. Biochem., 2004, 73, 791–836 CrossRef CAS PubMed.
- R. K. Saiki, D. H. Gelfand, S. Stoffel, S. J. Scharf, R. Higuchi, G. T. Horn, K. B. Mullis and H. A. Erlich, Science, 1988, 239, 487–491 CrossRef CAS PubMed.
- R. Stoltenburg, C. Reinemann and B. Strehlitz, Biomol. Eng., 2007, 24, 381–403 CrossRef CAS PubMed.
- M. U. Musheev and S. N. Krylov, Anal. Chim. Acta, 2006, 564, 91–96 CrossRef CAS PubMed.
- D. Aird, M. G. Ross, W.-S. Chen, M. Danielsson, T. Fennell, C. Russ, D. B. Jaffe, C. Nusbaum and A. Gnirke, Genome Biol., 2011, 12, R18 CrossRef CAS PubMed.
- I. P. Oscorbin and M. L. Filipenko, Comput. Struct. Biotechnol. J., 2021, 19, 6315–6327 CrossRef CAS PubMed.
- T. R. Battersby, D. N. Ang, P. Burgstaller, S. C. Jurczyk, M. T. Bowser, D. D. Buchanan, R. T. Kennedy and S. A. Benner, J. Am. Chem. Soc., 1999, 121, 9781–9789 CrossRef CAS PubMed.
- K. Matsunaga, M. Kimoto and I. Hirao, J. Am. Chem. Soc., 2017, 139, 324–334 CrossRef CAS PubMed.
- J. A. Latham, R. Johnson and J. J. Toole, Nucleic Acids Res., 1994, 22, 2817–2822 CrossRef CAS PubMed.
- K. Ohsawa, T. Kasamatsu, J. Nagashima, K. Hanawa, M. Kuwahara, H. Ozaki and H. Sawai, Anal. Sci., 2008, 24, 167–172 CrossRef CAS PubMed.
- A. Shoji, M. Kuwahara, H. Ozaki and H. Sawai, J. Am. Chem. Soc., 2007, 129, 1456–1464 CrossRef CAS PubMed.
- J. D. Vaught, C. Bock, J. Carter, T. Fitzwater, M. Otis, D. Schneider, J. Rolando, S. Waugh, S. K. Wilcox and B. E. Eaton, J. Am. Chem. Soc., 2010, 132, 4141–4151 CrossRef CAS PubMed.
- Y. Higashimoto, T. Matsui, Y. Nishino, J. Taira, H. Inoue, M. Takeuchi and S. Yamagishi, Microvasc. Res., 2013, 90, 64–70 CrossRef CAS PubMed.
- J. Kang, M. S. Lee, J. A. Copland, B. A. Luxon and D. G. Gorenstein, Bioorg. Med. Chem. Lett., 2008, 18, 1835–1839 CrossRef CAS PubMed.
- P. E. Burmeister, S. D. Lewis, R. F. Silva, J. R. Preiss, L. R. Horwitz, P. S. Pendergrast, T. G. McCauley, J. C. Kurz, D. M. Epstein, C. Wilson and A. D. Keefe, Chem. Biol., 2005, 12(1), 25–33 CrossRef CAS PubMed.
- E. Medina, E. J. Yik, P. Herdewijn and J. C. Chaput, ACS Synth. Biol., 2021, 10, 1429–1437 CrossRef CAS PubMed.
- C. Liu, C. Cozens, F. Jaziri, J. Rozenski, A. Maréchal, S. Dumbre, V. Pezo, P. Marlière, V. B. Pinheiro, E. Groaz and P. Herdewijn, J. Am. Chem. Soc., 2018, 140, 6690–6699 CrossRef CAS PubMed.
- H. Hoshino, Y. Kasahara, H. Fujita, M. Kuwahara, K. Morihiro, S. Tsunoda and S. Obika, Bioorg. Med. Chem. Lett., 2016, 26, 530–533 CrossRef CAS PubMed.
- C. Cozens, V. B. Pinheiro, A. Vaisman, R. Woodgate and P. Holliger, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 8067–8072 CrossRef CAS PubMed.
- A. I. Taylor and P. Holliger, Nat. Protoc., 2015, 10, 1625–1642 CrossRef CAS PubMed.
- H. Yu, S. Zhang, M. R. Dunn and J. C. Chaput, J. Am. Chem. Soc., 2013, 135, 3583–3591 CrossRef CAS PubMed.
- A. E. Rangel, Z. Chen, T. M. Ayele and J. M. Heemstra, Nucleic Acids Res., 2018, 46, 8057–8068 CrossRef CAS PubMed.
- M. R. Dunn, R. M. Jimenez and J. C. Chaput, Nat. Rev. Chem., 2017, 1, 0076 CrossRef CAS.
- A. C. Larsen, M. R. Dunn, A. Hatch, S. P. Sau, C. Youngbull and J. C. Chaput, Nat. Commun., 2016, 7, 11235 CrossRef CAS PubMed.
- N. Komarova and A. Kuznetsov, Molecules, 2019, 24, 3598 CrossRef PubMed.
- A. Lozoya-Colinas, Y. Yu and J. C. Chaput, J. Am. Chem. Soc., 2023, 145, 25789–25796 CrossRef CAS PubMed.
- N. Kundu, C. M. McCloskey, M. Hajjar and J. C. Chaput, Biochemistry, 2023, 62, 3245–3254 CrossRef CAS PubMed.
- G. J. Jaffe, K. Westby, K. G. Csaky, J. Monés, J. A. Pearlman, S. S. Patel, B. C. Joondeph, J. Randolph, H. Masonson and K. A. Rezaei, Ophthalmology, 2021, 128, 576–586 CrossRef PubMed.
- Z. Liu, T. Chen and F. E. Romesberg, Chem. Sci., 2017, 8, 8179–8182 RSC.
- K. Son, J. M. Gibson, J. Gerson, K. K. Leung, M. Stocco, Y. Lu, T. E. Kippin and K. W. Plaxco, J. Am. Chem. Soc., 2026, 148(9), 10016–10025 CrossRef PubMed.
- B. J. Hicke, C. Marion, Y.-F. Chang, T. Gould, C. K. Lynott, D. Parma, P. G. Schmidt and S. Warren, J. Biol. Chem., 2001, 276, 48644–48654 CrossRef CAS PubMed.
- C. P. Rusconi, E. Scardino, J. Layzer, G. A. Pitoc, T. L. Ortel, D. Monroe and B. A. Sullenger, Nature, 2002, 419, 90–94 CrossRef CAS PubMed.
- L. Gold, D. Ayers, J. Bertino, C. Bock, A. Bock, E. Brody, J. Carter, V. Cunningham, A. Dalby, B. Eaton, T. Fitzwater, D. Flather, A. Forbes, T. Foreman, C. Fowler, B. Gawande, M. Goss, M. Gunn, S. Gupta, D. Halladay, J. Heil, J. Heilig, B. Hicke, G. Husar, N. Janjic, T. Jarvis, S. Jennings, E. Katilius, T. Keeney, N. Kim, T. Kaske, T. Koch, S. Kraemer, L. Kroiss, N. Le, D. Levine, W. Lindsey, B. Lollo, W. Mayfield, M. Mehan, R. Mehler, M. Nelson, S. Nelson, D. Nieuwlandt, M. Nikrad, U. Ochsner, R. Ostroff, M. Otis, T. Parker, S. Pietrasiewicz, D. Resnicow, J. Rohloff, G. Sanders, S. Sattin, D. Schneider, B. Singer, M. Stanton, A. Sterkel, A. Stewart, S. Stratford, J. Vaught, M. Vrkljan, J. Walker, M. Watrobka, S. Waugh, A. Weiss, S. Wilcox, A. Wolfson, S. Wolk, C. Zhang and D. Zichi, Nat. Preced., 2010 DOI:10.1038/npre.2010.4538.1.
- Y. Kasahara, Y. Irisawa, H. Fujita, A. Yahara, H. Ozaki, S. Obika and M. Kuwahara, Anal. Chem., 2013, 85, 4961–4967 CrossRef CAS PubMed.
