Open Access Article
Open Access ArticleA computational framework for tuning intra- and intermolecular ductility in polyurethanes
Chenxi
Sheng†
ab,
Meng
Huang†
a,
Andrew P.
Dove
 *b and
Linjiang
Chen
*b and
Linjiang
Chen
 *ab
*ab
aState Key Laboratory of Precision and Intelligent Chemistry, University of Science and Technology of China, Hefei, China. E-mail: linjiangchen@ustc.edu.cn
bSchool of Chemistry, University of Birmingham, Birmingham, UK. E-mail: a.dove@bham.ac.uk
First published on 9th March 2026
Abstract
The development of sustainably sourced polymers with robust mechanical properties is an important aspect of the challenge to mitigate the impact of plastic waste on the environment. However, with a wide range of potential bio-derivable building blocks and chemicals with which to construct polymers, predicting their thermomechanical properties is challenging. To address this challenge, here we established an integrated computational framework to relate chemical design to mechanics in isohexide-based polyurethanes (PUs). We combined and varied three structural motifs: (i) dynamic-bond moieties in the backbone, (ii) hydrogen-bonding moieties that mediate interchain cohesion, and (iii) stereochemical ring configurations. Density functional theory (DFT) with the “External Force is Explicitly Included” (EFEI) formalism quantified how different dynamic bonds control single-chain scission forces, while semiempirical EFEI calculations and classical molecular dynamics (MD) revealed how the number and arrangement of hydrogen-bond sites govern double-chain shear forces. Reactive MD with ReaxFF was used to probe uniaxial tensile deformation of amorphous PU bulk systems. Sulfur-containing dynamic-bond moieties markedly reduced single-chain scission forces, consistent with their use in self-healing and reprocessable PUs, whereas nitrogen-containing motifs combined with highly multidentate hydrogen-bonding groups maximized both intrachain strength and interchain cohesion. A representative design (NO5M) achieved a substantially higher peak stress than a disulfide-rich analogue (SS4I) under tensile loading. This multiscale framework yields chemically interpretable design rules for high-performance, recyclable PUs and illustrates the synergistic use of EFEI and MD simulations in polymer mechanochemistry.
Introduction
Polymeric materials underpin technologies ranging from structural components and coatings to elastomers and biomedical devices,1–4 yet their persistence in the environment has created a pressing sustainability challenge.5,6 Designing polymers whose end-of-life fate can be controlled without sacrificing mechanical robustness is therefore a key goal in polymer chemistry. Dynamic covalent chemistry offers one powerful strategy: incorporating exchangeable bonds into polymer backbones enables reprocessing, depolymerization, and chemical recycling under appropriate triggers, while preserving covalent connectivity.5,7–9 However, many dynamic covalent bonds (for example, imines and disulfides) are intrinsically weaker than the C–C, C–O, or C–N bonds they replace, so their introduction often leads to trade-offs in mechanical properties.5,10,11 Understanding and mitigating these trade-offs is essential for translating dynamic polymers into demanding applications.Polyurethanes (PUs) are an especially important platform for such efforts.12 They are ubiquitous in foams, elastomers, and high-performance coatings,13,14 yet most commercial PUs are based on permanent urethane linkages that are difficult to recycle. Recent work has shown that embedding dynamic covalent motifs into PU backbones or cross-links—such as disulfides, imines, or other exchangeable linkages—can yield materials that are reprocessable, degradable, or self-healing, while strong hydrogen-bond interactions in the hard segments can partially compensate for the mechanical weakness of dynamic bonds in the soft segments.15–17 In these systems, the overall thermomechanical response is governed not only by the dynamic bond chemistry but also by supramolecular interactions and the underlying segmental architecture.
Stereochemical control adds a further, powerful lever for tuning the properties and degradability of such polymers. Isohexide-based monomers derived from sugar alcohols (isosorbide, isomannide, and isoidide) provide rigid, chiral bicyclic units whose configuration strongly influences chain packing, glass-transition temperature, and susceptibility to hydrolytic or enzymatic degradation.18–20 Building such isohexide units into polyesters, polycarbonates, or polyurethanes has produced families of materials in which mechanical properties and degradation profiles can be modulated simply by switching stereochemistry at the monomer level.12,21,22
Dove and co-workers recently reported a series of non-segmented, alternating isohexide-based polyurethanes (IIPU and IMPU) that display stereochemistry-dependent mechanical behavior and degradability.12 They showed that exchanging isoidide for isomannide converts a tough, semicrystalline plastic into a strong amorphous thermoplastic elastomer, while keeping the overall composition and stoichiometry nearly identical. Detailed rheology, tensile testing, and accelerated degradation experiments demonstrated that stereochemistry controls supramolecular hydrogen-bond networks, crystallinity, and hydrolytic stability; copolymerization and blending of the stereoisomers allowed independent tuning of toughness and degradation rate. Molecular dynamics (MD) simulations supported the experimental observation that the isohexide configuration directs the balance of intra- and intermolecular hydrogen bonds and thereby the bulk mechanical response.
The current work extends this isohexide PU platform by introducing dynamic covalent bonds and hydrogen-bonding motifs to create recyclable PUs with tunable mechanics. Our goal was to obtain a mechanistic, multiscale picture that connects local bond scission, supramolecular cohesion, and bulk stress–strain behavior, and to clarify how the choice of dynamic-bond moiety in the backbone and the number and arrangement of hydrogen-bond donors and acceptors in the side chains interplay with the isohexide stereochemical environment. From a polymer-mechanics perspective, macroscopic failure under tensile loading reflects competition between covalent scission along load-bearing chains and interchain debonding/chain pullout governed by supramolecular cohesion. Accordingly, we used single-chain scission forces and double-chain shear forces as complementary molecular-scale descriptors to quantify these two limiting contributions and to rationalize bulk stress–strain behavior.
To this end, we have developed a multiscale computational framework that links single-chain mechanochemistry, interchain cohesion, and bulk mechanical response in isohexide-based PUs. We focused on three modular, synthetically accessible features: dynamic-bond moieties in the backbone, which set the intrinsic strength of the chains; hydrogen-bonding moieties, which provide tunable supramolecular interactions between chains; and isoidide or isomannide rings, which influence chain packing. Density functional theory (DFT) with the “External Force is Explicitly Included” (EFEI) formalism was used to quantify single-chain scission forces.23–27 Semiempirical EFEI simulations and classical MD then determined how the number and spatial arrangement of hydrogen-bond sites govern double-chain shear forces and hydrogen-bond statistics. Finally, reactive MD with ReaxFF probed uniaxial tensile deformation of amorphous bulk systems, thereby connecting molecular-scale design parameters to macroscopic stress–strain behavior.28–31
Computational details
Quantum-chemical calculations with EFEI
All quantum-chemical calculations were performed with ORCA 5.0.32 For single, finite PU chains (Fig. 2a), we used the BP86 exchange–correlation functional33 with the def2-TZVP basis set, augmented by D3(BJ) dispersion corrections to account for long-range interactions. For double-chain systems (Fig. 2c), which comprise more than 300 atoms, we employed the GFN2-xTB semiempirical quantum mechanical method, which provides an efficient description of geometry and dispersion interactions in large organic systems.34To model mechanical deformation, we used the EFEI formalism.24,25,35 In this approach, an external force is applied along a specified vector between two atoms, and geometry optimization is performed under this constraint. For both single-chain and double-chain systems, the force was applied between terminal carbon atoms located at the two ends of the chain.35,36 After each force increment, a constrained geometry optimization was carried out. For single-chain calculations, the external force was initially increased in steps of 0.5 nN, and then refined to 0.1 nN near the scission point. For double-chain calculations, a constant increment of 0.02 nN was used until shear occurred.
To determine the scission point in single chains (Fig. 2a and b), we analyzed the force–extension curve between the two terminal atoms. The scission point was defined as the last converged geometry before irreversible breaking of one or more covalent bonds, as evidenced by a discontinuous jump in extension and subsequent structural fragmentation.
For double chains (Fig. 2c and d), we defined the shear point based on changes in extension under successive force increments. Shear was identified when an increase in force produced an elongation more than ten times larger than in the previous step, and when this accelerated elongation persisted for at least three consecutive steps. This behavior signals the loss of non-bonded interactions that maintain the close-contact geometries of the two chains. The last point prior to this rapid extension was taken as the shear point.
Classical MD simulations of double chains
Classical MD simulations of the double PU chains studied by EFEI were conducted using the LAMMPS package.37,38 Atomistic force-field parameters were generated with the LigParGen web server, which provides OPLS-AA/CM1A-LBCC parameters suitable for organic systems.37–41Each system was placed in a 100 × 100 × 100 Å3 simulation box with periodic boundary conditions. An initial NVT simulation at 500 K was run for 8 ns with a time step of 0.001 ps, using a Langevin thermostat for temperature control. During the production phase, configurations were saved every 2 ps over the last 4 ns, yielding 2000 configurations per system. Hydrogen bonds were identified geometrically: a hydrogen bond was counted when the hydrogen–acceptor distance was <2.0 Å and the donor–hydrogen–acceptor angle exceeded 135°.42
Reactive MD simulations of PU bulk materials
Bulk mechanical properties were investigated using the ReaxFF reactive force field, as implemented in LAMMPS.37,38 We employed the dispersion/CHONSSi-lg ReaxFF parameter set, which has been validated for polymers containing C, H, O, N, and S atoms.43–45For each PU design, we constructed a simulation box consisting of 24 polymer chains, each with four repeating units (n = 4 in Fig. 1), amounting to a total of approximately 8500 atoms. All chains were initially aligned parallel to the x-axis in a periodic simulation box of ∼170 × 150 × 200 Å3, corresponding to an initial density of ∼0.02 g cm−3.
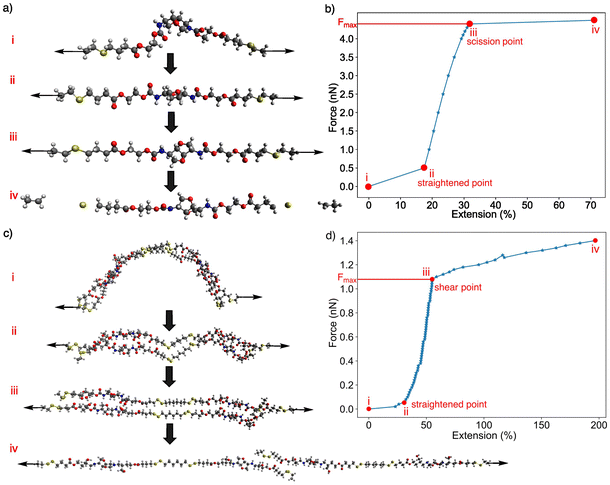 | ||
| Fig. 2 Quantum-chemical EFEI protocol for defining single-chain scission and double-chain shear points. (a) Representative geometries along tensile deformation of the single S2M chain: (i) relaxed, force-free structure; (ii) straightened chain under moderate load; (iii) configuration at the scission point, corresponding to the last intact structure before covalent bond scission; and (iv) fully scissile structure.36 (b) Calculated force–extension curve for S2M, with the four configurations in (a) indicated and the maximum tolerated force Fmax marking the scission point. (c) Representative geometries along tensile deformation of the SS5I double chains: (i) relaxed configuration; (ii) straightened pair of chains; (iii) configuration at the shear point, i.e., the last structure before loss of key non-bonded interactions between chains; and (iv) fully separated, sheared chains. (d) Corresponding force–extension curve for the SS5I double chains, showing the definition of Fmax used to quantify intermolecular ductility. | ||
Amorphous polymer structures were then prepared via the following protocol:
1. Energy minimization of the initial configuration using a conjugate-gradient algorithm.
2. Heating to 300 K at 1 atm using an NPT ensemble for 50 ps.
3. Sequential heating from 300 K to 800 K in 100 K increments, with 15 ps of NVT dynamics at each temperature.
4. Additional NPT equilibration at 800 K for 15 ps.
5. Cooling from 800 K to 300 K using a similar staged protocol.
This heating–cooling cycle was repeated until the density variation between consecutive cycles was less than 0.1% over five successive iterations. The final equilibrated densities were 1.25–1.35 g cm−3, consistent with dense amorphous polymers.
