Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Structural insights into ortho-aminophenol oxidases: kinetic and crystallographic characterization of SmNspF and SgGriF
Hoa
Le Xuan
 ab and
Annette
Rompel
ab and
Annette
Rompel
 *ab
*ab
aUniversität Wien, Fakultät für Chemie, Institut für Biophysikalische Chemie, Josef-Holaubek Platz 2, 1090 Vienna, Austria. E-mail: annette.rompel@univie.ac.at; Web: https://www.bpc.univie.ac.at
bUniversität Wien, Vienna Doctoral School in Chemistry (DoSChem), Währinger Straße 42, 1090 Vienna, Austria Web: https://doschem.univie.ac.at
First published on 30th January 2026
Abstract
Actinobacteria-derived o-aminophenol oxidases (AOs) represent a largely unexplored subclass of type-III copper enzymes with catalytic properties distinct from tyrosinases and catechol oxidases. The determination of the first crystal structure of an AO (SmNspF) displays unique loop insertions and important second-sphere amino acids in vicinity of the binuclear copper center. The substrate-guiding effect of the second activity controller (HisB2+1) influences the binding affinity for carboxyl-containing substrates in the AOs SmNspF and SgGriF. Thus, kinetic investigations reveal both overlapping and distinct substrate preferences for SmNspF and SgGriF: while both enzymes oxidize monophenols, o-aminophenols, and o-diphenols, they do so at significantly different reaction rates. SmNspF preferentially oxidizes carboxylated substrates such as 3,4-dihydroxybenzoic acid and 3-amino-4-hydroxybenzoic acid, whereas SgGriF exhibits higher activity toward para-methylated analogs, including 4-methylcatechol and 2-amino-4-methylphenol. Remarkably, both enzymes display enzymatic activities beyond the known AO reactivity spectrum by oxidizing 2-aminoresorcinol and o-phenylenediamine, which underlies the high versatility of the binuclear copper center. Altogether, these findings provide a structural basis for AO's enzymatic activity and broaden the known catalytic spectrum, which enables the prediction of catalytic properties in type-III copper proteins based on their amino acid sequence.
Introduction
Actinobacteria of the genus Streptomyces are renowned for their exceptional metabolic diversity and their ability to produce a wide range of bioactive natural products, including antibiotics, antifungals, immunosuppressants, antiparasitic agents and pigments.1–5 One example from this genus is Streptomyces murayamaensis, which produces the enzyme o-aminophenol oxidase (AO) SmNspF (Uniprot ID: D6RTB9), and another example is Streptomyces griseus, which produces the AO SgGriF (Uniprot ID: B1VTI5).6,7o-Aminophenol oxidases (AOs; EC 1.10.3.4) belong to the type-III copper enzyme family, a widespread group of metalloenzymes characterized by a binuclear copper center, each coordinated by three histidine residues. Type-III copper enzymes occur in all domains of life, spanning archaea, bacteria, fungi, plants, and animals.8–11 In all subclasses of type-III copper proteins, the active site is proposed to bind molecular oxygen in a symmetric side-on bridging configuration (μ-η2:η2), which facilitates the activation of O2 for subsequent substrate oxidation.12–14 The best-characterized representatives of type-III copper proteins include catechol oxidases (EC 1.10.3.1), which oxidize o-diphenols to o-quinones, and tyrosinases (TYR; EC 1.14.18.1), which possess both monophenolase activity (hydroxylation of monophenols to o-diphenols) and diphenolase activity (Fig. S1).12,13 The relatively underexplored AOs have been reported to exhibit both monophenolase and diphenolase activities, suggesting enzymatic overlap with TYRs.14 However, AOs uniquely catalyze the oxidation of o-aminophenols to nitrosophenols (nitroso-forming activity), a reaction not observed in catechol oxidases or TYRs (Fig. S1).11,14 The structural basis for the functional divergence between AOs and TYRs has been attributed to a key active-site residue (activity selector) with AOs featuring a conserved asparagine and TYRs a conserved tyrosine at a position near the active center.15 Substituting Asn43 in the SgGriF with the isoleucine present at the homologous position in TYR SzTYR, a tyrosinase originated from Streptomyces sp. ZL-24, abolishes the nitroso-forming activity, whereas introducing an asparagine in place of the homologous isoleucine in SzTYRs confers this function.15Deciphering the structural basis for the diverse substrate specificities among the wide variety of type-III copper enzymes present in nature has long been a challenge. Within this report, we aim to investigate the enzymatic divergence within the AO subclass by characterizing the catalytic activity of SmNspF and SgGriF, to evaluate the potential of AOs to oxidize a broad range of small phenolic and o-aminophenolic compounds, as well as non-phenolic substrates such as aromatic amines and diamines. To achieve a comprehensive functional and structural understanding of this largely unstudied enzyme subclass, we further determine, for the first time, the three-dimensional structure of an AO, confirming the coupled binuclear copper site and elucidating the role of second sphere residues in comparison to bacterial and plant TYR crystal structures. These insights support progress in green biotechnology by promoting selective catalytic systems that avoid harsh oxidants and energy-intensive processes and utilize biocatalysts containing Earth-abundant metals rather than rare or precious ones.
Results and discussion
Expression and purification
To achieve maximum yields and purity of SmNspF and SgGriF, expression conditions were examined by adjusting the medium composition, specifically the NaCl and CuSO4 concentrations to support proper copper ion incorporation into the type-III copper center, as well as by optimizing the incubation temperature and incubation time (for detailed description of the protein expressions see SI Section S1.2). Heterologous expression using a lac-operon expression system and a GST-tag for affinity purification resulted in much higher expression efficiency for SmNspF (yield ≈ 3.1 mg l−1 medium) compared to SgGriF (yield ≈ 0.8 mg l−1 medium) (Table S2). SmNspF also demonstrates higher purification efficiency than SgGriF following the first affinity chromatography step (Table S2, Fig. S2). Additionally, SgGriF requires an extra size-exclusion chromatography step (with a Superdex 75 10/300 GL; GE, Boston, MA, USA) to achieve purity levels higher than 95% (Table S2). The successful expression of purified SmNspF and SgGriF is verified by protein analysis with SDS-PAGE and intact protein mass spectrometry (SI Section S1.3, S1.4 and Fig. S4–S6). ESI-Orbitrap-MS revealed molecular masses of 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 837.41 ± 1.00 Da for SmNspF and 35
837.41 ± 1.00 Da for SmNspF and 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 736.16 ± 1.18 Da for SgGriF, which matches well with the calculated mass for the recombinantly expressed enzymes (SmNspF: 35
736.16 ± 1.18 Da for SgGriF, which matches well with the calculated mass for the recombinantly expressed enzymes (SmNspF: 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 838.36 Da, SgGriF: 35
838.36 Da, SgGriF: 35![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 737.28 Da). The successful incorporation of copper ions into the type-III copper center was confirmed by copper content analysis using 2,2′biquinoline, which revealed copper-to-enzyme ratios of 1.6
737.28 Da). The successful incorporation of copper ions into the type-III copper center was confirmed by copper content analysis using 2,2′biquinoline, which revealed copper-to-enzyme ratios of 1.6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for SmNspF and 1.5
1 for SmNspF and 1.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 for SgGriF.16 UV-VIS spectroscopic investigation of SgGriF after the addition of a 20-fold molar excess of hydroxylamine to an oxygen-saturated solution reveals the emergence of a band in the 325–355 nm range, indicating the formation of a metal–ligand charge transfer complex based on the met and/or oxy form (Fig. S7).17,18 The 280/345 nm absorption ratio of SgGriF is approximately 0.11, which is comparable to the ratio observed for SmNspF (approximately 0.12), as reported by Ginsbach et al. (2012), also following the addition of a 20-fold molar excess of hydroxylamine.14 Hydroxylamine reduces the resting met-form of the binuclear copper center, resulting in the formation of the deoxy-form, which enables dioxygen binding to generate the catalytically active oxy-form.14
1 for SgGriF.16 UV-VIS spectroscopic investigation of SgGriF after the addition of a 20-fold molar excess of hydroxylamine to an oxygen-saturated solution reveals the emergence of a band in the 325–355 nm range, indicating the formation of a metal–ligand charge transfer complex based on the met and/or oxy form (Fig. S7).17,18 The 280/345 nm absorption ratio of SgGriF is approximately 0.11, which is comparable to the ratio observed for SmNspF (approximately 0.12), as reported by Ginsbach et al. (2012), also following the addition of a 20-fold molar excess of hydroxylamine.14 Hydroxylamine reduces the resting met-form of the binuclear copper center, resulting in the formation of the deoxy-form, which enables dioxygen binding to generate the catalytically active oxy-form.14
Kinetic investigation
The monophenolase activity of SgGriF was investigated by Suzuki et al. (2006)19 using L-tyrosine, 4-hydroxybenzaldehyde and phenol, with no activity observed. In our study, both enzymes show no visible monophenolase activity toward tyramine but exhibit slow enzymatic activity toward the monophenol 4-methylphenol (Fig. 1, columns 4, 5, 8 and 9).6,14,19 Notably, the addition of 0.5 mM hydroxylamine accelerated the oxidation of 4-methylphenol with both AOs, as indicated by a slight color change from colorless to pinkish observed within 3 min of reaction time (Fig. 1, i column 8). However, enzymatic oxidation of 4-methylphenol after three hours of reaction time was clearly observed with either SmNspF or SgGriF, both in the presence and absence of hydroxylamine. This suggests the existence of the resting as well as deoxy-form after this enzyme purification process, which includes the elution of GST-tagged enzymes with reduced glutathione during the first GST-tag affinity chromatography step (Fig. S2). The presence of the enzyme in its deoxy-form is essential for initiating the monophenolase reaction cycle of type-III copper enzymes.20With regard to diphenolase activity, both the met and oxy forms can catalyze o-diphenol oxidation according to the reaction mechanism published in ref. 20. SmNspF and SgGriF show rapid diphenolase activity toward 4-methylcatechol (Fig. 1, columns 6 and 7; Table 1). Notably, SgGriF displays higher catalytic efficiency for 4-methylcatechol compared to SmNspF, indicated by a more intense color change and an approximately 2-fold higher kcat/Km ratio (Fig. 1, columns 6 and 7; Table 1). Both AOs show diphenolase activity towards catechol, dopamine, 4-methylcatechol and 3,4-dihydroxybenzoic acid (3,4DHBA) (Fig. 1, columns 6 and 7; Table 1). However, no activity is observed with DOPAC.
| Substrate | AO | K m [mM] | k cat [s−1] | k cat/Km [s−1 m M−1] |
|---|---|---|---|---|
| Experimental details are described in SI Section S1.7 and Fig. S12–14. SmNspF in the literature with Km values [mM]: 4-methylphenol = 0.70 ± 0.04 (O2 consumption at 298 K), catechol = no data, 3,4DHBA = no data, 3A4HBA = 4.1 ± 0.6, 2AP = 11 ± 4; kcat values [s−1]: 4-methylphenol = 2.22 ± 0.06 (O2 consumption at 298 K), catechol = no data, 3,4DHBA = no data, 3A4HBA = 3.5 ± 0.2, 2AP = 120 ± 30.6,14SgGriF in the literature with Km values [mM]: 4-methylphenol = no data, catechol = 2.5 ± 0.6/19 ± 2, 3,4DHBA = no data, 3,4-dihydroxybenzaldehyde = 0.41 ± 0.06, L-DOPA = 5.5 ± 1.8, 3A4HBA = no data, 2AP = 6.7 ± 1.5/3.5 ± 0.6, 2A4MP = 0.75 ± 0.06; kcat values [s−1]: 4-methylphenol = no data, catechol = 11 ± 1/12 ± 1, 3,4-dihydroxybenzaldehyde = 0.80 ± 0.09, L-DOPA = 0.066 ± 0.016, 3,4DHBA = no data, 3A4HBA = no data, 2AP = 320 ± 50/20 ± 3; 2A4MP = 18 ± 1.6,14a Kinetic parameters could not be determined due to the low aqueous solubility of 3A4HBA (<9 mM), which prevented adequate definition of the Michaelis–Menten curve for SgGriF.b Kinetic values could not be determined due to insufficient or absent enzymatic activity.c Kinetic parameters for o-aminophenol oxidation were determined by monitoring the formation of colored phenoxazinone oxidation products, reflecting the quinone imine-forming activity. | ||||
| o-Diphenols (Fig. S9) | ||||
| Benzene-1,2-diol (catechol) | SmNspF | 1.41 ± 0.20 | 1.275 ± 0.106 | 0.904 ± 0.149 |
| SgGriF | 2.58 ± 0.20 | 2.198 ± 0.107 | 0.852 ± 0.078 | |
| 4-Methylbenzene-1,2-diol (4-methylcatechol) | SmNspF | 1.73 ± 0.35 | 5.648 ± 0.633 | 3.265 ± 0.755 |
| SgGriF | 0.19 ± 0.01 | 1.140 ± 0.008 | 6.000 ± 0.319 | |
| 4-(2-Aminoethyl)benzol-1,2-diol (dopamine) | SmNspF | 10.04 ± 0.77 | 0.838 ± 0.048 | 0.083 ± 0.008 |
| SgGriF | 5.19 ± 0.17 | 1.054 ± 0.021 | 0.203 ± 0.008 | |
| 3,4-Dihydroxybenzoic acid (3,4DHBA) | SmNspF | 4.65 ± 0.44 | 0.723 ± 0.056 | 0.155 ± 0.019 |
| SgGriF | 5.47 ± 0.15 | 0.020 ± 0.001 | 0.004 ± 0.001 | |
| (3,4-Dihydroxyphenyl)acetic acid (DOPAC) | SmNspF | n. d.b | n. d.b | n. d.b |
| SgGriF | n. d.b | n. d.b | n. d.b | |
| o-Aminophenols (Fig. S10)c | ||||
| 2-Aminophenol (2AP) | SmNspF | 9.35 ± 3.27 | 14.092 ± 4.011 | 1.507 ± 0.680 |
| SgGriF | 2.30 ± 0.06 | 26.481 ± 0.395 | 11.513 ± 0.346 | |
| 2-Amino-4-methylphenol (2A4MP) | SmNspF | 6.13 ± 1.01 | 8.374 ± 0.103 | 1.366 ± 0.226 |
| SgGriF | 0.50 ± 0.03 | 6.385 ± 0.172 | 12.770 ± 0.840 | |
| 3-Amino-4-hydroxybenzoic acid (3A4HBA) | SmNspF | 4.50 ± 0.83 | 12.086 ± 1.406 | 2.686 ± 0.981 |
| SgGriF | n. d.a | n. d.a | n. d.a | |
| 4-Amino-3-hydroxybenzoic acid (4A3HBA) | SmNspF | n. d. | n. d. | n. d. |
| SgGriF | n. d. | n. d. | n. d. | |
| 2-Amino-1,3-benzenediol (2-aminoresorcinol) | SmNspF | 12.14 ± 0.98 | 2.315 ± 0.166 | 0.19 ± 0.02 |
| SgGriF | 8.83 ± 1.68 | 10.402 ± 1.689 | 1.18 ± 0.30 | |
| Aniline derivatives (Fig. S11) | ||||
| Aniline | SmNspF | n. d.b | n. d.b | n. d.b |
| SgGriF | n. d.b | n. d.b | n. d.b | |
| 1,2-Diaminobenzene (o-phenylenediamine) | SmNspF | 7.28 ± 0.24 | 0.018 ± 0.002 | 0.003 ± 0.001 |
| SgGriF | 3.03 ± 0.22 | 0.007 ± 0.002 | 0.002 ± 0.001 | |
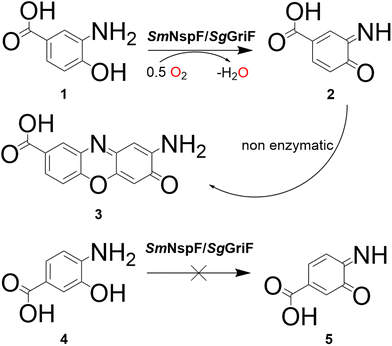
SmNspF and SgGriF catalyze the oxidation of o-aminophenols to phenoxazinone products (quinone imine-forming activity) with substrates including 2-aminophenol (2AP), 2-amino-4-methylphenol (2A4MP), 3-amino-4-hydroxybenzoic acid (3A4HBA, 1) and 2-aminoresorcinol (Fig. 1 and 2, column 1; Table 1). Despite both enzymes showing enzymatic activity toward 3A4HBA (1), no activity was observed with 4-amino-3-hydroxybenzoic acid (4), which contains a carboxylic acid group in the meta-position relative to the phenolic hydroxy group (Fig. 1, columns 1 and 2; Fig. 2). These findings indicate that AOs display substrate specificity for o-aminophenols with functional groups at the para-position, which supports effective substrate binding at the active center.
Remarkably, AOs are accepting dihydroxyaniline substrates with two hydroxy groups at the ortho-position relative to the amino group as they exhibit enzymatic activity toward 2-aminoresorcinol (Table 1 and Fig. S10). The catalytic efficiency of this oxidation is much lower compared to 2-aminophenol which is demonstrated by an approximately 10-fold lower activity with SmNspF and 4-fold lower activity with SgGriF. Whereas tyrosinases demonstrate their ability to hydroxylate aniline derivatives, SmNspF and SgGriF show no activity when incubated with aniline (Table 1).21–23 However, both AOs are capable of enzymatically oxidizing substrates beyond phenolic compounds, as they exhibit low activities toward o-phenylenediamine (Table 1 and Fig. 1, i column 3), similar to tyrosinases.21–23 This expands the known enzymatic versatility of AOs, demonstrating their ability to catalyze the oxidation of aromatic o-diamines for the first time. The oxidation of o-phenylenediamine with AOs proceeds more slowly compared to catechol (over 300-fold lower) and 2AP (over 600-fold lower). An explanation for the considerable difference between the substrate classes (aromatic o-diamines vs. o-diphenols vs. o-aminophenols) is their relative tendency to undergo deprotonation, given the significantly higher pKa of anilines (pKa ∼ 25) compared to phenols (pKa ∼ 10) and the amino group in aminophenols (pKa ∼ 5).22,24 Hence, substrate deprotonation is a pivotal and rate-determining step in the enzymatic reaction mechanism.22
Differences in enzymatic kinetics between SmNspF and SgGriF are most evident in their substrate preferences for para-substituted o-diphenols and o-aminophenols. Thus, SmNspF displays strikingly higher activity toward substrates with a carboxylic acid group (approximately 38-fold higher for 3,4DHBA), whereas SgGriF exhibits greater activity toward those substituted with a methyl group in the para-position (approximately 2-fold higher for 4-methylcatechol and 9-fold for 2A4MP) (Table 1). The exceptional preference of SmNspF for o-aminophenol substrates bearing a carboxylic acid group is evident not only in comparison to SgGriF but also to tyrosinases.6,21–23 Based solely on the observed reaction rates, a clear distinction emerges between AOs and tyrosinases. In contrast to kinetic data collected from two fungal tyrosinases (Neurospora crassa and Agaricus bisporus), which exhibit a strong preference for o-diphenolic substrates over their corresponding o-aminophenol analogs, both AOs examined in this study show predominantly higher or comparable catalytic efficiencies with o-aminophenol substrates (Table 1, Fig. S9–S10).21–23 For example, SgGriF oxidizes 2-aminophenol approximately 5-fold faster than catechol and SmNspF oxidizes 3A4HBA approximately 2-fold faster than the corresponding o-diphenol 3,4DHBA (Table 1). Enzymatic activities of mushroom tyrosinase published by Muñoz-Muñoz et al. (2011) show a 70-fold higher catalytic efficiency for catechol and 29-fold higher for 3,4DHBA oxidation in comparison to their o-aminophenol analogs.21
Structural evaluation
Protein crystals of SmNspF were obtained from a buffered solution at pH 7.5 containing 20 mM TRIS, 8% PEG 8000 and 800 mM Li2SO4 as the precipitating agent (SI Section S1.8). An in crystallo activity using 2-amino-4-methylphenol confirmed that the crystallized protein remained active, as evidenced by the rapid and intense coloration of the protein crystal (Fig. S15). The crystal structure of SmNspF represents the first experimental structural determination of an AO (PDB ID: 9T62). The SmNspF core architecture closely resembles previously reported polyphenol oxidase (PPO) structures.7,20 A great structural similarity is observed with the crystal structure of the tyrosinase from Streptomyces castaneoglobisporus (ScTYR, PDB ID:1WX2, Uniprot ID: Q83WS2), sharing 48.17% sequence identity and Streptomyces avermitillis (SaTYR, PDB ID: 6J2U, Uniprot ID: Q79Z38), sharing 43.17% sequence identity.13,25 A notable distinction between SmNspF and bacterial tyrosinases is the presence of three elongated loop regions, conserved among AOs and they are not present in tyrosinases (Fig. 3). One of these loops (Peptide Loop 1 (PL1)) lies near the active site and includes a 14-residue insertion in both SmNspF (Pro138–Thr151) and SgGriF (Pro138–Thr151) compared to the structure of ScTYR and SaTYR (Fig. 3).14 PL1 (Pro138–Thr151) and Peptide Loop 3 (PL3: Phe212–Gly219) could possibly restrict the active site and subsequently play a role in substrate recognition due to their proximity to the binding pocket (distance to CuA of PL1 ≈ 9.5 Å and PL3 ≈ 15.5 Å) (Fig. 3). In contrast, Peptide Loop 2 (PL2: Ser195–Asn200) exhibits a much larger distance from the active-site pocket (distance to CuA of PL3 ≈ 19.5 Å) (Fig. 3). Yet the function of this conserved AO loop in enzyme activity is still unclear. The monomeric SmNspF is ellipsoidal, measuring approximately 48 × 46 × 32 Å. The secondary structure is predominantly α-helical, with the enzyme core formed by a four α-helix bundle (α2, α3, α5, α6). Each of the two catalytic copper ions is coordinated by three histidines from the α-helices. CuA is ligated by His39, His58, and His67, with His39 positioned mid-helix on α2. CuB is coordinated by His222, His226, and His248, located in the middle of α6 and α7. The presence of an ‘oxygen moiety’ in the electron density map between the copper ions suggests that the enzyme was crystallized in its met-form. The dicopper center features a Cu–Cu distance of 3.8 Å, comparable to that of the met-form of ScTYR (3.7 Å).18Multiple structure to function relationships in polyphenol oxidases (PPOs) have been reported in earlier studies and comprehensively reviewed by Kampatsikas and Rompel (2021).20 Second-shell residues at the active-site play a crucial role in substrate specificity and reaction rates, despite only minor residue differences among PPOs.26,27 For example, the residue positioned after the second CuB-coordinating histidine, known as the second activity controller, alters the local electrostatic environment (Fig. 4).20 In tyrosinases ToTYR-02 and JrTYR, this position contains a small hydrophobic residue (Ile in ToTYR-02 and Leu in JrTYR), whereas in ToTYR-06, and in catechol oxidases IbCO, and VvCO, it is occupied by a long, positively charged residue (Arg or Lys) capable of stabilizing acidic groups on substrate's tail, as the HisB2+1 residue interacts directly with the substrate's tail.20,27,28 A positively charged amino acid at this position such as Arg enhances the active site's affinity for negatively charged substrates, whereas the opposite effect occurs for positively charged substrates.28 The substrate-guiding effect of the second activity controller (HisB2+1) can also help to explain the functional differences between SmNspF and SgGriF, given the high sequence homology between tyrosinases and AOs (Fig. 4).
 | ||
| Fig. 4 Structure alignment of SmNspF (PDB ID: 9T62) and SgGriF (AlphaFoldDB model B1VTI5) with PyMOL Molecular Graphics System, Version 3.0 Schrödinger, LLC.29,30 (A) Active center of SmNspF with coordinating histidines in cyan and the activity selector Asn43 in yellow. The seventh histidine is depicted in blue, and the second activity controller (HisB2+1) Gln227 is depicted in grey. Copper ions are depicted as brown spheres. (B) Active center of SgGriF with coordinating histidines in cyan and the activity selector Asn43 in yellow. SgGriF harbors Asn247, depicted in magenta, instead of the seventh histidine, and the second activity controller (HisB2+1) Val227 is shown in blue. Copper ions are depicted as brown spheres. The structure alignment of SmNspF (PDB ID: 9T62) and SmNspF AlphaFold model (AlphaFoldDB model D6RTB9) is shown in Fig. S16. | ||
In SmNspF, HisB2+1 is a polar Gln, which can act as a hydrogen-bond donor, potentially explaining its preference for substrates with carboxyl groups that serve as hydrogen-bond acceptors.31 In contrast, SgGriF contains a hydrophobic Val at this position, resulting in a significantly lower reaction rate toward carboxylic-tail substrates such as 3,4-dihydroxybenzoic acid and 3-amino-4-hydroxybenzoic acid. This divergence between the two AOs can be visualized in the electrostatic and hydrophobic surface maps (Fig. 5). On the one hand, residue Gln227 in SmNspF increases the electrostatic potential in the outer binding-pocket region in comparison to the surface map of SgGriF with Val227. On the other hand, Val227 in SgGriF significantly increases local hydrophobicity, which hinders polar interactions with substrates harboring a carboxylate tail. Please note that PPOs with a seventh histidine residue, such as SmNspF, exhibit high affinity for substrates with a carboxylic tail, whereas those containing leucine at this position show greater affinity for substrates with a decarboxylated tail.20,32,33 The binding-pocket volumes of both enzymes calculated with a maximum depth of 7.2 Å are very similar (approximately 218 Å3 in SmNspF and 230 Å3 in SgGriF). One possible cause of the slightly tighter binding pocket of SmNspF is the different orientation of Peptide Loop 1, which in SmNspF shows a greater interaction with the outer binding-pocket region (Fig. 3 and 5). Despite the divergent orientation of Peptide Loop 1, both enzymes exhibit nitroso-forming activity, indicating that this additional loop is likely not pivotal for this activity. The slightly smaller binding pocket in SmNspF compared to SgGriF is consistent with the observed kinetic values. On the one hand, SmNspF exhibits relatively high substrate specificity for 3A4HBA due to its tightened binding pocket; on the other hand, it shows much lower reaction rates (approximately tenfold lower) for the other investigated o-aminophenols compared to the larger binding pocket in SgGriF.
 | ||
| Fig. 5 Electrostatic and hydrophobic surface map of (A) SmNspF (PDB ID: 9T62) and (B) SgGriF (AlphaFoldDB model B1VTI5) with ChimeraX 1.11.29,30,34 Residues Gln227 (SmNspF) and Val227 (SgGriF) are highlighted in green. The negative potential surface is depicted in red, and the positive potential surface is depicted in blue. The hydrophilicity is depicted in cyan, and hydrophobicity is depicted in yellow. The calculated binding-pocket volume with a maximum depth of 7.2 Å is 218 Å3 in SmNspF and 230 Å3 in SgGriF. | ||
SgGriF lacks the seventh histidine and instead contains an asparagine in the homologous position (Fig. 4). In tyrosinases, the presence of amino acids Asp or Asn at the positions of the first (HisB1+1) and second (HisB2+1) activity controller residues creates an environment that enhances the basicity of the conserved HisB1 and HisB2 residues, enabling them to act as proton acceptors.20 This structural feature is an indicator of monophenolase activity, based on previous mutagenesis studies focusing on the first (HisB1+1) and second (HisB2+1) activity controllers.20 The thereby activated HisB1 and HisB2 can deprotonate monophenolic substrates, facilitating the monophenolase function.20 In SmNspF and SgGriF, the HisB1+1 position is occupied by a conserved Gly residue, while the HisB2+1 position contains a polar Gln in SmNspF and a nonpolar Val in SgGriF (Fig. 4). Despite this structural difference from tyrosinases, both enzymes display slow monophenolase activity toward 4-methylphenol (Fig. 1, ii columns 8 and 9).12 A hypothesis for the weak monophenolase activity of both enzymes is that the relatively flexible HisA2, in the absence of the thioether bridge, can move closer to the copper center. The thioether bridge is present in most structurally characterized plant and fungal PPOs.20 Without the thioether bond, as in SmNspF and SgGriF as well as in most bacterial tyrosinases, HisA2 could possibly interact with the acidic residue (water keeper Glu207), increasing its basicity enough to deprotonate monophenolic substrates (Fig. 1, columns 8 and 9).20,35 Recent experimental investigations combined with quantum mechanics/molecular mechanics studies by Kipouros et al. (2022) show an alternative mechanism, highlighting that the monophenolic hydrogen reacts directly with the μ:η2:η2-peroxide dicopper(II) center which acts as a proton acceptor.36–38
Conclusion
The investigation of the enzymatic activities of SmNspF and SgGriF, which belong to a highly versatile subclass of the binuclear copper protein family, confirmed their overall similarity in substrate specificity, while also revealing differences in catalytic efficiencies between them. This study provides new insight into the AOs by presenting crystal structure information about the overall structure and dicopper center of SmNspF and by elucidating how small sequence variations in the second coordination sphere can influence the distinct substrate preferences of SmNspF and SgGriF. The substrate-guiding effect of the second activity controller (HisB2+1) in SmNspF and SgGriF determines the binding affinity for carboxyl-containing substrates, thereby leading to increased or decreased reaction rates with compounds such as 3,4DHBA and 3A4HBA. Regarding monophenolase activity, SmNspF and SgGriF were found to be active without hydroxylamine, while the addition of hydroxylamine further accelerated this activity. Beyond the classical mono-, o-diphenolic, and o-aminophenolic substrates, both AOs are also capable of oxidizing 2-aminoresorcinol and o-phenylenediamine, albeit at markedly slower rates. This observation not only extends the known substrate spectrum of AOs, but also suggests a previously underappreciated flexibility in their enzymatic activities. Since aniline derivatives are widely used in dyes, pharmaceuticals, and synthetic polymers, exploiting AO reactivity toward o-phenylenediamines offers opportunities in the synthesis of functional polymers, sensors, and novel building blocks.39–42 Resorcinol derivatives are applied in pharmaceuticals, tire manufacturing, hair dyes, and dermatology, but pose serious toxicological risks as it can disrupt the central nervous system, impair red blood cell function, and act as an endocrine disruptor.43–50 They represent major phenolic wastewater contaminants from industries such as paper, resin, textile, plastic, coking, tanning, rubber, pharmaceuticals, herbicides, fungicides, and petroleum production.46 The ability of AOs to oxidize such pollutants underscores their potential for sustainable bioremediation, which utilizes biocatalysts containing Earth-abundant metals rather than rare or precious ones. Overall, these findings highlight the ability of AOs to oxidize a great variety of small compounds and provide the opportunity for rational engineering of catalytic properties from sequence-level information to enable reactions previously considered inaccessible for biocatalysts.Author contributions
Hoa Le Xuan: conceptualization, data curation, formal analysis, investigation, methodology, project administration, validation, visualization, and writing – original draft. Annette Rompel: conceptualization, funding acquisition, project administration, resources, supervision, and writing – original draft. All the authors have approved the final manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are openly available in phaidra at https://phaidra.univie.ac.at/o:2178036. Crystallographic dataset are openly available at European Synchrotron Radiation Facility DOI: https://doi.org/10.15151/ESRF-DC-2333941181. Ref. 51–65 are cited in the supplementary information (SI). Supplementary information containing detailed experimental procedures on heterologous expression, analytical characterization, kinetic investigation, and X-ray data collection is available. See DOI: https://doi.org/10.1039/d5qi02495a.Acknowledgements
The research was supported by the Austrian Science Fund (FWF): https://doi.org/10.55776/P32932 as well as the University of Vienna. The authors would like to express their gratitude to Dr Igor Melnikov for beamline support with X-ray diffraction at the ESRF Beamline ID30A-3 as part of the Austrian ACDC BAG Proposal MX2455. Moreover, the authors thank Anna Fabisikova, M.Sc. (Mass Spectrometry Center, Core Facility, Faculty of Chemistry, University of Vienna) for technical support with ESI-MS measurements and Elizabeta Paar, M.Sc. for her support with protein expression optimization.References
- E. A. Barka, P. Vatsa, L. Sanchez, N. Gaveau-Vaillant, C. Jacquard, J. P. Meier-Kolthoff, H.-P. Klenk, C. Clément, Y. Ouhdouch and G. P. van Wezel, Taxonomy, Physiology, and Natural Products of Actinobacteria, Microbiol. Mol. Biol. Rev., 2015, 80, 1–43, DOI:10.1128/MMBR.00019-15.
- A. van der Meij, S. F. Worsley, M. I. Hutchings and G. P. van Wezel, Chemical ecology of antibiotic production by actinomycetes, FEMS Microbiol. Rev., 2017, 41, 392–416, DOI:10.1093/femsre/fux005.
- T. Arai and Y. Mikami, Chromogenicity of Streptomyces, Microbiology, 1972, 23, 402–406, DOI:10.1128/am.23.2.402-406.19.
- A. A. Sarmiento-Tovar, L. Silva, J. Sánchez-Suárez and L. Diaz, Streptomyces-Derived Bioactive Pigments: Ecofriendly Source of Bioactive Compounds, Coatings, 2022, 12, 1858, DOI:10.3390/coatings12121858.
- C. Popa and G. Bahrim, Streptomyces Tyrosinase: Production and Practical Applications, Innovative Rom. Food Biotechnol., 2011, 8, 1–7 CAS.
- A. Noguchi, T. Kitamura, H. Onaka, S. Horinouchi and Y. Ohnishi, A copper-containing oxidase catalyzes C-nitrosation in nitrosobenzamide biosynthesis, Nat. Chem. Biol., 2010, 6, 641–643, DOI:10.1038/nchembio.418.
- UniProt Consortium, UniProt: the universal protein knowledgebase in 2021, Nucleic Acids Res., 2021, 49, D480–D489, DOI:10.1093/nar/gkaa1100.
- F. Aguilera, C. McDougall and B. M. Degnan, Origin, evolution and classification of type-3 copper proteins: lineage-specific gene expansions and losses across the Metazoa, BMC Evol. Biol., 2013, 13, 96, DOI:10.1186/1471-2148-13-96.
- M. Pretzler and A. Rompel, Tyrosinases: a family of copper-containing metalloenzymes, ChemTexts, 2024, 10, 12–59, DOI:10.1007/s40828-024-00195-y.
- I. Kampatsikas, M. Pretzler and A. Rompel, Identification of Amino Acid Residues Responsible for C−H Activation in Type-III Copper Enzymes by Generating Tyrosinase Activity in a Catechol Oxidase, Angew. Chem., Int. Ed., 2020, 59, 20940–20945, DOI:10.1002/anie.202008859.
- A. M. Mayer, Polyphenol oxidases in plants and fungi: going places? A review, Phytochemistry, 2006, 67, 2318–2331, DOI:10.1016/j.phytochem.2006.08.006.
- E. I. Solomon, D. E. Heppner, E. M. Johnston, J. W. Ginsbach, J. Cirera, M. Qayyum, M. T. Kieber-Emmons, C. H. Kjaergaard, R. G. Hadt and L. Tian, Copper Active Sites in Biology, Chem. Rev., 2014, 114, 3659–3853, DOI:10.1021/cr400327t.
- A. Rompel, H. Fischer, K. Büldt-Karentzopoulos, D. Meiwes, F. Zippel, H.-F. Nolting, C. Hermes, B. Krebs and H. Witzel, Spectroscopic and EXAFS studies on catechol oxidases with dinuclear copper centers of type 3: evidence for μ-η2:η2-peroxo-intermediates during the reaction with catechol, J. Inorg. Biochem., 1995, 59, 715, DOI:10.1016/0162-0134(95)97803-X.
- J. W. Ginsbach, M. T. Kieber-Emmons, R. Nomoto, A. Noguchi, Y. Ohnishi and E. I. Solomon, Structure/function correlations among coupled binuclear copper proteins through spectroscopic and reactivity studies of NspF, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 10793–10797, DOI:10.1073/pnas.1208718109.
- H. Le Xuan, F. Panis and A. Rompel, Identification of an Activity Selector for the Nitroso-Forming Activity in Bacterial Type-III Copper Enzymes, Angew. Chem., Int. Ed., 2025, 64, e202501560, DOI:10.1002/anie.202501560.
- P. M. Hanna, R. Tamilarasan and D. R. McMillin, Cu(I) analysis of blue copper proteins, Biochem. J., 1988, 256, 1001–1004, DOI:10.1042/bj2561001.
- K. D. Karlin, J. C. Hayes, Y. Gultneh, R. W. Cruse, J. W. McKown, J. P. Hutchinson and J. Zubieta, Copper-mediated hydroxylation of an arene: model system for the action of copper monooxygenases. Structures of a binuclear copper(I) complex and its oxygenated product, J. Am. Chem. Soc., 1984, 106, 2121–2128, DOI:10.1021/ja00319a036.
- R. S. Himmelwright, N. C. Eickman, C. D. LuBien, K. Lerch and E. I. Solomon, Chemical and spectroscopic studies of the binuclear copper active site of Neurospora tyrosinase: comparison to hemocyanins, J. Am. Chem. Soc., 1980, 102, 7339–7344, DOI:10.1021/ja00544a031.
- H. Suzuki, Y. Furusho, T. Higashi, Y. Ohnishi and S. Horinouchi, A Novel o-Aminophenol Oxidase Responsible for Formation of the Phenoxazinone Chromophore of Grixazone, J. Biol. Chem., 2006, 281, 824–833, DOI:10.1074/jbc.M505806200.
- I. Kampatsikas and A. Rompel, Similar but Still Different: Which Amino Acid Residues Are Responsible for Varying Activities in Type-III Copper Enzymes?, ChemBioChem, 2021, 22, 1161–1175, DOI:10.1002/cbic.202000647.
- J. L. Muñoz-Muñoz, F. Garcia-Molina, P. A. Garcia-Ruiz, R. Varon, J. Tudela, J. N. Rodriguez-Lopez and F. Garcia-Canovas, Catalytic oxidation of o-aminophenols and aromatic amines by mushroom tyrosinase, Biochim. Biophys. Acta, Proteins Proteomics, 2011, 1814, 1974–1983, DOI:10.1016/j.bbapap.2011.07.015.
- M. Pretzler and A. Rompel, Beyond Phenolics: Alternative Substrates for Type III Copper Enzymes, ChemBioChem, 2025, 26, e202400982, DOI:10.1002/cbic.202400982.
- O. Toussaint and K. Lerch, Catalytic oxidation of 2-aminophenols and ortho hydroxylation of aromatic amines by tyrosinase, Biochemistry, 1987, 26, 8567–8571, DOI:10.1002/cbic.202400982.
- I. A. Badea, L. Axinte and L. Vladescu, Monitoring of aminophenol isomers in surface water samples using a new HPLC method, Environ. Monit. Assess., 2013, 185, 2367–2375, DOI:10.1007/s10661-012-2717-7.
- Y. Matoba, T. Kumagai, A. Yamamoto, H. Yoshitsu and M. Sugiyama, Crystallographic Evidence That the Dinuclear Copper Center of Tyrosinase Is Flexible during Catalysis, J. Biol. Chem., 2006, 281, 8981–8990, DOI:10.1074/jbc.M509785200.
- Y. Wu, R. Feng, T.-P. Zhou, I. Y. Zhang and B. Wang, Deciphering the Coordination Environment’s Impact on O–O Activation in Dicopper Enzymes: Computational Insights from AhyBURP Peptide Cyclase, ACS Catal., 2025, 15, 17456–17466, DOI:10.1021/acscatal.5c05262.
- A. Bijelic, M. Pretzler, C. Molitor, F. Zekiri and A. Rompel, The Structure of a Plant Tyrosinase from Walnut Leaves Reveals the Importance of “Substrate-Guiding Residues, Angew. Chem., Int. Ed., 2015, 54, 14677–14680, DOI:10.1002/anie.201506994.
- S. M. Prexler, R. Singh, B. M. Moerschbacher and M. E. Dirks-Hofmeister, A specific amino acid residue in the catalytic site of dandelion polyphenol oxidases acts as ‘selector’ for substrate specificity, Plant Mol. Biol., 2018, 96, 151–164, DOI:10.1007/s11103-017-0686-5.
- J. Jumper, R. Evans, A. Pritzel, T. Green, M. Figurnov, O. Ronneberger, K. Tunyasuvunakool, R. Bates, A. Žídek, A. Potapenko, A. Bridgland, C. Meyer, S. Kohl, A. J. Ballard, A. Cowie, B. Romera-Paredes, S. Nikolov, R. Jain, J. Adler and D. Hassabis, Highly accurate protein structure prediction with AlphaFold, Nature, 2021, 596, 583–589, DOI:10.1038/s41586-021-03819-2.
- M. Varadi, S. Anyango, M. Deshpande, S. Nair, C. Natassia, G. Yordanova, D. Yuan, O. Stroe, G. Wood, A. Laydon, A. Zídek, T. Green, K. Tunyasuvunakool, S. Petersen, J. Jumper, E. Clancy, E. Green, A. Vora, M. Lutfi and S. Velankar, AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models, Nucleic Acids Res., 2022, 50, D439–D444, DOI:10.1093/nar/gkab1061.
- M. Goldfeder, M. Kanteev, S. Isaschar-Ovdat, N. Adir and A. Fishman, Determination of tyrosinase substrate-binding modes reveals mechanistic differences between type-3 copper proteins, Nat. Commun., 2014, 5, 4505, DOI:10.1038/ncomms5505.
- D. Hernández-Romero, A. Sanchez-Amat and F. Solano, A tyrosinase with an abnormally high tyrosine hydroxylase/dopa oxidase ratio, FEBS J., 2006, 273, 257–270, DOI:10.1111/j.1742-4658.2005.05038.x.
- C. Olivares, J. C. García-Borrón and F. Solano, Identification of active site residues involved in metal cofactor binding and stereospecific substrate recognition in Mammalian tyrosinase. Implications to the catalytic cycle, Biochemistry, 2002, 41, 679–686, DOI:10.1021/bi011535n.
- E. C. Meng, T. D. Goddard, E. F. Pettersen, G. S. Couch, Z. J. Pearson, J. H. Morris and T. E. Ferrin, UCSF ChimeraX: Tools for structure building and analysis, Protein Sci., 2023, 32, e4792, DOI:10.1002/pro.4792.
- M. Fekry, K. K. Dave, D. Badgujar, E. Hamnevik, O. Aurelius, D. Dobritzsch and U. H. Danielson, The Crystal Structure of Tyrosinase from Verrucomicrobium spinosum Reveals It to Be an Atypical Bacterial Tyrosinase, Biomolecules, 2023, 13, 1360, DOI:10.3390/biom13091360.
- I. Kipouros, A. Stańczak, J. W. Ginsbach, P. C. Andrikopoulos, L. Rulíšek and E. I. Solomon, Elucidation of the tyrosinase/O2/monophenol ternary intermediate that dictates the monooxygenation mechanism in melanin biosynthesis, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2205619119, DOI:10.1073/pnas.2205619119.
- A. Stańczak, I. Kipouros, P. Eminger, E. M. Dunietz, E. I. Solomon and L. Rulíšek, Coupled binuclear copper sites in biology: An experimentally-calibrated computational perspective, Coord. Chem. Rev., 2025, 525, 216301, DOI:10.1016/j.ccr.2024.216301.
- I. Kipouros, A. Stańczak, E. M. Dunietz, J. W. Ginsbach, M. Srnec, L. Rulíšek and E. I. Solomon, Experimental Evidence and Mechanistic Description of the Phenolic H-Transfer to the Cu2O2 Active Site of oxy-Tyrosinase, J. Am. Chem. Soc., 2023, 145, 22866–22870, DOI:10.1021/jacs.3c07450.
- D. Yang, J. Wang, Y. Cao, X. Tong, T. Hua, R. Qin and Y. Shao, Polyaniline-Based Biological and Chemical Sensors: Sensing Mechanism, Configuration Design, and Perspective, ACS Appl. Electron. Mater., 2023, 5, 593–611, DOI:10.1021/acsaelm.2c01405.
- M.-R. Huang, X.-G. Li and Y. Yang, Oxidative polymerization of o-phenylenediamine and pyrimidylamine, Polym. Degrad. Stab., 2000, 71, 31–38, DOI:10.1016/S0141-3910(00)00137-3.
- L. Li, M. Zheng, D. Jiang, S. Cao, K. Liu and J. Liu, Colorimetric and Fluorescent Probes Based on the Oxidation of o-Phenylenediamine for the Detection of Bio-Molecules, Prog. Chem., 2022, 34, 1815–1830, DOI:10.7536/PC210927.
- Q. Ye, S. Ren, H. Huang, G. Duan, K. Liu and J.-B. Liu, Fluorescent and Colorimetric Sensors Based on the Oxidation of o-Phenylenediamine, ACS Omega, 2020, 5, 20698–20706, DOI:10.1021/acsomega.0c03111.
- H. Dressler, Resorcinol: Its Uses and Derivatives, Springer US, Boston, 1994, in The Use of Resorcinol in Rubber Compositions, 59–83, DOI:10.1007/978-1-4899-0999-2_4.
- K. Ngamchuea, B. Tharat, P. Hirunsit and S. Suthirakun, Electrochemical oxidation of resorcinol: mechanistic insights from experimental and computational studies, RSC Adv., 2020, 10, 28454–28463, 10.1039/d0ra06111e.
- G. Manasa, A. K. Bhakta, Z. Mekhalif and R. J. Mascarenhas, Voltammetric study and rapid quantification of resorcinol in hair dye and biological samples using ultrasensitive maghemite/MWCNT modified carbon paste electrode, Electroanalysis, 2019, 31, 1363–1372, DOI:10.1002/elan.201900143.
- M. R. Gomez, R. A. Olsina, L. D. Martinez and M. F. Silva, Simultaneous determination of chloramphenicol, salicylic acid and resorcinol by capillary zone electrophoresis and its application to pharmaceutical dosage forms, Talanta, 2003, 61, 233–238, DOI:10.1016/S0039-9140(03)00267-4.
- R. Deepa, H. Manjunatha, V. Krishna and B. E. Kumara Swamy, Electrochemical Investigation of Resorcinol in Pterocarpus marsupium RoxB by Cyclic Voltammetric Study, J. Anal. Bioanal. Tech., 2014, 5, 218, DOI:10.4172/2155-9872.1000218.
- M. Khodari, G. A. M. Mersal, E. M. Rabie and H. F. Assaf, Electrochemical Sensor based on Carbon Paste Electrode Modified by TiO2 nano-particles for the Voltammetric Determination of Resorcinol, Int. J. Electrochem. Sci., 2018, 13, 3460–3474, DOI:10.20964/2018.04.04.
- E. Pasquier, C. Viguié, J.-B. Fini, S. Mhaouty-Kodja and C. Michel-Caillet, Limits of the regulatory evaluation of resorcinol as a thyroid disruptor: When limited experimental data challenge established effects in humans, Environ. Res., 2023, 222, 115330, DOI:10.1016/j.envres.2023.115330.
- I. V. Dingenen, E. Andersen, S. Volz, M. Christiansen, J. Novák, A.-C. Haigis, E. Stacy, B. R. Blackwell, D. L. Villeneuve, L. Vergauwen, K. Hilscherová, H. Holbech and D. Knapen, The thyroid hormone system disrupting potential of resorcinol in fish, Ecotoxicol. Environ. Saf., 2024, 284, 116995, DOI:10.1016/j.ecoenv.2024.116995.
- E. Gasteiger, C. Hoogland, A. Gattiker, S. Duvaud, M. Wilkins, R. Appel and A. Bairoch, in Proteomics Protocols Handbook, ed. J. Walker, Humana Press, Totowa 2005, The Use of Resorcinol in Rubber Compositions, 571–607, DOI:10.1385/1-59259-890-0:571.
- D. F. Swinehart, The Beer-Lambert Law, J. Chem. Educ., 1962, 39, 333, DOI:10.1021/ed039p333.
- D. Marquardt, An Algorithm for Least-Squares Estimation of Nonlinear Parameters, J. Soc. Ind. Appl. Math., 1963, 11, 431–441, DOI:10.1137/0111030.
- C. Vonrhein, C. Flensburg, P. Keller, A. Sharff, O. Smart, W. Paciorek, T. Womack and G. Bricogne, Data processing and analysis with the autoPROC toolbox, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2011, 67, 293–302, DOI:10.1107/S0907444911007773.
- I. J. Tickle, C. Flensburg, P. Keller, W. Paciorek, A. Sharff, C. Vonrhein and G. Bricogne, STARANISO, United Kingdom: Global Phasing Ltd, Cambridge, 2016. https://staraniso.globalphasing.org/cgi-bin/staraniso.cgi Search PubMed.
- A. Vagin and A. Teplyakov, MOLREP: an Automated Program for Molecular Replacement, Appl. Crystallogr., 1997, 30, 1022–1025, DOI:10.1107/S0021889897006766.
- E. Krissinel, A. A. Lebedev, V. Uski, C. Ballard, R. M. Keegan, O. Kovalevskiy, R. A. Nicholls, N. S. Pannu, P. Skubak, J. Berrisford, M. Fando, B. Lohkamp, M. Wojdyr, A. J. Simpkin, J. M. H. Thomas, C. Oliver, C. Vonrhein, G. Chojnowski, A. Basle, A. Purkiss, M. N. Isupov, S. McNicholas, E. Lowe, J. Trivino, K. Cowtan, J. Agirre, D. J. Rigden, I. Uson, V. Lamzin, I. Tews, G. Bricogne, A. G. W. Leslie and D. Brown, CCP4 Cloud for structure determination and project management in macromolecular crystallography, Acta Crystallogr., Sect. D:Struct. Biol., 2022, 78, 1079–1089, DOI:10.1107/S2059798322007987.
- P. Emsley and K. Cowtan, Coot: model-building tools for molecular graphics, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2004, 60, 2126–2132, DOI:10.1107/S0907444904019158.
- G. N. Murshudov, A. A. Vagin and E. J. Dodson, Refinement of macromolecular structures by the maximum-likelihood method, Acta Crystallogr., Sect. D:Biol. Crystallogr., 1997, 53, 240–255, DOI:10.1107/S0907444996012255.
- V. B. Chen, W. B. Arendall III, J. J. Headd, D. A. Keedy, R. M. Immormino, G. J. Kapral, L. W. Murray, J. S. Richardson and D. C. Richardson, MolProbity: all-atom structure validation for macromolecular crystallography, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2010, 66, 12–21, DOI:10.1107/S0907444909042073.
- C. K. Brown, J. F. Corbett and N. P. Loveless, Spectrophotometric studies on the protonation of hydroxy and aminophenazines in aqueous solution, Spectrochim. Acta, Part A, 1979, 35, 421–423, DOI:10.1016/0584-8539(79)80155-5.
- S. Fornera and P. Walde, Spectrophotometric quantification of horseradish peroxidase with o-phenylenediamine, Anal. Biochem., 2010, 407, 293–295, DOI:10.1016/j.ab.2010.07.034.
- G. A. Gamov, M. N. Zavalishin, A. Y. Khokhlova, A. V. Gashnikova, A. N. Kiselev, A. V. Zav'yalov and V. V. Aleksandriiskii, Kinetics of the Oxidation of Protocatechuic and Gallic Acids by Atmospheric Oxygen in the Presence of Laccase from T. versicolor, Russ. J. Phys. Chem., 2020, 94, 294–300, DOI:10.1134/S0036024420020119.
- W. J. Herbert, Calculating extinction coefficients for enzymatically produced o-quinones, Anal. Biochem., 1976, 75, 211–218, DOI:10.1016/0003-2697(76)90072-5.
- J. L. Muñoz, F. García-Molina, R. Varón, J. N. Rodriguez-Lopez, F. García-Cánovas and J. Tudela, Calculating molar absorptivities for quinones: Application to the measurement of tyrosinase activity, Anal. Biochem., 2006, 351, 128–138, DOI:10.1016/j.ab.2006.01.011.
| This journal is © the Partner Organisations 2026 |