- M. Kuwahara and S. Obika, Artif. DNA PNA XNA, 2013, 4, 39–48 CrossRef PubMed.
- K. P. Williams, X.-H. Liu, T. N. M. Schumacher, H. Y. Lin, D. A. Ausiello, P. S. Kim and D. P. Bartel, Proc. Natl. Acad. Sci. U. S. A., 1997, 94, 11285–11290 CrossRef CAS PubMed.
- J. T. Sczepanski and G. F. Joyce, J. Am. Chem. Soc., 2013, 135, 13290–13293 CrossRef CAS PubMed.
- B. Wlotzka, S. Leva, B. Eschgfäller, J. Burmeister, F. Kleinjung, C. Kaduk, P. Muhn, H. Hess-Stumpp and S. Klussmann, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 8898–8902 CrossRef CAS PubMed.
- A. Vater, F. Jarosch, W. Purschke, R. Reutter, C. Maasch, K. Buchner, D. Eulberg, M. Huber-Lang and S. Klussmann, Mol. Immunol., 2010, 47, 2290 CrossRef.
- K. Hoehlig, C. Maasch, N. Shushakova, K. Buchner, M. Huber-Lang, W. G. Purschke, A. Vater and S. Klussmann, Mol. Ther., 2013, 21, 2236–2246 CrossRef CAS PubMed.
- D. Faulhammer, B. Eschgfäller, S. Stark, P. Burgstaller, W. Englberger, J. Erfurth, F. Kleinjung, J. RUPP, S. D. Vulcu, W. Schröder, S. Vonhoff, H. Nawrath, C. Gillen and S. Klussmann, RNA, 2004, 10, 516–527 CrossRef CAS PubMed.
- S. Jhaveri, B. Olwin and A. D. Ellington, Bioorg. Med. Chem. Lett., 1998, 8, 2285–2290 CrossRef CAS PubMed.
- J. Kang, M. S. Lee, S. J. Watowich and D. G. Gorenstein, FEBS Lett., 2007, 581, 2497–2502 CrossRef CAS PubMed.
- D. J. King, J. G. Safar, G. Legname and S. B. Prusiner, J. Mol. Biol., 2007, 369, 1001–1014 CrossRef CAS PubMed.
- F. Leonard, N. P. Ha, P. Sule, J. F. Alexander, D. E. Volk, G. L. R. Lokesh, X. Liu, J. D. Cirillo, D. G. Gorenstein, J. Yuan, S. Chatterjee, E. A. Graviss and B. Godin, J. Controlled Release, 2017, 266, 238–247 CrossRef CAS PubMed.
- M. R. Dunn and J. C. Chaput, ChemBioChem, 2016, 17, 1804–1808 CrossRef CAS PubMed.
- M. R. Dunn, C. M. McCloskey, P. Buckley, K. Rhea and J. C. Chaput, J. Am. Chem. Soc., 2020, 142, 7721–7724 CrossRef CAS PubMed.
- C. Gasse, M. Zaarour, S. Noppen, M. Abramov, P. Marlière, S. Liekens, B. De Strooper and P. Herdewijn, ChemBioChem, 2018, 19, 754–763 CrossRef CAS PubMed.
- J. A. Latham, R. Johnson and J. J. Toole, Nucleic Acids Res., 1994, 22, 2817–2822 CrossRef CAS PubMed.
- M. Renders, E. Miller, C. H. Lam and D. M. Perrin, Org. Biomol. Chem., 2017, 15, 1980–1989 RSC.
- T. R. Battersby, D. N. Ang, P. Burgstaller, S. C. Jurczyk, M. T. Bowser, D. D. Buchanan, R. T. Kennedy and S. A. Benner, J. Am. Chem. Soc., 1999, 121, 9781–9789 CrossRef CAS PubMed.
- S. Arangundy-Franklin, A. I. Taylor, B. T. Porebski, V. Genna, S. Peak-Chew, A. Vaisman, R. Woodgate, M. Orozco and P. Holliger, Nat. Chem., 2019, 11, 533–542 CrossRef CAS PubMed.
- S. Ni, H. Yao, L. Wang, J. Lu, F. Jiang, A. Lu and G. Zhang, Int. J. Mol. Sci., 2017, 18, 1683 CrossRef PubMed.
- F. Tolle, G. M. Brändle, D. Matzner and G. Mayer, Angew. Chem., Int. Ed., 2015, 54, 10971–10974 CrossRef CAS PubMed.
- Q. Shao, T. Chen, K. Sheng, Z. Liu, Z. Zhang and F. E. Romesberg, J. Am. Chem. Soc., 2020, 142, 2125–2128 CrossRef CAS PubMed.
- J. S. Temme, I. S. MacPherson, J. F. DeCourcey and I. J. Krauss, J. Am. Chem. Soc., 2014, 136, 1726–1729 CrossRef CAS PubMed.
- S. Horiya, J. K. Bailey, J. S. Temme, Y. V. Guillen Schlippe and I. J. Krauss, J. Am. Chem. Soc., 2014, 136, 5407–5415 CrossRef CAS PubMed.
- K. D. Kovacevic, J. C. Gilbert and B. Jilma, Adv. Drug Delivery Rev., 2018, 134, 36–50 CrossRef CAS PubMed.
- S. Gupta, D. W. Drolet, S. K. Wolk, S. M. Waugh, J. C. Rohloff, J. D. Carter, W. S. Mayfield, M. R. Otis, C. R. Fowler, T. Suzuki, M. Hirota, Y. Ishikawa, D. J. Schneider and N. Janjic, Nucleic Acid Ther., 2017, 27, 345–353 CrossRef CAS PubMed.
- M. Shadid, M. Badawi and A. Abulrob, Expert Opin. Drug Metab. Toxicol., 2021, 17, 1281–1292 CrossRef CAS PubMed.
- J. M. Migliorati, S. Liu, A. Liu, A. Gogate, S. Nair, R. Bahal, T. P. Rasmussen, J. E. Manautou and X. Zhong, Drug Metab. Dispos., 2022, 50, 888–897 CrossRef CAS PubMed.
- Y. El Abd, A. Tabll, R. Smolic and M. Smolic, Hum. Antibodies, 2022, 30, 15–24 CAS.
- European Medicines Agency, Human Medicines Highlights 2022, 2022 Search PubMed.
- H. M. Berman, Nucleic Acids Res., 2000, 28, 235–242 CrossRef CAS PubMed.
- R. Troisi, N. Balasco, I. Autiero, L. Vitagliano and F. Sica, Int. J. Mol. Sci., 2023, 24, 16318 CrossRef CAS PubMed.
- H. Su, W. Wang, Z. Du, Z. Peng, S. Gao, M. Cheng and J. Yang, Adv. Sci., 2021, 8(24), 2102592 CrossRef CAS PubMed.
- Z. Peng, W. Wang, H. Wei, X. Li and J. Yang, Proteins: Struct., Funct., Bioinf., 2023, 91, 1704–1711 CrossRef CAS PubMed.
- R. Das, R. C. Kretsch, A. J. Simpkin, T. Mulvaney, P. Pham, R. Rangan, F. Bu, R. M. Keegan, M. Topf, D. J. Rigden, Z. Miao and E. Westhof, Proteins: Struct., Funct., Bioinf., 2023, 91, 1747–1770 CrossRef CAS PubMed.
- Y. El Abd, A. Tabll, R. Smolic and M. Smolic, Hum. Antibodies, 2022, 30, 15–24 CAS.
- Y. Aslan, M. Atabay, H. K. Chowdhury, I. Göktürk, Y. Saylan and F. Inci, Biosensors, 2023, 13, 569 CrossRef CAS PubMed.
- H. Kaur, J. G. Bruno, A. Kumar and T. K. Sharma, Theranostics, 2018, 8, 4016–4032 CrossRef CAS PubMed.
- J. He, Q. Duan, C. Ran, T. Fu, Y. Liu and W. Tan, Acta Pharm. Sin. B, 2023, 13, 1358–1370 CrossRef CAS PubMed.
- B. Schneider, B. A. Sweeney, A. Bateman, J. Cerny, T. Zok and M. Szachniuk, Nucleic Acids Res., 2023, 51, 9522–9532 CrossRef PubMed.
- J. Hennig, ChemBioChem, 2025, 26(7), 1439–4227 CrossRef PubMed.
- M. J. Forster, Micron, 2002, 33, 365–384 CrossRef CAS PubMed.
- K. Liebl and M. Zacharias, Biophys. J., 2023, 122, 2841–2851 CrossRef CAS PubMed.
- O. Love, R. Galindo-Murillo, M. Zgarbová, J. Šponer, P. Jurečka and T. E. Cheatham, J. Chem. Theory Comput., 2023, 19, 4299–4307 CrossRef CAS PubMed.
- M. Zgarbová, M. Otyepka, J. Šponer, A. Mládek, P. Banáš, T. E. Cheatham and P. Jurečka, J. Chem. Theory Comput., 2011, 7, 2886–2902 CrossRef PubMed.
- C.-A. Mattelaer, H.-P. Mattelaer, J. Rihon, M. Froeyen and E. Lescrinier, J. Chem. Theory Comput., 2021, 17, 3814–3823 CrossRef CAS PubMed.
- J. Rihon, S. Reynders, V. Bernardes Pinheiro and E. Lescrinier, J. Cheminf., 2025, 17, 53 Search PubMed.
- M. F. Sanner, A. J. Olson and J.-C. Spehner, Biopolymers, 1996, 38, 305–320 CrossRef CAS PubMed.
- R. M. Betz and R. C. Walker, J. Comput. Chem., 2015, 36, 79–87 CrossRef CAS PubMed.
- J. Rihon, C.-A. Mattelaer, R. W. Montalvão, M. Froeyen, V. B. Pinheiro and E. Lescrinier, Nucleic Acids Res., 2024, 52, 2836–2847 CrossRef CAS PubMed.
- C. I. Bayly, P. Cieplak, W. Cornell and P. A. Kollman, J. Phys. Chem., 1993, 97, 10269–10280 CrossRef CAS.
- S. Reynders, J. Rihon and E. Lescrinier, J. Chem. Theory Comput., 2025, 21, 2798–2814 CrossRef CAS PubMed.
- O. Love, R. Galindo-Murillo, D. R. Roe, P. D. Dans, T. E. Cheatham III and C. Bergonzo, J. Chem. Theory Comput., 2024, 20, 9354–9363 CrossRef CAS PubMed.
- S. Kirkpatrick, C. D. Gelatt and M. P. Vecchi, Science, 1983, 220, 671–680 CrossRef CAS PubMed.
- Y. Sugita and Y. Okamoto, Chem. Phys. Lett., 1999, 314, 141–151 CrossRef CAS.
- P. Schofield, A. I. Taylor, J. Rihon, C. D. Peña Martinez, S. Zinn, C.-A. Mattelaer, J. Jackson, G. Dhaliwal, G. Schepers, P. Herdewijn, E. Lescrinier, D. Christ and P. Holliger, Nucleic Acids Res., 2023, 51, 7736–7748 CrossRef CAS PubMed.
- M. Kohlberger and G. Gadermaier, Biotechnol. Appl. Biochem., 2022, 69, 1771–1792 CrossRef CAS PubMed.
- J. C. Cox, Nucleic Acids Res., 2002, 30, 108e–1108 CrossRef PubMed.
- R. Kirby, E. J. Cho, B. Gehrke, T. Bayer, Y. S. Park, D. P. Neikirk, J. T. McDevitt and A. D. Ellington, Anal. Chem., 2004, 76, 4066–4075 CrossRef CAS PubMed.
- D. T. Tran, K. P. F. Janssen, J. Pollet, E. Lammertyn, J. Anné, A. Van Schepdael and J. Lammertyn, Molecules, 2010, 15, 1127–1140 CrossRef CAS PubMed.
- L. C. Bock, L. C. Griffin, J. A. Latham, E. H. Vermaas and J. J. Toole, Nature, 1992, 355, 564–566 CrossRef CAS PubMed.
- L. C. Griffin, J. J. Toole and L. L. K. Leung, Gene, 1993, 137, 25–31 CrossRef CAS PubMed.
- F. Bracht and K. Schror, Biochem. Biophys. Res. Commun., 1994, 200, 933–937 CrossRef CAS PubMed.
- L. R. Paborsky, S. N. McCurdy, L. C. Griffin, J. J. Toole and L. L. Leung, J. Biol. Chem., 1993, 268, 20808–20811 CrossRef CAS PubMed.
- H. A. Levine and M. Nilsen-Hamilton, Comput. Biol. Chem., 2007, 31, 11–35 CrossRef CAS PubMed.
- M. McKeague, E. M. McConnell, J. Cruz-Toledo, E. D. Bernard, A. Pach, E. Mastronardi, X. Zhang, M. Beking, T. Francis, A. Giamberardino, A. Cabecinha, A. Ruscito, R. Aranda-Rodriguez, M. Dumontier and M. C. DeRosa, J. Mol. Evol., 2015, 81(5–6), 150–161 CrossRef CAS PubMed.
- M. Legiewicz, C. Lozupone, R. Knight and M. Yarus, RNA, 2005, 11(11), 1701–1709 CrossRef CAS PubMed.
- L. Chen, B. Zhang, Z. Wu, G. Liu, W. Li and Y. Tang, Comput. Struct. Biotechnol. J., 2023, 21, 1005–1013 CrossRef CAS PubMed.
- S. J. Lee, J. Cho, B.-H. Lee, D. Hwang and J.-W. Park, Biomedicines, 2023, 11, 356 CrossRef CAS PubMed.
- J. Rihon, C.-A. Mattelaer, R. W. Montalvão, M. Froeyen, V. B. Pinheiro and E. Lescrinier, Nucleic Acids Res., 2024, 52(6), 2836–2847 CrossRef CAS PubMed.
- F. Bottari, E. Daems, A.-M. de Vries, P. Van Wielendaele, S. Trashin, R. Blust, F. Sobott, A. Madder, J. C. Martins and K. De Wael, J. Am. Chem. Soc., 2020, 142, 19622–19630 CrossRef CAS PubMed.
- B. E. Eaton, L. Gold and D. A. Zichi, Chem. Biol., 1995, 2, 633–638 CrossRef CAS PubMed.
- J. M. Carothers, S. C. Oestreich and J. W. Szostak, J. Am. Chem. Soc., 2006, 128, 7929–7937 CrossRef CAS PubMed.
- J. Hoetzel and B. Suess, J. Mol. Biol., 2022, 434, 167631 CrossRef CAS PubMed.
- N. Nakatsuka, A. Faillétaz, D. Eggemann, C. Forró, J. Vörös and D. Momotenko, Anal. Chem., 2021, 93, 4033–4041 CrossRef CAS PubMed.
- H. Kawabe, C. A. Thomas, S. Hoshika, M.-J. Kim, M.-S. Kim, L. Miessner, N. Kaplan, J. M. Craig, J. H. Gundlach, A. H. Laszlo, S. A. Benner and J. A. Marchand, Nat. Commun., 2023, 14, 6820 CrossRef CAS PubMed.
- D. Kestemont, M. Renders, P. Leonczak, M. Abramov, G. Schepers, V. B. B. Pinheiro, J. Rozenski and P. Herdewijn, Chem. Commun., 2018,(49), 6408–6411 RSC.
- M. McKeague, V. Calzada, L. Cerchia, M. DeRosa, J. M. Heemstra, N. Janjic, P. E. Johnson, L. Kraus, J. Limson, G. Mayer, M. Nilsen-Hamilton, D. Porciani, T. K. Sharma, B. Suess, J. A. Tanner and S. Shigdar, Aptamers, 2022, 6, 10–18 Search PubMed.
Footnote |
| † Both authors contributed equally to the work. |
| This journal is © The Royal Society of Chemistry 2026 |