Uniaxial tensile deformation was then applied along each of the three Cartesian axes at 300 K and 1 atm. The deformation speed was set to 15 m s−1. Engineering stresses were computed via the Virial stress tensor.46 Engineering stress is the applied force divided by the original cross-sectional area of a material, which is a measure of internal resistance per unit area and is often called nominal stress. Simulations were continued until significant bond breaking occurred along the deformation direction or until delamination of chains was observed in the directions transverse to loading.
Results
Scission of single chains
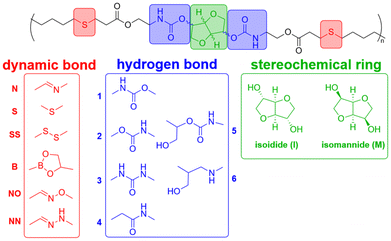
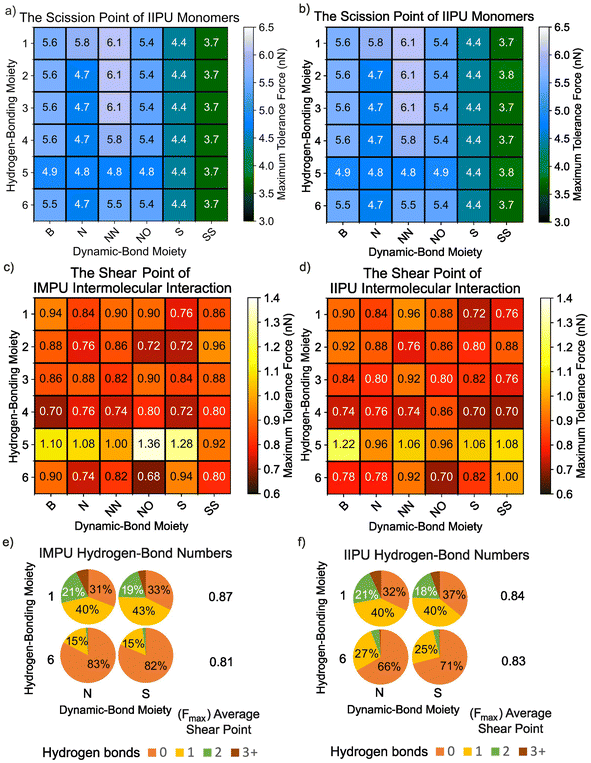
We first examined how dynamic-bond and hydrogen-bonding moieties influence the mechanical response of isolated PU chains. Six representative dynamic-bond motifs and six hydrogen-bonding motifs, selected from previously reported materials,7,47–54 were incorporated into isomannide-based (IMPU) and isoidide-based (IIPU) PU chains. The resulting maximum tolerated forces were obtained from DFT-EFEI calculations (Fig. 3a and b). IMPU and IIPU single chains displayed very similar scission forces, with differences of at most 0.1 nN for a given combination of dynamic-bond and hydrogen-bonding moieties. This suggests that the large experimental contrast in Young's modulus and tensile response between IMPU (elastomeric) and IIPU (tough plastic) reported previously does not originate from differences in covalent bond strength, but rather from supramolecular packing and morphology. Our results therefore support the experimental interpretation12 that isohexide stereochemistry primarily tunes network structure and hydrogen-bond organization, while the underlying backbone bonds remain comparably robust.Focusing on IMPU, we found that dynamic-bond moieties have a stronger effect on chain scission than hydrogen-bonding moieties. The scission force depends sensitively on which bond in the dynamic-bond motif breaks first (see Fig. S4). For most dynamic-bond moieties, the maximum tolerated force lies between 4.7 and 6.1 nN, indicating a relatively robust covalent network. In contrast, the original S-containing structure supports only 4.4 nN, and the disulfide-based dynamic-bond moiety (SS) exhibits even lower scission forces of 3.7–3.8 nN. These results indicate that sulfur-containing dynamic bonds substantially weaken the chains, as C–S and S–S bonds easily rupture under tensile loading. Hydrogen-bonding moieties exert a more subtle but non-negligible influence. Certain hydrogen-bonding motifs (notably moiety 5) break earlier, with maximum tolerated forces below 5.0 nN, due to cleavage of a C–O bond within the hydrogen-bonding group. This behavior can be attributed to hyperconjugation and electronic effects associated with the side chain. Among all tested combinations, IMPU and IIPU chains bearing the NN dynamic-bond moiety together with hydrogen-bonding moieties 1, 2, or 3 reach the highest scission forces of 6.1 nN. The NN dynamic-bond moiety here is so robust that the scission point is on the adjacent hydrogen-bonding moiety rather than moiety NN. These three robust IMPUs can compensate for the weakness of degradable linkages, while the choice of hydrogen-bonding group modulates but does not dominate the intra-chain-level mechanochemical response.
Intermolecular ductility of PU double chains
We next investigated how the same chemical modifications affect intermolecular cohesion between PU chains. Using semiempirical EFEI calculations, we computed the shear forces required to disrupt non-bonded interactions between chains for all combinations of dynamic-bond and hydrogen-bonding moieties (Fig. 3c and d).In contrast to the single-chain results, the strength of non-bonded interactions is influenced primarily by the hydrogen-bonding moiety rather than the dynamic-bond motif. Among all hydrogen-bonding moieties, moiety 5—which can form up to four hydrogen bonds—exhibits the highest shear forces, typically exceeding 1.0 nN. Moiety 4, which has only two hydrogen-bond sites, shows significantly weaker non-bonded interactions, generally below 0.8 nN. Moieties 1, 2, and 3, each offering three hydrogen-bond sites, yield intermediate shear forces in the 0.8–0.9 nN range. Moiety 6 also contains two hydrogen-bond sites but exhibits somewhat higher and more variable shear forces than moiety 4. This behavior can be attributed to outer hydroxyl groups, which can form relatively strong but geometrically sensitive hydrogen bonds.
Overall, the dynamic-bond moiety has only a minor influence on interchain shear force, and no systematic differences are observed between IMPU and IIPU, despite variations in stereochemical ring configurations. Within the IMPU set, the optimal PU design combines the NO dynamic-bond motif with hydrogen-bonding moiety 5, tolerating shear forces up to 1.36 nN. For IIPU, the best performer uses the B dynamic-bond motif with moiety 5, withstanding up to 1.22 nN. These results reinforce that hydrogen bond topology—number and spatial arrangement of donors and acceptors—is the primary determinant of intermolecular ductility in the double-chain system.
To capture the structural diversity accessible at finite temperature, we performed MD simulations for selected combinations and computed ensemble-averaged hydrogen-bond statistics (Fig. 3e and f). For hydrogen-bonding moiety 1, other structural variables (dynamic-bond type and stereochemical ring) exert little influence: both IMPU and IIPU chain pairs predominantly form a single interchain hydrogen bond (∼40%), followed by no hydrogen bond (30–37%), while configurations with two hydrogen bonds account for ∼20% and those with three or more are rare. Consistent with the similar hydrogen-bond populations, the mean shear forces are comparable (0.87 nN for IMPU and 0.84 nN for IIPU).
In contrast, for hydrogen-bonding moiety 6, hydrogen bonding is strongly suppressed. In IMPU chain pairs, >80% of configurations contain no interchain hydrogen bond and only ∼15% contain one, consistent with the reduced number of donor/acceptor sites (two vs. three in moiety 1) and the less favourable orientation of the terminal hydroxyl groups. This reduction in effective interchain cohesion is reflected in the lower mean shear force (0.81 nN). In IIPU chain pairs, the fraction of configurations without hydrogen bonds decreases to >60% and those with one hydrogen bond increases to ∼25%, indicating a modest stereochemical effect. Moreover, N6 forms a higher fraction of hydrogen-bonded configurations than S6, consistent with the additional acceptor capability of the nitrogen-containing dynamic-bond motif. Importantly, analysis of hydrogen-bond geometries (H⋯A distance and donor–H–acceptor angle distributions) shows that moiety 6 not only forms fewer hydrogen bonds but also samples more bent and longer hydrogen bonds on average, providing a mechanistic basis for its weaker resistance to shearing.
Taken together, these results show that hydrogen-bonding moiety is the primary control for inter-chain-level cohesion, with the number and geometry of hydrogen-bond sites setting the overall hydrogen-bond population. The strong correlation we observe between shear force and the number of accessible hydrogen-bond sites mirrors the conclusions of Stubbs et al.,12 who used MD simulations and tensile experiments to show that the isoidide and isomannide stereochemistries direct distinct intra- and intermolecular hydrogen-bonding patterns and thereby govern crystallinity and mechanical behavior. In our case, multidentate hydrogen-bonding motifs mimic the role of the IIPU “hardening” motif: they increase the population of interchain hydrogen bonds and yield higher shear force, whereas motifs with fewer or poorly oriented donors/acceptors resemble the IMPU-like situation with weaker interchain cohesion. Comparatively, dynamic-bond motifs and stereochemistry play secondary roles that become noticeable mainly when the hydrogen-bond network is intrinsically weak.
From chains to bulk mechanical response
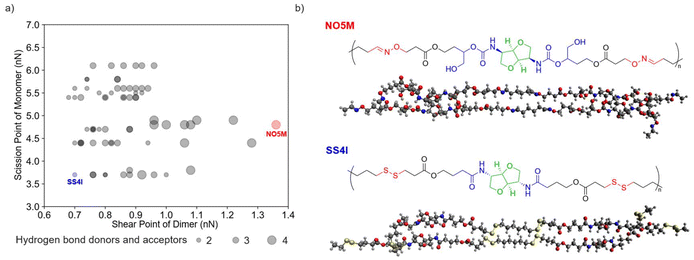
Connecting the single-chain and double-chain insights to bulk mechanical performance (Fig. 4), we summarize the calculated single-chain scission forces and double-chain shear forces across all 72 designed PU structures. In general, polymers with fewer hydrogen-bond donors and acceptors and weak dynamic bonds show poor performance at one or both scales, whereas structures with more hydrogen-bond sites and stronger dynamic bonds perform better in both single-chain scission and double chain shear force.Based on these trends, two representative PU polymers are selected for reactive MD tensile tests of amorphous bulk materials: (1) NO5M—an IMPU-based PU with the NO dynamic-bond moiety and hydrogen-bonding moiety 5, which exhibits strong performance at both the single-chain and double-chain levels; (2) SS4I—an IIPU-based PU with the disulfide (SS) dynamic-bond moiety and hydrogen-bonding moiety 4, which shows among the lowest scission and shear force.
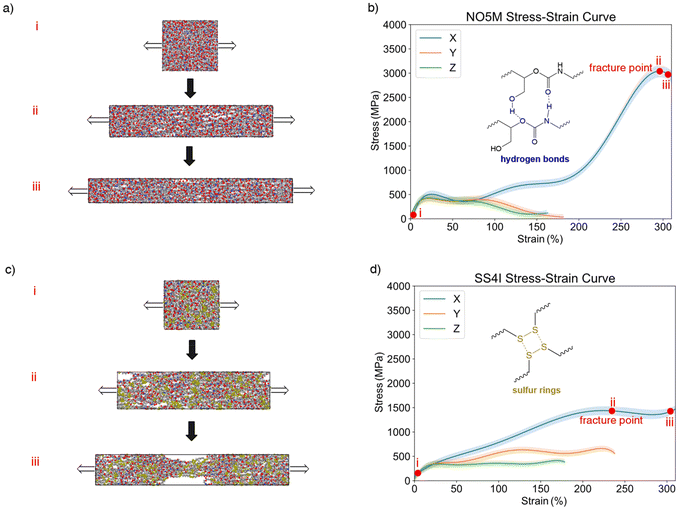
Fig. 5 presents the stress–strain curves obtained from ReaxFF tensile simulations. The maximum stress (peak of the curve) is taken as the apparent scission stress, while the subsequent decrease reflects continued bond breaking and delamination, indicative of material failure. When the external load is applied along the x-axis, parallel to the initial chain orientation, both covalent and non-bonded interactions contribute to the response, providing a reasonable representation of load transfer in aligned polymer domains. Given that MD simulations utilize a relatively small simulation box, we can efficiently determine the relative differences in mechanical responses of two materials.55,56
For NO5M, the stress–strain curve along x initially rises steeply. At stresses above ∼500 MPa, small cavities start to appear, leading to a plateau and fluctuations. As deformation proceeds and most non-bonded interactions are overcome, the covalent backbone bears the majority of the load, and the stress rapidly increases to a peak of ∼3500 MPa. Beyond this point, extensive bond scission triggers a continuous decline in stress, marking material failure.
For SS4I, the stress–strain curve increases more gradually but continues to rise as the simulation cell is elongated to roughly three times its original length. The maximum stress, however, is only ∼1500 MPa, less than half that of NO5M, underscoring the detrimental impact of weak S–S and C–S bonds on bulk strength. Notably, the stress does not drop sharply after the peak. Upon visual inspection of the relevant MD snapshots, it is found that dynamic exchange of disulfide bonds allows partial network reconfiguration (Fig. 5d, inset), which sustains load over a broader strain range despite substantial bond breakage.57,58
When deformation is applied along the y- and z-axes, the chains are predominantly loaded through non-bonded interactions, resulting in markedly lower stress levels and earlier onset of delamination for both systems. These directions thus report on the interchain cohesion established by the hydrogen-bond network and other non-covalent interactions.
Although our simulations are performed at much higher strain rates than experiment, the qualitative features of the NO5M and SS4I stress–strain curves resemble the behavior of isohexide PUs reported previously: a pronounced strain-hardening regime and high peak stress for materials with strong, well-organized hydrogen-bond networks, and a softer, more gradual response when supramolecular cohesion is weaker.12,59 In this sense, NO5M plays a role analogous to the semicrystalline, strongly H-bonded IIPU, whereas SS4I behaves more like a softer, strongly dynamic system in which weak S–S linkages cap the attainable strength.
Overall, the bulk simulations confirm the design rules inferred at smaller scales. First, dynamic-bond moieties control the intrinsic strength of the backbone and thus strongly influence single-chain scission forces and the maximum attainable bulk stress. Sulfur-based motifs (S, SS) substantially weaken the material, while nitrogen-containing motifs (NN, NO, B) are more robust. The reduction in simulated fracture stress for the disulfide-rich SS4I analogue is also consistent with numerous experimental studies on disulfide-containing PUs, where the introduction of S–S bonds improves self-healing or reprocessability but generally lowers modulus and strength compared to non-dynamic analogues.60 By contrast, the NO- and NN-based dynamic moieties provide higher single-chain scission forces and bulk stresses, in line with reports that imine-containing PUs can combine excellent mechanical properties with dynamic exchange and recyclability when placed in appropriately reinforced domains.61 Second, hydrogen-bonding moieties dictate the density and strength of interchain interactions, modulating inter-chain shear forces and affecting the onset of cavitation and delamination in bulk. Multidentate motifs with four hydrogen-bond sites (moiety 5) yield the strongest cohesion. Last, the NO5M system combines a stronger dynamic-bond motif with a highly multidentate hydrogen-bond network, leading to superior performance at all scales compared to the disulfide-rich, weakly hydrogen-bonded SS4I analogue.
Conclusion
We have developed a comprehensive multiscale methodology for probing and designing the mechanical properties of dynamic, recyclable polymers, illustrated here for isohexide-based polyurethanes. By integrating DFT-based and semiempirical EFEI calculations with classical and reactive MD simulations, we systematically connected single-chain-level mechanochemistry, in which dynamic-bond moieties control single-chain scission forces, to double-chain-level intermolecular cohesion, where hydrogen-bonding moieties determine non-bonded interaction strengths and hydrogen-bond statistics, and finally to bulk-scale mechanical response, where both factors combine to set the stress–strain behavior under uniaxial tensile loading.Our results show that dynamic-bond moieties dominate single-chain-level strength: sulfur-containing motifs (S, SS) significantly reduce scission forces, whereas nitrogen-containing motifs such as NN, NO, and B provide markedly stronger backbones. Hydrogen-bonding moieties emerge as the primary determinants of interchain cohesion, with the number and spatial arrangement of hydrogen-bond donors and acceptors controlling double-chain shear forces and hydrogen-bond populations. These two design dimensions remain important in the bulk, where polymers that combine robust dynamic bonds with multidentate hydrogen-bonding motifs (for example, NO5M) display significantly higher scission stresses than disulfide-rich, weakly hydrogen-bonded analogues such as SS4I. In combination with the experimental isohexide PU family,12,59 our results suggest a route to experimentally implement the NO- and NN-type dynamic motifs and multidentate hydrogen-bonding groups identified here, providing a clear set of molecular targets for future synthesis and mechanical testing.
Beyond the specific PU chemistries studied here, our framework provides a general strategy for relating chemical design to macroscopic mechanical performance in dynamic and degradable polymers. The workflow is compatible with high-throughput screening and could readily be coupled to machine learning or optimization algorithms to explore broader chemical spaces. Future work will focus on integrating such data-driven approaches, extending the methodology to more complex architectures (e.g., cross-linked networks and block copolymers), and refining the connection between simulation and experiment for predictive polymer design.
Author contributions
L. C. and A. P. D. conceived the project; L. C. supervised the research. C. S. developed and implemented all code and conducted the computational studies, with M. H.'s guidance and supervision. C. S. and L. C. led the preparation of the manuscript, with input from all authors. We thank the Hefei Advanced Computing Center for providing computational resources for this project. M. H. thanks the support from the National Natural Science Foundation of China 22503097.Conflicts of interest
The authors declare no competing interests.Data availability
The data supporting the findings of this study are available within the paper and its supplementary information (SI). The additional information of SI includes the description of computational methods, benchmark study of intramolecular and intermolecular ductility, fragments after the scission of single chains of benchmark molecules and PU, distributions of hydrogen-bond numbers across hydrogen-bonding moiety, statistics of hydrogen number of different smapshotted IMPU/IIPU polymer structures and statistics of hydrogen bonds in IMPU polymer structures. See DOI: https://doi.org/10.1039/d5py01221j.All code developed and used in this study is available at https://github.com/pic-ai-robotic-chemistry/poly-mech-props.
Acknowledgements
The AI-driven experiments, simulations and model training were performed on the robotic AI-Scientist platform of Chinese Academy of Sciences (CAS). L. C. acknowledges the University of Science and Technology of China (USTC) Startup Program for funding.References
- A. Kosmalska-Olańska, J. Olszewski and A. Masek, A brief review of optical polymers in material engineering, eXPRESS Polym. Lett., 2024, 18(12), 1291–1326 CrossRef.
- T. Gurunathan, S. Mohanty and S. K. Nayak, Hyperbranched Polymers for Coating Applications: A Review, Polym.-Plast. Technol. Eng., 2015, 55(1), 92–117 CrossRef.
- A. Ershad-Langroudi, N. Babazadeh, F. Alizadegan, S. M. Mousaei and G. Moradi, Polymers for implantable devices, J. Ind. Eng. Chem., 2024, 137, 61–86 CrossRef CAS.
- W. Wu, W. Wang and J. Li, Star polymers: Advances in biomedical applications, Prog. Polym. Sci., 2015, 46, 55–85 CrossRef CAS.
- G. X. Wang, D. Huang, J. H. Ji, C. Volker and F. R. Wurm, Seawater-Degradable Polymers-Fighting the Marine Plastic Pollution, Adv. Sci., 2020, 8(1), 2001121 CrossRef.
- T. Bond, V. Ferrandiz-Mas, M. Felipe-Sotelo and E. van Sebille, The occurrence and degradation of aquatic plastic litter based on polymer physicochemical properties: A review, Crit. Rev. Environ. Sci. Technol., 2018, 48(7–9), 685–722 CrossRef.
- L. Li, X. Peng, D. Zhu, J. Zhang and P. Xiao, Recent Progress in Polymers with Dynamic Covalent Bonds, Macromol. Chem. Phys., 2023, 224(20), 2300224 CrossRef CAS.
- R. Kaur, P. Singh, S. Tanwar, G. Varshney and S. Yadav, Assessment of Bio-Based Polyurethanes: Perspective on Applications and Bio-Degradation, Macromolecular, 2022, 2(3), 284–314 CrossRef CAS.
- R. Hatti-Kaul, L. J. Nilsson, B. Zhang, N. Rehnberg and S. Lundmark, Designing Biobased Recyclable Polymers for Plastics, Trends Biotechnol., 2020, 38(1), 50–67 CrossRef CAS.
- A. Sugimura, M. Asai, T. Matsunaga, Y. Akagi, T. Sakai, H. Noguchi and M. Shibayama, Mechanical properties of a polymer network of Tetra-PEG gel, Polym. J., 2012, 45(3), 300–306 CrossRef.
- S. Ma and D. C. Webster, Degradable thermosets based on labile bonds or linkages: A review, Prog. Polym. Sci., 2018, 76, 65–110 CrossRef CAS.
- C. J. Stubbs, J. C. Worch, H. Prydderch, Z. Wang, R. T. Mathers, A. V. Dobrynin, M. L. Becker and A. P. Dove, Sugar-Based Polymers with Stereochemistry-Dependent Degradability and Mechanical Properties, J. Am. Chem. Soc., 2022, 144(3), 1243–1250 CrossRef CAS PubMed.
- A. Kemona and M. Piotrowska, Polyurethane Recycling and Disposal: Methods and Prospects, Polymers, 2020, 12(8), 1752 CrossRef CAS.
- A. Delavarde, G. Savin, P. Derkenne, M. Boursier, R. Morales-Cerrada, B. Nottelet, J. Pinaud and S. Caillol, Sustainable polyurethanes: toward new cutting-edge opportunities, Prog. Polym. Sci., 2024, 151, 101805 CrossRef CAS.
- X. Zhu, W. Zhang, G. Lu, H. Zhao and L. Wang, Ultrahigh Mechanical Strength and Robust Room-Temperature Self-Healing Properties of a Polyurethane-Graphene Oxide Network Resulting from Multiple Dynamic Bonds, ACS Nano, 2022, 16(10), 16724–16735 CrossRef CAS.
- J. C. Worch and A. P. Dove, 100th Anniversary of Macromolecular Science Viewpoint: Toward Catalytic Chemical Recycling of Waste (and Future) Plastics, ACS Macro Lett., 2020, 9(11), 1494–1506 CrossRef CAS.
- Y. H. Hsu, D. Luong, D. Asheghali, A. P. Dove and M. L. Becker, Shape Memory Behavior of Biocompatible Polyurethane Stereoelastomers Synthesized via Thiol-Yne Michael Addition, Biomacromolecules, 2022, 23(3), 1205–1213 CrossRef CAS PubMed.
- B. Koo and C. Kim, Synthesis of Stereocontrolled Degradable Polymer by Living Cascade Enyne Metathesis Polymerization, Angew. Chem., Int. Ed., 2023, 62(46), e202312399 CrossRef CAS.
- J. R. Jagannathan and F. A. Leibfarth, Stereoconvergent Chain-Growth Polymerization, ACS Cent. Sci., 2025, 11, 797–804 CrossRef CAS PubMed.
- J. C. Worch, H. Prydderch, S. Jimaja, P. Bexis, M. L. Becker and A. P. Dove, Stereochemical enhancement of polymer properties, Nat. Rev. Chem., 2019, 3(9), 514–535 CrossRef CAS.
- W. Vogelzang, R. J. I. Knoop, D. S. van Es, R. Blaauw and E. Maaskant, Biobased high-performance polyesters: Synthesis and thermal properties of poly(isoidide furanoate) and co-polyesters, Eur. Polym. J., 2023, 200, 112516 CrossRef CAS.
- A. Saadaoui, R. Medimagh, S. Marque, D. Prim, S. Chatti, H. Casabianca and M. S. Zina, New biosourced AA and, AB monomers from 1,4:3,6-dianhydrohexitols, Isosorbide, Isomannide, and Isoidide, Des. Monomers Polym., 2017, 20(1), 221–233 CrossRef CAS.
- M. K. Beyer, The mechanical strength of a covalent bond calculated by density functional theory, J. Chem. Phys., 2000, 112(17), 7307–7312 CrossRef CAS.
- J. Ribas-Arino, M. Shiga and D. Marx, Understanding covalent mechanochemistry, Angew. Chem., Int. Ed., 2009, 48(23), 4190–4193 CrossRef CAS PubMed.
- I. M. Klein, C. C. Husic, D. P. Kovacs, N. J. Choquette and M. J. Robb, Validation of the CoGEF Method as a Predictive Tool for Polymer Mechanochemistry, J. Am. Chem. Soc., 2020, 142(38), 16364–16381 CrossRef CAS.
- Q. Mu and J. Hu, Polymer mechanochemistry: from single molecule to bulk material, Phys. Chem. Chem. Phys., 2024, 26(2), 679–694 RSC.
- C. R. Wick, E. Topraksal, D. M. Smith and A.-S. Smith, Evaluating the predictive character of the method of constrained geometries simulate external force with density functional theory, Forces Mech., 2022, 9, 100143 CrossRef.
- C. E. Petilla, C. J. Dela Cruz and C. L. Mahinay, Mechanical properties of Si(1–x)–C(x): strength and stiffness of materials using LAMMPS molecular dynamics simulation, Jpn. J. Appl. Phys., 2024, 63(8), 08SP09 CrossRef CAS.
- S. Taheri, Molecular dynamics simulation of mechanical properties of hafnium nanowires: the effects of size, temperature and strain rate, Phys. B: Condens. Matter, 2025, 711, 417290 CrossRef CAS.
- C. W. Jang, J. W. Mullinax and J. W. Lawson, Mechanical Properties and Failure of Aerospace-Grade Epoxy Resins from Reactive Molecular Dynamics Simulations with Nanoscale Defects, ACS Appl. Polym. Mater., 2022, 4(8), 5269–5274 CrossRef CAS.
- G. Aral, Oxide shell layer influences on size-dependent tensile and compressive mechanical properties of iron nanowires: A ReaxFF molecular dynamics study, J. Appl. Phys., 2019, 126(13), 135109 CrossRef.
- F. Neese, Software update: The ORCA program system—Version 5.0, WIREs Computational Molecular Science, 2022, 12 DOI:10.1002/wcms.1606.
- A. D. Becke, Density-functional exchange-energy approximation with correct asymptotic behavior, Phys. Rev. A: At., Mol., Opt. Phys., 1988, 38(6), 3098–3100 CrossRef CAS.
- C. Bannwarth, S. Ehlert and S. Grimme, GFN2-xTB-An Accurate and Broadly Parametrized Self-Consistent Tight-Binding Quantum Chemical Method with Multipole Electrostatics and Density-Dependent Dispersion Contributions, J. Chem. Theory Comput., 2019, 15(3), 1652–1671 CrossRef CAS PubMed.
- M. A. Cardosa-Gutierrez, G. D. Bo, A.-S. Duwez and F. Remacle, Steering the Mechanism of the Furan-Maleimide Retro-Diels-Alder Reaction to a Sequential Pathway with an External Mechanical Force, ChemRxiv, 2022, 2022(0304) DOI:10.26434/chemrxiv-2022-g64jp.
- J. Ribas-Arino and D. Marx, Covalent mechanochemistry: theoretical concepts and computational tools with applications to molecular nanomechanics, Chem. Rev., 2012, 112(10), 5412–5487 CrossRef CAS.
- D. M. Heyes, Pressure tensor of partial-charge and point-dipole lattices with bulk and surface geometries, Phys. Rev. B: Condens. Matter Mater. Phys., 1994, 49(2), 755–764 CrossRef CAS PubMed.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. in ’t Veld, A. Kohlmeyer, S. G. Moore, T. D. Nguyen, R. Shan, M. J. Stevens, J. Tranchida, C. Trott and S. J. Plimpton, LAMMPS - a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales, Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- L. S. Dodda, J. Z. Vilseck, J. Tirado-Rives and W. L. Jorgensen, 1.14*CM1A-LBCC: Localized Bond-Charge Corrected CM1A Charges for Condensed-Phase Simulations, J. Phys. Chem. B, 2017, 121(15), 3864–3870 CrossRef CAS.
- L. S. Dodda, C. d. Vaca, I. Tirado-Rives, J. Jorgensen and W. L, LigParGen web server: an automatic OPLS-AA parameter generator for organic ligands, Nucleic Acids Res., 2017, 45(W1), W331–W336 CrossRef CAS.
- W. L. Jorgensen, Tirado-Rives. Potential energy functions for atomic-level simulations of water and organic and biomolecular systems, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 6665–6670 CrossRef CAS.
- E. Arunan, G. R. Desiraju, R. A. Klein, J. Sadlej, S. Scheiner, I. Alkorta, D. C. Clary, R. H. Crabtree, J. J. Dannenberg and P. Hobza, et al., Definition of the hydrogen bond (IUPAC Recommendations 2011), Pure Appl. Chem., 2011, 83(8), 1637–1641 CrossRef CAS.
- M. Kowalik, C. Ashraf, B. Damirchi, D. Akbarian, S. Rajabpour and A. C. T. van Duin, Atomistic Scale Analysis of the Carbonization Process for C/H/O/N-Based Polymers with the ReaxFF Reactive Force Field, J. Phys. Chem. B, 2019, 123(25), 5357–5367 CrossRef CAS PubMed.
- Q. Mao, S. Rajabpour, M. Kowalik and A. C. T. van Duin, Predicting cost-effective carbon fiber precursors: Unraveling the functionalities of oxygen and nitrogen-containing groups during carbonization from ReaxFF simulations, Carbon, 2020, 159, 25–36 CrossRef CAS.
- A. K. Rappé, C. J. Casewit, K. S. Colwell, W. A. Goddard III and W. M. Skiff, UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations, J. Am. Chem. Soc., 1992, 114(25), 10024–10035 CrossRef.
- M. P. Allen and D. J. Tildesley, Computer simulation of liquids, Oxford university press, 2017 Search PubMed.
- R. Zhang, W. Huang, P. Lyu, S. Yan, X. Wang and J. Ju, Polyurea for Blast and Impact Protection: A Review, Polymers, 2022, 14(13), 2670 CrossRef CAS.
- J. A. Chiong, Y. Zheng, S. Zhang, G. Ma, Y. Wu, G. Ngaruka, Y. Lin, X. Gu and Z. Bao, Impact of Molecular Design on Degradation Lifetimes of Degradable Imine-Based Semiconducting Polymers, J. Am. Chem. Soc., 2022, 144(8), 3717–3726 CrossRef CAS PubMed.
- Y. Yin, Y. Xu, X. Zhang, B. Duan, Z. Xin and C. Bao, Mechanically Strong and Tough Poly(urea-urethane) Thermosets Capable of Being Degraded under Mild Condition, Macromol. Rapid Commun., 2023, 44(5), e2200765 CrossRef PubMed.
- M. B. Sims, K. Y. Patel, M. Bhatta, S. Mukherjee and B. S. Sumerlin, Harnessing Imine Diversity To Tune Hyperbranched Polymer Degradation, Macromolecules, 2018, 51(2), 356–363 CrossRef CAS.
- W. Cai, M. Hassani, R. Karki, E. D. Walter, K. H. Koelsch, H. Seradj, J. P. Lineswala, H. Mirzaei, J. S. York and F. Olang, et al., Synthesis, metabolism and in vitro cytotoxicity studies on novel lavendamycin antitumor agents, Bioorg. Med. Chem., 2010, 18(5), 1899–1909 CrossRef CAS PubMed.
- W. Fenzl, Boron compounds, XLVIII. O-Ethylboranediylation with ethylbis(2-isopropyl-6-methyl-4-pyrimidinyloxy)borane, Justus Liebigs Ann. Chem., 1978, 6, 1030–1036 CrossRef.
- J. Jia, J. Liu, Z. Q. Wang, T. Liu, P. Yan, X. Q. Gong, C. Zhao, L. Chen, C. Miao and W. Zhao, et al., Photoinduced inverse vulcanization, Nat. Chem., 2022, 14(11), 1249–1257 CrossRef CAS.
- F. Garcia and M. M. Smulders, Dynamic covalent polymers, J. Polym. Sci., Part A: Polym. Chem., 2016, 54(22), 3551–3577 CrossRef CAS.
- M. M. Afroz, Y. K. Shin, A. C. T. van Duin and K. D. Li-Oakey, ReaxFF reactive force field model enables accurate prediction of physiochemical and mechanical properties of crystalline and amorphous shape–memory polyurethane, J. Appl. Polym. Sci., 2023, 140(39), e54466 CrossRef CAS.
- G. Zhang, W. Zhang, Y. Li, H. Chen, J. Ge, Y. Pan and D. Hou, The effect of curing regimes on the hydration products of lightweight ultra-high performance concrete: From experiments to MD simulationsz, Constr. Build. Mater., 2024, 412, 134865 CrossRef.
- J. Canadell, H. Goossens and B. Klumperman, Self-Healing Materials Based on Disulfide Links, Macromolecules, 2011, 44(8), 2536–2541 CrossRef CAS.
- D. Giannantonio, A. Brandolese and A. P. Dove, Tuneable and degradable thermosets possessing dynamic aliphatic disulfide bonds via stereoselective thiol–yne polymerisation, Polym. Chem., 2025, 16(6), 734–741 RSC.
- S. R. Petersen, H. Prydderch, J. C. Worch, C. J. Stubbs, Z. Wang, J. Yu, M. C. Arno, A. V. Dobrynin, M. L. Becker and A. P. Dove, Ultra-Tough Elastomers from Stereochemistry-Directed Hydrogen Bonding in Isosorbide-Based Polymers, Angew. Chem., Int. Ed., 2022, 61(17), e202115904 CrossRef CAS PubMed.
- X. Li, R. Yu, Y. He, Y. Zhang, X. Yang, X. Zhao and W. Huang, Self-Healing Polyurethane Elastomers Based on a Disulfide Bond by Digital Light Processing 3D Printing, ACS Macro Lett., 2019, 8(11), 1511–1516 CrossRef CAS.
- X. Xu, X. Ma, M. Cui, H. Zhao, N. E. Stott, J. Zhu, N. Yan and J. Chen, Fully biomass-derived polyurethane based on dynamic imine with self-healing, rapid degradability, and editable shape memory capabilities, Chem. Eng. J., 2024, 479, 147823 CrossRef CAS.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |