Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Photobase generators for amino acid N-carboxyanhydride ring-opening photopolymerization: rapid access to degradable polypeptide-based networks
Robert D.
Murphy
 a,
Thiago Ouriques
Machado
a,
Carlo
Gonzato
cd,
Fernando
Vidal
b,
Ander
Leiza
b,
Oihane
Varela
b,
Haritz
Sardon
b,
Olivier
Soppera
a,
Thiago Ouriques
Machado
a,
Carlo
Gonzato
cd,
Fernando
Vidal
b,
Ander
Leiza
b,
Oihane
Varela
b,
Haritz
Sardon
b,
Olivier
Soppera
 cd and
Andreas
Heise
cd and
Andreas
Heise
 *a
*a
aDepartment of Chemistry, RCSI University of Medicine and Health Sciences, Dublin, Ireland. E-mail: andreasheise@rcsi.ie
bPOLYMAT and Department of Polymers and Advanced Materials: Physics, Chemistry and Technology, Faculty of Chemistry, University of the Basque Country UPV/EHU, Paseo Manuel de Lardizábal, 3, 20018, Donostia-San Sebastián, Spain
cUniversité de Haute-Alsace, CNRS, IS2M UMR 7361, Mulhouse 68100, France
dUniversité de Strasbourg, Strasbourg 67000, France
First published on 4th February 2026
Abstract
Conventional resin systems for 3D object fabrication rely predominantly on non-degradable (meth)acrylate networks, despite global concerns about microplastic persistence. Polypeptide-based networks offer an attractive alternative due to their inherent biocompatibility and degradability, yet efficient methods to photochemically access such materials remain severely underdeveloped. In this contribution, photobase generators (PBGs) are introduced as efficient initiating systems for photo-induced N-carboxyanhydride (NCA) ring-opening polymerization (ROP), enabling rapid (<10 min) and spatiotemporally controlled polypeptide synthesis using ultraviolet (365 nm) and visible (405 nm) light irradiation. Incorporation of disulfide-containing difunctional NCA monomers enables photostructuring of crosslinked polymer networks, which can be readily degraded on demand using chemical reductants. This strategy represents the first demonstration of light-triggered NCA ROP by PBGs to provide direct access to photocurable, degradable polypeptide networks. The combination of rapid photopolymerization and reductive degradability shown here may truly expand the utility of NCA ROP systems as next-generation resins for manufacture of 3D structures with on-demand degradability.
Introduction
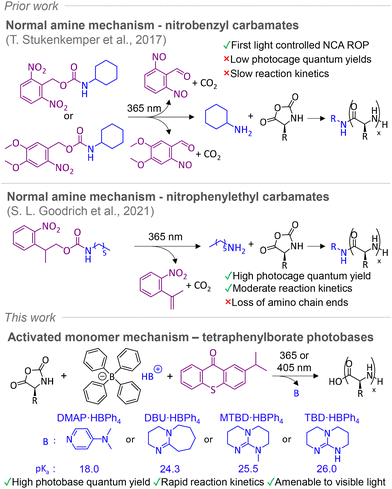
Light is a highly effective stimulus for chemical transformations, due to its low energy input and high spatial and temporal control.1 It represents a powerful tool for polymerization processes, allowing for high reactivity over amenable wavelengths and intensities through specific chromophore selection.2,3 Photopolymerization has revolutionalized processes such as lithographic patterning, coatings, as well as 3D printing.4,5 While most work has focused exclusively on free-radical and (more recently) controlled radical photopolymerization systems,6–8 the utility of photosensitizers has provided a basis for the design of photoacid generators (PAGs) and photobase generators (PBGs) for non-radical based photopolymerizations.9 PAG or PBG systems are designed to absorb at the wavelength of interest,10 liberating ionic species or free Lewis acids/bases which promote cationic or anionic type polymerization.11,12 PAGs for example have found use in ring opening polymerization (ROP) of cyclic monomers including cyclosiloxanes,13 and spiro-orthocarbonates,14 as well as in lithographic technologies.15,16In contrast to PAGs, PBGs are less explored, though significant advances over the past decade have broadened their scope, enhancing efficiency through synthetic development of both ionic and non-ionic systems.17–23 In terms of PBG adaption for ROP, tetraphenylborate salts (HBPh4) of organobases have been used to great effect. Wang and coworkers pioneered their use with 1,5,7-triaza-bicyclo [4.4.0]dec-5-ene (TBD·HBPh4), which undergoes a proton transfer rearrangement upon irradiation at 254 nm, releasing organobase TBD to promote photoinitiated ROP (photo-ROP) of ε-CL.24 Similar strategies have been used for L-lactide25 and cyclosiloxane.26 Despite the advances, there is room to expand the scope of PBGs for the synthesis of other biodegradable polymers, such as polypeptides, via ROP of corresponding amino acid N-carboxyanhydrides (NCAs).27 Synthetic advances over the past decades for NCA ROP,28–30 have presented a scalable, inexpensive method for generating bio-inspired polypeptides with modular side chains,31 which have utility across biomedicine,32–34 3D printing,35–37 and beyond.38,39 Concerning NCA photopolymerization, our group pioneered the use of photocaged amines, using UV light (365 nm) to liberate primary amines for controlled ROP via conventional normal amine mechanism (NAM) to generate polypeptides,40,41 with Goodrich et al. making other notable developments in recent years (Fig. 1).42 However, these reactions are kinetically slow, which is unsuitable for direct fabrication of polypeptide structures by NCA photo-ROP. PBGs as an alternative, may offer improved kinetics via the activated monomer mechanism (AMM), which may provide rapid access to defined polypeptides.43 This could potentially enable the photostructuring of polypeptides in a manner similar to current lithographic or 3D printing technologies.
Herein, we disclose for the first time the use of PBGs for NCA ROP, enabling the synthesis of well-defined synthetic polypeptides through photopolymerization. Through rapid photo-induced proton transfer of tetraphenylborate PBGs, transient organobases could initiate NCA ROP in remarkably short timeframes (min), affording polypeptides with controllable dispersities using UV (365 nm) and visible (405 nm) light. Beyond homopolypeptide formation, this photo-ROP strategy provides, for the first time, a direct route to access 2.5D photostructured polypeptide networks via photo-ROP of difunctional NCAs, which degradable using reductive chemical triggers.
Results and discussion
PBG NCA photo-ROP for synthetic polypeptides
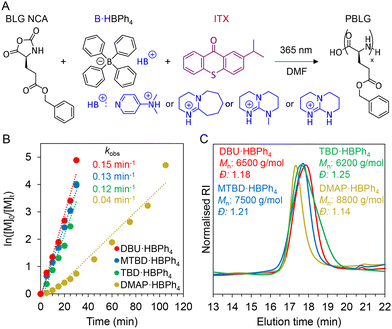
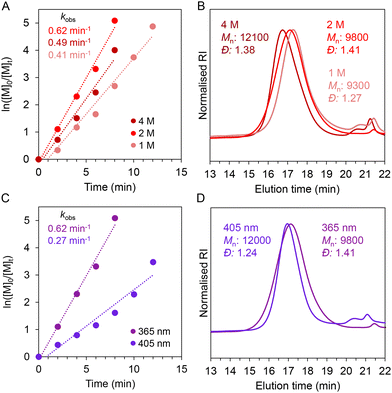
A range of PBGs were initially synthesised by complexing sodium tetraphenylborate (NaBPh4), with four organobases with variable pKa's,44 4-dimethylaminopyridine (DMAP, 18.0), 1,8-diazabicyclo[5.4.0]undec-7-ene (DBU, 24.3), 7-methyl-1,5,7-triazabicyclo[4.4.0]dec-5-ene (MTBD, 25.5), and 1,5,7-triazabicyclo[4.4.0]dec-5-ene (TBD, 26.0). The chemical structures of resulting PBGs – DMAP·HBPh4, DBU·HBPh4, MTBD·HBPh4, and TBD·HBPh4 were confirmed by 1H NMR spectroscopy (Fig. S1–S4). To identify a lead PBG system, benzyl-L-glutamate (BLG) NCA and carbobenzyloxy-L-lysine (ZLL) NCA were synthesized and characterized by 1H NMR spectroscopy (Fig. S5 and S6), before their subsequent use in photo-ROP. All reactions were conducted in a closed vial, using a mounted LED and no stirring (Fig. S7). Initially, variables such as solvent (dimethylformamide – DMF), monomer molarity (0.4 M), monomer/initiator ([M]0/[I]0) ratio (50), irradiation wavelength (365 nm) and intensity (8.0 mW cm−2) were kept constant to determine a lead PBG capable of promoting fastest ROP kinetics of BLG NCA (Fig. 2A). The molar concentration of each respective PBG and photosensitiser, isopropylthioxanthone (ITX), were also kept constant. For PBGs (Table S1, entries 1–4), DBU·HBPh4 promoted the fastest consumption of BLG NCA, confirmed by real-time FTIR spectroscopy, with >99% conversion noted after 30 min (Fig. S8). Both MTBD·HBPh4 and TBD·HBPh4 PBGs had similar conversion (98–99%) over the same timeframe, while 52% conversion was noted for DMAP·HBPh4, which further increased to >99% conversion after a prolonged time (105 min). DBU·HBPh4 exhibited the fastest rate for NCA photo-ROP, having an apparent first-order rate constant kobs, of 0.15 min−1, while MTBD·HBPh4 and TBD·HBPh4 presented similar rates of 0.13 min−1 and 0.12 min−1 respectively (Fig. 2B). For DMAP·HBPh4, a significantly slower kinetic profile was observed (kobs = 0.04 min−1). Nonetheless, formed poly(benzyl-L-glutamate)s (PBLG)s had similar characteristics as determined by 1H and DOSY NMR spectroscopy (Fig. S9–S16), while symmetrical traces and low dispersities (<1.3) were observed by size exclusion chromatography (SEC) (Fig. 2C).The DBU·HBPh4 PBG was then chosen for subsequent tests, to investigate the effect of monomer concentration on photo-ROP kinetics. At molarities of 1 M and 2 M, BLG NCA was fully converted in 12 and 8 min (Fig. S17), exhibiting kobs rates of 0.41 min−1 and 0.62 min−1 respectively (Fig. 3A). At 4 M, NCA conversion was >98% after 8 min and had a slower kobs of 0.49 min−1, likely due to precipitation of the polypeptide during polymerization (Fig. S18). SEC traces were similar for all tested concentrations, though higher dispersities were noted for higher molarities (Fig. 3B, Table S1, entries 5–7). An NCA concentration of 2 M was maintained for further experiments, as it could facilitate fast polymerization forming polypeptides with acceptable dispersities. Photo-ROP of ZLL NCA was also possible using DBU·HBPh4/ITX, undergoing full conversion to PZLL in 14 min (Fig. S19A), which afforded symmetrical SEC traces (Fig. S19B). The ITX photosensitiser enabled polymerization with visible light also (405 nm, 10.0 mW cm−2). Full conversion of NCAs took 4 min longer (12 min) than for experiments using 365 nm (8 min) (Fig. S20), with a kobs rate 0.27 min−1 noted for 405 nm photopolymerization (Fig. 3C). Though similar molecular weight and dispersity values (12.0 kg mol−1, 1.24) to those from UV irradiation (9.8 kg mol−1, 1.41) were observed (Table S1, entry 8, Fig. 3D), which is consistent with the absorption spectrum of ITX, which has a maximum at 385 nm and tails at around 410 nm.
Other solvents were also examined in order to further elucidate optimum conditions. Apart from DMF, dimethyl acetamide (DMAc), (DMSO), and N-methyl-2-pyrrolidone (NMP) could all enable NCA ROP (Table S1, entries 10–12). Photo-ROP of BLG NCA was slower for NMP (15 min) and DMAc (10 min), while it was faster in DMSO (6 min) (Fig. S21A), forming polypeptides with molecular weights of 10.4, 11.1, and 7.1 kg mol−1 and dispersities of 1.26, 1.49, and 1.42 respectively (Fig. S21B). Changing the [M]0/[I]0 ratio from 50 to 25, and 10 lead to more rapid ROP kinetics in both DMF (Table S1, entries 13 and 14, Fig. S22) and NMP (Table S1, entries 15 and 16, Fig. S23) as anticipated (Table S1, entries 13–16), with NCAs undergoing full conversion in 1 min at [M]0/[I]0 = 10 (Fig. S22A and S23A). However, SEC traces became bimodal at lower [M]0/[I]0 (Fig. S22B and S23B) for both solvents, with a [M]0/[I]0 = 10 in DMF providing a similar molecular weight (5.2 kg mol−1, 1.26) to that of [M]0/[I]0 = 25 (5.7 kg mol−1, 1.39). PBG/ITX/NCA was also then tested for stability during storage in the dark ([M] = 2, [M]0/[I]0 = 50). The system was found to slowly polymerize, with NCA full conversion noted 396 min after addition of PBG/ITX (Fig. S24), significantly slower than under 365 nm irradiation (8 min). This is ascribed to the PBG protonated DBU, as ammonium salts are known to have low NCA ROP capabilities,45 while NCAs also have stability issues in DMF.43 This competing process does not invalidate fast NCA photopolymerization claim by PBGs, though it suggests that freshly prepared solutions should be used to outpace this side reaction, a concept successfully applied for aqueous NCA polymerization.46–49
PBG NCA photo-ROP for degradable polymer networks
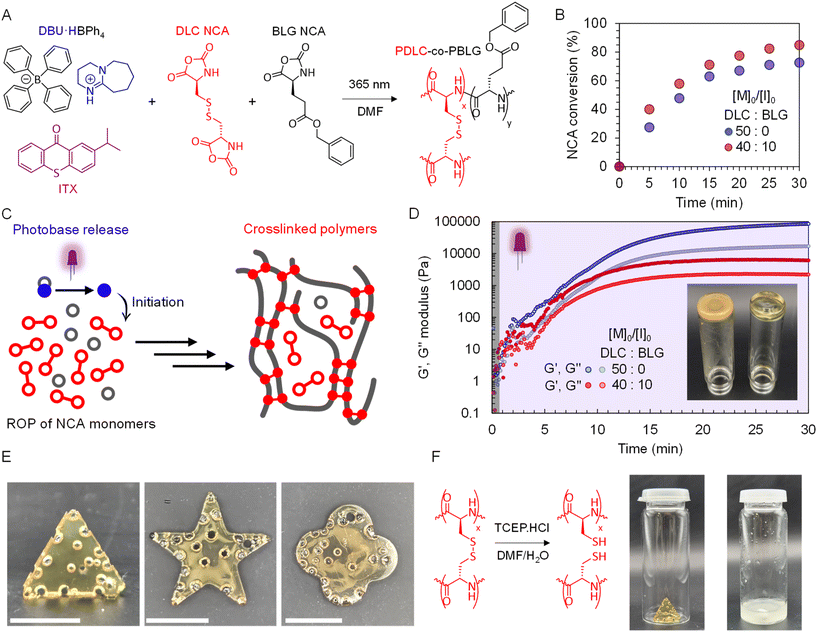
To demonstrate applicability of the photo-NCA process for polypeptide gel formation, PBG ROP with a difunctional NCA (di-NCA) was then explored. While our group and others have reported di-NCA use for crosslinked polypeptides,50–52 herein we demonstrate the first case of harnessing it within photo-ROP for network crosslinking. The di-NCA monomer, L-cystine, termed di-L-cysteine (DLC) herein, was firstly synthesized (Fig. S25). Optimized conditions for homopolymerization of BLG NCA were used for initial tests using DLC NCA alone, specifically solvent (DMF), monomer concentration (2 M), [M]0/[I]0 ratio (50), irradiation wavelength (365 nm) and intensity (8.0 mW cm−2). At this concentration, free standing gels could be formed, though bubbles from CO2 release were noted during formation of the poly(di-L-cysteine) (PDLC) gel network (Fig. S26A), with less bubbles were noted after switching to 1 M concentration (Fig. S26B). In terms of NCA monomer conversion (Fig. 4B), 27% consumption of DLC NCA was measured after 5 min, which plateaued at 66% from 25 min onwards, likely due to monomer immobilization into the PDLC gel networks. BLG NCA was then formulated with DLC, in an attempt to increase molecular weight between crosslinks (Fig. 4C), altering the kinetics for ROP. Maintaining the [M]0/[I]0 ratio at 50, a composition of DLC (40) to BLG (10) was trialed, which enhanced NCA consumption for the system, with 40% conversion noted after 5 min, and 74% conversion noted after 30 min (Fig. 4B). Real-time photo-rheology was then conducted in order to identify the mechanical property regimes PDLC networks (Fig. 4C). Initially, the resins were allowed to incubate in darkness for 30 seconds, before irradiating continuously with 365 nm light at an intensity of 5.0 mW cm−2 (Fig. 4D). An observable crossover between storage modulus (G′) and loss modulus (G″) was noted within 4 min for 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 and 40
0 and 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 formulations, signifying the gel point. While both underwent similar profiles, a distinct difference in plateaued G′ was noted 30 min after irradiation. A G′ of 84
10 formulations, signifying the gel point. While both underwent similar profiles, a distinct difference in plateaued G′ was noted 30 min after irradiation. A G′ of 84![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Pa was recorded for the 50
000 Pa was recorded for the 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 resin, while the diluted 40
0 resin, while the diluted 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 resin had a G′ of 6100 Pa, demonstrating a 13.8-fold increase for the full PDLC network. A control resin with 50
10 resin had a G′ of 6100 Pa, demonstrating a 13.8-fold increase for the full PDLC network. A control resin with 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0 composition was also analyzed in the absence of light irradiation (Fig. S27), displaying no evolution of G′ and G″, signifying the photoinitiated nature of the resins.
0 composition was also analyzed in the absence of light irradiation (Fig. S27), displaying no evolution of G′ and G″, signifying the photoinitiated nature of the resins.
To demonstrate the workability of NCA/PBG resins, photocrosslinked 2.5D networks were manufactured from DLC NCA, whereby the resin was filled into molds and irradiated at 365 nm (8.0 mW cm−2) for 2 h. Different high-fidelity shapes were photostructured from PDLC networks (Fig. 4E), albeit with CO2 bubbles present. Interestingly, the presence of disulfide bonds renders these PDLC networks de-crosslinkable, via disulfide reductive hydrolysis into thiols, providing a controllable and reversible route to polypeptide network disassembly. Upon immersion in a tris(2-carboxyethyl)phosphine hydrochloride (TCEP·HCl) solution, deconstruction of the network was indeed observed (Fig. 4F), rendering a 30 wt% recovery of the initial network after purification. The recovered polymer, anticipated to be poly(L-cysteine) (PLC), was analyzed by DOSY NMR (Fig. S28A), and compared with the DLC NCA monomer (Fig. S28B). A lower diffusion coefficient was observed for PLC (6.96 × 10−7) in contrast to DLC NCA (2.45 × 10−6), characteristic of a larger macromolecule, ultimately suggesting successful de-crosslinking of the photostructured PDLC network to a PLC homopolypeptide.
Conclusions
In summary, we have reported a highly efficient strategy for PBG promoted photo-ROP of NCAs. A range of conditions were trialed by varying molarity, monomer/initiator ratio, light wavelength and NCA monomer(s), indicating the high suitability of PBGs for NCA ROP, with rapid kinetics and good molecular weight control of corresponding polypeptides, including the first case of photopolymerization for fabricating polypeptide-based gel networks into 2.5D structures. The demonstrable synergy of NCAs/PBGs reported here may unlock a new avenue for the design of polypeptide materials by photopolymerization, whereby future design optimization may enable programmable properties via intrinsic secondary structures, or specific enzyme responsiveness for biodegradation, all the while expanding their capabilities in lithography and 3D printing.Author contributions
The project idea was conceived by A. H. The original manuscript and figures were drafted by R. D. M. Experiments were designed by R. D. M., T. O. M., C. G., F. V., H. S., O. S., and A. H., and performed by R. D. M., T. O. M., C. G., A. L., and O. V. All authors have provided input and given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the Supplementary Information.Supplementary information (SI): All synthetic procedures, spectra and characterisation data. See DOI: https://doi.org/10.1039/d6py00091f.
Acknowledgements
Funded by the European Union (GA 101129842). Views and opinions expressed are however those of the authors only and do not necessarily reflect those of the European Union or the European Innovation Council and SMEs Executive Agency (EISMEA). Neither the European Union nor the granting authority can be held responsible for them.References
- R. Göstl, A. Senf and S. Hecht, Chem. Soc. Rev., 2014, 43, 1982–1996 RSC.
- N. Corrigan, J. Yeow, P. Judzewitsch, J. Xu and C. Boyer, Angew. Chem., Int. Ed., 2019, 58, 5170–5189 Search PubMed.
- C. Wu, N. Corrigan, C. H. Lim, W. Liu, G. Miyake and C. Boyer, Chem. Rev., 2022, 122, 5476–5518 Search PubMed.
- A. Bagheri and J. Jin, ACS Appl. Polym. Mater., 2019, 1, 593–611 Search PubMed.
- C. Aydogan, G. Yilmaz, A. Shegiwal, D. M. Haddleton and Y. Yagci, Angew. Chem., Int. Ed., 2022, 61, e202117377 CrossRef CAS PubMed.
- B. P. Fors and C. J. Hawker, Angew. Chem., Int. Ed., 2012, 51, 8850–8853 CrossRef CAS PubMed.
- Z. Wu, K. Jung, C. Wu, G. Ng, L. Wang, J. Liu and C. Boyer, J. Am. Chem. Soc., 2022, 144, 995–1005 CrossRef CAS PubMed.
- S. Gillhuber, A. K. Finch, J. O. Holloway, H. Frisch, F. Weigend, C. Barner-Kowollik and P. W. Roesky, Angew. Chem., Int. Ed., 2025, 64, e202502890 CrossRef CAS PubMed.
- N. Zivic, P. K. Kuroishi, F. Dumur, D. Gigmes, A. P. Dove and H. Sardon, Angew. Chem., Int. Ed., 2019, 58, 10410–10422 CrossRef CAS PubMed.
- P. Utroša, J. A. Carroll, E. Žagar, D. Pahovnik and C. Barner-Kowollik, Chemistry, 2024, 30, e202400820 CrossRef PubMed.
- C. J. Martin, G. Rapenne, T. Nakashima and T. Kawai, J. Photochem. Photobiol., C, 2018, 34, 41–51 CrossRef CAS.
- H. W. Pei, K. Ye, Y. Shao, D. Chen, Z. Y. Sun, T. Gong, D. Liu and K. Sun, Polym. Chem., 2024, 15, 248–268 RSC.
- W. Zhang, S. Li, S. Liu, T. T. Wang, Z. H. Luo, C. Bian and Y. N. Zhou, JACS Au, 2024, 4, 4317–4327 CrossRef CAS PubMed.
- D. Kojic, R. Wolff, Y. Mete, T. Koch, J. Stampfl, S. Baudis, K. Ehrmann and R. Liska, Eur. Polym. J., 2024, 208, 112876 CrossRef CAS.
- C. Fu, J. Xu and C. Boyer, Chem. Commun., 2016, 52, 7126–7129 RSC.
- H. Lai, J. Zhang, F. Xing and P. Xiao, Chem. Soc. Rev., 2020, 49, 1867–1886 RSC.
- X. Zhang, W. Xi, C. Wang, M. Podgórski and C. N. Bowman, ACS Macro Lett., 2016, 5, 229–233 CrossRef CAS PubMed.
- S. Schandl, T. Koch, J. Stampfl, K. Ehrmann and R. Liska, React. Funct. Polym., 2023, 182, 105460 CrossRef CAS.
- A. Shiraishi and T. Yamashita, ChemistrySelect, 2020, 5, 2858–2863 Search PubMed.
- K. Y. Chung, A. Uddin and Z. A. Page, Chem. Sci., 2023, 14, 10736–10743 Search PubMed.
- M. T. Kiker, A. Uddin, L. M. Stevens, C. J. O'Dea, K. S. Mason and Z. A. Page, J. Am. Chem. Soc., 2024, 146, 19704–19709 Search PubMed.
- J. A. Vazquez, X. Lopez de Pariza, N. Ballinger, N. Sadaba, A. Y. Sun, A. O. Olanrewaju, H. Sardon and A. Nelson, Polym. Chem., 2024, 16, 589–597 Search PubMed.
- Y. Liu, E. C. Chang, N. W. Kang, Y. C. Lin and W. C. Chen, ACS Appl. Polym. Mater., 2025, 7, 5418–5428 CrossRef CAS.
- X. Sun, J. P. Gao and Z. Y. Wang, J. Am. Chem. Soc., 2008, 130, 8130–8131 CrossRef CAS PubMed.
- E. Placet, J. Pinaud, O. Gimello and P. Lacroix-Desmazes, ACS Macro Lett., 2018, 7, 688–692 Search PubMed.
- W. Zhang, T.-T. Wang, S. Li, C. Zhao, C. Bian, Y.-N. Zhou and Z.-H. Luo, Angew. Chem., Int. Ed., 2025, 64, e202503923 CrossRef CAS PubMed.
- Y. Wu, K. Chen, J. Wang, M. Chen, W. Dai and R. Liu, J. Am. Chem. Soc., 2024, 146, 24189–24208 CrossRef CAS PubMed.
- Y. Xia, Z. Song, Z. Tan, T. Xue, S. Wei, L. Zhu, Y. Yang, H. Fu, Y. Jiang, Y. Lin, Y. Lu, A. L. Ferguson and J. Cheng, Nat. Commun., 2021, 12, 828–832 CrossRef PubMed.
- Z. Y. Tian, Z. Zhang, S. Wang and H. Lu, Nat. Commun., 2021, 12, 1–11 Search PubMed.
- Q. Li, Y. Lan, W. Wang, G. Ji, X. Li and Z. Song, Macromolecules, 2023, 56, 7023–7031 Search PubMed.
- T. J. Deming, Chem. Rev., 2016, 116, 786–808 CrossRef CAS PubMed.
- P. Zhang, Y. Zhang, X. Ding, W. Shen, M. Li, E. Wagner, C. Xiao and X. Chen, Adv. Mater., 2020, 32, 1–11 Search PubMed.
- C. Ge, J. Yang, S. Duan, Y. Liu, F. Meng and L. Yin, Nano Lett., 2020, 20, 1738–1746 CrossRef CAS PubMed.
- C. Ge, H. Ye, F. Wu, J. Zhu, Z. Song, Y. Liu and L. Yin, J. Mater. Chem. B, 2020, 8, 6530–6547 RSC.
- R. D. Murphy, C. Delaney, S. Kolagatla, L. Florea, C. J. Hawker and A. Heise, Adv. Funct. Mater., 2023, 33, 1–9 CrossRef.
- R. D. Murphy, M. Cosgrave, N. Judge, E. Tinajero-Diaz, G. Portale, B. Wu and A. Heise, Small, 2024, 20, 1–6 CrossRef PubMed.
- R. D. Murphy, R. V. Garcia, A. Heise and C. J. Hawker, Prog. Polym. Sci., 2022, 124, 101487 CrossRef CAS.
- T. P. Nguyen, A. D. Easley, N. Kang, S. Khan, S. M. Lim, Y. H. Rezenom, S. Wang, D. K. Tran, J. Fan, R. A. Letteri, X. He, L. Su, C. H. Yu, J. L. Lutkenhaus and K. L. Wooley, Nature, 2021, 593, 61–66 Search PubMed.
- F. Meng, B. Ju, Z. Wang, R. Han, Y. Zhang, S. Zhang, P. Wu and B. Tang, J. Am. Chem. Soc., 2022, 144, 7610–7615 CrossRef CAS PubMed.
- T. Stukenkemper, J. F. G. A. Jansen, C. Lavilla, A. A. Dias, D. F. Brougham and A. Heise, Polym. Chem., 2017, 8, 828–832 Search PubMed.
- T. Stukenkemper, X. Paquez, M. W. G. M. Verhoeven, E. J. M. Hensen, A. A. Dias, D. F. Brougham and A. Heise, Macromol. Rapid Commun., 2018, 39, 1–5 CrossRef PubMed.
- S. L. Goodrich, M. R. Hill, R. A. Olson and B. S. Sumerlin, Polym. Chem., 2021, 12, 4104–4110 RSC.
- Y. Wu, D. Zhang, P. Ma, R. Zhou, L. Hua and R. Liu, Nat. Commun., 2018, 9, 1–10 Search PubMed.
- Y. Qi, W. Cheng, F. Xu, S. Chen and S. Zhang, Synth. Commun., 2018, 48, 876–886 CrossRef CAS.
- C. D. Vacogne and H. Schlaad, Chem. Commun., 2015, 51, 15645–15648 RSC.
- J. Jacobs, D. Pavlović, H. Prydderch, M. A. Moradi, E. Ibarboure, J. P. A. Heuts, S. Lecommandoux and A. Heise, J. Am. Chem. Soc., 2019, 141, 12522–12526 CrossRef CAS PubMed.
- C. Grazon, P. Salas-Ambrosio, E. Ibarboure, A. Buol, E. Garanger, M. W. Grinstaff, S. Lecommandoux and C. Bonduelle, Angew. Chem., Int. Ed., 2020, 59, 622–626 CrossRef CAS PubMed.
- A. H. Morrell, N. J. Warren and P. D. Thornton, Macromol. Rapid Commun., 2024, 45, 2400103 CrossRef CAS PubMed.
- E. Tinajero-Díaz, N. Judge, B. Li, T. Leigh, R. D. Murphy, P. D. Topham, M. J. Derry and A. Heise, ACS Macro Lett., 2024, 13, 1031–1036 CrossRef PubMed.
- E. D. Raftery, E. G. Gharkhanian, N. G. Ricapito, J. McNamara and T. J. Deming, Chem. – Asian J., 2018, 13, 3547–3553 CrossRef CAS PubMed.
- R. D. Murphy, E. Bobbi, F. C. S. de Oliveira, S. A. Cryan and A. Heise, J. Polym. Sci., Part A: Polym. Chem., 2019, 57, 1209–1215 CrossRef CAS.
- P. Utroša, E. Žagar and D. Pahovnik, Eur. Polym. J., 2024, 204, 112707 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |