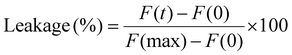Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Antimicrobial amphiphilic random copolymers from bio-based methacrylates: effect of chemical composition on activity and selectivity
Zao Cheng ab,
Erika Zaganelli
ab,
Erika Zaganelli ad,
Theo van Kootenb,
Caterina Presuttic,
Bert Poolmanc,
Laura Mazzocchetti
ad,
Theo van Kootenb,
Caterina Presuttic,
Bert Poolmanc,
Laura Mazzocchetti d and
Patrizio Raffa
d and
Patrizio Raffa *a
*a
aSmart and Sustainable Polymeric Products, Engineering and Technology Institute Groningen, University of Groningen, Nijenborgh 4, 9747 AG, The Netherlands. E-mail: p.raffa@rug.nl
bBiomaterials & Biomedical Technology, University Medical Center Groningen, Antonius Deusinglaan 1, 9713 AV Groningen, The Netherlands
cDepartment of Biochemistry, University of Groningen, Nijenborgh 3, 9747 AG, Groningen, The Netherlands
dDepartment of Industrial Chemistry, University of Bologna, Via Piero Gobetti 85, 40129, Bologna, Italy
First published on 22nd April 2026
Abstract
Antibiotic resistance is a growing global threat, prompting the development of novel synthetic antimicrobial agents. Many systems are inspired by antimicrobial peptides (AMPs), which are structurally cationic amphiphilic copolymers. To mimic this type of structure, using at the same time bio-based building blocks, in this study, we synthesized 15 amphiphilic random copolymers using amino acid-derived methacrylates with different side chains (Ala, Phe, and Lys) as the cationic components and bio-based tetrahydrogeranyl methacrylate (THGA) as the hydrophobic component via Reversible Addition–Fragmentation chain Transfer (RAFT) polymerization, yielding copolymers with varied cationic–hydrophobic balances. The antimicrobial activity of these copolymers was evaluated against E. coli and S. aureus, whereas cytotoxicity was evaluated using mouse fibroblasts (L929). The results indicate that Lys-based copolymers with higher cationic content are the most active and selective. Activity against S. aureus was influenced by the hydrophobic–hydrophilic balance, showing a non-linear trend. The cytotoxicity of these polymers also increased with the rise in cationic content. Live/dead staining, SEM, and calcein leakage assays confirmed the membrane-disrupting effects of random copolymers containing Lys as the cationic component. Furthermore, leakage was observed in liposomes mimicking bacterial membranes at concentrations below the minimum inhibitory concentration (MIC). These findings highlight the potential of bio-based amphiphilic copolymers as selective antimicrobial agents, paving the way for a more sustainable design of such pharmaceutical products.
1. Introduction
Controlling the proliferation of harmful microorganisms, identifying pathogenic bacteria, and minimizing their impact on human life have long been focal points of research.1 This issue has become even more critical with the overuse of antibiotics, which has led to the emergence and rapid spread of antibiotic-resistant bacteria, posing a serious threat to global health.2,3 In response, researchers are actively seeking novel structural strategies to create next-generation antimicrobial materials that combine high efficacy with excellent biosafety. These efforts extend to a diverse array of approaches, including metal nanoparticles,4,5 antimicrobial peptides,6,7 antimicrobial polymers,8–11 and emerging hybrid systems that integrate multiple mechanisms of action.12–14 In the search for effective antibiotic alternatives, researchers are focused on identifying critical factors that enhance antimicrobial efficacy while minimizing cytotoxicity. Antimicrobial peptides (AMPs) represent a promising class of molecules, typically composed of 10–50 amino acid residues.7,15 Although this may be an oversimplification, the accepted antibacterial mechanism of AMPs is attributed to their amphiphilic nature, characterized by cationic charges, hydrophobic chains, and hydrophilic groups, which enables them to interact with bacterial membranes, disrupting cell integrity.16 The positive charges allow AMPs to bind preferentially to the negatively charged components commonly found on bacterial surfaces, such as anionic lipids in the membranes or teichoic acids in Gram-positive bacteria.17,18 These electrostatic interactions promote selective accumulation on bacterial membranes over mammalian cells. In Gram-negative bacteria, AMPs often first interact with the outer membrane, which contains lipopolysaccharides (LPS), before reaching the inner membrane; in Gram-positive bacteria, although the absence of LPS allows direct access to the cytoplasmic membrane, the thick peptidoglycan layer can also constitute a substantial barrier to AMP penetration.19,20 Consequently, the efficiency of membrane disruption and pore formation depends strongly on the specific AMPs and bacterial strain, and membrane destabilization remains a common mode of action.21With advancements in controlled polymerization techniques, particularly Reversible Deactivation Radical Polymerization (RDRP), such as Atom Transfer Radical Polymerization (ATRP) and Reversible Addition–Fragmentation chain Transfer (RAFT) polymerization,22–25 scientists have been able to synthesize numerous amphiphilic antimicrobial polymers that mimic the above-mentioned functional and structural attributes of natural AMPs.26–33 Recent investigations on synthetic polymers revealed that the molar mass plays a crucial role in determining their biocompatibility. In many reported studies, particularly for linear polymers, an increase in molar mass is associated with heightened cytotoxicity.34–36 To achieve an optimal amphiphilic balance, hydrophobic and cationic monomers are frequently utilized, with hydrophilic neutral monomers sometimes included.37–39 Importantly, the addition of these hydrophilic monomers has been demonstrated to significantly enhance biocompatibility. As mentioned above, it is commonly accepted that cationic groups facilitate the adsorption of polymers onto anionic bacterial membranes via electrostatic interactions, which has led to extensive research on various cationic moieties to better understand their structural implications for the overall biological activity.40–43 On the other hand, hydrophobic groups are suggested to disrupt the integrity of bacterial and mammalian membranes, prompting researchers to screen various hydrophobic monomers (including differences in chain length and chemical structure) to improve selectivity.37,44–46 Overall, both the type and density of cationic groups, as well as the nature of hydrophobic monomers, play crucial roles in determining antimicrobial activity and cytotoxicity.39,47,48 To address these requirements, cationic amino acid-functionalized monomers have been widely employed in antimicrobial polymer design, owing to their ability to partially reproduce key structural and functional features of AMPs.49–51 However, two critical limitations remain. First, although amino acid-based monomers have been increasingly used, most synthetic amphiphilic copolymers reproduce only a limited subset of AMPs’ compositional features, making it difficult to systematically disentangle the specific contributions of individual amino acid side chains to antimicrobial activity, selectivity, and cytotoxicity. In natural AMP sequences, antimicrobial function arises from a complex interplay of multiple residues and motifs, which together govern membrane affinity, selectivity, and conformational stability.52 Replicating this level of functional complexity in synthetic polymer systems remains a significant challenge. Second, the hydrophobic components in the majority of reported antimicrobial polymers are still based on fossil-derived monomers, which limits the sustainability of these materials.37,38,44,47
With the goal of proposing a more sustainable design for peptide-inspired amphiphilic polymers, in this study, we prepared polymers based on cationic and hydrophobic bio-based monomers. We converted amino acids into the corresponding methacrylates and explored the impact of amino acid side chain variations on the antimicrobial activity of amphiphilic polymers, utilizing amino acids with different side chains (Ala, methyl; Phe, aromatic; Lys, cationic) as the cationic components. Additionally, bio-based tetrahydrogeraniol (THG) was incorporated as the hydrophobic component, via modification into the corresponding methacrylate.53 Beyond its renewable origin, THG provides a moderate and well-balanced hydrophobicity, comparable to that of commonly used short- to medium-chain linear alkyl methacrylates (e.g., C6–C8), while its branched aliphatic structure helps avoid excessive hydrophobicity, which often significantly increases the cytotoxicity of polymers due to excessively long hydrophobic chains.32,54 Through RAFT controlled polymerization, we successfully synthesized a library of 15 statistical amphiphilic random copolymers with varying ratios of cationic and hydrophobic components. The antimicrobial activity and cytotoxicity of these copolymers were evaluated against Gram-negative E. coli, Gram-positive S. aureus, and mouse fibroblast cells (L929). These findings contribute to a deeper understanding of amphiphilic antimicrobial polymers and may help develop agents with enhanced antimicrobial efficacy and reduced cytotoxicity toward mammalian cells, starting from more sustainable building blocks.
2. Materials and methods
Materials
2,2′-Azobisisobutyronitrile (AIBN), tetrahydrogeraniol (THG), triethylamine (TEA), methacryloyl chloride, Boc-Ala-OH, Boc-Phe-OH, Boc-Lys (Boc)-OH·DCHA, N,N′-dicyclohexylcarbodiimide (DCC), N,N-dimethylpyridin-4-amine (DMAP), trifluoroacetic acid (TFA), 2-hydroxyethyl methacrylate (HEMA), 4-cyano-4-[(dodecylsulfanylthiocarbonyl)sulfanyl]pentanoic acid, calcein, phosphatidylcholine (PC), sodium bicarbonate (NaHCO3), anhydrous sodium sulfate (Na2SO4), sodium chloride (NaCl), potassium bisulfate (KHSO4), hydrochloric acid (HCl), dimethylformamide (DMF), dichloromethane (DCM), and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich. Phosphate buffered saline (PBS) and Roswell Park Memorial Institute (RPMI) 1640 medium were purchased from Gibco. The Cell Proliferation Kit XTT was purchased from PanReac AppliChem, and a LIVE/DEAD BacLight Bacterial Viability Kit was bought from Thermo Fisher Scientific (catalog number L7012). The bacteria and cells were obtained from ATCC: E. coli (ATCC 25922); S. aureus (ATCC 6538); and mouse fibroblast cells (L929 (ATCC CCL-1)).Synthesis of monomers
1H NMR (400 MHz, CDCl3) δ 6.09 (dt, J = 2.0, 1.0 Hz, 1H), 5.54 (p, J = 1.6 Hz, 1H), 4.25–4.11 (m, 2H), 1.94 (dd, J = 1.6, 1.0 Hz, 3H), 1.70–1.66 (m, 1H), 1.55–1.48 (m, 3H), 1.34–1.10 (m, 6H), 0.89 (dd, J = 20.5, 6.6 Hz, 9H).
13C NMR (101 MHz, CDCl3) δ 167.56, 136.56, 125.11, 63.28, 39.17, 37.11, 35.47, 29.92, 27.93, 24.60, 22.66, 22.58, 19.57, 18.31.
The synthesis methods for Boc-Phe-HEMA (93% yield) and Boc-Lys(Boc)-HEMA (95% yield) are identical. Notably, for the purchased Boc-Lys(Boc)-OH·DCHA salt, a pre-treatment was required. The precursor was first dissolved in DCM, and the solution was then extracted three times with a cold KHSO4 aqueous solution. The organic layer was dried over Na2SO4 and filtered, and the solvent was removed under reduced pressure to obtain the free Boc-Lys(Boc)-OH acid.
13C NMR (101 MHz, CDCl3) δ 173.15, 167.03, 153.62, 135.81, 126.16, 125.99, 80.23, 62.82, 62.16, 50.14, 28.32, 28.28, 28.26, 18.59, 18.22.
13C NMR (101 MHz, CDCl3) δ 171.69, 167.03, 155.04, 135.87, 135.80, 129.26, 128.55, 127.07, 126.23, 79.95, 62.94, 62.17, 54.35, 38.27, 28.26, 18.25.
13C NMR (101 MHz, CDCl3) δ 176.22, 156.31, 155.81, 80.06, 79.35, 53.25, 40.06, 31.97, 29.51, 28.40, 28.32, 22.37.
13C NMR (101 MHz, CDCl3) δ 172.57, 167.00, 156.03, 135.81, 126.19, 79.89, 62.83, 62.17, 53.24, 40.01, 32.26, 29.56, 28.40, 28.29, 22.42, 18.23.
Synthesis of Boc-protected random copolymers
Typically, Boc-Ala-HEMA and THGA (in varying ratios, totaling 10 mmol) were dissolved in 5 mL of 1,4-dioxane, followed by the addition of the RAFT chain transfer agent 4-cyano-4-[(dodecylsulfanylthiocarbonyl)sulfanyl]pentanoic acid (5% of the total monomer amount, 0.5 mmol) and the initiator AIBN (8 mg, 0.05 mmol). The mixture was purged with nitrogen for 15 minutes and then stirred at 70 °C. After 18 hours, the reaction was terminated by cooling the vial in an ice bath. The copolymer was isolated by evaporating the dioxane under reduced pressure, dissolving the residue in dichloromethane, and precipitating twice in an excess of n-hexane (or in a 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ethanol–water mixture for polymers with high THGA content) to remove unreacted impurities. The sample was then dried overnight in a vacuum oven at 40 °C, yielding a light-yellow solid copolymer.
1 ethanol–water mixture for polymers with high THGA content) to remove unreacted impurities. The sample was then dried overnight in a vacuum oven at 40 °C, yielding a light-yellow solid copolymer.
Deprotection of copolymers
The Boc-protected random copolymer (1 g) was combined with TFA (1 mL).55 After stirring for 30 minutes, TFA was removed by purging with nitrogen. The remaining residue was dissolved in water, and impurities were removed using a 3500 Da dialysis bag for 3 days. The copolymer solution in the dialysis bag was then lyophilized using a freeze dryer to obtain a light, fluffy, white fibrous copolymer. NMR analysis was used to determine the removal of Boc protective groups.1H NMR spectroscopy
NMR spectroscopy was used to analyze all the Boc-protected and de-protected copolymers to analyze polymer composition and conversion. Analysis was performed to determine the mole percent of THGA (MPTHGA), degree of polymerization (DP), and number-average molecular weight (Mn). All experiments were performed using an Agilent 400 MHz WB NMR spectrometer. All experiments were run with a gas flow across the probes at 535 L h−1 with sample spinning and at a temperature of 25 °C.Size exclusion chromatography (SEC)
The copolymers were also characterized by SEC analysis to measure the Mn, weight-average molecular weight (Mw) and dispersity (Đ) using DMF (containing 0.01 M LiBr) as an eluent in a Viscotek GPCmax instrument equipped with a model 302 TDA detector based on the refractive index and intrinsic viscosity and two columns (Agilent Technologies – PolarGel-L and M, 8 μm, 30 cm) at a flow rate of 1.0 mL min−1 and a temperature of 50 °C. A universal calibration curve was prepared using narrow dispersion poly(methyl methacrylate) (PMMA) standards (Polymer Laboratories). Polymer solutions of 2 mg mL−1 in DMF were prepared in the eluent and filtered through 0.45 μm filters prior to injection.Dynamic light scattering (DLS) and zeta potential (ζ) measurements
For those polymers that could self-assemble into aggregates in aqueous solution, a Malvern Zetasizer Nano ZS (Malvern Instruments, Worcestershire, UK) instrument equipped with a HeNe laser was used with a wavelength of λ = 633 nm and a scattering angle of 173°. The data were computed using cumulant fit analysis and the hydrodynamic diameter size (Dh) was calculated using the Stokes–Einstein equation with the software. The ζ value was determined from the electrophoretic mobility of the particles by applying the Smoluchowski equation. All polymers were measured at a concentration of 1 mg mL−1 in PBS (the solutions were filtered using a 0.22 μm pore size filter). Final data for the size and ζ of the particles were calculated as the average of three consecutive measurements of the same sample.Transmission electron microscopy (TEM)
The morphology of the nanoscale aggregates was analyzed via TEM using a Philips CM120 operating at 120 keV. Images were recorded using a Gatan model UltraScan 4000SP CCD camera under low-dose conditions. All polymers were measured at concentrations of 0.1–0.5 mg mL−1 in PBS (using a 0.22 μm pore size filter) and 10 μl of the polymer solution was placed on a carbon-coated copper grid (Quantifoil 3.5/1, Quantifoil Micro Tools, Jena, Germany). Then, samples were negatively stained with 2% uranyl acetate, and the excess solution was blotted off and dried at room temperature for a few minutes before measurement.Zone of inhibition experiment
Briefly, a single colony of microorganisms was picked from the agar plate, inoculated into Mueller–Hinton broth (MHB; 50 mL) and cultured at 37 °C for 12 h. Fresh colonies were then collected by centrifugation (6500 rpm, 5 min) and washed three times with PBS to remove the remaining culture medium. Then, the microbial solution was diluted to 1.0 × 108 CFU mL−1 with PBS, 100 µL were taken and spread evenly on the agar plate, 10 µL of random copolymers (10 mg mL−1 in PBS or 10% DMSO in PBS was used as a solvent for insoluble samples) with the same content were added to different positions of the plate, and then the plate was incubated at 37 °C for 18 hours; the antibacterial activity was evaluated by measuring the inhibition zone.The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) tests
Random copolymers were initially dissolved in PBS to achieve a stock concentration of 20 mg mL−1. For polymers that were insoluble in PBS, PBS containing up to 10% DMSO was used to ensure complete dissolution. Serial 2-fold dilutions of polymers were prepared from the stock solutions in PBS and transferred to the 96-well sterile round-bottom polypropylene plate for final concentrations of 7.8–2000 μg mL−1 (10 μL per well). PBS and 10% DMSO in PBS were used as solvent controls. Bacteria were grown overnight (∼18 hours) in MHB at 37 °C, and fresh colonies were then collected by centrifugation (6500 rpm, 5 min) and washed three times with PBS to remove the remaining culture medium. Then, the microbial solution was diluted to 4 × 105 CFU mL−1 (the counting and the culture dilution were performed using Bürker-Türk counting chambers) with MHB. The bacterial suspension (90 μL per well) was transferred to a 96-well sterile round-bottom polypropylene plate. Then, the polymer solution was diluted to a final concentration of 7.8–2000 μg mL−1 (10 μL per well). The plate was sealed with parafilm and incubated for 18 hours at 37 °C without shaking. The MIC was defined as the lowest concentration of polymers to completely inhibit bacterial growth, as indicated by the lack of turbidity.After the MIC test, the MBC test was carried out directly. For each sample, from the concentration where no bacterial growth was observed to the concentration where turbid growth occurred, 10 μL of each test concentration was spotted on an agar plate and incubated at 37 °C for 24 hours. The lowest concentration where no bacteria survived was defined as the MBC.
Cytotoxicity assay
The Cell Proliferation Kit XTT56,57 (PanReac AppliChem) was used to assess in vitro cytotoxicity using mouse fibroblast cells (L929). L929 cells were grown in complete medium (RPMI 1640, Gibco), until reaching subconfluency at 37 °C and 5% CO2 in a cell culture incubator (Eppendorf CellXpert C170i). A 96-well cell culture plate was used to seed the cells (104 cells per well). After 24 h, the growth medium was replaced with RPMI 1640 medium consisting of different polymer concentrations, and two-fold serial dilutions were performed starting from 5000 µg mL−1 with/without the polymer (with PBS-treated wells as the negative control). Cell viability was normalized to the negative control and defined as 100%. After a further 24 hours of incubation in the incubator (the polymer continued to expose cells), 50 µl of reaction solution (from the kit) was added to each well and the plate was placed in the incubator for 2 hours. The absorbance of the samples relative to the background control was measured at 475 nm using a microplate reader (BioTek Synergy H1) at a wavelength of 475 nm. In order to measure reference absorbance (to measure non-specific readings), a wavelength of 660 nm was used and subtracted from the 475 nm measurement.The following formula was used to calculate the percentage of cell viability:
Live/dead membrane integrity assay
Bacterial cell membrane integrity was studied using the LIVE/DEAD BacLight Bacterial Viability Kit58,59 (bought from Thermo Fisher Scientific (catalog number L7012)). The kit utilized mixtures of the SYTO9 green fluorescent nucleic acid stain and the red-fluorescent nucleic acid stain, propidium iodide. Bacterial cells with a compromised membrane that were dead or dying would stain red with propidium iodide, whereas cells with an intact membrane would stain green with SYTO9. The bacterial suspension of 108 cells mL−1 in PBS was incubated in the absence (as control) or presence of different concentrations (MIC and 4× MIC) of polymer for 4 h at 37 °C. After that the bacteria were harvested by centrifugation and washed with PBS 3 times. Then, the bacteria were resuspended in a 0.9% salt solution to reach a final density of 1 × 106 cells mL−1. 10 µL of the bacterial suspension was added to 3 µL of the dye mixture (SYTO9 with propidium iodide 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Sample solutions were mixed well and incubated in the dark at room temperature for 15 min. Then, they were examined by confocal laser scanning microscopy (CLSM, DM4000B, Leica).
1). Sample solutions were mixed well and incubated in the dark at room temperature for 15 min. Then, they were examined by confocal laser scanning microscopy (CLSM, DM4000B, Leica).
Scanning electron microscopy (SEM)
Bacteria were collected by centrifugation, and washed cells (after washing with PBS three times) were adhered to slides and fixed with 2% glutaraldehyde for 10 min at 4 °C. They were then washed with 0.1 M cacodylate buffer, incubated with 1% OsO4 in 0.1 M cacodylate buffer for 1 h at room temperature, washed with water, and then dehydrated using a series of graded ethanol solutions (30, 50, 70%) for 15 min each at room temperature, and finally, they were dehydrated three times with 100% ethanol for 30 min each and then dried. The dehydrated samples were gold-coated for 20 s and imaged under high vacuum using a Supra 55 Scanning Electron Microscope (Zeiss).Lipid vesicle preparation
A lipid mixture composed of E. coli polar lipids and egg PC in a 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio was prepared for vesicle formation. E. coli polar lipids were obtained from a commercial total lipid extract (Avanti Polar Lipids) by acetone precipitation followed by diethyl ether extraction, as described in ref. 60. Both E. coli lipids and egg PC were dissolved in chloroform and mixed, and the solvent was evaporated using a rotary evaporator (Büchi Labortechnik AG). The resulting lipid film was washed with diethyl ether, then hydrated in 50 mM potassium phosphate (pH 7.0) to a final concentration of 20 mg mL−1. The lipid suspension was sonicated using a tip sonicator (70% amplitude, 15 s on/45 s off, 16 cycles) while kept on ice, and then subjected to three freeze–thaw cycles (liquid nitrogen and room-temperature water bath). The prepared large-unilamellar vesicles (LUVs) were aliquoted and stocked in liquid nitrogen to prevent oxidation.
1 molar ratio was prepared for vesicle formation. E. coli polar lipids were obtained from a commercial total lipid extract (Avanti Polar Lipids) by acetone precipitation followed by diethyl ether extraction, as described in ref. 60. Both E. coli lipids and egg PC were dissolved in chloroform and mixed, and the solvent was evaporated using a rotary evaporator (Büchi Labortechnik AG). The resulting lipid film was washed with diethyl ether, then hydrated in 50 mM potassium phosphate (pH 7.0) to a final concentration of 20 mg mL−1. The lipid suspension was sonicated using a tip sonicator (70% amplitude, 15 s on/45 s off, 16 cycles) while kept on ice, and then subjected to three freeze–thaw cycles (liquid nitrogen and room-temperature water bath). The prepared large-unilamellar vesicles (LUVs) were aliquoted and stocked in liquid nitrogen to prevent oxidation.
Liposome preparation for a calcein leakage experiment
A 100 mM calcein stock solution was prepared in 50 mM potassium phosphate buffer (KPi) and adjusted to pH 7.0 using 2 M NaOH (Sigma-Aldrich). Liposomes were resuspended in a solution containing 70 mM calcein and 20 mM KPi (pH 7.0) to a final lipid concentration of 5 mg mL−1. Calcein, at a self-quenching concentration, was encapsulated into the vesicles by subjecting the suspension to three freeze–thaw cycles at 30 °C. Following this, the vesicle suspension was extruded 13 times through a 200 nm polycarbonate membrane (Whatman, GE Healthcare) to obtain large-unilamellar vesicles (LUVs). Unencapsulated calcein was removed via size-exclusion chromatography using a 22 cm Sephadex G-75 column (Sigma-Aldrich) pre-equilibrated with an iso-osmotic buffer composed of 50 mM KPi (pH 7.0) plus 50 mM sucrose. Fractions (1 mL each) containing calcein-loaded liposomes were identified under ultraviolet illumination and pooled. The final vesicle suspension was diluted to a total volume of 12 mL in the same buffer (KPi 50 mM, pH 7.0, sucrose 50 mM).Calcein leakage experiment
The ability of the polymer to induce membrane lysis or pore formation was assessed using a calcein leakage assay. Calcein-loaded vesicles were incubated with increasing concentrations of the polymer (0–1000 µg mL−1), and the release of calcein was monitored as an increase in fluorescence intensity. Measurements were performed using a Spark Multimode Plate Reader (TECAN) at 25 °C, with an excitation wavelength of 495 nm and emission recorded at 517 nm. Fluorescence was measured every minute for a total of 30 minutes, until a fluorescence plateau was reached. Afterward, Triton X-100 (50% w/w) was added to fully disrupt the vesicles and determine the maximum fluorescence signal, F(max).The percentage of calcein leakage was calculated according to the following equation:
Each measurement was performed in triplicate.
3. Results and discussion
Polymer synthesis and characterization
In this study, the synthetic route shown in Fig. 1 was used to functionalize the bio-based starting material by introducing carbon–carbon double bonds into the Boc-protected amino acid moieties (Ala, methyl; Phe, aromatic; Lys, cationic) and THG. The 1H and 13C NMR characterization studies confirmed that the monomer was successfully synthesized (Fig. S1–S10). Then, a library of amphiphilic copolymers was successfully synthesized using Reversible Addition–Fragmentation chain-Transfer (RAFT) polymerization (Fig. 1).61 Specifically, to closely approximate the molar mass of antimicrobial peptides (several kDa) and avoid increased cytotoxicity associated with higher molar mass, all polymers were prepared with a target DP of 20, with the Chain Transfer Agent (CTA) constituting 5% of the total molar amount of monomers. The polymers include three components: (1) cationic groups, represented by three amino acids with different side chains – Ala (methyl side chain), Phe (aromatic phenyl ring), or Lys (positively charged amino side chain); (2) hydrophobic monomer, derived from bio-based THG and modified with a double bond to form THGA; and (3) end-groups from the CTA, specifically CDPA, which contains a long aliphatic chain. | ||
| Fig. 1 Schematic diagram of the synthesis of bio-based monomers and amphiphilic random copolymers by RAFT polymerization. | ||
By adjusting the ratio of cationic monomers to hydrophobic monomers, a series of amphiphilic random copolymers with varying molar ratios between the two components were synthesized to study their structure–activity relationships. Based on the cationic component, the polymers were classified into three major families: alanine group (A family), phenylalanine group (P family), and lysine group (L family). The hydrophobic monomer was THGA for all polymers. The polymers were named P(x-r-T)y, where x indicates the type of amino acid (A, P or L), and y is a number from 1 to 9, corresponding to the relative amount of cationic monomer (from 10 to 90%). For example, in P(A-r-T)1, A corresponds to the cationic monomer with Ala, r stands for random, T corresponds to the hydrophobic monomer THGA, and number 1 indicates 10 mol% of the cationic monomer (and 90 mol% of the hydrophobic one).
All the copolymers (with and without Boc deprotection) prepared via RAFT polymerizations were analyzed by 1H NMR and SEC to determine the monomer conversion (% conv.), molecular weights (Mn and Mw), copolymer composition (MPTHGA (%)), and Đ (Table 1). The 1H NMR analysis results of the reaction solutions before and after the reaction were compared, and the monomer conversion was calculated by comparing the vinyl proton signals at 5.5 and 6.3 ppm with the solvent peak (Fig. S11), indicating that the monomer conversion of all prepared polymers exceeded 80%, with only the reaction monomer conversion of P(A-r-T)3 being 75.2%. The Mn value was estimated by SEC and 1H NMR with good agreement between theoretical Mn and experimental values. The average DP was calculated by comparing the integral peak areas of the side chain terminal groups and main chain end groups (Fig. S13, S15 and S17). The MPTHGA value was determined by comparing the signals generated by the two different side chains in the 1H NMR spectra. For each copolymer series, the fraction of hydrophobic repeating units (THGA) in the copolymer, MPT, ranged from 10% to 90%, consistent with the monomer feed ratios. The SEC results (Fig. S21) showed that the molecular weights (3 to 4 kDa) and Đ values (1.2 to 1.4) indicate that the polymerization reaction was well controlled. After deprotection of the Boc group with TFA, the signal at about 1.4 ppm (attributed to the protons of the tert-butyl group) disappeared in the 1H NMR spectrum of the polymer, confirming successful removal of the Boc protecting group.
| Random copolymers | Theoretical composition (%) | % conv.a | Boc-protected | Deprotected | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
MPTHGA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b (%) b (%) |
Mn![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) c c |
DPd | Mn![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) e e |
Mw![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) e e |
Đe | MPTHGA![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b (%) b (%) |
Mn![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) c c |
DPd | Mn![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) e e |
Mw![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) e e |
Đe | H2O sol | Dh![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) f (nm) f (nm) |
ζg (mV) | |||
| a % conv. (% monomer conversion during polymerization) determined by 1H NMR peak integration analysis.b MPTHGA (mole percentage of hydrophobic side chains) determined by 1H NMR peak integration analysis.c Mn (number average molecular weight) determined by 1H NMR peak integration analysis.d DP (degree of polymerization) values determined by 1H NMR peak integration analysis.e Mn, Mw, and dispersity (Đ) determined by SEC analysis in DMF against PMMA standards.f Dh (hydrodynamic number average diameter size) determined using a Malvern Zetasizer Nano ZS at a concentration of 0.1 mg mL−1 in PBS.g ζ (zeta potential) determined using a Malvern Zetasizer Nano ZS at a concentration of 0.1 mg mL−1 in PBS. nd – cannot be determined, indicating one of the following: (i) the NMR peak assignment is unclear because the polymer aggregates in the deuterated reagent after deprotection; (ii) the insolubility of the polymer in THF prevents SEC measurement; or (iii) the insolubility of the polymer in PBS prevents DLS measurement. | |||||||||||||||||
| P(A-r-T)1 | 10(Ala)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90(THGA) 90(THGA) |
90.2 | 85 | 3750 | 14 | 3400 | 4080 | 1.20 | 93 | 3570 | 14 | 2800 | 3900 | 1.41 | ✗ | nd | nd |
| P(A-r-T)3 | 30(Ala)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 70(THGA) 70(THGA) |
75.2 | 72 | 3660 | 13 | 3900 | 5000 | 1.28 | 75 | 3320 | 13 | 3300 | 4400 | 1.35 | ✗ | nd | nd |
| P(A-r-T)5 | 50(Ala)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50(THGA) 50(THGA) |
90.9 | 49 | 4180 | 14 | 5000 | 6200 | 1.25 | 50 | 3350 | 14 | 4000 | 5800 | 1.45 | ✗ | nd | nd |
| P(A-r-T)7 | 70(Ala)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30(THGA) 30(THGA) |
83.9 | 30 | 5290 | 17 | 5000 | 6200 | 1.25 | 34 | 3960 | 17 | 3600 | 5000 | 1.38 | ✓ | 25.6 | nd |
| P(A-r-T)9 | 90(Ala)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10(THGA) 10(THGA) |
89.6 | 13 | 5709 | 18 | 5000 | 6200 | 1.23 | 26 | 3420 | 16 | 3500 | 4900 | 1.41 | ✓ | 53.9 | +8.2 |
| P(P-r-T)1 | 10(Phe)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90(THGA) 90(THGA) |
87.4 | 91 | 3620 | 15 | 3500 | 4400 | 1.25 | 93 | nd | nd | 3000 | 4000 | 1.35 | ✗ | nd | nd |
| P(P-r-T)3 | 30(Phe)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 70(THGA) 70(THGA) |
91.8 | 66 | 4180 | 14 | 4500 | 5500 | 1.22 | 76 | 4140 | 15 | 3400 | 4600 | 1.37 | ✗ | nd | nd |
| P(P-r-T)5 | 50(Phe)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50(THGA) 50(THGA) |
92.7 | 47 | 5190 | 15 | 4800 | 6100 | 1.27 | 54 | 4350 | 15 | 3700 | 4900 | 1.31 | ✗ | nd | nd |
| P(P-r-T)7 | 70(Phe)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30(THGA) 30(THGA) |
93.5 | 27 | 5170 | 14 | 5200 | 6400 | 1.25 | 34 | nd | nd | 3700 | 5000 | 1.35 | ✗ | nd | nd |
| P(P-r-T)9 | 90(Phe)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10(THGA) 10(THGA) |
94.3 | 11 | 5520 | 14 | 5500 | 6600 | 1.18 | 11 | nd | nd | 3400 | 4400 | 1.28 | ✗ | nd | nd |
| P(L-r-T)1 | 10(Lys)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 90(THGA) 90(THGA) |
91.6 | 84 | 5480 | 19 | 5100 | 5600 | 1.10 | 91 | 4980 | 20 | nd | nd | nd | ✗ | nd | nd |
| P(L-r-T)3 | 30(Lys)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 70(THGA) 70(THGA) |
94.2 | 71 | 6820 | 21 | 5200 | 5800 | 1.12 | 81 | 4560 | 18 | nd | nd | nd | ✗ | nd | nd |
| P(L-r-T)5 | 50(Lys)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50(THGA) 50(THGA) |
96.8 | 51 | 5760 | 16 | 5900 | 7100 | 1.09 | 53 | nd | nd | nd | nd | nd | ✓ | 48.1 | +21.6 |
| P(L-r-T)7 | 70(Lys)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30(THGA) 30(THGA) |
93.5 | 34 | 7070 | 18 | 6600 | 7200 | 1.08 | 29 | nd | nd | nd | nd | nd | ✓ | 59.0 | +18.5 |
| P(L-r-T)9 | 90(Lys)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10(THGA) 10(THGA) |
93.3 | 19 | nd | nd | 6300 | 7300 | 1.15 | 13 | nd | nd | nd | nd | nd | ✓ | 64.6 | +13.4 |
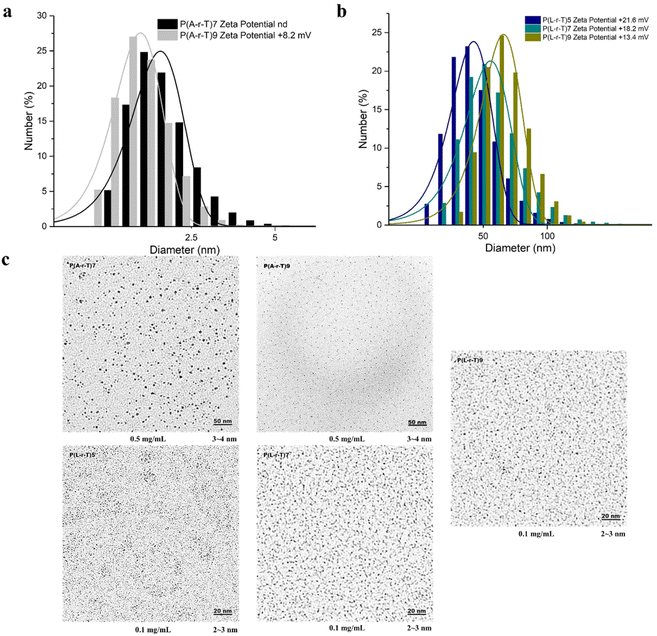
The water solubility of the random copolymers also correlates with the increasing content of cationic components. In the Ala series random copolymers, which have methyl side chains, the polymers become water-soluble when the cationic content reaches approximately 70%. In contrast, the Phe series polymers, bearing more hydrophobic phenyl side chains, remain insoluble regardless of cationic content. For the Lys series, which contains 2 cationic side chains per monomer, water solubility is achieved when the cationic component reaches around 50%. Subsequently, the amphiphilic polymers that could be dissolved in water were also characterized in aqueous media. All characterized polymers have positive ζ (8–21 mV) due to the cationic charge of the amino groups (Table 1). These low ζ values may be due to the difficulty of random copolymers in forming uniform and stable micelles and the partial shielding of cationic charges by polymer intra- and inter-chain interactions. It can be pointed out here that lower stability of the aggregates should be beneficial for the antimicrobial activity, as the polymers most likely interact with the membrane as single chains. Random copolymers with Ala and Lys as cationic components all showed a clear trend of increasing self-assembled aggregate size with the increase of cationic components (Ala or Lys) (Fig. 2a and b), suggesting that the more cationic polymers have larger aggregation numbers. Despite the lower ζ, TEM images confirmed the formation of sufficiently stable nanoscale aggregates (Fig. 2c). The aggregates seem irregular in shape and have sizes below 10 nm. This is obviously smaller than the hydrodynamic sizes obtained by DLS. This is because TEM is performed on dry samples under vacuum, and dehydration can also cause the nanoscale aggregates to shrink or disintegrate. Also, DLS measures the hydrodynamic diameter of the hydrated shell, so the measured diameter is larger than that observed by TEM.62 Furthermore, because the TEM-observed aggregates are only a few nanometers in size, this approaches the resolution limit of TEM, making it difficult to reliably compare aggregate sizes between samples.
Antimicrobial activity
The antimicrobial activity of the polymers was evaluated by determining the MIC (the lowest concentration of the polymer that inhibited visible bacterial growth) and the MBC (the lowest concentration of the polymer at which the bacteria could not survive): the Gram-negative bacterium E. coli (strain ATCC 25922) and the Gram-positive bacterium S. aureus (strain ATCC 6538) (Table 2) were tested. Due to the limited water solubility of the polymers, DMSO was used as a cosolvent.| Random copolymers | MIC (µg mL−1) | MBC (µg mL−1) | ||
|---|---|---|---|---|
| E. coli (ATCC 25922) | S. aureus (ATCC 6538) | E. coli (ATCC 25922) | S. aureus (ATCC 6538) | |
| P(A-r-T)1 | >5000 | 500 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(A-r-T)3 | >5000 | 2000 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(A-r-T)5 | >5000 | 2000 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(A-r-T)7 | >5000 | 500 | >5000 | 1000 |
| P(A-r-T)9 | 2500 | >2000 | 5000 | >2000 |
| P(P-r-T)1 | >5000 | 250 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(P-r-T)3 | >5000 | 2000 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(P-r-T)5 | >5000 | 2000 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(P-r-T)7 | >5000 | 2000 | >10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(P-r-T)9 | 5000 | 1000 | 5000 | >2000 |
| P(L-r-T)1 | >5000 | 500 | 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(L-r-T)3 | >5000 | 1000 | 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
>2000 |
| P(L-r-T)5 | 1000 | 1000 | 5000 | >2000 |
| P(L-r-T)7 | 250 | 2000 | 2000 | >2000 |
| P(L-r-T)9 | 125 | 500 | 2000 | 500 |
As previously mentioned, these kinds of antimicrobial polymers are believed to kill bacteria mainly via membrane disruption.63,64 These amphiphilic polymers contain cationic and hydrophobic groups, each serving a distinct function. The cationic groups attract anionic components on the surface of bacterial cell membranes through electrostatic interactions, promoting the adsorption of the polymer onto the membrane while compromising membrane integrity and disrupting the transport of compounds across it.65–67 Simultaneously, the interaction between the cationic groups and the cell membrane induces the polymer to adopt an overall amphiphilic conformation, allowing the hydrophobic groups to insert into the membrane, leading to membrane disruption and cell death.68,69
For the Gram-negative bacteria E. coli, among the random copolymers composed of Ala and Phe, only the samples with the highest cationic content (up to 90%) showed measurable antibacterial activity, which may be strictly correlated with their water solubility; however, their MICs were very high (2500 and 5000 µg mL−1, respectively), indicating that these polymers are unlikely to be suitable for prospective in vivo applications. In contrast, for the Lys-based copolymers, increasing the cationic monomer content gradually enhanced the polymer antibacterial activity, with the MICs decreasing from 1000 µg mL−1 for P(L-r-T)5 to 125 µg mL−1 for P(L-r-T)9. For the Gram-positive bacteria S. aureus, the balance between the cationic part and the hydrophobic chain with Ala, Phe and Lys had a more obvious effect on the MIC. As the content of the hydrophobic chain decreased, the antibacterial activity first weakened and then increased, with the exception of P(A-r-T)9. Among them, the most hydrophobic one, P(P-r-T)1, showed the lowest MIC (250 µg mL−1), followed by P(A-r-T)1, P(A-r-T)7, P(L-r-T)1, and P(L-r-T)9, with MICs of 500 µg mL−1. The MBCs of the random copolymers tested were all higher than the MIC. Only P(L-r-T)9 with the strongest antibacterial activity had the same MBC and MIC against S. aureus, which was 500 µg mL−1. As reported in the previous section, the studied polymers are either insoluble in water, or they form relatively stable aggregates. The low value of ζ suggests that the aggregates are labile enough to allow the polymers to interact with the bacterial membranes as single chains.
The results of the zone of inhibition experiments are shown in Fig. S22 and S23. For E. coli, it can be clearly seen that there was no growth in the areas where P(L-r-T)7 and P(L-r-T)9 were applied, while E. coli grew in the areas where the other samples were applied. For S. aureus, the inhibition zone of P(L-r-T)9 was very obvious, and the inhibition zone of P(P-r-T)1 could also be observed, while the inhibition zones of P(A-r-T)1, P(A-r-T)7 and P(L-r-T)1 needed to be observed very carefully, which may be related to the diffusion behavior of the samples on the agar plate. This also illustrates the limitation of this method, which is only suitable for samples with a strong antibacterial effect and easy diffusion on the agar plate.
Notably, increased hydrophobicity does not universally enhance antimicrobial activity in this system. While moderate amphiphilicity is sufficient for effective activity against S. aureus, overcoming the outer membrane barrier of E. coli requires a substantially higher hydrophilic and cationic content. This observation highlights the crucial role that the fundamental differences in the cell membrane structure of Gram-positive and Gram-negative bacteria play in determining the antimicrobial properties of polymers. For S. aureus, the absence of an outer membrane allows amphiphilic polymers with moderate hydrophobicity and cationic content to access and disrupt the cytoplasmic membrane relatively efficiently, leading to higher overall activity across much of the polymer library. In contrast, E. coli possesses a lipopolysaccharide-rich outer membrane that constitutes a significant permeability barrier. In this case, initial electrostatic interactions and charge accessibility dominate antimicrobial efficacy. Polymers with higher hydrophobic content may experience reduced aqueous dispersibility or partial self-association, limiting effective interaction with the negatively charged outer membrane. Only polymers with sufficiently high hydrophilic and cationic content remain well dispersed and exhibit enhanced activity against E. coli.
Cytotoxicity
Since there are many reports that cationic polymers can cause cytotoxicity by increasing the permeability of mammalian cell membranes, and the change in membrane permeability depends largely on the number of cationic charges on the polymer surface,70,71 in order to evaluate the toxicity characteristics of such polymers in future applications, the cell viability of mouse fibroblasts (L929) after 24 h of exposure to the random copolymers was tested, and the results are shown in Fig. 3. The cell viability after 24 hours showed the same pattern for random copolymers composed of all cationic species in this study (Ala, Phe and Lys). In random polymers with lower cationic component content, no cytotoxicity was shown at a concentration of up to 5000 µg mL−1, while the cytotoxicity gradually increased with the increase of cationic component content. Similar results have been reported in the literature previously.40,47 Polymers with higher cationic content contain more protonated units, which catalyze the acid hydrolysis of ester bonds in the phospholipid bilayer of the cell membrane to form single-chain lipids and fatty acids. Then, the single-chain lipids will quickly destroy the stability of the membrane and cause cytotoxicity.72,73 For random copolymers with Ala and Phe as cations, among the tested polymers, only the polymer with a cationic content of 90% showed certain cytotoxicity at concentrations of 5000 and 2500 µg mL−1, respectively. It is worth mentioning that for the P(P-r-T)9 sample, the cell viability at a concentration of 2500 µg mL−1 was just 50%. For random copolymers with Lys as the cation, copolymers with more than 50% cationic components began to show cytotoxicity at a concentration of 5000 µg mL−1, while cytotoxicity was reduced at 2500 µg mL−1, and cell viability remained above 50%.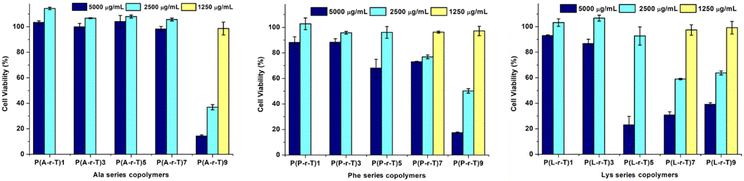 | ||
| Fig. 3 The cytotoxicity of random copolymers with different cationic monomers and monomer feed ratios on cultured mouse fibroblasts (L929) after 24 h of incubation. | ||
Selectivity to bacteria over mammalian cells
To examine the selective activity of random copolymers to bacterial cells over mammalian cells, we determined the IC50/MIC ratio as a selectivity measure (Table 3). In general, copolymers with Ala and Phe as cationic components have weak antibacterial activity against Gram-negative bacteria E. coli (MIC ≥ 2500 µg mL−1), and their selectivity indexes for E. coli are very small (≤0.5), indicating that these polymers have no selectivity for E. coli compared to animal cells. However, copolymers with Lys as the cationic component have obvious selectivity for E. coli over animal cells, which increases with the increase of the content of the cationic component. The selectivity coefficient increases from 2.5 for P(L-r-T)5 to 20 for P(L-r-T)9. For Gram-positive bacteria S. aureus, only P(A-r-T)9 and P(L-r-T)7 showed low or no selectivity among the tested polymers. The other samples showed certain selectivity; in particular, when the content of cationic components was small (10%), their selectivity was larger than 10 and the selectivity of Phe (with a hydrophobic benzene ring on the side chain) series polymers of the cationic component was larger than 20. It is worth mentioning that P(L-r-T)9 showed certain selectivity for both E. coli and S. aureus over L929 cells, with values of 20 and 5, respectively, indicating the possibility that it can be used to treat bacterial infections. Therefore, the subsequent antibacterial mechanism research was carried out using P(L-r-T)9.| Random copolymers | MIC (µg mL−1) | IC50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a (µg mL−1) a (µg mL−1) |
Selectivityb | ||
|---|---|---|---|---|---|
| E. coli (ATCC 25922) | S. aureus (ATCC 6538) | L929 | E. coli (ATCC 25922) | S. aureus (ATCC 6538) | |
| a Concentration for a 50% reduction in cell viability.b Selectivity was calculated as IC50/MIC. | |||||
| P(A-r-T)1 | >5000 | 500 | >5000 | — | >10 |
| P(A-r-T)3 | >5000 | 2000 | >5000 | — | >2.5 |
| P(A-r-T)5 | >5000 | 2000 | >5000 | — | >2.5 |
| P(A-r-T)7 | >5000 | 500 | >5000 | — | >10 |
| P(A-r-T)9 | 2500 | >2000 | 1250 | 0.5 | <0.625 |
| P(P-r-T)1 | >5000 | 250 | >5000 | — | >20 |
| P(P-r-T)3 | >5000 | 2000 | >5000 | — | >2.5 |
| P(P-r-T)5 | >5000 | 2000 | >5000 | — | >2.5 |
| P(P-r-T)7 | >5000 | 2000 | >5000 | — | >2.5 |
| P(P-r-T)9 | 5000 | 1000 | 2500 | 0.5 | 2.5 |
| P(L-r-T)1 | >5000 | 500 | >5000 | — | >10 |
| P(L-r-T)3 | >5000 | 1000 | >5000 | — | >5 |
| P(L-r-T)5 | 1000 | 1000 | 2500 | 2.5 | 2.5 |
| P(L-r-T)7 | 250 | 2000 | 2500 | 10 | 1.25 |
| P(L-r-T)9 | 125 | 500 | 2500 | 20 | 5 |
Bacterial membrane integrity assay
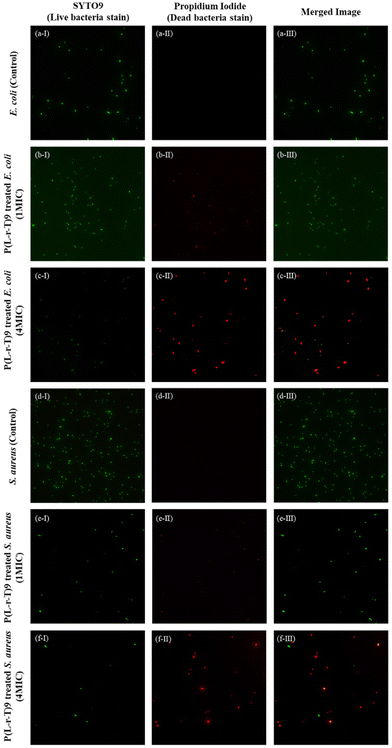
Several researchers have proposed that the bactericidal mechanism of these amphiphilic antimicrobial agents involves an initial electrostatic interaction between the cationic components and the microbial cell membrane. This is followed by the insertion of hydrophobic groups, which disrupts membrane permeability, leading to a breakdown of transmembrane potential and leakage of intracellular contents, ultimately resulting in cell death.65–67 To further investigate the said mechanism, bacterial viability was assessed using the Thermo Fisher Scientific LIVE/DEAD BacLight Kit to evaluate the integrity of bacterial cell membranes following treatment with the polymer P(L-r-T)9. This kit includes two dyes: green-fluorescent SYTO 9, which labels all bacterial cells in the sample, including those with intact and damaged membranes, and red-fluorescent propidium iodide (PI), which selectively penetrates only bacteria with compromised membranes.In the control group, E. coli and S. aureus cells not treated with the polymer P(L-r-T)9 exhibited strong green fluorescence with no red fluorescence, indicating that all bacteria were alive with intact cell membranes (Fig. 4a-I, a-II, a-III, d-I, d-II, and d-III). In contrast, cells exposed to a 1MIC of the polymer solution for 4 hours showed reduced green fluorescence along with some red fluorescence signals, suggesting partial membrane damage; however, a substantial number of bacteria remained viable at this concentration (Fig. 4b-I, b-II, b-III, e-I, e-II, and e-III). At the higher concentration of 4MIC, the red fluorescence became more prominent, indicating extensive membrane damage and bacterial cell death, although some cells still survived (Fig. 4c-I, c-II, c-III, f-I, f-II, and f-III). These observations suggest that the polymer exerts bactericidal activity by disrupting bacterial cell membranes, which was further confirmed with SEM.
SEM can be employed to analyze the antimicrobial effects of synthetic polymers by observing characteristic morphological changes induced on bacterial membrane surfaces (Fig. 5). Untreated E. coli and S. aureus control cells exhibited bright, smooth, and undamaged membrane surfaces, as shown in the SEM micrographs (Fig. 5a-I and b-I). However, after treatment with P(L-r-T)9 at a concentration equal to MIC for 4 hours, noticeable alterations were observed. The surface of E. coli showed distinct depressions and the presence of particulate structures, while the membrane of S. aureus appeared rough and wrinkled and featured some vesicle formation, with even visible membrane disruptions in some cases. At a higher concentration of 4MIC for 4 hours, the membranes of both E. coli and S. aureus exhibited widespread damage and rupturing, resulting in collapsed cells.
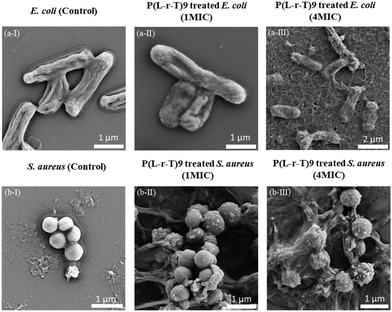 | ||
| Fig. 5 SEM images of E. coli (from a-I to a-III) and S. aureus (from b-I to b-III) treated with P(L-r-T)9 at different concentrations (MIC and 4× MIC) for 4 h. | ||
In summary, these observations indicate that P(L-r-T)9 induces significant characteristic changes in the bacterial membrane surfaces, as evident from the SEM and live/dead staining. This leads to membrane rupture and the leakage of intracellular contents, ultimately causing bacterial cell death. To further support the cell membrane breaking mechanism, a study of the interaction of the most active polymers, P(L-r-T)7 and P(L-r-T)9, with E. coli![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) PC vesicles was investigated using a calcein leakage experiment as described in the Materials and methods. Both polymers, P(L-r-T)7 and P(L-r-T)9, induced a concentration-dependent increase in calcein leakage (Fig. 6), indicating their ability to permeabilize the lipid bilayer and promote pore formation or membrane destabilization. However, their leakage profiles differed in magnitude and threshold of activity.
PC vesicles was investigated using a calcein leakage experiment as described in the Materials and methods. Both polymers, P(L-r-T)7 and P(L-r-T)9, induced a concentration-dependent increase in calcein leakage (Fig. 6), indicating their ability to permeabilize the lipid bilayer and promote pore formation or membrane destabilization. However, their leakage profiles differed in magnitude and threshold of activity.
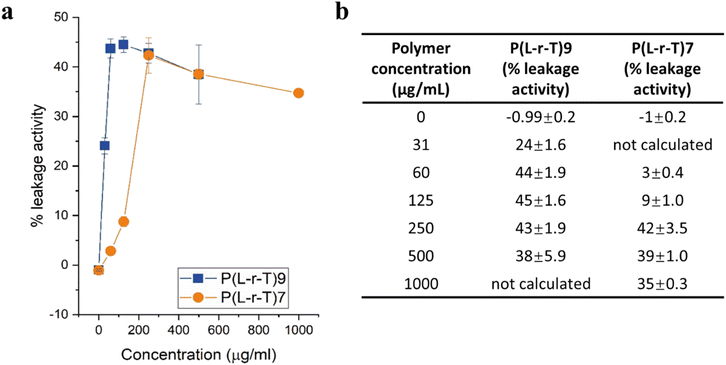
For P(L-r-T)7, the leakage activity progressively increased with concentration, reaching a maximum of 42% at 250 µg mL−1. Notably, this concentration corresponds to the previously determined minimum inhibitory concentration (MIC) for this polymer, suggesting that membrane permeabilization may be directly linked to its antibacterial efficacy.
In contrast, P(L-r-T)9 showed significant leakage (44%) already at 60 µg mL−1, below its MIC of 250 µg mL−1. At 125 and 250 µg mL−1, the leakage values remained similar (45% and 43%, respectively), indicating an early saturation of membrane-disruptive activity. This earlier onset of leakage may reflect a more specific or efficient interaction of P(L-r-T)9 with the lipid membrane compared to P(L-r-T)7. Interestingly, at concentrations above 250 µg mL−1, both polymers showed a plateau or slight decrease in leakage activity. This non-linear trend suggests that optimal membrane interaction occurs at intermediate concentrations. Several mechanisms may underlie the reduced leakage at higher polymer doses, including polymer aggregation, limiting effective interaction with the bilayer; membrane rigidification, leading to stabilization rather than disruption; and direct interaction with calcein, potentially quenching fluorescence without additional leakage. Further investigation of this phenomenon is outside the scope of this work but may represent a follow-up study.
Overall, these results confirm that both polymers can disrupt lipid membranes in a concentration-dependent manner, with P(L-r-T)9 displaying earlier and more potent activity, possibly due to its enhanced membrane targeting properties. In contrast, P(L-r-T)7 exhibits its maximum leakage activity at 250 µg mL−1, which coincides with its MIC, suggesting a direct correlation between membrane permeabilization and antimicrobial efficacy for this polymer.
4. Conclusions
In summary, we successfully synthesized a library of 15 statistical amphiphilic random copolymers by using bio-based monomers, utilizing amino acids with different side chains (as cationic components) and tetrahydrogeraniol (as the hydrophobic component) as starting materials, turning them into methacrylates, and employing RAFT polymerization. These copolymers were screened against Gram-negative E. coli and Gram-positive S. aureus, as well as mouse fibroblast cells (L929), to evaluate their antimicrobial activity and cytotoxicity. We systematically examined the impact of the monomer ratio and the type of amino acid side chain (Ala, methyl; Phe, aromatic; Lys, cationic) on both antimicrobial activity and cytotoxicity. The results demonstrated that for E. coli, only the copolymers with the highest cationic content (up to 90%) from Ala and Phe exhibited antimicrobial activity. In contrast, the random copolymers composed of Lys showed gradually enhanced antimicrobial activity as the cationic content increased. For E. coli, there seems to be a direct correlation between water solubility and antimicrobial activity, but not for S. aureus. In this case, the balance between cationic components and hydrophobic chains influenced the MIC, with antimicrobial activity first decreasing and then increasing as the hydrophobic chain content decreased, with P(A-r-T)9 being the only outlier. In general, low solubility and polymer aggregation may be limiting factors in antimicrobial activity. Cytotoxicity also increased with higher cationic content. Among all the copolymers, P(L-r-T)9 exhibited notable antimicrobial activity against both E. coli and S. aureus, while demonstrating optimal selectivity for mammalian cells, and it is therefore the best candidate for practical applications. Live/dead staining and SEM results confirmed that its antimicrobial effect was mediated by disruption of the bacterial cell membrane. Consistently, calcein leakage assays in model liposomes composed of E. coli polar lipids and phosphatidylcholine (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) revealed that P(L-r-T)9 induces significant membrane permeabilization already at concentrations below its MIC, supporting a mechanism of action based on membrane destabilization and subsequent bacterial cell lysis. These findings provide valuable insights for the design and synthesis of more sustainable and effective antimicrobial amphiphilic polymers with strong biocompatibility.
1) revealed that P(L-r-T)9 induces significant membrane permeabilization already at concentrations below its MIC, supporting a mechanism of action based on membrane destabilization and subsequent bacterial cell lysis. These findings provide valuable insights for the design and synthesis of more sustainable and effective antimicrobial amphiphilic polymers with strong biocompatibility.
Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including GPC, NMR, SEM, cytotoxicity, and dead cell staining, are available at Figshare at https://doi.org/10.6084/m9.figshare.31852735.Supplementary information (SI): experimental section and characterization details; supplementary synthetic routes of amphiphilic antimicrobial random copolymers; 1H and 13C NMR spectra and analyses; and SEC and inhibition zone test results of polymers. See DOI: https://doi.org/10.1039/d6py00298f.
Acknowledgements
The first author of this work was financially supported by the China Scholarship Council (CSC) under Grant Number 202204910075. We thank the European Union's Horizon 2020 research and innovation program, under the Marie Skłodowska-Curie grant agreement BioInspireSensing No. 955643, for supporting the research of Caterina Presutti.References
- P. Sarmah, M. M. Dan, D. Adapa and T. Sarangi, Electron. J. Biol., 2018, 14, 50–58 Search PubMed.
- D. M. Brogan and E. Mossialos, Global. Health, 2016, 12, 8 CrossRef.
- M. Kumar, D. K. Sarma, S. Shubham, M. Kumawat, V. Verma, P. B. Nina, D. Jp, S. Kumar, B. Singh and R. R. Tiwari, Front. Microbiol., 2021, 12, 609459 CrossRef PubMed.
- S. Shahzadi, N. Zafar and R. Sharif, in Bacterial Pathogenesis and Antibacterial Control, Intechopen, London, 2018, 51, DOI:10.5772/intechopen.72526.
- R. Rawashdeh and Y. Haik, Dyn. Biochem. Process Biotechnol. Mol. Biol., 2009, 3, 12–20 Search PubMed.
- Y. Yan, Y. Li, Z. Zhang, X. Wang, Y. Niu, S. Zhang, W. Xu and C. Ren, Colloids Surf., B, 2021, 202, 111682 CrossRef CAS PubMed.
- J. K. Boparai and P. K. Sharma, Protein Pept. Lett., 2020, 27, 4–16 CrossRef CAS.
- S. Duan, R. Wu, Y. Xiong, H. Ren, C. Lei, Y. Zhao, X. Zhang and F. Xu, Prog. Mater. Sci., 2022, 125, 100887 CrossRef CAS.
- K. Huang, C. Yang, S. Huang, C. Chen, Y. Lu and Y. Lin, Int. J. Mol. Sci., 2016, 17, 1578 CrossRef PubMed.
- L. Timofeeva and N. Kleshcheva, Appl. Microbiol. Biotechnol., 2011, 89, 475–492 CrossRef CAS.
- J. Chen, F. Wang, Q. Liu and J. Du, Chem. Commun., 2014, 50, 14482–14493 RSC.
- T. I. Shabatina, O. I. Vernaya and M. Y. Melnikov, Molecules, 2023, 28, 1603 CrossRef CAS PubMed.
- U. Patel and E. C. Hunt, J. Nanotheranostics, 2023, 4, 429–462 CrossRef.
- M. Manoswini, D. Bhattacharya, P. Sen, N. Ganguly and P. S. Mohanty, Adv. Nat. Sci.: Nanosci. Nanotechnol., 2021, 12, 025003 Search PubMed.
- L. J. Zhang and R. L. Gallo, Curr. Biol., 2016, 26, R14–R19 CrossRef CAS PubMed.
- W. C. Wimley, ACS Chem. Biol., 2010, 5, 905–917 CrossRef CAS PubMed.
- K. Matsuzaki, Biochim. Biophys. Acta, Biomembr., 1999, 1462, 1–10 CrossRef CAS.
- Y. Q. Xiong, K. Mukhopadhyay, M. R. Yeaman, J. Adler-Moore and A. S. Bayer, Antimicrob. Agents Chemother., 2005, 49, 3114–3121 CrossRef CAS PubMed.
- N. Malanovic and K. Lohner, Pharmaceuticals, 2016, 9, 59 CrossRef PubMed.
- V. I. Band and D. S. Weiss, Antibiotics, 2015, 4, 18–41 CrossRef PubMed.
- A. H. Benfield and S. T. Henriques, Front. Med. Technol., 2020, 2, 2020 Search PubMed.
- G. Moad, E. Rizzardo and S. H. Thang, Chem.– Asian J., 2013, 8, 1634–1644 CrossRef CAS PubMed.
- F. L. Hatton, Polym. Chem., 2020, 11, 220–229 RSC.
- N. P. Truong, G. R. Jones, K. G. Bradford, D. Konkolewicz and A. Anastasaki, Nat. Rev. Chem., 2021, 5, 859–869 CrossRef CAS PubMed.
- N. Corrigan, K. Jung, G. Moad, C. J. Hawker, K. Matyjaszewski and C. Boyer, Prog. Polym. Sci., 2020, 111, 101311 CrossRef CAS.
- D. S. Uppu, S. Samaddar, C. Ghosh, K. Paramanandham, B. R. Shome and J. Haldar, Biomaterials, 2016, 74, 131–143 CrossRef CAS PubMed.
- K. E. Locock, T. D. Michl, J. D. Valentin, K. Vasilev, J. D. Hayball, Y. Qu, A. Traven, H. J. Griesser, L. Meagher and M. Haeussler, Biomacromolecules, 2013, 14, 4021–4031 CrossRef CAS PubMed.
- K. Kuroda and W. F. DeGrado, J. Am. Chem. Soc., 2005, 127, 4128–4129 CrossRef CAS PubMed.
- K. Kuroda, G. A. Caputo and W. F. DeGrado, Chem. – Eur. J., 2009, 15, 1123–1133 CrossRef CAS PubMed.
- K. Lienkamp and G. N. Tew, Chem. – Eur. J., 2009, 15, 11784–11800 CrossRef CAS PubMed.
- K. Lienkamp, A. E. Madkour, A. Musante, C. F. Nelson, K. Nusslein and G. N. Tew, J. Am. Chem. Soc., 2008, 130, 9836–9843 CrossRef CAS PubMed.
- G. J. Gabriel, J. A. Maegerlein, C. F. Nelson, J. M. Dabkowski, T. Eren, K. Nüsslein and G. N. Tew, Chem. – Eur. J., 2009, 15, 433–439 CrossRef CAS PubMed.
- H. Takahashi, G. A. Caputo, S. Vemparala and K. Kuroda, Bioconjugate Chem., 2017, 28, 1340–1350 CrossRef CAS PubMed.
- K. Lienkamp, K. N. Kumar, A. Som, K. Nüsslein and G. N. Tew, Chem. – Eur. J., 2009, 15, 11710–11714 CrossRef CAS PubMed.
- J. Pachla, R. J. Kopiasz, G. Marek, W. Tomaszewski, A. Głogowska, K. Drężek, S. Kowalczyk, R. Podgórski, B. Butruk-Raszeja, T. Ciach, J. Mierzejewska, A. Plichta, E. Augustynowicz-Kopeć and D. Jańczewski, Biomacromolecules, 2023, 24, 2237–2249 CrossRef CAS PubMed.
- Z. Cheng and P. Raffa, Mater. Adv., 2025, 6, 4939–4968 RSC.
- P. Pham, S. Oliver, E. H. Wong and C. Boyer, Polym. Chem., 2021, 12, 5689–5703 RSC.
- R. Namivandi-Zangeneh, R. J. Kwan, T.-K. Nguyen, J. Yeow, F. L. Byrne, S. H. Oehlers, E. H. Wong and C. Boyer, Polym. Chem., 2018, 9, 1735–1744 RSC.
- R. Bhat, L. L. Foster, G. Rani, S. Vemparala and K. Kuroda, RSC Adv., 2021, 11, 22044–22056 RSC.
- S. Schaefer, D. Melodia, C. Pracey, N. Corrigan, M. D. Lenardon and C. Boyer, Biomacromolecules, 2024, 25, 871–889 CrossRef CAS PubMed.
- Y. Hu, J. Zhao, J. Zhang, Z. Zhu and J. Rao, ACS Macro Lett., 2021, 10, 990–995 CrossRef CAS PubMed.
- A. Tyagi and A. Mishra, ACS Omega, 2021, 6, 34724–34735 CrossRef CAS PubMed.
- S. Hong, H. Takahashi, E. T. Nadres, H. Mortazavian, G. A. Caputo, J. G. Younger and K. Kuroda, PLoS One, 2017, 12, e0169262 CrossRef PubMed.
- P. T. Phuong, S. Oliver, J. He, E. H. H. Wong, R. T. Mathers and C. Boyer, Biomacromolecules, 2020, 21, 5241–5255 CrossRef CAS PubMed.
- V. Sambhy, B. R. Peterson and A. Sen, Angew. Chem., Int. Ed., 2008, 47, 1250–1254 CrossRef CAS PubMed.
- I. Sovadinova, E. F. Palermo, M. Urban, P. Mpiga, G. A. Caputo and K. Kuroda, Polymers, 2011, 3, 1512–1532 CrossRef CAS.
- E. F. Palermo and K. Kuroda, Biomacromolecules, 2009, 10, 1416–1428 CrossRef CAS PubMed.
- Y. Oda, S. Kanaoka, T. Sato, S. Aoshima and K. J. B. Kuroda, Biomacromolecules, 2011, 12, 3581–3591 CrossRef CAS PubMed.
- T. Maity, S. Paul and P. Y. De, J. Macromol. Sci., Part A:Pure Appl. Chem., 2023, 60, 2–17 CrossRef CAS.
- I. Mukherjee, A. Ghosh, P. Bhadury and P. De, ACS Omega, 2017, 2, 1633–1644 CrossRef CAS PubMed.
- J. Zhu, H. Han, F. Li, X. Wang, J. Yu, C.-C. Chu and D. Wu, J. Colloid Interface Sci., 2019, 540, 634–646 CrossRef CAS PubMed.
- J. Koehbach and D. J. Craik, Trends Pharmacol. Sci., 2019, 40, 517–528 CrossRef CAS PubMed.
- S. Noppalit, A. Simula, N. Ballard, X. Callies, J. M. Asua and L. Billon, Biomacromolecules, 2019, 20, 2241–2251 CrossRef CAS PubMed.
- J. Tan, Y. Zhao, J. L. Hedrick and Y. Yang, Adv. Healthcare Mater., 2022, 11, 2100482 CrossRef CAS PubMed.
- S. G. Roy and P. De, Polymer, 2014, 55, 5425–5434 CrossRef CAS.
- L. M. Jost, J. M. Kirkwood and T. L. Whiteside, J. Immunol. Methods, 1992, 147, 153–165 CrossRef CAS PubMed.
- D. A. Scudiero, R. H. Shoemaker, K. D. Paull, A. Monks, S. Tierney, T. H. Nofziger, M. J. Currens, D. Seniff and M. R. Boyd, Cancer Res., 1988, 48, 4827–4833 CAS.
- M. A. Auty, G. E. Gardiner, S. J. McBrearty, E. O. O'Sullivan, D. M. Mulvihill, J. K. Collins, G. F. Fitzgerald, C. Stanton and R. P. Ross, Appl. Environ. Microbiol., 2001, 67, 420–425 CrossRef CAS PubMed.
- L. Boulos, M. Prévost, B. Barbeau, J. Coallier and R. Desjardins, J. Microbiol. Methods, 1999, 37, 77–86 CrossRef CAS PubMed.
- M. J. Newman and T. H. Wilson, J. Biol. Chem., 1980, 255, 10583–10586 CrossRef CAS PubMed.
- E. F. Palermo, S. Vemparala and K. Kuroda, Biomacromolecules, 2012, 13, 1632–1641 CrossRef CAS PubMed.
- S. C. Owen, D. P. Y. Chan and M. S. Shoichet, Nano Today, 2012, 7, 53–65 CrossRef CAS.
- A. Jain, L. S. Duvvuri, S. Farah, N. Beyth, A. J. Domb and W. Khan, Adv. Healthcare Mater., 2014, 3, 1969–1985 CrossRef CAS PubMed.
- H. Takahashi, G. A. Caputo and K. Kuroda, Biomater. Sci., 2021, 9, 2758–2767 RSC.
- R. F. Epand, B. P. Mowery, S. E. Lee, S. S. Stahl, R. I. Lehrer, S. H. Gellman and R. M. Epand, J. Mol. Biol., 2008, 379, 38–50 CrossRef CAS PubMed.
- H. Murata, R. R. Koepsel, K. Matyjaszewski and A. J. Russell, Biomaterials, 2007, 28, 4870–4879 CrossRef CAS PubMed.
- J. Y. Huang, R. R. Koepsel, H. Murata, W. Wu, S. B. Lee, T. Kowalewski, A. J. Russell and K. Matyjaszewski, Langmuir, 2008, 24, 6785–6795 CrossRef CAS PubMed.
- N. Kawabata and M. Nishiguchi, Appl. Environ. Microbiol., 1988, 54, 2532–2535 CrossRef CAS PubMed.
- P. Pham, S. Oliver and C. Boyer, Macromol. Chem. Phys., 2023, 224, 2200226 CrossRef CAS.
- S. Hong, P. R. Leroueil, E. K. Janus, J. L. Peters, M. M. Kober, M. T. Islam, B. G. Orr, J. R. Baker Jr. and M. M. Banaszak Holl, Bioconjug. Chem., 2006, 17, 728–734 CrossRef CAS PubMed.
- F. Unger, M. Wittmar and T. Kissel, Biomaterials, 2007, 28, 1610–1619 CrossRef CAS PubMed.
- D. M. L. Morgan, V. L. Larvin and J. D. Pearson, J. Cell Sci., 1989, 94, 553–559 CrossRef CAS PubMed.
- D. M. L. Morgan, J. Clover and J. D. Pearson, J. Cell Sci., 1988, 91, 231–238 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |