Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Fluorescently labelled polypropylene as a model microplastic for cellular imaging
Alexander Evans†
 a,
Ríona M. Devereux†
a,
Ríona M. Devereux† a,
Zoë R. Turner
a,
Zoë R. Turner a,
Angela J. Russell
a,
Angela J. Russell *ab and
Dermot O'Hare
*ab and
Dermot O'Hare *a
*a
aChemistry Research Laboratory, Department of Chemistry, University of Oxford, 12 Mansfield Road, OX1 3TA Oxford, UK. E-mail: dermot.ohare@chem.ox.ac.uk; angela.russell@chem.ox.ac.uk
bDepartment of Pharmacology, University of Oxford, Mansfield Road, OX1 3QT, Oxford, UK
First published on 1st May 2026
Abstract
Polypropylene (PP) is the second most produced plastic globally and is environmentally pervasive in the form of microplastics (MPs). The cellular localisation and fate of MPs within biological systems remain poorly understood. We present a method for the covalent functionalisation of PP with the fluorescent label rhodamine B (Rb) and its subsequent MP evaluation in a human embryonic kidney model cell line (HEK293T). Rb was successfully coupled into PP (PPRb) under Steglich esterification conditions. PPRb is stable under physiological pH and so can be utilised in vitro, offering a significantly improved MP model for conventional staining methods that can definitively locate MPs in human cells. Cellular uptake of PPRb was rapid (≤1 h), and PPRb was non-toxic up to 70 µM and was not observed to localise within the nucleus. Increased mean lysosomal fluorescence intensity suggested an endosomal uptake mechanism followed by significant clearance over a 48 h period. This approach provides a robust platform for definitive microplastic tracking, enabling reliable, standardised studies of polymer–cell interactions across biological systems.
Introduction
Commercially, polyolefins (PO), namely polypropylene (PP)1 and polyethylene (PE),2 are synthesised at scale to meet the needs of a broad array of applications, with PP accounting for 17% of global plastic production.3 Desirable and tuneable properties, such as mechanical strength or creep resistance, as a result of branching, molecular mass (Mn), and dispersities (ĐM), coupled with low production costs are amongst the driving forces behind their widespread utility.4 Although PP is recyclable, a significant proportion enters landfills or the environment.5–7 A large amount of PP waste is converted to microplastics (MPs),8 or indeed nanoplastics,9 via slow degradation due to (a)biotic factors.7 Detection of PO MPs is widely reported, and they are highly prevalent.10,11 Plastic debris is an environmentally persistent and complex contaminant (e.g. varying in size, morphology, and chemical constituency),12 whilst MP detection remains challenging.13,14 Consequently, recent legislative efforts have been introduced in Europe to curtail the prevalence of single use plastics, given the emergence of MPs as a pressing environmental concern due to their ubiquity, potential ecological ramifications, and associated risk.10,14–17The impact of MPs on macroscopic organisms has been extensively studied, with recent reports evaluating their effects on mice,18,19 fish,20–22 and humans.23 There are reports detailing MP entry into aquatic and terrestrial food chains via biomagnification and bioaccumulation,12,24–26 and their detection in fish, birds, and mammals feeding on aquatic organisms or living in aquatic environments has been the subject of further investigation.22,27 Perturbation of ecological processes may align with other anthropogenic pollutants despite the lack of lethality; reduced fecundity and growth at the first trophic level may cause persistent reduction of biomass.28,29 Additionally, investigations into which animals can ingest MPs and the mechanism by which this occurs have been reported.30,31 However, the marked chemical inertness severely hampers evaluation on living species; the incorporation of a fluorescent probe circumvents MP detection issues in organic tissues.32–37
However, the effects on single-celled organisms, which constitute fundamental components of aquatic ecosystems, remain relatively underexplored. These minute organisms, encompassing diverse taxa such as phytoplankton and protozoa, play pivotal roles in nutrient cycling and food web dynamics.29,38–40 The ingestion of microplastics by single-celled organisms can lead to various detrimental outcomes, including altered feeding behaviours, impaired metabolic functions, and compromised reproductive success.29,38–40 Furthermore, the sorption of toxic chemicals onto microplastics introduces an additional layer of risk, as the bioavailability of these pollutants increases upon ingestion by these microorganisms,13,16,28,33,41 although disputed.10 Despite the limited research in this area, the potential for cascading effects throughout aquatic ecosystems underscores the urgency to comprehensively elucidate the consequences of microplastic exposure on single-celled organisms. Ascertaining an understanding of the posed risk is crucial for the educated development of policy. Current methods for evaluating the role of MPs with single-celled organisms predominantly utilise staining,42 due to the relative synthetic simplicity,43 although some dyes are covalently bound.37 In vitro assays on mice44 and human45 cell lines utilise polystyrene (PS) for this purpose. However, this methodology has drawbacks, primarily leaching of free dye, potentially leading to the presence of artefacts, confounding the interpretation of the results.46–48 Subsequently, reports with claims of polymer sample purity, or indeed covalent coupling of dye units to the polymer chain, are not supported by adequate rigorous characterisation data leading to the ambiguous interpretation of results.
In the field of mechanical recycling, separation techniques have been extended to include fluorescent labelling, enabling separation by specific polymeric materials;49–51 recent reports utilise fluorescently tagged PE in this way.52 Notably, the recycling process typically involves high temperatures and consequently careful selection of incorporated spectroscopic markers is required to avoid degradation.52 To the best of our knowledge, the recent literature presents a few examples of synthesised isotactic-PP copolymers containing fluorescent units, and, so far, only the work by Pragliola et al. has evaluated them as a fluorescent marker of PP.32,53 Fluorescent probe coupled PP may be used to mimic the in vivo behaviour of PP-derived MP pollutants on a cellular level.
Several Norwegian institutions and initiatives have pioneered extensive studies in evaluating MP research;54 despite a plethora of published articles, it is posited that a lack of standardisation and validated methods, particularly for the smallest plastic particles, hampers conclusive and harmonised research.55 Given the interdisciplinary nature of the problem, it is well within the remit of polymer scientists to design, synthesise, and provide specific model polymers for MP analysis in collaboration with relevant academic disciplines. We have recently reported a general two-step methodology to functionalise PP, creating a wide range of covalent linkages, something traditionally difficult to achieve in a controlled manner.56 Consequently, we present a method for the covalent fluorescent labelling of rhodamine B (Rb) functionalised PP, PPRb, and subsequent MP evaluation in a human kidney cell line. We believe that this work provides a new methodology for covalently bound fluorescent tags on a PO system which allows confident analysis of any polymer–cell interaction. This has the potential to offer a harmonised approach for the study of related polymeric materials and enable conclusive evaluation of the interaction of such ubiquitous materials within cells and organisms.
Results and discussion
Synthesis and characterisation of PPRb
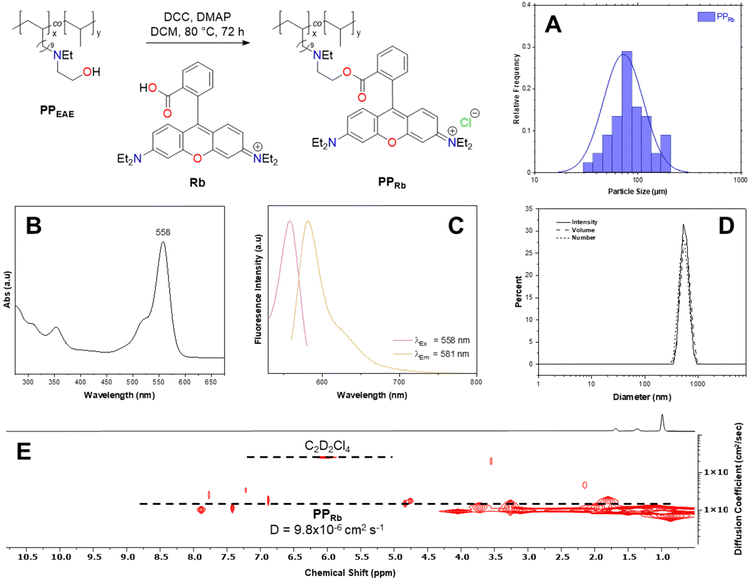
PPRb was synthesised via the quantitative transformation of a previously reported alcohol-functionalised PP copolymer (PPEAE) under Steglich esterification conditions (DMAP 0.3 eq., DCC 2 eq., DCM, 80 °C, 72 h) with Rb (2 eq.) (Fig. 1).56,57Following extensive purification by water Soxhlet extraction, PPRb with 1.87 mol% covalently-bound dye incorporation was isolated and fully characterised. Key 1H NMR spectroscopic handles evidence the presence of the dye; six aromatic resonances between 6.37 and 7.91 ppm integrate appropriately (Fig. S1). Convection compensated diffusion experiments support the conclusion that Rb is bound to a higher molecular weight species with a diffusion coefficient of 9.8 × 10−6 cm2 s−1 (Fig. 1E), which is comparable to the parent copolymer (ca. 9.9 × 10−6 cm2 s−1) and considerably different from the free dye (ca. 6.2 × 10−6 cm2 s−1) as tracked by aromatic protons and a distinct quartet assigned to –NCH2CH3 (3.44 ppm) (Fig. S1 and S2). PPRb solution in chloroform (0.5 mg mL−1) exhibits an excitation wavelength (λEx) of 558 nm by ultraviolet–visible (UV-Vis) spectroscopy (Fig. 1B). Significantly, the polymer was also shown to be fluorescent with an emission wavelength (λEm) of 581 nm (Fig. 1C). Crucially, λEx and λEm are distinctly altered when compared to derivative Rb (λEx = 569 nm, λEm = 592 nm) in a comparable molar concentration of the fluorophore in chloroform (Fig. S8).
The polymer was further characterised by Fourier transform infrared spectroscopy (ν = 1736 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) and 808 cm−1 (C
O) and 808 cm−1 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) CAr)) and differential scanning calorimetry (Tm = 113 °C), and a single decomposition step was evidenced (Tmax = 431 °C) by thermogravimetric analysis (Fig. S6). Multiple complementary techniques were employed to assess the polymer particle size, as variations in measurement conditions can substantially influence the observed dimensions. Mn analysis by size exclusion chromatography (SEC) was unsuccessful, with PPRb not eluting within the calibration window; however, the theoretical molar mass (Mn, th) is predicted to be ca. 14
CAr)) and differential scanning calorimetry (Tm = 113 °C), and a single decomposition step was evidenced (Tmax = 431 °C) by thermogravimetric analysis (Fig. S6). Multiple complementary techniques were employed to assess the polymer particle size, as variations in measurement conditions can substantially influence the observed dimensions. Mn analysis by size exclusion chromatography (SEC) was unsuccessful, with PPRb not eluting within the calibration window; however, the theoretical molar mass (Mn, th) is predicted to be ca. 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 g mol−1 based on the parent copolymer experimental molar mass (Mn, SEC = 13
700 g mol−1 based on the parent copolymer experimental molar mass (Mn, SEC = 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 g mol−1) (Fig. S6). Since the Mark–Houwink correlation is not known for functionalized PP, and the presence of a polar group is likely to affect the radius of gyration in solution, it is reasonable to assume that the backbone chain length remains unchanged during post polymerization modification.58 Dynamic light scattering (DLS) suggests that PPRb aggregates in the solution state, common for amphiphilic copolymers and potentially induced by the ionic nature of the fluorophore.59 PPRb in toluene solution (1 mg mL−1) exhibited good agreement between number, volume, and intensity values for the hydrodynamic diameter (Dh), ca. 570 nm, which increased from that of the parent copolymer, ca. 140 nm, supporting a micellar arrangement in solution with increasing hydrophilic character (Fig. 1D, Fig. S7, and Table S1). It is worth noting that this may subsequently affect the permeability of PPRb towards the cell membrane; however, existing in vitro experiments employ charged, colloidally stable PS particles44 and we believe that our approach offers an alternative with significantly reduced impact (vide infra). Additionally, some PPRb was cryofractured (cPPRb) in an attempt to simulate accelerated weathering (Fig. S9 and S10), such that the irregularities in size and shape of MPs found within the environment could be rudimentarily assessed.32
800 g mol−1) (Fig. S6). Since the Mark–Houwink correlation is not known for functionalized PP, and the presence of a polar group is likely to affect the radius of gyration in solution, it is reasonable to assume that the backbone chain length remains unchanged during post polymerization modification.58 Dynamic light scattering (DLS) suggests that PPRb aggregates in the solution state, common for amphiphilic copolymers and potentially induced by the ionic nature of the fluorophore.59 PPRb in toluene solution (1 mg mL−1) exhibited good agreement between number, volume, and intensity values for the hydrodynamic diameter (Dh), ca. 570 nm, which increased from that of the parent copolymer, ca. 140 nm, supporting a micellar arrangement in solution with increasing hydrophilic character (Fig. 1D, Fig. S7, and Table S1). It is worth noting that this may subsequently affect the permeability of PPRb towards the cell membrane; however, existing in vitro experiments employ charged, colloidally stable PS particles44 and we believe that our approach offers an alternative with significantly reduced impact (vide infra). Additionally, some PPRb was cryofractured (cPPRb) in an attempt to simulate accelerated weathering (Fig. S9 and S10), such that the irregularities in size and shape of MPs found within the environment could be rudimentarily assessed.32
Cellular studies
Incorporating fluorophores directly into MP models marks a critical advancement in visualising and understanding MP interactions within biological systems. Fluorophore incorporation on PP enabled an in vitro investigation in a human embryonic kidney (HEK293T) model cell line, where PPRb is utilised as a model for MPs, overcoming imaging issues suffered by traditional polyolefins. Through the covalent linkage of the dye moiety, it can be concluded that any observations arise due to the presence of the dye-bound polymer and not through leaching; other methods previously reported suffer from this drawback.42,48,60 A control experiment was performed to measure polymer–dye stability using a citrate–phosphate buffer solution, mimicking the acidic lysosomal compartments (pH 4, 38 °C), as detailed in the SI.61 Detected fluorescence correlated to a maximum hydrolysis of 1.6% over 3 days, and was deemed not statistically significant, suggesting minimal degradation in a physiologically relevant pH range (Fig. S17).HEK293T cells were chosen as the model cell line to investigate uptake and localisation via confocal microscopy. The primary entry points for MPs into the human body are ingestion, inhalation, and dermal contact.62,63 Once internalised, MPs can enter the circulatory system, potentially accumulating in organs such as the gut, liver, and kidneys.64 Given the kidney's role in filtering and detoxifying various substances, including environmental contaminants like MPs, the HEK293T cell line serves as a relevant model for examining cellular defence mechanisms in response to MPs.65
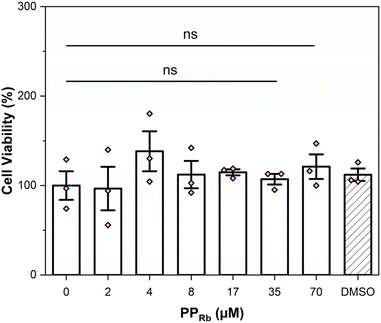
The acute cytotoxicity of PPRb was assessed in vitro using the AlamarBlue™ Cell Viability Assay in HEK293T cells. Cells were exposed to a range of [PPRb] for 24 h, with cell viability measured across the concentration gradient. No significant cytotoxic effects were observed up to the highest concentration tested (PPRb = 70 µM), as compared to the control group (PPRb = 0 µM) (Fig. 2). Additionally, bright-field microscopy of cells treated with 35 µM PPRb for 24 h revealed no signs of cell death or structural abnormalities relative to untreated control cells (Fig. 2 and S13). It is worth noting that PPRb was rigorously purified prior to cellular treatment. In environmental contexts, however, MPs often act as sorbents, potentially concentrating harmful and toxic materials on their surfaces, which may influence their biological effects.33,66–68
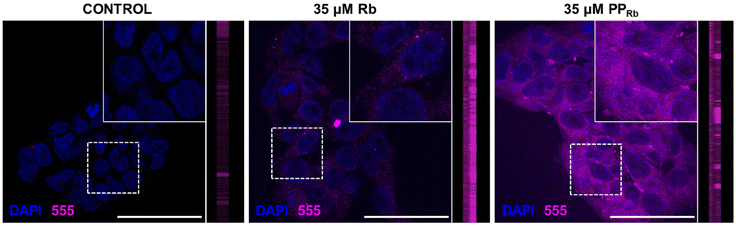
To assess the uptake and localisation of PPRb in vitro, HEK293T cells were incubated with either 35 µM of unbound fluorophore (Rb) or fluorophore-bound PPRb for 1 h and compared with untreated control cells (Fig. 3). A detectable signal was observed in the 555 nm channel for both Rb and PPRb, with minimal background fluorescence in the negative control group. Notably, cells treated with PPRb exhibited a higher signal intensity compared to those treated with free Rb, indicating a consistent cellular uptake of PPRb without removal upon washing. The 1 h treatment period demonstrates rapid cellular uptake and accumulation of PPRb within HEK293T cells. The dye-bound polymer, PPRb, exhibits localisation within cells due to its polymeric structure, in contrast to the free Rb dye, which results in a more diffuse cellular distribution. Accumulation of PPRb within cells was evident, as confirmed by z-stack imaging (Fig. 3), highlighting the polymer's capacity for localised uptake and retention compared with the less structured, widespread localisation pattern observed with free Rb. The different observed cellular behaviours of both Rb and PPRb are consistent with the absence of ester hydrolysis and dye liberation from PPRb (Fig. S17).69
After 24 h, cellular localisation of PPRb is reduced compared to the 1 h time point, suggesting a potential decrease in intracellular retention (Fig. 4B). This reduction in cellular localisation may indicate that PPRb is being actively removed from the cell through a cellular clearance pathway, possibly via an endosomal recycling pathway, although a metabolic degradation pathway cannot be ruled out.70
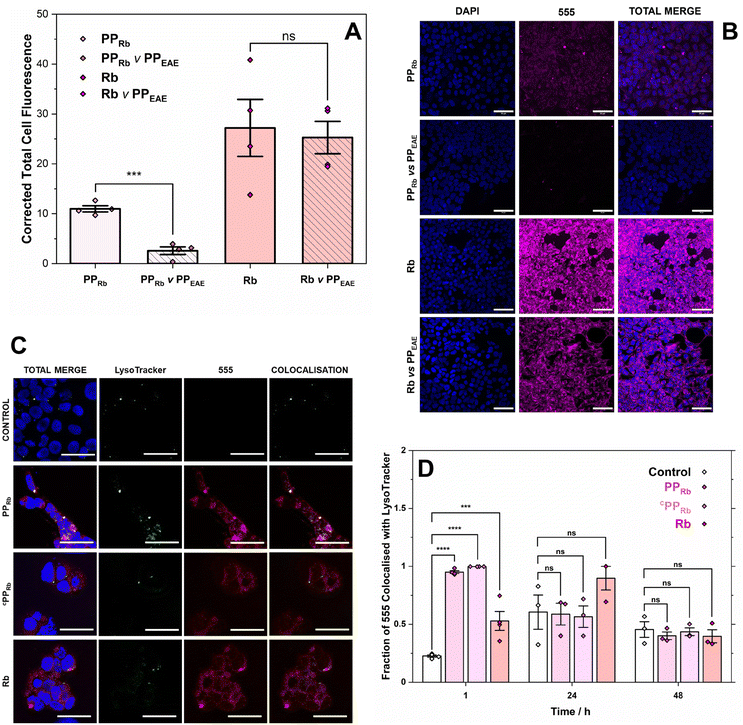
A competition assay was conducted to assess the influence of the parent alcohol-functionalised copolymer, PPEAE, on the cellular localisation of PPRb after 24 h (Fig. 4A and B). The results indicated that the parent polymer directs cellular localisation of PPRb, as fluorescence was significantly decreased upon co-incubation with 35 µM PPEAE. In contrast, the localisation of Rb remained unaffected by the presence of PPEAE, supporting the conclusion that PPEAE and PPRb localise to the same cellular compartments, while Rb does not. This further supports the limited impact of the charged fluorophore driving cellular uptake, trafficking and compartmentalisation, strengthening the role of PPRb as a model MP. Notably, this competitive reduction in fluorescence was not observed with the Rb fluorophore alone, underscoring that PPEAE, and not Rb, modulates the intracellular dynamics of PPRb localisation.
Furthermore, a sample of PPRb was subjected to cryogenic fracturing to produce cPPRb, a model intended to simulate accelerated environmental degradation.32 However, data showed no significant difference in cellular localisation between cPPRb and the original PPRb sample (Fig. S11). Importantly, across all timepoints, there was no detectable co-localisation of PPRb and the nuclear stain DAPI (Fig. S11). Excluding certain molecules or structures from the nucleus can prevent potential disruptions to nuclear architecture or DNA stability, safeguarding essential cellular processes.71
It was hypothesised that, due to the polymer size and relatively low Mn, the polymer, or any nanoparticles formed through self-assembly, would enter the cells through an energy-dependent trafficking mechanism via an endosomal pathway.72 It is, therefore, reasonable to infer possible trafficking of PPRb to the lysosomes upon cellular entry. To investigate this, HEK293T cells were stained with LysoTracker, an acidic vesicle-labelling dye, and incubated with PPRb for 1, 24, and 48 h (Fig. 4C, S14–S16). The mean LysoTracker fluorescence intensity for PPRb, cPPRb and free Rb increased relative to the control at 1 and 24 h, suggesting a potential endosomal uptake mechanism. By 48 h, however, the mean fluorescence intensity of LysoTracker in the PPRb and cPPRb treated cells returned to control levels, while the fluorescence for free Rb remained elevated. Following this, the colocalization of the 555 channels with LysoTracker was investigated for PPRb, cPPRb, and Rb (Fig. 4C and D and S14 and S15). Incubation of PPRb and cPPRb showed a significant fraction of channel overlap at 1 h, confirming colocalization of the polymer with lysosomes in vitro at this early timepoint. However, this overlap was insignificant at the 24 and 48 h timepoints. These colocalization results further support a lysosomal uptake mechanism. Free Rb showed no significant overlap at all tested timepoints, suggesting that the increase in lysosomes may not be directly associated with free Rb and could be a secondary effect of its incubation. These observations and earlier dosing data showing reduced PPRb uptake by 24 h (Fig. 4B) suggest that cells may actively work to clear the polymer over time.
Concluding remarks
Polypropylene with the covalently-bound fluorescent rhodamine B tag, PPRb, was synthesised and characterised for in vitro studies against a HEK293T cell line and evaluated by confocal microscopy. The two-step copolymerization–post-modification strategy employed is facile and, importantly, allows the preparation of rigorously purified model polymer samples that do not undergo dye leaching.Incubation with PPRb resulted in rapid cellular uptake (1 h) whilst remaining non-toxic up to 70 µM. Crucially, no localisation within the nucleus was observed under the limits of detection. Initial investigations were undertaken into the uptake mechanism and efficacy as a PP MP model. Increased mean lysosomal fluorescence intensity following PPRb treatment suggests an endosomal uptake mechanism with significant clearance implied over a 48 h period. Cellular localisation of PPRb is directed by the parent alcohol-functionalised copolymer, as evidenced by competition studies, indicating that PPRb is a valuable model for PP cell interaction. Optimistically, accounting for the constraints of our investigation, PPRb acts as an inert orthogonal species within the cell. However, it is important to investigate this further, especially by probing the polymer species as a vector for harmful environmental substances.
We believe that this method for the synthesis and application of covalently bound dye moieties can elucidate the degree to which concern can be levied against the developing environmental plastic pollution. More broadly, this blueprint can enable a precise and confident interlaboratory analysis of polymer–cell interactions through the targeted synthesis of dye-modified polymers of interest. Additionally, this method offers many investigative opportunities, be it an in vitro cell line or in vivo studies of more complex organisms, including their ecological surroundings.
Author contributions
A. E. performed the investigation and formal analysis of polymer-related experiments. R. M. D. undertook the investigation and formal analysis of biological-related experiments. A. E. conceived and managed the project and drafted the manuscript. All authors revised the manuscript. D. O. H. and A. J. R. supervised the research and acquired the funding.Conflicts of interest
There are no conflicts to declare.Data availability
The supporting data have been provided as part of the supplementary information (SI). Supplementary information: experimental and data analysis methods, supporting polymer characterization, and images and data pertaining to the cell studies. See DOI: https://doi.org/10.1039/d6py00089d.Acknowledgements
A. E. thanks the EPSRC for funding under the Impact Acceleration Awards Doctoral Impact Scheme. The research reported in this work was (in part) supported financially by SCG Chemicals PLC (Thailand). R. M. D. thanks Wellcome Trust (218514/Z/19/Z) for studentship funding, established with support from Merck Sharp and Dohme Corp and Janssen Pharmaceutica NV. We also thank Dr Coral Mycroft (Oxford NMR Submission Service) for running various NMR experiments and Ms Liv Thobru, Ms Sara Rund Herum, and Ms Rita Jenssen (Norner AS) for running GPC analysis.References
- G. Natta, P. Pino, P. Corradini, F. Danusso, E. Mantica, G. Mazzanti and G. Moraglio, J. Am. Chem. Soc., 1955, 77, 1708–1710 CrossRef CAS.
- K. Ziegler, E. Holzkamp, H. Breil and H. Martin, Angew. Chem., 1955, 67, 426 CrossRef CAS.
- A. H. Westlie, E. Y.-X. Chen, C. M. Holland, S. S. Stahl, M. Doyle, S. R. Trenor and K. M. Knauer, Macromol. Rapid Commun., 2022, 43, 2200492 CrossRef CAS PubMed.
- L. Resconi, L. Cavallo, A. Fait and F. Piemontesi, Chem. Rev., 2000, 100, 1253–1346 CrossRef CAS PubMed.
- R. Francis, Recycling of Polymers: Methods, Characterization and Applications, John Wiley & Sons, 2016 Search PubMed.
- V. V. Narmadha, J. Jose, S. Patil, M. O. Farooqui, B. Srimuruganandam, S. Saravanadevi and K. Krishnamurthi, Int. J. Environ. Res., 2020, 14, 629–640 CrossRef CAS.
- M. Potrykus, V. Redko, K. Głowacka, A. Piotrowicz-Cieślak, P. Szarlej, H. Janik and L. Wolska, Sci. Total Environ., 2021, 758, 143649 CrossRef CAS PubMed.
- J. P. G. L. Frias and R. Nash, Mar. Pollut. Bull., 2019, 138, 145–147 CrossRef CAS PubMed.
- J. Gigault, A. T. Halle, M. Baudrimont, P. Y. Pascal, F. Gauffre, T. L. Phi, H. El Hadri, B. Grassl and S. Reynaud, Environ. Pollut., 2018, 235, 1030–1034 CrossRef CAS PubMed.
- E. E. Burns and A. B. A. Boxall, Environ. Toxicol. Chem., 2018, 37, 2776–2796 CrossRef CAS PubMed.
- J. Sun, X. Dai, Q. Wang, M. C. M. van Loosdrecht and B.-J. Ni, Water Res., 2019, 152, 21–37 CrossRef CAS PubMed.
- A. A. Horton, A. Walton, D. J. Spurgeon, E. Lahive and C. Svendsen, Sci. Total Environ., 2017, 586, 127–141 CrossRef CAS PubMed.
- S. O. Fakayode, T. F. Mehari, V. E. Fernand Narcisse, C. Grant, M. E. Taylor, G. A. Baker, N. Siraj, M. Bashiru, I. Denmark, A. Oyebade, D. K. Bwambok, C. Kuedukey, T. Alonge and D. Anum, Appl. Spectrosc. Rev., 2024, 59, 1183–1277 CrossRef CAS.
- G. Chen, Q. Feng and J. Wang, Sci. Total Environ., 2020, 703, 135504 CrossRef CAS PubMed.
- A. A. Koelmans, P. E. Redondo-Hasselerharm, N. H. M. Nor, V. N. de Ruijter, S. M. Mintenig and M. Kooi, Nat. Rev. Mater., 2022, 7, 138–152 CrossRef.
- P. Liu, X. Zhan, X. Wu, J. Li, H. Wang and S. Gao, Chemosphere, 2020, 242, 125193 CrossRef CAS PubMed.
- K. Liu, X. Wang, T. Fang, P. Xu, L. Zhu and D. Li, Sci. Total Environ., 2019, 675, 462–471 CrossRef CAS PubMed.
- Y. Jin, L. Lu, W. Tu, T. Luo and Z. Fu, Sci. Total Environ., 2019, 649, 308–317 CrossRef CAS PubMed.
- L. Lu, Z. Wan, T. Luo, Z. Fu and Y. Jin, Sci. Total Environ., 2018, 631–632, 449–458 CrossRef CAS PubMed.
- L. Lei, S. Wu, S. Lu, M. Liu, Y. Song, Z. Fu, H. Shi, K. M. Raley-Susman and D. He, Sci. Total Environ., 2018, 619–620, 1–8 CrossRef CAS PubMed.
- A. Batel, F. Linti, M. Scherer, L. Erdinger and T. Braunbeck, Environ. Toxicol. Chem., 2016, 35, 1656–1666 CrossRef CAS PubMed.
- L. C. de Sá, M. Oliveira, F. Ribeiro, T. L. Rocha and M. N. Futter, Sci. Total Environ., 2018, 645, 1029–1039 CrossRef PubMed.
- N. Chartres, C. B. Cooper, G. Bland, K. E. Pelch, S. A. Gandhi, A. BakenRa and T. J. Woodruff, Environ. Sci. Technol., 2024, 58(52), 22843–22864 CrossRef CAS PubMed.
- Y. Huang, Q. Liu, W. Jia, C. Yan and J. Wang, Environ. Pollut., 2020, 260, 114096 CrossRef CAS PubMed.
- F. S. Hamid, M. S. Bhatti, N. Anuar, N. Anuar, P. Mohan and A. Periathamby, Waste Manage. Res., 2018, 36, 873–897 CrossRef PubMed.
- A. A. de Souza Machado, W. Kloas, C. Zarfl, S. Hempel and M. C. Rillig, Glob. Chang. Biol., 2018, 24, 1405–1416 CrossRef PubMed.
- N. Baranzini, L. Pulze, C. Bon, L. Izzo, S. Pragliola, V. Venditto and A. Grimaldi, Fish Shellfish Immunol., 2022, 127, 492–507 CrossRef CAS PubMed.
- T. S. Galloway, M. Cole and C. Lewis, Nat. Ecol. Evol., 2017, 1, 0116 CrossRef PubMed.
- P. Raju, P. Santhanam, S. S. Pandian, M. Divya, A. Arunkrishnan, K. N. Devi, S. Ananth, J. Roopavathy and P. Perumal, Arch. Microbiol., 2021, 204, 84 CrossRef CAS PubMed.
- S. Anbumani and P. Kakkar, Environ. Sci. Pollut. Res., 2018, 25, 14373–14396 CrossRef CAS PubMed.
- H. S. Auta, C. U. Emenike and S. H. Fauziah, Environ. Int., 2017, 102, 165–176 CrossRef CAS PubMed.
- S. Pragliola, F. Grisi, V. Vitale, O. Sacco, V. Venditto, L. Izzo, A. Grimaldi and N. Baranzini, Polym. Chem., 2022, 13, 2685–2693 RSC.
- C. Campanale, C. Massarelli, I. Savino, V. Locaputo and V. F. Uricchio, Int. J. Environ. Res. Public Health, 2020, 17, 1212 CrossRef CAS.
- T. Maes, R. Jessop, N. Wellner, K. Haupt and A. G. Mayes, Sci. Rep., 2017, 7, 44501 CrossRef CAS PubMed.
- L. Nalbone, A. Panebianco, F. Giarratana and M. Russell, Mar. Pollut. Bull., 2021, 172, 112888 CrossRef CAS PubMed.
- E. G. Karakolis, B. Nguyen, J. B. You, P. J. Graham, C. M. Rochman and D. Sinton, Environ. Sci. Technol., 2018, 5, 62–67 CAS.
- B. Nguyen and N. Tufenkji, Environ. Sci. Technol., 2022, 56, 6426–6435 CrossRef CAS PubMed.
- S. Rist, A. Baun and N. B. Hartmann, Environ. Pollut., 2017, 228, 398–407 CrossRef CAS PubMed.
- M. Long, I. Paul-Pont, H. Hégaret, B. Moriceau, C. Lambert, A. Huvet and P. Soudant, Environ. Pollut., 2017, 228, 454–463 CrossRef CAS PubMed.
- Y. Mao, H. Ai, Y. Chen, Z. Zhang, P. Zeng, L. Kang, W. Li, W. Gu, Q. He and H. Li, Chemosphere, 2018, 208, 59–68 CrossRef CAS PubMed.
- M. Gao, Y. Liu and Z. Song, Chemosphere, 2019, 237, 124482 CrossRef CAS PubMed.
- E. G. Karakolis, B. Nguyen, J. B. You, C. M. Rochman and D. Sinton, Environ. Sci. Technol., 2019, 6, 334–340 CAS.
- S. Morgana, B. Casentini, V. Tirelli, F. Grasso and S. Amalfitano, Trends Anal. Chem., 2024, 172, 117559 CrossRef CAS.
- B.-K. Jung, S.-W. Han, S.-H. Park, J.-S. Bae, J. Choi and K.-Y. Ryu, Neurotoxicology, 2020, 81, 189–196 CrossRef CAS PubMed.
- Y. Zhang, S. Wang, V. Olga, Y. Xue, S. Lv, X. Diao, Y. Zhang, Q. Han and H. Zhou, Chemosphere, 2022, 291, 132714 CrossRef CAS PubMed.
- M. T. Sturm, E. Myers, D. Schober, A. Korzin and K. Schuhen, Analytica, 2023, 4, 27–44 CrossRef CAS.
- W. J. Shim, S. H. Hong and S. E. Eo, Anal. Methods, 2017, 9, 1384–1391 RSC.
- G. Malafaia, T. M. d. Luz, M. A. I. Ahmed, S. Karthi and A. P. d. C. Araújo, J. Hazard. Mater. Adv., 2022, 6, 100054 CAS.
- D. Jubinville, E. Esmizadeh, S. Saikrishnan, C. Tzoganakis and T. Mekonnen, Sustainable Mater. Technol., 2020, 25, e00188 CrossRef CAS.
- E. Maris, A. Aoussat, E. Naffrechoux and D. Froelich, Miner. Eng., 2012, 29, 77–88 CrossRef CAS.
- S. Brunner, P. Fomin and C. Kargel, Waste Manage., 2015, 38, 49–60 CrossRef CAS PubMed.
- A. Arenas-Vivo, F. R. Beltrán, V. Alcázar, M. U. de la Orden and J. Martinez Urreaga, Mater. Today Commun., 2017, 12, 125–132 CrossRef CAS.
- R. Shang, H. Gao, F. Luo, Y. Li, B. Wang, Z. Ma, L. Pan and Y. Li, Macromolecules, 2019, 52, 9280–9290 CrossRef CAS.
- A. L. Lusher, R. Hurley, H. P. H. Arp, A. M. Booth, I. L. N. Bråte, G. W. Gabrielsen, A. Gomiero, T. Gomes, B. E. Grøsvik, N. Green, M. Haave, I. G. Hallanger, C. Halsband, D. Herzke, E. J. Joner, T. Kögel, K. Rakkestad, S. B. Ranneklev, M. Wagner and M. Olsen, Environ. Int., 2021, 157, 106794 CrossRef CAS PubMed.
- A. Bakir, A. R. McGoran, B. Silburn, J. Russell, H. Nel, A. L. Lusher, R. Amos, R. S. Shadrack, S. J. Arnold, C. Castillo, J. F. Urbina, E. Barrientos, H. Sanchez, K. Pillay, L. Human, T. Swartbooi, M. R. Cordova, S. Y. Sani, T. W. A. W. Wijesinghe, A. A. D. Amarathunga, J. Gunasekara, S. Somasiri, K. Mahatantila, S. Liyanage, M. Müller, Y. Y. Hee, D. F. Onda, K. M. Jansar, Z. Shiraz, H. Amir and A. G. Mayes, Sci. Rep., 2024, 14, 12714 CrossRef CAS PubMed.
- A. Evans, O. Casale, L. J. Morris, Z. R. Turner and D. O'Hare, Macromolecules, 2024, 57, 10778–10791 CrossRef CAS.
- B. Neises and W. Steglich, Angew. Chem., Int. Ed. Engl., 1978, 17, 522–524 CrossRef.
- L. K. Kostanski, D. M. Keller and A. E. Hamielec, J. Biochem. Biophys. Methods, 2004, 58, 159–186 CrossRef CAS PubMed.
- C. J. Kay, P. D. Goring, C. A. Burnett, B. Hornby, K. Lewtas, S. Morris, C. Morton, T. McNally, G. W. Theaker, C. Waterson, P. M. Wright and P. Scott, J. Am. Chem. Soc., 2018, 140, 13921–13934 CrossRef CAS PubMed.
- Z. Gao, K. Wontor and J. V. Cizdziel, Molecules, 2022, 27, 7415 CrossRef CAS PubMed.
- S. Jordans, S. Jenko-Kokalj, N. M. Kühl, S. Tedelind, W. Sendt, D. Brömme, D. Turk and K. Brix, BMC Biochem., 2009, 10, 23 CrossRef PubMed.
- J. C. Prata, Environ. Pollut., 2018, 234, 115–126 CrossRef CAS PubMed.
- M. Revel, A. Châtel and C. Mouneyrac, Curr. Opin. Environ. Sci. Health, 2018, 1, 17–23 CrossRef.
- R. Gautam, J. Jo, M. Acharya, A. Maharjan, D. Lee, P. B. KC, C. Kim, K. Kim, H. Kim and Y. Heo, Sci. Total Environ., 2022, 838, 156089 CrossRef CAS PubMed.
- N. Jourde-Chiche, F. Fakhouri, L. Dou, J. Bellien, S. Burtey, M. Frimat, P.-A. Jarrot, G. Kaplanski, M. Le Quintrec, V. Pernin, C. Rigothier, M. Sallée, V. Fremeaux-Bacchi, D. Guerrot and L. T. Roumenina, Nat. Rev. Nephrol., 2019, 15, 87–108 CrossRef PubMed.
- K. Lu, R. Qiao, H. An and Y. Zhang, Chemosphere, 2018, 202, 514–520 CrossRef CAS PubMed.
- L. G. A. Barboza, L. R. Vieira, V. Branco, N. Figueiredo, F. Carvalho, C. Carvalho and L. Guilhermino, Aquat. Toxicol., 2018, 195, 49–57 CrossRef CAS PubMed.
- L. Bradney, H. Wijesekara, K. N. Palansooriya, N. Obadamudalige, N. S. Bolan, Y. S. Ok, J. Rinklebe, K.-H. Kim and M. B. Kirkham, Environ. Int., 2019, 131, 104937 CrossRef CAS PubMed.
- J. Tanaka, A. Evans, P. Gurnani, A. Kerr and P. Wilson, Polym. Chem., 2020, 11, 2519–2531 RSC.
- L. Shang, K. Nienhaus and G. U. Nienhaus, J. Nanobiotechnol., 2014, 12, 5 CrossRef PubMed.
- B. M. Webster and C. P. Lusk, Trends Cell Biol., 2016, 26, 29–39 CrossRef CAS PubMed.
- G. J. Doherty and H. T. McMahon, Annu. Rev. Biochem., 2009, 78, 857–902 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |